Abstract
As the prevalence of cardiovascular risk factors in children continues to rise globally, identifying modifiable lifestyle factors becomes increasingly important for early prevention. Here, using data from a large-scale cohort (2021–2024) involving 7459 Chinese children from three provinces, we examine the association between eating timing patterns and multiple cardiovascular parameters. Participants are classified into three eating patterns: extended eating window (duration more than 12.5 hours), late eating window (duration 12.5 hours or less with last meal after 20:00), and early eating window (duration 12.5 hours or less with last meal at or before 20:00). Early eating window is associated with a 26% lower risk of elevated blood pressure (odds ratio: 0.74, 95% confidence interval: 0.65–0.84, p < 0.001) compared to extended eating window. We conduct a nested case-control study (n = 51) with comprehensive lipidomic and proteomic analyses. Proteomics identifies 83 differentially expressed proteins, with enrichment in cardiac muscle contraction and lipid metabolism pathways. Mediation analysis demonstrates that triacylglycerols mediate 64.8% of the effect between eating patterns and blood pressure. Our findings suggest associations between early eating windows and protective cardiovascular parameters in children, potentially mediated through lipid metabolism pathways.
Similar content being viewed by others
Introduction
Cardiovascular diseases (CVDs) remain the leading cause of global mortality, with their pathophysiological foundations often established during childhood and adolescence1,2. The alarming rise in cardiovascular risk factors among youth worldwide represents a critical public health challenge requiring innovative preventive approaches beyond traditional interventions3. While nutritional research has historically emphasized dietary composition, emerging evidence suggests that the temporal pattern of energy intake—specifically when we eat—may be as important as what we eat for cardiometabolic health4,5.
This temporal dimension of nutrition aligns with our expanding understanding of circadian biology—the intricate system of molecular clocks that orchestrates physiological processes, including vascular tone, cardiac contractility, glucose homeostasis, and lipid metabolism5. These endogenous circadian rhythms are synchronized by both photic cues (light exposure) and non-photic zeitgebers, particularly feeding patterns, creating an intricate bidirectional relationship between meal timing and cardiometabolic function6,7,8. Time-restricted eating (TRE), the practice of limiting food intake to specific daily time windows without necessarily restricting calories, has emerged as a promising strategy for improving cardiovascular health in adults9,10,11. Compelling evidence from clinical investigations demonstrates that later last meal timing is consistently linked with elevated blood pressure (BP), obesity, and adverse lipid profiles12. Building on these findings, recent studies have revealed that restricting food intake to earlier parts of the day confers enhanced cardiometabolic benefits compared to later time-restricted eating patterns5,13,14, suggesting that both the duration and timing of eating are critical determinants of cardiovascular health.
Despite this robust evidence in adults, research examining meal timing patterns and cardiovascular health in pediatric populations remains notably limited. This research deficit is particularly significant in light of the global shift in youth eating patterns, which increasingly feature extended grazing behaviors and progressively delayed last meal timing15,16. These temporal shifts in energy intake are occurring during a pivotal developmental window when vascular structure, autonomic regulation, and metabolic programming are being established, potentially creating a compounding effect with dietary composition that may influence cardiovascular health trajectories throughout the lifespan17. Furthermore, while adult studies have begun to identify candidate molecular pathways underlying the cardiovascular effects of eating timing18, the specific mechanisms by which eating timing characteristics influence cardiovascular health in developing children remain largely unexplored, limiting our ability to develop targeted preventive strategies for this vulnerable population.
This study aims to investigate the relationships between eating timing characteristics and comprehensive cardiovascular parameters in a large cohort of children and adolescents. By integrating detailed cardiovascular assessments with lipidomics and proteomics analyses, we seek to elucidate the molecular mechanisms underlying these relationships. This integrative approach may provide insights into how eating patterns are associated with cardiovascular health during critical developmental periods, potentially informing evidence-based recommendations for early cardiovascular risk prevention through optimized meal timing strategies.
Results
Baseline characteristics and study population
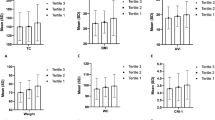
Among the 7459 participants (3848 boys and 3611 girls), significant sex differences were observed in anthropometric and cardiovascular parameters (Table 1). Boys demonstrated higher body mass index (BMI) (18.2 ± 3.6 vs 17.9 ± 3.5 kg/m², P < 0.01), waist-to-height ratio (WHtR) (0.4 ± 0.1 vs 0.4 ± 0.0, P < 0.01), systolic BP (SBP, 102.7 ± 11.9 vs 100.5 ± 10.3 mmHg, P < 0.01), higher daily physical activity levels (2.3 ± 1.0 vs 1.9 ± 0.8 metabolic equivalent (MET)-hours, P < 0.01), and daily energy intake (1583.4 ± 431.2 vs 1548.8 ± 413.7 kcal, P < 0.01). Cardiovascular parameters showed that boys had marginally higher carotid intima-media thickness (CIMT) bilaterally (right: 0.49 ± 0.06 vs 0.48 ± 0.06 mm, P < 0.05; left: 0.49 ± 0.06 vs 0.48 ± 0.07 mm, P < 0.05) and larger cardiac dimensions.
Associations between eating window and last meal timing with cardiovascular parameters
After comprehensive covariate adjustment, participants with longer eating windows showed significantly adverse cardiovascular parameters (Fig. 1a, Supplementary Table 1). Each additional hour of eating window duration was associated with higher systolic BP (β = 0.312, 95% confidence interval (CI): 0.177–0.446, P.adj < 0.001) and diastolic BP (β = 0.354, 95% CI: 0.246–0.462, P.adj < 0.001). Eating window duration positively associated with bilateral CIMT (right: β = 0.010, 95% CI: 0.008–0.012, P < 0.001; left: β = 0.010, 95% CI: 0.007–0.012, P.adj<0.001). Cardiac structural parameters demonstrated that longer eating windows were associated with larger left ventricular dimensions and reduced cardiac function indices. Brachial-ankle pulse wave velocity (baPWV) measurements did not show statistically significant associations (right: β = −4.866, 95% CI: −11.486–1.753, P.adj = 0.15; left: β = −3.757, 95% CI: −11.146-3.632, P.adj = 0.32).
a Restricted cubic spline analysis showing the non-linear relationships between eating window (left) or last meal time (right) and hypertension risk. Histograms show the distribution of exposures. Solid red lines indicate odds ratios (ORs); dashed lines represent 95% confidence intervals (CIs). P10, P50 and P90 represent the 10th, 50th and 90th percentiles, respectively. Statistical analysis was performed using restricted cubic spline analysis with mixed-effects logistic regression (two-sided). b Forest plot of hypertension risk stratified by eating patterns. Results are presented as ORs and 95% CIs. Eating timing patterns were categorized as: Extended EW (eating window >12.5 h), Late EW (eating window ≤12.5 h with last meal after 20:00), and Early EW (eating window ≤12.5 h with last meal no later than 20:00). The Extended EW group served as the reference. All models were adjusted for sex, age, height Z-score, waist-to-height ratio, area of residence, physical activity, total caloric intake, macronutrient composition, dietary quality score, parental education, parental occupation, annual household income, parental marital status, and family history of hypertension. Results are presented as odds ratios and 95% confidence intervals, where the odds ratio represents the point estimate and the confidence interval bars represent the uncertainty range. Statistical analysis was performed using mixed-effects logistic regression with false discovery rate correction (two-sided).
Later last meal timing was independently associated with adverse cardiovascular profiles (Fig. 1b, Supplementary Fig. 2b, Supplementary Table 1). Each hour delay in last meal timing was associated with higher systolic BP (β = 0.360, 95% CI: 0.208–0.512, P < 0.001) and diastolic BP (β = 0.294, 95% CI: 0.172–0.416, P < 0.001). Structural cardiovascular parameters showed that later meal timing was associated with increased bilateral CIMT and larger cardiac dimensions.
Eating timing pattern classification and cardiovascular outcomes
Restricted cubic spline analysis identified optimal thresholds at 12.5 hours for the eating window and 20:00 for last meal timing (Fig. 1a, Supplementary Table 2, 3). Binary threshold analysis revealed that an eating window of more than 12.5 hours was associated with significantly higher risk of elevated BP compared to 12.5 hours or less (odds ratio (OR) = 1.32, 95% CI: 1.17–1.48, P < 0.001). Similarly, last meal timing after 20:00 was associated with increased risk compared to meals at or before 20:00 (OR = 1.53, 95% CI: 1.37–1.72, P < 0.001) (Fig. 1b).
Based on identified thresholds, participants were classified into three distinct eating timing patterns. Using the Extended Eating Window (Extended EW) as the reference group, both Late EW and Early EW demonstrated protective associations. Late EW was associated with reduced risk of elevated BP (OR = 0.83, 95% CI: 0.69–0.99, P = 0.04), while Early EW showed the strongest protective effect (OR = 0.74, 95% CI: 0.65–0.84, P < 0.001) after comprehensive covariate adjustment (Fig. 1b).
Subgroup analyses of eating timing patterns
Stratified analyses demonstrated consistent protective associations of Early EW compared to Extended EW across diverse population subgroups (Supplementary Table 4). Early EW showed protective effects in both males (OR = 0.67, 95% CI: 0.56–0.80, P < 0.001) and females (OR = 0.81, 95% CI: 0.67–0.97, P = 0.025), across age groups, and in both normal weight and overweight participants.
Sensitivity analyses
We conducted sensitivity analyses using alternative thresholds (Supplementary Table 5). Using a 12-hour threshold for eating window duration yielded consistent results, with extended eating windows (>12 h) showing significantly higher risk compared to restricted windows (≤12 h) (OR = 1.24, 95% CI 1.08–1.42, P = 0.003). Alternative last meal timing thresholds of 19:00 (>19:00 vs ≤19:00: OR = 1.24, 95% CI: 1.05–1.47, P < 0.05) and 21:00 (>21:00 vs ≤21:00: OR = 1.11, 95% CI: 0.96–1.23, P = 0.16) showed consistent directional effects. Analysis without multiple imputation for missing values yielded similar associations (Early EW vs Extended EW: OR = 0.78, 95% CI: 0.68–0.89, P < 0.001; Late EW vs Extended EW: OR = 0.85, 95% CI: 0.70–1.03, P = 0.09).
Proteomic signatures associated with eating timing patterns
In the nested case-control component (n = 51), we observed the extended EW group exhibited significantly higher BP (SBP: 127.2 ± 12.8 mmHg; DBP: 74.5 ± 8.8 mmHg) and arterial stiffness compared to both early (SBP: 104.9 ± 10.0 mmHg; DBP: 63.2 ± 7.7 mmHg) and late EW groups (P < 0.001) (Supplementary Table 9). Proteomic analysis identified 112 differentially expressed proteins (58 up-regulated, 54 down-regulated) across the three eating timing patterns (Source Data File). The volcano plot analysis (Fig. 2a) revealed significant differential protein expression. Notable differentially expressed proteins included myosin light chain 1 (MYL1), insulin-like growth factor binding protein 1 (IBP1), parvalbumin alpha (PRVA), apolipoprotein H (APOH), and alcohol dehydrogenase 4 (ADH4). WGCNA identified multiple protein modules associated with eating timing patterns, as visualized in the cluster dendrogram (Fig. 2b and Supplementary Fig. 3). The analysis identified 1597 proteins (total 1958 proteins quantified) organized into 36 modules showing strong associations with cardiovascular parameters, particularly SBP and DBP (Supplementary Fig. 4). The Venn diagram (Fig. 2c) demonstrates the substantial overlap (83 proteins: 47 up-regulated, 36 down-regulated) between DEGs and Weighted gene co-expression network analysis (WGCNA) module proteins. Functional enrichment analyses further demonstrated that early eating windows were associated with favorable regulation of cardiac muscle contraction pathways and lipid metabolic processes (e.g., fatty acid degradation, cholesterol metabolism), while extended windows activated pathways implicated in cardiomyopathy and cellular stress responses (Fig. 2e and Supplementary Fig. 5). Protein-protein interaction network analysis highlighted apolipoprotein A4 (APOA4), cholesteryl ester transfer protein (CETP), and phospholipid transfer protein (PLTP), as hub proteins with high connectivity, which are critical for lipid transport and metabolism, suggesting their central regulatory roles in the molecular networks associated with different eating timing patterns (Fig. 2d).
a Volcano plot showing differential protein expression across three comparison groups. The y-axis represents log2 fold-change (negative values indicate down-regulation, positive values indicate up-regulation). Red dots represent significantly differentially expressed proteins (FDR-adjusted P < 0.05), while blue dots represent non-significant proteins. Differential expression analysis was performed using DESeq2 with false discovery rate adjustment (two-sided). b Weighted Gene Co-expression Network Analysis (WGCNA) cluster dendrogram showing module assignment based on proteomic data from 1958 quantified proteins. Colors represent different protein modules (n = 36 modules) identified by hierarchical clustering, with the y-axis showing the dissimilarity between proteins based on their co-expression patterns. c Venn diagram illustrating the overlap between differentially expressed proteins (DEPs) and proteins identified in significant WGCNA modules. d Protein-protein interaction network of key differentially expressed proteins. e Functional enrichment analysis of the differentially expressed proteins from the intersection analysis. The dot plot shows significantly enriched Gene Ontology (GO) terms (adjusted P < 0.05), with dot size representing gene count and color intensity representing statistical significance level. The GO analysis is categorized into three main domains: Biological Process (BP), Cellular Component (CC), and Molecular Function (MF).
Lipidomic profiles across eating timing patterns
Principal component analysis demonstrated clear separation among the three eating patterns, with PC1 and PC2 explaining 43.7% and 9.4% of the total variance, respectively (Fig. 3a). The extended EW group exhibited a lipid profile distinctly separated from both early and late EW groups along PC1, while early and late EW groups showed partial overlap with distinguishable cluster centers. Altered lipid classes included phosphatidylethanolamines (PE), sphingomyelins (SM), and triacylglycerols (TAG), with participants in the extended EW group exhibiting elevated levels of specific SMs and TAGs compared to the early EW group (Fig. 3b). Hierarchical clustering of the top differential lipids revealed distinct patterns across the three eating patterns (Fig. 3c). Two major lipid clusters were identified: one predominantly comprising TAGs that were elevated in the extended EW group, and another mainly consisting of PEs that were downregulated in the extended EW group compared to early and late EW groups. Notably, TAG species with longer carbon chains and higher unsaturation (TAGs 52-58) showed the most pronounced elevation in the extended EW group. Pathway enrichment analysis of differential metabolites revealed significant perturbations in multiple metabolic pathways (Fig. 3d). The most significantly enriched pathways were related to obesity, hypertension, and glycerophospholipid metabolism.
a Principal component analysis (PCA) score plot. Principal Component 1 (PC1) and PC2 explain 43.7% and 16.6% of the total variance, respectively. b Bubble plot of differential lipid species. Lipid classes are defined as: BMP bis(monoacylglycero)phosphate, CE cholesteryl ester, CER ceramide, DAG diacylglycerol, DGDG digalactosyldiacylglycerol, FFA free fatty acid, HCER hexosylceramide, LPC lysophosphatidylcholine, LPE lysophosphatidylethanolamine, LPI lysophosphatidylinositol, LPS lysophosphatidylserine, PC phosphatidylcholine, PE phosphatidylethanolamine, PG phosphatidylglycerol, PI phosphatidylinositol, SM sphingomyelin, TAG triacylglycerol. c Heatmap of the top differential lipids. d Pathway enrichment analysis for differential metabolites. The outer circle represents the total metabolites in each pathway, and the inner circles show hit counts and enrichment factors. The outer ring represents the total number of metabolites in each pathway, and inner rings show hit counts and enrichment factors, with color intensity representing statistical significance levels.
Hub metabolites
From lipidomic profiles comparing early versus extended eating windows, we identified blood pressure-associated lipid biomarkers through consensus analysis of four machine learning algorithms (Supplementary Figs. 6–11). Six lipids were consistently selected as core hubs by all algorithms, with an additional 21 high-confidence hubs identified by ≥3 algorithms. (Fig. 4). This heatmap demonstrates clear separation between case (elevated BP) and control groups, with multiple TAGs consistently upregulated in participants with elevated BP (Fig. 4c). Box plots of core hub metabolites selected by all four machine learning models further confirmed these patterns (Fig. 4d). TAG(52:2)_FA14:0, TAG(56:6)_FA18:1, TAG(56:6)_FA20:5, and TAG(56:6)_FA22:5 showed significantly higher expression levels in participants with elevated BP, while PC(18:0/18:0) and PE(O/16:0/18:1) showed lower levels. Parallel mediation analysis revealed that down-regulated lipid metabolites had a significant total indirect effect of −0.229 (95% CI: −0.403, −0.054), with these mediators accounting for 64.8% of the total effect between dietary pattern and elevated BP. (Supplementary Fig. 12).
a Down-regulated Lipid Features Selected by 4 Models. Venn diagram showing overlap of down-regulated lipid features (from Extended Eating Window (EW) versus Early EW) that were selected by four machine learning algorithms as associated with elevated blood pressure (HBP) (Random Forest (RF), Support Vector Machine (SVM), XGBoost, Elastic Net). The intersection of all four methods identifies the most robust down-regulated biomarkers with consensus support across all analytical approaches. b Up-regulated Lipid Features Selected by 4 Models. Venn diagram showing overlap of up-regulated lipid features (from Early EW versus Extended EW comparison) that were selected by four machine learning algorithms as associated with elevated blood pressure. c Heatmap of high-confidence hub lipid metabolites selected by at least three algorithms. d Core Hub Metabolites Expression by HBP Status. Box plots comparing the expression levels of the six core hub metabolites between normotensive controls (n = 25) and children with elevated blood pressure (n = 26). Box plots show the median (center line), interquartile range (box bounds: 25th–75th percentile), whiskers extend to 1.5× interquartile range from box bounds, and outliers are shown as individual points. Statistical analysis was performed using a two-sided Wilcoxon rank-sum test. Exact P values are displayed on the figure. *P < 0.05, **P < 0.01, ***P < 0.001. TAG triacylglycerol, FA fatty acid, PC phosphatidylcholine, PE phosphatidylethanolamine, Hex2Cer dihexosylceramide, HexCer hexosylceramide.
Discussion
This study provides a comprehensive investigation of associations between eating timing patterns and cardiovascular health in children through an integrative approach combining clinical parameters with molecular profiling. Our findings demonstrate that eating timing characteristics—specifically duration and last meal timing—independently influence cardiovascular parameters, with extended eating windows (>12.5 hours) showed a 32% higher risk of elevated blood pressure compared to those with shorter eating windows (≤12.5 hours), and last meal timing after 20:00 showed a 53% higher risk compared to ≤20:00. These associations were accompanied by measurable differences in subclinical cardiovascular parameters, including elevated carotid intima-media thickness and modest changes in cardiac dimensions. While these differences remain within clinically normal ranges, their presence during childhood may have implications for long-term cardiovascular health trajectories, though longitudinal studies are needed to establish this relationship.
Our findings align with emerging evidence from adult chronobiology research, where extended eating windows and delayed meal timing have been consistently linked to elevated blood pressure and metabolic dysregulation9,10,19,20,21,22,23,24,25. Despite these compelling adult findings, pediatric populations have remained conspicuously underrepresented in cardiovascular-focused chronutrition research, with existing studies primarily examining metabolic outcomes in adolescents with established obesity or Type 2 diabetes26,27. Our investigation addresses this critical gap by examining temporal eating patterns in healthy children, revealing that these behavioral dimensions may represent previously underrecognized modifiable factors for early cardiovascular risk stratification. The remarkable consistency of associations across diverse demographic subgroups—including sex, age, weight status, and residential location—underscores the generalizability of these relationships within similar populations.
The temporal association between late last meal timing (after 20:00) and elevated blood pressure risk merits particular emphasis within the pediatric developmental context. This finding extends adult epidemiological studies demonstrating 55% increased coronary heart disease odds with late-night eating28, and 17% higher metabolic syndrome prevalence among individuals consuming late dinners29. Our timing-dependent effects during childhood suggest that dietary temporal organization may establish cardiovascular risk trajectories during critical developmental windows, when metabolic programming and vascular architecture undergo fundamental maturation processes.
The elevated CIMT observed among children with extended eating windows represents our most clinically significant finding, given its established predictive value for future cardiovascular events30. CIMT serves as a validated surrogate marker for systemic atherosclerotic burden, with even modest pediatric elevations predicting accelerated disease progression and premature cardiovascular morbidity in young adults31. The detection of measurable CIMT differences during childhood—when atherosclerotic processes are typically nascent—suggests that eating timing patterns may influence vascular health through mechanisms that manifest earlier in the life course than previously recognized. These structural vascular changes likely reflect early endothelial dysfunction and adaptive arterial wall remodeling in response to chronic circadian misalignment-induced metabolic stress. The parallel observation of subtle but significant cardiac dimensional changes further supports this interpretation, as these modifications—while remaining within normal pediatric ranges—have been prospectively linked to adverse ventricular remodeling and cardiovascular complications in longitudinal studies. Collectively, these subclinical findings underscore the potential long-term cardiovascular implications of eating timing patterns established during formative childhood years.
Our integrated multi-omics analysis unveiled distinct molecular signatures associated with different eating patterns, offering preliminary mechanistic insights into the pathways potentially mediating our observed clinical associations. Notably, the decreased expression of APOH in early eating window participants warrants attention, given APOH’s established associations with metabolic syndrome, diabetes, and cardiovascular disease risk32. While this expression pattern aligns with potential cardioprotective mechanisms, experimental confirmation of causality remains essential. Conversely, the enhanced expression of muscle-contractile proteins (MYL1, MYH1) in early eating window participants suggests augmented skeletal muscle function and metabolic capacity33,34. Pathway enrichment analysis revealed that early eating windows were associated with favorable modulation of cardiac muscle contraction and lipid metabolic pathways, while extended eating windows activated stress response and cardiomyopathic signaling cascades. Network topology analysis identified key hub proteins—including APOA4, CETP, and PLTP—that occupy central positions in lipid transport and reverse cholesterol transport networks, suggesting that eating timing may influence cardiovascular risk through modulation of lipoprotein metabolic efficiency.
The lipidomic profiles revealed distinct patterns associated with different eating window categories, with the most striking differences observed in PE and TAG. Early eating window participants showed enrichment of phosphatidylethanolamines—essential membrane constituents critical for mitochondrial bioenergetics and cellular energy homeostasis35,36,37. This lipid profile suggests that optimal eating timing may enhance mitochondrial function and reduce oxidative stress, contributing to improved cardiovascular health through fundamental cellular metabolic mechanisms. Conversely, extended eating window participants exhibited elevated concentrations of longer-chain, highly unsaturated triacylglycerols (particularly TAGs 52-58), species previously implicated in arterial stiffness and hypertension pathogenesis38. Our mediation analysis demonstrated that specific triacylglycerol species collectively mediated 64.8% of the total effect linking eating timing patterns to elevated blood pressure risk—a substantial proportion suggesting that lipid metabolic alterations may serve as a primary pathway through which eating timing influences pediatric cardiovascular health. However, these observational findings cannot establish causality; experimental validation through controlled intervention studies is required to confirm whether modifying eating timing directly induces these metabolic and cardiovascular changes.
From a clinical perspective, our findings suggest that systematic assessment of eating timing patterns could augment conventional pediatric cardiovascular risk management. Current clinical recommendations for childhood cardiovascular risk reduction emphasize dietary quality, physical activity, and weight management. Our findings suggest that temporal eating organization—specifically, encouraging earlier meal completion and avoiding extended eating windows—may represent an additional dimension worthy of clinical consideration. Given the high prevalence of cardiovascular risk factors in children globally and the challenges associated with implementing dietary composition changes3, optimizing meal timing represents a complementary and potentially more feasible intervention strategy. Additionally, through consensus machine learning analysis incorporating four independent algorithms, we identified six core hub metabolites: TAG(52:2), TAG(56:6), TAG(56:6)_FA20:5, TAG(56:6)_FA22:5, PC(18:0/18:0), and PE(O-16:0/18:1). These species demonstrated consistent discriminatory capacity for eating timing-associated cardiovascular risk, showing promise as intervention monitoring biomarkers. However, translating these research findings into clinical practice requires several critical validation steps, including replication in larger, ethnically diverse cohorts and demonstration of intervention efficacy through randomized controlled trials.
This study’s strengths include its large-scale cohort, comprehensive cardiovascular phenotyping including subclinical markers, and multi-omics analyses in pediatric chronunutrition research. The consistent associations observed across demographic subgroups suggest potential robustness of the primary findings. However, several limitations must be acknowledged. The observational design precludes causal inference, and residual confounding by unmeasured factors cannot be excluded, including unmeasured dietary components (specific bioactive compounds, food processing methods), timing-specific physical activity patterns, sleep behaviors, and genetic background. The omics component’s sample size, while providing valuable exploratory insights, limits generalizability and necessitates external validation. Our study population comprised Chinese children from southwest provinces, potentially limiting applicability to other populations given ethnic, cultural, and lifestyle differences in dietary behaviors and circadian biology. Future research should prioritize larger multi-omics studies with external validation, randomized controlled trials testing eating timing interventions in diverse pediatric populations, and longitudinal investigations examining whether childhood eating patterns influence adult cardiovascular outcomes.
Methods
Study design and participants
Cohort study
This research complies with all relevant ethical regulations. Ethical approval was obtained from the Ethics Committee of the Children’s Hospital of Chongqing Medical University (No. 2021.97), and all procedures were conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from all participants and their legal guardians. Participants did not receive monetary compensation but received free health examination results and personalized health recommendations. A large-scale cohort study was conducted to investigate the impact of eating timing patterns on cardiovascular health in children and adolescents. A two-stage cluster sampling method was employed for participant recruitment. Participants aged 6–17 years old with continuous residence in the target region for at least six months were eligible for inclusion, and both parents and children signed the informed consent form. Exclusion criteria included: (1) congenital heart disease or other cardiovascular anomalies; (2) kidney disease or endocrine disorders including diabetes mellitus; (3) current use of medications affecting blood pressure or lipid metabolism; (4) acute infectious illness within two weeks prior to assessment; (5) serious digestive system diseases or metabolic disorders affecting nutritional metabolism; or (6) inability to complete assessments due to physical or cognitive limitations. The cohort was established between 2021 and 2023, with staggered recruitment timelines across three provinces in Southwest China: Sichuan Province (n = 2111) in 2021, Chongqing Municipality (n = 5232) in 2022, and Guizhou Province (n = 2506) in 2023, yielding an initial cohort of 9849 participants. After applying exclusion criteria, our final analytical sample comprised 7459 participants. Follow-up assessments were conducted in Chongqing in 2023 (first follow-up, n = 3830) and 2024 (second follow-up, n = 2314), and in Guizhou and Sichuan in 2024 (first follow-up, n = 3274). Follow-up assessments yielded 15,662 blood pressure measurements, 4920 pulse wave velocity assessments, 2862 echocardiographic evaluations, and 3203 biomedical analyses. Supplementary Fig. 1 presented the detailed participant flow diagram. Detailed information about the study design has been published previously39.
Nested case-control study
For the nested case-control study, participants were selected during the second follow-up period using a systematic protocol. Eligibility criteria included completion of all available assessment points, consistent eating pattern across baseline and follow-up assessments, no disease or use of medications affecting metabolic parameters, and agreement to provide additional blood samples for omics analyses. Sample size was calculated using the formula nA = κnB and nB = (1 + 1/κ)(σ(Z1-α/2 + Z1-β)/(μA-μB))2. With parameters α = 0.0167 (adjusted for multiple comparisons), statistical power=93%, and anticipated effect size based on pilot data (σ = 4.0), we determined that 12 participants per group would provide sufficient statistical power. To account for potential sample quality issues, we increased the sample size to 17 participants per group. From eligible participants, we randomly selected 17 individuals from each eating pattern category (Extended EW, Late EW, and Early EW) using computer-generated random numbers, verifying balance in age, sex, and height distribution across groups. The final nested case-control sample comprised 51 participants, who provided additional fasting blood samples for comprehensive lipidomics and proteomics analyses.
Measurement of eating timing characteristics
Eating timing information was collected using a standardized protocol that remained consistent across all study sites40,41. Validated eating pattern questionnaires were administered to all participants at baseline and during subsequent follow-up visits. For children aged 6–11 years, parents/guardians completed the questionnaires with input from the children, while for participants aged 12–17 years, questionnaires were self-administered with parental verification. Participants reported their typical first and last mealtimes over the past month preceding each assessment, including any caloric beverages or snacks. The primary eating timing parameters were: (1) eating window duration, calculated as the time interval between the first and last energy intake of the day; and (2) last meal timing, recorded as the clock time (24-hour format) of the final energy intake each day. To determine optimal thresholds for eating window categorization, we employed restricted cubic spline analysis with model selection criteria (Akaike Information Criterion and Bayesian Information Criterion) to identify natural inflection points in the relationship between eating window and cardiovascular outcomes. The analysis consistently identified the primary inflection point at 12.5 hours (corresponding to the 50th percentile of our eating window distribution), indicating a meaningful change in the dose-response relationship. Similarly, spline analysis for last meal timing identified 20:00 as the optimal threshold, representing the median last meal timing in our population. Based on these parameters, participants were categorized into three distinct eating pattern groups: (1) Extended Eating Window (Extended EW): eating window >12.5 hours, regardless of last meal timing (e.g., first meal at 07:00, last meal at 20:30, duration = 13.5 hours), (2) Late Eating Window (Late EW): eating window ≤12.5 hours with last meal after 20:00 (e.g., first meal at 08:00, last meal at 20:30, duration = 12.5 hours), and (3) Early Eating Window (Early EW): eating window ≤12.5 hours with last meal no later than 20:00 (e.g., first meal at 07:00, last meal at 19:30, duration = 12.5 hours).
Measurement of cardiovascular indicators
Blood pressure
BP was measured using an arm-type electronic sphygmomanometer (HEM-7051, OMRON, Japan) following standardized protocols42. Three consecutive measurements were taken at 2-minute intervals with participants remaining silent and still, and the average of the three measurements was used for analysis. Elevated BP was defined as mean systolic and/or diastolic blood pressure ≥90th percentile based on age-, sex-, and height-specific reference standards for Chinese children43.
Cardiovascular structure
Cardiac structure and function indicators were evaluated using standardized echocardiography (Acuson Sequoia 512 system, Siemens, Germany) by certified sonographers who were blinded to participants’ clinical information. Parasternal long-axis, short-axis, and apical views were obtained following American Society of Echocardiography guidelines. Left ventricular end-diastolic diameter (LVDd) and end-systolic diameter (LVDs) were measured from M-mode recordings, with measurements averaged across three cardiac cycles. Ejection fraction (EF) and fractional shortening (FS) were calculated using standard formulas. For CIMT measurement, high-resolution B-mode ultrasound images were obtained from both common carotid arteries, with measurements taken 1 cm proximal to the carotid bulb. The mean value of three measurements on each side was used for analysis.
Arterial stiffness
Arterial stiffness was assessed using brachial-ankle pulse wave velocity (baPWV) measured with an automatic arteriosclerosis detection device (BP-203RPE III, OMRON, Japan). Measurements were performed in a temperature-controlled room (22–25 °C) after participants rested in a supine position for 10 minutes. BP cuffs were applied to both brachial and ankle arteries, and baPWV was calculated as the distance between measurement points divided by pulse wave transit time. The average of left and right baPWV values was used for analysis. Anthropometric measurements were performed by trained nurses using standardized procedures.
Biochemical Parameters
Fasting blood samples were drawn by trained nurses between 07:00 and 09:00 AM after an overnight fast of at least 8 hours. The samples were processed within 2 hours of collection. Plasma lipid profile (Total cholesterol (TC), triglycerides (TG), low-density lipoprotein cholesterol (LDL-C), high-density lipoprotein cholesterol (HDL-C)), fasting glucose, and liver function tests were analyzed using a fully automatic biochemical analyzer (ADVIA 2400, Siemens, Germany). Non-HDL-C was calculated as the difference between total cholesterol and HDL-C. Quality control-maintained CV < 10% with external assessment through Fuwai Hospital, Beijing.
Lipidomics analysis
Fasting blood samples were collected between 07:00 and 09:00 AM. After 1 h stratification, samples were centrifuged at 1000 × g for 10 min at room temperature. The supernatants were immediately frozen in liquid nitrogen for 15 min and stored at −80 °C. Hemolyzed samples were excluded. Lipids in plasma samples were extracted using a modified Bligh and Dyer method, followed by analysis on an ultra-HPLC-tandem mass spectrometry system (Q Exactive HF - X, Thermo Fisher Scientific). Quality control (QC) was strictly implemented, with QC samples analyzed every 10 samples. A total of 683 lipid species from 24 lipid classes were quantified, with relative standard deviation <30% in QC samples and detection rate > 80% across all study samples. The detailed lipid extraction, chromatographic separation, and mass spectrometry conditions were in accordance with the methods described in published literature, ensuring the reliability and reproducibility of the lipidomic data44.
Proteomics analysis
High - abundance proteins were depleted using the Multiple Affinity Removal System to improve the detection of low - abundance proteins. The remaining proteins were then subjected to tryptic digestion. Peptide samples were analyzed using a nano-UPLC system (Evosep One) coupled with a high - resolution mass spectrometer (timsTOF Pro2, Bruker). To ensure reproducibility45, a pooled quality control (QC) sample was created from equal volumes of all digested samples and analyzed repeatedly throughout the analytical sequence. QC samples demonstrated technical reproducibility with Pearson correlation coefficients >0.95. Data processing was performed using Spectronaut software against the UniProt human database, with the false discovery rate (FDR) controlled at 1% for both peptides and proteins. In total, 1958 proteins were identified and quantified. The proteomics data analysis, including peptide identification, protein quantification, and FDR control, was based on the methods described in published literature44,46,47,48. Detailed methods for lipidomics and proteomics analysis were provided in the Supplementary Methods. All lipidomics and proteomics measurements were conducted by Shanghai Biotree Biomedical Technology Co., Ltd.
Anthropometric measurements
Height was measured to the nearest 0.1 cm using a stadiometer, and weight was measured to the nearest 0.1 kg using a calibrated electronic scale, with participants wearing light clothing and no shoes. Waistline was measured at the midpoint between the lower border of the rib cage and the iliac crest to the nearest 0.1 cm. BMI was calculated as weight (kg)/height(m)2, and WHtR was calculated as waist circumference(cm)/height(cm). Overweight and obesity were defined using age- and sex-specific BMI cutoffs based on Chinese national standards49, where ≥85th percentile indicates overweight and ≥95th percentile indicates obesity.
Assessment of covariates
Demographic and lifestyle information was collected through standardized questionnaires administered by trained interviewers. Physical activity was evaluated using the Physical Activity Questionnaire for Adolescents (PAQ-A), which provided detailed information on frequency and duration of activities based on MET values50. Comprehensive dietary assessment used validated semi-quantitative food frequency questionnaires adapted for Chinese pediatric populations, capturing total caloric intake, macronutrient composition, and dietary quality scores based on the Chinese Residents’ Balanced Diet Pagoda, which captured the frequency and portion size of commonly consumed foods51,52,53. Dietary variables included: (1) total daily caloric intake (kcal/day); (2) macronutrient composition expressed as percentages of total calories from carbohydrates, fats, and proteins; (3) dietary quality score based on adherence to the Chinese Healthy Eating Index for children. The area of residence was classified as urban or rural based on the current living address. Socioeconomic status indicators included parental education level (categorized as junior high school and below, high school/vocational school, or college and above), occupation (categorized as workers/migrant workers, farmers, management/soldiers/technicians, or others), and annual household income. Family history of hypertension and parental marital status (married, divorced, or other) were also recorded.
Sex and gender considerations
Sex was considered in the study design with stratified analyses by sex. Biological sex was determined based on legal identification documents (household registration) as verified by parents/guardians. All analyses were adjusted for sex, and sex-stratified results are presented in the subgroup analyses. Sex-disaggregated data are provided in Table 1 (baseline characteristics separately for boys n = 3848 and girls n = 3611) and in Source Data files.
Quality control and standardization
Comprehensive quality control measures were implemented across all measurement sites. All personnel underwent standardized training programs including theoretical instruction (2 hours) and hands-on practice (24 hours). Equipment was calibrated daily (blood pressure monitors, scales) or weekly (ultrasound systems, pulse wave velocity devices) using certified standards. For cardiovascular measurements, 1% of assessments were repeated by independent observers to ensure reliability. All blood samples were processed locally within 2 hours and analyzed at a central laboratory to ensure consistency. External quality assessment through the National Center for Cardiovascular Diseases. Cross-site standardization was maintained through inter-site reliability assessments with standardized operators, and comprehensive equipment maintenance by certified technicians.
Statistical analysis
Descriptive statistics were presented as means ± standard deviations for continuous variables and frequencies (percentages) for categorical variables. Between-group differences were assessed using Student’s t-tests for continuous variables and chi-square tests for categorical variables. Height Z-scores were calculated to standardize values across age and sex groups. Missing data patterns were assessed, with missing rates of socioeconomic status, dietary intake questionnaires, and physical activity assessments. Missing data (<30% for any variable) were handled using multiple imputations with the expectation-maximization algorithm (five imputations). Mixed-effects models were employed to account for repeated measurements and within-individual variations in our longitudinal data. Participant ID was included as a random effect for within-person correlations over time. Covariates included sex, age group, height z-score, waistline-to-height ratio, area of residence, total daily caloric intake, macronutrient composition, dietary quality scores, physical activity levels, parental education and occupation, annual household income, parental marital status, and family history of hypertension. Mixed-effects linear regression models assessed associations between eating timing patterns and cardiovascular indicators. FDR correction using the Benjamini-Hochberg method was systematically applied across all cardiovascular outcomes within each analysis domain. Mixed-effects logistic regression models evaluated relationships with elevated BP risk, estimating odds ratios (OR) and 95% CIs. All mixed-effects models underwent a comprehensive diagnostic evaluation. Restricted cubic spline analysis explored non-linear associations, with knots placed at the 10th, 50th, and 90th percentiles of the exposure distribution. Subgroup analyses stratified the sample by sex, age (<12 years/≥12 years), obesity status (normal weight/overweight or obese based on age- and sex-specific BMI cutoffs), dietary quality (above/below median Chinese Healthy Eating Index score), and residence (urban/rural). Sensitivity analyses examined robustness of findings using alternative thresholds: eating window duration (≤11 h vs >11 h, ≤13 h vs >13 h) and last meal timing (≤19:00 vs >19:00, ≤21:00 vs >21:00). Additional sensitivity analysis excluded multiply imputed data to assess the impact of missing data handling.
For the nested case-control component (n = 51), comprehensive multiomics analyses were performed to investigate molecular mechanisms. Proteomic analysis included differential expression analysis (DESeq2) to compare the three eating window groups (Extended EW, Late EW, and Early EW) (FDR-adjusted P < 0.05 considered significant). WGCNA was applied to identify protein co-expression modules, and module-trait relationships were calculated to associate these modules with cardiovascular parameters using Pearson correlation coefficients (Supplementary Fig. 13). Functional enrichment analyses (GO terms, KEGG pathways) of differentially expressed proteins were performed to identify significantly enriched biological pathways. Protein-protein interaction networks were constructed using the STRING database to visualize functional relationships among differentially expressed proteins. Lipidomic analysis employed a comprehensive two-stage analytical framework: (1) differential expression analysis to identify eating pattern-associated lipids, and (2) machine learning-based feature selection to identify blood pressure-related lipid biomarkers. In the first stage, differential expression analysis was performed using the limma package with empirical Bayes moderation to compare lipid profiles among eating timing groups. Extended EW served as the reference group representing the most circadian-misaligned pattern. Significantly different lipids from the Early EW vs Extended EW comparison were classified as up-regulated or down-regulated and carried forward for machine learning analysis. In the second stage, four optimized machine learning algorithms were applied to identify lipids predictive of elevated blood pressure status (elevated vs. normal): Random Forest using random Forest package with systematic parameter optimization (mtry: 1–15, ntree: 100–1500) and features selected by variable importance >0.05; Elastic Net using glmnet package with alpha optimization (0-1, increment 0.1) via 10-fold cross-validation and features selected at lambda.1se; SVM-RFE using the e1071 package with cost (0.001–1000) and gamma (0.00001-10) optimization employing adaptive recursive feature elimination; and XGBoost using the xgboost package with comprehensive hyperparameter tuning and 5-fold cross-validation. Lipid species selected by all four methods were defined as “core hub” biomarkers, while those selected by ≥ 3 methods were classified as “high confidence hub” biomarkers. Mediation analysis was conducted using the mediation package to examine the mediating effects of selected lipids on the relationship between eating patterns and blood pressure. Mediation analyses using bootstrapping methods (500 iterations) examined whether specific lipid species potentially mediated the relationships between eating timing patterns and cardiovascular outcomes. These analyses explored potential pathways based on the biological assumption of temporal ordering (eating patterns → metabolic changes → cardiovascular outcomes). All mediation models included identical covariate adjustment as primary analyses, and the proportion of effect mediated was calculated for each significant mediator. Following the same machine learning framework as the lipidomic analysis, proteins selected by all four methods were defined as “core hub” biomarkers. All statistical analyses were performed using R software (version 4.3.2) and SAS (version 9.4). Two-sided p-values < 0.05 were considered statistically significant, with appropriate corrections for multiple comparisons where applicable.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data generated in this study have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository under accession code PXD06646654. [https://www.ebi.ac.uk/pride/archive/projects/PXD066466]. The proteomics data are available under restricted access for research purposes that comply with the informed consent and ethical approval of this study; access can be obtained by submitting a data request to the corresponding author (Dr. Xiaohua Liang, xiaohualiang@hospital.cqmu.edu.cn) and signing a data use agreement. Requests will be responded to within 4 weeks. The raw lipidomics data generated in this study are protected and available under controlled access due to ethical restrictions concerning sensitive health information from minors and compliance with Chinese data protection regulations (Personal Information Protection Law of the People’s Republic of China). Access to raw lipidomics data requires: (1) submission of a research proposal to the corresponding author (Dr. Xiaohua Liang, xiaohualiang@hospital.cqmu.edu.cn); (2) approval by the Ethics Committee of Children’s Hospital of Chongqing Medical University; (3) execution of a data use agreement that specifies permitted uses, prohibits re-identification attempts, and requires data destruction upon project completion. Requests will be responded to within 4 weeks. The processed lipidomics data (lipid intensities and differential abundance results) are provided in the Source Data file. The summary statistics and processed datasets supporting the findings of this study are available in the Supplementary Information file and the Source Data file. De-identified participant-level data (clinical, anthropometric, and dietary data) are available from the corresponding author upon request, subject to review and approval of a research proposal and a signed data access agreement. Source data are provided with this paper.
Code availability
The source code, including scripts for cardiovascular data processing, mixed-effects modeling, differential expression analysis, machine learning feature selection, mediation analysis, and data visualization, is publicly available on GitHub at (https://github.com/LQ-doctor/Eating-Time-Patterns) and has been archived on Zenodo with DOI: (https://doi.org/10.5281/zenodo.17996048).
References
Martin, S. S. et al. 2024 Heart Disease and Stroke Statistics: A Report of US and Global Data From the American Heart Association. Circulation 149, e347–e913 (2024).
Jacobs, D. R. Jr. et al. Childhood cardiovascular risk factors and adult cardiovascular events. N. Engl. J. Med. 386, 1877–1888 (2022).
López-Bueno, R. et al. Global prevalence of cardiovascular risk factors based on the Life’s Essential 8 score: an overview of systematic reviews and meta-analysis. Cardiovasc. Res. 120, 13–33 (2024).
Gupta, C. C. et al. A time to rest, a time to dine: sleep, time-restricted eating, and cardiometabolic health. Nutrients 14, 420 (2022).
Raji, O. E., Kyeremah, E. B., Sears, D. D., St-Onge, M. P. & Makarem, N. Chrononutrition and cardiometabolic health: an overview of epidemiological evidence and key future research directions. Nutrients 16, 2332 (2024).
Ansu Baidoo, V. & Knutson, K. L. Associations between circadian disruption and cardiometabolic disease risk: A review. Obesity 31, 615–624 (2023).
Wehrens, S. M. T. et al. Meal timing regulates the human Circadian system. Curr. Biol. 27, 1768–1775.e1763 (2017).
Lewis, P., Oster, H., Korf, H. W., Foster, R. G. & Erren, T. C. Food as a circadian time cue - evidence from human studies. Nat. Rev. Endocrinol. 16, 213–223 (2020).
Liang, X. et al. The optimal time restricted eating interventions for blood pressure, weight, fat mass, glucose, and lipids: A meta-analysis and systematic review. Trends Cardiovasc. Med. 34, 389–401 (2024).
Chew, H. S. J. et al. Umbrella review of time-restricted eating on weight loss, fasting blood glucose, and lipid profile. Nutr. Rev. 81, 1180–1199 (2023).
Melkani, G. C. & Panda, S. Time-restricted feeding for prevention and treatment of cardiometabolic disorders. J. Physiol. 595, 3691–3700 (2017).
Yoshida, J., Eguchi, E., Nagaoka, K., Ito, T. & Ogino, K. Association of night eating habits with metabolic syndrome and its components: a longitudinal study. BMC Public Health 18, 1366 (2018).
Palomar-Cros, A. et al. Dietary circadian rhythms and cardiovascular disease risk in the prospective NutriNet-Santé cohort. Nat. Commun. 14, 7899 (2023).
Jamshed, H. et al. Effectiveness of early time-restricted eating for weight loss, fat loss, and cardiometabolic health in adults with obesity: a randomized clinical trial. JAMA Intern. Med. 182, 953–962 (2022).
Sjöholm, P., Pahkala, K., Davison, B., Juonala, M. & Singh, G. R. Early life determinants of cardiovascular health in adulthood. The Australian Aboriginal Birth Cohort study. Int. J. Cardiol. 269, 304–309 (2018).
Zou, M., Northstone, K., Perry, R., Johnson, L. & Leary, S. The association between later eating rhythm and adiposity in children and adolescents: a systematic review and meta-analysis. Nutr. Rev. 80, 1459–1479 (2022).
McMillen, I. C. & Robinson, J. S. Developmental origins of the metabolic syndrome: prediction, plasticity, and programming. Physiol. Rev. 85, 571–633 (2005).
Rothschild, J., Hoddy, K. K., Jambazian, P. & Varady, K. A. Time-restricted feeding and risk of metabolic disease: a review of human and animal studies. Nutr. Rev. 72, 308–318 (2014).
Nie, Z. et al. Effects of time-restricted eating with different eating windows on human metabolic health: pooled analysis of existing cohorts. Diabetol. Metab. Syndr. 15, 209 (2023).
Regmi, P. & Heilbronn, L. K. Time-Restricted Eating: Benefits, Mechanisms, and Challenges in Translation. iScience 23, 101161 (2020).
Petersen, M. C. et al. Complex physiology and clinical implications of time-restricted eating. Physiol. Rev. 102, 1991–2034 (2022).
Asher, G. & Sassone-Corsi, P. Time for food: the intimate interplay between nutrition, metabolism, and the circadian clock. Cell 161, 84–92 (2015).
BaHammam, A. S. & Pirzada, A. Timing matters: the interplay between early mealtime, circadian rhythms, gene expression, circadian hormones, and metabolism-a narrative review. Clocks Sleep. 5, 507–535 (2023).
Romero-Cabrera, J. L. et al. Chronodisruption and diet associated with increased cardiometabolic risk in coronary heart disease patients: the CORDIOPREV study. Transl. Res. 242, 79–92 (2022).
Wilkinson, M. J. et al. Ten-hour time-restricted eating reduces weight, blood pressure, and atherogenic lipids in patients with metabolic syndrome. Cell Metab. 31, 92–104.e105 (2020).
Bakhsh, J. A., Vu, M. H., Salvy, S. J., Goran, M. I. & Vidmar, A. P. Effects of 8-h time-restricted eating on energy intake, dietary composition and quality in adolescents with obesity. Pediatr. Obes. 19, e13165 (2024).
Hegedus, E. et al. Randomized controlled feasibility trial of late 8-hour time-restricted eating for adolescents with Type 2 diabetes. J. Acad. Nutr. Diet. 124, 1014–1028 (2024).
Cahill, L. E. et al. Prospective study of breakfast eating and incident coronary heart disease in a cohort of male US health professionals. Circulation 128, 337–343 (2013).
Kutsuma, A., Nakajima, K. & Suwa, K. Potential association between breakfast skipping and concomitant late-night-dinner eating with metabolic syndrome and Proteinuria in the Japanese population. Science 2014, 253581 (2014).
Ling, Y. et al. Varying definitions of Carotid intima-media thickness and future cardiovascular disease: a systematic review and meta-analysis. J. Am. Heart Assoc. 12, e031217 (2023).
Epure, A. M. et al. Risk factors during the first 1000 days of life for carotid intima-media thickness in infants, children, and adolescents: A systematic review with meta-analyses. PLoS Med. 17, e1003414 (2020).
Crook, M. A. Apolipoprotein H: its relevance to cardiovascular disease. Atherosclerosis 209, 32–34 (2010).
Giebelstein, J. et al. The proteomic signature of insulin-resistant human skeletal muscle reveals increased glycolytic and decreased mitochondrial enzymes. Diabetologia 55, 1114–1127 (2012).
Moriggi, M. et al. Skeletal muscle proteomic profile revealed gender-related metabolic responses in a diet-induced obesity animal model. Int. J. Mol. Sci. 22 (2021).
Becker, T. et al. Role of phosphatidylethanolamine in the biogenesis of mitochondrial outer membrane proteins. J. Biol. Chem. 288, 16451–16459 (2013).
Patel, D. & Witt, S. N. Ethanolamine and Phosphatidylethanolamine: Partners in Health and Disease. Oxid. Med. Cell Longev. 2017, 4829180 (2017).
Mejia, E. M. & Hatch, G. M. Mitochondrial phospholipids: role in mitochondrial function. J. Bioenerg. Biomembr. 48, 99–112 (2016).
Kulkarni, H. et al. Plasma lipidomic profile signature of hypertension in Mexican American families: specific role of diacylglycerols. Hypertension 62, 621–626 (2013).
Liu Qin, P. Z. & Shuanggui, Y. uan Association between body composition and blood pressure among school-age children in two southwestern provinces and one municipality in China: A cross-sectional survey. Chin. J. Evid. -Based Pediatr. 19, 360–366 (2024).
Liang, X. et al. The impact of dietary and sleep rhythms on blood pressure in children and adolescents: a cross-sectional study. Hypertens. Res. 47, 649–662 (2024).
Ren, Y. et al. Analysis of dietary patterns on cardiovascular risks in children: from a cross-sectional and a longitudinal study. Public Health 220, 35–42 (2023).
Liang, X. et al. The impact of PM2.5 on children’s blood pressure growth curves: A prospective cohort study. Environ. Int. 158, 107012 (2022).
Hui, F., Yin-Kun, Y., Jie, M. I. & Epidemiology, D. O. Updating blood pressure references for Chinese children aged 3-17 years. Chin. J. Hypertens. (2017).
He, J. et al. Serum proteome and metabolome uncover novel biomarkers for the assessment of disease activity and diagnosing of systemic lupus erythematosus. Clin. Immunol. 251, 109330 (2023).
Ashkarran, A. A. et al. Measurements of heterogeneity in proteomics analysis of the nanoparticle protein corona across core facilities. Nat. Commun. 13, 6610 (2022).
Blume, J. E. et al. Rapid, deep and precise profiling of the plasma proteome with multi-nanoparticle protein corona. Nat. Commun. 11, 3662 (2020).
Ferdosi, S. et al. Enhanced Competition at the Nano-Bio Interface Enables Comprehensive Characterization of Protein Corona Dynamics and Deep Coverage of Proteomes. Adv. Mater. 34, e2206008 (2022).
Xu, H. et al. Lysozyme modulates inflammatory responses to exacerbate the severity of rheumatoid arthritis. Int. Immunopharmacol. 152, 114427 (2025).
Commission, N. H. Screening for overweight and obesity among school-age children and adolescents. Beijing, China: National Health Commission (2018).
Wang, Q. et al. Associations between physical activity and hypertension in Chinese Children: a cross-sectional study from Chongqing. Front. Med. 8, 771902 (2021).
Feng, Y. et al. Protective effects of appropriate amount of nuts intake on childhood blood pressure level: a cross-sectional study. Front. Med. 8, 793672 (2021).
Ren, Y. et al. The correlation between nuts and algae-less diet and children’s blood pressure: from a cross-sectional study in Chongqing. Clin. Exp. Hypertens. 45, 2180024 (2023).
Chen, J., Luo, S., Liang, X., Luo, Y. & Li, R. The relationship between socioeconomic status and childhood overweight/obesity is linked through paternal obesity and dietary intake: a cross-sectional study in Chongqing, China. Environ. Health Prev. Med. 26, 56 (2021).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–d553 (2025).
Acknowledgements
We thank all the children, their families, and the staff of the participating schools in Sichuan, Chongqing, and Guizhou for their cooperation and support. We also acknowledge the medical teams from the respective local centers for their assistance in data and sample collection. This work was supported by the Natural Science Foundation Project of China (No. 82574113, 82373590, to X.L.), the Joint Key Project of Chongqing Science and Technology Bureau and Health Commission (No. 2025ZDXM008, to X.L.), the National Key Research and Development Program of China (No. 2024YFC2707605, to X.L.), the Program for Youth Innovation in Future Medicine from Chongqing Medical University (No. W0088, to X.L.), the General Project of Clinical Medical Research from National Clinical Research Center for Child Health and Disorders (No. NCRCCHD-2022-GP-01, to X.L.), the Young and Middle-aged Medical Outstanding Expert Project of Chongqing Municipal Health Commission (No. 78, to X.L.), the Major Health Project of Chongqing Science and Technology Bureau (No. CSTC2021jscx-gksb-N0001, CSTB2024NSCQ-MSX0180, 2023MSXM036, to X.L.), and the Intelligent Medicine Project of Chongqing Medical University (No. ZHYX202109, to X.L.). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
X.L. and Q.L. conceived and designed the study. Q.L., J.C., X.A., L.L.C. (Lanling Chen), L.J.C. (Lijing Chen), D.H., J.M., L.X., S.Y., and J.X. acquired the data. Q.L., X.A., and J.C. performed the statistical and bioinformatic analyses. Q.L. drafted the manuscript. X.L., B.B., and J.X. provided critical revision of the manuscript. X.L. supervised the project and acquired funding. All authors reviewed, edited, and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Angelo Pietrobelli and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Q., Chen, J., An, X. et al. Multiomics insights into eating time patterns and cardiovascular risk among Chinese children. Nat Commun 17, 1891 (2026). https://doi.org/10.1038/s41467-026-68617-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68617-8