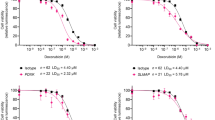
Abstract
Childhood cancer survivors face increased cardiometabolic risks from cancer treatment exposures, yet mechanisms remain unclear. Here, epigenome-wide analysis identifies 1893 DNA methylation (DNAm) sites in peripheral-blood-mononuclear-cells (PBMCs) associated with at least one cardiometabolic risk factor (CMRF), including obesity (n = 1720), abnormal glucose (n = 201), hypertriglyceridemia (n = 145), hypercholesterolemia (n = 38) and hypertension (n = 34) in 2938 survivors from the St. Jude Lifetime Cohort. A core set of five DNAm sites near CPT1A and LMNA is associated with all CMRFs. Mediation analyses identify 24 sites mediating associations between treatments and CMRFs, implicating inflammatory and metabolic pathways. Notably, cg20370568, a cis-expression quantitative trait methylation site for ANTXR2, mediates 20% of the effect of body-trunk-radiotherapy on abnormal glucose. These findings suggest that prior genotoxic cancer treatments may become biologically embedded through DNAm variations that could contribute to cardiometabolic dysfunction and highlight candidate biomarkers for refining risk stratification and guiding intervention strategies in survivorship care.
Similar content being viewed by others
Introduction
Cardiovascular disease (CVD)-related death is the leading cause of non-cancer late mortality (i.e., 5 or more years after diagnosis of primary cancer) in childhood cancer survivors, accounting for ~26% of deaths within 45 years post diagnosis1. Survivors face a substantially higher burden of CVD than individuals without a history of childhood cancer, with an earlier age of onset and a well-established association with prior exposures to genotoxic cancer treatments, including anthracycline chemotherapy, and chest-directed radiotherapy2. Importantly, cardiometabolic risk factors (CMRFs)—including abnormal glucose, obesity, hypertension, hypertriglyceridemia, and hypercholesterolemia—potentiate risks of future CVD and late mortality3,4. While treatment-related CMRFs are highly prevalent, the underlying biological mechanisms linking cancer treatments to cardiometabolic dysfunctions remain poorly understood. Elucidation of these molecular mechanisms is critical to advance the precision medicine paradigm of personalized risk stratifications and targeted interventions, ultimately improving long-term survivorship outcomes.
DNA methylation (DNAm) is a dynamic and environmentally responsive epigenetic mechanism that integrates signals from internal and external stressors, including genotoxic cancer therapies such as chemotherapy and radiotherapy (RT). Although inter-individual variability in DNAm is common, individuals who received specific cancer therapies can exhibit persistent shifts relative to those without exposures5. Because these DNAm changes may endure for years, they represent a plausible link in the causal pathway between prior cancer treatment and subsequent long-term adverse health outcomes. Our previous study in the well-characterized St. Jude Lifetime Cohort (SJLIFE) identified 935 5′-cytosine-phosphate-guanine-3′ (CpG) sites exhibiting persistent DNAm variations associated with various cancer treatments5. Moreover, a subset of these CpGs mediated the association between specific cancer treatments and CMRFs, suggesting that DNAm may serve as a molecular intermediary linking cancer therapy exposures to long-term cardiometabolic dysfunctions. Notably, DNAm at four independent CpGs (cg06963130, cg21922478, cg22976567, cg07403981) mediated 70.3% of the association between abdominal-RT and hypercholesterolemia. Two of these CpGs reside near genes with well-established roles in lipid metabolism: cg21922478 (ITGA1) and cg22976567 (LMNA), suggesting the biological plausibility of treatment-induced DNAm variations contributing to CMRFs.
Beyond its role in the biological embedding of cancer treatment effects, DNAm is a putative regulator of gene expression, influencing metabolic pathways implicated in cardiometabolic dysfunction6. DNAm-regulated genes are involved in fatty acid synthesis, lipid transport, and storage7,8, as well as inflammatory pathways9,10 that contribute to insulin resistance, abnormal glucose, and hypertension in the general population. Among long-term survivors of childhood cancer, we previously conducted another outcome-centric epigenome-wide association study (EWAS) that complemented our exposure-centric5 EWAS approach, identifying CpGs associated with circulating lipid levels11. This analysis identified 106 lipid-associated CpGs, reinforcing the functional relevance of DNAm in metabolic dysfunction and highlighting its potential as a biomarker for identifying survivors with existing lipid abnormalities or those at higher risk for developing metabolic disorders.
Despite these insights, the biological mechanisms linking cancer treatment exposures to cardiometabolic toxicities remain incompletely understood. Survivors exhibit substantial variability in the timing of onset, clinical presentation, and long-term outcomes of CMRFs, highlighting the need for mechanistic studies that move beyond the statistical association. While our prior work demonstrated that DNAm may mediate the effects of specific treatments on select CMRFs, those analyses were designed around treatment-associated CpGs, assessed a limited subset of treatment-CMRF pairs, and did not examine transcriptional consequences.
Here, we adopt an outcome-first design to assess whether peripheral-blood DNAm mediates associations between prior treatments and CMRFs (Fig. 1). We first map CpGs associated with each CMRF via an epigenome-wide analysis, then evaluate their links to treatment exposures, and estimate mediation for treatment-DNAm-CMRF trios. Additionally, we consolidate uncorrelated CpGs into composite DNAm scores for composite mediation analysis, and we integrate cis-eQTM analyses to connect mediator CpGs to nearby gene expression. By integrating EWAS, causal mediation analysis, and cis-expression quantitative methylation (cis-eQTM) mapping, we identify CpGs that are not only statistically associated with CMRFs but also biologically informative. This comprehensive framework advances our understanding of DNAm as one type of molecular intermediary between cancer treatments and late effects and highlights candidate biomarkers with potential utility for refining risk stratification and guiding future strategies to mitigate CMRFs and further CVD risk as well as premature mortality in survivors of childhood cancer.
a DNAm was measured in peripheral blood mononuclear cells (PBMCs) from 2938 survivors of childhood cancer from the SJLIFE cohort using methylation arrays. b EWAS analyses identified CpGs associated with five CMRFs. Toy example boxplot of CMRF-methylation association. c CMRF-associated CpGs were tested for their associations with nine cancer treatment exposures (toy example boxplot of a treatment-methylation association). d CpGs associated with both a CMRF and cancer treatment underwent causal mediation analysis to infer whether DNAm explained part of the treatment effect on CMRFs. e Multiple mediators of the same treatment-CMRF pair were aggregated into a composite methylation score, enabling a combined mediation analysis. f An eQTM analysis identified CpGs whose methylation levels correlate with nearby gene expression (toy example scatter plot). CMRF cardiometabolic risk factor, DNAm DNA methylation, EWAS epigenome-wide association study, eQTM expression quantitative trait methylation, PBMC peripheral blood mononuclear cells, RT radiotherapy, SJLIFE St Jude Lifetime Cohort Study.
Results
Characteristics of the study population
The study included 2938 childhood cancer survivors (53% male) from the SJLIFE cohort (Fig. 1a). The median age at primary cancer diagnosis was 6.1 (interquartile range [IQR], 2.8–12.6) years, with blood sampling for DNAm conducted at a median age of 30.1 (IQR, 21.6–38) years, and the most recent clinical follow-up occurring at a median age of 33.4 (IQR, 25.4–41.8) years (Table 1). Survivors represented a diverse range of primary cancer diagnoses, including leukemia (34.0%), lymphoma (18.6%), embryonal tumors (13.4%), central nervous system (CNS) tumors (12.7%), sarcoma (12.3%), and other malignancies (9.0%).
Survivors underwent multimodality therapy. Most received vinca alkaloids (71.4%), anthracyclines (57.0%), and alkylating agents (56.7%), while antimetabolites (48.3%), corticosteroids (45.5%), and epipodophyllotoxins (35.2%) were also common. Given the strong correlations among chest, abdominal, and pelvic RT exposures (Phi coefficient > 0.65) and their likely shared systemic effects on peripheral-blood-mononuclear-cells (PBMCs) derived DNAm, we defined a composite body-trunk-RT variable as irradiation to any of these body regions. Overall, 26.7% of survivors received body-trunk-RT and 26.5% received brain-RT. Treatment exposures were frequently concurrent, with strong correlations between corticosteroids and antimetabolites (Phi = 0.77), asparaginase enzymes and antimetabolites (0.71), corticosteroids and asparaginase enzymes (0.69), and corticosteroids and vinca alkaloids (0.57; see Supplementary Fig. 1).
CMRFs were highly prevalent, affecting 58% of survivors. Prevalent CMRFs were defined using the highest Common Terminology Criteria for Adverse Events (CTCAE) grade recorded before or after the blood draw for DNAm. Obesity was most common (41.0%) and defined as CTCAE grade ≥3, (BMI ≥ 30 kg/m² in adults or ≥95th percentile for their age and sex in children). Hypertension (29.2%), hypertriglyceridemia (20.4%), hypercholesterolemia (16.4%), and abnormal glucose (10.4%) were defined as CTCAE grade ≥2, reflecting moderate or more severe diseases. Despite the high prevalence of CMRFs, co-occurrence was modest (pairwise Phi <0.35), except for hypertriglyceridemia and hypercholesterolemia (Phi = 0.78; Supplementary Fig. 2). Survivors had a median of two CMRFs, with 1242 exhibiting none, 827 one, 406 two, 235 three, 178 four, and 78 all five CMRFs.
To support causal inference in downstream mediation analyses, we separately defined incident cases as those developing or worsening after DNAm sampling. Incidence was lower than prevalence given short duration of follow-up since blood draw for DNAm; obesity (8.7%), hypertension (7%), hypertriglyceridemia (2.7%), hypercholesterolemia (1.7%), and abnormal glucose (2.5%). These incident cases provided temporal alignment for investigating DNAm mediation of treatment-related cardiometabolic risk.
DNAm signatures of CMRFs reveal distinct and shared epigenetic associations
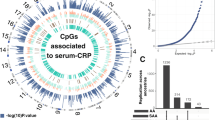
To delineate the epigenetic landscape underlying post-treatment cardiometabolic risks, we performed separate EWAS analyses for each of the five CMRFs using prevalent case status (Fig. 1b). Across 752,683 CpGs, we identified 1894 unique CpGs significantly associated with one or more CMRF (p < 9 × 10−8), yielding a total of 1912 CpG-CMRF associations. Obesity, the most prevalent CMRF, had the largest number of associated CpGs (n = 1720), followed by abnormal glucose (n = 201), and hypertriglyceridemia (n = 145), whereas hypercholesterolemia (n = 38) and hypertension (n = 34) yielded fewer associations (Fig. 2a). These CpGs showed distinct distributions of effect sizes by CMRF (Fig. 2b) and were enriched in epigenetically dynamic regions, particularly open sea regions and 5′ untranslated regions (UTRs), while being significantly less common in CpG islands and near transcription start sites (TSSs) (Supplementary Fig. 3).
a Circle Manhattan plots for each of the five CMRFs. Each plot displays EWAS results across autosomal chromosomes, arranged clockwise. Red points denoting CpGs significantly associated with the corresponding CMRF (p < 9 × 10−8), based on logistic regression models with two-sided hypothesis testing. The outer circular heatmap represents the local density of significant CpG associations across the genome, with darker shading indicating a higher number of significant CpGs within the genomic bins. Inner concentric tracks correspond to individual CMRFs, color-coded by outcome as indicated by the legend. b Boxplots showing the distribution of effect sizes (absolute log odds ratio) for each CMRF-DNAm association, ordered by median effect size. Effect sizes are shown for CpGs associated with abnormal glucose (n = 201), hypercholesterolemia (n = 38), hypertriglyceridemia (n = 144), hypertension (n = 34), and obesity (n = 1716). Boxes indicate the interquartile range (IQR), center lines denote medians, and whiskers extend to 1.5× the IQR. c UpSet plot illustrating the number of unique and overlapping CpGs across the five CMRFs. The large obesity-specific cluster (n = 1574 CpGs) was omitted to highlight remaining intersections. For example, 27 CpGs overlapped between obesity and hypertriglyceridemia. Set size bars on the right shows the number of significantly associated CpGs per CMRF. Source data are provided as a Source Data file. CMRF cardiometabolic risk factor, DNAm DNA methylation, EWAS epigenome-wide association study.
Most CMRF-associated CpGs (91%; 1715/1893) were specific to a single condition, while 9% overlapped across two or more CMRFs (Fig. 2c). The largest overlaps occurred between obesity and abnormal glucose (n = 74 CpGs), obesity and hypertriglyceridemia (n = 27), and obesity, abnormal glucose, and hypertriglyceridemia (n = 16). Genes mapped to shared CpGs were enriched in metabolic and inflammatory pathways, including the FTO (fat mass and obesity associated) Obesity Variant Mechanism12, JAK-STAT13, and IL-2/STAT514 signaling. Additionally, hypertriglyceridemia and obesity showed co-enrichment in fatty acid beta-oxidation and integrin signaling, suggesting a unified metabolic–inflammatory axis (Supplementary Data 1).
Strikingly, five CpGs (cg00574958, cg03725309, cg05325763, cg17058475, cg22976567) were associated with all five CMRFs, suggesting potential regulatory hubs. Three (cg00574958, cg05325763, cg17058475) mapped to CPT1A, a gene central to fatty acid metabolism15,16, while cg22976567 mapped to LMNA, a gene implicated in heart disease and dyslipidemia17,18. These shared loci suggest an interconnection across key biological pathways underlying cardiometabolic dysfunction in survivors. The presence of these shared CpGs highlights possible convergence points across metabolic and cardiovascular pathways, underscoring the biological interconnectedness of post-treatment cardiometabolic dysfunction in survivors.
To separate total from lifestyle-independent associations, we refit EWAS with prespecified additions of smoking and physical activity, then body mass index (BMI) (non-obesity outcomes), and diet quality (Health Eating Index [HEI]) in a HEI-complete subset of survivors. Across conditions, smoking/physical activity introduced little changes, whereas most loss of statistical significance occurred when BMI was added. Retention of significant results from the baseline model ranged from high for obesity (no BMI; 81% retained) to low for hypertension (8%), with intermediate retention for hypertriglyceridemia (48%), hypercholesterolemia (37%), and abnormal glucose (16%; Supplementary Fig. 4). Effect sizes for retained loci were highly concordant across models (Spearman ρ = 0.75–0.96), indicating mainly uniform shrinkage rather than sign reversals (Supplementary Fig. 5); median absolute attenuation of the regression β coefficient was modest (2–11%; up to 18% for the single hypertension locus; Supplementary Fig. 6). In the HEI-complete subset, adding HEI had negligible additional impact (ρ = 0.99 vs. the BMI+smoking+physical activity model; Supplementary Fig. 7). Notably, multi-phenotype “hub” CpGs, including three in CPT1A, remained robust for obesity and hypertriglyceridemia but were particularly weakened for hypertension after BMI adjustment (Supplementary Fig. 8).
Relevance of CMRF-associated CpGs in survivors relative to the general population
To assess whether CMRF-associated CpGs identified in survivors reflect broader cardiometabolic biology, we cross-referenced them with the EWAS Catalog19, which aggregates findings from primarily non-cancer populations. We observed significant enrichment for CpGs associated with C-reactive protein (CRP)—a biomarker of systemic inflammation20 (one-sided hypergeometric test, Benjamini–Hochberg [BH] p < 9.29 × 10−81, Supplementary Data 2). CRP-associated CpGs were highly represented among those linked to hypertension (90.9%), hypercholesterolemia (81.6%), hypertriglyceridemia (81.4%), and abnormal glucose (85.1%), yet notably lower in obesity (57.3%).
While many CMRF-associated CpGs overlapped with loci reported in general population EWAS, a substantial subset, particularly among obesity-associated CpGs, appeared unique to survivors. Obesity had the highest proportion of CpGs not reported for any trait in the EWAS Catalog (14%), compared to abnormal glucose and hypercholesterolemia (8%), hypertriglyceridemia (6%), and hypertension (3%), suggesting survivor-specific epigenetic signatures. Among the 1720 obesity-associated CpGs, 155 (9%) had been linked to BMI, waist circumference, or obesity-related traits (Supplementary Data 3). While some CpGs align with established obesity phenotypes, a substantial fraction appears unique to this cohort, potentially reflecting survivor-specific epigenetic signatures (Supplementary Data 4). These findings underscore both shared and unique components of cardiometabolic risk in survivors.
Cancer treatment exposure is associated with widespread DNAm variations
We next examined the extent to which cancer treatments were associated with CMRF-related CpGs, focusing on seven chemotherapy agents (asparaginase enzymes, alkylating agents, antimetabolites, anthracyclines, corticosteroids, epipodophyllotoxins, and vinca alkaloids) and two site-specific radiotherapies (body-trunk-RT and brain-RT; Fig. 1c). We did not examine associations between primary cancer diagnosis and DNAm, as cancer diagnoses are highly collinear with treatment exposures in this cohort (See “Methods”; Supplementary Data 5). Among 1893 CpGs associated with prevalent CMRFs, 64% (1218 CpGs) were also associated with at least one treatment (BH-adjusted p < 0.05), resulting in 2574 significant treatment-CpG associations (Fig. 3).
Heatmap illustrating CpGs significantly associated with cancer treatment exposure. Associations between treatments and DNAm were tested using linear regression models with two-sided hypothesis testing, and multiple testing was controlled using the BH false discovery rate (FDR < 0.05; N = 2938 survivors). Rows represent CpG sites and columns represent treatment exposures; blue indicates significant associations after FDR correction. Bar plots above the heatmap indicate the number of significant CpG associations per treatment. Columns on the right summarize CMRF associations for each CpG, with CpGs sorted by the number of CMRFs to which they are associated. Source data are provided as a Source Data file. BH Benjamini–Hochberg, CMRF cardiometabolic risk factor, RT radiotherapy.
RT exposures had the strongest epigenetic signatures, with body-trunk-RT (n = 603) and brain-RT (n = 499), accounting for most associations. Among chemotherapies, alkylating agents (n = 346) and asparaginase enzymes (n = 319) had the most CpG associations, while vinca alkaloids (n = 67) had the fewest. Over half (55%) of treatment-associated CpGs were linked to multiple treatments; 12 CpGs were associated with seven or more treatments. One CpG, cg04880464, was associated with eight therapies (all but vinca alkaloids) and also associated with prevalent abnormal glucose, hypertriglyceridemia, and obesity; this CpG has previously been linked to CRP levels in the EWAS Catalog.
Overlap between treatment- and CMRF-associated CpGs varied by condition (Supplementary Fig. 9). Body-trunk-RT showed the highest median overlap across CMRFs (25.1%), followed by alkylating agents (16.8%), while vinca alkaloids showed minimal overlap (1.5%). Of note, obesity-associated CpGs were less frequently associated with treatments; 23.4% of obesity-associated CpGs showed no treatment associations, compared to 7.5% for hypertension, 4.3% for hypertriglyceridemia, 2.4% for abnormal glucose, and 1.9% for hypercholesterolemia, indicating that a subset of obesity-related DNAm variations may be independent of treatment exposures.
DNAm partially mediates the effects of cancer treatments on CMRFs
To investigate whether DNAm acts as an intermediary between cancer treatments and CMRFs, we examined the three-part axis of treatment, DNAm, and CMRF (Fig. 1d). To preserve the temporal sequence necessary for causal inference, we limited this analysis to incident CMRF events arising after DNAm blood sampling (Table 1). CpGs were selected for mediation testing if they were significantly associated with both a treatment and a CMRF in incident-only models; some prevalent associations did not replicate, likely reflecting reduced statistical power (Supplementary Fig. 10). We applied a counterfactual-based mediation framework21 to each of eleven treatment-CMRF pairs. Following prior methodology5, we defined “strong mediators” as CpGs with BH-adjusted average causal mediation effect (ACME) p < 0.05 that explained more than 10% of the total effect. Using this criterion, we identified 24 CpGs mediating five treatment-CMRF associations (Fig. 4a and Supplementary Data 6). Mediation proportions ranged up to 24%, suggesting that DNAm variation may partially explain the cardiometabolic toxicity associated with cancer treatment exposure.
a Heatmap summarizing treatment-CMRF pairs with evidence of DNAm mediation (N = 2938 survivors). “Strong mediators” are defined as CpG sites with a statistically significant average causal mediation effect (ACME; two-sided BH p < 0.05) and a mediation proportion >10%, estimated using regression-based mediation analysis. Each cell is annotated with the maximum proportion of the total treatment effect mediated and the number of mediator CpGs identified (total: independent), where independent CpGs represent correlation-pruned mediators. b, c Composite mediation models for abnormal glucose (N = 2938 survivors). For each analysis, correlated mediator CpGs were grouped by correlation structure and four representative CpGs were selected. Composite mediators were constructed as the sum of centered and scaled DNAm values across representative CpGs. Mediation was assessed using regression-based mediation analysis with two-sided hypothesis testing, in which path a was estimated using linear regression of the composite mediator on treatment, path b using logistic regression of abnormal glucose on the composite mediator conditional on treatment, and paths c and c’ using logistic regression models of abnormal glucose on treatment without and with adjustment for the mediator, respectively. Path coefficients (a, b, c, c’), effect estimates, and exact p values are shown in the schematic. The composite mediator exhibited a statistically significant ACME (two-sided p < 0.05), accounting for b 11.7% of the total brain-RT effect and c 22.4% of the total body-trunk-RT effect on abnormal glucose. Source data are provided as a Source Data file. CMRF cardiometabolic risk factor, Max. maximum, RT radiotherapy.
Several CpGs mediated more than one treatment-CMRF association, suggesting shared epigenetic pathways. Five CpGs (cg00574958, cg05325763, cg09737197, cg17058475, cg26731839) mediated the effects of both brain-RT and body-trunk-RT on abnormal glucose. Another CpG (cg09921385) mediated the effect of both body-trunk-RT and anthracyclines on hypertriglyceridemia, highlighting potential convergence of chemotherapy and radiotherapy pathways in driving metabolic dysregulation.
Of particular note, cg20370568 mediated 20% of the effect of body-trunk-RT on abnormal glucose (ACME p = 1.0 × 10−3). This CpG is located near ANTXR2, a gene implicated in extracellular matrix (ECM) remodeling, inflammatory regulation, and vascular homeostasis22, supporting its potential relevance in therapy-related metabolic dysregulation.
Core DNAm mediators preserved despite lifestyle and outcome adjustments
Restricting the mediation analysis to incident-only CMRFs (i.e., those developed strictly after the DNAm blood draw) produced modest reductions in proportion mediated (<5% attenuation) for most condition-treatment pairs, with the largest decrease for abnormal glucose (4.2%) and a small increase for hypercholesterolemia (+2.0%; Supplementary Fig. 11). By treatment, attenuation was 4.9% for brain-RT, 1.8% for anthracyclines, and 1.6% for body-trunk-RT. ACME estimates were highly consistent across definitions (Spearman ρ = 0.89, p = 3.3 × 10−7, Supplementary Fig. 12), and effect directions were preserved for all 30 trios. Retention was full for abnormal glucose with body-trunk-RT (7/7 retained) and hypercholesterolemia with brain-RT (1/1 retained), partial for hypertriglyceridemia with anthracyclines (6/8), and attenuated for abnormal glucose with brain-RT (5/12). All five CpGs mediating radiotherapy effects on abnormal glucose in the primary analysis (cg00574958, cg05325763, cg09737197, cg17058475, cg26731839) remained significant with at least one exposure, and cg20370568 near ANTXR2 retained robust mediation (20.6%, p < 1 × 10−3, Supplementary Data 6).
We next evaluated robustness to lifestyle covariates adjustment, sequentially adding smoking/physical activity and then BMI. For abnormal glucose and hypercholesterolemia, proportion mediated was largely unchanged after smoking/physical activity (<0.2% attenuation) but decreased after BMI (2.7 and 3.5%, respectively, Supplementary Fig. 13), with similar trends for ACME (Supplementary Fig. 14). In contrast, hypertriglyceridemia showed greater reduction at the smoking/physical activity step (4.78%) with little further change after BMI (0.44%). ACME rankings were strongly preserved across models (Spearman ρ = 0.92, p < 1 × 10−6; ρ = 0.72, p < 1 × 10−5 after sequential adjustment), and effect directions were unchanged. Retention patterns mirrored these shifts, with some loss for brain-RT mediators after BMI (abnormal glucose 3/12 retained; hypercholesterolemia 0/1 retained; Supplementary Fig. 15), while trunk-RT mediators were fully retained. Notably, CpGs at CPT1A and ANTXR2 (cg20370568) remained significant, including cg20370568 mediating 15% after BMI adjustment (5% attenuation, p = 0.006; Supplementary Data 6).
Composite methylation scores capture cumulative mediation effects
Among the five treatment-CMRF pairs with strong mediators, four involved multiple CpGs that met mediation criteria, with a median of seven mediators per pair (range, 2–12), representing 23 unique CpGs in total (Supplementary Data 6). Using the correlation-pruned mediator sets (defined in “Methods”), we constructed a composite DNAm score for each treatment-CMRF pair and re-estimated mediation. The composite analyses yielded a single DNAm signature-level indirect effect (Fig. 1e).
Two treatment-CMRF pairs—body-trunk-RT and brain-RT with abnormal glucose—each had four independent mediators based on pairwise correlation. Mediation analysis using the resulting composite scores revealed significant effects: 12% of the association between brain-RT and abnormal glucose (ACME p < 1 × 10−3; Fig. 4b) and 22% for body-trunk-RT and abnormal glucose (ACME p < 1 × 10−3; Fig. 4c) were mediated by DNAm.
Sensitivity analyses using the minimum (rather than the maximum) ACME estimate to select representative CpGs yielded similar results, with mediation proportions of 13% for brain-RT and 20% for body-trunk-RT, both significant at ACME p < 1 × 10−3. These results support the robustness of the composite methylation score approach in capturing multi-CpG mediation effects and provide further evidence that DNAm may play a role in the biological embedding of treatment exposures contributing to CMRFs.
DNAm associated with expression of genes in CMRF-related pathways
To examine whether CMRF-associated CpGs influence gene expression, we conducted a cis-eQTM analysis in 199 survivors with matched blood-based transcriptomic data (Fig. 1f). Using linear regression, we tested CpG-gene pairs within ±1 megabase (Mb) of a gene’s TSS, identifying 79 CpGs (4.2%) significantly associated with expression of 76 genes (eGenes, BH-adjusted p < 0.05), yielding 83 unique eQTM pairs. Most eGenes (91%) were associated with a single CpG (eCpG). Mapping these eQTMs to their respective CMRFs revealed 100 CMRF-eQTM associations, predominantly from obesity (n = 72 eQTMs), followed by abnormal glucose (n = 18), hypertriglyceridemia (n = 5), and hypercholesterolemia (n = 3), and hypertension (n = 2). Many eCpGs had prior associations with metabolic and inflammatory traits in the EWAS Catalog (Supplementary Data 7), suggesting functional relevance and possible causal contributions to CMRFs.
We also identified 144 treatment-eQTM associations (Fig. 5a), with body-trunk-RT (n = 31) and brain-RT (n = 25) showing the most associations, followed by alkylating agents (n = 24), asparaginase enzymes (n = 21), and anthracyclines (n = 14). Several CpGs were associated with multiple treatments. Notably, the eQTM cg06096184-LRIG1, was associated with seven treatments, implicating LRIG1—a negative regulator of epidermal growth factor receptor (EGFR) signaling23—in shared epigenetic response to therapy-induced cellular stress. This CpG has been associated with smoking, aging, COPD, CRP levels, type 2 diabetes, and nitrogen dioxide exposure in the EWAS Catalog.
a Number of CpGs associated with both CMRFs and treatments that were identified as eQTMs (N = 199 survivors). b Scatter plot showing the association between DNAm at cg20370568 and ANTXR2 expression (N = 191 survivors). The solid line represents the ordinary least-squares linear regression fit, and the shaded band indicates the 95% confidence interval of the fitted mean. Pearson correlation coefficients and two-sided p-values are shown. c Mediation diagram showing cg20370568 partially mediates the effect of body-trunk-RT on abnormal glucose (N = 191 survivors), with a 20.3% proportion of the effect mediated. Mediation was assessed using regression-based mediation analysis with two-sided hypothesis testing, in which path a was estimated using linear regression of the CpG mediator on treatment, path b using logistic regression of abnormal glucose on the CpG mediator conditional on treatment, and paths c and c’ using logistic regression models of abnormal glucose on treatment without and with adjustment for the mediator, respectively. Path coefficients (a, b, c, c’), effect estimates, and exact p values are shown in the schematic. d Stratified scatter plots showing the association between cg20370568 DNAm and ANTXR2 expression among survivors exposed or unexposed to body-trunk-RT (N = 191 survivors). Solid lines represent ordinary least-squares linear regression fits and shaded bands indicate the 95% confidence intervals of the fitted means. Pearson correlation coefficients, two-sided p-values, and sample size (N) for each group are shown. Source data are provided as a Source Data file. CMRF cardiometabolic risk factor, RT radiotherapy, TPM transcripts per million.
Finally, cg20370568, identified as a mediator of body-trunk-RT on abnormal glucose, was significantly associated with expression of ANTXR2 (β = −0.179, BH p = 0.048; Fig. 5b, c). The inverse DNAm-expression association was substantially stronger in individuals exposed to body-trunk-RT (Pearson r = −0.48, p = 2.49 × 10−5) compared to unexposed individuals (Pearson r = −0.11, p = 0.21), and a formal interaction test confirmed a significant DNAm-by-RT interaction on gene expression (p = 0.0035; Fig. 5d). This CpG has also been associated with CRP levels, type 2 diabetes, and COPD in the EWAS Catalog (Supplementary Data 7). Its dual identification as both a treatment-related mediator and an eQTM highlights it as a compelling candidate for future mechanistic investigations of therapy-related cardiometabolic risk.
Discussion
In this comprehensive epigenetic analysis of survivors of childhood cancer, we identified DNAm signatures that may explain how cancer treatments become biologically embedded and contribute to long-term cardiometabolic risk. Our findings highlight five key insights. First, nearly 2000 CpGs were associated with at least one CMRF, revealing widespread DNAm variation tied to metabolic dysfunction. Second, over half of these CpGs were also associated with prior cancer therapies—particularly radiotherapy—suggesting persistent treatment-induced epigenetic remodeling. Third, causal mediation analysis identified 24 CpGs that mediated treatment-CMRF associations, with mediation proportions up to 24%, suggesting that DNAm partially accounts for the cardiometabolic consequences of therapeutic exposures. Fourth, composite methylation scores integrating independent mediator CpGs captured robust, multi-site mediation effects for body-trunk- and brain-RT on abnormal glucose. Finally, cis-eQTM analysis linked several CMRF-associated CpGs to gene expression, including cg20370568, a potential regulator of ANTXR2 and glucose metabolism through vascular and ECM pathways. Together, these findings support a model in which persistent DNAm variations encode the biological impact of treatment exposures, ultimately shaping transcriptional activity and disease risk. By identifying CpGs that mediate treatment effects and affect gene expression, this study provides mechanistic insights and a foundation for future work to validate DNAm markers and assess their potential clinical utility in survivorship care.
Our findings suggest that cardiometabolic dysfunction in survivors is driven by both shared and condition-specific DNAm signatures. Obesity accounted for the largest fraction, but substantial overlap across abnormal glucose and hypertriglyceridemia points to interconnected pathways. Enrichment in open sea regions and regulatory elements, rather than CpG islands, highlights distal regulation in post-treatment metabolic risk. Cross-referencing with the EWAS Catalog further revealed that while many signals coincide with inflammatory pathways, especially CRP-related CpGs, a notable subset, particularly obesity loci, appeared unique to survivors. Five CpGs, including sites in CPT1A and LMNA, were shared across all CMRFs, suggesting regulatory hubs underlying broad cardiometabolic vulnerability in survivors. Together, these patterns support inflammation as a core axis while also pointing to survivor-specific DNAm that warrant further mechanistic and translational investigation.
Treatment exposures were strongly associated with DNAm signatures, with radiotherapy and alkylating agents producing the broadest effects. Over half of treatment-associated CpGs were linked to multiple exposures, suggesting convergent epigenetic responses. At the same time, subsets of CpGs remained condition-specific, pointing to distinct treatment pathways. Together, these patterns highlight that both shared inflammatory mechanisms and treatment-specific DNAm alterations shape long-term cardiometabolic risk in childhood cancer survivors.
Many obesity-associated CpGs were not linked with treatment exposures, suggesting pre-existing susceptibilities or alternative pathways unrelated to therapy. Indeed, a subset of obesity-related CpGs overlapped with signals from the non-cancer EWAS population in the EWAS Catalog, indicating potential commonality with the general population. Nevertheless, a considerable fraction was unique to survivors, implying additional or distinct epigenetic influences on adiposity in this cohort. One possibility is that these CpGs capture baseline host genetic or lifestyle factors, such as diet24, physical inactivity25,26, or altered endocrine function27. Future longitudinal studies and replication in independent cohorts will be needed to clarify whether these CpGs represent predisposition to obesity, reflect broader environmental influences, or constitute novel survivor-specific modifications. Understanding these mechanisms will be crucial for developing targeted interventions to mitigate long-term excess adiposity in survivors of childhood cancer.
Our mediation analysis supports a model in which DNAm acts as a molecular intermediary linking cancer therapies to long-term cardiometabolic risk in survivors. Multiple CpGs significantly mediated treatment-CMRF associations, suggesting convergence on shared molecular pathways. For instance, five CpGs jointly mediated the effects of both brain-RT and body-trunk-RT on abnormal glucose, implicating common radiation-induced epigenetic mechanisms. Composite methylation scores, which aggregated independent mediator CpGs, explained up to 22% of the treatment-CMRF association; consistent with or exceeding mediation proportions reported in prior studies across metabolic, environmental, and psychosocial exposures (typically <1% to 20%28). These findings highlight the potential of DNAm to encode the lasting biological impact of therapy and identify epigenetic signatures as mechanistic biomarkers and possible targets for risk reduction.
Although only a subset of CMRF-associated CpGs showed significant cis-eQTMs (likely due to limited sample size), these highlight potential regulatory pathways through which DNAm influence cardiometabolic risk. Obesity showed the highest number of CpG-gene expression associations (i.e., eQTMs), consistent with its widespread DNAm variations, while body-trunk-RT had the most treatment-related eQTMs. Notably, cg06096184, associated with LRIG1, a negative regulator of EGFR signaling23, was associated with seven treatments, suggesting a broad epigenetic response to therapy-induced stress. Given the role of LRIG1 in cancer progression, therapy resistance, and long-term toxicity23,29, variations in its DNAm and expression may carry important implications for survivorship care, particularly in mitigating late cardiometabolic effects.
One of the most compelling findings was the dual role of cg20370568 as both a treatment-related mediator and an eQTM. This CpG accounted for 20% of the association between body-trunk-RT and abnormal glucose and was inversely associated with ANTXR2 (also known as CMG2) expression—particularly among survivors exposed to body-trunk-RT. ANTXR2 is involved in vascular homeostasis, ECM remodeling, and collagen VI turnover22,30, processes essential for maintaining insulin delivery and glucose uptake31,32. While loss of ANTXR2 function promotes fibrosis and insulin resistance33,34, our mediation model suggests that increased ANTXR2 expression (linked to lower DNAm) was associated with greater risk of abnormal glucose. While this may seem counterintuitive, it may reflect a maladaptive or compensatory response to radiation-induced tissue injury, where elevated ANTXR2 signals persistent ECM remodeling or inflammation. Alternatively, blood-based expression may not fully reflect the regulatory role of ANTXR2 in insulin-sensitive tissues such as adipose, liver, or muscle. Prior associations of cg20370568 and CRP levels, type 2 diabetes, and COPD in the EWAS Catalog further support its relevance to systemic inflammation and metabolic disease. Together, these findings suggest cg20370568 may serve as an epigenetic bridge between cancer therapy and long-term cardiometabolic risk in survivors.
Lifestyle factors such as smoking, physical activity, and BMI are well-established determinants of both DNAm and cardiometabolic health and thus represent important potential confounders. Their centrality also underscores the translational relevance of our findings, as these modifiable exposures may themselves become targets for intervention. To evaluate robustness, we adjusted for these factors in EWAS and mediation sensitivity analyses. Attenuation was modest overall and greatest in models with BMI adjustment (notably for abnormal glucose and hypercholesterolemia), yet many associations and mediation findings persisted. These results suggest that our findings of underlying molecular mechanisms linking treatment exposures to cardiometabolic risk are not due to confounding of lifestyle effects.
Despite the strengths of this study, several limitations should be acknowledged. First, because prevalent CMRFs at blood collection were included to maximize EWAS discovery power, reverse causality cannot be excluded; mediation analysis restricted to incident events helped mitigate this, but future longitudinal studies are needed to establish temporality. Second, although CMRFs were defined using standardized, clinically accepted thresholds and adjudicated by a clinical panel, some misclassifications are possible due to borderline measurements or incomplete ascertainment. Attrition contributes to the latter, though returnees and non-returnees were largely comparable35. Follow-up time was also modestly longer in some incident cases; we therefore adjusted for the follow-up duration in all models, and incident-only analyses yielded consistent results. Loss to follow-up differed by baseline status; survivors with a baseline CMRF had lower loss than those without (21.5% vs. 35.4%; Chi-square p < 1 × 10−3). While SJLIFE does not select participants for return core visits based on cardiometabolic risk, ancillary studies sometimes target survivors with specific high-risk profiles36,37. Survivors with baseline CMRFs may therefore have been preferentially invited for additional assessments in these ancillary studies, contributing to their longer average follow-up. This pattern reduces concern that attrition obscured progression captured by our primary (increase/worsening) endpoint. Conversely, because baseline-negative survivors were more often lost, estimated incident rates are likely conservative. Taken together, adjustment for follow-up time and concordant findings across worsening and incident-only endpoints argue against serious bias due to attrition. Third, residual confounding from lifestyle or other metabolic factors may remain, despite directed acyclic graph (DAG)-guided covariate adjustment and lifestyle sensitivity analyses. Fourth, generalizability is limited: SJLIFE is a single-center, predominantly European-ancestry cohort, where referral patterns, treatment protocols, and supportive care resources may not reflect community or resource-limited settings35. Enrollment beginning in 2007 also introduces potential survival bias by excluding earlier-treated survivors who did not survive to study onset. While participation rates are high, underrepresentation of those with the greatest burden of late effects remains possible. We therefore interpret our mediation estimates as internally valid and hypothesis-generating, with replication needed in independent, more diverse cohorts; where feasible, standardization and transportability methods will be used to map effects to target populations. Fifth, exclusions for missing genotype or radiotherapy data were rare (<1%) and not associated with measured demographic or treatment characteristics, making substantial selection bias unlikely, though residual bias from unmeasured factors cannot be excluded. Finally, DNAm was measured in peripheral blood, which may not fully reflect epigenetic or transcriptional variability in metabolically active organs like the liver, adipose tissue, or vascular endothelium, though it remains highly clinically translatable.
Our mediation framework rests on several assumptions common to causal inference. First, there should be no major uncontrolled exposure–outcome confounding, and second, no major uncontrolled mediator–outcome confounding. We addressed these through covariate adjustment informed by a DAG, but acknowledge that residual confounding may remain. Third, the analysis assumes no mediator–outcome confounders that are themselves affected by the exposure, which is plausible in our setting but cannot be directly verified. Finally, causal interpretation assumes no exposure–mediator interaction; evaluating this requires sufficient power and may be feasible in future studies with larger sample size and longer follow-up (i.e., more incident CMRFs). Although these assumptions cannot be directly tested, sensitivity analyses (incident-only restriction and lifestyle adjustment) yielded broadly concordant results. While our results are consistent with a causal framework in which treatment influences CMRFs through DNAm, the observational design and single-time-point DNAm measurement preclude definitive causal inference. Accordingly, these findings should be interpreted as evidence of statistical mediation consistent with, but not conclusive proof of, a causal pathway. Future work should benchmark high-throughput, correlation-aware mediation frameworks that jointly model many CpGs and incorporate formal sensitivity analyses to further solidify causal interpretations.
Our findings lay the foundation for a new paradigm in survivorship research—one that leverages epigenetic biomarkers to explore how treatment exposures shape long-term health. Mediator CpGs not only reveal biological mechanisms but also have potential for risk prediction and monitoring. These epigenetic markers could guide earlier identification of high-risk individuals and inform surveillance strategies and targeted interventions. DNAm signatures may also serve as surrogate endpoints for evaluating interventions aimed at reducing cardiometabolic risk. To clarify the temporal and functional relevance of these findings, longitudinal studies with pre-treatment DNAm profiling and multi-omics integration will be critical. Ultimately, incorporating DNAm markers into risk models and exploring their modifiability may enhance risk assessment and survivorship care, with the goal of improving long-term outcomes for survivors of childhood cancer.
Methods
Statistics & reproducibility
This study is an observational analysis of childhood cancer survivors enrolled in the SJLIFE cohort. Sample sizes were determined by the availability of DNAm, genotype, gene expression, and clinically ascertained cardiometabolic phenotype data within the cohort. No statistical method was used to predetermine sample size. Primary analyses included 2938 survivors of European ancestry with available DNAm data and clinically ascertained CMRFs. Sex was determined by clinical records and was included as a covariate in all relevant models; sex-stratified analyses were not performed due to limited statistical power. Age and other demographic characteristics are summarized in the cohort description.
Participants were not financially compensated beyond standard SJLIFE clinical evaluations. To minimize population stratification, analyses were restricted to survivors of European ancestry and adjusted for genetic principal components. Survivors were excluded if genotype data for ancestry inference, clinical CMRF ascertainment, or detailed radiotherapy exposure information were unavailable, yielding a final analytic sample of 2938 participants. No additional data were excluded, and no evidence of systematic selection bias was observed.
All statistical analyses, including epigenome-wide association analyses, mediation analyses, and eQTM analyses, were performed in R (version 4.4.1). Logistic and linear regression models were used where appropriate, with two-sided hypothesis testing. Multiple testing was controlled using the Benjamini–Hochberg false discovery rate. Additional statistical tests included Fisher’s exact, hypergeometric, and Kruskal–Wallis tests. All tests were two-sided and p-values < 0.05 were considered statistically significant unless otherwise noted. Randomization and blinding were not applicable since this is an observational study.
Study population and study design
This study utilized data from SJLIFE, a retrospectively constructed cohort with prospective follow-up to assess long-term health outcomes in survivors of childhood cancer treated at the SJCRH. Enrollment of individuals who survived 5+ years from diagnosis began in 2007 and is currently ongoing. The study design, eligibility criteria, and follow-up protocols have been described in detail previously35. Study procedures were approved by the SJCRH Institutional Review Board (IRB), and all participants provided written informed consent prior to participation.
The overall study design is illustrated in a conceptual framework (Fig. 1) with the following procedures: First, we profiled DNAm of PBMCs in SJLIFE survivors using the Illumina EPIC v1 BeadChip (Fig. 1a). Second, we mapped CpGs associated with each of five CMRFs in cross-sectional analyses of prevalent status as of the most recent follow-up by epigenome-wide association (EWAS; Fig. 1b). Third, we assessed whether these CMRF-associated CpGs were also associated with cancer treatments, including seven chemotherapy agents (asparaginase enzymes, alkylating agents, antimetabolites, anthracyclines, corticosteroids, epipodophyllotoxins, and vinca alkaloids) and two body region-specific radiotherapies (body-trunk-RT and brain-RT; Fig. 1c). Fourth, CpGs linked to both a treatment and a CMRF were carried forward to causal mediation analysis to estimate indirect effects along the treatment-DNAm-CMRF pathway (Fig. 1d). Fifth, for treatment-CMRF pairs with multiple CpG mediators, we clustered correlated CpGs (r2 ≥ 0.05), retained the most significant mediator from each cluster, and then created a composite DNAm signature-level mediator by summing their z-score standardized DNAm values (Fig. 1e). Finally, in a subset with matched transcriptomes of PBMCs (n = 199 survivors), we performed cis-eQTM analyses to connect mediator CpGs to nearby gene expression (Fig. 1f). Full model specifications, covariates, multiple-testing procedures, and CMRF endpoint definitions are detailed in “Methods” subsections further below.
Participants
Given the SJLIFE cohort’s demographic composition (>80% are individuals of European ancestry), the current availability of DNAm data, and the need to minimize population stratification, analyses were restricted to survivors of European ancestry. Among 3059 survivors with DNAm data, 3044 had full genotype data (for genetic ancestry), and 2966 had CMRFs clinically ascertained. We further excluded 28 participants without radiotherapy details, yielding a final sample of 2938. Potential selection bias arising from these exclusions was evaluated as described in the “Methods: Statistical Analysis”; no systematic associations between exclusion and measured pre-exposure characteristics were detected (Supplementary Data 8).
CMRFs
CMRFs were ascertained within the SJLIFE cohort using a standardized protocol that integrates retrospective medical record abstraction, patient- or proxy-reported health questionnaires, and comprehensive in-person clinical evaluations conducted at baseline and approximately every 5 years thereafter. These evaluations included physical examinations, laboratory testing, and diagnostic assessments, supplemented by external medical records to verify interim medical events. This ensured consistent ascertainment across baseline and follow-up.
Outcomes were graded using a modified version of the National Cancer Institute CTCAE (version 4.03)38, with grades ranging from 0 (no problem) to 4 (life-threatening). A multidisciplinary clinical review panel developed a standardized rubric to harmonize definitions and ensure clinical relevance. As previously described39, grading thresholds were adapted from established guidelines, including the National High Blood Pressure Education Program (hypertension) and Centers for Disease Control and Prevention definitions for BMI percentiles (obesity), with pediatric-specific thresholds applied where appropriate. Abnormal glucose metabolism, hypertriglyceridemia, and hypercholesterolemia were defined using CTCAE laboratory and grading criteria (see Supplementary Methods).
To account for potential bias from differential follow-up time, we defined follow-up time as the interval from blood draw for DNAm to each participant’s last clinical assessment and included this variable as a covariate in all models (see “Statistical analysis: Mediation analysis subsection”). Prior work in SJLIFE has shown that survivors who return for follow-up are largely comparable to those who do not across demographic and clinical characteristics, with only modest differences in diagnostic group distribution35. Attrition in follow-up based on the baseline cardiometabolic risk was evaluated and incorporated as detailed in the “Methods: Statistical Analysis” section, comparing follow-up by incident status and baseline CMRF and adjusting all models for follow-up duration.
For analysis, each CMRF was coded as a binary variable, indicating the presence or absence of the condition. The highest recorded CTCAE grade was used to determine case status for each participant. A CTCAE grade of ≥2 indicated clinically significant abnormal glucose, hypercholesterolemia, hypertriglyceridemia, and hypertension. A more stringent threshold of CTCAE grade ≥3 was applied for obesity (consistent with the clinical definition of BMI ≥ 30 kg/m2 in adults and BMI ≥95th percentile for their age and sex in children). These thresholds were selected based on clinical relevance, ensuring that cases captured reflect clinically significant disease burden (see Supplementary Methods for grading criteria).
For EWAS analyses, CMRFs were classified as prevalent events, meaning they were considered present if they had occurred at any time before or after blood collection. This approach maximized statistical power and allowed for comprehensive assessment of DNAm correlates of disease status. For mediation analyses, in order to maintain temporal alignment with causal mediation assumptions, we restricted our analysis to incident CMRF events, meaning they were considered present only if they developed or worsened after the blood collection (i.e., an increase in CTCAE grade following baseline reaching grade ≥2; Table 1).
Cancer treatments
Cancer treatment histories were abstracted from medical records, capturing both chemotherapy and radiotherapy (RT) exposures with potential relevance to cardiometabolic risk. Chemotherapeutic agents included alkylating agents (classic), anthracyclines, antimetabolites, asparaginase enzymes, corticosteroids, epipodophyllotoxins, and vinca alkaloids. RT exposures were classified based on the primary exposed body region, including brain-, chest-, abdominal-, and pelvic-RT. Due to strong correlations among abdominal-, pelvic-, and chest-RT (Phi coefficient > 0.65), and their systemic effects on peripheral blood-derived DNAm, we generated a composite variable body-trunk-RT to indicate exposure to any of these three regions. This approach reduced collinearity while preserving biological relevance in analyses of RT-associated DNAm variations. All chemotherapy classes and RT exposures were defined as binary variables indicating whether a given treatment was administered within the first 5 years following cancer diagnosis, consistent with the categorization shown in Table 1. Treatments administered beyond this period were uncommon (<6% of survivors), indicating that nearly all exposures occurred during the initial treatment phase. A sensitivity analysis excluding a small proportion of survivors who received any treatment beyond 5 years post diagnosis yielded results consistent with the primary analysis. Each of the eleven treatment classes were considered separately for all statistical analyzss.
Covariates
To account for potential confounders, all analyses were adjusted for key demographic, genetic, and leukocyte composition variables. Age at blood draw was included for models where DNAm was the dependent variable, while age at the most recent clinical visit was used for models assessing CMRFs to ensure alignment with the most updated health status and minimize biases due to variation in follow-up time. Sex was included as a binary covariate. To adjust for population stratification, we included the top ten principal components (PCs) derived from principal component analysis of common genetic variants in whole-genome sequencing (WGS) data. Global DNAm variability was accounted for by including the top four DNAm PCs, selected based on a scree plot and the proportion of variance explained, ensuring that key axes of variation were captured while minimizing overfitting. Leukocyte subtype compositions were estimated from the DNAm data using the Houseman method40, including proportions for B cells, CD4+ T cells, CD8+ T cells, monocytes, neutrophils, and natural killer (NK) cells. Additionally, experimental batch effects were assessed and adjusted in models incorporating gene expression data.
Lifestyle variables (smoking, physical activity, BMI, and diet quality by the Healthy Eating Index [HEI-2010]) were incorporated in prespecified sensitivity models. Smoking was ascertained by survey and coded as ever vs. never (≥100 lifetime cigarettes). Physical activity was coded as ≥150 vs. <150 min/week of at least moderate activity. BMI was modeled categorically; adults (underweight <18.5 kg/m2; normal 18.5–24.9 kg/m2; overweight 25.0–29.9 kg/m2; obese ≥30 kg/m2) and youth (<20 years old) using CDC growth-chart41 percentiles (underweight <5th; normal weight 5–85th; overweight 85–95th; obese ≥95th). HEI-2010 was derived from a nutrition questionnaire using standard scoring which assesses adherence to the 2010 Dietary Guidelines for Americans42. To reflect potential confounding at the time of DNAm blood draw, we used lifestyle measurements obtained at or nearest to the DNAm visit, preferring pre-draw values when available; measurements within 12 months prior to and up to 3 months after the DNAm draw were eligible, with the value closest in time selected when multiple observations were available. Due to missing HEI, two analysis cohorts were used; a larger cohort with smoking, physical activity, and BMI (N = 2037) and an HEI-complete subset with all four lifestyle variables (N = 1382).
DNAm bioinformatic analysis
Genome-wide DNAm data were generated using Illumina Infinium MethylationEPIC BeadChip v1 (Illumina, San Diego, CA, USA), with DNA extracted from PBMCs. Preprocessing was conducted using the minfi v1.46.0 package43 in R, following established quality control procedures including a detection p-value ≤ 0.01. CpGs were excluded if they were located on sex chromosomes, contained single nucleotide polymorphisms, or were previously identified as cross-reactive or non-specific. Quantile normalization was applied at the probe level to ensure comparability across samples. After these steps, 752,683 CpGs remained.
To adjust for potential confounding, we next performed a CpG-specific regression for each of the 752,683 CpGs. Specifically, the M-value at each CpG was regressed on age at blood draw for DNAm, sex, leukocyte subtype proportions, the top ten genetic PCs, and the top four DNAm PCs (as described above). The resulting residuals were then used in subsequent analyses. Age at DNAm assessment was included because it is highly correlated with both time since primary cancer diagnosis and time since treatment completion (Pearson r = 0.88 and r = 0.85, respectively; both p < 2.2 × 10−16), thereby serving as an effective proxy for latency since diagnosis and treatment. Including age at DNAm in the residualization step helps account for interindividual differences in latency, ensuring subsequent models capture variation not attributable to heterogeneity in time at blood draw for DNAm since diagnosis.
Gene expression bioinformatic analysis
Gene expression data were generated from PBMC RNA sequencing, with raw counts quantified using htseq-count44 and aligned to the hg38 human reference genome. Gene expression levels were converted to transcripts per million (TPM), and genes with a median TPM ≤ 1.0 across samples were excluded (considered to have very low or negligible expression). Matched DNAm and RNA sequencing data from the same blood draw were available for 199 individuals. To ensure consistency in downstream analyses, log2-transformed quantile-normalized TPM values were residualized using the same covariates applied in DNAm analyses, including age at blood draw, sex, leukocyte subtype proportions, and genetic ancestry PCs, with an additional adjustment for RNA sequencing batches, and top five gene expression PCs.
CpG-gene mapping
For EWAS functional analyses, CpGs were mapped to their nearest genes using annotations from the Illumina EPIC v1 manifest. For expression quantitative trait methylation (eQTM) analysis, CpGs were mapped to genes based on proximity to the TSS within a ±1 megabase (Mb) window, a threshold commonly used in eQTM studies to capture potential cis-regulatory interactions45,46. Of 12,231 genes analyzed, 7099 contained at least one CpG within the 1 Mb TSS window. In total, 14,513 CpG-gene pairs were identified, with each CpG mapped to a median of two genes (mean 2.05, range [1-9]). Conversely, each gene was associated with a median of six CpGs (mean 8.79, range [1-42]).
Statistical analysis
Selection bias assessment
We evaluated potential selection bias arising from exclusions by modeling the probability of exclusion (1 = no WGS or missing RT details; 0 = included) using logistic regression with pre-exposure covariates (sex, age at diagnosis, diagnosis group). In a secondary analysis, we additionally included chemotherapy class indicators to test whether exclusion varied by treatment exposure or follow-up time. No measured covariate was associated with exclusion (all two-sided p > 0.10, Supplementary Data 8), and estimates were essentially unchanged after adding treatment indicators (Supplementary Data 9).
CMRF EWAS
To identify differentially methylated CpGs associated with CMRFs, we performed an EWAS for each condition using logistic regression models. Analyses included all 752,683 CpGs that passed quality control. Each model treated the binary CMRF outcome as the dependent variable, with the residualized M-value for each CpG standardized (i.e., centered and scaled across samples) as the independent variable, ensuring effect sizes are expressed per one standard deviation change in the methylation level for each CpG. Covariate adjustments included age at the most recent follow up, sex, and relevant treatment exposures. Each treatment class was considered individually and was modeled as a binary variable (1 = exposed, 0 = unexposed), with survivors unexposed to that treatment serving as the reference group. Because many survivors received multiple treatments, other treatment exposures were included as covariates in the same model to account for co-exposure and ensure that associations reflect the independent effect of each treatment class. To minimize collinearity and over-adjustment, co-exposures were included only if (i) they were associated with the respective CMRF in the baseline clinical model (see below) and (ii) showed low correlation with the treatment of interest (Phi < 0.4). Childhood cancer treatments are often administered in standardized combinations determined by diagnosis and protocol and including all treatment classes simultaneously would include high collinearity and obscure interpretable associations. Genome-wide significance was set at p < 9 × 10−8, consistent with established threshold in EWAS to account for multiple testing while preserving statistical power. A summarized statistical analysis workflow can be found in Supplementary Fig. 16.
A sensitivity analysis with lifestyle variables was performed for each CMRF. Because lifestyle data were incomplete, we evaluated the impact of lifestyle adjustments in two cohorts: a larger lifestyle cohort with smoking, physical activity, and BMI available (N = 2037) and an HEI cohort with all four variables including the HEI (N = 1382). Building on top of the same base model for the primary EWAS, we fit a prespecified sequence of nested models that added lifestyle covariates sequentially—smoking, then physical activity, then BMI (BMI omitted when the outcome was obesity); in the HEI cohort, HEI was added last. Lifestyle measurements followed the timing rules described above (closest to DNAm; ≤12 months prior and ≤3 months after). We summarized the impact of lifestyle by (i) retention of genome-wide significant CpG-CMRF associations at each step (same threshold as primary EWAS: p < 9 × 10−8), and (ii) concordance of effect estimates between adjacent sequential models using Spearman correlations.
CMRF-treatment associations
To assess associations between cancer treatments and CMRFs, we performed logistic regression analyses for each treatment-CMRF pair, with the CMRF as the dependent variable and the treatment of interest as the independent variable. A model adjusting for sex and age at the most recent follow-up, and for other treatments whose Phi coefficient was below 0.4 with the exposure treatment, thereby minimizing collinearity while preserving distinct treatment effects5. Because the variables were coded as binary, the Phi coefficients here are numerically equivalent to the Pearson correlation coefficients (r). Significant treatment-CMRF associations were used for subsequent analyses (Supplementary Fig. 10).
Pathway enrichment analysis
Pathways enrichment analysis was performed to identify biological pathways associated with CMRF-associated CpGs and eQTMs using the enrichR 3.2 R package47. To maximize biological interpretability, we incorporated multiple curated pathway databases covering diverse cellular functions and disease mechanisms, including Reactome Pathways 2024, WikiPathways 2024 Human, BioPlanet 209, KEGG 2021 Human, Elsevier Pathway Collection, MSigDB Hallmark 202, BioCarta 2016, HumanCyc 2016, GO Biological Process 2023, GO Cellular Component 2023, GO Molecular Function 2023, MSigDB Oncogenic Signatures, and dbGaP. These databases were selected for their relevance to molecular signaling, metabolic processes, transcriptional regulation, and disease-related pathways, ensuring comprehensive coverage of potential mechanisms linking DNAm to CMRFs. Pathways were considered significantly enriched if they met a Benjamini–Hochberg (BH)-adjusted p-value threshold of <0.05.
CpG enrichment in genomic features
To evaluate whether CMRF-associated CpGs were over- or underrepresented in specific genomic regions, we performed enrichment analyzss using gene annotations from the Illumina EPIC v1 manifest. This analysis aimed to determine whether differential methylation preferentially occurred in regulatory regions, potentially influencing gene expression and cardiometabolic risk. We tested six CpG island annotations (island, north shore, north shelf, south shelf, south shore, open sea) and eight genomic functional regions (TSS1500, TSS200, 5′UTR, first exon, gene body, 3′UTR, exon-intron boundaries, intergenic). Enrichment was assessed using Fisher’s exact test, comparing the distribution of CMRF-associated CpGs to all CpGs included on the EPIC array as the background set. Odds ratios and 95% confidence intervals were calculated to quantify effect sizes. Multiple-testing correction was applied using the BH procedure across all 14 tested regions to account for multiple comparisons.
Targeted analysis of treatment-associated DNAm
To assess the effect of cancer treatments on DNAm, we performed an association analysis restricted to the CMRF-associated CpGs. Associations between residualized DNAm and treatment exposures were tested using linear regression models, with each treatment modeled separately as a binary independent variable (treated vs. untreated). Models adjusted for sex and non-correlated treatments (Phi coefficient <0.4) to account for potential confounding while minimizing collinearity. BH correction was applied to adjust for multiple comparisons across all CpG-treatment tests, with a significance threshold set at FDR < 0.05.
We focused on treatment rather than primary cancer diagnosis because in SJLIFE, treatment regimens are strongly determined by primary diagnosis, yielding high collinearity between diagnosis and treatment exposures. Logistic regression models predicting each treatment from diagnosis yielded highly significant associations (BH-adjusted p < 1 × 10−30; Supplementary Data 5), indicating that diagnosis and treatment are not independently distributed. To avoid multicollinearity, we focused our analysis on treatment-specific DNAm signatures, consistent with previous literature48.
Mediation analysis
To test whether DNAm mediated the effect of treatment on CMRF risk, mediation analysis was performed on treatment-CMRF-CpG trios that met three criteria: the CpG was significant in both the CMRF EWAS and treatment EWAS, and the treatment was significantly associated with the CMRF in a clinical model (described earlier).
Mediation analyses were performed to evaluate whether DNAm statistically mediated the associations between treatment exposures and subsequent cardiometabolic outcomes. All treatments occurred in childhood or adolescence, DNAm was measured years later in adulthood (median 21.5 years after treatment, IQR 13–30 years), and outcomes were restricted to incident cases that developed after the DNAm assessment (i.e., they were not present prior to or at DNAm assessment and were first detected at a subsequent clinical visit). This temporal structure establishes a clear sequence from treatment to DNAm to outcome.
Confounder choice was guided by a DAG from the treatment (X) to DNAm (M) and further to CMRF (Y) (Supplementary Fig. 17). The DAG defined two sets: exposure-outcome confounders denoted as C(XY) and mediator-outcome confounders denoted as C(MY). C(XY) included sex, age at most recent follow-up, relevant treatment co-exposures (treatments associated with the CMRF but not highly correlated with the exposure of interest; Phi <0.4; same as in the CMRF EWAS), and genetic ancestry. Primary cancer diagnosis was not included because it is highly collinear with treatment modality. C(MY) included age at DNAm blood draw, sex, relevant treatment co-exposures, leukocyte subtype proportions, and genetic ancestry. Lifestyle factors (smoking, physical activity, BMI) measured before DNAm were treated as potential C(MY); to avoid blocking putative indirect pathways, these were evaluated in additional prespecified sensitivity analyses rather than in the primary mediation models. This DAG-based approach aims to control confounding while avoiding over-adjustment for other variables plausibly on the causal pathway or affected by the exposure, ensuring valid estimation of direct and indirect effects.
Follow-up time was included as a covariate to account for variation in observation windows. We defined follow-up time as the interval (in years) from the date of blood draw for DNAm to each participant’s last clinical assessment. To assess whether differential follow-up time could bias incident-event analyses, we compared follow-up time by incident status (1 vs. 0) within each CMRF using Kruskal–Wallis tests with BH adjustment. Participants who went on to develop a new or worsening CMRF after blood draws were, on average, followed longer than those who have not yet for several conditions (e.g., abnormal glucose metabolism, obesity: BH-adjusted p = 0.041). To mitigate potential bias, models included follow-up duration, defined as the interval between blood draw for DNAm and the most recent clinical visit, ensuring that estimated indirect effects via DNAm were not driven by unequal at-risk periods.
We quantified loss to follow-up (no clinical visit after DNAm) by baseline CMRF status (present vs. absent) using two-sample tests of proportions within each condition and overall; participants with a baseline CMRF were consistently less likely to be lost to follow-up (overall 21.5% vs. 35.4%, Chi-square p = 2.2 × 10−36), with significant differences for abnormal glucose, hypertriglyceridemia, hypercholesterolemia, and hypertension, but not obesity.
Mediation was performed using the mediate function in the mediation v4.5.0 package in R21, with 1000 bootstrap simulations to estimate confidence intervals. The mediation framework combines two interrelated regression models (mediator and outcome) to estimate indirect (ACME) and direct (average direct effect, ADE) effects. The mediator model, capturing how DNAm depends on treatment exposure, was a linear regression with DNAm as the dependent variable and treatment of interest, sex, and relevant co-exposures (as used in the CMRF EWAS) as predictors. The outcome model, capturing how the CMRF depends on both treatment and DNAm, was a logistic regression with the binary incident CMRF outcome as the dependent variable. Predictors included DNAm residuals, sex, age at the most recent clinical follow-up, follow-up duration (included as a covariate to adjust for unequal observation time), and relevant co-exposures. The two models were jointly analyzed using the mediation framework21, which integrates their parameter estimates to quantify how much of the treatment-CMRF association operates indirectly through DNAm versus directly through pathways independent of DNAm.
Multiple testing was controlling using the BH procedure. A CpG was considered a strong mediator if it had a BH-adjusted p-value for the ACME < 0.05 and a proportion mediated > 10%. The 10% threshold was selected to balance sensitivity and biological relevance.
To assess the robustness of mediation findings, we conducted two sensitivity analyses. First, we repeated mediation after restricting outcomes to incident-only events, defined as CMRFs that developed strictly after the DNAm blood draw, thereby excluding worsened cases. Incident-only models included the same covariates as the primary analysis. Second, we evaluated robustness to lifestyle adjustments. Guided by EWAS sensitivity results (which showed modest, similar attenuation for smoking and physical activity and the largest attenuation for BMI), we fit nested mediation models: (i) base model (primary analysis), (ii) base + smoking + physical activity, and (iii) base + smoking + physical activity + BMI (applied to non-obesity outcomes). Diet quality (HEI) was not included due to minimal impact in EWAS as well as substantial missingness (n = 655 [32%] missing HEI data). For trios significant in the primary analysis, robustness was evaluated by changes in proportion mediated, relative change in ACME (primary-incident)/primary, Spearman correlation of estimates, and concordance and retention of significant trios.
Composite mediation analysis
For treatment-CMRF pairs with multiple strong mediating CpGs, a composite mediation analysis was performed to assess the cumulative effects of DNAm. CpGs were first ranked by the estimated ACME in descending order. To reduce redundancy, CpGs were pruned using pairwise Pearson correlation (r2 < 0.05), retaining the most statistically significant mediator within each correlated cluster to ensure independent signals. Following pruning, composite DNAm scores were generated by summing z-score standardized M-values of the remaining independent CpGs within each treatment-CMRF pair. These composite scores were then used as single mediators in models with the same structure as the individual CpG analyses, adjusting for age, sex, and treatment co-exposures associated with the CMRF but not the exposure treatment. Multiple comparisons were controlled using BH correction.
eQTM analysis
To investigate potential gene regulatory mechanisms, an eQTM analysis was conducted using 199 participants with matched DNAm and RNA sequencing data from the same blood draw. All CMRF-associated CpGs were tested for association with expression levels of nearby genes, defined using a ±1 Mb window around the gene’s TSS. This window was chosen based on prior studies45,46 demonstrating that the majority of cis-acting methylation-expression relationships occur within this range. Linear models were used to test associations with residualized gene expression as the dependent variable and residualized DNAm as the independent variable. Multiple testing correction was applied across all eQTM tests using the BH procedure, with significance defined as FDR < 0.05.
Statistical software
All analyses were conducted in R 4.4.1, with visualizations generated using the ggplot2 v3.5.1 package49. Circle Manhattan plots were creating using CMplot from the rMVP v4.5.1 package50, while UpSet plots and heatmaps were generated using ComplexHeatmap v1.3.351. Logistic regression models were fitted using the glm function in R with a logit link function, and linear regression models were fitted using the lm function in R.
Ethics & inclusion statement
This study complies with all relevant ethical regulations. Research involving human participants was approved by Institutional Review Board (IRB) at St. Jude Children’s Research Hospital (SJCRH). The St. Jude Lifetime Cohort Study (SJLIFE) protocol was reviewed and approved by the SJCRH IRB, and all participants provided written informed consent for clinical evaluation, longitudinal follow-up, and collection of biospecimens for research use. Consent for participants younger than 18 years of age was provided by a parent or legal guardian. Genomic and clinical data were generated, stored, and analyzed under institutional policies that ensure protection of participant privacy and confidentiality. This research did not involve any stigmatizing, incriminating, or discriminatory risk to participants, and analyses were conducted with attention to equitable and respectful representation of all participants enrolled in the SJLIFE cohort.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
DNAm data generated and analyzed in this study are available through the NCBI Gene Expression Omnibus website under the accession number GSE314261. Parts of these data have been previously published with the accession number GSE16915652. Processed, aggregate-level summary statistics generated in this study have been deposited in Zenodo, including epigenome-wide association results for CMRFs (https://doi.org/10.5281/zenodo.17969754)53 and DNAm-treatment association results (https://doi.org/10.5281/zenodo.17970459)54. Source Data underlying Fig. 2a are provided via Zenodo (https://doi.org/10.5281/zenodo.17970924)55, while source data for all remaining figures are included with this paper. Mediation analysis results are provided in Supplementary Data 6, and eQTM results are provided in Supplementary Data 7.
Individual-level demographic, cancer treatment, and clinical phenotype data from participants in the St. Jude Lifetime Cohort (SJLIFE) are available under controlled access through the St. Jude Cloud Cancer Survivorship research domain (http://survivorship.stjude.cloud/). Access is restricted due to participant privacy protections and informed consent requirements and is subject to approval of a standard data-use agreement. Data access requests are reviewed on a rolling basis, with response times dependent on request complexity and committee review schedules. Approved access is granted for the duration specified in the data-use agreement and is limited to qualified researchers for cancer survivorship research purposes. Additional guidance on the application process is available on the St. Jude Cloud website, with support provided by the St. Jude Cloud team (support@stjude.cloud). Publicly accessible summary information for SJLIFE publications since January 2023 is available via the St. Jude Survivorship Portal (https://viz.stjude.cloud/community/cancer-survivorship-community~4/publications). The remaining data are available within the Article, Supplementary Information, or Source Data file. Source data are provided with this paper.
Code availability
No dedicated software package was developed for this study. All code used to generate the analyses reported in this paper is publicly available via Zenodo (v.1.0.0, https://doi.org/10.5281/zenodo.17992009)56.
References
Reulen, R. C. et al. Long-term cause-specific mortality among survivors of childhood cancer. JAMA 304, 172–179 (2010).
Lipshultz, S. E., Franco, V. I., Miller, T. L., Colan, S. D. & Sallan, S. E. Cardiovascular disease in adult survivors of childhood cancer. Annu. Rev. Med. 66, 161–176 (2015).
Hammoud, R. A. et al. Modifiable cardiometabolic risk factors in survivors of childhood cancer. JACC CardioOncol. 6, 16–32 (2024).
Dixon, S. B. et al. Specific causes of excess late mortality and association with modifiable risk factors among survivors of childhood cancer: a report from the Childhood Cancer Survivor Study cohort. Lancet 401, 1447–1457 (2023).
Song, N. et al. Persistent variations of blood DNA methylation associated with treatment exposures and risk for cardiometabolic outcomes in long-term survivors of childhood cancer in the St. Jude Lifetime Cohort. Genome Med. 13, 53 (2021).
Kirk, E. P. & Klein, S. Pathogenesis and pathophysiology of the cardiometabolic syndrome. J. Clin. Hypertens. 11, 761–765 (2009).
Pfeiffer, L. et al. DNA methylation of lipid-related genes affects blood lipid levels. Circ. Cardiovasc. Genet. 8, 334–342 (2015).
Irvin, M. R. et al. Epigenome-wide association study of fasting blood lipids in the genetics of lipid-lowering drugs and diet network study. Circulation 130, 565–572 (2014).
Gomez-Alonso, M. D. C. et al. DNA methylation and lipid metabolism: an EWAS of 226 metabolic measures. Clin. Epigenet. 13, 7 (2021).
Wielscher, M. et al. DNA methylation signature of chronic low-grade inflammation and its role in cardio-respiratory diseases. Nat. Commun. 13, 2408 (2022).
Dong, Q. et al. Distinct DNA methylation signatures associated with blood lipids as exposures or outcomes among survivors of childhood cancer: a report from the St. Jude lifetime cohort. Clin. Epigenet. 15, 32 (2023).
Laber, S. et al. Linking the FTO obesity rs1421085 variant circuitry to cellular, metabolic, and organismal phenotypes in vivo. Sci. Adv. 7, eabg0108 (2021).
Hu, X., Li, J., Fu, M., Zhao, X. & Wang, W. The JAK/STAT signaling pathway: from bench to clinic. Signal Transduct. Target. Ther. 6, 1–33 (2021).
Lin, J.-X. & Leonard, W. J. The role of Stat5a and Stat5b in signaling by IL-2 family cytokines. Oncogene 19, 2566–2576 (2000).
Tang, M. et al. CPT1A-mediated fatty acid oxidation promotes cell proliferation via nucleoside metabolism in nasopharyngeal carcinoma. Cell Death Dis. 13, 1–13 (2022).
Qu, Q., Zeng, F., Liu, X., Wang, Q. J. & Deng, F. Fatty acid oxidation and carnitine palmitoyltransferase I: emerging therapeutic targets in cancer. Cell Death Dis. 7, e2226–e2226 (2016).
Crasto, S., My, I. & Di Pasquale, E. The broad spectrum of LMNA cardiac diseases: from molecular mechanisms to clinical phenotype. Front. Physiol. 11, 761 (2020).
Rohde, K. et al. Genetics and epigenetics in obesity. Metabolism 92, 37–50 (2019).
Battram, T. et al. The EWAS Catalog: a database of epigenome-wide association studies. Wellcome Open Res. 7, 41 (2022).
Black, S., Kushner, I. & Samols, D. C-reactive Protein *. J. Biol. Chem. 279, 48487–48490 (2004).
Tingley, D., Yamamoto, T., Hirose, K., Keele, L. & Imai, K. Mediation: R package for causal mediation analysis. J. Stat. Softw. 59, 1–38 (2014).
Park, S. Y. et al. ANTXR2 is a potential causative gene in the genome-wide association study of the blood pressure locus 4q21. Hypertens. Res. 37, 811–817 (2014).
Billing, O., Holmgren, Y., Nosek, D., Hedman, H. & Hemmingsson, O. LRIG1 is a conserved EGFR regulator involved in melanoma development, survival and treatment resistance. Oncogene 40, 3707–3718 (2021).
Lan, T. et al. Plant-based, fast-food, Western-contemporary, and animal-based dietary patterns and risk of premature aging in adult survivors of childhood cancer: a cross-sectional study. BMC Med. 23, 120 (2025).
Hocking, M. C. et al. Prospectively examining physical activity in young adult survivors of childhood cancer and healthy controls. Pediatr. Blood Cancer 60, 309–315 (2013).
Chung, O. K. J., Li, H. C. W., Chiu, S. Y., Ho, K. Y. E. & Lopez, V. The impact of cancer and its treatment on physical activity levels and behavior in Hong Kong Chinese childhood cancer survivors. Cancer Nurs. 37, E43–51 (2014).
Casano-Sancho, P. & Izurieta-Pacheco, A. C. Endocrine late effects in childhood cancer survivors. Cancers 14, 2630 (2022).
Fujii, R., Sato, S., Tsuboi, Y., Cardenas, A. & Suzuki, K. DNA methylation as a mediator of associations between the environment and chronic diseases: a scoping review on application of mediation analysis. Epigenetics 17, 759–785 (2022).
Stutz, M. A., Shattuck, D. L., Laederich, M. B., Carraway, K. L. & Sweeney, C. LRIG1 negatively regulates the oncogenic EGF receptor mutant EGFRvIII. Oncogene 27, 5741–5752 (2008).
Bürgi, J. et al. CMG2/ANTXR2 regulates extracellular collagen VI which accumulates in hyaline fibromatosis syndrome. Nat. Commun. 8, 15861 (2017).
Pi, X., Xie, L. & Patterson, C. Emerging roles of vascular endothelium in metabolic homeostasis. Circ. Res. 123, 477–494 (2018).
Vicent, D. et al. The role of endothelial insulin signaling in the regulation of vascular tone and insulin resistance. J. Clin. Invest. 111, 1373–1380 (2003).
Khan, T. et al. Metabolic dysregulation and adipose tissue fibrosis: role of collagen VI. Mol. Cell. Biol. https://doi.org/10.1128/MCB.01300-08 (2009).
Dankel, S. N. et al. COL6A3 expression in adipocytes associates with insulin resistance and depends on PPARγ and adipocyte size. Obesity 22, 1807–1813 (2014).
Howell, C. R. et al. Cohort profile: the St. Jude Lifetime Cohort Study (SJLIFE) for paediatric cancer survivors. Int. J. Epidemiol. 50, 39–49 (2021).
Maharaj, A. et al. Design and methods of a randomized telehealth-based intervention to improve fitness in survivors of childhood cancer with exercise intolerance. Contemp. Clin. Trials 133, 107339 (2023).
St. Jude Children's Research Hospital. Complementary Behavioral Interventions To Remediate Cognitive Impairment or Emotional Distress in Cancer Survivors: A Pilot Study. https://clinicaltrials.gov/study/NCT06989463 (2025).
Common Terminology Criteria for Adverse Events (CTCAE) | Protocol Development | CTEP. https://ctep.cancer.gov/protocoldevelopment/electronic_applications/ctc.htm.
Hudson, M. M. et al. Approach for classification and severity grading of long-term and late-onset health events among childhood cancer survivors in the St. Jude Lifetime Cohort. Cancer Epidemiol. Biomarkers Prev. 26, 666–674 (2017).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinform. 13, 1–16 (2012).
Growth Charts - Percentile Data Files with LMS Values. https://www.cdc.gov/growthcharts/cdc-data-files.htm (2024).
Guenther, P. M. et al. The Healthy Eating Index-2010 Is a valid and reliable measure of diet quality according to the 2010 Dietary Guidelines for Americans123. J. Nutr. 144, 399–407 (2014).
Aryee, M. J., Jaffe, A. E. & Corrada-Bravo, H. Minfi: a flexible and comprehensive bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Keshawarz, A. et al. Expression quantitative trait methylation analysis elucidates gene regulatory effects of DNA methylation: the Framingham Heart Study. Sci. Rep. 13, 12952 (2023).
Kober, K. M., Berger, L., Roy, R. & Olshen, A. Torch-eCpG: a fast and scalable eQTM mapper for thousands of molecular phenotypes with graphical processing units. BMC Bioinform. 25, 71 (2024).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Im, C. et al. Trans-ancestral genetic risk factors for treatment-related type 2 diabetes mellitus in survivors of childhood cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 42, 2306–2316 (2024).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer New York, NY, 2016).
rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genomics Proteomics Bioinform. https://academic.oup.com/gpb/article/19/4/619/7230384?login=false.
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Song, N. GSE169156, NCBI Gene Expression Omnibus. Persistent Variations of Blood DNA Methylation Associated with Treatment Exposures and Risk for Cardiometabolic Outcomes in Childhood Cancer Survivors https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE169156 (2021).
Eulalio, T. SJLIFE cardiometabolic risk factor EWAS summary results. Zenodo https://doi.org/10.5281/zenodo.17969754 (2025).
Eulalio, T. Cancer treatment–associated DNA methylation signatures in childhood cancer survivors (SJLIFE). Zenodo https://doi.org/10.5281/zenodo.17970459 (2025).
Eulalio, T. Source Data for Fig. 2a: circle Manhattan plot of CMRF-associated DNA methylation in childhood cancer survivors. Zenodo https://doi.org/10.5281/zenodo.17970924 (2025).
Eulalio, T. tyeulalio/Survivorship_DNAm_mediators_of_CMRFs: Analysis pipeline and figure generation for DNAm mediators of cardiometaoblic risk in childhood cancer survivors. Zenodo https://doi.org/10.5281/ZENODO.17992009 (2025).
Acknowledgements
We thank all the participants in the St. Jude Lifetime Cohort. This research was supported by funding from the American Lebanese Syrian Associated Charities to St. Jude Children’s Research Hospital and by grants including V-Foundation (DT2020-014 [ZW]) and National Institutes of Health (CA290112 [ZW], CA279520 [ZW], and CA195547 [MMH and KKN]). We also thank Dr. Yadav Sapkota for providing constructive feedback on the revised manuscript.
Author information
Authors and Affiliations
Contributions
Z.W. conceived and supervised the study. T.E. and Z.W. designed and conducted statistical analyses, interpreted results, prepared figures, and drafted the manuscript. M.M.H. and K.K.N. oversaw recruitment of study participants and clinical data collection. K.S. curated clinical data and maintained SJLIFE data infrastructure used for analyses. H.M., E.P., and J.E. supervised and/or performed DNA extraction. E.W. and G.N. supervised and/or performed EPIC array scanning. X.M., Y.K., N.M.P., X.C., J.Z., M.N., J.T.L., N.C., ZiqiaoW, D.S., B.K., S.B.D., G.T.A., and all other authors contributed to interpretation of data and to reviewing and revising the manuscript. Primary funding for this project was acquired by Z.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Eulalio, T., Kim, Y., Meng, X. et al. Epigenome-wide analysis identifies DNA methylation mediators of treatment-related cardiometabolic risk in survivors of childhood cancer. Nat Commun 17, 1979 (2026). https://doi.org/10.1038/s41467-026-68689-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68689-6