Abstract
Drug-resistant bacterial infections pose a serious threat to global health, driving the development of antibacterial strategies beyond classic antibiotics. Host defense peptide mimetic polymeric antibiotics have emerged as promising candidates to combat drug resistance, however, navigating the vast chemical space of polymers remains a significant challenge due to complex structure–activity relationships, while data-driven approaches are further constrained by polymer complexity and scarce labeled data. To address this, we develop PolyCLOVER, a framework that integrates multi-stage self-supervised learning, active learning, and high-throughput experimentation to iteratively discover polymeric antibiotics with potent antibacterial activity and low toxicity. Applied to a combinatorial library of ~100,000 poly(β-amino ester)s, the framework uncovers three lead compounds that self-assemble into stable nanoparticles (SANPs) with minimum inhibitory concentrations of 4 μg/mL and 8 μg/mL against multidrug-resistant S. aureus and A. baumannii, respectively. These SANPs also serve as adjuvant antibiotic carriers, restoring bacterial sensitivity to penicillin G. In vivo studies demonstrate their therapeutic efficacy both as monotherapies and in combination therapies with antibiotics. PolyCLOVER may become a powerful framework for discovery of new polymeric biomaterials without reliance on external datasets.
Similar content being viewed by others
Introduction
Antimicrobial resistance (AMR), particularly of bacterial origin, poses an escalating global health threat, with 4.71 million associated and 1.14 million attributable deaths reported in 20211,2,3. Current strategies to combat bacterial AMR primarily focus on two approaches: developing novel antibiotics to bypass resistance4,5,6, and employing combination therapies to restore efficacy, including combinations of existing antibiotics and co-administration of antibiotic adjuvants7,8,9,10. With antibiotic discovery becoming increasingly challenging11, host defense peptides (HDPs), or antimicrobial peptides (AMPs), have attracted attention as alternative therapeutics owing to their ability to disrupt membranes, modulate immunity, and regulate inflammation, thereby slowing resistance development12,13,14. However, their clinical translation is limited by inherent biological toxicity, poor in vivo stability, and high production costs15,16,17. In this context, synthetic polymeric antibiotics inspired by AMP mechanisms can achieve comparable antimicrobial efficacy, while their flexible and scalable synthesis addresses the limitations of AMPs in stability and manufacturability18,19,20,21,22,23. Moreover, their broad chemical design space enables antibacterial activity as standalone agents and synergistic effects as antibiotic adjuvants, together with extendable capabilities such as drug loading, barrier penetration, and targeted delivery24,25. These characteristics make synthetic polymeric antibiotics a particularly promising platform for next-generation antimicrobial design.
Current strategies for polymeric antibiotic discovery remain predominantly empirical, relying on modulation of charge and amphiphilicity coupled with inefficient trial-and-error screening26,27. The complexity of antimicrobial mechanisms and the structural diversity of candidate polymers hinder mechanistic understanding, limiting the elucidation of structure–activity relationships and posing a major barrier to the systematic exploration of this vast chemical space. Deep learning (DL), with its capacity to model complex, nonlinear relationships, has emerged as a powerful tool for data-driven prediction and optimization. DL has achieved notable success in AMP discovery28,29,30, including our previous work on empirical 6–9-mer peptide sequences31, and has further accelerated omics research by improving the throughput and efficiency of genome and proteome mining32,33,34,35,36. Despite this, DL applications to polymeric antibiotics remain scarce37, primarily due to challenges in data availability, structural diversity, and molecular representation. AMPs, derived from diverse natural sources and extensively studied, are supported by large, labeled datasets (typically >105 sequences). Their conserved backbone and standardized sequence representation further facilitate model learning. Polymeric antibiotics, by contrast, suffer from limited and fragmented data resources; existing data often vary in synthesis conditions, characterization protocols, and structural distributions, making it difficult to support the development of generalizable models. The high tunability of polymer backbones and side chains further complicates feature extraction and demands greater model generalization, while current representations remain inadequate for capturing their structural complexity. Nonetheless, this flexibility presents a unique opportunity to design multifunctional polymeric antibiotics.
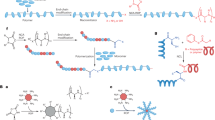
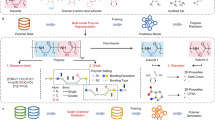
Herein, we propose a comprehensive framework (PolyCLOVER) for the design of polymeric antibiotics with integrated adjuvant effects to combat antibacterial drug resistance. To overcome the challenges posed by data availability, structural diversity, and molecular representation, we employ two critical designs in the framework. 1) A graph-based polymer foundation model, developed in our recent work38, is applied to capture the intricate structural characteristics of polymer systems, thereby enhancing predictive performance. 2) Multi-stage self-supervised learning, active learning, and high-throughput experiments are integrated to form an autonomous evolution workflow, eliminating the need for a pre-established dataset. Within a combinatorial library, PolyCLOVER can identify polymeric antibiotics that exhibit both potent antibacterial activity and low toxicity, enabling efficient navigation in a large chemical space without reliance on external datasets. The ensemble multi-task predictor recommends candidate combinations based on properties, diversity, and uncertainty, which are then labeled through experiments. The resulting data are used to update the pretrained model, which then predicts the next round of candidates. (Fig. 1a). When applied to a combinatorial poly(β-amino ester) library comprising about 100,000 candidates (Fig. 1b), three polymers with outstanding antibacterial efficacy against a wide range of clinically isolated drug-resistant pathogens and low toxicity are successfully identified. These leading compounds self-assemble into stable nanoparticles (SANPs), which facilitate in vivo transportation and distribution. Their potent therapeutic efficacy as polymeric antibiotics is validated in a mouse pneumonia model. Importantly, the candidate polymer showed ultra-high encapsulation efficacy and loading capacity when co-assembled with penicillin G. The co-assembled NPs showed a synergistic antibacterial effect and restored the sensitivity of drug-resistant Gram-positive pathogens to penicillin G, demonstrating a synergistic therapeutic effect in a mouse peritonitis model (Fig. 1c). Taken together, the discovered polymers are promising candidates as next-generation dual-functional antibiotics, and suggest that PolyCLOVER can be a viable approach for the efficient discovery of polymeric nanomedicine.
a PolyCLOVER workflow. A graph encoder was pretrained using a two-stage, multi-task self-supervised learning based on the constructed library. An initial subset of the library was sampled, followed by high-throughput synthesis and characterization of antibacterial activity and hemolytic rate to generate a labeled dataset. The graph encoder was then fine-tuned to form an ensemble predictor consisting of 20 base learners. The predictor estimated the activity and predictive uncertainty of unlabeled SANPs in the library, guiding the iterative selection of new candidates via UCB acquisition and K-Means clustering. This process was iteratively repeated. Created in BioRender. Ji, J. (2025) https://BioRender.com/egwvanc. b Combinatorial library construction. The library was constructed based on Michael addition between diacrylate and amine monomers. 8 diacrylates and 38 amines were selected based on solubility and cost from commercially available sources, including 16 amines with tertiary amine side groups or Boc-protected primary amine groups, 16 with hydrophobic side groups, and 6 with hydrophilic side groups. The ratios of B, C, and D were controlled within appropriate ranges to drive self-assembly, resulting in the formation of ~100,000 poly(β-amino esters) SANPs. c In vitro/in vivo validation. Additional in vitro experiments were conducted to characterize the detailed antibacterial activity, cytotoxicity, and particle size of the selected SANPs. The in vivo therapeutic efficacy of the SANPs, either alone or in combination with penicillin G, was validated in pneumonia and peritonitis models.
Results
Multi-stage self-supervised learning boosts model performance on limited data
Previous studies have demonstrated that positive charges present in polymers can effectively interact with the negatively charged bacterial cell membrane, while groups such as alkyl chains can insert into the bacterial cell membrane and disrupt its normal physiological function, thereby achieving a bactericidal effect39,40,41. Accordingly, positively charged and hydrophobic units were incorporated into polymers to impart potential antibacterial activity. We also introduced hydrophilic units and regulated the ratios of each component in a suitable range to promote the self-assembly of the polymers into stable SANPs. Guided by this design rationale, a combinatorial library comprising 98,304 (~100,000) poly(β-amino ester) SANPs was constructed via a multi-component Michael addition reaction (Supplementary Tables S1 and 2).
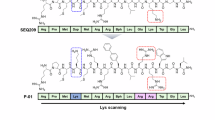
Next, we randomly selected 220 samples from the constructed library for laboratory synthesis and testing of their antibacterial and hemolytic properties to form the initial dataset. The initial dataset spans a vast chemical space, incorporating all monomer types presented in the library (Fig. 2a, Supplementary Fig. S1). We chose Methicillin-resistant Staphylococcus aureus (MRSA) as the model pathogen because it stands as one of the most prevalent pathogens, capable of causing life-threatening diseases such as sepsis, pneumonia, and multi-organ failure42. The antibacterial activity against MRSA and hemolytic behavior of the samples were quantitatively assessed using optical density (OD) values at 595 nm and the hemolysis ratio, respectively. The results reveal that the probability of identifying effective SANPs through random sampling in the extensive combinatorial library is very low. Most SANPs exhibit weak antibacterial activity, while a few samples with high antibacterial activity show high hemolytic toxicity (Fig. 2b). These findings underscore the considerable challenge of achieving a balance between high antibacterial activity and low hemolytic toxicity, highlighting the importance of ML in guiding the design of antibacterial SANPs.
a The number of different types of building blocks in the initial labeled dataset. The structures corresponding to the building block IDs can be found in Supplementary Table 1. b The distribution of antibacterial activity and hemolytic property in the initial dataset. Marginal histograms along the top and right axes show the distributions of each property. Contour lines were plotted using kernel density estimation with a threshold of 0.15. Lower values on axes indicate stronger antibacterial activity and better hemocompatibility. c Average test performance of different backbones on antibacterial and hemolytic properties. The mean value of the test root mean square error on three independent runs is reported. d Data preprocessing and model architecture. The Graph Transformer was pretrained sequentially on an unlabeled polymer dataset and the constructed unlabeled NP dataset to learn rich structural features. It was then fine-tuned using the labeled NP dataset to predict antibacterial and hemolytic properties. e t-SNE visualization of representations generated by the pretrained and unpretrained graph encoder. Each data point is colored according to the type of diacrylate used. f Test performance of the proposed approach. Pearson correlation coefficients are shown as 1-R to facilitate visualization on the radar plot, where a smaller enclosed area indicates better performance. The mean value of the test root mean square error on three independent runs is reported. g The measured value versus predicted plots for the proposed model. The dashed line represents the ideal case, where y = x. The Pearson correlation coefficient is shown in the lower right corner. Source data are provided as a Source Data file.
After collecting the initial dataset, we employed graph neural networks (GNNs) for modeling, due to their strong capability in capturing molecular structural features43,44. However, the intrinsic complexity of copolymers, particularly the absence of a well-defined and consistent topology, makes it challenging to represent them using the vanilla molecular graph. To enable effective message passing between subgraphs corresponding to individual building blocks, virtual nodes were introduced as communication bridges, following our previously reported approach38. In addition, three local virtual nodes were connected to their respective subgraphs to encode the copolymerization ratios (see ‘Dataset construction and preprocessing’ section in Methods). We evaluated the average test performance of 10 commonly used GNN backbones on antibacterial and hemolytic tasks and finally selected a graph transformer45 with multi-head attention (Fig. 2c, Supplementary Table S3).
Self-supervised learning has demonstrated remarkable success across various domains of artificial intelligence by leveraging intrinsic structural information in large-scale unlabeled data, enabling models to learn general representations that can be effectively transferred to downstream tasks46,47,48,49. This paradigm significantly reduces reliance on manual annotations and improves model generalization. To this end, we employed a periodicity-aware pretraining strategy to learn rich and generalizable representations from a large corpus of unlabeled general polymer data38. Building on this, our pool-based screening strategy enables the predictive model to access candidate SANPs that will be evaluated during the inference phase. That is, the model is exposed to the candidate set during training, effectively establishing a transductive learning scenario. Therefore, inspired by domain-specific large language models, we here propose a two-stage pretraining approach, in which the model undergoes continual pretraining on the entire accessible unlabeled library of poly(β-amino ester)s to further exploit the structural priors inherent in the screening space and to bridge the gap between pretraining and fine-tuning domains (see ‘Model construction’ section in Methods for full details). The ratio information of individual building blocks was extracted using a ratio embedding layer and assigned as the initial features of the corresponding virtual nodes (Fig. 2d).
Subsequently, we analyzed the model’s latent space to evaluate the effect of the two-stage pretraining. To visualize the learned polymer representations in two-dimensional space, we applied t-SNE—a widely used dimensionality reduction algorithm that preserves local similarities—to both the pretrained and unpretrained models (Fig. 2e; Supplementary Fig. S2–6). It can be observed that the representations generated by the pretrained model are evenly distributed in the latent space and exhibit clear clustering patterns, where structurally similar SANPs (the same diacrylates) show high consistency in their representations. In the unpretrained model, the representations are compressed into narrow regions, indicating limited structural awareness and reduced information retention. These results demonstrate that the pretrained model effectively captures key structural features while preserving maximal information50.
The pretrained model was then fine-tuned on the initial dataset to predict the antibacterial and hemolytic properties of SANPs. We compared the test performance of the two-stage self-supervised learning approach with baseline methods (Fig. 2f, g, Supplementary Figs. S7, 8). Due to the extreme scarcity of data, the randomly initialized model fails to capture structural patterns, resulting in poor performance. The weighted sum approach used ratio-weighted building block representations extracted by the pretrained encoder, followed by a prediction head. Its performance showed a slight improvement but remained inadequate. In contrast, our proposed approach achieves significantly better performance across all metrics, demonstrating its effectiveness under data-limited conditions. Moreover, the fine-tuned latent space reveals an ordered, continuous distribution that aligns with activity levels, reflecting better capture of the underlying structure–activity relationship (Supplementary Figs. S9, 10). Therefore, we adopt this model as the base learner for subsequent structure optimization of SANPs.
PolyCLOVER drives iterative improvements in antibacterial activity and hemocompatibility
Although the model was optimized, the limited amount of training data still poses a significant challenge for generalizing across the full combinatorial library. To address this, PolyCLOVER constructs an ensemble predictor composed of 20 homogeneous base learners, which is then embedded into a Bayesian optimization-based active learning framework. This pipeline facilitates efficient exploration of the vast chemical design space of SANPs to identify structures with potentially optimal properties.
Then, the ensemble predictor was employed to predict the activities of unlabeled SANPs in the remaining combinatorial library. The mean prediction across base learners was used as the activity estimate, while the standard deviation served as an estimate of predictive uncertainty. Each candidate was then scored using the Upper Confidence Bound (UCB) acquisition function, which balances exploration of uncertain regions with exploitation of potentially optimal candidates. To further improve optimization efficiency, we applied the K-means algorithm to cluster the top-ranked SANPs and selected representative structures from each cluster to enhance data diversity. PolyCLOVER then recommended 60 candidate SANPs for wet-lab synthesis and characterization. The newly generated data were subsequently incorporated into the initial dataset to support the next round of optimization (Fig. 3a; see ‘Active learning strategy’ section in Methods for full details).
a Time spent in each step of PolyCLOVER for identified SANPs screening. Heat maps of antibacterial (b) and hemolytic (c) properties in each round. d Density plots of antibacterial and hemolytic properties in each round. Lower values represent better antibacterial activity and improved hemocompatibility. Each round includes n = 60 (initial round n = 220) structurally distinct samples. The contour shows the kernel density estimation, with a white dot for the median, a thick bar for the interquartile range, and thin lines for the 95% confidence intervals. This visualization scheme is also applied in (e) and (f). The evolution of average LogP (e) and positive charge ratio (f) over iterative optimization rounds. The LogP value of each NP is calculated as a weighted sum of the LogP values of its building blocks according to their compositional ratios. g The frequency of individual building blocks over iterative optimization rounds. Boxed insets show the structures of B10 and C16. h The MIC value and selectivity index of the SANPs that meet the criterion. Samples above the dashed line represent MIC greater than 32 µg/mL. The selectivity index is calculated as the ratio of HC50 to MIC. Source data are provided as a Source Data file.
After four rounds of iterative optimization, the activities of the recommended SANPs plateaued, suggesting convergence of the active learning loop (Fig. 3b–d, Supplementary Figs. S11–15). At this stage, a greedy acquisition strategy was employed to further exploit the most promising candidates, without considering uncertainty or diversity. Compared to the initial dataset, the antibacterial activity of the selected SANPs showed a substantial improvement. Although the hemolytic rate increased slightly in the intermediate rounds due to the enhancement of antibacterial potency, it was successfully reduced to an acceptable level in the final round of optimization. As a result, a substantial number of SANPs identified in the final round exhibited both high antibacterial efficacy and low hemolytic toxicity.
To gain deeper insight into the model’s behavior, we further investigated the evolution of chemical structures and key physicochemical properties of the SANPs recommended by PolyCLOVER. In the first round, an increase in LogP indicated a shift toward higher hydrophobicity, suggesting that the model initially prioritized antibacterial activity. However, excessive hydrophobicity (e.g., from longer alkyl chains) was often associated with a higher potential for cell toxicity21, the dual-objective task subsequently regulated this property to balance antibacterial activity and biocompatibility (Fig. 3e). Additionally, the initially low ratio of positive charges, which may have limited interactions with negatively charged bacterial membranes, was substantially increased during optimization (Fig. 3f). A sharp increase in the de-protected primary amine of B10 suggested that the model favored a short chain length of positive charge (Fig. 3g), a tendency that has also been noted in some other polymer systems41,51. We also found the unique integration of polyfluorocarbon unit (C16) effectively preserved antibacterial efficacy while reduced toxicity, which aligned with previous studies52. Nevertheless, this moiety falls under the category of per- and polyfluoroalkyl substances (PFAS), and warrants attention in practical applications.
To further identify SANPs, we established a preliminary criterion for high-potential candidates: an antibacterial optical density (OD) value of less than 0.15 and a hemolytic rate of less than 50%. All SANPs meeting this criterion were synthesized and purified in a single batch, followed by detailed evaluation of their antibacterial and hemolytic properties. The minimum inhibitory concentration (MIC) and half-hemolytic concentration (HC50) were determined for each polymer, and the selectivity index was calculated as the ratio of HC50 to MIC (Fig. 3h, Supplementary Table S4). The results showed that most SANPs exhibited strong antibacterial activity and low hemolytic toxicity, with selectivity indices exceeding eight. Finally, samples with a selectivity index greater than 20 were selected, resulting in the identification of three optimal SANPs. Notably, PolyCLOVER enabled the discovery of three highly antibacterial and low-toxic SANPs from a combinatorial library of ~100,000 polymer candidates in just 20 days, demonstrating its remarkable efficiency in accelerating the discovery of new functional materials within large design spaces.
Identified SANPs exhibit promising in vitro performance
The identified SANPs were designated as H1, H2, and H3 (Fig. 4a, Supplementary Fig. S16). The positively charged amines (B) used in the formulation consisted of deprotected primary amines with short chains. The hydrophobic amines (C) featured either a relatively short tertiary alcohol or a short chain with fluorinated side groups, which likely mitigated the hemolytic toxicity associated with longer hydrophobic chains. The hydrophilic amines (D) included oligomeric polyethylene glycol (PEG) side groups in H1 and H3, while H2 contained a unique amide side group. Notably, the proportion of hydrophilic components was the lowest among the three segments, while the proportion of positively charged groups was the highest, which is crucial for interaction with negatively charged bacterial cell membranes. Additionally, due to an excess of diacrylate during polymer preparation, all three SANPs retained double bond residues at both ends, allowing for potential future functional modifications. The MIC of all three SANPs were 8 μg/mL against MRSA ATCC 43300. Both the HC50 (H1 = 168 μg/mL, H2 > 512 μg/mL, H3 = 266 μg/mL) and the half-maximal inhibitory concentration (IC50; H1 = 53 μg/mL, H2 = 162 μg/mL, H3 = 109 μg/mL; assessed in L929 fibroblast cells) followed a consistent trend, which was also observed in the MLE-12 lung epithelial cell line, with H2 exhibiting the lowest toxicity among the three (Supplementary Fig. S17). These findings underscore the necessity of leveraging a dual-objective optimization strategy to achieve a balance between antibacterial efficacy and toxicity. The self-assembly properties of H1–3 were confirmed by dynamic light scattering (DLS) and transmission electron microscopy (TEM). Three SANPs exhibited particle diameters of ~100 nm, with H2 having a larger diameter compared to H1 and H3, and all three SANPs carried positive charges (Fig. 4b).
a Detailed information of the selected identified SANPs (H1, H2, H3), including molecular structure, monomer ratio, MIC values, and hemolytic toxicity. b Size distribution, zeta potentials, and TEM images of identified SANPs. Scale bar: 1 μm. c Bactericidal performance of SANPs and SM at 32 µg/mL for 9 h incubation. Statistical analysis was performed using one-way ANOVA. d Changes in MIC values of identified SANPs in TSB and FBS. e Bactericidal kinetics of identified SANPs and SM at 32 µg/mL within 100 min. f Evolution of resistance to identified SANPs and SM in MRSA after 28 days of passaging in TSB media. g MIC values (µg/mL) of the identified SANPs against 3 reference strains (S. aureus ATCC 29213, S. epidermidis ATCC 35984, and A. baumannii ATCC 19606) and 11 clinically isolated drug-resistant pathogens (S. aureus-MRSA, VISA, S. capitis-MRSC, E. faecalis-VRE, A. baumannii-MDRAB, P. aeruginosa-CRPA, K. pneumoniae-CRKP). Gram-positive and Gram-negative bacteria are shown on the left and right, respectively. Each test was performed in three independent experiments. h TEM images of MRSA after incubation with identified SANPs for 10 min at 64 µg/mL. Scale bar: 200 nm. i Live/dead bacterial staining of MRSA after incubation with identified SANPs for 15 min at 64 µg/mL. Control: PBS treatment only; no antimicrobial agent applied. Scale bar: 20 μm. j Fluorescence analysis of membrane potential disruption induced by identified SANPs and SM at 64 µg/mL. Statistical analysis was conducted using one-way ANOVA. k Molecular dynamics simulation of the interaction between H2 and the cell membrane. In panels (b, h, i), experiments were performed in triplicate, yielding consistent results, and a representative image is shown. In panels (b, c, e, f, j), data are shown as mean ± s.d. (n = 3 independent replicates). Source data are provided as a Source Data file.
We further evaluated the bactericidal efficacy of the identified SANPs by benchmarking them against streptomycin (SM), a broad-spectrum antibiotic. After 9 h of co-incubation at 32 μg/mL, H1 and H2 reduced MRSA viability by 3 orders of magnitude, while H3 achieved a 4-order reduction (Fig. 4c). SM exhibited the strongest antibacterial effect, with a 5-order reduction. Meanwhile, when the tryptic soy broth (TSB) medium was replaced with fetal bovine serum (FBS), a more complex environment, we found that the SANPs maintained their antibacterial efficacy, with MIC values fluctuating by only 1-2 fold. This stability may be attributed to the biologically inert fluorinated and PEG side chains (Fig. 4d). Next, we assessed the bactericidal kinetics of the SANPs, which is another key metric in the practical application of antibacterials. (Fig. 4e). All three SANPs demonstrated rapid bactericidal action. Notably, H1 and H2 eliminated 90% of bacteria within 60 min, while H3 achieved this reduction in just 4 min. In contrast, SM exhibited significantly slower killing kinetics. As an antibiotic that inhibits bacterial protein synthesis by targeting ribosomes53, SM requires prolonged exposure, aligning with prior reports that antibiotics generally exert bactericidal effects over several hours or longer54. These findings highlighted that the identified SANPs not only outperformed antibiotics in bactericidal speed but also likely operated via a different antibacterial mechanism. Furthermore, compared to SM, the SANPs exhibited a substantially lower tendency for resistance development. Over a 28-day testing period, the MIC values for H1–3 fluctuated modestly (1–4 fold), while SM showed a significant 28-fold increase in MIC (Fig. 4f). Among 11 clinically isolated drug-resistant pathogens, the SANPs exhibited antibacterial efficacy comparable to that observed in reference strains. In particular, the SANPs exhibited potent activity against all tested Gram-positive bacteria, whereas activity among Gram-negative strains was restricted to A. baumannii. These results indicate that the antibacterial spectrum of the SANPs is selective and biased towards Gram-positive pathogens. (Fig. 4g).
Subsequently, we investigated the antibacterial mechanism of the identified SANPs. TEM images were obtained after 5 min of co-incubation with the identified SANPs (Fig. 4h). In the control group, the MRSA membrane maintained a clear and intact structure. In contrast, after interaction with the SANPs, significant membrane disintegration was observed, leading to the release of bacterial cell contents and resulting in bacterial death. Live/dead bacterial staining assays were performed after 5 min of exposure to SANPs H1–3 (Fig. 4i). The assays employed N01, which penetrated both live and dead bacteria, binding to nucleic acids and emitting green fluorescence, while propidium iodide (PI) entered only damaged or dead bacteria, binding to DNA and emitting red fluorescence. Confocal fluorescence microscopy revealed intense red fluorescence in MRSA exposed to SANPs H1–3, even after a short incubation period. To assess their effect on bacterial membrane potential, we used the fluorescent probe DiSC3(5) (Fig. 4j). This probe accumulates in intact bacterial membranes, leading to fluorescence quenching. When the membrane is disrupted, the probe leaks from the membrane, causing fluorescence emission as its concentration decreases. Bacterial suspensions without SANPs exhibited negligible fluorescence intensity. In contrast, the addition of Triton X-100, a strong surfactant, resulted in a significant increase in fluorescence emission due to membrane destabilization. SANPs H1–3 induced robust fluorescence signals, confirming their ability to disrupt bacterial membrane potential. SM, however, did not induce fluorescence, consistent with its non-membrane-targeting antibacterial mechanism. The membrane-attaching mechanism of the SANPs was further revealed by molecular simulations. All-atom molecular dynamics results demonstrate that when the SANPs approach the cell membrane, they rapidly adsorb to the surface. This process is primarily driven by electrostatic interactions. (Fig. 4k, Supplementary Fig. S18). Analysis of these results suggests that the identified SANPs likely exert their bactericidal effects through strong interactions with the cell membranes of MRSA in a short timeframe.
Identified SANPs eradicate lung infection with high efficacy and biocompatibility
The therapeutic efficacy of the identified SANPs was further investigated in a mouse lung infection model. MRSA-induced acute pneumonia is among the most encountered lung infections, featuring a high incidence rate and significant mortality55. Female mice were inoculated with MRSA to establish an acute pneumonia model by intratracheal instillation (Fig. 5a). The identified SANPs were administered via tail intravenous injection (IV injection) on the following day. Mice were euthanized one day post-treatment, and lung tissues were collected for morphological observation and bacterial counting. The PBS-treated group presented unhealthy dark red lung appearances. In contrast, the lungs of the identified SANPs-treated group appeared healthy and pink. Bacterial counts from tissue homogenates indicated that the SANPs-treated group demonstrated significant improvement, reducing bacterial counts by three orders of magnitude, comparable to the results observed in the SM-treated group (Fig. 5b, c). The results of H&E staining revealed that the alveolar structure was collapsed in the PBS treatment group. In contrast, it remained porous in the SANPs-treated group, indicating the relief of lung infection. Additionally, Masson staining showed no significant blue fiber deposition, further suggesting that H1–3 exhibited a favorable therapeutic effect. And CD68 staining demonstrated that H1–3 modulated the excessive inflammatory response. (Fig. 5d).
a Schematic illustration of the acute pneumonia model construction and treatment protocol in mice. On day 0, mice were anesthetized with isoflurane, and MRSA suspension (108 CFU/mL, 60 μL) was administered intratracheally to induce infection. On day 1, the identified polymers H1, H2, H3 and SM were administered via tail vein injection at a dose of 5 mg/kg. On day 2, the mice were euthanized, and their lungs were excised for morphological observation. The lung tissue was homogenized for further bacterial colony counts. Created in BioRender. Ji, J. (2025) https://BioRender.com/qnaeui1. b Visual examination of mouse lung appearance and bacterial colony culture from tissue homogenate to assess infection. c Quantification of lung bacterial load through colony counting in infected mice. d Histological analysis of lung tissues using H&E, Masson, and CD68 staining to evaluate the extent of tissue damage, fibrosis, and inflammation. Scale bar: 300 nm. e, f Quantitative results and fluorescence imaging of the distribution of SANPs within the mice to determine tissue targeting efficiency. In panels (b) and (d), experiments were performed in triplicate, with consistent results, and one representative image is shown. In panels (c) and (f), data are expressed as mean ± s.d. (n = 3 independent replicates). Statistical analysis was performed using one-way ANOVA. Source data are provided as a Source Data file.
To examine the distribution of the SANPs in mice post-injection, rhodamine B was conjugated to the SANPs through a reaction between isothiocyanate and the amine groups (Supplementary Fig. S19). Statistical results and fluorescence images of mouse organs were captured 2 h post-injection (Fig. 5e, f). The results revealed that most of the fluorescence was concentrated in the liver, with a notable presence in the lungs, indicating that the SANPs effectively reached lung tissue via the tail vein injection. Next, we assessed the tissue toxicity of the SANPs (Supplementary Fig. S20). Histological analysis of heart, liver, spleen, lung, and kidney samples from SANPs-treated mice revealed no significant tissue necrosis compared to the PBS control group. The lung sections exhibited a healthy, loose alveolar structure, and no damage was observed in the glomeruli or renal tubules of the kidney sections. In conclusion, the identified SANPs demonstrated no evident tissue toxicity.
Identified SANPs restore MRSA susceptibility to penicillin G
The above investigation demonstrated the intrinsic antibacterial activity and self-assembly properties of the identified SANPs. Furthermore, we evaluated the potential of the SANPs as antibiotic adjuvants. It was anticipated that abundant functional groups within the SANPs could be integrated with antibiotics to form nanocomposites, thus enhancing the penetration of antibiotics through bacterial membranes. MRSA resistance primarily arises from the acquisition of the mecA gene, which encodes penicillin-binding protein 2a (PBP2a)56. PBP2a has a significantly reduced affinity for β-lactam antibiotics, such as penicillin G (PG), making these agents largely ineffective against MRSA. The integrated nanocomposites held promise for bypassing this resistance barrier by facilitating more effective antibiotic delivery.
We demonstrated that the SANPs efficiently encapsulated PG, thereby reinstating the susceptibility of MRSA to this antibiotic. Checkerboard assays revealed a clear synergistic effect between the identified SANPs and PG, with all fractional inhibitory concentration index (FICI) values less than 1. Notably, the FICI value between H2 and PG reached 0.375. At sub-MIC concentrations of H2, the MIC of PG was reduced from 128 μg/mL to 4 μg/mL (Fig. 6a). At optimal ratios, the SANPs and PG formed stable spherical nanocomposites with diameters ranging from ~100 to 200 nm. TEM images obtained in the dry state clearly showed PG crystals encapsulated within the nanocomposites (Fig. 6b, Supplementary Fig. S21). Interestingly, in contrast to previous observations, the H2-PG nanocomposites exhibited the smallest size and a more uniform distribution among the three (Supplementary Fig. S22). Furthermore, H2 almost completely encapsulated PG, with a drug loading rate close to 50%, indicating a strong interaction between the two components (Supplementary Fig. S23). Molecular dynamics simulations further confirmed the presence of electrostatic interactions between H2 and PG, with a distinct assembly trend observed over 50 ns (Fig. 6c, Supplementary Fig. S24). Based on the above findings, we speculate that, due to the cationic and hydrophobic properties, the SANP nanocomposites can act as membrane-disruptive carriers that preferentially accumulate and integrate into bacterial membranes57. This process leads to passive enrichment of PG at the membrane interface, thereby increasing its local concentration. At the same time, the membrane-disruptive effect of SANPs may generate additional cell wall fragments in the surrounding environment, which could kinetically facilitate the acylation of the active-site serine of PBP2a by β-lactams58, ultimately overcoming the low-affinity resistance mechanism. For subsequent in vivo evaluations, we selected the H2-PG nanocomposite, which exhibited optimal synergy (FICI = 0.375), the highest encapsulation efficiency (EE = 97%), and a smaller particle size (106 nm).
a Checkerboard assays assessing the antibacterial synergy between identified SANPs and penicillin G (PG) against MRSA. FICI values are indicated in the upper right corner. b Characterization of SANPs-PG nanocomposites, including size distribution (by DLS), morphology (by TEM; scale bar: 1 μm), encapsulation efficiency (EE%), and loading capacity (LC%) determined by UV–Vis spectroscopy. c Molecular dynamics simulation of the assembly process of H2 and PG. d Schematic illustration of the lethal MRSA peritonitis model and corresponding treatment strategy. Cyclophosphamide was administered intraperitoneally (150 mg/kg on day 1 and 100 mg/kg on day 3) to induce immunosuppression. On day 4, mice were intraperitoneally infected with 1 × 107 CFU of MRSA. Post-infection, mice were treated intraperitoneally with H2 (10 mg/kg), PG (10 mg/kg), SM (10 mg/kg), H2-PG nanocomplex (5 mg/kg), or PBS (100 μL, control). After 24 h, the mice were euthanized, and samples of serum, spleen, lung, liver, and kidney were collected. Created in BioRender. Ji, J. (2025) https://BioRender.com/9obgfww. e Plate culture images of bacterial colonies obtained from liver tissue homogenates. f Quantification of bacterial loads in liver, kidney, spleen, and lung tissues. g Concentrations of inflammatory cytokines in mouse serum. h Histological analysis of liver, kidney, spleen, and lung sections stained with H&E. Scale bar: 300 nm. In panels (b, e, h), experiments were performed in triplicates with consistent results, and one representative image is shown. In panels (f, g), data are expressed as mean ± s.d. (n = 4 independent replicates). Statistical analysis was performed using one-way ANOVA. Source data are provided as a Source Data file.
To evaluate the in vivo therapeutic potential of H2 as a penicillin enhancer, we tested H2-PG nanocomposites in a lethal MRSA peritonitis model, comparing the results with treatments using H2 and PG alone. This model induces systemic diseases, such as sepsis, more readily than respiratory infection models, making it a stringent test for therapeutic strategies. We used a neutropenic mouse model, which is well-established for evaluating antibiotic efficacy59. Immunosuppression in mice was induced by cyclophosphamide injection prior to infection. On day 4, mice were infected via intraperitoneal injection with MRSA (1 × 107 CFU), using a lab-induced strain highly resistant to PG in vitro (MIC > 64 μg/mL), followed by the administration of various treatments. After 24 h, the mice were euthanized. Blood and tissues (liver, kidney, spleen, and lung) were harvested for bacterial quantification (Fig. 6d). Treatment with H2 (10 mg/kg), PG (10 mg/kg), SM (10 mg/kg), and H2-PG nanocomposites (5 mg/kg) resulted in a reduction of the average bacterial loads in these organs by 1.2, 0.9, 2.3, and 2.5 log units, respectively, compared to untreated controls (Fig. 6e, f). This result suggests that in the context of MRSA infection, PG exhibits minimal therapeutic efficacy. In contrast, the H2-PG nanocomposites exhibited anti-infective efficacy comparable to that of streptomycin. Serum inflammatory cytokine analysis revealed a similar trend, with significant reductions in pro-inflammatory markers observed in the H2-PG nanocomposite-treated group (Fig. 6g). Histological examination through H&E staining further confirmed that the H2, SM, and H2-PG treatment groups significantly ameliorated tissue damage compared to untreated controls (Fig. 6h). Acute toxicity evaluation in healthy mice showed no significant changes in body weight, organ indices, or serum biochemical markers after treatment with H2 or H2-PG, indicating the absence of detectable systemic toxicity in vivo (Supplementary Fig. S25). These results collectively demonstrate that H2 has substantial potential as an antibiotic adjuvant, capable of restoring bacterial susceptibility to antibiotics.
Discussion
In summary, we develop PolyCLOVER, a deep learning-guided framework for the design of polymeric antibiotics with integrated adjuvant effects to combat antibacterial drug resistance. By integrating multi-stage self-supervised learning, active learning, and high-throughput experimentation, PolyCLOVER enables efficient navigation of a vast polymeric chemical space without reliance on external datasets. Within 20 days, the framework successfully identified three lead SANPs from a library of ~100,000 candidates. These SANPs exhibited strong antibacterial activity against drug-resistant pathogens while maintaining low hemolytic toxicity. Remarkably, these three SANPs not only function as stand-alone antibacterials but also possess antibiotic delivery capacity that restores bacterial susceptibility to antibiotics, as confirmed by both in vitro and in vivo validation. These results demonstrate the therapeutic potential of the identified SANPs in combating drug-resistant bacterial infections. At present, the optimization objective is limited to planktonic bacteria; however, it can be naturally extended to more challenging targets, such as bacterial biofilms, in future work.
To alleviate data scarcity in polymer science, inspired by the development of domain-specific large language models, we propose a multi-stage learning pipeline involving sequential training on an unlabeled general polymer dataset, the unlabeled poly(β-amino ester) library, and the small, labeled dataset. Notably, the goal of PolyCLOVER is to search and optimize candidates with desired properties within a predefined combinatorial chemical space, rather than to directly apply the fine-tuned model to other polymer families. Therefore, although our approach exhibits a transductive nature, it allows for more effective utilization of unlabeled structural information in data-scarce settings, thereby improving model performance within the targeted candidate library. Collectively, this multi-stage training strategy provides a data-efficient paradigm for screening in large chemical libraries.
More broadly, we established PolyCLOVER as a generalizable and scalable platform not only applicable to poly(β-amino ester)s but also potentially extensible to a broad range of polymer backbones and complex topologies, including branched or star-shaped architectures. Through continuous interaction with internal experimental data, it enables autonomous evolution of functional polymers within a large chemical space, achieving complex design objectives without relying on external data. As such, PolyCLOVER offers a blueprint for next-generation polymeric biomaterial discovery frameworks that are adaptive, continuously learning, and extensible to the design of materials with precisely tailored functions.
Methods
Construction of SANPs library
Poly(β-amino ester)s can be synthesized through the Michael addition between diacrylate and amine monomers, enabling extensive customization of the polymer chemical properties60. Such polymers have been widely explored for their ability to deliver DNA and proteins due to their inherent positive charge61,62. It can also be expected to establish antibacterial libraries based on poly(β-amino ester)s, given their positive charge nature. In this study, 8 diacrylates with distinct structures were selected from commercially available sources based on solubility and cost considerations. 16 amines with tertiary amine side groups or Boc-protected amino groups were selected as the positively charged components. Additionally, 16 amines with hydrophobic side groups and 6 with hydrophilic side groups were prepared (Supplementary Table S1). Eight different feeding ratios were set to ensure its self-assembly capacity (Supplementary Table S2).
Chemical synthesis of poly(β-amino ester)s
The step-growth polymerization of Michael addition produces linear polymers with ester and tertiary amine structures in their skeleton. In this study, reaction occurred between diacrylates and three types of amines to produce random copolymers (Supplementary Fig. S26). The polymer is formed by alternating diacrylate and amine units, with three types of amines randomly distributed along the chains. The molar ratio of diacrylate to amine was set at 1.2:1. Initially, the amine monomer solution (3 M in DMF) was pipetted to a reaction vial containing 48 μL of diacrylate (3 M in DMF) according to the specified ratio. The reaction mixture was then heated to 90 °C and stirred magnetically for 24 h. For combinations with Boc-protected amine monomers, 80 μL of trifluoroacetic acid was added after polymerization. The solution was stirred overnight at room temperature to remove the protective group. Finally, the crude product was purified by precipitating in ether/dichloromethane (1:1 in vol%) solution three times. All samples involved in the screening process are summarized in Supplementary Data 1.
Antibacterial and hemolytic activity labeling
The synthesized and purified polymer was dissolved in DMSO at the concentration of 1.28 mg/mL. Fresh MRSA (ATCC 43300) was incubated in TSB overnight (120 rpm, 37 °C). The bacterial suspension was then diluted and calibrated to a concentration of 105 CFU/mL. 195 μL bacterial solution was added to each well in the 96-well plate, followed by addition of 5 μL polymer solution (final concentration at 32 μg/mL). The bacterial suspension was further incubated at 37 °C for 9 h. OD values at 595 nm of each well were by the microplate reader.
For the hemolysis assay, 4% red blood cell (RBC) suspension obtained from New Zealand White rabbits (purchased from Dr. Can Bioscience Inc., Hangzhou, Zhejiang, China) was used. First, 195 μL suspension was added to each well of the 96-well plate, and then 5 μL polymer solution was added to achieve a final polymer concentration of 128 μg/mL. The negative control group was added with 5 μL PBS, and the positive control group was added with 5 μL 0.4% Triton X-100. The plates were incubated at 37 °C for 1 h. After incubation, the plates were centrifuged at 4 °C (200 × g, 10 min). After centrifugation, the 100 μL supernatant was gently transferred into a new 96-well plate. OD values at 576 nm were measured, and the hemolysis ratio was calculated according to the following formula:
Dataset construction and preprocessing
In this study, a combinatorial polymer library of 98,304 poly(β-amino ester) SANPs is constructed via the Cartesian product of 8 diacrylates (A), 16 cationic amines (B), 16 hydrophobic amines (C), 6 hydrophilic amines (D), and 8 specific formulation ratios. Given the high-throughput nature of this study, direct characterization of actual monomer incorporation ratios for each candidate would be neither cost-effective nor necessary. Instead, feeding ratios—combined with chemical structures—are used as practical and consistent model inputs. This design enables efficient, data-driven prediction of biological activity, while maintaining alignment between training and inference conditions. The four-component Michael addition polycondensation product is treated as a random arrangement of three building blocks (A−B, A−C, A−D). The structure of each building block is generated by the reaction module of the RDKit63 package according to the general formulas:
‘[NH2:1][C;!$(C = O):6].[C:2] = [C:3][C:4] = [O:5]»[*]-[N:1]([C;!$(C = O):6])-[C:2][C:3][C:4] = [O:5]’ and ‘[C:1] = [C:2][C:3] = [O:4]»[*]-[C:1][C:2][C:3] = [O:4]’.
The copolymers are further converted into graph representations38. Specifically, repeating units of the copolymer is first transformed into three separate molecular line graphs \({\{{G}_{i}^{s}\}}_{i=1}^{3}\). The atom and bond features are summarized in Supplementary Tables S5, 6. The initialization of node features in the line graphs is defined as:
where \({W}_{v}\in {{\mathbb{R}}}^{0.5d\times {d}_{{{\rm{atom}}}}}\) and \({W}_{e}\in {{\mathbb{R}}}^{0.5d\times {d}_{{{\rm{bond}}}}}\) are trainable weights, \(d\) denotes the dimensionality of the embedding space, \({{{\bf{x}}}}_{i,j}\) is the feature of a bond represented as a node in the line graph, and \({{{\bf{x}}}}_{i}\), \({{{\bf{x}}}}_{j}\) are the features of the atoms at its two ends. Then, a global virtual node \({v}^{{{\rm{global}}}}\) is introduced and connected to all nodes across subgraphs to facilitate inter-subgraph message passing. Three local virtual nodes \({\{{v}_{i}^{{{\rm{local}}}}\}}_{i=1}^{3}\) are connected to their corresponding components to specify the ratios of individual building blocks \({\{{r}_{i}\}}_{i=1}^{3}\). The features of these virtual nodes are initialized according to the following equation:
where \({{\rm{embedding}}}:\{{r}_{i}\}\to {{\mathbb{R}}}^{d}\) is a trainable embedding function that maps discrete indices to continuous vector, \({{\boldsymbol{{h}}}}^{{{\rm{global}}}}\in {{\mathbb{R}}}^{d}\) is a trainable vector. Furthermore, to enrich the graph representation, we incorporate RDKit and Mordred64 descriptors of each repeating unit into each subgraph via additional virtual nodes \({\{{v}_{i}^{{{\rm{add}}}}\}}_{i=1}^{3}\). As a result, we use \(G=(V,E)\) to denote the ensembled graph of poly(β-amino ester)s in the following.
Model construction
Model architecture
We selected graph transformer (GT) architecture as the backbone of neural network, specifically adopting LiGhT45, due to its superior performance among ten commonly used GNN variants. To encode graph structural information into Transformer, Path Encoding (PE) and Distance Encoding (DE) are employed. Formally, the path feature \({a}_{i,j}^{p}\) between \({v}_{i}\) and \({v}_{j}\) in the path attention matrix \({A}^{p}\in {{\mathbb{R}}}^{{N}_{v}\times {N}_{v}}\) is defined as follow:
where \({{\boldsymbol{{h}}}}_{{v}_{n}}\) represents the feature of the n-th node in the shortest path between \({v}_{i}\) and \({v}_{j}\), \({N}_{p}\) represents the number of nodes in the path, \({W}_{n}^{p}\in {{\mathbb{R}}}^{{d}_{p}\times d}\) and \({W}^{p}\in {{\mathbb{R}}}^{1\times {d}_{p}}\) are trainable weights. Similarly, the distance feature \({a}_{i,j}^{d}\) between \({v}_{i}\) and \({v}_{j}\) in the distance attention matrix \({A}^{d}\in {{\mathbb{R}}}^{{N}_{v}\times {N}_{v}}\) is calculated as follow:
where \({d}_{i,j}\) represents the distance between \({v}_{i}\) and \({v}_{j}\), \({W}_{1}^{d}\in {{\mathbb{R}}}^{{d}_{d}\times 1}\) and \({W}_{2}^{d}\in {{\mathbb{R}}}^{1\times {d}_{d}}\) are trainable weights. Based on the structural encodings, GT processes the graphs through stacked attention layers. For a given feature matrix \({H}^{(l)}\in {{\mathbb{R}}}^{{N}_{v}\times d}\), the backbone of GT updates node representations using the following formulation:
where \({W}_{k}^{Q},{W}_{k}^{K},{W}_{k}^{V}\in {{\mathbb{R}}}^{d\times {d}_{k}}\) are trainable weights of k-th head, \({W}_{1}\in {{\mathbb{R}}}^{4d\times d}\), \({W}_{2}\in {{\mathbb{R}}}^{d\times 4d}\) are trainable weights of the feed-forward network, \(\sigma (\cdot )\) is the GELU activation function, \({d}_{k}=d/h\) is the dimension of each attention head, \(h\) stands for the number of heads. After the final GT layer, the learned node representations are aggregated into a graph-level embedding using a readout operator:
Finally, the graph-level embedding \({{\boldsymbol{{h}}}}_{G}\) is then passed through a projection head to produce the final predictions:
where \(\hat{{{\bf{y}}}}\in {{\mathbb{R}}}^{2}\) denotes the predicted antibacterial and hemolytic properties, \({W}_{1}^{{{\rm{proj}}}}\in {{\mathbb{R}}}^{d\times 4d}\) and \({W}_{2}^{{{\rm{proj}}}}\in {{\mathbb{R}}}^{2\times d}\) are trainable weights.
Training objectives
In this work, we propose a multi-stage learning strategy designed to fully leverage the structural priors of polymers. This strategy consists of three sequential stages: (1) general polymer knowledge learning, (2) continual pretraining to capture structural features specific to poly(β-amino ester)s, and (3) fine-tuning for activity prediction.
In the first stage, we adopt the pretraining approach proposed in our previous work38, performing SSL on about one million unlabeled structures to capture general knowledge of polymers. Specifically, given a batch of N polymers, Periodicity Augmentation is employed to generate different views as positive pairs (different head-to-tail orders and numbers of repeating units), while different polymers are treated as negative pairs. Next, random masking is applied to 50% of the nodes in the constructed graphs. 80% of the selected nodes are replaced with a special [MASK] token, 10% are replaced with random nodes, and the remaining 10% are left unchanged. We adopt a multi-task SSL strategy to capture the hierarchical structural features of polymers. A contrastive loss \({ {\mathcal L} }_{{{\rm{CL}}}}\) (InfoNCE Loss) is applied to pull positive pairs closer and push negative pairs apart, while a reconstruction loss \({ {\mathcal L} }_{{{\rm{recon}}}}\) (Cross Entropy Loss) is applied to recover the masked nodes:
where \({{{\bf{z}}}}_{i}\) and \({{{\bf{z}}}}_{i}^{{\prime} }\) are the latent embeddings of two augmented views of the same polymer, \(\gamma\) is a temperature parameter, \({ {\mathcal L} }_{{{\rm{InfoNCE}}}}^{{\prime} }(i)\) denotes the symmetric loss \({ {\mathcal L} }_{{{\rm{InfoNCE}}}}(i)\) for the paired positive sample.
However, there exists a substantial gap between the pretraining dataset and the downstream finetuning domain. In this work, we propose to adopt a continual pretraining strategy, in which the graph encoder is further pretrained on the constructed unlabeled library of poly(β-amino ester)s to fully exploit the structural priors inherent in the screening space. In this stage, two independent random masks are applied to the same poly(β-amino ester) to form a positive pair, while different poly(β-amino esters) serve as negative pairs. The pretraining objective remains consistent with Eq. (9).
After the two-stage pretraining, the model is finetuned to predict antibacterial and hemolytic activities. For this purpose, a randomly initialized projection head is attached to the pretrained graph encoder to output the predicted activity values. We optimize the model using the MSE Loss, where the objective is to minimize the difference between the predicted activities \(\hat{{{\bf{y}}}}\) and the experimentally measured activities \({{\bf{y}}}\) for each sample. The loss is defined as:
Training details
In pretraining stage, the model is trained using the Adam optimizer with a learning rate of 2 × 10−4, a weight decay of 1 × 10−6, and a batch size of 96 for 10,000 steps. In finetuning stage, the model is trained with a batch size of 32. Hyperparameters, including learning rate, dropout, and weight decay, are selected via grid search based on validation performance (Supplementary Table S7). The learning rate is dynamically adjusted using the PolynomialDecayLR scheduler. Early stopping with a patience of 20 epochs is employed based on validation performance. All experiments are implemented in PyTorch with DGL and trained on three NVIDIA RTX 4090 GPUs.
Evaluation metrics
To quantitatively assess model performance on each tasks, we employ the following metrics: root mean squared error (r.m.s.e.), mean absolute error (m.a.e.), Pearson coefficient (R), and top-n r.m.s.e. for test samples with the highest activity. These metrics are defined as follows:
where \(y\) is the measured value, \(\hat{y}\) is the predicted value, \(\bar{y}\) is the mean of measured values, \(\overline{\hat{y}}\) is the mean of predicted values, \({{\rm{T}}}_{n}\) denotes the indices of the top-n test samples with the highest activity.
Active learning strategy
An active learning framework is applied over multiple rounds to optimize antibacterial and hemolytic properties of candidates, formulated as a dual-objective regression task. The optimization target for each sample is defined as a scalar score, computed as a weighted sum of the predicted antibacterial activity and hemolysis ratio:
where λi is the weighting coefficient that balances the relative importance of the two objectives.\({\hat{y}}_{i}\) is the prediction for the i-th task. In this work, λ for antibacterial activity was empirically set to 0.7, while λ for hemolytic activity was set to 0.3. To estimate the model’s prediction μ(x) and uncertainty σ(x), an ensemble learning approach is employed. Specifically, 20 independent models are trained, and for each sample, the predicted mean and standard deviation are calculated from the outputs of the ensemble members. To select candidate SANPs in each round, the UCB acquisition function is employed, balancing exploitation of high-performing samples and exploration of uncertain regions. The acquisition function is defined as:
where β is a hyper-parameter controlling the trade-off between exploration and exploitation (β = 2).
In each round, K-Means algorithm is applied to group the top-ranked samples (γ = 0.1) into 20 clusters. From each cluster, the top 3 samples based on are selected, yielding a total of 60 candidates (Supplementary Table S8). These samples are then experimentally tested for antibacterial activity and hemolysis ratio. After several iterations of this process, the entire library is predicted and ranked using the final model. The top 60 candidates, based on their predicted scores s(x), are synthesized and characterized again to validate the model’s predictive performance and assess the effectiveness of the active learning approach.
MIC assay
MRSA was incubated in TSB overnight (120 rpm, 37 °C). The bacterial suspension was then diluted and calibrated to a concentration of 105 CFU/mL. Purified SANPs were dissolved in DMSO to a concentration of 2.56 mg/mL. 195 μL MRSA suspension was added to the 96-well plate, and 5 μL tested SANPs solution was added to each well. The SANPs concentration was serially diluted two-fold, ranging from 32 μg/mL to 4 μg/mL. The plate was incubated for 9 h. TSB solution served as the control group. The MIC was determined as the lowest concentration that resulted in a completely clear solution.
HC50 assay
RBC suspension (195 μL) was added to each well of a 96-well plate, followed by the addition of 5 μL of purified SANPs solution. The final SANPs concentrations tested were 8, 16, 32, 64, 128, 200, 256, 400, and 512 μg/mL, respectively. The negative control group was added with 5 μL PBS, and the positive control group was added with 5 μL 0.4% Triton X-100. The plates were incubated at 37 °C for 1 h, then centrifuged at 4 °C (200 × g, 10 min). After centrifugation, 100 μL supernatant was transferred into a new 96-well plate. OD values at 576 nm were measured using a microplate reader, and the hemolysis ratio for each SANPs concentration was calculated. HC50 was determined by fitting the hemolysis ratio-concentration curve to identify the SANPs concentration at which 50% hemolysis occurred.
Bacterial strains
The model strain used for high-throughput screening, S. aureus ATCC 43300 (MRSA), was purchased from the Shanghai Bioresource Collection Center (Shanghai, China). For subsequent characterization assays, three reference strains (S. aureus ATCC 29213, S. epidermidis ATCC 35984, and A. baumannii ATCC 19606) and eleven clinically isolated drug-resistant pathogens, including S. aureus-MRSA, VISA, S. capitis-MRSC, E. faecalis-VRE, A. baumannii-MDRAB, P. aeruginosa-CRPA, K. pneumoniae-CRKP, were obtained from the Department of Infectious Diseases, Sir Run Run Shaw Hospital, School of Medicine, Zhejiang University.
Checkerboard assay
A checkerboard assay was performed to evaluate the synergistic effects of penicillin G sodium (PG) and the identified polymers (H1–3) against MRSA. The MICs of H1–3 were first determined, followed by the checkerboard assay conducted in 96-well microtiter plates. PG was serially diluted along the vertical axis from 128 µg/mL to 8 µg/mL, while the polymers were diluted along the horizontal axis: H1 and H3 ranged from 16 µg/mL to 1 µg/mL, and H2 ranged from 32 µg/mL to 2 µg/mL. The final row and column contained wells where both the polymer and PG concentrations were 0 µg/mL as controls. The plates were incubated at 37 °C for 9 h, and all experiments were performed in triplicate to ensure reproducibility and reliability.
Preparation and characterization of H2-PG nanocomposites
The optimal formulation ratio for H2-PG nanocomposites was determined based on checkerboard assays. A stock solution of PG was prepared at 10 mg/mL in deionized water, while the polymer stock solution was 10.24 mg/mL in DMSO. Prior to use, the polymer solution was diluted with deionized water. For NC preparation, PG was diluted to 1 mg/mL in deionized water, and the polymer solution was diluted to the optimal concentration. Equal volumes of the PG and polymer solutions were mixed, vortexed for 30 s, and incubated at room temperature for 5 min to allow nanocomplex formation.
The hydrodynamic diameter of the NCs was measured using DLS on a Malvern Zetasizer Advance Pro (Blue Label). The zeta potential was also determined using the same instrument. The morphology of the NCs was examined by TEM.
The encapsulation efficiency (EE) and drug loading capacity (LC) of the antibiotic-loaded NCs were quantified using UV–Vis spectroscopy. NCs were prepared as described above and subsequently subjected to ultrafiltration using a 3 kDa molecular weight cutoff (MWCO) membrane to separate free PG from the nanocomplexes. The filtrate was collected and diluted appropriately, and its absorbance spectrum was recorded in the wavelength range of 185–220 nm using a UV–Vis spectrophotometer. Due to interference from the mixture, a slight shift in the position of the maximum absorption peak was observed; thus, the peak exhibiting the highest absorbance within this range was used to determine the concentration of unencapsulated PG. The EE and LC were calculated according to the following equations:
where\({W}_{{{\rm{total}}}}\) represents the initial total amount of PG, \({W}_{{{\rm{free}}}}\) is the amount of unencapsulated PG quantified from the filtrate, and \({W}_{{{\rm{polymer}}}}+{W}_{{{\rm{encapsulated}}}}\) corresponds to the total mass of the nanocomplexes.
Ethical statement
Mice were obtained and maintained in pathogen-free conditions at the Laboratory Animal Centre of Zhejiang University and raised at 20–22 °C and 40–70% humidity, with a dark-light cycle of 12 h. All animal studies were performed in compliance with the guide for care and use of laboratory animals and adopted the protocol that has been approved by Institutional Animal Care and Use Committee of CDSER of Zhejiang University (IACUC-25-S257).
Mouse pneumonia model
On day 0, healthy female ICR mice (18–20 g) were anesthetized with isoflurane. MRSA suspension (108 CFU/mL, 60 μL) was administered intratracheally. Successfully infected mice were randomly divided into five groups (n = 3). On day 1, polymers H1, H2, H3, and SM were administered via tail vein injection (5 mg/kg). PBS served as the control. On day 2, the mice were euthanized, their lungs were excised for morphological observation. The lung tissue was homogenized for further bacterial colony counts.
Lethal MRSA peritonitis model
A lethal MRSA peritonitis model was established in 20 male C57BL/6 mice (18–20 g), randomly assigned to five groups (n = 4). Cyclophosphamide was administered intraperitoneally (150 mg/kg on day 1 and 100 mg/kg on day 3) to induce immunosuppression. On day 4, mice were intraperitoneally infected with 1 × 107 CFU of MRSA. Post-infection, mice received intraperitoneal treatments: H2 (10 mg/kg), PG (10 mg/kg), SM (10 mg/kg), H2-PG nanocomplex (5 mg/kg), or PBS (100 μL, control). After 24 h, mice were euthanized, and serum, spleen, lung, liver, and kidney samples were collected. Serum samples were used for the evaluation of inflammatory cytokine levels, while the harvested organs were analyzed for bacterial load and histopathological examination.
Statistics and reproducibility
Statistical analyses were performed using GraphPad Prism (version 10.1.2). One-way ANOVA followed by post hoc multiple-comparison tests was used. Data are presented as mean ± standard deviation (SD). Exact p values are shown on the figures. No statistical method was used to predetermine sample size, no data were excluded from the analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available within the main text and the Supplementary Information. The pretrained checkpoints and processed datasets are available at https://doi.org/10.6084/m9.figshare.2887696165. Source data are provided with this paper.
Code availability
The code of the work is available via GitHub at https://github.com/wuyuhui-zju/PolyCLOVER and via figshare at https://doi.org/10.6084/m9.figshare.2887696165.
References
Naghavi, M. et al. Global burden of bacterial antimicrobial resistance 1990-2021: a systematic analysis with forecasts to 2050. Lancet 404, 1199–1226 (2024).
Ho, C. S. et al. Antimicrobial resistance: a concise update. Lancet Microbe 6, 100947 (2024).
Martins, A. et al. Antibiotic candidates for Gram-positive bacterial infections induce multidrug resistance. Sci. Transl. Med. 17, eadl2103 (2025).
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Hutchings, M. I., Truman, A. W. & Wilkinson, B. Antibiotics: past, present and future. Curr. Opin. Microbiol. 51, 72–80 (2019).
Shi, Z. et al. A comprehensive overview of the antibiotics approved in the last two decades: retrospects and prospects. Molecules 28, 1762 (2023).
Bognár, B., Spohn, R. & Lázár, V. Drug combinations targeting antibiotic resistance. npj Antimicrob. Resist. 2, 29 (2024).
Tyers, M. & Wright, G. D. Drug combinations: a strategy to extend the life of antibiotics in the 21st century. Nat. Rev. Microbiol. 17, 141–155 (2019).
Joseph, J., Boby, S., Mooyottu, S. & Muyyarikkandy, M. S. Antibiotic potentiators as a promising strategy for combating antibiotic resistance. npj Antimicrob. Resist. 3, 53 (2025).
Song, M. et al. A broad-spectrum antibiotic adjuvant reverses multidrug-resistant Gram-negative pathogens. Nat. Microbiol. 5, 1040–1050 (2020).
Lewis, K. The Science of Antibiotic Discovery. Cell 181, 29–45 (2020).
MacNair, C. R., Rutherford, S. T. & Tan, M. W. Alternative therapeutic strategies to treat antibiotic-resistant pathogens. Nat. Rev. Microbiol. 22, 262–275 (2024).
Magana, M. et al. The value of antimicrobial peptides in the age of resistance. Lancet Infect. Dis. 20, E216–E230 (2020).
Mookherjee, N., Anderson, M. A., Haagsman, H. P. & Davidson, D. J. Antimicrobial host defence peptides: functions and clinical potential. Nat. Rev. Drug Discov. 19, 311–332 (2020).
Fernandes, S. & Cavalieri, F. Antimicrobial peptides boosted by ultrasound. Nat. Biomed. Eng. 9, 1177–1179 (2025).
Hancock, R. E. & Sahl, H. G. Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat. Biotechnol. 24, 1551–1557 (2006).
Yuan, Y. et al. Stable peptide-assembled nanozyme mimicking dual antifungal actions. Nat. Commun. 15, 5636 (2024).
Gelman, M. A., Weisblum, B., Lynn, D. M. & Gellman, S. H. Biocidal activity of polystyrenes that are cationic by virtue of protonation. Org. Lett. 6, 557–560 (2004).
Lam, S. J. et al. Combating multidrug-resistant Gram-negative bacteria with structurally nanoengineered antimicrobial peptide polymers. Nat. Microbiol. 1, 16162 (2016).
Nederberg, F. et al. Biodegradable nanostructures with selective lysis of microbial membranes. Nat. Chem. 3, 409–414 (2011).
Wu, Y. et al. Host defense peptide mimicking antimicrobial amino acid polymers and beyond: Design, synthesis and biomedical applications. Prog. Polym. Sci. 141, 101679 (2023).
Xie, J. et al. Addressing MRSA infection and antibacterial resistance with peptoid polymers. Nat. Commun. 12, 5898 (2021).
Qian, Y. et al. Secondary amine pendant β-peptide polymers displaying potent antibacterial activity and promising therapeutic potential in treating MRSA-induced wound infections and keratitis. J. Am. Chem. Soc. 144, 1690–1699 (2022).
Maset, R. G. et al. Combining SNAPs with antibiotics shows enhanced synergistic efficacy against S. aureus and P. aeruginosa biofilms. npj Biofilms Microbomes 9, 36 (2023).
Schaefer, S. et al. A synthetic peptide mimic kills Candida albicans and synergistically prevents infection. Nat. Commun. 15, 6818 (2024).
Wu, Y. M. et al. Host defense peptide mimicking antimicrobial amino acid polymers and beyond: design, synthesis and biomedical applications. Prog. Polym. Sci. 141, 47 (2023).
Shabani, S. et al. Synthetic peptide branched polymers for antibacterial and biomedical applications. Nat. Rev. Bioeng. 2, 343–361 (2024).
Shen, X. et al. AI-driven discovery and design of antimicrobial peptides. Sci. China Technol. Sci. 68, 2200205 (2025).
Wan, F., Torres, M. D. T., Guan, C. & de la Fuente-Nunez, C. Tutorial: guidelines for the use of machine learning methods to mine genomes and proteomes for antibiotic discovery. Nat. Protoc. 20, 2685–2697 (2025).
Wan, F., Wong, F., Collins, J. J. & de la Fuente-Nunez, C. Machine learning for antimicrobial peptide identification and design. Nat. Rev. Bioeng. 2, 392–407 (2024).
Huang, J. et al. Identification of potent antimicrobial peptides via a machine-learning pipeline that mines the entire space of peptide sequences. Nat. Biomed. Eng. 7, 797–810 (2023).
Chen, J. W. et al. Global marine microbial diversity and its potential in bioprospecting. Nature 633, 371–379 (2024).
Santos, C. D. Jr. et al. Discovery of antimicrobial peptides in the global microbiome with machine learning. Cell 187, 35 (2024).
Torres, M. D. T. et al. Mining human microbiomes reveals an untapped source of peptide antibiotics. Cell 187, 31 (2024).
Maasch, J., Torres, M. D. T., Melo, M. C. R. & de la Fuente-Nunez, C. Molecular de-extinction of ancient antimicrobial peptides enabled by machine learning. Cell Host Microbe 31, 22 (2023).
Torres, M. D. T. et al. Mining for encrypted peptide antibiotics in the human proteome. Nat. Biomed. Eng. 6, 67–75 (2022).
Wu, T. et al. AI-guided few-shot inverse design of HDP-mimicking polymers against drug-resistant bacteria. Nat. Commun. 15, 6288 (2024).
Wu, Y. et al. Periodicity-aware deep learning for polymers. Nat. Comput. Sci. 5, 1214–1226 (2025).
Makabenta, J. M. V. et al. Nanomaterial-based therapeutics for antibiotic-resistant bacterial infections. Nat. Rev. Microbiol. 19, 23–36 (2021).
Pham, P., Oliver, S., Nguyen, D. T. & Boyer, C. Effect of cationic groups on the selectivity of ternary antimicrobial polymers. Macromol. Rapid Commun. 43, 2200377 (2022).
Phuong, P. T. et al. Effect of hydrophobic groups on antimicrobial and hemolytic activity: Developing a predictive tool for ternary antimicrobial polymers. Biomacromolecules 21, 5241–5255 (2020).
Turner, N. A. et al. Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nat. Rev. Microbiol. 17, 203–218 (2019).
Wieder, O. et al. A compact review of molecular property prediction with graph neural networks. Drug Discov. Today.: Technol. 37, 1–12 (2020).
Reiser, P. et al. Graph neural networks for materials science and chemistry. Commun. Mater. 3, 93 (2022).
Li, H., Zhao, D. & Zeng, J. KPGT: knowledge-guided pre-training of graph transformer for molecular property prediction. In Proceedings of the 28th ACM SIGKDD Conference on Knowledge Discovery and Data Mining (ACM, 2022).
Wang, Y., Wang, J., Cao, Z. & Barati Farimani, A. Molecular contrastive learning of representations via graph neural networks. Nat. Mach. Intell. 4, 279–287 (2022).
Hu, W. et al. Strategies for pre-training graph neural networks. In Proceedings of the 8th International Conference on Learning Representations (ICLR, 2020).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Qiang, B. et al. Bridging the gap between chemical reaction pretraining and conditional molecule generation with a unified model. Nat. Mach. Intell. 5, 1476–1485 (2023).
Wang, T. & Isola, P. Understanding contrastive representation learning through alignment and uniformity on the hypersphere. In Proceedings of the 37th International Conference on Machine Learning (ACM, 2020).
Frederiksen, N., Hansen, P. R., Björkling, F. & Franzyk, H. Peptide/peptoid hybrid oligomers: The Influence of hydrophobicity and relative side-chain length on antibacterial activity and cell Selectivity. Molecules 24, 4429 (2019).
Zhang, W. et al. Host defense peptide mimicking cyclic peptoid polymers exerting strong activity against drug-resistant bacteria. Biomater. Sci. 10, 4515–4524 (2022).
O’Sullivan, M. E. et al. Aminoglycoside ribosome interactions reveal novel conformational states at ambient temperature. Nucleic Acids Res. 46, 9793–9804 (2018).
Zhou, M. et al. Poly(2-oxazoline)-based functional peptide mimics: eradicating MRSA infections and persisters while alleviating antimicrobial resistance. Angew. Chem. Int. Ed. 59, 6412–6419 (2020).
Rubinstein, E., Kollef, M. H. & Nathwani, D. Pneumonia caused by methicillin-resistant Staphylococcus aureus. Clin. Infect. Dis. 46, S378–S385 (2008).
Adedeji-Olulana, A. F. et al. Two codependent routes lead to high-level MRSA. Science 386, 573–580 (2024).
Williams, S. C. et al. Polyacrylamide-based antimicrobial copolymers to replace or rescue antibiotics. ACS Cent. Sci. 11, 486–496 (2025).
Fuda, C. et al. Activation for catalysis of penicillin-binding protein 2a from methicillin-resistant Staphylococcus a ureus by bacterial cell wall. J. Am. Chem. Soc. 127, 2056–2057 (2005).
Mishra, B., Lakshmaiah Narayana, J., Lushnikova, T., Wang, X. & Wang, G. Low cationicity is important for systemic in vivo efficacy of database-derived peptides against drug-resistant Gram-positive pathogens. Proc. Natl. Acad. Sci. 116, 13517–13522 (2019).
Dosta, P. et al. Investigation of the enhanced antitumour potency of STING agonist after conjugation to polymer nanoparticles. Nat. Nanotechnol. 18, 1351–1363 (2023).
Lynn, D. M. & Langer, R. Degradable poly (β-amino esters): synthesis, characterization, and self-assembly with plasmid DNA. J. Am. Chem. Soc. 122, 10761–10768 (2000).
Ben-Akiva, E. et al. Biodegradable lipophilic polymeric mRNA nanoparticles for ligand-free targeting of splenic dendritic cells for cancer vaccination. Proc. Natl. Acad. Sci. 120, e2301606120 (2023).
Landrum, G. RDKit: Open-source cheminformatics. Retrieved from https://www.rdkit.org (2013).
Moriwaki, H., Tian, Y.-S., Kawashita, N. & Takagi, T. Mordred: a molecular descriptor calculator. J. Cheminform. 10, 4 (2018).
Wu, Y. Pretrained checkpoints and source code. figshare https://doi.org/10.6084/m9.figshare.28876961 (2025).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (52293381), Zhejiang Provincial Natural Science Foundation of China (LR25E030001), the “Pioneer” and “Leading Goose” R&D Program of Zhejiang (2025C04012), and the National Key Research and Development Program of China (2022YFB3807300). This work was also supported by Transvascular Implantation Devices Research Institute China (TIDRIC) under Grant No. KY012024007 and KY012024009.
Author information
Authors and Affiliations
Contributions
P.Z. and J.J. conceptualized and supervised the project. P.Z., Y.W., C.W., and X.S. designed the experiments, analyzed the data, and wrote the paper. Y.W. and P.Z. designed the overall framework. Y.W. was responsible for the design, training, and analysis of the deep learning model. C. W. carried out the chemical synthesis and in vitro assays. X.S. investigated the self-assembly behavior of SANPs and their synergistic therapeutic effects with antibiotics. Y.C. and H.W. evaluated the performance of SANPs against clinically isolated multidrug-resistant bacteria. B.X. participated in the mouse peritonitis model experiments. Z.Z. participated in the toxicity evaluation. Y.F.C. assisted with the experiments. W.D. and Y.H. contributed to the mouse pneumonia model experiments. L.Z. participated in experimental discussions. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Eleftherios Mylonakis, Fangping Wan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Y., Wang, C., Shen, X. et al. Iterative discovery of potent polymeric antibiotics via multi-stage and multi-task learning against antimicrobial resistance. Nat Commun 17, 1878 (2026). https://doi.org/10.1038/s41467-026-68682-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68682-z