Abstract
The thymus educates thymocytes through a selection process mediated by thymic epithelial cells (TECs). Recent advances have made the generation of T lymphocytes from induced pluripotent stem cells (iPSc) a promising therapeutic strategy. However, current approaches often fail to replicate the thymic niche, leading to impaired T cell generation. Here we address the production of functional mature iPSc-derived TECs supporting in vitro T cell generation. We optimize thymic lineage differentiation through an unbiased multifactorial experimental design. By modulating specific signaling pathways, we generate progenitors that mature into medullary and cortical TECs. Co-culture with primary hematopoietic progenitors in a 3D thymic organoid setup induces their differentiation into CD4+ and CD8+ T cells. Importantly, thymic organoids support multilineage differentiation, with dendritic cell populations also emerging. Thus, the presented thymic organoid model provides a practical platform for studying thymic cellular interactions and thymopoiesis in vitro, and opens further research perspectives towards cell-based therapies.
Similar content being viewed by others
Introduction
The thymus plays a crucial role in the establishment of central immune self-tolerance. Its main function is the generation of a diverse yet non-autoreactive T lymphocyte repertoire1. The thymus is subdivided into two lobes and structured into two principal niches, a peripheral cortical area surrounding a central medulla. Distinct cellular populations shape the specific microenvironment of these niches. Thymic epithelial cells (TECs) are crucial actors that regulate the maturation of developing T cells, the thymocytes2. Two main populations of TECs have been identified: the cortical TECs (cTECs) and the medullary TECs (mTECs), characterized in humans by the KRT8/18 and KRT5/14 cytokeratin expression3. Although KRT8/18 tags all epithelial cells, KRT5/14 are restricted to the medullary subpopulation. In addition, the expression of the surface marker CD205 is specific to cTECs2,4. Recently, single-cell studies have revealed a higher level of heterogeneity among TECs with the identification of new mTEC subpopulations that have various functional roles such as thymocyte homing, commitment, proliferation regulation and selection3,4,5. TECs are central mediators of thymocyte selection by enforcing a functional and non-autoreactive T repertoire through several steps. Thymocytes originate from CD34+CD7+ early-thymic progenitors (ETPs), which derive from circulating thymus-seeding progenitors (TSPs), and differentiate through several stages characterized by CD4 and CD8 expression6,7,8. Positive selection is mediated by cTEC at the double positive (DP) stage and leads to the selection of thymocytes with a functional T cell receptor (TCR)4. Negative selection filters out potential autoreactive thymocytes at the single-positive (SP) stage in the medulla9,10 and is mediated by HLA-DRhi mTECs (mTEChi) expressing a great diversity of peripheral tissue antigens (PTAs) under the regulation of AIRE4,11,12. Mature CD4 and CD8 T lymphocytes are called recent thymic emigrants and migrate to the periphery13. Recent research has shown that parallel intrathymic dendritic cell (DC) differentiation occurs from the same ETPs14. Interestingly, crosstalk with this thymic DC population has been shown to support thymocyte maturation and selection14,15.
Although the mechanisms supporting the differentiation of mTECs and cTECs remain elusive, recent studies have identified one or several bipotent thymic epithelial progenitor (TEP) populations capable of giving rise to complete and functional cortical and medullary compartments16,17,18. Studies have identified a bipotent TEP population exhibiting a cortical-like phenotype19,20. NOTCH21,22 and RANK-CD40-LTB23,24,25 signaling appear to play a crucial role in regulating TEC fate decisions. The regulation of thymus organogenesis has also seen remarkable progress in recent years, notably due to the publication of the single-cell RNA sequencing (scRNA-seq) thymic cell atlas at different stages of development26,27,28. TEP identity is notably characterized by the expression of FOXN1, the master regulator that induces the expression of the TEC-specific transcriptional program29,30,31,32. TEPs emerge from the third pharyngeal pouch endoderm (3PPE), a transient embryonic structure formed by the involution of the endoderm into the foregut tube between weeks 3 and 4 in humans33,34,35. A gene cascade involving TBX1, PAX9, and EYA1 has been identified as a crucial regulator of 3PPE formation. The 3PPE derives from the ventral pharyngeal endoderm, which originates from the anterior foregut endoderm (AFE)33,36. The regulation of fate acquisition in these structures has also shown advances at the single-cell level32,37,38.
Mimicking thymus organogenesis in vitro to generate TECs from embryonic (ESc) and induced pluripotent stem cells (iPSc) has been a long-standing objective over the last decade, using both reprogramming and direct differentiation approaches29,39,40,41,42,43,44. However, the complexity of regulatory mechanisms in the differentiation process and our limited understanding of in vivo thymic development have hindered the achievement of this goal45. Indeed, thymic differentiation often yields immature and non-functional cells with TEP identity that require grafting in vivo to complete maturation40,41,42. Moreover, the available protocols for iPSc differentiation into TEPs have not been optimized by unbiased systematic approaches able to robustly assess the effects of the supplemented factors at each stage46,47,48. Reliable methods for iPSc differentiation into functional, mature TECs in vitro and their long-term maintenance in a culture system that reproduces the thymic niche are yet to be developed. Such systems could enable the generation of iPSc-derived TECs, opening promising perspectives for regenerative medicine49. Indeed, treatment of congenital pathologies affecting TECs, such as DiGeorge’s syndrome or severe combined immunodeficiency (SCID) caused by gene mutations impairing TEC development, could benefit from corrected TECs to restore thymic function. Finally, iPSc-derived TECs could be used to improve ex vivo T cell generation to counteract the dramatic loss of thymus function linked to a weakened immune response during aging39,50.
In vitro T Lymphocyte generation has been greatly improved using feeder stromal cells such as OP9-DLL1 or MS5-hDLL4, which have been modified to express NOTCH ligands51,52,53. Formation of artificial thymic organoids has improved the maturation status of the obtained T cells, and partially replicated thymic function39,54. However, these setups only partially reproduce thymic selection, as they lack the self-antigen presentation ability of mTEChi.
Because of the complexity of thymic organoid co-cultures and iPSc thymic differentiation, due to the multiplicity of factors involved, precise optimization of the protocols is difficult to achieve. Here, we used the optimal design of experiments (DOE), a systematic statistical framework for experimental design, to collect the most information with the fewest experiments. A transcriptomics readout was used to optimize iPSc thymic differentiation based on recent single-cell datasets of thymus organogenesis. We propose a two-week protocol that efficiently yields TEPs expressing thymic identity markers. To overcome the difficulty of maturing TEPs ex vivo, we develop a 3D co-culture system with CD34+CD7+ thymic progenitor cells that have not yet passed the T lineage commitment checkpoint. These cells will be referred to as ETPs in a broader sense, encompassing TSP2 cells arriving in the thymus, ETPs and their immediate progeny at the double negative (DN)1 stage14,55,56. We set up a fibrin hydrogel air-liquid interface culture that results in the formation of human thymic organoids (hTOs). hTOs could be maintained for up to 5 weeks, and display a phenotype similar to mature TECs, particularly HLA-DRhi medullary-like TECs, which exhibit higher AIRE expression than in a 2D setup. TEC-like cells and thymocytes show some degree of co-localization in the organoids, and hTOs demonstrate the ability to drive the maturation of diverse progenitors into SP CD4+ and CD8+ T lymphocytes. Further characterization of hTO products by scRNA-seq confirms the mature state of the generated T lymphocytes. Finally, we report the secondary differentiation of ETPs into the dendritic cell lineage in line with recent in vivo observations, thus demonstrating a complex in vitro organoid model capable of both cell maturation and multilineage potential.
Results
Optimization of directed iPSc differentiation toward a TEC fate using DOE-based factor screening and bulk transcriptomics
To set up an optimized differentiation protocol of iPSc into TEPs, we followed an approach of directed differentiation that recapitulates the events of thymic organogenesis. Recently, the understanding of thymic identity acquisition and its underlying regulation at the cellular and molecular level has greatly improved through the use of single-cell-omics approaches32,37,38,57. Definitive endoderm (DE) undergoes anteriorization into anterior foregut endoderm (AFE) from which emerges the third pharyngeal pouch endoderm (3PPE), a transitory structure that gives rise to TEPs (Fig. 1a). We aimed to recapitulate these events in vitro by modulating the main pathways involved in thymic organogenesis, such as BMP, WNT, Hedgehog (SHH) and FGF.
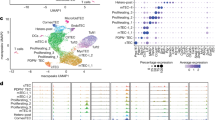
a Differentiation itinerary of the thymic epithelium showing DE (Definitive Endoderm), AFE (Anterior Foregut Endoderm), 3PPE (Third Pharyngeal Pouch Endoderm) and TEP (Thymic Epithelial Progenitor) stages (Created in BioRender. Guillonneau, C. (2025) https://BioRender.com/r5v7uln). b Factors tested in DOE (abbreviations), their associated pathways and the timings corresponding to each tested stage transition (D5-D7: DE-to-AFE, D7-D11: AFE-to-3PPE and D11-D13: 3PPE-to-TEP). Plackett–Burman designs were used to estimate factor effects on differentiation at the three transitions, with two dose levels (−1/+1) per factor. c UMAP representations of early (Left) and late (Right) pharyngeal development scRNA-seq reference datasets, with clusters corresponding to stages of the thymic differentiation trajectory shown in color. Reference datasets: Han et al. (E8.5–E9.5) and Magaletta et al. (E9.5–E12.5). d Bulk RNA-seq results for each sample (vertical) treated with combinations of factors tested in DOE in four experiments (D5-D7, Han E8.5; D7-D11, Han and Magaletta E9.5; D11-D13, Magaletta E11.5/12.5) showing expression scores of marker genes for pharyngeal development clusters (horizontal). Cluster names corresponding to the thymic differentiation trajectory are highlighted in bold. The upper part of each heatmap shows the factor combinations applied, with high doses highlighted in orange. Statistical significance was assessed by ANOVA (Supplementary Fig. 3). Significant factors are shown in red or blue, indicating those to be supplemented or excluded in the optimized protocol. Han et. al. dataset: D5 to D7: p = 0.02338 (Noggin); 4.246e-05 (IWR1); 0.05526 (LY3); 4.345.e-07 (RA)/D7 to D11: p = 5.233.e-05 (CHIR99); 0.0002875 (FLI06); 0.0015895 (IWR1). Magaletta et. al. dataset: D7 to D11: p = 1.877.e-05 (CHIR99); 1.877.e-05 (FLI06); 0.0003179 (IRW1)/D11 to D13: dots indicate suggestive p values, p = 0.089927 (BMP4); 0.079926 (IGF1); 0.086185 (RANKL). e Summary of pathway modulation: black, DOE-identified pathways to be activated (+) or inhibited (−); blue, pathways neutral in DOE but enhancing FOXN1 expression (BMP/FGF) or proliferation (FGF/EGF) at the TEP stage; green, the FGF pathway (not tested in DOE, post-DOE), activating FOXN1 at the TEP stage. DE Definitive Endoderm, AFE Anterior Foregut Endoderm, 3PPE Third Pharyngeal Pouch Endoderm, TEP Thymic Epithelial Progenitor. See corresponding factor names in (b) (Created in BioRender. Guillonneau, C. (2025) https://BioRender.com/ht4edy0). f Summary of the optimized protocol with full factor list, doses and exposure windows, DE Definitive Endoderm, AFE Anterior Foregut Endoderm, 3PPE Third Pharyngeal Pouch Endoderm, TEP Thymic Epithelial Progenitor.
The exit from the pluripotency state and DE induction have been thoroughly studied58, and the process of iPSc differentiation into DE has been precisely characterized. We performed this step by stimulating Nodal through Activin A exposure in combination with a 24-h pulse of WNT through CHIR99. The differentiating cells showed a peak of expression for the DE marker gene SOX17 at days (D) 3–5 (Supplementary Fig. 1a). In addition, a vast majority of cells stained positive for SOX17 and another DE marker FOXA2 at D5 (Supplementary Fig. 1b). These results reflect the high efficiency of the iPSc to DE differentiation, which does not require further optimization.
We applied DOE to test the effect of a series of factors that impact differentiation pathways and are known to be potentially active in the transitions toward AFE, 3PPE and TEP. The modulation of RA, TGFb, BMP, and WNT pathways on definitive endoderm anteriorization between D5 and D7 was first studied. Then the effect of FGF8, NOTCH, Hedgehog (SHH), BMP and WNT modulation of 3PPE was assessed between D7 and D11. Lastly, the effect of BMP, FGF10, EGF, IGF1, and RANKL modulation on TEP fate induction between D11 and D13 was investigated (Fig. 1b). These pathways were selected based on studies of thymic organogenesis regulation45,59,60,61,62 and previous thymic differentiation of hESc40,41. A Plackett-Burman combinatorial screening design was performed for each of the three transitions by testing multiple combinations of factors with two concentration levels (Supplementary Fig. 1c, d). The choice of this experimental design is justified by its ability to test a maximum number of factors within a limited number of runs, which is adapted to the experimental constraints of differentiating iPSc.
Using a low-dimensionality readout, such as measuring the expression of one or a few marker genes, can yield imprecise results due to a marked lack of specificity and gene expression variability. To overcome this weakness and robustly assess factor effects, we performed bulk RNA-seq on the DOE samples and compared their transcriptome to two scRNA-seq atlases of pharyngeal development. The atlas by Han et al.37 corresponds to early stages of development from E8.5 to 9.5, covering the AFE and 3PPE transitions. The second by Magaletta et al.32 focuses on later stages of development from E9.5 to 12.5, including the 3PPE and TEP transitions. Raw data reanalysis from these two datasets, applying uniform manifold approximation and projection (UMAP) (Supplementary Fig. 2a), led to the delineation of transcriptomic maps of the thymic differentiation trajectory from E8.5 to 9.5 (Han) and E9.5 to 12.5 (Magaletta). This was achieved by studying the pattern of expression of known genes specific to thymic precursor cell populations (Fig. 1c and Supplementary Fig. 2b). We then scored the transcriptome similarity of each DOE sample to the different cell populations at each relevant stage of pharyngeal development, i.e., E8.5 for anterior foregut formation, E9.5 for pharynx formation and E11.5/E12.5 for TEP transition, by using the mean expression of the top marker genes of each cell population (Supplementary Data 1 and 2).
The differentiation conditions for AFE (D5 to D7) were optimized using the Han dataset. Unsupervised hierarchical clustering of DOE iPSc-derived samples and reference progenitor cell populations identified two clusters of samples with high expression of anterior foregut markers (Fig. 1d, Top Left). These samples also show strong expression of markers for developmentally close populations (lateral foregut and ventral foregut) and low expression of midgut and dorsal lateral foregut markers, confirming DE anteriorization. To identify the effect of the tested factors on anterior foregut induction, we computed their statistical main effects using factorial ANOVA on the mean expression of the anterior foregut gene markers. RA, NOGGIN and LY3 supplementation, as well as the absence of IWR1, had the most impact on DE anteriorization (Supplementary Fig. 3). Hence, BMP inhibition by NOGGIN and, to a lesser extent, TGFβ inhibition by LY3 promoted anteriorization, whereas WNT inhibition by IWR1 was globally detrimental at this step.
We applied a similar strategy to the 3PPE transition (D7 to D11) and found a cluster of DOE samples with the highest expression of pharynx markers (Fig. 1d, Bottom Left). ANOVA of the tested factors on the mean expression of the pharynx gene markers identified a statistically significant effect for the addition of CHIR99, as well as the deprivation of FLI06 and IWR1 (Supplementary Fig. 3). We then used the Magaletta dataset as reference, as it also covers the transition to 3PPE with immature pharynx 1 and 2 populations that correspond to the developmental stage of the pharynx population from the Han dataset. We confirmed the positive effect of CHIR99, as well as the negative effect of FLI06 and IWR1 on 3PPE formation (Fig. 1d, Top Right, and Supplementary Fig. 3). Hence, WNT activation by CHIR99, in contrast to its inhibition by IWR1, greatly promotes pharyngeal formation of the 3PPE, whereas NOTCH inhibition by FLI06 is detrimental. Interestingly, contrary to the Parent et al. study40, we observed no effect of Hedgehog inhibition by cyclopamine at this stage. Similarly, BMP4 and FGF8 supplementation had no significant effect on pharyngeal induction, despite being commonly used in previous studies40,42,43. We therefore specifically tested these two conditions by measuring D15 FOXN1 expression by qPCR to directly quantify differentiation efficiency, rather than using the intermediate 3PPE as a proxy. Interestingly, we observed a significantly higher expression of FOXN1 in TEP samples exposed to FGF8 and BMP4 at the earlier 3PPE stage (Supplementary Fig. 4a). Thus, the combination of these two factors at this stage seems beneficial for later TEP differentiation, but does not directly affect differentiation efficiency at 3PPE, thereby justifying keeping them in the protocol.
Finally, we optimized the differentiation conditions for the transition to TEP (D11 to D13), leveraging the Magaletta et al. dataset, which covers late stages of pharyngeal development, including early cTEC and mTEC, even though no bona fide transitory TEP population was identified. Since cTECs are the natural and most direct product of TEP differentiation, we focused on them as the target population for this step of differentiation. Hierarchical clustering identified a cluster of DOE samples that had the highest expression of cTEC markers, along with markers of its closely related mTEC and thyroid populations (Fig. 1d, Bottom Right). Although our statistical approach could not discriminate cTEC from mTEC or thyroid cells for their resemblance to TEPs, we took this population cluster as a distant proxy for TEPs. The ANOVA on the mean expression of cTEC markers showed that the factors with the most impact were the addition of BMP4 and IGF1, as well as the deprivation of RANKL. We also chose to retain FGF10 and EGF since we observed a direct positive effect of these two factors on the growth rate of the differentiation end product (Supplementary Fig. 4b).
Finally, we performed a post-DOE optimization considering recent publications on thymic differentiation, highlighting the benefits of FGF8 supplementation for AFE induction45. We confirmed similar results in our protocol by measuring a significant increase in FOXN1 expression at the end of differentiation in samples exposed to FGF8 from D4 to D6 (Supplementary Fig. 4c).
Results of the DOE optimization are synthesized in Fig. 1e, describing selected inhibited (−) or activated (+) pathways. We summarized our iPSc differentiation protocol to TEPs in a table displaying doses and duration of differentiation factor supplementation (Fig. 1f).
Characterization of the differentiation product demonstrates efficient differentiation into TEPs
To follow the thymic differentiation process at the transcriptomic level, we sequenced samples at multiple time points across our differentiation protocol by bulk RNA-seq. Human primary TECs, sorted as CD45-EPCAM+ and originating from thymic samples of pediatric patients undergoing cardiac surgery, were included to assess the state of maturation of the differentiation product. iPSc samples were included as the starting point of the differentiation trajectory. We performed differential gene expression analysis of each time-point sample against D0 iPSc and identified nine gene modules enriched at distinct differentiation stages and comprising known markers of thymic differentiation stages, such as SOX17 (DE), HOXA3 (AFE), EYA1 (3PPE), or EPCAM (TEP) that hit in the same module as CXCL12, whose expression is known to be a common feature of cTECs and which attracts DP thymocytes for positive selection63 (Fig. 2a, Left). However, markers associated with a more mature TEC phenotype, such as cortical proteasome components encoded by PSMB11 and PRSS16, or genes involved in antigen presentation (HLA-DRA, CD80, and CD86), were not differentially expressed in late differentiation cultures compared to D0 iPSc at the significance threshold of adjusted p-value = 0.01, consistent with their immature thymic epithelial state. As a control, we showed that, apart from PRSS16, these TEC marker genes were overexpressed in primary TECs compared to D0 iPSc (Supplementary Fig. 5a, b). Furthermore, a direct comparison between primary TECs and TEPs highlighted the upregulation in TECs of maturation markers such as HLA-DRA, HLA-DRB1, and HLA-DQB2, and chemokines (CCL3, CCL19, and CCL25) together with CCR9, while TEPs were enriched for PRSS16, PRSS23, Integrins (ITGA5, ITGA6, and ITGAV), and KRT19, consistent with the acquisition in TECs of a thymic functional program, distinct from the immature epithelial state of TEPs (Supplementary Fig. 5c). To determine the biological processes enriched in the nine gene modules, Gene Ontology (GO) terms were associated using ClusterProfiler (v3.18.1). Ordering GO terms by time of differentiation showed a shift from the pluripotency state and cell proliferation to embryogenesis and organ morphogenesis (Fig. 2a, Right). GO terms in late differentiation samples and primary TEC controls were associated with lymphocyte interaction and immune response regulation, indicating the enrichment of our differentiation product in TEC-associated genes. A detailed list of GO terms is available in Supplementary Fig. 6. Together, these results show the progressive loss of pluripotency and acquisition of TEC fate through modulation of the pathways we identified using the DOE-based screen.
a Bulk RNA-seq across iPSc-to-TEP differentiation, with samples from D0 (iPSc) to D14–D22 (TEPs). Heatmap shows log2-normalized expression of genes grouped into nine co-regulated modules defined by differential expression versus D0 iPSc (Left). Primary human TECs were included as controls. GO terms enriched in each gene module were identified using ClusterProfiler (v3.18.1) and arranged by shared gene similarity (Right). The network was plotted in Cytoscape (v3.8.x) with orientation forced along the differentiation timeline, yielding a graph of significant biological processes enriched at each stage. DE Definitive Endoderm, AFE Anterior Foregut Endoderm, 3PPE Third Pharyngeal Pouch Endoderm, TEP Thymic Epithelial Progenitor. (Created in BioRender. Guillonneau, C. (2025) https://BioRender.com/cx6622l). b Cell-type deconvolution of the D16 differentiation end product against the Magaletta et al. scRNA-seq atlas of pharyngeal organogenesis. Deconvolution method: MuSiC (MuSiC_prop(), default parameters), outputs shown as estimated proportions per reference cell type (n = 6, from independent differentiations, statistics: mean + standard deviation). Source data are provided as a Source Data file.
To finally assess the efficiency of the optimized TEP differentiation protocol, we performed a deconvolution analysis of bulk RNA-seq data from the end product against the Magaletta dataset. This approach identified cTEC as the closest reference population to our differentiation product, confirming that we generated TEPs with high efficiency (Fig. 2b). In addition, we showed that these cells express PAX9, associated with early pharyngeal development and indicative of pre-thymic identity, as well as FOXN1, which is progressively upregulated during thymic epithelial lineage commitment and maturation consistent with acquisition of a thymic epithelial fate (Supplementary Fig. 7a). These cells are generated in large quantities, yielding ~2.1 million cells from 100,000 iPSc per P12 well at the end of differentiation (Supplementary Fig. 7b). We then benchmarked our protocol against two state-of-the-art protocols from Sun et al.41 and Parent et al. studies, by measuring the expression of PAX9 and FOXN1. We found that our protocol yielded cells expressing significantly higher levels of PAX9 and FOXN1 (Supplementary Fig. 7c). To test whether these results were due to protocol overfitting to our iPSc cell line LON71 or represented a true optimization of the biological process, we differentiated two other iPSc lines (MIPS203 and LON80) and found no significant differences in FOXN1 and PAX9 expression, except for the MIPS203, which showed lower FOXN1 expression (Supplementary Fig. 7d, e).
CD205+ TEPs are the main product of differentiation, exhibit mTEC fate orientation and express NOTCH functional ligands
TEPs exhibit significant cTEC/mTEC lineage plasticity, and studies have shown that these progenitors express CD20532,64. Thus, we asked whether the iPSc-derived TEPs we generated expressed the CD205 surface marker and in what proportion. We confirmed by flow cytometry that almost all cells expressed the epithelial marker EPCAM and found that a large majority of them were positive for CD205, as shown in Fig. 3a, Left. Controls also attested to the specificity of the staining (Fig. 3a, Middle). To generalize this result, we performed a series of independent differentiations and assessed the proportions of EPCAM+ and CD205+ cells at the TEP stage and the earlier 3PPE stage for comparison. We found that the median percentage of cells positive for EPCAM at the TEP stage was close to 100%, and ~70% expressed the additional CD205 (Fig. 3a, Right). As expected, CD205 expression was not detected at the 3PPE stage, in contrast to EPCAM, which marks the epithelial cell fate.
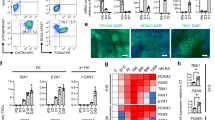
a Flow cytometry quantification of differentiation end product TEPs expressing EPCAM together with the TEP/cTEC lineage marker CD205. HLA-DRA staining was included to assess MHCII expression. Source data are provided as a Source Data file. b Heatmaps showing mean-centered normalized gene expression of thymic crosstalk genes (Left) and NOTCH ligand genes (Right) in unsorted TEPs and CD205+ TEPs. D0 iPSc and primary TECs are included as controls. Source data are provided as a Source Data file. c Cell type deconvolution of CD205+ sorted TEPs from n = 6 independent differentiations against the Magaletta et al. scRNA-seq atlas of pharyngeal organogenesis (statistics: mean + standard deviation). Source data are provided as a Source Data file. d Normalized expression of cTEC cluster marker genes from the Magaletta dataset in unsorted TEPs and CD205+ TEPs. Source data are provided as a Source Data file. e Confocal microscopy immunofluorescence of DLL4 and DLL1 in TEPs (n = 4). The magenta signal is shown in red and the green channel intensity was slightly enhanced for visibility in the merge. HEK293 cells were used as a negative control for DLL4 and DLL1 expression (n = 2). Nuclei were stained with DAPI. Normalized expression of DLL4 and DLL1 is shown for TEPs, CD205+ TEPs and primary TECs as a positive control (Right).
We then asked whether the bona fide CD205+ TEPs are equipped to sense maturation signals from developing thymocytes and to provide signals to promote their differentiation. To this end, we performed RNA-seq of sorted CD205+ TEP populations and analyzed the expression of a set of genes encoding proteins involved in the responsiveness of maturing TECs to signals provided by the developing thymocytes as well as in promoting thymocyte development. We identified a set of genes showing preferential expression in CD205+ TEPs and coding for proteins involved in the classical and alternative NF-κB signaling pathway, with RELA and RELB constituting the NF-κB molecular complexes, TRAF5 mediating signal transduction and RANK (TNFRSF11A) receiving RANKL signals from thymocytes and activating the NF-κB pathway (Fig. 3b, Left). Activation of the alternative NF-κB pathway is crucial for acquiring an mTEC fate and features that enable mature mTECs to sustain self-antigen expression and presentation to developing thymocytes by MHCII molecules. Thus, the CD205+ TEP population appears preferentially oriented toward an mTEC fate. We then performed a deconvolution analysis against the Magaletta dataset32 and observed that 17% of these cells with inferred TEC signature were medullary oriented (Fig. 3c). Moreover, comparison of classical cTEC marker expression between CD205+ sorted and unsorted TEPs showed lower expression in the CD205+ TEPs (Fig. 3d). We identified several NOTCH ligands, including DLL4, JAG1, and JAG2, whose coding genes showed robustly higher expression in CD205+ TEPs compared to iPSc, as well as DLL1, which was expressed at comparable levels in the two populations due to its role in maintaining pluripotent stem cells, explaining its high expression in iPSc65,66 (Fig. 3b, Right). Activation of the NOTCH pathway is crucial for promoting thymocyte development. To assess the expression of the key NOTCH ligands DLL1 and DLL4 at the protein level, we performed immunofluorescence experiments on D14 TEPs (Fig. 3e, Left), which revealed consistent surface staining supporting their potential to deliver NOTCH signaling to developing thymocytes. This was further consistent with transcript detection of DLL1 and DLL4 in our RNA-seq data (Fig. 3e, Right).
Interestingly, we observed that all CD205+ TEPs have an enhanced expression of KRT19, which has recently been identified as a marker of a separate mTEC-fated progenitor32,64,67. Together, our findings indicate that the CD205+ TEPs we generated are likely capable of sensing signals that activate medullary differentiation programs in TEPs.
Expression of mature and medullary TEC markers in TEP-derived cells after 2D co-culture with ETPs
While the maturation into functional TECs in vitro has proven challenging, with most studies grafting TEPs into mice40,41,42,43, we aimed to achieve full TEP maturation in vitro. Thymocytes have been identified as a crucial source of signaling that regulates thymic medulla formation and maturation, especially for the mTEChi compartment68.
Therefore, we investigated the capacity of early thymocytes to promote TEP maturation. ETPs from human thymus samples were harvested and sorted by flow cytometry using the gating strategy CD3-CD4-CD8-CD14-CD19-CD56-(Lin-) CD34+CD7+ (Supplementary Fig. 8). This rare population constitutes multipotent hematopoietic progenitors upstream of T-cell differentiation7,56. We characterized the sorted cell population by flow cytometry and validated their ETP phenotype (Supplementary Fig. 9a). These cells were negative for CD3, double negative for CD4 and CD8, and positive for CD44, a marker of early progenitor cells uncommitted to the T-cell lineage shared by TSP2, ETP, and DN1 cell populations55. Of note, the purity of the ETP sorting strategy may vary, likely due to the heterogeneity of early thymocyte populations in donors and experimental variability. This is exemplified in Supplementary Fig. 9b, where a minor CD44− population, potentially comprising committed cells, remains.
To test whether the TEPs we generated have the potential to phenocopy TECs from the medullary lineage in addition to the cortical lineage, we performed 2D co-cultures of TEPs and primary ETPs using 20,000 TEPs and 2500 ETPs per well, supplementing the medium with cytokines stimulating TEC and T-cell growth and maturation, namely, RANKL, IL7, SCF, and FLT3L56,69 (Fig. 4a). We selected these factors due to their well-established role in supporting early thymocyte survival and expansion, to complement the endogenous secretion by differentiating TECs. Recent advances indicate that RANKL drives the expansion of the already committed mTEP population64, which is compatible with a beneficial role of RANKL in increasing medullary cell growth. Interestingly, from a seemingly random distribution shortly after co-culture setup, ETPs were nearly exclusively located at the surface of the large, differentiated epithelial cells after one week (Fig. 4b). Quantification of the co-localized cells confirmed a significant increase after one week of co-culture (Fig. 4c). Given the ability of TECs to interact with thymocytes and even to incorporate them, forming thymic nurse cell complexes, it is possible that this physical interaction is concurrent with cell-cell signaling during thymopoiesis. However, we did not investigate further whether thymocytes were incorporated by TECs or merely located at their surface.
a 2D co-culture of D13 iPSc-derived TEPs with primary human ETPs in medium supplemented with RANKL, SCF, IL7 and FLT3L (Created in BioRender. Guillonneau, C. (2025) https://BioRender.com/at0ys62). b Bright-field microscopy images of TEPs co-cultured with ETPs after 24 h and 1 week. Two independent experiments are shown. c Quantification of ETP/TEP co-localization using the FIJI software. The co-localization ratio was defined as the number of thymocytes located within the membrane perimeter of adherent epithelial cells divided by the total number of thymocytes. *p = 0.0159 (two-sided Wilcoxon rank sum exact test), n = 2 (two independent experiments) for both time-points (statistics: mean + standard deviation). Source data are provided as a Source Data file. d Confocal microscopy immunofluorescence of mature and medullary TEC markers in D15 TEPs co-cultured with ETPs (n = 4). Nuclei were stained with DAPI.
Finally, immunofluorescence revealed surface expression of KRT5 and KRT14 in a subset of cells, highlighting the emergence of medullary features in a minor subpopulation (Fig. 4d). Indeed, in humans, KRT5 and KRT14 are mTEC markers, with KRT5 broadly expressed across all mTECs and KRT14 more restricted to mTEC(I), which represent an early stage of mTEC differentiation, as shown in the human TEC atlas by Park et al. (https://cellatlas.io/studies/thymus-development/dataset/132/cherita) (Supplementary Fig. 10a). Since CLDN3 appears to be more restricted to the mature medullary compartments (mTECII–mTECIII) in the human TEC atlas (Supplementary Fig. 10a), we performed CLDN3 staining to assess medullary maturation and observed expression with a typical focal pattern in a sparse subset of cells (Fig. 4d). We counted CLDN3 positive cells in a larger field IF and found that they comprised 14% of the total cell population, indicating heterogeneity and relatively low maturation of TECs in 2D co-culture (Supplementary Fig. 10b). In addition, we found the presence of a few cells positive for HLA-DR expression, some of them showing co-expression with CLDN3 (Fig. 4d). Since HLA-DR positive cells were sparse in immunofluorescence assays, we performed FACS analysis and identified a minority of 7% of the epithelial cells (EPCAM+) being HLA-DR+ (Supplementary Fig. 10c). Together these results show that the generated TEPs have the potential to differentiate into TEC-like cells, with a small fraction expressing markers associated with maturation (HLA-DR) and medullary fate (CLDN3), thereby strengthening the potential of the TEPs we generated to respond to signals driving medullary differentiation, albeit with a low efficiency in 2D co-culture.
Finally, we observed fewer CD45+ ETP-derived cells in the 2D co-culture than expected, potentially due to cell death or insufficient stimulation during culture. In addition, we detected minimal differentiation into CD3+ cells, most of which were CD4-CD8+ cells, suggesting that the 2D co-culture supports only very limited ETP maturation (Supplementary Fig. 11).
A 3D thymic organoid system further promotes medullary differentiation
Classical monolayer culture is known to be unsuitable for TEC maintenance, leading to a rapid loss of TEC functional markers69. Considerable progress has been made recently, with the development of 3D culture systems in hydrogels and the setup of artificial thymic organoids52,70,71,72. We developed a human thymic organoid (hTO) co-culture system by reaggregating human primary ETPs with the iPSc-derived TEPs and seeding the cell mass into a fibrin hydrogel (Fig. 5a). Organoids were cultured at air-liquid interfaces on cell inserts, using the same XVIVO10 base culture medium, as other tested media, such as DMEM, did not support comparable cell growth. As for the 2D co-culture, we supplemented this medium with RANKL, IL7, SCF and FLT3L. After 24 h, organoids exhibited a spheroid shape with a crown-like structure of early thymocytes (Fig. 5b, Left). Organoids showed signs of growth from D4 post-seeding, with an increase in total size and the formation of cellular projections into the hydrogel (Fig. 5b, Right).
a Reaggregation of iPSc-derived TEPs with primary human ETPs to form hTOs (human Thymic Organoids) cultured in a 3D fibrin hydrogel at an air-liquid interface in medium supplemented with RANKL, IL7, SCF, and FLT3L (Created in BioRender. Guillonneau, C. (2025) https://BioRender.com/apk8n61). b Bright-field images of hTOs (human Thymic Organoids) at 24 h and D4 showing a compact spheroid core with a peripheral cellular crown (identical magnification) (n = 12 from independent differentiations and ETP donors). c scRNA-seq of EPCAM+CD45− cells from D7 organoids with UMAP embedding and cell-cycle scoring (10x Genomics; Seurat workflow). d Module score for a TEP gene set: PDPN, PAX9, IL7, EBF1, FN1. e Integration of the EPCAM+CD45− scRNA-seq dataset (Clusters 1, 2, and 3) with the Bautista human TEC atlas; with reference cells in gray. f Cell type deconvolution of TEPs and CD205+ TEPs against the EPCAM+CD45− scRNA-seq dataset. g Distribution of the Tau tissue-specific variability index within clusters 3 and 1. Tau was computed from GTEx tissue expression data; TRA enrichment was assessed on the top-200 cluster-specific genes. Source data are provided as a Source Data file. h Assessment of the TRA repertoire in Cluster 3, showing enrichment for brain antigens. Source data are provided as a Source Data file.
We first validated the increased FOXN1 expression by qPCR throughout differentiation, culminating in hTOs (Supplementary Fig. 12a). We then leveraged the ease of harvesting hTO cells with low mortality in the early growth phase to perform scRNA-seq experiments and monitor early TEP differentiation at D7. We isolated the EPCAM+CD45- population from D7 organoids, likely corresponding to TEP-derived cells, and performed scRNA-seq. Analysis of the generated data revealed one main cluster 1, a smaller cluster 2 close to the former and composed of cells showing cycling features, and a distant smaller cluster 3 (Fig. 5c). To track the fate of the TEP population in the organoids, we selected a module of five common TEP marker genes, i.e., PDPN, PAX9, IL7, EBF1, and FN1, from the Magaletta dataset and Farley et al.64. We computed the expression score of this gene module and found strong expression in the main cluster 1 and in the cycling cells, revealing the strong TEP identity of these cells. In contrast, the more distant cluster 3 and a fraction of cycling cells showed low TEP identity, indicating that these cells may stem from the iPSc-derived TEPs in the organoids (Fig. 5d). Monocle3 (v1.3.3) trajectory analysis, rooted in cycling cells, revealed a fully connected graph where all cells were developmentally linked, with two trajectories: one toward high and another toward low TEP identity cells (Supplementary Fig. 12b).
To identify the nature of the low TEP identity cluster, we integrated the organoid scRNA-seq data with the human TEC atlas from Bautista et al.3. Remarkably, we found that the low TEP identity cells from cluster 3 were located within the medullary region of the UMAP, in close vicinity of mTEChi, whereas the high TEP identity cells from cluster 1 lay in the cortical region of the graph next to immature cycling cTECs, without clustering perfectly with the latter (Fig. 5e). This indicates that the induced TECs share transcriptomic similarities with immature cTECs, but are not sufficient per se to confirm a definitive cTEC identity. In addition, the absence of detected expression of the functional markers HLA-DRA, CD80, or CD86 indicates that these cells remain at an immature stage.
To confirm that the low TEP identity cells arose after organoid formation rather than being already present in iPSc-derived TEPs, we performed a deconvolution analysis of the bulk RNA-seq data from iPSc-derived TEPs and sorted CD205+ TEPs against the three clusters identified in organoid scRNA-seq. We found that the main high TEP identity cluster 1 was the closest to iPSc-derived TEPs, which were negative for the mixture of cells with low TEP identity (Fig. 5f). Furthermore, we also showed that sorted CD205+ TEPs correspond to the high TEP identity clusters. Finally, we assessed the enrichment of TRA gene expression among the genes that are most specific to the low TEP identity cluster 3. To this end, we computed for each gene, the tissue specificity index Tau73, by implementing its calculation algorithm in a Shiny R application that we developed to parse the comprehensive human protein atlas database, including the human gene tissue expression database, GTEx (https://www.proteinatlas.org/about/download). We found a shift toward higher Tau values in the low TEP identity cluster 3 in comparison to cluster 1, with almost 40% of the top 200 genes classified as tissue specific (Tau > 0.7) (Fig. 5g). In addition, most identified TRA genes were of brain origin (Fig. 5h), consistent with previous findings that neural programs are activated early during the immature mTEC stage in differentiation3.
Together our findings show that upon 3D interaction with primary ETPs, the iPSc-derived TEPs initiate differentiation toward a medullary fate and exhibit TRA-favored expression in the early phase of hTO growth.
TEC maturation in the late phase of organoid differentiation
hTOs could be maintained for several weeks in the 3D culture system, reaching 5 mm in size. At D11, organoid morphology evolved from a spheroid to a complex structure with multiple projections colonizing the gel (Fig. 6a). To assess the level of medullary differentiation and the interaction of these cells with thymocytes (CD45+) in the late phase of organoid development, we performed confocal microscopy at D28 (Fig. 6b and Supplementary Fig. 12c). We observed extensive KRT5 staining in the cellular projections indicative of medullary-oriented cells. Importantly, we confirmed that KRT5 was not already present at the TEP stage (Supplementary Fig. 12d). Physical interactions between thymocytes and TECs could also be observed in the organoids with some degree of colocalization, even though only a slight majority of thymocytes (µ = 65%, n = 3) were located around organoid projections. Interestingly, distant z-slices from the organoid, in the hydrogel volume, showed dispersed thymocytes, highlighting their ability to freely circulate in the gel, which was originally devoid of cells during its casting (Supplementary Fig. 12c, Left). This property was also observed for KRT5+ cells, which migrated far from the initial seeded micromass (Supplementary Fig. 12c, Right).
a Bright-field images of hTOs (human Thymic Organoids) at D11 and D25 showing outgrowths into the hydrogel (n = 6 from independent differentiations and ETP donors) (Created in BioRender. Guillonneau, C. (2025) https://BioRender.com/7oyshki). b 3D immunofluorescence confocal imaging of the medullary epithelial compartment (KRT5) and the hematopoietic compartment (CD45) in D28 hTOs. c Flow cytometry of D28 organoids. Black squares indicate the TEC (CD45−EPCAM+) and thymocyte (CD45+EPCAM-) fractions, along with an undefined (CD45+EPCAMlo) population. d FPKM expression of genes encoding HLA-DR, HLA-DM, HLA-DO, and the co-stimulatory molecule CD86 from bulk RNA-seq of sorted HLA-DRlo and HLA-DRhi TECs from D28 hTOs (n = 1). Source data are provided as a Source Data file. e Deconvolution of HLA-DRhi and HLA-DRlo TECs against the Bautista human TEC atlas. Primary human mTEChi, mTEClo and cTEC samples were included as controls. Proportions were estimated using MuSiC (v0.2.0). f qPCR quantification of AIRE expression in D23 hTO TEP/ETP co-cultures (3D) and in conventional 2D monolayer co-cultures. Relative quantification to GAPDH. Error bars show mean ± SD (n = 6, from independent differentiations and ETP donors). **p = 0.001332 (two-sided Wilcoxon rank sum exact test). Source data are provided as a Source Data file. g Proportion of CD205+ and CD205− cells within the CD45−EPCAM+ fraction of D7, D14 and D21 hTOs (human Thymic Organoids) (Left), their respective percentage of HLA-DR cells (Middle), and mean fluorescence intensity (MFI) of HLA-DR and HLA-DP (Right). Source data are provided as a Source Data file.
We then investigated the cellular composition of D28 organoids by flow cytometry and identified three main populations that likely correspond to the TEC compartment (EPCAM+CD45−), thymocytes (EPCAM−CD45+) and an uncharacterized EPCAMloCD45+ population (Fig. 6c). A stromal cell population (EPCAM⁻ CD45⁻), present in a lower proportion than epithelial cells, was also identified (Fig. 6c and Supplementary Fig. 12e). These cells could either be an unintended byproduct of TEP differentiation or result from TEC dedifferentiation and phenotype loss.
Further analysis of the EPCAM+CD45− compartment composition, based on HLA-DR and CD205 expression, revealed a global decrease in CD205 and a large fraction of HLA-DR+ cells with high and low HLA-DR expression (Fig. 6c). To characterize this population further, we isolated HLA-DRhi and HLA-DRlo cells and performed highly sensitive bulk RNA-seq. We found strong expression of HLA-DR encoding genes, namely, HLA-DRA and HLA-DRB1, with higher levels in HLA-DRhi than HLA-DRlo cells, confirming HLA-DR complex expression in the organoid at D28 (Fig. 6d). HLA-DRhi cells exhibited robust expression of CD86, an essential costimulatory molecule for the priming and activation of naive T cells74. In addition, we found that the genes coding for HLA-DM but not HLA-DO were expressed, indicating that HLA-DRhi cells are able to present processed self-antigen peptides via MHCII molecules75,76, and provide activation signals to developing thymocytes, thereby recapitulating antigen presenting cell features of mature TECs.
Finally, to evaluate the cellular composition of HLA-DRhi and HLA-DRlo fractions, we performed deconvolution analysis against the human TEC atlas as an alternative to scRNA-seq and direct cell population identification. Indeed, harvesting the tightly imbricated cells from late organoids suffers from poor recovery and high mortality, hindering scRNA-seq approaches. Inference of the relative abundance of the known human TEC subpopulations in HLA-DRhi and HLA-DRlo TEC samples revealed a mixture of cells that included terminally differentiated mTEC populations, i.e., corneo-like mTECs, myoid cells and ciliated cells also referred as mTEC mimetic cells, as well as mature cTEC populations (Fig. 6e). The absence of immature TEC populations indicates that the iPSc-derived TEPs we used to form the organoids kept differentiating after D7 and lost their immature phenotype. However, despite the observation of higher levels of AIRE expression by qPCR compared to the 2D co-culture system (Fig. 6f), we did not detect its transcripts in RNA-seq data nor hints of bona fide mTEChi by deconvolution, suggesting that AIRE expression might either not reach levels sufficient for its transactivation function or be temporally desynchronized among the maturing TECs.
To monitor the dynamics of EPCAM+ CD205+ and HLA-DR+ cell populations during hTO differentiation, we analyzed samples at D7, D14, and D21. We found that the proportion of CD205+ and CD205− cells within the EPCAM+ population was balanced, with a stable 45% of CD205+ cells at D7 and D14, before decreasing to 20% at D21 (Fig. 6g, Left and Supplementary Fig. 13). This reduction aligns with the overall decrease of CD205 observed at D28 (Fig. 6c). Notably, we observed that a large majority of these CD205+ and CD205- cells are positive for HLA-DR expression at D14 and even more markedly at D21 (Fig. 6g, Middle), consistent with the enhanced HLA-DR and HLA-DP fluorescence at D21 (Fig. 6g, Right). hTOs were also formed with TEPs generated from a slightly modified differentiation protocol that modulated the interplay between BMP4 and Activin A, known to regulate TEP differentiation and maturation77,78. BMP4 has been shown to promote the maintenance of TEPs in a progenitor state, while Activin A directs differentiation towards a medullary fate77. The inhibition of BMP4 and addition of Activin A to our protocol at the end of the AFE phase introduced a differentiation bias towards a medullary fate, with markedly reduced proportions of EPCAM+CD205+ cells in hTOs at each time point (Fig. 6g, Left and Supplementary Fig. 13).
Together, these findings show that our differentiation product acquired a phenotype close to that of human TECs, with differentiation into distinct subpopulations exhibiting cortical and medullary-like features, as well as the capacity to present self-antigen peptides to developing thymocytes via high MHCII expression.
hTOs support thymopoiesis and mature into SP CD4+ and CD8+ T lymphocytes with thymic emigrant phenotypes
To investigate the capacity of hTOs to differentiate ETPs into mature SP T cells, hTOs were dissociated and analyzed by flow cytometry. Soft mechanical dissociation yielded mostly non-adherent CD45+ hematopoietic cells. CD34 expression was absent, confirming the loss of hematopoietic progenitor identity and thus ETP differentiation (Supplementary Fig. 14a). Importantly, we observed a CD3+ population arising at D17 and rapidly expanding at D21, reaching up to 60% of the hematopoietic population at D35, confirming active T cell differentiation in our system (Fig. 7a, b). We also note substantial variation in the proportion of CD3+ cells between experiments at D21 and in later hTOs.
a Flow cytometry of D17, D21, and D35 hTOs showing increasing percentages of CD3+ T cells; representative plots. b Summary of CD3⁺ T cell proportions across independent experiments, with a marked increase observed after 21 days of co-culture. Source data are provided as a Source Data file. c Flow cytometry of D21 hTOs showing αβ and γδ TCR expression within the CD3⁺ fraction (TCRαβ and TCRγδ antibodies). d Flow cytometry of the CD45+CD3+ fraction of D35 hTOs showing SP CD8+ T cells and a less abundant SP CD4+ T cell population. A subset of SP cells displays a CCR7+CD62L+ phenotype, typical of thymic emigrants.
TCR expression was analyzed by flow cytometry to distinguish αβ vs γδ T lineage orientation in hTOs (Fig. 7c). Both αβ and γδ populations were detected in CD3+ thymocytes. The anti-TCRαβ and anti-TCRγδ antibodies used in this study detect assembled heterodimers rather than individual chains, consistent with complete V(D)J recombination. However, the αβ lineage predominated, in contrast to recent results using a similar system79. A large CD3+ population lacking αβ and γδ staining remained and may represent immature pre-TCR-bearing DN T cells. Indeed, CD3 has been reported to be part of the early pre-TCR complex, composed of a TCR β chain and a pre-α chain, appearing as early as the DN2 stage80,81.
Although most CD3+ cells are CD4-CD8- DN thymocytes, we found that hTOs also contained CD4+CD8− SP4 and CD4−CD8+ SP8 cells, albeit in lesser proportions (Fig. 7d). While CD4+CD8+ double-positive (DP) cells were not observed, other hTO experiments have shown their presence, albeit at low frequency within the CD3+ fraction (Fig. 8a, b). In this hTO experiment, a prominent SP CD8+ population, along with a smaller SP CD4+ population, was detected, revealing SP CD8+ and CD4+ thymocyte differentiation and a preferential maturation towards the CD8 fate. We then assessed the maturation state of the SP cells by quantifying the expression of CCR7 and CD62L. Importantly, this analysis revealed populations of SP T cells within hTOs that displayed a mature CCR7+CD62L+ phenotype, demonstrating the ability of hTOs to drive thymopoiesis. This is further supported by the observation that ETPs cultured without TEPs but in the same culture medium as hTOs mostly died by D15, as shown by the culture of three independent ETP isolations, compared to ETP-derived thymocytes within the organoids (Supplementary Fig. 14b). The survival rate can be as low as 0.6% living cells, as illustrated in Supplementary Fig. 14c, Top.
a Flow cytometry of CD45+ cells and their CD3+ subset (CD45+CD3+) for CD4 and CD8 expression in D27 to D38 organoids (five independent organoids/timepoints). Stacked bars show the proportion of CD3+ cells within the DN population (CD4−CD8−). Comparison of SP4 (CD4⁺), SP8 (CD8⁺) and DP (CD4⁺CD8⁺) proportions within total CD45⁺ cells and the CD45⁺CD3⁺ subset across five hTO time points (Bottom Right). Source data are provided as a Source Data file. b Flow cytometry of CD3+ cells gated from total hTOs and analyzed for CD4 and CD8 expression in D10 to D42 organoids. c Summary of CD3⁺ thymocyte subsets (DN, DP, SP4, SP8) from the experiments in (a, b), showing their proportions (Left) and estimated absolute cell numbers (Right) over time, with loess-fitted regression curves for each group. Yellow shading indicates 95% confidence interval (CI) of the loess regression. Source data are provided as a Source Data file.
In addition to ETPs, we isolated already committed DN2/3 thymocytes (CD3-CD4-CD8−CD34−CD7+) (Supplementary Fig. 8) and cultured them alone in the same medium as hTOs (Supplementary Fig. 14c, Middle). Although very low, cell survival was higher than with ETPs (11.3% living cells). Among these cells, differentiation into CD3+ SP8 T cells was observed, indicating preexisting T-lineage commitment. However, they did not exhibit a mature CCR7+CD62L+ phenotype. Hence, the lack of similar differentiating thymocyte populations from ETPs cultured alone confirms the absence of committed DN thymocytes in the isolated ETPs and underscores the reliability of our ETP isolation strategy.
To track CD4+ and CD8+ thymocyte populations throughout organoid differentiation, we analyzed hTOs at several time points from D10 to D42. In a first set of experiments covering four time points from D27 to D38 (Fig. 8a and Supplementary Figs. 15 and 17), we observed an overrepresentation of CD4+ and DP cells within the CD45+ fraction compared to the CD45+CD3+ subset, most pronounced at D27 and gradually diminishing (Fig. 8a, Bottom Right). This shows that ISP cells (CD4+)82 and their immediate DP progeny are captured in the CD45+ fraction before surface CD3 upregulation, but only rarely complete the transition to CD3+ DP cells. It also reveals that engagement into the ISP stage strongly decreases in later hTOs, while CD3+ DN thymocytes accumulate (Fig. 8a), suggesting a direct progression at a limited yield into CD3+ DP cells followed by rapid selection into CD3+ SP8 and SP4 cells, leaving little intermediate accumulation of DP cells within the CD3+ fraction.
We complemented this analysis with a second set of experiments covering five time points from D10 to D42, in which CD4 and CD8 expression was directly assessed within the CD3⁺ fraction of living cells from hTOs (Fig. 8b and Supplementary Figs. 16 and 17). Note that CD3+ SP4 and SP8 cells were already detectable at D10, consistent with previous reports that the DN-to-SP transition can occur within 4–12 days in experimental systems that promote thymocyte differentiation, such as intrathymic transfers or fetal thymus organ cultures83,84. Analyses of the combined sets of hTOs revealed a substantial variability in the proportion of CD4+ and CD8+ SP T cells within the organoids. To characterize the impact of differentiation time on T cell population dynamics within hTOs, we performed loess regression analyses on the proportions of each population (Fig. 8c, Left). We found stable proportions of DN thymocytes throughout differentiation, with a slight decrease of SP4, and stable proportions of SP8 thymocytes. However, the number of generated SP8 and SP4 cells increased over time (Fig. 8c, Right), indicating that their stable or reduced proportion among all CD3+ cells results from the accumulation of CD3+ DN thymocytes, thus masking the parallel increase of SP4 and SP8 T cells. Finally, we showed that the absolute yield of T cell generation from 2500 ETPs in a single organoid at D40 was ~15,000 CD3+ cells, including ~4000 CD8+ T cells and ~500 CD4+ T cells (Fig. 8c, Right).
scRNA-seq demonstrates multilineage differentiation of dendritic and mature T cells
To investigate the CD45+EPCAMlo compartment of hTOs, we performed highly sensitive bulk RNA-seq on the sorted population (Fig. 9a). Gene ontology analysis of differentially expressed genes, relative to iPSc, showed enrichment of terms linked to antigen processing and presentation by the MHCII complex, as well as interaction with lymphocytes (Fig. 9b). In addition, deconvolution analysis of the CD45+EPCAMlo transcriptome against the human thymic cell atlas from Park et al.26, which includes the CD45+ cell fraction, showed that dendritic cells 2 (DC2) likely constitute the majority of the CD45+EPCAMlo population (Fig. 9c). Notably, one of the two tested CD45+EPCAMlo samples showed slight positivity for DC1, indicating that this population may comprise a mixture of DCs, including an important proportion of DC2 and potentially a small number of DC1.
a The CD45+ EPCAMlo population in D28 hTOs was sorted by flow cytometry, and its transcriptome was sequenced using ultra-high-sensitive bulk RNA-seq. b GO term analysis of differentially expressed genes in CD45+EPCAMlo samples (n = 2), shows enrichment of biological processes associated with antigen processing and presentation. c Cell deconvolution of CD45+EPCAMlo from D28 hTOs (n = 2, Replicate (Rep)1 and Rep2) against the human thymic cell atlas from Park et al. d scRNA-seq analysis of the hematopoietic compartment of D28 hTOs (human Thymic Organoids). The UMAP projection reveals three main cell clusters (T(P): proliferating T cells, T(Q): quiescent T cells and DC: dendritic cells), along with an additional stromal cluster. Cell origin, either from the iPSc line (Individual 1) or the primary ETPs (Individual 2) was assessed by SNP comparison using scSplit (v1.0.8). Cluster identity was assigned based on differentially expressed marker genes. e Expression of DC, stromal, and T cell marker genes in the D28 hTO (human Thymic Organoids) clusters defined in d (Left). CD3 and CD7 expression identify T lymphocytes, LY86 and PLEK identify dendritic cells (DCs) and KRT8/18 mark a rare stromal population. Expression of NKT cell marker genes in the same clusters (Right). No enrichment of NKT cells was detected. f For refined cell cluster annotation, D28 hTO (human Thymic Organoids) scRNA-seq data were projected onto the human thymic cell atlas from Park et al. using Seurat label transfer. The reference UMAP (Left) and the projection of D28 hTO cells (Right) are shown; with reference cells in gray. g Violin plots of normalized expression of CD34, FLT3, a TCR-signaling/response module (LAT, ZAP70, LCK, CD5), and a CD3 module (CD3D, CD3E, CD3G, CD247) in hTO cells mapped to DN, SP, DC1, and DC2 clusters. Each dot represents one cell; n indicates the number of cells per cluster. Statistical comparisons (DN vs SP), TCR signaling/response: p = 0.02494 and CD3 genes: p = 0.04041 (Welch’s one-sided t-test).
To further characterize the state of maturation of thymocytes in hTOs and explore their heterogeneity, we performed scRNA-seq of cells harvested at D28 by soft mechanical dissociation for CD45+ cell isolation. Three clusters were identified via UMAP projection (Fig. 9d). Two of these clusters exhibited hematopoietic identity, defined by CD45 expression, while the third and minor one was negative for CD45 expression, consistent with a stromal identity. Differential gene expression analysis pinpointed KRT8, KRT18, FN1, and PDPN as markers of the stromal cluster, indicative of a TEP-derived origin. Identification of key markers for each hematopoietic cell cluster enabled us to identify a dendritic cell population (PLEK, LY86, and HLA-DRA) of which approximately 40% expressed CD14, and a T cell population (CD3E, CD7, and TCF7) (Fig. 9e, Left). We ruled out the presence of NK and NKT cells based on the minimal expression of classical NK lineage markers (Fig. 9e, Right). T cells were divided into an actively proliferating minor cluster “T(P)”, and a larger, quiescent cluster “T(Q)” (Fig. 9d). To further refine cluster annotation and identify their in vivo counterparts, we projected our dataset onto the human thymic cell atlas of Park et al.26 using Seurat label transfer (Fig. 9f). This analysis confirmed the presence of two major DC populations (DC1 and DC2) and SP mature T cells confirming that hTOs support multilineage hematopoietic differentiation. It also identified DN thymocytes, which showed absence of CD34 and FLT3 and expression of CD3 module genes (CD3D, CD3E, CD3G, and CD247) together with upregulation of a pre-TCR/TCR signaling/response module (LAT, ZAP70, LCK, and CD5); both modules were silent in DCs (Fig. 9g). These data indicate post-commitment DN cells, with signaling activity consistent with the early assembly of TCRβ–pre-Tα with CD3 (cytoplasmic in CD3− DN; surface in CD3+ DN). In SP cells, we observed a further significant upregulation of the CD3 module and the TCR signaling/response module, which, together with our previous observation that a subset of SP cells displays a CCR7+CD62L+ phenotype (Fig. 7d), is consistent with the expectation that SP cells express a mature TCR.
As expected, single cell transcriptomic data confirmed the maturation state of hTO-derived T cells, showing expression of markers such as CCR7, CD27, or IL2RA (CD25) (Fig. 9e, Left and Supplementary Fig. 18). To validate that the DCs originated from ETPs rather than from iPSc differentiation, we performed a single nucleotide polymorphism (SNP) analysis using scSplit (v1.0.8) on the dissociated hTO scRNA-seq data. Each cell SNP profile was compared and grouped according to its donor of origin, the ETP donor or the donor from whom the iPSc were derived. As expected, the SNP analysis confirmed that cells originated from two individuals, with the stromal cluster deriving from iPSc and the hematopoietic compartment from ETPs (Fig. 9d). Therefore, both DC and T cell populations generated in hTOs originated from ETPs, revealing the hTO ability to support multilineage hematopoietic differentiation.
Finally, as a control to assess the ability of hTOs to support T cell commitment and maturation, we generated organoids by aggregating iPSc-derived TEPs and hematopoietic stem and progenitor cells (HPSCs) differentiated from the same iPSc line used for TEP generation following the protocol established by Flippe et al.85, which is devoid of any NOTCH signaling induction (Supplementary Fig. 19a). We identified two distinct CD34+ populations: a CD34hi compartment (mainly in the embryoid bodies), and a non-adherent CD34int compartment resembling cord blood (CB) hematopoietic stem cells (HSCs), predominantly found in the culture medium (Supplementary Fig. 20a). A large subset of these CD34+ cells also expressed CD7 (~40% in embryoid bodies and ~60–90% in culture medium) and CD44 (~46% in culture medium and 28% in embryoid bodies), resembling the phenotype of primary ETPs used in our hTO system (Supplementary Fig. 20b, c). In contrast to CD34+CD7- HSPCs, which do not support hTO aggregation (Supplementary Fig. 19a), the CD34+CD7+ HSPCs successfully aggregated (Supplementary Fig. 20d) and differentiated, giving rise at D35 to ~28% CD45+CD3+ cells composed almost entirely of TCRαβ+CD8+ T cells (Supplementary Fig. 21). Interestingly, we observed that hTOs at D28 generated both CD45+CD3+ SP4 and SP8 cells, suggesting that the SP4 population observed at D28, which disappears by D35, corresponds to ISP cells that would entirely mature into SP8 mature T cells at D35. We also observed myeloid lineage differentiation at levels comparable to those in hTOs (Supplementary Fig. 22). Consistent with impaired hTO formation observed with CD34+CD7− HSPCs, attempts to reaggregate CB-derived HSCs, predominantly CD34+CD7−, similarly failed to generate organoids (Supplementary Fig. 19b). Together, these experiments further confirm our model’s ability to support the differentiation of immature, uncommitted hematopoietic progenitors (CD34+CD7+) into both T and myeloid lineages.
Discussion
This work provides an additional protocol for the directed differentiation of iPSc into TEPs and their further maturation into functional thymic organoids supporting thymopoiesis in vitro. Our original approach relies on the combination of an experimental design-based optimization of differentiation factors and the development of a 3D organoid model to favor thymocyte crosstalk within a three-dimensional hydrogel structure.
Previous thymic differentiation protocols were established by testing only a few distinct factor combinations, often iteratively and without a robust statistical approach40,41,42,43,44,86. While this methodology can improve differentiation efficiency, it may also lead to suboptimal results or confounding factor effects. A DOE-based optimization approach reduces bias by simultaneously testing multiple factor combinations. Moreover, the quality of the readout is crucial for optimization. Instead of relying solely on the expression of a few markers, we utilized bulk RNA-seq and publicly available scRNA-seq data from thymic and pharyngeal organogenesis to classify differentiation efficiency. This multidimensional readout enabled us to measure the impact of each differentiation factor on the sample transcriptomes and to select those with the highest similarity to in vivo developing pharyngeal populations. Although the public data used as references were from mice, thymus organogenesis was found to be mostly conserved in humans57. Moreover, the positive results of the optimization advocate for the relevance of the approach. Since we used only one iPSc line for DOE optimization, there was a risk of overfitting the protocol to this specific line. However, we showed that our protocol was also able to differentiate two additional iPSc cell lines into thymic epithelial progenitor cells, confirming its robustness.
In addition to previously identified regulators of AFE and 3PPE induction, such as RA and WNT, we identified a positive effect of IGF1 supplementation on TEP differentiation. We also demonstrated the effect of EGF and FGF10 on iPSc-derived TEP growth. Although our study does not directly demonstrate BMP4 effect on 3PPE induction, we highlighted that it could “prime” cells for improved TEP differentiation at the later stage, measured by increased FOXN1 expression. Recently, several groups reported the generation of human TEPs from iPSc42,43. By testing the effect of various factor combinations on marker gene expression, they observed similar results regarding BMP4 and RA supplementation compared to our study. Interestingly, Gras-Pena et al.43 showed that BMP signaling is time dependent and that switching from activation at D2 to inhibition at D15 benefits TEP induction. This is reflected in our protocol by an increase of the BMP4 dose between the 3PPE and TEP stages. Our findings are also consistent with the protocol proposed by Ramos et al. regarding the WNT-BMP-FGF signaling axis at the TEP stage. However, both Ramos and Gras-Pena studies note the importance of the SHH axis during TEP differentiation. We studied the effect of SHH inhibition using cyclopamine at the AFE stage and did not observe significant results. However, given the similar observations reported by Ramos et al. and Gras-Pena et al., our protocol would benefit from a systematic study of precisely timed SHH modulation, using more potent factors than cyclopamine, such as SAG (SHH agonist) and SANT-1 (SHH inhibitor), to improve TEP differentiation. Finally, our study suggests that a systematic, robust experimental design is well-suited for iPSc differentiation studies due to their inherent variability, temporal sensitivity, multifactorial regulation, and intense experimental workload requirements. Broader DOE designs, testing more factors with more informative readouts, such as single-cell omics, could provide powerful insights into gene regulation during cell differentiation. In addition, we observed high variability both within and between experiments, leading to fluctuating differentiation yields. The state of iPSc during seeding could be an important source of variation. Reagent variability, such as that of Matrigel, is another potential source that must be carefully controlled. Therefore, rigorous standardization and quality control measures are essential for reliable iPSc thymic differentiation.
Characterization of the differentiation product showed expression of TEP markers at both the RNA and protein levels. However, weak expression of TEC markers indicates immaturity, consistent with previous studies in which TEP grafting in vivo was required for the acquisition of functionality40,41. Developing an accessible and practical platform for thymopoiesis studies while ensuring compatibility with future clinical applications necessitates a fully in vitro culture system. A key contribution of this work is the establishment of a 3D co-culture system to mature TEPs into HLA-DRhi mTECs. Thymic crosstalk has been shown to be crucial for mTEC maturation and medulla structuring via signaling from thymocytes, involving the RANK-LTB-CD40 pathways. Furthermore, classical 2D culture has been shown to be detrimental to primary TEC functionality, whereas 3D culture promotes maintenance and maturation of primary TECs69. Therefore, we adapted previous methodologies52,69 to develop a fibrin-based hydrogel seeded with ETP-TEP reaggregates. This approach resulted in the formation of hTOs that grow at the air-liquid interface within 3D hydrogels. Since thymopoiesis in vivo relies on the sequential migration of cells through cortical and medullary niches, achieving proper cortico-medullary segregation within hTOs must be pursued. Modulating matrix properties and minimizing handling steps during reaggregation could be promising strategies to achieve this goal.
Although we did not provide direct evidence of thymic selection occurring in hTOs, we observed the generation of SP T lymphocytes with mature CCR7+CD62L+ phenotypes from ETPs. Since the acquisition of these markers relies on survival signals through interaction with TECs, this indirectly demonstrates the ability of hTOs to support thymic selection. The presence of SP CD4+ T lymphocytes in approximately the same proportion as SP CD8+ T cells suggests that DP thymocytes undergo CD4+ selection within hTOs. However, the underrepresentation of DP cells and the accumulation of CD3+ DN thymocytes compared to the fetal thymus point to a DN-to-DP bottleneck, likely due to limiting interaction of developing thymocytes with Notch-ligand-positive TECs at later stage of hTO formation, highlighting the need to fine-tune the synchronization between TEC and thymocyte maturation throughout hTO development. In addition, the variability in CD3+ frequencies at comparable time points (e.g., D21) likely reflects variation in hTO development, with CD3⁺ DN cells accumulating earlier in some organoids and later in others. Interestingly, flow cytometry confirmed the ability of our system to generate both αβ and γδ T lineages, in contrast to studies showing an abnormal restriction to γδ T generation79. We also observed that a substantial fraction of the CD45+ compartment consisted of CD3+ TCRαβ− TCRγδ− cells, consistent with DN thymocytes expressing CD3 and associated with the pre-TCR complex (β chain with pre-Tα and CD3 subunits), whose net abundance likely reflects DN proliferation combined with limited progression to the DP stage. A deeper analysis of the hTO hematopoietic compartment using scRNA-seq revealed the presence of dendritic cells (DC). This population corresponds to the EPCAMloCD45+ population observed in hTO flow cytometry data. Given recent reports highlighting the critical role of intrathymic DCs in complementary thymocyte selection, this population may contribute to T cell differentiation in hTOs, potentially promoting commitment to the SP CD4+ fate.
Ramos et al. reaggregated and grafted the TEPs they had differentiated into murine models to promote their maturation into TECs. They reported KRT5 and KRT8 positive cells but did not provide a quantitative assessment of the maturation level of the TEC products. In contrast, our study demonstrates the presence of a substantial HLA-DR positive epithelial population. However, the swine thymus graft proposed by Gras-Pena et al. successfully reproduced cortico-medullary segregation, possibly due to the pre-existing ECM structure, which may influence TEP fate decisions. In terms of T cell generation, both systems support thymopoiesis, with Ramos et al. generating both innate immune cells and mature T cells, while Gras-Pena et al. primarily observed DP cells. This highlights the closer resemblance of thymic grafts to the human thymus, whereas our ex vivo system exhibits lower DP and SP4 production. However, our system is more compatible with scalable, off-the-shelf therapies and carries a lower risk of rejection compared to swine thymus grafts.
At the current scale, T-cell output remains modest mainly because the number of hematopoietic progenitors available from primary thymic samples restricts input and thus limits the size of organoid batches. iPSc-derived HSPCs have proven compatible with hTOs, offering a potentially unlimited input source that overcomes this limitation and enables scale-up within our established system. While the DN-to-DP bottleneck also constrains T-cell generation, it appears less pronounced in hTOs seeded with CD34+CD7+ iPSc-derived HSPCs (Supplementary Fig. 21). Thus, the combination of iPSc-derived HSPC input and refinements, including automated hydrogel seeding and tuning of gel stiffness and aggregate size to improve culture-medium diffusion and cell interactions, will deliver higher and more reproducible T-cell output, enabling comprehensive functional assays and translational studies.
With the aforementioned necessary improvements and additional studies, we foresee two main clinical applications for this system. The first leverages the differentiated TEC compartment to remediate rare pathologies where deficiencies in crucial genes involved in thymic organogenesis (such as FOXN1 or PAX1 in SCID and TBX1 in DiGeorge’s syndrome) lead to thymic hypoplasia and immunodeficiency. Autologous transplantation with gene-corrected iPSc-derived TECs could correct this deficiency and restore, at least partially, T cell populations with self-tolerance. The second application leverages the ability of hTOs to mature T cells derived from iPSc banks for heterologous, off-the-shelf cell therapy, which also enables cell editing, such as CAR modification, for immuno-oncology applications.
Our study provides insights into the regulation of iPSc thymic differentiation and advocates for the application of robust statistical tools for this purpose. It establishes a fully in vitro human thymic organoid model derived from iPSc. This multilineage organoid includes TECs, DCs, T cells, and a potential fibroblastic population, demonstrating thymic functionality through its ability to generate mature SP T lymphocytes. Thus, it provides a crucial resource for modeling thymopoiesis in vitro. The use of iPSc opens promising research perspectives for long-term applications in regenerative medicine and cellular therapies, particularly via the in vitro generation of engineered T lymphocytes.
Methods
Isolation of human primary ETPs and TECs
Postnatal human thymic samples were obtained as surgical waste from routine pediatric cardiac surgeries performed at CHU Nantes. The study was conducted under institutional declaration DC-2017-2987, in compliance with the French CODECOH regulatory framework. Within this framework, patients’ legal guardians were informed by the clinical team, and their non-opposition to research use was obtained. Procedural oversight and authorization were formalized through an institutional agreement (“charte d’utilisation”) between CHU Nantes and the clinical and research teams. All samples were fully anonymized prior to being provided to the research team.
ETPs isolation was performed as described in Lavaert et al.56. Briefly, fresh thymus samples were dissected on the same day in 1 mm3 fragments in RPMI1640 and dissociated by mechanical pipetting to release thymocytes. A 5 min incubation in red blood cell lysis solution (Miltenyi, 130-094-183) was performed for erythrocyte depletion. Cells were passed through 70 µm meshes and incubated with CD3 labeled Dynabeads (Thermofisher, 11031) to deplete most DP and SP thymocytes. ETP-enriched cell suspension was stained for flow cytometry and ETPs sorted with a FACS ARIA (BD Biosciences) using phenotype CD3−CD4−CD8−CD14−CD19−CD56−CD34+CD7+ and immediately used for reaggregation culture. For TEC isolation, thymic fragments were digested using a 0.5 mg mL−1 collagenaseD (Roche, 11088866001), 1 mg mL−1 dispase (Roche, 04942078001) and 0.5 mg mL−1 DNAseI (Roche, 11284932001) solution in RPMI1640 for 45 min at 37 °C and dissociated using GentleMACS (Miltenyi, 130-093-235) and C tubes (Miltenyi, 130 093 237). Cell suspension was deposited on a 21% Optiprep (Sigma Aldrich, D1556 250 ML) gradient and centrifuged 20 min at 500 g for TEC enrichment. Cell suspension was filtered on a 100 µm mesh and sorted based on the CD45−EPCAM+ phenotype.
Stem cell culture and banking
iPSc lines were provided by the Nantes iPSC core facility (L.D.). The LON71 and LON80 are human iPSc generated from healthy fibroblasts, from a 51-year-old male and a 57-year-old female, respectively87. The MIPS203 is a human iPSc generated from fibroblasts from a 30-year-old female88. Cells were banked at P21 and used between P21 and P35. The banks were validated by SNP arrays.
iPSc culture and differentiation into thymic epithelial progenitors (TEPs)
iPSc lines were maintained on Matrigel coated 6-well plates (StemCell Technologies, 354277) in mTeSR1 medium (StemCell Technologies). iPSc cultures were passaged every 5 to 6 days, using xeno-free passaging solution (StemCell Technologies) and were seeded in clumps. In case of abnormal colony morphology in cultures, daily cleaning was performed to guarantee optimal iPSc purity. iPSc were routinely cultured at 37 °C in normoxic conditions. To control cell state variability, iPSc were passaged 4 days before induction of differentiation, then harvested 24 h before the D0 in single cell suspension using TrypLE (ThermoFisher) with 5 min incubation at 37 °C. Cells were resuspended in mTeSR1 medium with 10 µM ROCK inhibitor Y27632 (Sigma Aldrich Y0503) and seeded on Matrigel-coated 12-well plates at 37 × 103 cells/cm2. Differentiation was induced at D0 with XVIVO10 medium (Lonza, BE04-380Q) supplemented with 5 µM CHIR99 (Miltenyi, 130-106-539) and 100 ng mL−1 ActivinA (BiotechneR&D, 338-AC). Culture medium changes were performed at fixed hours, with quick rinsing by warm PBS to eliminate potential differentiation medium leftovers in wells. D1 and D2 medium contained only 50 ng mL−1 Activin A. At D3, cells were passaged in single cell as described above and seeded at low density (14.2-20 × 103 cells/cm2 depending on iPSc line) in Matrigel coated 12-well plate with 10 µM Y27 and 50 ng mL−1 ActivinA. Culture medium was changed at D4, D5, D7, D9, D10, D11, and D13 as previously described with supplementation as detailed in Fig. 1. The supplementation included Activin A, CHIR99, BMP4 (Miltenyi, 130-111-167), retinoic acid (Sigma Aldrich, 302-79-4), FGF8 (BiotechneR&D, 423-F8), Noggin (Miltenyi 130-103-455), LY-364947 (Sigma Aldrich L6293-5MG), FGF10 (Miltenyi, 130-127-858), IGF1 (Miltenyi, 130-093-886), EGF (Miltenyi, 130-097-751), and RANK-L (BiotechneR&D: 6449-TEC).
TEP reaggregation with ETPs and hTO formation
TEPs were harvested at D13 using TrypLE with a 7 min incubation at 37 °C and mechanical flushing. TEPs were aggregated in low binding 96-well plates (ThermoFisher, 174929), at an 8:1 ratio with fresh ETP sorted as described above, using 20,000 TEPs and 2,500 ETPs per well. Cultures were maintained for 24 h in XVIVO10-based medium supplemented with 1% Glutamax (Gibco, 35050-38/2165269), 1% non-essential amino acids (Gibco, 11140-035), 30 mM L-ascorbic acid, 0.1 µM retinoic acid, 50 ng mL−1 BMP4, 50 ng mL−1 RANKL, 10 ng ml−1 FGF8, 10 ng ml−1 FGF10, 10 ng ml−1 IGF1, 10 ng ml−1 EGF, 10 ng ml−1 SCF (Peprotech AF-300-07), 5 ng ml−1 IL7 (Peprotech AF-200-07), and 5 ng ml−1 FLT3L (Peprotech AF-300-19). Cell micromasses should show a typical spheroid morphology with a compact central cell mass surrounded by a crown composed mainly of ETPs. Hydrogels were formed by mixing a 8 mg mL−1 Fibrinogen (Sigma Aldrich, 341578) solution heated at 37 °C with 10 U mL−1 Thrombin (Sigma Aldrich, 605190) and 26,000 U mL−1 Aprotinin (Sigma Aldrich, 616370), and immediately casting it in the superior compartment of 12-well inserts (Merk PTHT12H48). 2–4 spheroids were individually collected using cut 200 µL pipette tips pre-coated with anti-adherence solution and seeded delicately on the hydrogels. The same culture medium described above was added in a way to form an air-liquid interface without drowning the hydrogels and changed every three days during the first week. Then retinoic acid, BMP4, RANKL, FGF8, FGF10, IGF1, and EGF were removed from the culture medium. To harvest the cells, the hydrogels were carefully collected using cut 1000 µL pipette tips and incubated 15 min at 37 °C in a 0.5 mg mL−1 Collagenase (Roche, 11088866001) solution in DMEM/F12. Organoids were subsequently processed with a 10 min incubation in TrypLE at 37 °C with frequent mechanical disruption by pipetting, then filtered on a 100 µm mesh.
iPSc to HSPC directed differentiation
iPSc (LON71) were differentiated toward a HSPC identity following a protocol described in ref. 85. Briefly, iPScs were incubated with Dispase (StemCell Technologies, 07913) at 37 °C for 20 min, then rinsed with DMEM-F12 (Gibco, 11330). The colonies were lifted in mTeSR1 with 10 µM of Y-27632 using a 5 mL pipette placed perpendicularly to the bottom of the well, and deposited into low-attachment MW6 plates (Corning, 003471) to form embryoid bodies (EBs) for 18 h. The next day the EBs were collected, deposited into 15 mL polystyrene tubes and allowed to sediment. The medium was aspirated and the EBs were resuspended in EB medium [StemPro-34 SFM (Gibco, 10640) + 2 mM Glutamax (Gibco, 35050-38/2165269) + 50 µM 2-mercaptoethanol (Gibco, 31350010) + 1% non-essential amino acids (Gibco, 11140-035) + 50 µg mL−1 L-ascorbic acid (Sigma Aldrich, A7506-100)] with D0 cytokines (Table 1). This technique was employed for medium changes until D9 to D12, with one modification: as we observed the progressive migration of the CD34+ precursor cells from the EBs into the culture medium, we harvested those cells during medium changes at D5, D7, and D9 when applicable, by collecting and spinning the culture medium at 200 G for 5 min. The pellet was resuspended in the fresh culture medium and reintroduced into the culture wells. Cells were differentiated for 9 to 12 days to reach the HSPC stage, an early progenitor state uncommitted to the T-cell lineage and free of NOTCH signaling induction.
HSPC harvesting and iPSc-derived hTO formation
On D12 of iPSc to HSPC differentiation, the organoid bodies were dissociated in TrypLE for 15 min at 37 °C, with mechanical disruption by vortexing every 5 min. The cells present in the culture medium underwent the same treatment for 5 min. The dissociated cells were spun at 300 G for 5 min, resuspended in FACS buffer and stained against CD34 and CD7 and with DAPI. The cells were sorted on an Aurora CS (Cytek Biosciences), gating on living CD34+CD7+ and CD34+CD7− cells. The iPSc-derived TEP cells were harvested as described above. TEP and CD7+ or CD7− HSP cells were aggregated in low binding 96-well plates at a ratio of 3000 HSPCs and 20,000 TEPs per well. Culture was maintained for 24 h as described above. After 24 h, we observed the successful formation of spheroids in the CD7+ condition, and an absence of aggregation in the CD7− condition. The CD7+ hTOs were seeded on hydrogels and cultured for 5 weeks as described above, and FACS analyses were performed on D28 and D35 on an Aurora (Cytek Biosciences N7-00003).
The same protocol was followed using cord blood CD34+ cells (StemCell technologies ref 70008.1, lot #1602190124) and a ratio of 3000 cord blood cells and 20,000 TEPs per well. After 24 h, we observed the successful formation of spheroids, which were seeded on hydrogels and cultured as described above. The culture was maintained for three weeks, at which point we observed the spontaneous lysis of the hTOs. The gels and remaining cells were treated as previously described and analyzed on an Aurora (Cytek Biosciences N7-00003).
RNA-extraction, RT and qPCR
Cell samples were lysed using RLT Lysis buffer (Qiagen, 79216). RNA extraction was performed using RNAeasy Mini Kit (Qiagen, 74104) and retrotranscription with Superscript III (Thermofisher, 18080093) following the manufacturer’s guidelines. qPCR was performed with SYBR Green Fast kit (ThermoFisher, 4385612) on a ViiA 7 Real-Time PCR System (ThermoFisher) using the primers listed in Table 2. Relative quantification of target gene expression was measured using the delta CT method with GAPDH as the endogenous control.
Bulk RNA sequencing by DGE-seq and SMART-seq
For 3′ Digital gene expression (DGE) bulk RNA sequencing, the protocol was performed according to our implementation of previously developed protocols88. Ten nanograms of total RNA were used for library preparation. mRNA was tagged via poly(A) tails-specific adapters, well-specific barcodes and universal molecular identifiers (UMI) with template-switching retrotranscription. cDNAs were pooled, amplified and tagmented by transposon-fragmentation to enrich 3′ ends. Sequencing was performed on an Illumina Novaseq 6000. Kits used were Zymo purification kit, Research D4004-1-L, Kit Advantage 2 PCR Enzyme System (Clontech, 639206), QIAquick Gel Extraction, Kit AgencourtAMPure XP magnetic beads and Nextera DNA (FC-121-1031). For highly sensitive bulk RNA-seq, we performed SMART-seq RNA sequencing. RNA was processed with the SMART-Seq v4 Ultra Low Input RNA kit and the libraries were prepared with Illumina Nextera XT DNA Library Preparation Kit. An Illumina NextSeq500 was used for paired-end sequencing.
RNA-seq data processing and analysis
DGE RNA-seq data were processed using the SRP pipeline v1.089 (https://gitlab.univ-nantes.fr/bird_pipeline_registry/srp-pipeline), a Snakemake-based workflow for 3’ sequencing RNA profiling analysis. Read pairs used for analysis matched the following criteria: all 16 bases of the first read had quality scores of at least 10 and the first 6 bases correspond exactly to a designed well-specific barcode. The second reads were aligned to RefSeq human mRNA sequences (hg19) using BWA version 0.7.17. Reads mapping to several transcripts of different genes or containing more than three mismatches with the reference sequences were filtered out from the analysis. DGE profiles were generated by counting for each sample the number of unique UMIs associated with each RefSeq gene. Sequenced samples with at least 50,000 counts and 6000 expressed genes were retained for further analysis.
For SMART-seq data, reads were aligned to the human reference genome (hg19) using TopHat2 (v2.0.12) and Bowtie2 (v2.2.3) with default parameters. FPKM quantification was performed using Cufflinks (v2.2.1), and read counts were obtained using HTSeq-count (v0.11.x).
Batch correction and differential expression were performed with DESeq2 R package (v1.30.1), with the threshold of 1 log2FC and 0.05 Benjamini–Hochberg p-value to classify a gene as differentially expressed vs D0 iPSc controls. Gene Ontology (GO) analysis of differentially expressed gene modules was performed using the ClusterProfiler R package (v3.18.1) on the KEGG database of biological pathways. The GO network was plotted using Cytoscape (v3.8.x). Data visualization relied on custom pipelines based on the ComplexHeatmap (v2.6.2) and ggplot2 (v3.3.3) R packages. To predict cell type proportions in the bulk samples, deconvolution was performed against multiple scRNA-seq atlases as reference (from Magaletta et al., Bautista et al., and Park et al.), applying the MuSiC_prop() function with default parameters (MuSiC v0.2.0). Bulk RNA-seq data from human mTEChi, mTEClo, and cTEC populations, used as controls in the deconvolution of hTO-derived TECs, were generated by the Peterson lab and are available in GEO under accession number GSE176445.
scRNA-seq data generation
D7 dissociated hTOs were sorted using a FACS ARIA (BD Biosciences) to purify EPCAM+CD45− cell population and remove dead cells and debris. Cells from D28 hTOs were harvested by soft mechanical dissociation to capture those that dissociate from the 3D co-culture, which are mostly CD45+. Cells were loaded on the Chromium instrument (10X Genomics). Libraries were prepared with Chromium Single-Cell 3′ Reagent v2 Kit (10× Genomics, Pleasanton, CA) as per the manufacturer’s protocol. Illumina adapters were incorporated during library preparation. For D28 hTOs, cell samples were incubated with TotalSeqB antibodies (BioLegend) prior to droplet generation. cDNA synthesis was conducted to simultaneously capture both mRNA and hashtag oligonucleotide sequences. A targeted PCR amplification of the hashtag-derived cDNA was then performed to generate a discrete library, which was pooled with the mRNA-derived library. We considered the cell sample labeled with TotalSeq-B0252 for further analysis. Sequencing was performed on an Illumina Novaseq 6000.
scRNA-seq data processing and analysis
Primary data analysis was performed on CellRanger (v5). Cell count matrices were exported and secondary analysis was performed on R using Seurat v4 package90. Quality control was defined by keeping cells with gene numbers between 2000 and 15,000 and fewer than 0.1 proportion of mitochondrial genes. Variable gene identification, data scaling and dimension reduction used functions integrated in Seurat. Dimensionality reduction used UMAP and tSNE embedding on the first 10 dimensions. Clustering used the Louvain algorithm implemented in the FindCluster function with a resolution of 0.3. Visualization and marker determination relied on default Seurat functions. To determine whether D28 hTO-harvested cells originated from one or two genetically distinct human samples, we used scSplit (v1.0.8) with the options -n0 -s2, which infers sample identity based on SNP patterns91. For the analysis of public sc datasets of thymus and pharyngeal organogenesis, raw data were downloaded at the accession provided in the data availability section of the relevant papers26,32,37. Park’s dataset was downsized by random sampling by a factor of 0.25 to ease data handling. Quality control was set at 500/10,000/10 and 500/4000/10 for Han’s and Magaletta’s datasets. Dimension reduction was performed by UMAP on first 10 (Park), 21 (Magaletta) and 25 (Han) dimensions. Clustering was performed with a resolution of 0.75 (Park), 0.5 (Magaletta), 0.6 (Han). Visualization and marker identification were performed as described above. Projection of D28 hTO-harvested cells onto the Park thymic atlas was performed using Seurat’s label transfer workflow (Seurat v4.x). Anchors were computed with FindTransferAnchors (PCA reduction, 30 dimensions), and cell type labels were transferred with TransferData. Query cells were then mapped onto the UMAP structure of the Park reference with MapQuery using the stored UMAP model, to enable visualization of predicted identities in the reference topological space. The datasets from Magaletta and Han were obtained from the GEO databases (GSE182135 and GSE136689, respectively). Park’s dataset was obtained from the human thymic cell atlas portal: https://cellatlas.io/studies/thymus-development#resources.
We used scRNA-seq data from Bautista (GSE147520) as provided, for integration with our scRNA-seq data and for deconvolution of bulk RNA-seq profiles, both generated from hTO-derived TECs. For the integration, anchors were computed with FindIntegrationAnchors (2000 features, PCA reduction, 30 dimensions), and datasets were harmonized with IntegrateData. The integrated object was processed with ScaleData, PCA (30 dimensions), and UMAP embedding, followed by clustering using FindNeighbors and FindClusters, enabling joint visualization and annotation of TEC subsets across the two datasets
Flow cytometry and immunofluorescence imaging
Cells were stained with the antibodies listed in Supplementary Data 3 according to the supplier’s instructions and analyzed on the FACS ARIA (BD Biosciences), the FACS Celesta (BD Biosciences), and the spectral FACS Aurora (Cytek). Flow cytometry data was processed on FlowJo (vX.0.7) and Cytek Spectroflo (v3.3.0). For 2D IF, cells were cultured on Ibidi 8-well plates (Ibidi, 80806), fixed with 4% PFA for 15 min and permeabilized with IF buffer (PBS, 0.2% Triton, 10% inactivated FBS) for 1 h. Cells were incubated in IF buffer overnight at 4 °C with primary antibodies and for 2 h at room temperature for secondary antibodies. Plates were imaged on a confocal SIM microscope. Modifications were introduced to the protocol for DLL1, 4, PAX9 and FOXN1 staining, consisting of using 0.1% Triton and no inactivated FBS in the IF buffer, as well as omitting Triton during antibody incubation. For hTO staining, the hydrogels were harvested by using a widely cut 1000 µL pipette tip pre-coated with anti-adherence solution and carefully deposited in Ibidi 8-well plates in cold PBS. Most of the gel matrix was removed by delicate flushing and several PBS washing steps, without damaging the main cellular structures. hTOs were fixed with 4% PFA for 20 min and permeabilized with IF buffer (PBS, 0.2% Triton, 10% inactivated FBS) for 2 h. Primary staining was performed in IF buffer for 24 h and secondary staining for 4 h, and imaged on SIM confocal microscope in Z-stack mode. Images were processed on FIJI (v2.16).
Gene tissue specificity calculation and measure of TRA enrichment
Calculation of gene tissue specificity scores was performed using a tool developed internally, that implements the methods proposed by Yanai et al.73 and Pan et al.92. The Tau was calculated for a list of genes using the gene tissue expression data (GTEx) from the Human Protein Atlas database (https://www.proteinatlas.org/about/download). The Tau index varies between 0 and 1, with higher values indicative of higher gene tissue specificity. The tool was deployed into a user-friendly and free to use dashboard built in Shiny for diffusion to the community and allowing customized analyses. The Shiny application is available at https://nathanprovin.shinyapps.io/GeneTissueSpe/.
Statistics and data analysis
Statistical tests were performed in R. Mean comparison significance on qPCR graphs was tested using the non-parametric Wilcoxon tests with a significance p-value threshold of p < 0.05. DOE experimental plans were designed using R DOE base package (v1.1.x). DOE results were analyzed by factorial ANOVA as detailed in figure legends. All statistical tests were two-sided unless specified. Graphs and statistical analyses were generated on GraphPad Prism (v10.4.1).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The DGE-seq RNA sequencing data related to the DOE-based factor screening and generated in this study have been deposited in the GEO database under accession code GSE302942. The DGE-seq data related to the iPSc to TEP differentiation and generated in this study have been deposited in the GEO database under the accession code GSE304598. The CD205 TEP RNA-seq data generated in this study have been deposited in the GEO database under accession code GSE294118. The D7 hTO TEC scRNA-seq data generated in this study have been deposited in the GEO database under accession code GSE293928. The RNA-seq data of HLA-DRhi, HLA-DRlo TECs and thymic DCs in D28 hTOs generated in this study have been deposited in the GEO database under accession code GSE294061. The scRNA-seq data of D28 hTO detaching cells generated in this study have been deposited in the GEO database under accession code GSE293930. The scRNA-seq data of early pharyngeal development (Han) reused in this study are available in the GEO database under accession code GSE136689. The scRNA-seq data of late pharyngeal development (Magaletta) reused in this study are available in the GEO database under accession code GSE182135. Park’s scRNA-seq data of human thymic cells reused in this study are available at [https://cellatlas.io/studies/thymus-development#resources]. Bautista’s scRNA-seq data of human TECs reused in this study are available in the GEO database under accession code GSE147520. RNA-seq data from human mTEChi, mTEClo, and cTEC populations, used as controls in this study, are available in the GEO database under accession code GSE176445. All other data are available in the article and its Supplementary files or from the corresponding author upon request. Source data are provided with this paper.
Code availability
We have provided the documented R code that we developed for the calculation of gene tissue specificity scores93 as a Git repository available from https://github.com/TeamGiraud/GeneTissueSpe and as a Shiny application at https://nathanprovin.shinyapps.io/GeneTissueSpe.
References
Miller, J. F. A. P. The function of the thymus and its impact on modern medicine. Science 369, eaba2429 (2020).
Abramson, J. & Anderson, G. Thymic epithelial cells. Annu Rev. Immunol. 35, 1–34 (2016).
Bautista, J. L. et al. Single-cell transcriptional profiling of human thymic stroma uncovers novel cellular heterogeneity in the thymic medulla. Nat. Commun. 12, 1096–15 (2021).
Kadouri, N., Nevo, S., Goldfarb, Y. & Abramson, J. Thymic epithelial cell heterogeneity: TEC by TEC. Nat. Rev. Immunol. 20, 239–253 (2019).
Bornstein, C. et al. Single-cell mapping of the thymic stroma identifies IL-25-producing tuft epithelial cells. Nature 559, 622–626 (2018).
Carpenter, A. C. & Bosselut, R. Decision checkpoints in the thymus. Nat. Immunol. 11, 666–673 (2010).
Deng, Y. et al. Leaving no one behind: tracing every human thymocyte by single-cell RNA-sequencing. Semin. Immunopathol. 43, 29–43 (2021).
Chopp, L. B. et al. An integrated epigenomic and transcriptomic map of mouse and human αβ T cell development. Immunity 53, 1182–1201.e8 (2020).
Klein, L., Klugmann, M., Nave, K. A., Tuohy, V. K. & Kyewski, B. Shaping of the autoreactive T-cell repertoire by a splice variant of self protein expressed in thymic epithelial cells. Nat. Med. 6, 56–61 (2000).
Klein, L., Kyewski, B., Allen, P. M. & Hogquist, K. A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don’t see). Nat. Rev. Immunol. 14, 377–391 (2014).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401 (2002).
Cumano, A. et al. New molecular insights into immune cell development. Annu. Rev. Immunol. 37, 497–519 (2019).
Cosway, E. J., James, K. D., Lucas, B., Anderson, G. & White, A. J. The thymus medulla and its control of αβT cell development. Semin. Immunopathol. 43, 15–27 (2021).
Liang, K. L. et al. Intrathymic dendritic cell-biased precursors promote human T cell lineage specification through IRF8-driven transmembrane TNF. Nat. Immunol. 24, 474–486 (2023).
Spidale, N. A., Wang, B. & Tisch, R. Cutting edge: antigen-specific thymocyte feedback regulates homeostatic thymic conventional dendritic cell maturation. J. Immunol. 193, 21–25 (2014).
Nusser, A. et al. Developmental dynamics of two bipotent thymic epithelial progenitor types. Nature 606, 165–171 (2022).
Bleul, C. C. et al. Formation of a functional thymus initiated by a postnatal epithelial progenitor cell. Nature 441, 992–996 (2006).
Ulyanchenko, S. et al. Identification of a bipotent epithelial progenitor population in the adult thymus. Cell Rep. 14, 2819–2832 (2016).
Alves, N. L. et al. Serial progression of cortical and medullary thymic epithelial microenvironments. Eur. J. Immunol. 44, 16–22 (2014).
Baik, S., Jenkinson, E. J., Lane, P. J. L., Anderson, G. & Jenkinson, W. E. Generation of both cortical and Aire+ medullary thymic epithelial compartments from CD205+ progenitors. Eur. J. Immunol. 43, 589–594 (2013).
Li, J. et al. NOTCH1 signaling establishes the medullary thymic epithelial cell progenitor pool during mouse fetal development. Development 147, dev178988 (2020).
Liu, D. et al. Canonical Notch signaling controls the early thymic epithelial progenitor cell state and emergence of the medullary epithelial lineage in fetal thymus development. Development 147, dev178582 (2020).
Irla, M. RANK signaling in the differentiation and regeneration of thymic epithelial cells. Front. Immunol. 11, 623265 (2021).
Irla, M. et al. Autoantigen-specific interactions with CD4+ thymocytes control mature medullary thymic epithelial cell cellularity. Immunity 29, 451–463 (2008).
Lopes, N. et al. Thymocytes trigger self-antigen-controlling pathways in immature medullary thymic epithelial stages. eLife 11, e69982 (2022).
Park, J.-E. et al. A cell atlas of human thymic development defines T cell repertoire formation. Science 367, eaay3224 (2020).
Gao, H. et al. The lineage differentiation and dynamic heterogeneity of thymic epithelial cells during thymus organogenesis. Front. Immunol. 13, 805451 (2022).
Zeng, Y. et al. Single-cell RNA sequencing resolves spatiotemporal development of pre-thymic lymphoid progenitors and thymus organogenesis in human embryos. Immunity 51, 930–948.e6 (2019).
Bredenkamp, N. et al. An organized and functional thymus generated from FOXN1-reprogrammed fibroblasts. Nat. Cell Biol. 16, 902–908 (2014).
Nowell, C. S. et al. Foxn1 regulates lineage progression in cortical and medullary thymic epithelial cells but is dispensable for medullary sublineage divergence. PLoS Genet. 7, e1002348 (2011).
Blackburn, C. C. et al. The nu gene acts cell-autonomously and is required for differentiation of thymic epithelial progenitors. Proc. Natl. Acad. Sci. USA 93, 5742–5746 (1996).
Magaletta, M. E. et al. Integration of single-cell transcriptomes and chromatin landscapes reveals regulatory programs driving pharyngeal organ development. Nat. Commun. 13, 457 (2022).
Farley, A. M. et al. Dynamics of thymus organogenesis and colonization in early human development. Development 140, 2015–2026 (2013).
Gordon, J., Bennett, A. R., Blackburn, C. C. & Manley, N. R. Gcm2 and Foxn1 mark early parathyroid- and thymus-specific domains in the developing third pharyngeal pouch. Mech. Dev. 103, 141–143 (2001).
Gordon, J. & Manley, N. R. Mechanisms of thymus organogenesis and morphogenesis. Development 138, 3865–3878 (2011).
Figueiredo, M., Zilhão, R. & Neves, H. Thymus inception: molecular network in the early stages of thymus organogenesis. Int. J. Mol. Sci. 21, 5765 (2020).
Han, L. et al. Single cell transcriptomics identifies a signaling network coordinating endoderm and mesoderm diversification during foregut organogenesis. Nat. Commun. 11, 4158 (2020).
Nowotschin, S. et al. The emergent landscape of the mouse gut endoderm at single-cell resolution. Nature 569, 361–367 (2019).
Chhatta, A., Mikkers, H. M. M. & Staal, F. J. T. Strategies for thymus regeneration and generating thymic organoids. J. Immunol. Regen. Med. 14, 100052 (2021).
Parent, A. V. et al. Generation of functional thymic epithelium from human embryonic stem cells that supports host T cell development. Cell Stem Cell 13, 219–229 (2013).
Sun, X. et al. Directed differentiation of human embryonic stem cells into thymic epithelial progenitor-like cells reconstitutes the thymic microenvironment in vivo. Cell Stem Cell 13, 230–236 (2013).
Ramos, S. A. et al. Generation of functional human thymic cells from induced pluripotent stem cells. J. Allergy Clin. Immun. 149, 767–781.e6 (2022).
Gras-Peña, R. et al. Human stem cell-derived thymic epithelial cells enhance human T-cell development in a xenogeneic thymus. J. Allergy Clin. Immun. 149, 1755–1771 (2022).
Soh, C.-L. et al. FOXN1 (GFP/w) reporter hESCs enable identification of integrin-β4, HLA-DR, and EpCAM as markers of human PSC-derived FOXN1(+) thymic epithelial progenitors. Stem Cell Rep. 2, 925–937 (2014).
Provin, N. & Giraud, M. Differentiation of pluripotent stem cells into thymic epithelial cells and generation of thymic organoids: applications for therapeutic strategies against APECED. Front. Immunol. 13, 930963 (2022).
Callaghan, N. I. et al. Harnessing conserved signaling and metabolic pathways to enhance the maturation of functional engineered tissues. NPJ Regen. Med. 7, 44 (2022).
Toms, D., Deardon, R. & Ungrin, M. Climbing the mountain: experimental design for the efficient optimization of stem cell bioprocessing. J. Biol. Eng. 11, 35 (2017).
Yasui, R., Sekine, K. & Taniguchi, H. Clever experimental designs: shortcuts for better iPSC differentiation. Cells 10, 3540 (2021).
Besnard, M., Padonou, F., Provin, N., Giraud, M. & Guillonneau, C. AIRE deficiency, from preclinical models to human APECED disease. Dis. Model Mech. 14, dmm046359 (2021).
Chaudhry, M. S., Velardi, E., Dudakov, J. A. & Brink, M. R. M. Thymus: the next (re)generation. Immunol. Rev. 271, 56–71 (2016).
Bosticardo, M. et al. Artificial thymic organoids represent a reliable tool to study T-cell differentiation in patients with severe T-cell lymphopenia. Blood Adv. 4, 2611–2616 (2020).
Montel-Hagen, A. et al. In vitro recapitulation of murine thymopoiesis from single hematopoietic stem cells. Cell Rep. 33, 108320–108320 (2020).
Seet, C. S. et al. Generation of mature T cells from human hematopoietic stem and progenitor cells in artificial thymic organoids. Nat. Methods 14, 521–530 (2017).
Iriguchi, S. et al. A clinically applicable and scalable method to regenerate T-cells from iPSCs for off-the-shelf T-cell immunotherapy. Nat. Commun. 12, 430 (2021).
Canté-Barrett, K. et al. Loss of CD44dim expression from early progenitor cells marks T-cell lineage commitment in the human thymus. Front. Immunol. 8, 32 (2017).
Lavaert, M. et al. Integrated scRNA-Seq identifies human postnatal thymus seeding progenitors and regulatory dynamics of differentiating immature thymocytes. Immunity 52, 1088–1104.e6 (2020).
Li, L.-C. et al. Single-cell patterning and axis characterization in the murine and human definitive endoderm. Cell Res. 31, 326–344 (2021).
D’Amour, K. A. et al. Efficient differentiation of human embryonic stem cells to definitive endoderm. Nat. Biotechnol. 23, 1534–1541 (2005).
Davenport, C., Diekmann, U., Budde, I., Detering, N. & Naujok, O. Anterior–posterior patterning of definitive endoderm generated from human embryonic stem cells depends on the differential signaling of retinoic acid, Wnt-, and BMP-signaling. Stem Cells 34, 2635–2647 (2016).
Balciunaite, G. et al. Wnt glycoproteins regulate the expression of FoxN1, the gene defective in nude mice. Nat. Immunol. 3, 1102–1108 (2002).
Barbarulo, A. et al. Hedgehog signalling in the embryonic mouse thymus. J. Dev. Biol. 4, 22 (2016).
Bleul, C. C. & Boehm, T. BMP signaling is required for normal thymus development. J. Immunol. 175, 5213–5221 (2005).
Lucas, B. et al. Progressive changes in CXCR4 expression that define thymocyte positive selection are dispensable for both innate and conventional αβT-cell development. Sci. Rep. 7, 5068 (2017).
Farley, A. M. et al. Thymic epithelial cell fate and potency in early organogenesis assessed by single cell transcriptional and functional analysis. Front. Immunol. 14, 1202163 (2023).
Fang, X. et al. MIR148A family regulates cardiomyocyte differentiation of human embryonic stem cells by inhibiting the DLL1-mediated NOTCH signaling pathway. J. Mol. Cell. Cardiol. 134, 1–12 (2019).
Ivey, K. N. et al. MicroRNA regulation of cell lineages in mouse and human embryonic stem cells. Cell Stem Cell 2, 219–229 (2008).
Lucas, B. et al. Embryonic keratin19+ progenitors generate multiple functionally distinct progeny to maintain epithelial diversity in the adult thymus medulla. Nat. Commun. 14, 2066 (2023).
Lopes, N., Sergé, A., Ferrier, P. & Irla, M. Thymic crosstalk coordinates medulla organization and T-cell tolerance induction. Front. Immunol. 6, 365 (2015).
Pinto, S. et al. An organotypic coculture model supporting proliferation and differentiation of medullary thymic epithelial cells and promiscuous gene expression. J. Immunol. 190, 1085–1093 (2013).
Campinoti, S. et al. Reconstitution of a functional human thymus by postnatal stromal progenitor cells and natural whole-organ scaffolds. Nat. Commun. 11, 6372 (2020).
Hun, M. et al. Native thymic extracellular matrix improves in vivo thymic organoid T cell output, and drives in vitro thymic epithelial cell differentiation. Biomaterials 118, 1–15 (2017).
Fan, Y. et al. Bioengineering thymus organoids to restore thymic function and induce donor-specific immune tolerance to allografts. Mol. Ther. 23, 1262–1277 (2015).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
Williams, J. A., Tai, X. & Hodes, R. J. CD28-CD80/86 and CD40-CD40L interactions promote thymic tolerance by regulating medullary epithelial cell and thymocyte development. Crit. Rev. Immunol. 35, 59–76 (2015).
Yang, S., Fujikado, N., Kolodin, D., Benoist, C. & Mathis, D. Immune tolerance. Regulatory T cells generated early in life play a distinct role in maintaining self-tolerance. Science 348, 589–594 (2015).
Mellins, E. D. & Stern, L. J. HLA-DM and HLA-DO, key regulators of MHC-II processing and presentation. Curr. Opin. Immunol. 26, 115–122 (2014).
Lepletier, A. et al. Interplay between follistatin, activin A, and BMP4 signaling regulates postnatal thymic epithelial progenitor cell differentiation during aging. Cell Rep. 27, 3887–3901.e4 (2019).
Ramos, S. A. et al. Generation of functional thymic organoids from human pluripotent stem cells. Stem Cell Rep. 18, 829–840 (2023).
Hosaka, N., Kanda, S., Shimono, T. & Nishiyama, T. Induction of γδT cells from HSC-enriched BMCs co-cultured with iPSC-derived thymic epithelial cells. J. Cell Mol. Med. 25, 10604–10613 (2021).
Azzam, H. S. et al. CD5 expression is developmentally regulated by T cell receptor (TCR) signals and TCR avidity. J. Exp. Med. 188, 2301–2311 (1998).
von Boehmer, H. & Fehling, H. J. Structure and function of the pre-T cell receptor. Immunology 15, 433–452 (1997).
Staal, F. J. T., Weerkamp, F., Langerak, A. W., Hendriks, R. W. & Clevers, H. C. Transcriptional control of T lymphocyte differentiation. Stem Cells 19, 165–179 (2001).
Douagi, I., André, I., Ferraz, J. & Cumano, A. Characterization of T cell precursor activity in the murine fetal thymus: evidence for an input of T cell precursors between days 12 and 14 of gestation. Eur. J. Immunol. 30, 2201–2210 (2000).
Kraft, D. L., Weissman, I. L. & Waller, E. K. Differentiation of CD3-4-8- human fetal thymocytes in vivo: characterization of a CD3-4+8- intermediate. J. Exp. Med. 178, 265–277 (1993).
Flippe, L. et al. Rapid and reproducible differentiation of hematopoietic and t cell progenitors from pluripotent stem cells. Front. Cell Dev. Biol. 8, 577464 (2020).
Inami, Y. et al. Differentiation of induced pluripotent stem cells to thymic epithelial cells by phenotype. Immunol. Cell Biol. 89, 314–321 (2011).
Gaignerie, A. et al. Urine-derived cells provide a readily accessible cell type for feeder-free mRNA reprogramming. Sci. Rep. 8, 14363 (2018).
Kilens, S. et al. Parallel derivation of isogenic human primed and naive induced pluripotent stem cells. Nat. Commun. 9, 360 (2018).
Charpentier, E. et al. 3’ RNA sequencing for robust and low-cost gene expression profiling. https://doi.org/10.21203/rs.3.pex-1336/v1 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Xu, J. et al. Genotype-free demultiplexing of pooled single-cell RNA-seq. Genome Biol. 20, 290 (2019).
Pan, J.-B., Hu, S.-C., Wang, H., Zou, Q. & Ji, Z.-L. PaGeFinder: quantitative identification of spatiotemporal pattern genes. Bioinformatics 28, 1544–1545 (2012).
Provin, N. GeneTissueSpe: combinatory differentiation of human induced pluripotent stem cells generates functional thymic epithelium driving dendritic- and CD4/CD8 T-cell development. https://doi.org/10.5281/zenodo.17674360 (2025).
Acknowledgements
This work was supported by the EJP-Rare Disease JTC2019 program TARID project (ANR-19-RAR4-0011-05), “la Région Pays de la Loire” through the RFI Bioregate grant (ThymIPS), the NORD (The APS Type 1 Foundation grant) ID: 22001 and the “SATT Ouest Valorisation” project OrgaTreg to M.G., as well as “la Fondation pour la Recherche Médicale” (FRM) under grant Equipe FRM EQU202303016266 to C.G. and M.G. This work was also partially supported by the ANR SelfExpress (ANR-22-CE15-0045-01) and by "la Région Pays de la Loire" through the Trajectoire Nationale program (2023_01170) to M.G. N.P. was supported by “la Fondation d’entreprise ProGreffe”. M.d.A. received PhD fellowship support from the FRM (grant no. 17366). This work was partially funded by the Labex IGOprogram supported by the National Research Agency via the investment program ANR-11-LABX-0016-01. We thank the members of the “Genomic’IC” core facility headed by Franck Letourneur at Cochin Institute, Paris, France, for bulk RNA-seq data production, as well as, Drs Pärt Peterson, Julia Maslovskaja and Kai Kisand from Tartu University, Estonia, for human TEC RNA-seq datasets. We thank the iPSC and the MicroPICell core facilities of Nantes supported by IBiSA and Biogenouest, for the use of their resources and technical support. We specifically thank Caroline Chariau and Isabelle Leray (iPSC facility), as well as Anne Gaignerie (MicroPICell) for her advice on iPSc-to-HSPC differentiation, and Laurence Delbos for her advice on flow cytometry. We also thank Ella Beaumann for bioinformatics assistance and Léana Lecompte and Teresa Rodriguez-Trenchs for their help in iPSc culture and differentiation.
Author information
Authors and Affiliations
Contributions
N.P. and M.G. designed the study and wrote the manuscript; N.P., M.d.A. performed most of the experimental work; A.L.B. performed key HSPC experiments; A.B., L.B., C.F., J.P., and P.H. performed experiments; N.P., E.K., and M.G. performed bioinformatics analyses; P.M., O.B., and X.S. provided key material; C.G. and L.D. contributed to the study design and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kaomei Guan and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Provin, N., d’Arco, M., Le Bozec, A. et al. Combinatory differentiation of human induced pluripotent stem cells generates functional thymic epithelium driving dendritic cell and CD4/CD8 T cell development. Nat Commun 17, 1969 (2026). https://doi.org/10.1038/s41467-026-68675-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68675-y