Abstract
Colorectal cancer responds poorly to immune checkpoint blockade in most patients with microsatellite-stable (MSS) tumors, highlighting the need for alternative targets. B7H3 (CD276) is an immune checkpoint protein that is frequently overexpressed in tumors, but how it is maintained at high protein levels is unclear. Here we show that palmitic acid (PA) promotes B7H3 palmitoylation by the palmitoyltransferase ZDHHC24 at cysteine 496. This modification prevents B7H3 from binding to sequestosome 1 (SQSTM1, also called p62), limiting autophagic degradation and stabilizing B7H3, which suppresses CD8+ T cell antitumor activity. Disrupting this pathway by mutating cysteine 496, or by deleting Zdhhc24 in a colitis-associated colorectal cancer (CAC) mouse model, enhances CD8+ T cell responses. We also develop a cell-penetrant peptide that blocks the ZDHHC24–B7H3 interaction, boosts antitumor immunity, and synergizes with blockade of programmed cell death protein 1 (PD-1). These findings identify B7H3 palmitoylation as a targetable metabolic–immune node for colorectal cancer immunotherapy.
Similar content being viewed by others
Introduction
Colorectal cancer is the third most common cancer worldwide1. Traditionally, surgery has been the primary treatment, but recent advances suggest that immunotherapy may offer durable and potentially curative outcomes2. Immune checkpoints, such as programmed death-1 (PD-1), programmed death-ligand 1 (PD-L1), and cytotoxic T-lymphocyte–associated antigen 4 (CTLA-4), play crucial roles in maintaining self-tolerance and regulating antitumor immunity3,4,5. These checkpoint pathways involve receptor–ligand interactions that deliver co-inhibitory signals to T cells, thereby suppressing their activation and effector function. Immune check point inhibitors (ICIs) targeting PD-1/PD-L1 are effective primarily in microsatellite instability-high (MSI-H) colorectal cancers, which constitute only about 15% of cases6,7,8,9. In contrast, these treatments show limited success in the majority of colorectal cancers that are microsatellite-stable (MSS)10,11. Therefore, it is imperative to conduct an intensive search for alternative immune checkpoints to apply in the treatment of MSS tumors.
The B7 family, comprising a group of structurally related transmembrane proteins, including B7-1, B7-2, B7-H1 (also known as PD-L1), B7-H3, and others, interacts with CD28 family receptors to either stimulate or suppress immune response12,13,14. B7H3 (CD276) has recently emerged as a compelling immunotherapy target due to its role in suppressing cytotoxic T cell and natural killer cell activity15,16,17,18. B7H3 is highly expressed in various malignancies, and its elevated expression is associated with poor prognosis and adverse clinical outcomes19,20,21,22. Although B7H3 mRNA is widely detectable across various tissues and cell types, protein expression is comparatively limited, suggesting that posttranscriptional mechanisms tightly regulate its abundance23. Posttranslational modifications (PTMs) of B7H3 remain underexplored, yet they are likely to play important roles in modulating its function in cancer. While phosphorylation and ubiquitination sites have been predicted, N-glycosylation plays a pivotal role in stabilizing B7H3 and ensuring its stable expression on the plasma membrane, thereby facilitating its immunosuppressive function24,25,26. However, it remains mostly unknown whether other post-translational modifications apart from glycosylation contribute to regulating the expression or function of B7H3. Elucidating these unexplored modifications and their regulatory mechanisms will be critical for developing more effective B7H3-targeted therapies.
S-acylation (often referred to as S-palmitoylation or simply palmitoylation) is an important post-translational modification in which a thioester bond is formed between the sulfhydryl group of a cysteine residue and, most commonly, a 16-carbon palmitate27,28,29,30. This PTM leads to an increase in the lipophilicity of the modified target protein, which can affect its stability, function, or subcellular trafficking31,32,33,34,35. Palmitoylation is mainly catalyzed by a relatively conserved family of zinc-finger DHHC (Asp-His-His-Cys) type-containing (ZDHHC) protein acyl-transferases (PATs)36,37. Accumulating evidence suggests that immune-related molecules such as PD-L138 and NOD1/239 can be palmitoylated, indicating that palmitoylation plays an important role in the immune response.
Protein palmitoylation is regulated not only by ZDHHC family enzymes but also by the availability of its active substrate, palmitoyl-coenzyme A. Palmitoyl-coenzyme A, the active metabolite of palmitic acid (PA), reportedly initiates palmitoylation modifications in a variety of proteins40,41,42. PA is a naturally occurring saturated fatty acid that has been reported to play important roles in immune evasion, metabolism, and inflammation43,44,45. However, whether PA contributes to the establishment of an immunosuppressive microenvironment in MSS colorectal cancer remains unclear.
In this work, we identify that the palmitoylation of B7H3 triggered by PA is a key driving factor in the formation of the immune-suppressive microenvironment in colorectal cancer. Mechanistically, we show that ZDHHC24 catalyzes B7H3 palmitoylation at cysteine 496 (C496), preventing its interaction with the autophagy receptor SQSTM1/p62 and blocking its autophagic degradation. Biologically, we demonstrate that palmitoylation of B7H3 C496 remodels the immune microenvironment and specifically promotes tumor immune escape by inhibiting the antitumor effects of CD8+ T cells. Based on these findings, we design a peptide (E-peptide) that binds competitively to ZDHHC24. Both in vivo and in vitro experiments demonstrate that E-peptide reduces the palmitoylation of B7H3, promotes its autophagic degradation, and ultimately enhances the anti-tumor activity of CD8+ T cells. Notably, combining E-peptide with anti-PD-1 therapy produces a synergistic effect, significantly improving therapeutic outcomes. These findings provide important insights into immunotherapy for colorectal cancer and identify B7H3 palmitoylation as a promising therapeutic target.
Results
PA-triggered B7H3 palmitoylation enhances B7H3 protein stability
To identify the key immune checkpoints regulating the tumor microenvironment in MSS colorectal cancer, we performed an immunohistochemical analysis of B7 family members using formalin-fixed paraffin-embedded (FFPE) tumor samples from both MSS and MSI-H colorectal cancer patients (cohort 1, Table S1). The results indicated that the B7H3 protein was markedly overexpressed in MSS tumors (Figs. 1A, B and S1A). As an immune checkpoint molecule, B7H3 is expressed not only on tumor cells but also on various immune cell populations. Immunofluorescence staining showed strong co-localization of B7H3 with the tumor marker epithelial cell adhesion molecule (EPCAM) in tumor tissues (Fig. S1B), indicating prominent expression in tumor cells. Flow cytometry further confirmed that more than 95% of tumor cells are B7H3–positive, compared to about 30% of immune cells (Fig. S1C, D) (cohort 2, Table S2). When normalized to 100,000 total cells, B7H3⁺ tumor cells accounted for approximately 90,000 (Fig. S1E), highlighting tumor cells as the predominant source of B7H3 in MSS tumors. Interestingly, subsequent qPCR analysis revealed no significant alterations in B7H3 mRNA levels in MSS samples compared to MSI-H samples (Fig. S1F) (cohort 3, Table S3), suggesting that PTMs likely play a critical role in regulating B7H3 expression specifically in MSS tumors. Since it has been reported that the FUT8-catalyzed glycosylation of B7H3 can stabilize B7H3 and ensure its stable expression on the plasma membrane25, we next assessed FUT8 expression in MSS and MSI-H colorectal tumor samples via immunohistochemistry. However, no significant difference in FUT8 expression was observed between these groups (Fig. S1G, H). To better understand the underlying mechanisms, we performed RNA sequencing using fresh colon tumor samples (cohort 4, Table S4). Gene Ontology (GO) enrichment analysis revealed that several genes regulating metabolites, such as carboxylic acids, fatty acids, and vitamins, were aberrantly expressed in MSS tumors (Fig. 1C and Supplementary Data 1). Therefore, we speculated that there were underlying metabolic abnormalities in these two types of tumors. We then collected fresh CRC samples for nontargeted metabolite analysis (Figs. 1D and S1I) (cohort 5, Table S5 and Supplementary Data 2) and conducted Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of the metabolomics data (Fig. 1E). For the metabolites increased in MSS CRC, we observed the enrichment of pathways related to fatty acid degradation (Fig. 1F). Notably, fatty acid metabolism-associated pathways were also enriched in the GSEA results of the RNA-seq data (Fig. S1J–L). The findings suggest that abnormalities in fatty acid metabolism may be characteristic of MSS tumors. On the basis of those findings, we treated RKO cells with five metabolites associated with fatty acid degradation—glutaric acid, palmitoyl carnitine, hexadecanal, Palmitic acid (PA), and coenzyme A—identified in our metabolomic analysis. Among these, only PA significantly increased B7H3 expression (Fig. 1G, H). To further confirm this observation, we performed dose-dependent experiments in RKO cells and found that increasing PA concentrations led to a concentration-dependent increase in B7H3 protein levels (Fig. S1M). Since PA has been widely reported to trigger palmitoylation, we employed an acyl-biotin exchange (ABE) assay46 to determine whether B7H3 undergoes palmitoylation (Fig. S1N). Our results demonstrated that B7H3 palmitoylation levels increased in response to higher concentrations of PA (Fig. 1I). Moreover, treatment with 2-bromopalmitate (2-BP), a pan-palmitoylation inhibitor, significantly reduced B7H3 palmitoylation (Figs. 1J and S1O). Like that of another member of the B7 family, B7H1 (PD-L1)38, the protein level of B7H3 was closely related to its palmitoylation status (Fig. S1P). Notably, the qPCR results indicated that neither 2-BP nor palmostatin B (an inhibitor of depalmitoylation) affected B7H3 transcript levels (Fig. S1Q). When 2-BP was applied to inhibit palmitoylation, we observed a dose- and time-dependent decrease in B7H3 expression, whereas palmostatin B treatment led to the opposite effect (Figs. 1K and S1R). Given that palmitoylation can also affect protein membrane association, we conducted cytoplasmic and membrane fractionation assays, which revealed a reduction in B7H3 expression within both the cytoplasmic and membrane fractions following 2-BP treatment (Fig. S1S). Immunofluorescence analysis confirmed this decrease in B7H3 expression (Fig. S1T). Additionally, using a cycloheximide (CHX) chase assay, we assessed the impact of B7H3 palmitoylation on protein stability. Treatment with 2-BP markedly accelerated B7H3 degradation (Fig. 1L, M), whereas inhibition of depalmitoylation with palmostatin B slowed down B7H3 degradation (Fig. S1U, V). Taken together, these findings suggest that the PA-triggered palmitoylation contributes to B7H3 stability.
A Heatmap of B7 family immunohistochemical (IHC) staining score in MSI-H and MSS tumors. B Statistical results of IHC staining scores of B7 family in MSI-H and MSS tumors. (n = 10). C RNA-seq of MSS and MSI-H tumor samples followed by Gene Ontology (GO) enrichment analysis of differentially expressed genes. D Heatmap of the top 50 representative metabolites in MSS and MSI-H tumor samples. E Kyoto encyclopedia of genes and genomes (KEGG) circos plots illustrating differential metabolites identified through untargeted metabolomics in MSS and MSI-H tumor samples. F KEGG-enriched pathway of metabolites with increased levels in MSS tumors compared to MSI-H. Immunoblot (IB) analysis of B7H3 in RKO cells treated with DMSO, glutaric acid (1 mM), palmitoylcarnitine (5 μM), hexadecanal (10 μM), palmitic acid (100 μM), or coenzyme A (3 mM) for 24 h (G) and quantification (H) (n = 3). I Acyl-biotin exchange (ABE) assay of B7H3 palmitoylation in RKO or SW480 cells after PA treatment. PA palmitic acid. HAM hydroxylamine. The samples derive from the same experiment, but different gels for B7H3, and another for Palm-B7H3 were processed in parallel. J ABE assay of B7H3 palmitoylation in RKO or SW480 cells after 2-bromopalmitate (2-BP, 50 μM) treatment. The samples derive from the same experiment, but different gels for B7H3, and another for Palm-B7H3 were processed in parallel. K IB analysis of B7H3 after 2-BP treatment at indicated doses/times. IB analysis of B7H3 stability after CHX (100 μg/ml) treatment with 2-BP (50 μM) (L) and quantification (M) (n = 3). Data indicate the mean ± SD, by unpaired 2-tailed Student’s t test (B), One-way ANOVA with Dunnett’s multiple comparisons test (H), and two-way ANOVA with Tukey’s test (M). n = 3 independent experiments (G, I–L). Source data are provided as a Source Data file.
B7H3 C496 palmitoylation inhibits SQSTM1/p62-mediated autophagic degradation
B7H3, as a membrane protein, can in principle be degraded via the ubiquitin–proteasome system (UPS), autophagy–lysosome pathway, or endocytosis to lysosomes47,48. We hypothesized that palmitoylation primarily modulates one of these routes to stabilize B7H3. Therefore, we applied pathway-specific modulators together with 2-BP. Pharmacological agents that disrupt distinct stages of the autophagy–lysosome pathway-3-MA (which inhibits class III PI3K and blocks autophagosome initiation), bafilomycin A1(Baf A1) and chloroquine (both block lysosomal degradation)—effectively blocked the 2-BP-induced degradation of B7H3, whereas the proteasome inhibitor MG132 had no such effect (Fig. 2A, B). Moreover, autophagy induction via EBSS significantly accelerated B7H3 degradation, which was further enhanced by 2-BP (Fig. S2A, B). Together, these observations indicate that B7H3 degradation in this system is predominantly mediated by the autophagy–lysosome pathway. The essential autophagy proteins ATG5 and BECN1 are required for autophagosome formation, and B7H3 degradation was markedly reduced in ATG5- or BECN1-knockout (KO) cells (Figs. 2C, D and S2C, D). Impaired autophagy also counteracted the 2-BP-induced reduction in B7H3 expression (Fig. S2E, F). Selective autophagy relies on cargo receptors that transport specific substrates to autophagosomes for degradation49,50. Among the several cargo receptors tested (SQSTM1, OPTN, NDP52, NBR1, TOLLIP, and NIX), B7H3 exhibited a strong affinity for SQSTM1, as shown by immunoprecipitation assays (Fig. 2E). Additionally, cycloheximide (CHX) chase assays revealed that B7H3 degradation was nearly completely halted in RKO cells lacking SQSTM1 (Fig. 2F, G) and that SQSTM1 knockout prevented the 2-BP-induced degradation of B7H3 (Fig. 2H). These findings collectively demonstrate that B7H3 degradation occurs primarily through SQSTM1-mediated selective autophagy in the cell models examined. To identify the specific palmitoylation site on the B7H3 protein, we examined all 13 of its cysteine residues, five of which were predicted to be palmitoylated using GPS-palm51 and GPS-lipid52 (Fig. 2I). Analysis with the Swiss-Palm database, based on known palmitoyl-proteomes, also revealed a palmitoylation site (Fig. 2I). To validate these predictions experimentally, we generated site-specific cysteine-to-alanine mutants and assessed their palmitoylation levels using ABE assays. Among the mutants, the palmitoylation of C496A was almost undetectable compared with that of wild-type B7H3 (Fig. 2J), highlighting the essential role of C496. C496 is located in the intracellular domain of B7H3 (Fig. S2G) and is highly conserved across species based on multiple sequence alignment analysis performed in Jalview53 (Fig. 2K). Consistently, the C496A mutation at the B7H3 palmitoylation site also showed greater affinity for the SQSTM1/p62 cargo receptor than WT B7H3 (Fig. 2L). To explore the functional role of C496 palmitoylation, we utilized CRISPR/Cas9 to generate B7H3-knockout RKO cells (RKOKO) (Fig. S2H) and subsequently established stable cell lines expressing the C496A mutant (RKOC496A) (Fig. S2I). ABE assays confirmed that B7H3 palmitoylation was scarcely detectable in RKOC496A cells compared with that in wild-type (RKOWT) cells (Fig. 2M). A CHX assay revealed that B7H3 protein degradation was accelerated in RKOC496A cells (Fig. 2N, O), similar to the effects of palmitoylation inhibition by 2-BP. Immunofluorescence staining further demonstrated a marked decrease in fluorescence intensity in the RKOC496A cells (Fig. S2J). In addition, we found that treatment with rapamycin, which inhibits mTORC1 activity and thereby induces autophagy, led to accelerated degradation of B7H3 protein in RKOWT cells (Fig. S2K). In contrast, treatment with 3-MA, bafilomycin A1, or chloroquine effectively prevented B7H3 degradation in RKOC496A cells (Fig. S2L). Moreover, confocal microscopy revealed that RKOC496A cells displayed greater colocalization of B7H3 with LC3 and formed more distinct B7H3–SQSTM1/p62 aggregates compared with RKOWT cells (Figs. 2P and S2M), indicating that C496 palmitoylation restrains the autophagic degradation of B7H3. In conclusion, these findings demonstrate that the palmitoylation of B7H3 at cysteine 496 reduces its interaction with the cargo receptor SQSTM1/p62, thereby hindering its selective autophagic degradation.
IB analysis of B7H3 after treatment with MG132 (10 μM), bafilomycin A1 (Baf A1, 0.2 μM), 3-methyladenine (3-MA, 5 mM), or chloroquine (CQ, 50 μM) for 8 h (A) and quantification (B). (n = 3). IB analysis of B7H3 stability after CHX (100 μg/ml) treatment in WT and BECN1 KO RKO cells (C) and quantification (D). (n = 3). E Co-immunoprecipitation (co-IP) of HA-B7H3 with indicated Flag-tagged cargo receptors in HEK293T cells. The samples derive from the same experiment, but different gels for Flag, and another for HA and GAPDH, were processed in parallel. IB analysis of B7H3 stability after CHX (100 μg/ml) treatment in WT and SQSTM1 KO RKO cells (F) and quantification (G). (n = 3). H IB analysis of B7H3 in WT and SQSTM1 KO RKO cells treated with DMSO or 2-BP (100 μM). I Schematic of the predicted B7H3 palmitoylation site. J ABE assay of palmitoylation of B7H3 WT and indicated site mutants. The samples derive from the same experiment, but different gels for B7H3, and another for Palm-B7H3 were processed in parallel. K Multiple sequence alignment of B7H3 across species by Jalview. L Co-IP of Flag-SQSTM1 with HA-B7H3 WT or C496A in HEK293T cells. M ABE assay of palmitoylation in RKO stable cell lines expressing B7H3 WT or C496A. The samples derive from the same experiment, but different gels for B7H3, and another for Palm-B7H3 were processed in parallel. IB analysis of B7H3 stability after CHX (100 μg/ml) treatment in RKOWT and RKOC496A cells (N) and quantification (O). (n = 3). P Representative images of confocal microscopy after treatment of RKOWT or RKOC496A with EBSS and CQ (50 μM) for 8 h. Scale bars, 5 μm. Data indicate the mean ± SD, by unpaired 2-tailed Student’s t test (B) and two-way ANOVA with Tukey’s test (D, G, O). n = 3 independent experiments (A, C, E, F, H, J, L–N, P). Source data are provided as a Source Data file.
B7H3 palmitoylation inhibits the antitumor activity of CD8+ T cells
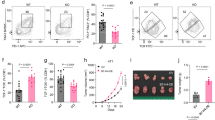
To investigate the function of B7H3 palmitoylation in vivo, we generated B7h3-knockout CT26 (CT26KO) and MC38 (MC38KO) cell lines of mouse colon cancer (Fig. S3A). Since murine B7H3 comprises only a single pair of IgC-like and IgV-like domains24, we engineered C278A mutants (CT26C278A and MC38C278A) that stably express palmitoylation-deficient B7H3, corresponding to the human modification site (Fig. S3B). Palmitoylation in these mutants was barely detectable, as confirmed by ABE assays (Fig. S3C). We then transplanted these modified mouse colon cancer cells subcutaneously into mice. In immunocompetent BALB/c mice, tumor progression was significantly inhibited in the palmitoylation-deficient B7H3 (CT26C278A) group compared with the CT26WT group (Figs. 3A, B and S3D). However, in highly immunodeficient NCG (NOD ShiLtJ Gpt Prkdcem26Cd52 Il2rgem26Cd22/Gpt) mice, no significant differences in tumor growth were observed between CT26WT and CT26C278A, as evidenced by comparable tumor weights and volumes (Figs. 3C, D and S3E). Similar results were obtained with the MC38 model, where there was no significant difference between MC38WT and MC38C278A in NCG mice (Fig. S3F–H). These findings suggest that B7H3 palmitoylation does not affect the proliferation of tumor cells directly; rather, the antitumor effects of B7H3 palmitoylation deficiency depend on the host immune system. To investigate the impact of B7H3 palmitoylation on the tumor microenvironment (TME), we conducted a comprehensive analysis of CD45+ immune cells infiltrating two types of CT26 tumors in BALB/c mice using spectral flow cytometry. In summary, we identified 14 unique cell subpopulations (Fig. 3E, F). The results revealed that some immunosuppressive cells, including monocytic myeloid-derived suppressor cells (M-MDSCs) and polymorphonuclear myeloid-derived suppressor cells (PMN-MDSCs), were reduced in palmitoylation-deficient (C278A) tumors (Figs. 3G and S3I). In contrast, some cell populations with antitumor effects, including CD8+ T, CD4+ T, and natural killer (NK) cells, were increased in C278A tumors (Figs. 3G and S3I). These results indicate that the mutation of B7H3 palmitoylation substantially alters the immune landscape within the TME, particularly by promoting CD8+ T cell infiltration (Figs. 3G, H and S3I). To test whether CD8⁺ T cells are essential mediators of antitumor effects caused by B7H3 palmitoylation deficiency, we depleted CD8⁺ T cells in vivo using an anti-CD8α antibody (Fig. 3I). Strikingly, depletion of CD8⁺ T cells completely abrogated the tumor growth difference between wild-type B7H3 and the palmitoylation-deficient C278A mutant (Figs. 3J, K and S3J), indicating that the antitumor effect associated with loss of B7H3 palmitoylation is primarily dependent on CD8⁺ T cells. To validate these findings, we subcutaneously transplanted the MC38 B7h3 knockout cell line (MC38KO) and the stable MC38WT and MC38C278A cell lines into C57BL/6J mice. The results revealed that the mutation of the palmitoylation site significantly suppressed tumor growth, an effect that was even more pronounced in the B7h3 knockout (MC38KO) group (Figs. 3L, M and S3K). An analysis of CD8+ T cell infiltration revealed that the MC38C278A group presented a significantly greater proportion of CD8+ T cells than the MC38WT group did, a trend that was further amplified in MC38KO tumors (Figs. 3N and S3L). Immunofluorescence staining of tumor tissues corroborated those findings (Fig. 3O, P). Furthermore, the expression of effector molecules of CD8+ T cells, such as granzyme B (GZMB) and interferon-gamma (IFN-γ), was higher in MC38C278A tumors compared with MC38WT tumors (Figs. 3Q, R and S3M, N). To confirm these observations, we performed a T cell-mediated tumor-killing assay by coculturing activated human primary T cells with the RKOKO, RKOWT, and RKOC496A cell lines. The RKOC496A mutation markedly enhanced the cytotoxicity of T cells to tumor cells (Figs. 3S and S3O). Consistent with the flow cytometry findings, the lactate dehydrogenase (LDH) cytotoxicity assay yielded similar results (Fig. 3T). Finally, using tissue microarrays of colorectal cancer samples, we examined the correlation between B7H3 expression and CD8+ T cell infiltration. Patients with high B7H3 expression had shorter overall survival and significantly fewer CD8+ T cells than did those with low B7H3 expression (Fig. S3P, Q). Consistently, Kaplan–Meier Plotter analysis further confirmed that high B7H3 expression is associated with poorer overall survival (Fig. S3R), and TIMER database analysis showed that high B7H3 (CD276) expression correlated with lower CD8⁺ T cell infiltration (Fig. S3S). Furthermore, gene set enrichment analysis (GSEA) based on RNA-seq data from colorectal tumor samples (cohort 4, Fig. 1C) revealed that the gene set negative regulation of T cell activation was significantly enriched in MSS tumors (Fig. S3T). In contrast, pathways related to T cell function, including CD8-positive, alpha-beta T cell differentiation and the T cell receptor signaling pathway, were negatively enriched in MSS tumors (Fig. S3U, V), suggesting impaired cytotoxic T cell functionality. Collectively, these findings suggest that B7H3 palmitoylation enhances its immunosuppressive function by limiting CD8+ T cell infiltration and activity, thereby contributing to tumor immune evasion.
Tumor growth (A) and tumor weight (B) of CT26WT versus CT26C278A tumors in BALB/c mice (n = 5). Tumor growth (C) and tumor weight (D) of CT26WT versus CT26C278A tumors in NCG mice (n = 5). Spectral flow cytometry analysis of CD45⁺ immune infiltrates in CT26WT and CT26C278A tumors from BALB/c mice (n = 5): UMAP clustering (E), immune composition (F), proportions of immune subpopulations (G), and CD8⁺ T-cell density map (H). I–K C57BL/6J mice were intraperitoneally injected with 200 μg of anti-CD8α depleting antibody or isotype control per mouse one day before subcutaneous inoculation of MC38WT or MC38C278A cells, and then once weekly thereafter for a total of three doses (n = 5). Schematic (I), tumor growth curves (J), and tumor weights (K). (L to R). MC38KO, MC38WT, and MC38C278A cells were injected subcutaneously into C57BL/6 J mice (n = 5). L Tumor growth curves. M Tumor weights. N Relative levels of tumor-infiltrating CD8+ T by flow cytometry. O Representative images of immunofluorescence of CD8+ T cells and CD45+ cells in subcutaneous graft tumors. Scale bars, 50 μm. P Quantification of CD8⁺ cells among CD45⁺ tumor-infiltrating leukocytes (TILs) from (O) (n = 5). For each tumor, three fields were quantified and averaged to obtain one value per tumor. Q Flow cytometry analysis of Granzyme B (GZMB) expression in tumor-infiltrating CD8+ T cells. R Flow cytometry analysis of Interferon-γ (IFN-γ) expression in tumor-infiltrating CD8+ T cells. S, T RKOKO, RKOWT, and RKOC496A cells were co-cultured with activated human T lymphocytes (n = 5). S Statistical analysis shows the percentage of caspase-3 in tumor cells by flow cytometry. T Cell-free supernatants were collected, and LDH release was measured to calculate percent cytotoxicity. Data indicate the mean ± SD, by unpaired 2-tailed Student’s t test (B, D), one-way ANOVA with Tukey’s test (K, M, N, P, Q, R), and two-way ANOVA with Tukey’s test (A, C, J, L, S, T). Source data are provided as a Source Data file.
ZDHHC24 mediates the palmitoylation of B7H3 at C496 and is correlated with poor survival in CRC patients
To identify which ZDHHC(s) catalyze B7H3 palmitoylation, we co-expressed each of the 23 FLAG-tagged ZDHHC enzymes with HA-B7H3. ZDHHC24 significantly increased the palmitoylation level of B7H3, whereas ZDHHC3, ZDHHC8, and ZDHHC13 only slightly increased the B7H3 palmitoylation level (Fig. S4A). Subsequent transfection of RKO cells with ZDHHC3, ZDHHC8, ZDHHC13, or ZDHHC24 indicated that only ZDHHC24 substantially increased B7H3 expression (Fig. S4B–E). Those findings led us to hypothesize that ZDHHC24 is the primary enzyme responsible for B7H3 palmitoylation. To confirm the interaction between B7H3 and ZDHHC24, we conducted coimmunoprecipitation (co-IP) assays, which revealed a physical interaction at the endogenous level in RKO and SW480 cells (Fig. 4A). Consistent with these findings, we observed the colocalization of B7H3 and ZDHHC24 through immunofluorescence (Fig. 4B). To assess the functional role of ZDHHC24, we used short hairpin RNA (shRNA) to stably knock down ZDHHC24 in human RKO cells and Zdhhc24 in murine MC38 cells (Fig. S4F, G). ABE assays subsequently revealed a significant decrease in B7H3 palmitoylation levels following ZDHHC24 or Zdhhc24 knockdown (Figs. 4C and S4H). In line with the above results, ZDHHC24 or Zdhhc24 knockdown also led to a reduction in endogenous B7H3 protein expression (Fig. 4D, E), an effect further validated by confocal microscopy (Fig. 4F). Notably, the transcript levels of B7H3 showed minimal changes following ZDHHC24 knockdown (Fig. S4I). To confirm this, we performed a CHX assay and found that ZDHHC24 knockdown accelerated the degradation of B7H3 protein (Fig. 4G, H). Additionally, when Flag-ZDHHC24 was transfected into RKOWT and RKOC496A cells, an increase in endogenous B7H3 expression was observed exclusively in RKOWT cells but not in palmitoylation-deficient RKOC496A cells (Fig. 4I). Moreover, we assessed the palmitoylation levels of RKOWT and RKOC496A following ZDHHC24 transfection. The results indicated that ZDHHC24 increased palmitoylation of RKOWT, but not that of the palmitoylation site-deficient RKOC496A variant (Fig. 4J). Interestingly, co-IP experiments also revealed that in the palmitoylation-deficient mutant RKOC496A, B7H3 exhibited a stronger interaction with SQSTM1/p62 proteins compared to RKOWT, and that ZDHHC24 reduced this interaction in RKOWT but not in RKOC496A cells (Fig. 4K). In addition, ZDHHC24 knockdown promoted punctate accumulation and colocalization of B7H3 and p62, as demonstrated by immunofluorescence (Fig. 4L). This enhanced interaction was further confirmed by proximity ligation assay (PLA) (Fig. 4M). An analysis of the Gene Expression Profiling Interactive Analysis (GEPIA) database revealed substantial ZDHHC24 overexpression in multiple tumor types, particularly in colorectal cancer (Fig. S4J). Tissue microarray analysis of colorectal cancer specimens revealed that patients with high ZDHHC24 expression had shorter overall survival (Fig. 4N), a finding further supported by Kaplan–Meier Plotter database analysis, which linked high ZDHHC24 expression to worse relapse-free survival in colon cancer (Fig. S4K). Moreover, ZDHHC24 expression was markedly higher in tumor tissues than in adjacent normal tissues and was positively correlated with clinical tumor stage (Fig. 4O, P). It is worth noting that no significant differences in B7H3 or ZDHHC24 protein levels were detected between MSS and MSI-H subtypes in colorectal cancer cell lines (Fig. S4L). However, patient-derived tumor specimens showed that ZDHHC24 expression was significantly higher in MSS tumors than in MSI-H tumors (Fig. S4M). This suggests that the in vivo tumor microenvironment, which is absent in standard cell culture systems, may modulate ZDHHC24 expression. Importantly, ZDHHC24 expression was positively correlated with B7H3 levels in tissue microarray samples (Fig. 4Q). Together, these findings highlight the critical role of ZDHHC24-mediated B7H3 C496 palmitoylation in maintaining B7H3 protein stability and underscore its association with poor survival outcomes in patients with colorectal cancer.
A Co-IP showing endogenous B7H3–ZDHHC24 interaction. B Confocal images showing co-localization of B7H3 and ZDHHC24 in RKO cells. Scale bars, 5 μm. C ABE assay of B7H3 palmitoylation in ZDHHC24-knockdown RKO cells. IB of B7H3 in RKO (D) and MC38 (E) cells with control or ZDHHC24/Zdhhc24 knockdown. F Confocal images of B7H3 staining in RKO shNC or shZDHHC24 cells. Scale bars, 5 μm. IB of B7H3 after CHX (100 μg/mL) for the indicated times (G) and quantification (H) (n = 3). IB of B7H3 protein levels (I) and ABE assay of B7H3 palmitoylation (J) after ZDHHC24 overexpression in RKOB7H3 WT and RKOB7H3 C496A cells. K Co-IP of B7H3 with SQSTM1 in RKOB7H3 WT and RKOB7H3 C496A cells with or without Flag-ZDHHC24 expression. L Confocal images of RKO shNC or shZDHHC24 cells after EBSS and CQ (50 μM) treatment for 8 h. Scale bars, 5 μm. M Duolink PLA showing B7H3–p62 interactions. Scale bars, 5 μm. N Kaplan–Meier overall survival stratified by ZDHHC24 expression (n = 94; log-rank test). O ZDHHC24 IHC staining in paired adjacent and primary tissues (n = 86 pairs). P ZDHHC24 IHC staining scores across tumor stages (n = 94). Q Association between ZDHHC24 and B7H3 IHC staining (n = 94). The samples (D, E, G, I, K) derive from the same experiment, but different gels for ZDHHC24/Flag, and another for B7H3 and GAPDH were processed in parallel. The samples (C, J) derive from the same experiment, but different gels for ZDHHC24, another for GAPDH, another for B7H3, and another for Palm-B7H3 were processed in parallel. Data indicate the mean ± SD, by two-way ANOVA with Tukey’s test (H), paired 2-tailed Student’s t test (O), chi-square test (P), and two-sided chi-square test (Q). n = 3 independent experiments (A–G, I–M). Source data are provided as a Source Data file.
ZDHHC24 promotes immunosuppression in colorectal cancer
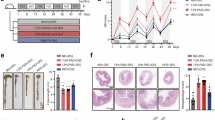
To examine the biological function of ZDHHC24 in vivo, we generated Zdhhc24 floxed (Zdhhc24fl/fl) mice (Fig. S5A) and crossed them with Villin-Cre transgenic animals (Fig. S5B) to generate intestinal epithelial cell (IEC) specific Zdhhc24-deficient (Zdhhc24ΔIEC) models (Fig. S5C). The efficacy of Cre recombinase was confirmed by crossing the Villin-Cre line with Rosa26-CAG-LSL-tdTomato reporter (Fig. S5D) mice. As expected, intestinal epithelial cells presented specific red fluorescence under the microscope (Fig. S5E). We further isolated IECs to verify the specific deletion of Zdhhc24 in Zdhhc24ΔIEC mice (Fig. S5F). Subsequently, we investigated the role of ZDHHC24 in CRC malignancy by establishing a colitis-associated colorectal (CAC) carcinogenesis mouse model (Fig. 5A). During CAC induction, Zdhhc24ΔIEC mice presented less weight loss than Zdhhc24fl/fl mice did (Fig. 5B). At the end of the observation period, the Zdhhc24fl/fl mice had slightly shorter colons than their Zdhhc24ΔIEC littermates did (Figs. 5C and S5G). Additionally, Zdhhc24ΔIEC mice developed fewer tumors (Fig. 5D, E) and presented significantly fewer large tumors (> ~1 mm) (Fig. 5F, G). We collected tumor tissues from Zdhhc24ΔIEC and Zdhhc24fl/fl and assessed B7H3 protein expression and palmitoylation levels. B7H3 protein expression and palmitoylation were both reduced in Zdhhc24ΔIEC mice (Fig. S5H, I). Histological examination revealed that Zdhhc24ΔIEC mice presented reduced colon thickening, inflammation, hyperplasia, and injury in the affected regions (Fig. 5H). Immunofluorescence analysis also revealed increased CD8+ T cell infiltration in tumors from Zdhhc24ΔIEC mice (Fig. 5I). These findings suggest that ZDHHC24 modulates antitumor immunity during colon tumor development. To validate these results, we transplanted Zdhhc24-knockdown mouse colon cancer cells (MC38) subcutaneously into C57BL/6J mice. Tumor growth was markedly reduced in the Zdhhc24-knockdown group compared with the control group, as shown by tumor growth curves and final tumor weights (Figs. 5J, K and S5J). Flow cytometry of single-cell suspensions from these tumors revealed a substantial increase in CD8+ T cell infiltration in the Zdhhc24-knockdown group (Figs. 5L and S5K), an effect that was corroborated by immunofluorescence analysis (Fig. S5L, M). Furthermore, the levels of functional markers, including GZMB and IFN-γ, were significantly greater in tumor-infiltrating CD8+ T cells in the Zdhhc24-knockdown group than in those in the control group (Figs. 5M, N and S5N, O). We subsequently established an in vitro cytotoxic T-cell killing assay for validation. Compared with control cells, cells with shRNA-mediated ZDHHC24 knockdown displayed increased susceptibility to apoptosis when activated T cells derived from human peripheral blood mononuclear cells were exposed (Fig. S5P, Q). This finding was supported by increased LDH levels in the culture supernatants of the respective cell populations (Fig. S5R). Additionally, IHC analysis of CRC tissue microarrays revealed that patients with high ZDHHC24 expression had lower CD8+ T cell counts in tumor tissues (Fig. 5O). A three-dimensional scatter plot further illustrated the correlation among ZDHHC24, B7H3, and CD8+ T cells (Fig. 5P). Collectively, these results suggest that ZDHHC24 promotes immunosuppression in colorectal cancer by inhibiting CD8+ T cell-mediated antitumor responses.
A Schematic of azoxymethane (AOM)/ dextran sulfate sodium (DSS)-induced colitis-associated colorectal cancer (CAC) model. Body weight change (B) and colon length (C) of Zdhhc24fl/fl and Zdhhc24ΔIEC mice during AOM/DSS treatments (n = 8). D Representative images of Zdhhc24fl/fl and Zdhhc24ΔIEC mice after longitudinal sectioning of the colon. Tumor number (E) and tumor size distributions (F, G) in Zdhhc24fl/fl and Zdhhc24ΔIEC mice (n = 8). Representative H&E staining (H) and immunofluorescence staining (I) of colon tissues/tumors in Zdhhc24fl/fl and Zdhhc24ΔIEC mice. Scale bars, 0.25 mm (H), 400 μm (top in I), and 50 μm (bottom in I). J–N MC38 shNC, MC38 shZdhhc24 (#1 and #2) cells were injected subcutaneously into C57BL/6J mice (n = 5). J Tumor growth curves. K Tumor weights. L Relative levels of tumor-infiltrating CD8+ T by flow cytometry. M Flow cytometry analysis of GZMB expression in tumor-infiltrating CD8+ T cells. N Flow cytometry analysis of IFN-γ expression in tumor-infiltrating CD8+ T cells. O Quantification of CD8+ T cell percentage in CRC tissue microarrays stratified by ZDHHC24 IHC expression (ZDHHC24Low, n = 48; ZDHHC24High, n = 46). Violin plots show the data distribution. Box plots show the median (center marker), the 25th and 75th percentiles (box limits) and whiskers extending to 1.5 × IQR; points beyond the whiskers represent outliers. P Correlation of ZDHHC24, B7H3 and CD8⁺ T cell IHC staining scores in CRC tissue microarrays (n = 94) assessed by Pearson’s correlation (two-tailed); r and exact P values are shown (no adjustment for multiple comparisons was applied). Data indicate the mean ± SD, by unpaired 2-tailed Student’s t test (B, C, E, F, O), one-way ANOVA with Tukey’s test (K, L, M, N). Two-way ANOVA with Tukey’s test (J). n = 3 independent experiments (H, I). Source data are provided as a Source Data file.
B7H3 palmitoylation is necessary for the ZDHHC24-mediated suppression of tumor immunity
To further investigate the relationship between B7H3 palmitoylation and ZDHHC24 in modulating the tumor immune microenvironment, we stably overexpressed Zdhhc24 in MC38WT and MC38C278A cells (Fig. S6A) and subcutaneously implanted these cells into C57BL/6J mice. ZDHHC24 significantly promoted tumor growth in the MC38WT group; however, this effect was nearly absent in the MC38C278A group (Figs. 6A, B and S6B). An analysis of immune cell infiltration revealed a greater proportion of CD8+ T cells in MC38C278A tumors, whereas ZDHHC24 overexpression reduced CD8+ T cell levels in MC38WT tumors but had no effect on MC38C278A tumors (Figs. 6C and S6C). These results were consistent with the immunofluorescence findings (Fig. 6D, E). Furthermore, CD8+ T cells expressing GZMB and IFN-γ were more common in MC38C278A tumors than in MC38WT tumors (Fig. 6F, H). While ZDHHC24 decreased the levels of GZMB- and IFN-γ-expressing CD8+ T cells in MC38WT tumors, it had no effect on MC38C278A tumors (Fig. 6F–I). In vitro, cytotoxic T cell assays with RKOWT and RKOC496A cells overexpressing ZDHHC24 (Fig. S6D) demonstrated that ZDHHC24 did not affect apoptosis in the absence of T cells (Figs. 6J and S6E). However, in the presence of T cells, ZDHHC24 significantly reduced apoptosis in RKOWT cells but not in RKOC496A cells (Figs. 6J, K and S6E). Collectively, these findings confirm that ZDHHC24-mediated tumor immunosuppression depends on B7H3 palmitoylation.
A–I MC38WT+Vector, MC38WT+Zdhhc24, MC38C278A+Vector, and MC38C278A+ Zdhhc24 cells were injected subcutaneously into C57BL/6J mice (n = 5). A The volume of the subcutaneous graft tumors was measured on the indicated days, and growth curves were drawn. B Quantification of measured weights of subcutaneous graft tumors. C Relative levels of tumor-infiltrating CD8+ T cells by flow cytometry. D Representative immunofluorescence images of tumor tissues. Scale bars, 50 μm. E Quantification of CD8⁺/CD45⁺ TILs from (D) (n = 5). For each tumor, three fields were quantified and averaged to obtain one value per tumor. F Flow cytometry analysis of IFN-γ expression in tumor-infiltrating CD8+ T cells. G Representative images of tumor-infiltrating IFN-γ+ CD8+ T cells analyzed by flow cytometry. H Flow cytometry analysis of GZMB expression in tumor-infiltrating CD8+ T cells. I Representative images of tumor-infiltrating GZMB+ CD8+ T cells analyzed by flow cytometry. J, K RKOWT + Vector, RKOWT + ZDHHC24, RKOC496A + Vector, and RKOC496A + ZDHHC24 cells were co-cultured with activated human T lymphocytes (n = 5). J Statistical analysis shows the percentage of caspase-3+ in tumor cells by flow cytometry. K Cell-free supernatants were collected, and LDH release was measured to calculate percent cytotoxicity. Data indicate the mean ± SD, by one-way ANOVA with Tukey’s test (B, C, E, F, H) and two-way ANOVA with Tukey’s test (A, J, K). Source data are provided as a Source Data file.
Inhibition of B7H3 palmitoylation with synthetic peptides
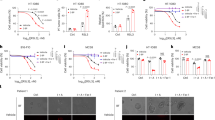
Given the critical role of ZDHHC24-mediated palmitoylation of B7H3 at C496 in promoting an immunosuppressive microenvironment in colorectal cancer, we further investigated the therapeutic potential of targeting B7H3 palmitoylation. To better understand the binding interaction between ZDHHC24 and B7H3, we analyzed the structural domains of B7H3. As an immune checkpoint protein, the extracellular Ig-like domain of B7H3 likely contributes to antigen recognition and immunomodulation. Thus, we hypothesized that the primary binding sites for ZDHHC24 are located within the transmembrane and intracellular regions. To test this hypothesis, we generated B7H3 constructs with targeted deletions in specific structural domains and conducted immunoprecipitation with ZDHHC24 (Fig. 7A). Our results revealed that amino acids 488–497 in the B7H3 sequence are essential for its interaction with ZDHHC24 (Fig. 7B). Notably, this region is highly conserved in human and mouse and encompasses the palmitoylation site in B7H3, underscoring its functional relevance. We subsequently designed a peptide (E-peptide) corresponding to the B7H3 488–497 amino acid sequence, and a control peptide (C-peptide) in which the same amino acids were randomly scrambled to disrupt the sequence-specific structure and abolish potential binding, following a previously described strategy54. Both peptides were fused with an N-terminal transcriptional activator of transcription (TAT) tag to facilitate cellular uptake (Fig. 7C). We assessed the cytotoxicity of the peptides in vitro and determined that cytotoxicity was observed only at concentrations exceeding 50 μM (Fig. S7A). We then attached a fluorescent motif to the first lysine of both the E-peptide and C-peptide (Fig. S7B) and confirmed its cellular entry via fluorescence microscopy (Fig. S7C). To evaluate the functionality of the synthesized peptides, we treated cells with E-peptide and C-peptide. In contrast to C-peptide, E-peptide significantly inhibited the binding of ZDHHC24 to B7H3, suggesting that E-peptide effectively competes with B7H3 for binding to ZDHHC24 (Fig. 7D). In parallel, we employed surface plasmon resonance (SPR) analysis to confirm the direct interaction between the E-peptide and ZDHHC24 (Fig. S7D, E). We further assessed B7H3 palmitoylation levels and found a significant reduction in palmitoylation with E-peptide treatment but not with C-peptide treatment (Fig. 7E). Notably, the decrease in B7H3 protein levels was dose dependent in RKOWT cells treated with E-peptide, whereas neither peptide affected B7H3 expression in RKOC496A cells (Fig. 7F). Additionally, E-peptide, but not C-peptide, markedly reduced B7H3 palmitoylation in RKO control cells, with limited effects in ZDHHC24 knockdown cells (Figs. 7G and S7F). Similarly, E-peptide treatment also reduced B7H3 protein levels in RKO control cells, but the decrease was not significant in ZDHHC24 knockdown cells (Fig. 7H, I). Moreover, co-immunoprecipitation analysis showed that E-peptide treatment enhanced the interaction between B7H3 and p62 (Fig. 7J). To determine the impact of synthesized peptides on immune function in vitro, RKO cells were treated with either E-peptide or C-peptide and subsequently coincubated with activated human CD8+ T cells. Both flow cytometry and lactate dehydrogenase toxicity assays demonstrated that E-peptide effectively enhanced CD8+ T cell killing (Figs. 7K, L and S7G). In conclusion, these data suggest that E-peptide inhibits B7H3 palmitoylation and promotes its degradation by disrupting the interaction between B7H3 and ZDHHC24, thereby effectively enhancing CD8+ T cell-mediated cytotoxicity in vitro.
A Schematic representations of various B7H3 deletion mutants. B Co-IP of Flag-ZDHHC24 with HA-B7H3 deletion mutants in HEK293T cells. The samples derive from the same experiment, but different gels for Flag, another for HA, and another for GAPDH were processed in parallel. C Schematic amino acid sequences of E-peptide and C-peptide. D Co-IP of HA-B7H3 with Flag-ZDHHC24 in HEK293T cells after E-peptide or C-peptide treatment (10 μM). The samples derive from the same experiment, but different gels for Flag, another for HA and GAPDH were processed in parallel. E ABE assay of B7H3 palmitoylation in HA-B7H3–transfected HEK293T cells after E-peptide or C-peptide treatment (10 μM). The samples derive from the same experiment, but different gels for B7H3, and another for Palm-B7H3 were processed in parallel. F IB analysis of B7H3 in RKOWT or RKOC496A cells treated with indicated concentrations of E-peptide or C-peptide. G ABE assay of B7H3 palmitoylation in RKO shNC, RKO shZDHHC24 (#1 and #2) cells after E-peptide (10 μM) treatment. The samples derive from the same experiment, but different gels for ZDHHC24, another for GAPDH, another for B7H3, and another for Palm-B7H3 were processed in parallel. IB analysis of B7H3 after E- or C-peptide treatment (10 μM) in RKO shNC, or shZDHHC24 (#1 and #2) cells (H) and quantification (I) (n = 3). The samples derive from the same experiment, but different gels for ZDHHC24, another for B7H3 and GAPDH were processed in parallel. J Co-IP of HA-B7H3 with Flag-SQSTM1 in HEK293T cells after E-peptide or C-peptide treatment (10 μM). K, L RKO cells were treated with E-peptide and C-peptide (10 μM) and then co-cultured with activated human T lymphocytes (n = 5). NS: normal saline. K Statistical analysis of the percentage of caspase-3+ in tumor cells by flow cytometry. L Cell-free supernatants were collected, and LDH release was measured to calculate percent cytotoxicity. Data indicate the mean ± SD, by two-way ANOVA (I, K, L). n = 3 independent experiments (B, D–H, J). Source data are provided as a Source Data file.
E-peptide enhances antitumor immunity in vivo
To further evaluate the in vivo efficacy of the synthesized small-molecule peptides, we conducted a toxicity assessment. Mice received five different concentrations of peptides via intraperitoneal injection (Fig. S8A). Only one mouse died at each of the 25 and 50 mg/kg doses, while all others survived throughout the observation period (Fig. S8B). At the end of the experiment, the surviving mice were sacrificed, and peripheral blood and organ tissues were collected for analysis. Hematoxylin–eosin (HE) staining revealed mild hepatic steatosis at the 50 mg/kg dose, but no significant histological abnormalities were observed in other organs (Fig. S8C). Additionally, no notable changes were observed in the levels of creatinine (CRE), alanine aminotransferase (ALT) or aspartate aminotransferase (AST) in the peripheral blood (Fig. S8D–F), indicating minimal systemic toxicity at lower concentrations. To evaluate hemocompatibility, we performed in vitro hemolysis assays and measured prothrombin time (PT) and activated partial thromboplastin time (APTT). The E-peptide did not induce hemolysis (Fig. S8G, H) or prolong PT or APTT (Fig. S8I, J), demonstrating good blood compatibility under the tested conditions. Next, we subcutaneously transplanted MC38 cells into C57BL/6J mice and administered the peptides intraperitoneally (10 mg/kg) after transplantation, with normal saline serving as a control (Fig. 8A). To verify intratumoral peptide infiltration, a separate cohort received 5′-FITC-conjugated peptides via intraperitoneal injection. Immunofluorescence of tumor sections confirmed effective infiltration of the peptides into the tumor, establishing a spatial basis for their biological activity (Fig. S8K). We then assessed the effect of E-peptide treatment on B7H3 expression and its palmitoylation in tumor tissues. E-peptide administration significantly reduced both total B7H3 protein and its palmitoylation levels (Fig. S8L, M). Importantly, compared with NS and C-peptide, E-peptide significantly slowed tumor growth (Fig. 8B–D), and similar results were observed in BALB/c mice (Fig. 8E–G). Flow cytometry analysis of CD8+ T cells in BALB/c mice revealed a significant increase in CD8+ T cell infiltration within the tumor microenvironment following E-peptide treatment, whereas no such increase was observed with C-peptide (Figs. 8H and S8N). Moreover, the proportion of IFN-γ+ CD8+ T cells and GZMB+ CD8+ T cells—key indicators of tumor-killing activity—was significantly elevated after E-peptide treatment (Fig. 8I–L). To further evaluate the therapeutic efficacy of the E-peptide in a human immune context, we employed immune-reconstituted humanized mice bearing subcutaneous SW480 human colorectal tumors. Mice were treated with either the control C-peptide or E-peptide (Fig. 8M). E-peptide treatment significantly suppressed tumor growth (Fig. 8N–P), enhanced CD8⁺ T cell infiltration (Fig. 8Q, R), and increased the expression of cytotoxic effector molecules GZMB and IFN-γ in the tumor microenvironment (Figs. 8S–V). Together, these results demonstrate that the E-peptide effectively enhances CD8⁺ T cell-mediated antitumor immunity in vivo.
A–D MC38 tumors in C57BL/6J mice treated with E-peptide or C-peptide (10 mg/kg) (n = 5). A Treatment scheme. B Tumor growth curves. C Tumor weights. D Representative tumor images. E–G CT26 tumors in BALB/c mice treated with E-peptide or C-peptide (10 mg/kg) (n = 5). E Tumor growth curves. F Tumor weights. G Representative tumor images. H–L Flow cytometry analysis of tumor-infiltrating lymphocytes from (G) (n = 5). H Proportion of CD8⁺ T cells among CD45⁺ cells. I GZMB expression in tumor-infiltrating CD8+ T cells. J Representative images of GZMB+CD8+ T cells. K IFN-γ expression in tumor-infiltrating CD8+ T cells. L Representative images of IFN-γ+ CD8+ T cells. M–P Human umbilical cord blood-derived CD34⁺ hematopoietic stem cells were intravenously injected into C-NKG mice via the superficial temporal vein. After 12 weeks, engraftment was confirmed by flow cytometry of peripheral blood, followed by subcutaneous inoculation of SW480. Mice were treated with either C-peptide or E-peptide on the indicated days (n = 5). M Schematic illustration of the humanized mouse model and treatment schedule. N Tumor growth curves. O Representative tumor images. P Tumor weights. Q–V Flow cytometry analysis of tumor-infiltrating lymphocytes from (O) (n = 5). Q Ratio of h-CD8+ T cells to h-CD45+ cells. R Representative images of tumor-infiltrating h-CD8+ T cells. S h-GZMB expression in tumor-infiltrating h-CD8+ T cells. T Representative images of h-GZMB+ h-CD8+ T cells. U h-IFN-γ expression in tumor-infiltrating h-CD8+ T cells. V Representative images of h-IFN-γ+ h-CD8+ T cells. The prefix h- denotes human-derived immune markers detected in humanized mice. Data indicate the mean ± SD, by unpaired 2-tailed Student’s t test (P, Q, S, U), one-way ANOVA with Tukey’s test (C, F, H, I, K), and two-way ANOVA with Tukey’s test (B, E, N). Source data are provided as a Source Data file.
The combination of E-peptide and anti-PD-1 therapy exerts a significant antitumor effect
A single immune checkpoint inhibitor may be insufficient to address the complexities of the tumor immune microenvironment. Increasing evidence suggested that combination therapies involving immune checkpoint inhibitors yield better clinical outcomes than monotherapies do55,56. Therefore, we hypothesized that E-peptide synergizes with PD-1 blockade, leading to more effective tumor suppression. We initially subcutaneously injected BALB/c mice with luciferase-expressing CT26 cells, followed by the intraperitoneal administration of anti-PD-1, E-peptide, or a combination of anti-PD-1 and E-peptide, with the IgG group serving as the control (Fig. S9A). The results demonstrated that monotherapies with either E-peptide or anti-PD-1 antibodies had moderate antitumor activity. However, the combination treatment significantly enhanced tumor inhibition, reducing the tumor size to a minimal level (Fig. 9A, B). To validate these findings, we expanded the sample size for each group and conducted a 60-day observation. The results confirmed that, compared to E-peptide or anti-PD-1 monotherapy, the combination therapy significantly inhibited tumor growth and even led to complete tumor regression in one mouse (Fig. 9C–G). Survival data (Fig. 9G) show a mean survival of 18.1 days in the IgG control, versus 24.4 days with anti-PD-1 ( + 34.8%) and 25.2 days with E-peptide (+39.2%). The combination of E-peptide and anti-PD-1 extended mean survival to 40.5 days (~124% over IgG). To assess interaction under the independent-action assumption, we estimated the Bliss-expected combination effect for E-peptide plus anti-PD-1 from their single-agent effects (defined as the normalized improvement in mean survival over IgG). This yielded an expected additive improvement of 60.4%, corresponding to a predicted mean survival of 29.0 days. The observed mean survival (40.5 days) exceeds the Bliss expectation, consistent with a synergistic interaction. Additionally, we validated the findings in C57BL/6J mice, which yielded results consistent with those observed in the BALB/c model (Figs. 9H, I and S9B). Moreover, flow cytometry analysis was performed to assess the impact of different treatments on tumor-infiltrating CD8+ T cells. The results demonstrated that the combination of E-peptide and anti-PD-1 antibodies significantly increased the number of CD8+ T cells and enhanced their functional activity (Figs. 9J–L and S9C–E). In summary, our findings emphasize the pivotal role of ZDHHC24-mediated B7H3 palmitoylation in promoting immune escape in colorectal cancer and demonstrate that targeting this modification could enhance antitumor immune responses, suggesting a promising strategy for improving the efficacy of immunotherapy in patients with colorectal cancer.
A, B CT26-luciferase tumors in BALB/c mice treated with anti-PD-1 (10 mg/kg), E-peptide (10 mg/kg), or the combination (n = 5). A Representative bioluminescence images. B Quantification of luminescence intensity. C–G BALB/c mice were injected subcutaneously with CT26 cells. Once tumors became measurable, mice were randomized into treatment groups and received IgG, anti-PD-1, E-peptide, or the combination by intraperitoneal injection every 3 days for a total of five doses (n = 10 mice per group). Mice were monitored until endpoint criteria were met (tumor volume > 2000 mm³ or >60 days). Tumor growth curves for the IgG (C), anti-PD-1 (D), E-peptide (E), and combination (F) groups. G Overall survival (Log-rank test). H–L MC38 cells were injected subcutaneously into C57BL/6J mice, and 1 week later, PD-1 (10 mg/kg), E-peptide (10 mg/kg), or a combination of both was administered intraperitoneally every 3 days for a total of 3 injections (n = 5). H Tumor growth curves. I Tumor weights. J Tumor-infiltrating CD8+ T cells by flow cytometry. K GZMB expression in tumor-infiltrating CD8+ T cells by flow cytometry. L IFN-γ expression in tumor-infiltrating CD8+ T cells by flow cytometry. M Working model illustrating that PA-triggered, ZDHHC24-mediated palmitoylation of the B7H3 at C496 stabilizes the protein by inhibiting its binding to the autophagy receptor SQSTM1/p62, thereby suppressing the antitumor activity of CD8+ T cells. Data indicate the mean ± SD, by one-way ANOVA with Tukey’s test (B, I, J, K, L) and two-way ANOVA with Tukey’s test (H). Source data are provided as a Source Data file.
Discussion
Various mechanisms of immunosuppression and evasion of malignant tumor cells are considered hallmarks of malignancy57. Immune checkpoint inhibitors, including anti-PD-1, anti-PD-L1, and anti-CTLA-4 antibodies, have shown durable efficacy in treating and potentially curing many types of tumors, ushering in a new era of tumor treatment58. However, the current approval of immune checkpoint inhibitors for the treatment of colorectal cancer is limited to a subset of MSI-H tumors, suggesting the potential involvement of alternative immune checkpoints in regulating immune escape in MSS colorectal cancer7. B7H3, an emerging immune checkpoint, is highly expressed across various human malignancies but is expressed at low levels in normal tissues, making it a promising target for the development of future immune checkpoint inhibitors24,59. Notably, despite low protein expression in normal tissues, B7H3 mRNA is detectable in a wide range of normal tissues and organs, suggesting the presence of significant posttranscriptional regulation23.
Metabolites not only provide energy for life activities but also regulate the PTM of proteins through derivation into metabolically active products60,61,62. PA, a long-chain saturated fatty acid, has been extensively reported as a metabolite that facilitates the crosstalk between metabolic processes and posttranslational protein modifications41,42,63,64. Consistent with the B7 family immunohistochemistry results, we demonstrated that PA triggered the palmitoylation of B7H3, which played a crucial role in the formation of an immunosuppressive microenvironment in MSS colorectal cancer. Although our non-targeted metabolomic profiling revealed elevated levels of PA in MSS tumors compared to MSI-H tumors, the precise mechanisms underlying this difference remain unclear. However, several plausible explanations may account for this observation. First, fatty acid-binding protein (FABP1), a key transporter of long-chain fatty acids, is preferentially lost in MSI-H tumors due to IFN-γ-driven immune signaling, but retained in MSS tumors, potentially facilitating lipid accumulation65. In addition, compared with MSS tumors, MSI-H tumors are enriched with metabolically active immune cells, which may consume fatty acids to support their effector functions66. Furthermore, fatty acid β-oxidation is suppressed in MSS tumors, leading to reduced lipid degradation and a possible increased intracellular accumulation of saturated fatty acids67. Notably, analyses of TCGA and GEO datasets also revealed that fatty acid-related metabolic pathways—including fatty acids, lipid localization, and lipid transport—are significantly downregulated in MSI-H colorectal cancers, consistent with our findings66. In our study, we found that PA were elevated in MSS tumors and promoted palmitoylation of B7H3 at the C496 residue. This modification hindered the binding of B7H3 to the autophagy receptor SQSTM1/p62, thereby reducing its autophagic degradation and enhancing B7H3 protein stability. Moreover, we employed an exhaustive screening approach to identify ZDHHC24-a DHHC family member with a conserved DHHC Cys domain32—as the enzyme that catalyzes B7H3 palmitoylation. Previously, ZDHHC24 has been shown to palmitoylate AKT, enabling membrane anchoring independent of PIP341, and to palmitoylate MAVS, promoting its aggregation and mitochondrial localization42. In our study, we found that ZDHHC24 catalyzed the palmitoylation of the B7H3 protein and that its expression level was positively correlated with that of B7H3. Furthermore, the expression level of ZDHHC24 is positively associated with advanced cancer grade and serves as a prognostic biomarker for poor clinical outcomes in patients with colorectal cancer.
Multiple types of cancer are categorized as “cold” tumors, characterized by a low tumor mutational burden, minimal immune cell infiltration, and limited expression of PD-L111,68, which correlate with poor responses to PD-1/PD-L1 monotherapy69. In addition, compensatory mechanisms between immune checkpoints may also limit the efficacy of monotherapy. In many tumors, inhibition of one pathway leads to upregulation of another. For example, Shi et al. found that inhibiting B7H3 leads to compensatory upregulation of PD-L1 on myeloid cells and increases Treg cell infiltration in tumors70. Similarly, Altan et al. found that PD-L1-negative breast cancer can evade immune surveillance by upregulating B7H4, and the expression level of B7H4 is often negatively correlated with PD-L1 levels71. Such feedback emphasizes that single-agent blockade can be undermined by alternative suppressive pathways. Consequently, combination immunotherapies targeting multiple inhibitory pathways have been pursued. Indeed, combined checkpoint blockade has already proven effective: the anti-LAG-3 antibody Relatlimab plus Nivolumab (anti-PD-1) was FDA-approved for advanced melanoma72. In addition, in patients with colorectal cancer, adding a CTLA-4 inhibitor to PD-1 blockade helps improve long-term efficacy8. In our study, we found that B7H3 palmitoylation inhibited the antitumor activity of CD8⁺ T cells. We synthesized an E-peptide to block B7H3 C496 palmitoylation, which increased intratumoral CD8⁺ T cell infiltration and enhanced their expression of effector molecules such as GZMB and IFN-γ, thereby exerting notable antitumor effects. Anti-PD-1 therapy similarly functions at the effector phase of T cell response by blocking the PD-1/PD-L1 axis and reinvigorating exhausted T cells. Therefore, it is mechanistically reasonable to hypothesize that dual targeting of B7H3 palmitoylation and PD-1 could result in additive—or potentially synergistic—enhancement of CD8⁺ T cell function. Our experimental data support this notion, as we observed significantly improved antitumor efficacy with the combination of E-peptide and anti-PD-1 compared to either monotherapy. Beyond PD-1, other inhibitory receptors, such as CTLA-4, LAG-3, TIM-3, and TIGIT also play critical roles in modulating T cell responses. Prior studies have shown that combining anti-B7H3 with anti-CTLA-4, which primarily reduces Treg-mediated suppression, may be more effective than combinations involving PD-1 blockade in certain tumor contexts70. Mechanistically, this may be explained by the distinct immune regulatory roles of CTLA-4 and PD-1: while anti-PD-1 relieves inhibition on already-activated effector T cells, CTLA-4 blockade preferentially reduces the suppressive activity of Tregs, which constitutively express high levels of CTLA-4 and play a central role in maintaining an immunosuppressive tumor microenvironment. Additionally, tumor-derived factors, such as VEGF can upregulate multiple immune checkpoint molecules (including PD-L1, CTLA-4, TIM-3, and LAG-3), suggesting that compensatory mechanisms among these pathways may limit the efficacy of monotherapies73,74. Thus, combining B7H3 palmitoylation inhibition with agents targeting additional checkpoints, such as LAG-3, TIM-3, or TIGIT may further enhance antitumor immunity. Although we have not yet directly tested combinations of E-peptide with inhibitors of CTLA-4 or other checkpoints, our findings lay a mechanistic foundation for such future explorations. Ongoing and future studies will focus on evaluating such multi-checkpoint strategies in preclinical models to identify synergistic combinations capable of overcoming immune resistance.
In conclusion, our findings suggest that PA-triggered B7H3 palmitoylation may be a key contributor to the immunosuppressive microenvironment in MSS CRC. Specifically, ZDHHC24-induced B7H3 palmitoylation facilitates the ability of malignant cells to evade CD8+ T cell attack. We also showed that the pharmacological inhibition of B7H3 palmitoylation using a membrane-permeating peptide effectively enhances the antitumor effects of CD8+ T cells both in vitro and in vivo (Fig. 9M). We believe that this study provides valuable insights into the mechanisms of immune escape in colorectal cancer. Taken together, B7H3 palmitoylation represents a critical regulatory process that enhances its immunosuppressive effect, and targeting this modification may offer a promising therapeutic strategy to overcome immune resistance in cancers.
Methods
Cell lines and cell culture
The human colon cancer cell lines RKO and SW480, the human embryonic kidney cell line HEK293T, and the murine colon cancer cell lines MC38 and CT26 were purchased from the American Type Culture Collection (ATCC). RKO and HEK293T cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, USA), whereas SW480 cells were cultured in Leibovitz’s L-15 medium (Gibco, USA). MC38 and CT26 cells were cultured in RPMI-1640 medium (Gibco, USA). All culture media were supplemented with 10% fetal bovine serum (Thermo Fisher, USA). RKO, HEK293T, MC38, and CT26 cells were maintained at 37 °C in a humidified incubator with 5% CO2, whereas SW480 cells were maintained at 37 °C in a CO2-free atmosphere. All cell lines were routinely tested for mycoplasma contamination using a mycoplasma PCR detection kit (BMC1040, Abbkine).
Antibodies and chemicals
The following commercially available primary antibodies were used: anti-B7H3 (ab134161, abcam, [WB: 1:200]; ab227679, abcam [IF: 1:100]; 66481-1-Ig, proteintech, [WB: 1:1000; IHC: 1:2500]); anti-ZDHHC24 (ARP55989_P050, aviva systems biology, [WB: 1:1000]; HPA016619, Atlas antibodies, [IHC: 1:50]; and ABIN6743942, antibodies online, [WB: 1:1000]); anti-Flag tag (14793S, cell signaling technology, [WB: 1:1000]); anti-HA tag (3724T, cell signaling technology, [WB: 1:1000]); anti-GAPDH (sc-47724, santa cruz biotechnology, [WB: 1:200] and A19056, ABclonal, [WB: 1:5000]); caveolin-1 (A19006; ABclonal, [WB: 1:5000]); anti-PD-L1 (29122, cell signaling technology, [WB: 1:1000]); horseradish peroxidase (HRP)-streptavidin(A21000, abbkine, [WB: 1:10000]); anti-SQSTM1/p62 (HA721171, HUABIO, [WB: 1:5000; IHC: 1:1000]); anti-CD326/EpCAM (A25344, ABclonal, [IF: 1:200]); anti-ATG5 (ET1611-38, HUABIO, [WB: 1:5000]); anti-Beclin 1 (R381896, zen-bioscience, [WB: 1:1000]). The following secondary anti-bodies were used in Western blot assays: goat anti-rabbit IgG secondary antibody (HRP) (SSA004; Sino Biological Inc), goat anti-mouse IgG secondary antibody (HRP) (SSA007; Sino Biological Inc), HRP-conjugated goat anti-mouse IgG light chain (AS0-62; ABclonal), and HRP-conjugated mouse anti-rabbit IgG light chain (AS061; ABclonal). All Western blot secondary antibodies were used at a dilution of 1:5000. The secondary antibodies used for immunofluorescence (IF) assays were as follows: Cy3, goat anti-rabbit IgG (A22220, Abbkine); Cy3, goat anti-mouse IgG (A22210, Abbkine); D-light 488, goat anti-rabbit IgG (A23220, Abbkine) and DyLight 488, goat anti-mouse IgG (A23210, Abbkine); goat anti-human IgG H&L (FITC) (HY-P83649, MCE); goat anti-human IgG H&L (AF594) (HY-P83648, MCE). All IF secondary antibodies were used at a dilution of 1:200. Antibodies for flow cytometry were listed: Zombie NIR Fixable Viability Kit (423106, BioLegend), BD Pharmingen™ FITC mouse anti-human CD45 (560976, BD), BD OptiBuild™ BV421 mouse anti-human NCAM-1 (CD56) (744217, BD), BD OptiBuild™ BV510 mouse anti-human CD11b (742638, BD), BD OptiBuild™ BV650 mouse anti-human CD14 (740633, BD), PE anti-human CD68 antibody (333808, BioLegend), BD pharmingen™ PerCP-Cy™5.5 mouse anti-human CD3 (552852, BD), BD OptiBuild™ RB780 mouse anti-human CD276 (B7-H3) (755486, BD), BD OptiBuild™ R718 mouse anti-human HLA-DR (752501, BD), BD horizon™ BV605 mouse anti-human CD19 (569363, BD), BD OptiBuild™ BV786 mouse anti-human CD16 (741042, BD), BD pharmingen™ APC mouse anti-human CD326 (EpCAM) (566842, BD), BD OptiBuild™ BV650 Mouse Anti-Human CD8 (743067, BD), APC anti-human/mouse Granzyme B (372204, BioLegend), BD pharmingen™ PE mouse anti-human IFN-γ (554552, BD), brilliant violet (BV)785-conjugated anti-mouse CD45 antibody (103149, BioLegend), BV510-conjugated anti-mouse CD3 antibody (100234, BioLegend), BV785-conjugated anti-mouse CD3 antibody (100232, BioLegend), BV650-conjugated anti-mouse CD8 antibody (100742, BioLegend), PE/cyanine7-conjugated anti-human/mouse Granzyme B recombinant antibody (372214, BioLegend), PerCP/Cyanine5.5-conjugated anti-mouse IFN-γ antibody (505822, BioLegend), FITC-conjugated anti-active Caspase-3(559341, BD), PE hamster anti-mouse γδ T-cell receptor (553178, BD), Alexa Fluor™ 647 rat anti-mouse CD206 (568808, BD), PE-CF594 rat anti-mouse CD49b (562453, BD), BUV661 rat anti-mouse CD93 (741574, BD), BUV737 rat anti-CD11b (612800, BD), BV510 rat anti-mouse F4/80 (743280, BD), PE-Cy7 rat anti-mouse Ly-6G (560601, BD), R718 rat anti-mouse Ly-6C (566987, BD), BUV496 rat anti-mouse CD4 (741050, BD), BUV563 hamster anti-mouse CD3e (741235, BD), BUV805 rat anti-mouse CD45 (748370, BD), BV421 rat anti-mouse Foxp3 (562996, BD), V450 rat anti-mouse CD45R (560472, BD), BV711 rat anti-mouse CD86 (740688, BD), BV650 hamster anti-mouse CD11c (564079, BD), Zombie Yellow™ fixable viability kit (423104, BioLegend), FITC anti-mouse Ly-6G/Ly-6C (Gr-1) antibody (108406, BioLegend), PE/Fire™ 640 anti-mouse I-A/I-E antibody (107664, BioLegend), PE/cyanine7 anti-mouse CD8a antibody (100722, BioLegend). For flow cytometry staining, viability dyes were used at 1:500 (per 2 × 106 cells in 100 μL), and all other fluorochrome-conjugated antibodies were used at 2.5 μL per 1 × 106 cells in 100 μL. The reagents used as follows: phenylmethylsulfonyl fluoride (PMSF) (HY-B0496; MCE), phosphatase inhibitor cocktail I (HY-K0021; MCE), phosphatase inhibitor cocktail II (HY-K0022; MCE), chloroquine (HY-17589A; MCE) 2-bromohexadecanoic acid (HY-111770; MCE), palmostatin B (HY-120911; MCE), N-ethylmaleimide (04260; Millipore Sigma), hydroxylamine solution (467804; Millipore Sigma), 3-MA (HY-19312; MCE), MG-132 (HY-13259; MCE), cycloheximide (HY-12320; MCE), EZ-Link TM BMCC-biotin (21900; Thermo Fisher Scientific), NH4Cl (HY-Y1269; MCE), anti-fade reagent (with DAPI) (HY-K1048; MCE), PMA (HY-18739; MCE), ionomycin (HY-13434A; MCE), monensin (HY-N0150; MCE), brefeldin A(HY-16592; MCE).
Plasmids
23 Flag-tagged ZDHHC family protein-encoding plasmids (n = 23) were obtained from MiaoLingBio, China (catalog numbers: P44768, P44768, P44770, P41465, P43454, P44771, P43506, P43467, P44775, P44767, P44779, P44777, P44769, P44783, P44773, P44780, P44390, P41826, P43407, P44772, P41821, P41823, P44-776). Expression vectors encoding pcDNA3.1-HA-B7H3, pcDNA3.1-Flag-SQSTM1/p62, pcDNA3.1-Flag-OPTN, pcDNA3.1-Flag-NDP52, pcDNA3.1-Flag-TOLLIP, pcDNA3.1-Flag-NIX and pcDNA3.1-Flag-NBR1 plasmids were constructed by insertion of the indicated DNA into the indicated vector. The pcDNA3.1-HA-B7H3-C496A was performed by using Mut Express II Fast Mutagenesis Kit V2 (C21401 Vazyme) according to the manufacturer’s instructions. The sequences are listed in Table S6 (Supplementary Information). All plasmids were confirmed by sequencing. Plasmids were first transformed into competent DH5α Escherichia coli (TSC-C14, Beijing Tsingke Biotech Co., Ltd.), and plasmid DNA for transfection was prepared using a plasmid miniprep kit (D6904, OMEGA) according to the manufacturer’s instructions. Cells cultured in dishes were starved with serum-free medium for 8 h. The indicated plasmids and transfection reagents (HighGene plus, RM09014P, abclonal) were then mixed well and waited for 20 min before being uniformly transferred to the culture dish. After 8 h, the complete medium containing serum was replaced and waited for 48 h.
RNA sequencing
Total RNA was isolated using TRIzol reagent (Invitrogen, CA, USA) according to the manufacturer’s protocol. The RNA’s purity and concentration were assessed via a NanoDrop 2000 spectrophotometer (Thermo Scientific, USA), while RNA integrity was confirmed using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). For transcriptome sequencing, libraries were prepared using the VAHTS Universal V6 RNA-seq Library Prep Kit as per the manufacturer’s guidelines. Sequencing was conducted on an Illumina Novaseq 6000 platform, generating 150 bp paired-end reads. Initial raw reads in FASTQ format underwent quality filtering with fast to yield clean reads. Clean reads were aligned to the reference genome using HISAT2, with gene expression levels quantified in fragments per kilobase million (FPKM) and gene read counts obtained through HTSeq-count. Differential expression analysis was performed using DESeq2, identifying differentially expressed genes (DEGs) with a threshold of p < 0.05 and fold change ≥ 2; genes with baseman < 10 were excluded. Functional enrichment analysis of DEGs was performed using a hypergeometric test to identify significantly enriched Gene Ontology (GO) terms.
Non-targeted metabolomics analysis
Non-targeted LC–MS-based metabolomics was performed on 18 colorectal tumor samples (MSS, n = 8; MSI-H, n = 10), each representing an independent biological replicate. In addition, pooled quality-control (QC) samples were prepared by combining equal aliquots of all extracts and were injected at regular intervals to monitor instrument stability by principal component analysis (PCA). All tumor samples used for non-targeted metabolomic analysis were collected under strictly standardized conditions to ensure experimental consistency. Tissues were rapidly excised, snap-frozen in liquid nitrogen, and stored at –80 °C until analysis. To minimize batch-related variability, all samples were processed simultaneously and analyzed within a single LC–MS/MS run. This unified workflow effectively eliminated the need for additional batch effect correction in downstream data analysis. Weigh 30 mg of sample into a 1.5 mL EP tube, followed by the addition of two small steel beads and 400 μL of methanol-water (4:1, v/v) containing an internal standard (4 μg/mL). The sample was pre-cooled at −40 °C for 2 min, then ground at 60 Hz for 2 min. After grinding, the sample underwent ultrasonic extraction in an ice-water bath for 10 min and was then left to stand at −40 °C for 2 h. The mixture was centrifuged for 20 min at 15,000 × g and 4 °C. Subsequently, 150 μL of the supernatant was drawn with a syringe and transferred to an LC injection vial. Liquid chromatography–mass spectrometry (LC–MS) analysis was conducted using a Waters ACQUITY UPLC I-Class plus/Thermo QE HF ultra-high-performance liquid chromatography (UHPLC) tandem high-resolution mass spectrometer (HRMS) at OE Biotech, Inc., Shanghai, China. The system was equipped with a heated electrospray ionization (ESI) source (Thermo Fisher Scientific, Waltham, MA, USA) to analyze metabolic profiles in both positive and negative ESI modes. An ACQUITY UPLC HSS T3 column (1.8 μm, 2.1 × 100 mm) was used for both ionization modes; the column temperature was maintained at 45 °C, and the flow rate was 0.35 mL/min. Gradient elution was performed with solvent A (water with 0.1% formic acid, v/v) and solvent B (acetonitrile with 0.1% formic acid, v/v) using the following program: 0–2 min, 95% A/5% B; 2–4 min, linear to 70% A/30% B; 4–8 min, linear to 50% A/50% B; 8–10 min, linear to 20% A/80% B; 10–14 min, 0% A/100% B; 14–15 min, 0% A/100% B; 15.1–16 min, re-equilibration at 95% A/5% B. Samples were kept at 10 °C during analysis, with an injection volume of 4 μL. Detection was performed over a mass range of 70–1050 m/z, using a primary scan resolution of 60,000 and a secondary scan resolution of 15,000. Collision energies of 10, 20, and 40 eV were applied. The mass spectrometer’s operating conditions were as follows: spray voltage of 3800 V (positive mode) and 3200 V (negative mode); sheath gas flow rate of 35 units; auxiliary gas flow of 8 units; capillary temperature of 320 °C; auxiliary gas heater temperature of 350 °C; and S-lens RF level of 50. Data preprocessing was performed using Progenesis QI v3.0 (Nonlinear Dynamics, Newcastle, UK) for baseline correction, peak identification, integration, retention time alignment, and intensity normalization across samples. Following missing value imputation and filtering, the data were further normalized via log-transformation to approximate a Gaussian distribution and reduce heteroscedasticity, ensuring suitability for downstream statistical analysis. Compound identification was based on accurate mass-to-charge ratios (m/z), secondary fragments, and isotopic distributions, utilizing databases such as the Human Metabolome Database (HMDB), Lipidmaps (V2.3), Metlin, and self-built databases. Peaks with over 50% missing values (ionic intensity = 0) were excluded, and zero values were replaced by half the minimum value. Data from positive and negative ion modes were merged into a single data matrix for further analysis. Differential metabolites between MSS and MSI-H groups were selected using Student’s t test with P < 0.05 and fold change (FC) ≥ 1.2 or ≤ 1/1.2.
Construction of stable cell lines
The lentiviral plasmids pLVX-ZDHHC24 or pLVX-Zdhhc24 were constructed by cloning ZDHHC24 or Zdhhc24 genes into the pLVX vector. We used the pLVX, pMD2.G, and psPAX2 three-plasmid packaging systems to generate the highly expressed viruses. Short hairpin RNA (shRNA) targeting the human ZDHHC24 or murine Zdhhc24 genes was expressed in the pLKO.1 vector to obtain knockdown (KD) clones. The pLKO.1-shRNAs, pMD2.G, and psPAX2 triple packaging systems were then used to construct silencing expression lentiviruses. The sequences are listed in Table S7 (Supplementary Information). Similarly, the three of them were mixed in 500 μL of serum-free medium and 40 μl of Lipofectamine 2000 in a mass ratio of 2:1:2. The plasmid mixture was then transfected into HEK293T cells. Harvested viral supernatant was filtered through a sterilized polyethersulfone PES syringe filter (SFPES025022SI, membrane solutions LLC) and used to infect the indicated cells. Stabilize cell lines by screening with 1 μg/ml puromycin for 48 h or 0.8 mg/mL G418 for 20 days. The CRISPR/Cas9 system was used to generate B7H3, SQSTM1, ATG5, and BECN1 KO cells. The single guide RNAs were designed from a public website, CHOPCHOP, and then cloned into the PX459 plasmid (62988, Addgene) or the PX462 plasmid (62987, Addgene). The sgRNA sequences are listed in Table S7 (Supplementary Information).
Cellular membrane and cytoplasmic protein isolation
Separation of cytosolic and cytoplasmic proteins of the indicated cells was performed using the ExKine™ membrane and cytoplasmic protein extraction kit (KTP3005; Abbkine).
Cell counting kit-8 assay
To assess peptide cytotoxicity in vitro, cells were seeded into 96-well plates (100 μL complete medium) and allowed to adhere overnight. Cells were then treated with E-peptide or C-peptide at the indicated concentrations (2.5, 5, 10, 25, 50, 100, and 200 μM) for 24 h. Subsequently, 10 μL of cell counting kit-8 (C6005, new cell and molecular biotech) reagent was added to each well and incubated at 37 °C for 1–4 h protected from light. Absorbance was measured at 450 nm using a microplate reader. Wells containing medium plus reagent without cells served as blanks, and cell viability was normalized to the vehicle-treated control (set to 100%).
Mouse serum biochemistry
Peripheral blood was collected from mice and allowed to clot at room temperature for 30 min. Serum was separated by centrifugation at 4 °C at 3000 × g for 10 min, and analyzed immediately or stored at −80 °C until use. Serum creatinine (CRE) was measured using the creatinine (CRE) content assay kit (AKNM008M, Beijing Boxbio Science and Technology Co., Ltd). The microplate reader was pre-warmed for ≥30 min and set to 546 nm. Reactions were carried out at 37 °C according to the manufacturer’s protocol, absorbance was recorded, and CRE concentration was calculated against the kit standard/blank using the provided formula. Serum alanine aminotransferase (ALT/GPT) and aspartate aminotransferase (AST/GOT) activities were determined using the GPT activity assay kit (AKAM006M, Beijing Boxbio Science and Technology Co., Ltd) and the GOT activity assay kit (AKAM019M, Beijing Boxbio Science and Technology Co., Ltd), respectively. The instrument was pre-warmed for ≥30 min and set to 505 nm. A matched control and a standard curve were included as instructed. After incubation and color development at 37 °C, absorbance was measured, and enzyme activities were calculated and reported as U/L according to the kit-provided formula.
Immunoprecipitation
Different magnetic beads (HAK21042, HAK21011, HUABIO) were selected according to the experimental needs. If the magnetic beads (HY-K0202, MCE) did not have a coupled antibody, it was necessary to mix the beads with the corresponding primary antibody for half an hour at room temperature or 2 h at 4 °C before harvesting the cells. Subsequently, start lysing the cells to harvest the proteins. Proteins are extracted in NP40 cell lysis buffer containing the protease cocktail inhibitor for 30 min. Next, the magnetic beads that have been coupled to the antibody are transferred to the supernatant into the cell lysate and rotated at 4 °C overnight. The following day, the supernatant was discarded, and the magnetic beads were washed five times with 500 μL of NP40 cell lysis buffer. Between each wash, magnetic separation gently. Finally, the supernatant was discarded, and 40 μl SDS loaded buffer was mixed with magnetic beads and boiled in a water bath for 10 min before being used for Western blotting.
Protein extraction and Western blotting
For cultured cells, discard the medium and wash the cells three times with cold PBS. Cells were lysed for 30 min using NP-40 buffer containing phenylmethylsulfonyl fluoride (HY-B0496, MCE) and cocktail protease inhibitor (HY-K0010, MCE). For tissue samples, total protein was extracted using the Total Protein Extraction Kit (EX1100, Solarbio) according to the manufacturer’s instructions. Lysates were centrifuged at 15,000 × g for 15 min at 4 °C. Supernatants were collected, and protein concentrations were quantified by BCA kit (23227; Thermo Fisher). Subsequently, the protein supernatants acquired were mixed with 5× concentrated sample buffer containing 250 mM Tris-HCl (pH 6.8), 5% β-mercaptoethanol, 10% sodium dodecyl sulfate (SDS), 30% glycerol, and bromophenol blue, followed by heating at 95–100 °C for 10 min. Protein samples were separated by electrophoresis in 8–10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). After electrophoresis, the separated protein bands were transferred to a polyvinylidene fluoride (PVDF) membrane. Subsequently, the membrane was then blocked by immersing in 5% skimmed milk in TBST or rapid blocking buffer (SW162-01, Seven, Beijing, China). Next, the membranes were incubated overnight at 4 °C with primary antibodies and then exposed to HRP-linked secondary antibodies for 2 h at ambient temperature. Finally, the membrane bands were washed three times with TBST and then visualized using an efficient chemiluminescence kit (34096; Thermo Fisher).
Acyl-biotin exchange (ABE) assay
The ABE assay was performed based on established protocols33,39,46, and the key experimental steps are detailed below. 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM MgCl2, 1% NP-40, 10% glycerol, protease inhibitor, and phosphatase inhibitor were added to NP-40 to configure the lysis buffer. Fresh N-ethylmaleimide (NEM) solution was prepared and added to the lysis buffer to a final concentration of 50 mM. The indicated cells were lysed in a buffer containing 50 mM NEM for 1 h on ice. Collect the cell suspension and mix it with magnetic beads precoated with the appropriate antibody for 3 h at 4°C. After 5 washes with lysis buffer (pH = 7.5), the beads were washed again 3 times with lysis buffer (pH = 7.2). Fresh hydroxylamine (HAM) solution was prepared in lysis buffer at pH 7.2 to a final concentration of 1 M. The beads were then incubated with lysis buffer containing 1 M HAM for 1 h at room temperature. Wash 3 times with Lysis Buffer (pH = 6.2) to remove residual HAM. Subsequently, the beads were incubated with lysis buffer (pH = 6.2) containing Biotin-BMCC (5 μM) at 4 °C for 1 h. The beads were washed twice with buffer (pH = 6.2) and then three times with lysis buffer (pH = 7.5). Excess buffer was removed and added to 50 μL sample buffer (5% SDS, 5% glycerol, 125 mM Tris-HCL (pH = 6.8), 0.01% bromophenol blue) containing 5 mM DL-Dithiothreitol (DTT). Samples were boiled at 80 °C for 10 min and then cooled to room temperature for 15 min.
Immunofluorescence assays
Cells were inoculated onto slides of 24-well culture plates, allowed to adhere to the wall, and then subjected to various treatments. The slides were rinsed three times with precooled PBS, followed by fixation in 4% paraformaldehyde for 20 min at room temperature. After removing the paraformaldehyde, the slides were washed three times with PBS. Added 0.3% Triton-X-100 to the wells and permeabilized for 15 min at room temperature. The slides were blocked with 3% BSA for 1 h at room temperature after washing three times with PBS. Subsequently, they were incubated with primary antibodies at 4 °C overnight. The following day, fluorochrome-labeled secondary antibodies were applied in the dark for 2 h at room temperature, and the samples were then stained with DAPI for 10 min. Tissue samples were treated similarly as described above. Finally, we obtained representative images using a Japanese Olympus FLUOVIEW FV1000 confocal scanning microscope or a Japanese Nikon ECLIPSE Ti2 fluorescence microscope.
Tumor-infiltrating lymphocyte isolation and flow cytometry analysis
For tumor-infiltrating immune cells, all mice were isolated immediately after sacrifice at the indicated times, and tumors were placed on ice and carefully cut into small pieces of approximately 1 mm. These fragmented tumor tissues were then digested in serum-free 1640 medium containing collagenase IV (1 mg/ml, 17104019; Invitrogen), deoxyribonuclease I (100 μg/ml, HY-108882; MCE), and hyaluronidase (250 μg/ml, BS171; Biosharp) for 90 min at 37 °C. The cell suspension was then filtered using a cell strainer. A Percoll gradient cell separation solution (17089101, Cytiva) was used to isolate tumor cells and immune cells infiltrating the tumor. For cytokine analysis, the obtained immune cells were incubated with PMA, Brefeldin A, monensin, and ionomycin for 6 h and then stained with the corresponding antibodies. Cytofix/Cytoperm™ fixation and permeabilization solution (554722, BD) was used for intracellular staining. Transcription factor buffer set (562574, BD) used for nuclear staining. Raw FCS files were analyzed using FlowJo software (v10.8.1).
T-cell killing assay
The experimental methodology was adapted from published protocols75,76, and the main procedures are described below. Peripheral blood mononuclear cells (PBMCs) were isolated from anticoagulated whole blood using human lymphocyte separation medium (7111011, Dakewe). Briefly, whole blood was diluted 1:1 with PBS and carefully layered onto the separation medium. Samples were centrifuged at room temperature at 800 × g for 30 min using a horizontal rotor with low acceleration and deceleration settings. After centrifugation, the opaque mononuclear cell layer at the plasma-medium interface was carefully collected into a new tube. Cells were washed 2 times with RPMI-1640 by centrifugation at 20 °C and 250 × g for 10 min, and the cell pellet was resuspended in normal saline or appropriate culture medium for downstream isolation. Human CD8+ T cells and human DCs were isolated from PBMCs using the CD8+ T cell isolation kit, human (130-096-495, Miltenyi Biotec), and the Dynabeads® Human DC Enrichment Kit (11308D, Thermo Fisher Scientific), respectively. The isolated DCs were first co-incubated with the indicated tumor cells, and then the pretreated DCs were then used to activate CD8+ T cells. To evaluate the tumor-killing effect mediated by T cells, the preconditioned CD8+ T cells were reactivated with anti-CD3 antibody (2 μg/ml), anti-CD28 antibody (2 μg/ml), and IL-2 (100 IU/ml). The activated CD8+ T cells were then co-incubated with the indicated tumor cells at a ratio of 10:1 for 24–48 h. Detection of T-cell mediated cytotoxicity using caspase-3 cleavage assay and lactate dehydrogenase (LDH) release assay. After centrifugation, the supernatant of each well was collected, and the amount of LDH released in the culture medium was determined using the LDH Cytotoxicity Assay Kit (HY-K1090, MCE). Cell clusters obtained after centrifugation were washed three times with PBS and then stained with FITC-coupled activated caspase-3 antibody, followed by flow cytometric analysis.
Immunohistochemistry (IHC)
The embedded tissue specimens were cut into sections at a thickness of 4 μm. Following sequential processes of dewaxing, rehydration, antigen retrieval, and blocking, the sections were treated with the specified primary antibody overnight and then incubated with the secondary antibody for 1 h at room temperature. Immunohistochemical results were quantified by the density and intensity of positive cells. The density of positive staining was categorized into 4 grades based on the intensity of positive staining as follows: 1 (<5%), 2 (5–30%), 3 (30–70%), and 4 (>70%). The intensity of positive staining was categorized into 4 grades, 0 (unstained), 1 (weak), 2 (moderate), and 3 (strong). The density and intensity scores were then multiplied to obtain the IHC score, which ranged from 0 to 12. To evaluate the CD8 staining score, three random fields of view were chosen from each tissue section and analyzed at a magnification of ×400. The mean number of positively stained cells observed in these fields was calculated to represent each case. IHC staining was independently scored by two experienced pathologists, and the final IHC score for each case was defined as the average of the two scores.
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
RNA was initially extracted from cells using AG RNAex Pro RNA extraction reagent (AG21101, Accurate Biotechnology (Hunan) Co., Ltd) following the manufacturer’s protocol. The extracted RNA was then reverse-transcribed into cDNA using the Reverse Transcription Kit (HiScript III RT SuperMix for qPCR, Vazyme). Finally, qPCR was performed using the SupRealQ Ultra Hunter SYBR qPCR Master Mix(U+) (Q713-02, Vazyme Biotech Co., Ltd) reagent on QuantStudio 3 instrument (Thermo Fisher). All primer sequences are provided in Table S8.
Animal study
The mice used in this study (Mus musculus) included C57BL/6J, BALB/c, NCG, Zdhhc24 fl/fl, Villin-cre, and Rosa26-CAG-LSL-tdTomato, which were purchased from GemPharmatech (Nanjing, China). The Zdhhc24 fl/fl mouse line was generated using CRISPR/Cas9 on a C57BL/6J genetic background (gRNA sequences are provided in Table S9) and maintained in our facility. The huHSC-C-NKG-ProF mice were generated by Cyagen Biosciences (Suzhou, China). All animals used in this study were female. Unless otherwise indicated in the figure legends, mice were 6 weeks old at the start of each experiment, and the number of animals used for each experiment (n) and the corresponding strain/substrain are reported in the figure legends. Sex was not included as a biological variable because all experiments were performed using female mice to reduce variability and maintain consistency across cohorts; therefore, no sex-based comparisons were performed, and this choice was made solely for experimental design and reproducibility considerations and does not reflect any sex bias or discrimination. All animal experiments were conducted with approval from the Animal Care and Use Committee of Tongji Hospital (TJH-202303006). Mice were housed under specific pathogen-free conditions with a 12 h light/12 h dark cycle, an ambient temperature of 22–24 °C, and relative humidity of 40–60%, with food and water available ad libitum.
For xenograft tumor assays, cells were digested, washed three times with PBS, and mixed with Matrigel (abs9493, Absin) at a 1:1 ratio to a final density of 2 × 106 cells/mL. A total of 100 μL of cell suspension was injected subcutaneously into 6-week-old female mice. Tumor volume was measured at the indicated time points and calculated as V = L × D2/2 (L: long axis, D: short axis). In the approved protocol, the maximal tumor burden was a tumor volume of 2000 mm3, and mice were euthanized when tumors reached this limit or when other humane endpoints were met. In addition to the predefined maximal tumor burden (tumor volume of 2000 mm3), mice were euthanized if any of the following humane endpoint criteria were met: (i) ulceration, necrosis, or bleeding at the tumor site; (ii) impaired mobility, inability to access food or water, or persistent abnormal posture/behavior; (iii) rapid weight loss (≥20% of baseline) or severe dehydration; (iv) signs of systemic distress such as labored breathing, hypothermia, or marked lethargy; or (v) any other condition judged by the animal care staff/veterinarian to require euthanasia to prevent undue suffering. Mice were monitored at least every 1–2 days, and more frequently when tumors approached the endpoint. Overall, the study did not substantially exceed the permitted limit. In the CT26 survival experiment (Fig. 9C–G), a small number of mice showed calculated tumor volumes slightly above 2000 mm3 at the scheduled measurement time points, reflecting caliper measurement variability and the estimated nature of the volume formula, together with rapid tumor growth between two consecutive monitoring time points; mice were euthanized immediately upon detection in accordance with the approved protocol. For therapeutic agents, anti-PD-1 antibody (10 mg/kg), E-peptide, and C-peptide (10 mg/kg) were injected into mice by intraperitoneal injection at the indicated time intervals. For Zdhhc24fl/fl mice, CRISPR/Cas9 technology was used to modify the Zdhhc24 gene (gRNA sequences are provided in Table S9). Briefly, the CRISPR/Cas9 system and donor were microinjected into fertilized eggs of C57BL/6 J mice. Transplantation of fertilized eggs yielded positive F0 mice, which were confirmed by PCR and sequencing. Positive F0 generation mice were backcrossed with C57BL/6 J for at least two generations to obtain a stable Zdhhc24fl/fl mouse model. Villin-cre and Rosa26-CAG-LSL-tdTomato are well-established commercial mice. Validation of PCR primers for Zdhhc24fl/fl and Villin-cre mice is provided in Table S10.
Hemolysis assay
Approximately 0.6 mL of blood was collected from anesthetized mice via cardiac puncture into a centrifuge tube pre-coated with sodium citrate as an anticoagulant. The sample was centrifuged at 2500 × g for 10 min to remove the plasma. The remaining red blood cells (RBCs) were washed repeatedly with normal saline until the supernatant became colorless. After discarding the supernatant, the RBCs were resuspended in 1 mL of normal saline to obtain an RBC suspension. Then, 100 μL of RBC suspension was added to 1 mL of E-peptide solution (dissolved in saline) at various concentrations. Saline alone was used as a negative control, and distilled water was used as a positive control. The mixtures were incubated at 37 °C for 2 h, followed by centrifugation at 2500 × g for 10 min. The optical density (OD) of the supernatant was measured at 541 nm. The hemolysis rate was calculated using the following formula:
Coagulation assays: prothrombin time (PT) and activated partial thromboplastin time (APTT)
PT and activated partial thromboplastin time APTT were measured using commercial assay kits (Bioswamp Life Science Lab, Wuhan, China) according to the manufacturer’s instructions. Briefly, whole blood was centrifuged at 2500 × g for 10 min to obtain platelet-poor plasma. Two hundred microliters of plasma was mixed with 10 μL of E-peptide solution (dissolved in normal saline) to achieve final peptide concentrations of 5, 10, 25, and 50 μM. An equal volume of normal saline was added to the plasma as the control. For PT testing, 70 μL of peptide-treated or control plasma and 140 μL of PT reagent were pre-incubated separately in a 37 °C water bath for 3 min. The two components were then mixed rapidly, and the clotting time was recorded upon cessation of flow during gentle tilting of the reaction tube. For APTT testing, 100 μL of peptide-treated or control plasma was mixed with 100 μL of APTT reagent (containing ellagic acid) and incubated at 37 °C for 5 min. Subsequently, 100 μL of 25 mM CaCl2 solution (pre-warmed to 37 °C) was added to initiate coagulation. The tube was gently agitated in a 37 °C water bath, and the time to visible fibrin formation was recorded as the APTT value.
Peptide synthesis
The C-peptide (YGRKKRRQRRRRKWEISQCKC) and E-peptide (YGRKKRRQ-RRRCWRKIKQSCE) were synthesized by QYAOBlO (ChinaPeptides Co., Ltd.) using standard Fmoc-based solid-phase peptide synthesis (SPPS) protocols on 2-chlorotrityl chloride resin. The synthesis followed a stepwise procedure involving resin swelling in DMF, sequential coupling of Fmoc-protected amino acids using HBTU and DIEA, and Fmoc deprotection with 20% piperidine in DMF. Coupling efficiency was monitored by Kaiser tests. Upon completion of chain assembly, the peptide was cleaved from the resin using a TFA-based cleavage cocktail (TFA/H₂O/EDT/TIS, 95:2:2:1, v/v) and precipitated in cold ether. The crude product was purified by preparative reversed-phase high-performance liquid chromatography (RP-HPLC) on a C18 column. Final peptide identity and purity were confirmed by mass spectrometry (MS) and analytical HPLC. The purified peptide was lyophilized and stored at −20 °C until use.
Proximity ligation assay (PLA)
Proximity ligation assay was detected using the Duolink® in situ red starter kit mouse/rabbit (Sigma-Aldrich) following the manufacturer’s protocol. Briefly, fixed and permeabilized samples on glass slides were blocked with Duolink® Blocking Solution at 37 °C for 60 min, then incubated overnight at 4 °C with primary antibodies: anti-CD276 (Abcam, ab134161) and anti-SQSTM1/p62 (CST, 88588). After washing in 1× Wash Buffer A, PLUS and MINUS PLA probes (1:5 dilution) were applied for 1 h at 37 °C, followed by ligation (1:40 ligase, 30 min at 37 °C) and amplification (1:80 polymerase, 100 min at 37 °C, in the dark). Slides were washed, mounted with Duolink® in situ mounting medium containing DAPI, and imaged using a confocal microscope (LSM900 Airyscan).
Surface plasmon resonance (SPR)
Binding measurements between ZDHHC24 and E-peptide were performed by surface plasmon resonance (SPR) on a Biacore 8 K instrument (GE Healthcare/Cytiva) using CM5 sensor chips, based on established protocols77,78 with all key experimental parameters provided below. In brief, experiments were conducted at 25 °C with a blank activated/deactivated reference flow cell. ZDHHC24 was immobilized on CM5 via amine coupling using 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide/N-hydroxysuccinimide (EDC/NHS), and residual esters were blocked with ethanolamine. The running/immobilization buffer was 1× phosphate-buffered saline with surfactant P20 (PBS-P+, pH 7.4). Interaction analyses were carried out in PBS-P+ (pH 7.4) supplemented with 5% (v/v) dimethyl sulfoxide (DMSO); an eight-point solvent-correction curve bracketing 5.0% DMSO (prepared by mixing 4.5% and 5.8% DMSO stocks) was used for bulk-refractive index correction. E-peptides were injected as analytes from low to high concentration (0.3125–10 µM) at a flow rate of 30 µL min−1 with a 150 s association phase, followed by surface regeneration with 10 mM glycine–HCl (pH 2.0) for 5 min between injections. Binding parameters were obtained by global fitting of double-referenced, solvent-corrected sensorgrams to a 1:1 Langmuir model in Biacore Insight (Cytiva), yielding the equilibrium dissociation constant (Kd).
Statistics and reproducibility
All assays were performed in at least three independent experiments. Sample sizes were determined based on prior experience or published literature. No statistical method was used to predetermine sample size. No data were excluded from the analyses. Human specimens diagnosed with colorectal cancer were randomly selected from eligible hospital patients. Animals were randomly assigned to control and experimental groups using a simple randomization method. Investigators were blinded to group allocation during data collection and analysis, including outcome assessment. Data are presented as mean ± SD. Statistical analyses were performed using GraphPad Prism (version 9.5) and Origin (version 2021). Before statistical analysis, tests for normality and homogeneity of variance were performed. Two-tailed unpaired or paired Student’s t tests were used for comparisons between two groups. For multiple-group comparisons, one-way ANOVA with Tukey’s or Dunnett’s multiple-comparisons tests, or two-way ANOVA with Tukey’s multiple-comparisons test were used. Categorical variables were analyzed using chi-square tests. Correlations were assessed using Pearson’s correlation (two-tailed). Survival analyses were performed using the Kaplan–Meier method and compared by the log-rank test. Statistical significance was set at p < 0.05.
Ethics statement
The clinical specimens were obtained from the Department of Gastrointestinal Surgery, Tongji Hospital. All patients were informed about the study and provided written informed consent prior to surgery. Consent was obtained for the acquisition, reporting, and sharing of patient clinical information at the individual level. No participant compensation was provided. Patients’ microsatellite status (MSS or MSI-H) and clinical characteristics were obtained from their medical records. Patient information, including age and sex, is shown in Tables S1–S5 (Supplementary Information). Tissue microarrays of colorectal tumors and paracancers were purchased from the Shanghai Outdo Biotech Co. Ltd (HColA180Su21). For these commercial tissue microarrays, demographic information was provided by the vendor and is reported in the Source Data. Sex- and/or gender-stratified analyses were not performed because the study was not designed to detect sex/gender differences, and the human analyses primarily focused on tumor molecular subtype (MSS vs MSI-H). The study was approved by the Ethics Committee of Tongji Hospital (TJ-IRB20220723). All animal experiments were conducted with approval from the Animal Care and Use Committee of Tongji Hospital (TJH-202303006).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The analysis of CD8+ T cell infiltration levels based on CD276 (B7H3) mRNA levels was performed using data from the TIMER database (https://cistrome.shinyapps.io/timer/). Kaplan–Meier analysis of overall survival (OS) and relapse-free survival (RFS) based on B7H3 and ZDHHC24 mRNA levels was conducted using the Kaplan–Meier Plotter database (https://kmplot.com/analysis/). The gene expression profile across all tumor samples and paired normal tissues based on ZDHHC24 mRNA levels was obtained from the Gene Expression Profiling Interactive Analysis (GEPIA) database (http://gepia.cancer-pku.cn/index.html). RNA-sequencing data for MSS and MSI-H tumors have been deposited in the Sequence Read Archive (SRA) under BioProject accession PRJNA1233603. Non-targeted metabolomics sequencing data for MSS and MSI-H tumors have been deposited in the MetaboLights database under accession MTBLS12300. All data in this study are provided in the supplementary table and source data. Source data are provided with this paper.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Dekker, E., Tanis, P. J., Vleugels, J. L. A., Kasi, P. M. & Wallace, M. B. Colorectal cancer. Lancet 394, 1467–1480 (2019).
Sharma, P. et al. Immune checkpoint therapy-current perspectives and future directions. Cell 186, 1652–1669 (2023).
Topalian, S. L., Taube, J. M., Anders, R. A. & Pardoll, D. M. Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat. Rev. Cancer 16, 275–287 (2016).
Sun, Q. et al. Immune checkpoint therapy for solid tumours: clinical dilemmas and future trends. Signal Transduct. Target. Ther. 8, 320 (2023).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Overman, M. J. et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): an open-label, multicentre, phase 2 study. Lancet Oncol. 18, 1182–1191 (2017).
André, T. et al. Nivolumab plus low-dose ipilimumab in previously treated patients with microsatellite instability-high/mismatch repair-deficient metastatic colorectal cancer: 4-year follow-up from CheckMate 142. Ann. Oncol. J. Eur. Soc. Med. Oncol. 33, 1052–1060 (2022).
Zhao, P., Li, L., Jiang, X. & Li, Q. Mismatch repair deficiency/microsatellite instability-high as a predictor for anti-PD-1/PD-L1 immunotherapy efficacy. J. Hematol. Oncol. 12, 54 (2019).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Schildberg, F. A., Klein, S. R., Freeman, G. J. & Sharpe, A. H. Coinhibitory pathways in the B7-CD28 ligand-receptor family. Immunity 44, 955–972 (2016).
Ceeraz, S., Nowak, E. C. & Noelle, R. J. B7 family checkpoint regulators in immune regulation and disease. Trends Immunol. 34, 556–563 (2013).
Burke, K. P., Chaudhri, A., Freeman, G. J. & Sharpe, A. H. The B7:CD28 family and friends: unraveling coinhibitory interactions. Immunity 57, 223–244 (2024).
Lee, Y. H. et al. Inhibition of the B7-H3 immune checkpoint limits tumor growth by enhancing cytotoxic lymphocyte function. Cell Res. 27, 1034–1045 (2017).
Peuker, K. et al. Microbiota-dependent activation of the myeloid calcineurin-NFAT pathway inhibits B7H3- and B7H4-dependent anti-tumor immunity in colorectal cancer. Immunity 55, 701–717.e7 (2022).
Tyagi, A. et al. Evidence supporting a role for the immune checkpoint protein B7-H3 in NK cell-mediated cytotoxicity against AML. Blood 139, 2782–2796 (2022).
Wang, C. et al. CD276 expression enables squamous cell carcinoma stem cells to evade immune surveillance. Cell Stem Cell 28, 1597–1613 (2021).
Ingebrigtsen, V. A. et al. B7-H3 expression in colorectal cancer: nuclear localization strongly predicts poor outcome in colon cancer. Int. J. Cancer 131, 2528–2536 (2012).
Zang, X. et al. Tumor-associated endothelial expression of B7-H3 predicts survival in ovarian carcinomas. Mod. Pathol. 23, 1104–1112 (2010).
Zang, X. et al. B7-H3 and B7x are highly expressed in human prostate cancer and associated with disease spread and poor outcome. Proc. Natl. Acad. Sci. USA 104, 19458–19463 (2007).
Ingebrigtsen, V. A. et al. B7-H3 expression in colorectal cancer: associations with clinicopathological parameters and patient outcome. BMC Cancer 14, 602 (2014).
Picarda, E., Ohaegbulam, K. C. & Zang, X. Molecular pathways: targeting B7-H3 (CD276) for human cancer immunotherapy. Clin. Cancer Res. J. Am. Assoc. Cancer Res. 22, 3425–3431 (2016).
Getu, A. A. et al. New frontiers in immune checkpoint B7-H3 (CD276) research and drug development. Mol. Cancer 22, 43 (2023).
Huang, Y. et al. FUT8-mediated aberrant N-glycosylation of B7H3 suppresses the immune response in triple-negative breast cancer. Nat. Commun. 12, 2672 (2021).
Chen, R. et al. Glycoproteomics analysis of human liver tissue by combination of multiple enzyme digestion and hydrazide chemistry. J. Proteome Res. 8, 651–661 (2009).
Jiang, H. et al. Protein lipidation: occurrence, mechanisms, biological functions, and enabling technologies. Chem. Rev. 118, 919–988 (2018).
Chamberlain, L. H. & Shipston, M. J. The physiology of protein S-acylation. Physiol. Rev. 95, 341–376 (2015).
Mesquita, F. S. et al. Mechanisms and functions of protein S-acylation. Nat. Rev. Mol. Cell Biol. 25, 488–509 (2024).
Resh, M. D. Covalent lipid modifications of proteins. Curr. Biol. 23, R431–R435 (2013).
Rocks, O. et al. The palmitoylation machinery is a spatially organizing system for peripheral membrane proteins. Cell 141, 458–471 (2010).
Linder, M. E. & Deschenes, R. J. Palmitoylation: policing protein stability and traffic. Nat. Rev. Mol. Cell Biol. 8, 74–84 (2007).
Zhang, M. et al. A STAT3 palmitoylation cycle promotes TH17 differentiation and colitis. Nature 586, 434–439 (2020).
Hao, J. W. et al. CD36 facilitates fatty acid uptake by dynamic palmitoylation-regulated endocytosis. Nat. Commun. 11, 4765 (2020).
Huang, X. et al. S-acylation of p62 promotes p62 droplet recruitment into autophagosomes in mammalian autophagy. Mol. Cell 83, 3485–3501 (2023).
Rana, M. S. et al. Fatty acyl recognition and transfer by an integral membrane S-acyltransferase. Science 359, eaao6326 (2018).
Elliot Murphy, R. & Banerjee, A. In vitro reconstitution of substrate S-acylation by the zDHHC family of protein acyltransferases. Open Biol. 12, 210390 (2022).
Yao, H. et al. Inhibiting PD-L1 palmitoylation enhances T-cell immune responses against tumours. Nat. Biomed. Eng. 3, 306–317 (2019).
Lu, Y. et al. Palmitoylation of NOD1 and NOD2 is required for bacterial sensing. Science 366, 460–467 (2019).
Chen, S. et al. Palmitoylation-dependent activation of MC1R prevents melanomagenesis. Nature 549, 399–403 (2017).
Bu, L. et al. High-fat diet promotes liver tumorigenesis via palmitoylation and activation of AKT. Gut 73, 1156–1168 (2024).
Bu, L. et al. Targeting APT2 improves MAVS palmitoylation and antiviral innate immunity. Mol. Cell 84, 3513–3529 (2024).
Tall, A. R. & Yvan-Charvet, L. Cholesterol, inflammation and innate immunity. Nat. Rev. Immunol. 15, 104–116 (2015).
Pascual, G. et al. Dietary palmitic acid promotes a prometastatic memory via Schwann cells. Nature 599, 485–490 (2021).
Zheng, S. et al. ZDHHC5-mediated NLRP3 palmitoylation promotes NLRP3-NEK7 interaction and inflammasome activation. Mol. Cell 83, 4570–4585 (2023).
Brigidi, G. S. & Bamji, S. X. Detection of protein palmitoylation in cultured hippocampal neurons by immunoprecipitation and acyl-biotin exchange (ABE). J Vis. Exp. 50031, https://doi.org/10.3791/50031 (2013).
Pohl, C. & Dikic, I. Cellular quality control by the ubiquitin-proteasome system and autophagy. Science 366, 818–822 (2019).
Cullen, P. J. & Steinberg, F. To degrade or not to degrade: mechanisms and significance of endocytic recycling. Nat. Rev. Mol. Cell Biol. 19, 679–696 (2018).
Gatica, D., Lahiri, V. & Klionsky, D. J. Cargo recognition and degradation by selective autophagy. Nat. Cell Biol. 20, 233–242 (2018).
Vargas, J. N. S., Hamasaki, M., Kawabata, T., Youle, R. J. & Yoshimori, T. The mechanisms and roles of selective autophagy in mammals. Nat. Rev. Mol. Cell Biol. 24, 167–185 (2023).
Ning, W. et al. GPS-palm: a deep learning-based graphic presentation system for the prediction of S-palmitoylation sites in proteins. Brief. Bioinform. 22, 1836–1847 (2021).
Xie, Y. et al. GPS-lipid: a robust tool for the prediction of multiple lipid modification sites. Sci. Rep. 6, 28249 (2016).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview version 2—a multiple sequence alignment editor and analysis workbench. Bioinform. Oxf. Engl. 25, 1189–1191 (2009).
Zhang, W. et al. Lactate is a natural suppressor of RLR signaling by targeting MAVS. Cell 178, 176–189.e15 (2019).
Wang, F. et al. Combined anti-PD-1, HDAC inhibitor and anti-VEGF for MSS/pMMR colorectal cancer: a randomized phase 2 trial. Nat. Med. 30, 1035–1043 (2024).
Ahmed, K. A. et al. Clinical outcomes of melanoma brain metastases treated with stereotactic radiosurgery and anti-PD-1 therapy, anti-CTLA-4 therapy, BRAF/MEK inhibitors, BRAF inhibitor, or conventional chemotherapy. Ann. Oncol. J. Eur. Soc. Med. Oncol. 27, 2288–2294 (2016).
Hegde, P. S. & Chen, D. S. Top 10 challenges in cancer immunotherapy. Immunity 52, 17–35 (2020).
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015).
Zhou, W. T. & Jin, W. L. B7-H3/CD276: an emerging cancer immunotherapy. Front. Immunol. 12, 701006 (2021).
Diskin, C., Ryan, T. A. J. & O’Neill, L. A. J. Modification of proteins by metabolites in immunity. Immunity 54, 19–31 (2021).
Qin, W., Yang, F. & Wang, C. Chemoproteomic profiling of protein-metabolite interactions. Curr. Opin. Chem. Biol. 54, 28–36 (2020).
Shang, S., Liu, J. & Hua, F. Protein acylation: mechanisms, biological functions and therapeutic targets. Signal Transduct. Target. Ther. 7, 396 (2022).
Peng, F. et al. Oncogenic fatty acid oxidation senses circadian disruption in sleep-deficiency-enhanced tumorigenesis. Cell Metab. 36, 1598–1618 (2024).
Du, A. et al. Fatty acids in cancer: metabolic functions and potential treatment. MedComm – Oncol. 2, e25 (2023).
Wood, S. M. et al. Fatty acid-binding protein 1 is preferentially lost in microsatellite instable colorectal carcinomas and is immune modulated via the interferon γ pathway. Mod. Pathol. 30, 123–133 (2017).
Lin, A., Zhang, J. & Luo, P. Crosstalk between the MSI status and tumor microenvironment in colorectal cancer. Front. Immunol. 11, 2039 (2020).
Akbari, V., Kallhor, M., Mollashahi, B. & Abolfazl, A. Transcriptome and network dissection of microsatellite stable and highly instable colorectal cancer. Asian Pac. J. Cancer Prev. 20, 2445–2454 (2019).
Galon, J. & Bruni, D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov. 18, 197–218 (2019).
Herbst, R. S. et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 515, 563–567 (2014).
Shi, W. et al. Immune checkpoint B7-H3 is a therapeutic vulnerability in prostate cancer harboring PTEN and TP53 deficiencies. Sci. Transl. Med. 15, eadf6724 (2023).
Altan, M. et al. Association of B7-H4, PD-L1, and tumor infiltrating lymphocytes with outcomes in breast cancer. NPJ Breast Cancer 4, 40 (2018).
Tawbi, H. A. et al. Relatlimab and nivolumab versus nivolumab in untreated advanced melanoma. N. Engl. J. Med. 386, 24–34 (2022).
Griffioen, A. W., Damen, C. A., Martinotti, S., Blijham, G. H. & Groenewegen, G. Endothelial intercellular adhesion molecule-1 expression is suppressed in human malignancies: the role of angiogenic factors. Cancer Res. 56, 1111–1117 (1996).
Wang, R., Shi, T. & Chen, W. Combination therapies with anti-angiogenesis and B7-H3 blockade in cancers. J. Cancer Immunol. 2, 17–21 (2020).
Huang, C. et al. PD-L1 methylation restricts PD-L1/PD-1 interactions to control cancer immune surveillance. Sci. Adv. 9, eade4186 (2023).
Zhu, Q. et al. O-GlcNAcylation promotes tumor immune evasion by inhibiting PD-L1 lysosomal degradation. Proc. Natl. Acad. Sci. USA 120, e2216796120 (2023).
Devlin, J. R. et al. Structural dissimilarity from self drives neoepitope escape from immune tolerance. Nat. Chem. Biol. 16, 1269–1276 (2020).
Chandran, S. S. et al. Immunogenicity and therapeutic targeting of a public neoantigen derived from mutated PIK3CA. Nat. Med. 28, 946–957 (2022).
Acknowledgements
We acknowledge the equipment support from the Experimental Medicine Center of Tongji Hospital, Tongji Medical School, Huazhong University of Science and Technology, and the technical support from the Medical Subcenter of HUST Analytical and Testing Center. This work was supported by the National Natural Science Foundation of China (no. 82472829 (X. Luo), no. 82425041 (G. Wang), no. 82330084 (G. Wang)), the National Key Research and Development Program of China (2022YFA1105303 (G. Wang), 2023YFC3402100 (G. Wang)), the Basic Research Support Program of Huazhong University of Science and Technology (2023BR036 (G. Wang)), and the Noncommunicable Chronic Diseases—National Science and Technology Major Project (no. 2024ZD0520600 (J. Hu)).
Author information
Authors and Affiliations
Contributions
X. Luo and J. Hu conceived and designed the study. Z. Rao performed most experiments and analyzed the data. C. Huang, Q. Wu, and M. Li performed the animal experiments. A. Liu, T. Zhu, and W. Yang collected biological samples. L. Yin, S. Zhu, X. She, and C. Yu performed data analysis. Lang Liu, P. Li, Y. Bai, and D. Zhang curated and compiled patients’ clinical and pathological data. T. Xie and X. Liu reviewed and edited the manuscript. Lu Liu and F. Xu provided experimental support. G. Wang provided project-level guidance, contributed to study design and data interpretation, and critically revised the manuscript with constructive suggestions. Z. Rao and X. Luo drafted the manuscript and revised it. X. Luo and J. Hu supervised the project. All authors reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sérgio Veloso and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rao, Z., Huang, C., Wu, Q. et al. Palmitic acid-triggered B7H3 palmitoylation promotes immune escape. Nat Commun 17, 1810 (2026). https://doi.org/10.1038/s41467-026-68525-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68525-x