Abstract
While neurogenesis is largely restricted to early life stages in animals, some taxa (including cnidarians, planarians and acoels) display lifelong neurogenic abilities. The cellular lineages and transcriptional programs underlying this process remain poorly understood in cnidarians. Combining reporter tracing and single-cell transcriptomics, we identify adult neurogenic trajectories in the sea anemone Nematostella vectensis. We uncover two distinct mechanisms: direct differentiation of peptidergic neurons from a multipotent progenitor pool, with identities specified proportionally to existing populations, and a stepwise maturation of cnidocytes (specialized cnidarian neural cells), marked by a transcriptionally distinct intermediate stage. Neural fate commitment is characterized by transient SoxC expression, with a homeodomain code associated with neural identities. These modular strategies support continuous neurogenesis and suggest that neural fate specification in cnidarians may use ancestral principles shared with bilaterians. Our study provides a foundational framework for understanding the mechanisms underlying the adult specification of neural cells and their evolutionary diversification across animal lineages.
Similar content being viewed by others
Introduction
Neurons are the fundamental units of nervous systems across animals. In most bilaterians (e.g., humans, mice, and flies), neural development is predominantly restricted to early life stages, with limited renewal capacities in adulthood. In contrast, lineages with extensive regenerative abilities and less complex nervous systems, such as cnidarians, acoels, and planarians, maintain ongoing neuronal renewal throughout life1. Cnidarians (e.g., sea anemones, corals, hydra, jellyfish), the outgroup to Bilateria, lack a centralized nervous system, with neurons embedded in diffuse nerve nets2,3. However, cnidarian and bilaterian neurons share key effectors, including conserved neuropeptide-processing machinery, ion channels, and synaptic proteins, as well as early neurogenic transcription factors like proneural basic helix-loop-helix (bHLH) and SoxB genes, pointing to a common evolutionary origin4,5,6,7,8,9,10. Major diversifications of transcription factor (TF) families central to bilaterian neural programs—such as Homeodomain, Sox, and bHLH—occurred early in animal evolution, with substantial cnidarian–bilaterian-shared expansions, which may have contributed to the diversification of neural identities across both groups11,12,13. Thus, investigating the cell lineages and transcriptional programs involved in neurogenesis in cnidarians could provide valuable insights into the mechanisms of lifelong generation of molecularly diverse neural cells across animals.
Cnidarian neural cells encompass ganglion neurons, sensory cells, and lineage-restricted cnidocytes (mechano-sensory neural cells). In the genetically tractable sea anemone Nematostella vectensis, over 20 subtypes have been molecularly characterized2,4,14,15,16,17,18,19, and its nervous system shows localized condensation of neurite tracts along the primary body axis, indicating a degree of morphological organization2,20. Neurons primarily employ a specialized neuropeptide repertoire4,15,16,17,18,19, along with other neurotransmitters (e.g., glutamate, GABA, but not monoamines)21,22,23. Akin to other highly regenerative species such as planarians, acoels, and annelids24,25, cnidarians like Nematostella exhibit feeding-dependent indeterminate growth and shrinkage, as well as asexual reproduction26,27. Notably, a population of peptidergic neurons (GLWa+) scales with body size15, suggesting that extensive adult neurogenesis occurs. However, the progenitor cell population(s) that enable adult neurogenesis and the transcriptional trajectories underlying this adult program of neural cell fate specification remain poorly understood.
Adult generation of neurons can occur via two distinct yet non-mutually exclusive mechanisms. Each of these mechanisms is associated with different degrees of fate restriction and nervous system organization. In the first, associated with highly organized centralized nervous systems, as in the rodent olfactory system or the zebrafish telencephalon, pools of lineage-restricted neural stem cells maintained within spatially delineated niches can generate a limited variety of neural subtypes that are integrated into nearby structures28. In the second mechanism, associated with more diffuse centralized nervous systems and nerve nets, as in planarians, acoels, and hydra, multipotent migratory stem cells can produce diverse somatic lineages, including most to all neuronal subtypes, under both homeostatic and injury-induced conditions29,30. In the adult sea anemone Nematostella vectensis, mesenterial Piwi1+ cells can give rise to cells expressing the conserved broadly neuronal marker Elav31, suggesting that they are a source for new neurons. It remains unclear how these putative stem cells contribute to adult neurogenesis in cnidarians, including which neurons they give rise to, whether intermediate lineage restriction occurs, and what molecular mechanisms accompany fate restriction.
In animals with continuous neural cell renewal—like cnidarians or planarians—the simultaneous presence of all cellular states (from stem cells to intermediate and mature neurons) presents a significant challenge for identifying cells along one or multiple lineage trajectories. This inability to distinguish temporal intermediates precludes trajectory inference and examination of their transcriptional maturation. Single-cell transcriptomics (scRNA-seq) can generate unbiased static snapshots of gene expression profiles across entire nervous systems, but lineage inference to investigate cell fate acquisition remains challenging32. Imaging of genetic drivers can capture cell dynamics through reporter inheritance, but reconstructing complex trajectories requires multiple targeted genetic tools. These issues are compounded in emerging model systems with limited genetic tools and sparse a priori knowledge needed for lineage reconstruction.
To address these challenges, we integrated temporal reporter tracing and single-cell transcriptome profiling to map the lineages and transcriptional programs underlying adult neurogenesis in the sea anemone Nematostella vectensis. We find that distributed neurogenesis scales the nerve net with peptidergic neurons generated directly from non-neural-exclusive progenitors, and cnidocytes maturing through a transcriptional intermediate stage before initiating their secondary neural program. These parallel transcriptional lineage strategies employ bilaterian-shared transcription factor classes (bHLH, homeodomain) across neural identities, suggesting common neural development principles for the generation of neural diversity.
Results
Distributed peptidergic neuron generation underlies adult nerve net expansion
To support indeterminate growth, an adult neurogenic program involving continuous generation and integration of neurons into the nerve net would be required. To identify such adult-born neural cells in vivo, we employed a characterized Elav regulatory sequence active in neurons from embryonic to adult stages to drive the expression of a photoconvertible KikumeGreenRed (KikGR) fluorescent reporter20,33. We labeled the standing population of Elav+ cells by photoconverting this reporter (day 0) in sexually immature adults and analyzed the body wall nerve net after 4 days. New Elav+ neurons arising from non-expressing cells would lack the photoconverted tracer and display new reporter expression (hereafter referred to as green cells) (Supplementary Fig. 1 and Fig. 1A). Thus, discriminating green neurons from previously generated stably expressing neurons (red and green co-expressing, orange cells hereafter) or cells that have ceased expressing the reporter (red cells hereafter), allowed us to investigate the dynamics of neurogenesis in situ.
A Active adult neurogenesis highlighted by Elav>KikGR reporter photoconversion and in situ imaging (see Supplementary Fig. 1). At 2, 4, and 7 days post-photoconversion, neural cells with the new green reporter signal and lacking the day 0 photoconverted reporter signal are newly expressing-only cells (green cells). Cells with the photoconverted tracer either also display new KikGR expression (ongoing expression, orange cells) or lack new KikGR expression (progeny cells, red cells). B Elav>KikGR+ body wall nerve net at 4 days post-photoconversion (5× view (left) and 40× single-neuron resolution (right)). Photoconverted KikGR day 0 tracer signal is shown in red, and the new KikGR expression in cyan. Arrows in insets highlight green neurons (in white), orange neurons (in orange and red neurons (in magenta). Scale bars are 500 µm (left), 50 µm (right) and 10 µm (insets). Experiment was performed three times with similar results. C Proportion of green, orange, and red Elav>KikGR+ cells detected in the oral and aboral regions (with hachure pattern) at 4 days post-photoconversion. For each image, cell counts are normalized by the total number of Elav>KikGR+ cells detected (n = 3 animals, mean ± SD). D Generation of Elav+ neurons from Elavnegative cells. Data is provided as Source Data File 1.
Imaging Elav>KikGR+ animals 4 days post-photoconversion revealed green neurons interspersed with orange neurons in the body wall nerve net (Fig. 1B), across the epidermal and subepidermal layers (Supplementary Movies 1 and 2), indicating ongoing neurogenesis contributing new neurons to the nerve net. Newly born neurons are dispersed throughout the epidermal and sub-epidermal layers, including in the neurite tracts at the base of mesenteries (Supplementary Movies 1 and 2). The majority of detected cells (~80%) are green or orange (Fig. 1C), with a small fraction red (Fig. 1B and C); the latter category suggests that a subset of differentiated neurons stop expressing the Elav>KikGR driver. Overall, the large fraction of green cells detected at 4 days post-photoconversion is consistent with active neurogenesis contributing new neurons to the body wall nerve net through the generation of Elav+ neurons from Elavneg cells (Fig. 1D).
To evaluate whether newly generated neurons are exclusively incorporated into specific regions of the nerve net, which could point to a spatially restricted growth zone on the oral side of the primary body axis, we examined both the oral and aboral regions. Green neurons are found in both regions at 2, 4, and 7 days post-photoconversion with similar proportions in each region (Fig. 1C and Supplementary Fig. 2). The synchronous presence of green cells in both regions, coupled with their high fraction, suggest that nerve net expansion occurs through highly active spatially distributed adult neurogenesis, with Elavneg cells differentiating into Elav+ neurons within one week.
Elav+ neurons encompass a diversity of neural cell states4,20. To investigate the molecular identity of the newly-generated Elav+ neural cells, we combined our photoconversion-based temporal recording with single-cell RNA-Sequencing (scRNA-Seq) of green, orange and red Elav>KikGR+ cells (Fig. 2A and Supplementary Figs. 3, 4). We identified 1331 peptidergic neurons from the 2764 Elav>KikGR+ sorted cells, highlighted by their shared expression of neuropeptide-processing genes (prohormone convertases34, amidating enzymes). These encompass over a dozen molecularly distinct subtypes, delineated by their restricted expression of neuropeptide precursors and neuropeptide G-protein coupled receptors (GPCR) (Fig. 2B, C and Supplementary Fig. 5)16,18. Notably, we recover all previously described neural populations in the body wall, including a Prdm14d+ population (Neural 0)14,15,16,18. As expected, we do not recover FoxQ2d+ putative sensory cells and mature cnidocytes, which are not labeled by the Elav>KikGR driver35. At 7 days post-photoconversion, we find a majority of the Elav>KikGR+ neural cells to be newly generated green cell (58%), consistent with our in situ nerve net imaging (Supplementary Fig. 6). Green cells are distributed among all peptidergic neuronal subtypes (Fig. 2D). Specifically, the number of green cells relative to the combined number of orange and red cells for each cluster was strongly correlated (R2 = 0.96, Pearson’s ρ). This suggests that Elav+ neuron subtypes, from the rarer to the more abundant, are generated concomitantly in vivo, at a frequency matching their overall proportions in the animal (Fig. 2E).
A Molecular identification of newly-generated Elav+ neurons. Plate-based scRNA-Seq (MARS-Seq protocol) allows recording of reporter status post-photoconversion (green, orange, or red cell) by flow cytometry for each cell sampled from Elav>KikGR+ adult body regions (body wall with mesenteries) at 2, 4, and 7 days post-photoconversion. scRNA-Seq transcriptional profiles are then matched to each cell’s reporter status color (Supplementary Figs. 3 and 4). B UMAP projection of the 1335 peptidergic neurons from the 2764 Elav>KikGR+ sampled cells from 2, 4, and 7 days post-photoconversion with their cluster identifier. C Scaled neural marker gene expression across Elav>KikGR+ peptidergic neuron clusters. Broadly expressed neural markers and neuropeptide processing genes (top) and specifically expressed neuropeptide precursor genes from15,16,17,18 (bottom). Dot size represents the percentage of cells in this cluster that express the gene. Neural cluster identifiers as in (B). See Supplementary Fig. 5 for expression across all Elav>KikGR+ cells. D Elav+ peptidergic neuron scRNA-seq UMAP projection highlighting the overlapping identity distribution between the previously labeled peptidergic neurons (orange and red cells) (left), and the newly expressing neurons (green) (right) at 7 days post-photoconversion. E Cell counts by reporter status (green vs. red and orange cells) across peptidergic cell clusters in 7-day photoconverted Elav>KikGR animals, showing a Pearson’s correlation R2 of 0.95. Line is the fitted linear regression with the 95% confidence interval in shaded gray. Cluster identifiers as in (B) and (C). Data is provided as Source Data File 2.
Progeny of FoxL2 + and SoxC + cells encompass neurons, cnidocytes and secretory cells
In silico trajectory inference approaches suggest that in the adult, peptidergic neurons, secretory cells, and cnidocytes (cnidarian-specific neural effector cells) are all part of a shared lineage36. However, it is unknown whether these populations derive from a common progenitor pool and what molecular features are associated with neural lineage restriction during adult growth. To better understand neural cell fate acquisition, we generated photoconvertible reporter lines for SoxC and FoxL2, two transcription factors expressed in putative progenitor cells in adults, to track their cellular progeny. Earlier work suggests SoxC plays a role in the differentiation of cnidocytes, secretory cells, and neurons during embryogenesis, and SoxC reporter expression has been noted across these cells in the adult36. FoxL2 is a forkhead box transcription factor highly expressed in progenitor cells and tentacular cnidocytes and lowly expressed in some neurons4,37. Together, these two reporters allowed us to probe the trajectories of early undifferentiated cell populations that may give rise to diverse neural cells in the adult.
We first investigated the temporal activity of SoxC during adult homeostatic growth. Imaging the body wall reveals SoxC>KikGR reporter expression in morphologically diverse cells, across the epidermal and subepidermal layers, including neurons, developing and mature cnidocytes, muscle cells, and secretory cells. To determine whether the neural cells might be part of a SoxC+-derived lineage, we assessed the presence of red cells post-photoconversion (Fig. 3A, B and Supplementary Movies 3 and 4). Thus, at 7 days post-photoconversion, the presence of both red (non-expressing progeny) and green (newly generated) mature cnidocytes and other neurons shows that these are derived from SoxC+ cells. This suggests that SoxC+ progenitor cells transiently express the driver early during differentiation and give rise to SoxCneg derivatives, including neurons and cnidocytes. Specifically, red neurons and cnidocytes represent cells generated prior to “day 0” of photoconversion, while green cnidocytes and neurons are those that developed within 7 days of the photoconversion “chase”. The recovery of primarily red and green cells at earlier timepoints (2 and 4 days post-photoconversion), in similar proportions in oral and aboral regions (Supplementary Fig. 7 and Supplementary Movies 5–7), further supports the conclusion that early expressing SoxC+ cells give rise to SoxCneg derivatives, including neurons and cnidocytes.
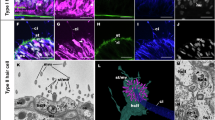
SoxC>KikGR+ (A) and FoxL2>KikGR+ (C) adult polyp body wall at 7 days post reporter photoconversion. Global field of view (5×, left) and high magnification image (40× extended depth of focus (A) or single image (C), right) with photoconverted KikGR signal shown in red and newly expressed KikGR in cyan. SoxC>KikGR+ cells encompass green, orange, and red mature cnidocytes, green, orange, and red neurons (respectively white, yellow, and magenta arrows), and orange secretory cells (yellow filled arrowhead). FoxL2>KikGR+ cells encompass orange (yellow arrow) and red (magenta arrows) neurons among other cells. Scale bars are 500 µm (left), 50 µm (right), and 10 µm in the insets. See Supplementary Figs. 8 and 9 and Supplementary Movies 3–9. Experiment performed at least two times with similar results. Proportion of red, orange and green SoxC>KikGR+ (B) and FoxL2>KikGR+ (D) cell bodies detected at 7 days post-photoconversion. Aboral end body wall images are indicated with a hachure pattern (n = 5, 6, 2, 2, respectively, for oral, SoxC>KikGR; aboral, SoxC>KikGR; and oral and aboral FoxL2>KikGR, average ± SD). E Global scRNA-Seq UMAP projection of 12,352 KikGR+ cells (2764 Elav>KikGR+ cells, 3451 SoxC>KikGR+ cells, and 6137 FoxL2>KikGR+ cells) at 2, 4, and 7 days post-photoconversion, colored by broad cluster identity. F Marker gene expression across clusters. Gene expression is normalized to the highest expression across clusters. Colors correspond to broad cluster annotations, as in (E). Dot size represents the proportion of cells expressing the gene in the cluster. Marker genes names are curated from refs. 13,14,15,16,17,18,39,42,43,47,89 (see Supplementary Data 1). G Cell identity distribution by driver and FACS gating strategy across the SoxC>KikGR, FoxL2>KikGR, and Elav>KikGR drivers. For the FoxL2>KikGR driver, the “KikGR+ no epi.” FACS gate depleted the epithelial epidermis cell fraction. Data is provided as Source Data File 3.
We next analyzed the expression of the FoxL2>KikGR driver. The reporter is expressed in multiple cell types in the body wall, across both epidermal and subepidermal layers, including neurons, mature cnidocytes and epithelial cells, and uncharacterized cells, with sparse secretory and muscle cells (Fig. 3C and Supplementary Figs. 8, 9). We thus investigated whether these differentiated cells derived from a FoxL2+ lineage (presence of red cells) or were actively expressing the reporter (orange cells). At 7 days post-photoconversion, we detect diverse red cells, including neurons and mature cnidocytes, suggesting these are non-expressing cells arising from FoxL2>KikGR+ cells (Fig. 3C). The sparse green cells present are mature cnidocytes and undetermined cells (Supplementary Figs. 8 and 9 and Supplementary Movies 8 and 9). This shows that FoxL2>KikGR+ cells give rise to multiple non-expressing derivatives within 7 days, including neurons. Accordingly, a large fraction of red cells (non-expressing progeny cells) are detected at 2, 4, and 7 days post-photoconversion (Supplementary Fig. 7). And this adult developmental process is spatially distributed across the primary body axis, as evidenced from the similar fraction of green/orange/red cells across both the oral and aboral regions (Fig. 3D and Supplementary Fig. 7). Overall, this separately highlights the presence of SoxC>KikGR+ and FoxL2>KikGR+ neural progeny cells encompassing both neurons and mature cnidocytes. These red non-expressing progeny cells thus arise from expressing cells (see below).
To investigate the molecular signatures of these adult neural developmental lineages, we combined the temporal recording of reporter activity with scRNA-Seq (workflow as in Fig. 2A). We sampled SoxC>KikGR+ and FoxL2>KikGR+ cells at 2, 4, and 7 days post-photoconversion (Supplementary Figs. 3, 4, and 10) and linked the molecular identity of each cell to its recorded reporter status (green, orange or red cell). We pooled 12,352 KikGR⁺ cells sorted from the SoxC>KikGR, FoxL2>KikGR, and Elav>KikGR drivers across 84 distinct MARS-Seq scRNA-Seq libraries from 27 independent samples and annotated 45 clusters using published marker genes4,5,14,16,18,35,36,38,40,41,42,42 (Fig. 3E–G and Supplementary Fig. 10). Clusters were then grouped into 13 broad cell classes, with groups Neural 0–4 representing Prdm14d+ neurons, Elavhigh peptidergic neurons, Elavlow peptidergic neurons, and other neural cells including FoxQ2d+ sensory cells and Eag+ mature cnidocytes43, respectively. Overall, our cluster resolution is consistent with subtype-specific gene expression patterns within peptidergic neurons, cnidocytes, and secretory cells18,40. We recover developing cnidocytes which express cnidarian-specific capsule proteins and known regulators, along with mucous cells, and digestive secretory cells (with distinct trypsins, chitinases, and pancreatic lipases)40. We identify epidermal epithelial cells, mesenteric retractor and parietal muscle cells4,44, cnidarian-specific gastrodermis cells, undetermined cells, and putative progenitor cells (SoxB(2)ahigh/SoxChigh). Peptidergic neurons are recovered from all three drivers (Elav>KikGR, FoxL2>KikGR, and SoxC>KikGR), while cnidocytes, mucous secretory cells, and epithelial epidermis cells are mostly absent from the Elav driver (Fig. 3G), consistent with the in situ imaging. The partially overlapping molecular identities of the neural populations suggest that peptidergic neurons represent a more restricted lineage distinct from other neural fates.
Differentiation of Piwi1 + progenitor cells
As putative progenitor cells display the highest SoxC levels (Fig. 3F and G), and neural cells can arise from unknown SoxC>KikGR+ cells (Fig. 3A), we examined these undifferentiated “progenitor” cells recovered from all drivers (in purple in Fig. 3E–G) as a source population for neural cell derivatives. These “progenitors” specifically express genes associated with proliferative pluripotent cells across metazoans, such as transcription factors from the bHLH (including Myc and atonal-related genes), Sox B and C families, RNA helicases (PL10, vasa1 and vasa2), and RNA-associated proteins (nanos1, nanos2, piwi1, piwi2) (Fig. 4A and Supplementary Figs. 11 and 12)30. Progenitors also display the strongest enrichment of both S-phase and G2/M cycling modules (Fig. 4A). In contrast to progenitors, neurons, cnidocytes, and secretory cells have no cell cycle module enrichment, suggesting they are primarily post-mitotic. Epidermis epithelial cells also display a cell cycle module enrichment, consistent with cell division in a subset of epidermis cells31,45,46.
A Molecular profiling of the progenitor cell pool recovered from the Elav>KikGR, FoxL2>KikGR and SoxC>KikGR drivers. (Top) S- phase and G2/M cycling genes module score across all clusters (same clustering and cluster colors as in Fig. 3E–G). (Bottom) Gene expression across all clusters. Dot size represents the proportion of cells in each cluster expressing the gene. Diverging temporal fates of progenitors. Ternary diagrams (B) and barplots with the recovered total cell numbers (C) showing the proportion of red, orange and green progenitor cells from the Elav>KikGR, SoxC>KikGR and FoxL2>KikGR drivers at 2, 4, and 7 days post-photoconversion. D Schematics of Piwi1+ progenitors giving rise to differentiated cells (top), with two possible expression dynamics: either expression of the driver occurs only in committed, differentiation-bound progenitors (middle) or already in self-renewing progenitors (bottom). For both scenarios, the corresponding progenitor cell color proportions at 7 days post-photoconversion are indicated on the right, with a majority of orange progenitor cells at 7 days indicative of driver expression in self-renewing progenitor cells, while a majority of green progenitor cells at 7 days is indicative of driver expression in differentiating and exit-committed progenitor cells. E Integrated model of progenitor cell temporal dynamics. Progenitors are a stable, self-renewing pool that expresses the FoxL2>KikGR driver; initiation of SoxC>KikGR or Elav>KikGR driver expression in progenitors is tied to differentiation initiation within days in vivo. Data is provided as Source Data File 4.
Subclustering of these progenitor cells uncovered both Piwi1high undifferentiated as well as putative differentiating states (Piwi1low) (Supplementary Fig. 12A–E). These Piwi1low cells also express either peptidergic neural markers, digestive secretory cell markers, or cnidocyte marker genes, respectively, in subclusters 7, 9, and 8, termed “precursors” (Supplementary Fig. 12A–C). These putative precursor states express lower levels of Piwi1 relative to the more undifferentiated states (clusters 0–6) (Supplementary Fig. 12C and D). A subset of the undifferentiated clusters displays a clear cycling signature enrichment (undifferentiated clusters 5, 3, and 6), seen across FoxL2>KikGR+ and Elav>KikGR+ cells. The low overall number of SoxC>KikGR+ cells sampled and the high fraction of putative cnidocyte precursors for this driver limit our analysis (Supplementary Fig. 12E). Both undifferentiated and putative precursor cells are labeled by the Elav>KikGR and FoxL2>KikGR drivers. Putative cnidocyte precursors are absent from Elav>KikGR⁺ cells.
Recent work on Nematostella stem cells showed that Piwi1+ cells can differentiate into Piwi1neg Elav+ neural cells in the body wall31. Here, we show that Piwi1+ progenitor cells are also recovered from the broadly neuronal Elav>KikGR driver20 (Fig. 3G). As Elav is highly expressed in peptidergic neurons and at low levels in progenitors (Fig. 3F), we hypothesized that the Elav+ progenitors identified here are early differentiation-committed cells. If so, progenitor cells recovered at 7 days post-photoconversion should be green, as the labeled progenitors present at day 0 would have differentiated (Fig. 4B–D). We thus evaluated whether the red and orange cell fractions were depleted in 7-day-post-photoconversion Elav>KikGR+ progenitors. We find that the majority of Elav>KikGR+ progenitors at 7 days (88%) are green. Accordingly, earlier timepoints (2 and 4 days) have intermediate red and orange progenitor cell fractions (Fig. 4B and C). This is consistent with a scenario in which Elav+ progenitors are differentiation-committed exit-bound cells (Fig. 4E).
Progenitor cells express SoxC and FoxL2 (Fig. 3F). Our detection of SoxC>KikGR+ and FoxL2>KikGR+ non-expressing neural derivatives with overlapping class identities (Fig. 3G) suggests a common progenitor pool (see below). However, the presence of green neurons labeled by the SoxC>KikGR driver but only orange and red neurons with the FoxL2>KikGR driver at 7 days post-photoconversion (Fig. 3A and C) reveals distinct temporal expression patterns during differentiation. To better understand this neural differentiation process, we evaluated when progenitors express SoxC relative to the initiation of differentiation. Specifically, as endogenous SoxC is recovered at a high level in progenitor cells (Fig. 3F and Supplementary Figs. 11 and 12), we assessed whether progenitors stably express this driver. If this were the case, using our photoconversion-based temporal recording, we would expect SoxC>KikGR+ progenitors to be mostly orange cells. Instead, we find that the majority of sampled SoxC>KikGR+ progenitor cells are green cells, at 2-, 4-, and 7-day post-photoconversion (Fig. 4B and C). This pattern suggests that SoxC driver initiation in progenitors is linked to a rapid differentiation initiation (Fig. 4D and E).
In marked contrast with the previous differentiation-committed dynamics observed for progenitors with both the Elav>KikGR and SoxC>KikGR drivers, progenitors recovered from the FoxL2>KikGR driver retain their photoconverted label and are either orange or red cells (>87% of progenitors) at 2, 4 and 7 days post-photoconversion (Fig. 4B). This suggests a stable self-renewing pool of progenitors where the FoxL2>KikGR driver is continuously expressed in cells that can differentiate (Fig. 4C and D), while expression of SoxC>KikGR or Elav>KikGR is associated with differentiation onset. This is further supported by the high endogenous expression of FoxL2 in progenitors (Fig. 3F and Supplementary Figs. 11 and 12), and that, even after 7 days post-photoconversion, FoxL2>KikGR+ progenitors remain overwhelmingly orange (Fig. 5B and C). Accordingly, at 7 days post-photoconversion, morphologically undifferentiated FoxL2>KikGR+ orange cells are detected in the body wall (Supplementary Figs. 8 and 9), whereas undifferentiated Elav>KikGR+ cells are green (Supplementary Fig. 13). Thus, the different drivers highlight that the population of progenitor cells is both self-renewing and differentiates into other cell states, consistent with a putative stem cell pool.
A Lineage tracing of peptidergic neurons from continuously labeled FoxL2 > KikGR+ Piwi1+ progenitors. These progenitors give rise to orange and red FoxL2 > KikGR+ peptidergic neurons at 2, 4, and 7 days post-photoconversion. Transiently expressing SoxC > KikGR+ progenitors generate peptidergic neurons with green, orange, and red reporter states present at 2, 4, and 7 days photoconversion, reflecting predicted differences between continuous (FoxL2 > KikGR) and transient (SoxC > KikGR) driver expression in the peptidergic lineage. Data shown in (B, C). B and C Proportion of green, orange and red Elav+ peptidergic neurons (Neural 1 class) recovered from FoxL2 > KikGR+ and SoxC > KikGR+ cells at 2, 4, and 7 days photoconversion. D Peptidergic neuron development: (top, scenario 1) neuron clusters encompass both differentiation intermediates and end states or (bottom, scenario 2) neuron clusters are distinct terminally differentiated subtypes generated in parallel. SoxC > KikGR labels early intermediate states (green or orange cells, no red cells), and late differentiated states accumulate red cells (scenario 1, see Supplementary Fig. 14). The ratio of green + orange to red cells across cell clusters distinguishes between intermediates or concurrent differentiated states within a cellular lineage. Letters indicate distinct neuron clusters. E Distribution of SoxC > KikGR+ peptidergic neuron clusters at 7 days post-photoconversion shows similar green + orange to red cells proportions across clusters, supporting that they are terminally differentiated neurons (scenario 2 in D) without intermediates among these. Additional data in Supplementary Fig. 14. F Proportional abundance of SoxC > KikGR⁺ and FoxL2 > KikGR⁺ peptidergic neural subtypes (in blue, R2 = 0.66). For comparison, digestive secretory cells are shown in orange and differentiated cell types that do not fit either proportional relationship within dashed lines. Additional data in Supplementary Fig. 14E. E, F 95% confidence interval of the fitted linear regression is shown in shaded gray. G Model of peptidergic neuron development: peptidergic-committed Piwi1⁺ multipotent cells differentiate into distinct peptidergic subtypes within 4 days in adults. Data provided in Source Data File 5.
Furthermore, while progenitors display molecular heterogeneity (Supplementary Fig. 12), undifferentiated Piwi1high progenitors are nevertheless recovered across both Elav>KikGR+ and FoxL2>KikGR+ cells, alongside Piwi1low peptidergic-specific precursor populations, among others. However, reporter tracing demonstrates that once the Elav>KikGR reporter is activated, progenitor cells no longer maintain the undifferentiated state but instead proceed toward differentiation (Fig. 4). Thus, activation of Elav>KikGR in undifferentiated progenitors is specifically linked to peptidergic neuronal differentiation, consistent with the absence of cnidocyte precursors among Elav>KikGR⁺ cells, and the lack of mature Elav>KikGR+ cnidocytes (also consistent with20,31). Overall, this uncovers a common pool of Piwi1+ progenitors with a stem cell-like molecular identity that self-renews and differentiates into multiple cell types (see below).
Direct differentiation of peptidergic neurons from SoxC +/FoxL2 + progenitors
We then examined the adult developmental lineage of peptidergic neurons, the largest neural class, from these Piwi1+ progenitors. Elav+ peptidergic neurons are recovered from all three drivers (Elav>KikGR, SoxC>KikGR and FoxL2>KikGR) (Neural 1 in Fig. 3E and G). To assess whether Piwi1+/SoxC+/Elavlow progenitors differentiate into peptidergic neurons, we tracked the derivatives of this population using the FoxL2>KikGR driver (Fig. 5A), as FoxL2>KikGR+ progenitors carry the photoconverted label even at 7 days post-photoconversion. We found that over 90% of peptidergic neurons recovered from the FoxL2>KikGR driver are either red or orange cells at 2, 4, and 7 days post-photoconversion (Fig. 5B and C). Thus, the presence of the photoconverted tracer label in peptidergic neurons after 7 days post photoconversion, together with the large fraction of red cells, is consistent with our proposed model of active neuron generation from FoxL2+/Piwi1+ progenitor cells. As SoxC expression in FoxL2+/Piwi1+ progenitors is transient and linked to differentiation initiation, if these Piwi1+/SoxC+ cells develop into peptidergic neurons, we predicted that the peptidergic neurons recovered from the SoxC>KikGR driver should encompass both red cells (cells generated prior to photoconversion), and an increasing fraction of orange and green cells (newly generated cells) at later timepoints (Fig. 5A). Therefore, we examined whether we could recover both green, orange and red peptidergic SoxC>KikGR+ neurons at all days post-photoconversion sampled (Fig. 5B and C). Overall, the simultaneous presence of red, orange, and green peptidergic neurons from the SoxC>KikGR driver, consistent with our in situ nerve net imaging data (Fig. 3A and C), supports our model of peptidergic neurons differentiating from FoxL2+/Piwi1+ progenitors, with a transient early expression of SoxC.
To identify the neural lineage(s) giving rise to peptidergic neuron subtypes, we next investigated whether the distinct peptidergic neurons (Fig. 2C) are terminally differentiated, or whether some are temporal intermediates. We thus compared the distribution of the newly generated cells (green and orange cells from the SoxC>KikGR driver) to the standing cell pool (red cells) across peptidergic neuron clusters (Fig. 5D and E). As the SoxC>KikGR driver is transiently expressed during differentiation (Supplementary Fig. 14), intermediate cell states would be mostly orange or green cells, while terminally differentiated end states would be mostly red cells, with increasing green/orange fraction as newly generated cells accumulate (Fig. 5D, scenario 1). In contrast, if clusters are distinct terminally differentiated states without clear intermediate states among them, they would share a similar green+orange to red cells proportion (Fig. 5D, scenario 2). Thus, the ratio of green+orange to red cells across clusters within a cellular lineage can discriminate between intermediate versus terminal states. At 7 days post-photoconversion, longer than the peptidergic maturation timeline, we found a strong linear relationship between newly and previously generated neurons across Neural 1 peptidergic subtypes (R2 = 0.92, Pearson’s ⍴, Fig. 5E, scenario 2 from Fig. 5D). Although the number of sampled cells for the less abundant subtypes limits conclusive findings across all clusters, our analysis of the more abundant clusters supports peptidergic neurons as distinct terminal states, rather than intermediate states (Supplementary Fig. 14).
Overall, our data are consistent with closely related peptidergic subtypes being generated from the Piwi1+ multipotent population, with their relative ratio fixed at an early peptidergic-committed stage as supported by multiple independent lines of evidence. First, the proportional ratio of old to new (red cells to green+orange cells) SoxC>KikGR+ peptidergic neurons (scenario 2 in Fig. 5D and Supplementary Fig. 14A) is specific to these cells and not seen across digestive secretory cells, which display a separate proportional relationship (Supplementary Fig. 14). Then, if there is a shared peptidergic-committed lineage, as the FoxL2>KikGR, SoxC>KikGR and Elav>KikGR drivers label the Piwi1+ source population, and thus also its derivatives, then the relative abundance between peptidergic subtypes would be constrained by this shared lineage and would thus correlate across drivers, but not for cells generated through distinct cellular lineages. Indeed, we find that the relative abundance of peptidergic cells subtypes is correlated across drivers and that the relationship is specific to peptidergic neurons (excluding neural cluster 22, see below) (Fig. 5F and Supplementary Fig. 15); while FoxQ2d+ sensory cells, cnidocytes and secretory cells do not share the same relationship. This early peptidergic-specific lineage restriction is further independently supported by reanalysis of RNA-Seq data from a Prdm14d>GFP reporter line, which labels both Prdm14d+ neurons (Neural 0 in Fig. 3F), and uncharacterized progenitors at low levels14. Genes enriched in Prdm14d>GFP+ cells are expressed here in Prdm14d+ neurons, as well as across non-Prdm14d-expressing peptidergic subtypes; and not in secretory cells or other neurons (Supplementary Fig. 15A–C), suggesting that peptidergic-committed Piwi1+/Prdm14dlow progenitors are indeed restricted to peptidergic neural fates. Accordingly, genes enriched in Prdm14d>GFPneg cells are expressed across secretory cells and other neural cells including cluster 22 neurons, cnidocytes (developing and mature) and FoxQ2d+ sensory cells, further supporting that these cells are not part of the shared peptidergic lineage. And digestive secretory cells (excluding cluster 12) also display correlated abundance across drivers (Fig. 5F and Supplementary Fig. 15D and E), with a proportional ratio distinct from that of peptidergic neurons, further supporting a separate digestive-secretory-cell specific lineage.
Taken together, the data supports a model in which direct generation of distinct Elav+ peptidergic neuron subtypes from peptidergic-restricted Piwi1+/SoxC+/Elavlow cells underlies homeostatic nerve net scaling (Fig. 5G).
Two-step transcriptional maturation of cnidocytes
Sea anemone neural cells encompass peptidergic neurons, putative sensory cells, and cnidocytes, cnidarian-restricted mechano-sensory neural cells2,47. Having addressed the progression of peptidergic Elav+ neurons, from Elavlow/SoxC+/FoxL2+/Piwi1+ progenitors to differentiated cells, we examined whether neurons and cnidocytes share a similar differentiation trajectory. We find that cnidocytes, like peptidergic neurons, develop from transiently expressing SoxC+ cells (Supplementary Fig. 16). Indeed, only orange and green developing SoxC>KikGR+ cnidocytes are detected in situ and in the scRNA-Seq data at 2, 4, and 7 days post-photoconversion (Supplementary Fig. 16A, C, and D). Mature cnidocytes do not actively express SoxC (Fig. 3A, F and Supplementary Fig. 16E, F). Accordingly, we find red mature SoxC>KikGR+ cnidocytes at 2 days post-photoconversion, with newly generated orange and green mature cnidocytes appearing at 4 and 7 days, respectively (Supplementary Fig. 16C, D and Supplementary Movies 3–7). Thus, both peptidergic neurons and cnidocytes transiently express SoxC during early differentiation. Furthermore, while developing cnidocytes do not express FoxL2 (Fig. 3G and Supplementary Figs. 11, 16E, F), they arise from undetermined FoxL2>KikGR+ cells, as evidenced by the presence of red and orange developing cnidocytes at 2, 4, and 7 days post-photoconversion (Supplementary Fig. 16B and C). We also note that FoxL2>KIKGR is secondarily re-activated in mature cnidocytes, consistently with the presence of green mature cnidocytes at 4 and 7 days post-photoconversion alongside the majority of red and orange ones (Supplementary Fig. 16B, C and 8, 9). Thus, while the cellular origin(s) of cells that give rise to developing cnidocytes is unclear, with isolated developing cnidocytes present at the base of the epidermis layer in the body wall (Supplementary Fig. 16D), these cells also develop from FoxL2+ cells, and developing cnidocytes transiently express SoxC during their maturation (Supplementary Fig. 16E, F). We also note that a distinct subset of progenitor cells show specific expression of known cnidocyte regulators (cnidocytes “precursors” in Supplementary Fig. 12C), and could indicate that Piwi1+ progenitors also give rise to developing cnidocytes. However, while peptidergic neurons (Neural 0/1/2 in Fig. 3G) are recovered across all drivers (FoxL2>KikGR, Elav>KikGR, SoxC>KikGR), the late cnidocytes (Neural 4 class) are not recovered from Elav>KikGR+ sorted cells (Fig. 3G, also in ref. 20). Thus, SoxC is transiently expressed in developing cnidocytes and separately in precursors of peptidergic neurons, and our findings are incompatible with a shared SoxC+ neural-specific bipotent intermediate state (see also Supplementary Fig. 15).
We then investigated the transcriptional signatures of neural fate acquisition in peptidergic neurons and cnidocytes, two distinct cellular lineages (Fig. 6A). Cnidocytes mature through two sequential transcriptional phases (Supplementary Fig. 16E, F). Initially, during the developing cnidocyte stage, cnidarian-specific genes are used to build their specialized capsule organelle (intracellular harpoon-like structure). Subsequently, developing cnidocytes initiate a neural transcriptional program, which includes the expression of neural effector genes, such as the Eag (Ether-à-go-go family) voltage-gated K+ channel43 and a voltage-dependent L-type calcium channel subunit beta-2 (cacnb2)47. This biphasic cnidocyte effector program is characterized by a shift in transcription factor usage. Early in development, cnidocytes express PaxA, cnidoJun, NR12, ZNF845, and other zinc finger and basic leucine zipper (bZIP) transcription factors, which are associated with capsule formation38,39,48. In contrast, the secondary stage is marked by a distinct regulatory signature defined by expression of nk6, uncharacterized Sox and Myc genes, and additional bZIP genes (Supplementary Fig. 16F), indicative of the onset of the neural program. Thus, cnidocytes undergo two distinct transcriptional phases, with the sequential deployment of a cnidarian-specific capsule program, followed by a neural program. This two-phase transcriptional process is supported by reanalysis of RNA-Seq data from a PouIV knockout (KO) mutant49, in which immature cnidocytes accumulate. Thus, genes upregulated in PouIV KO juvenile polyps correspond to those expressed in developing cnidocytes, whereas PouIV KO downregulated genes overlap with those expressed in both developing and late cnidocytes (Supplementary Fig. 17). These findings support a biphasic transcriptional maturation and suggest that when the developing cnidocyte program is blocked, the secondary neural program is not initiated. Thus, this sequential maturation is a two-step process with a clear transcriptional intermediate program that is deployed, then cleared prior to the initiation of the neural program, revealing a different neurogenic strategy from that of peptidergic neurons.
A Neural cell differentiation in adults. Elav+ peptidergic neurons (top) and cnidocytes (bottom) acquire their mature neural identities through distinct cellular trajectories with an initial transient expression of SoxC. B Neural effector differential expression between Elav+ peptidergic clusters and late cnidocytes. Average expression log2 fold-change enrichment between the late cnidocyte cluster and the maximum expressing peptidergic cluster. The dashed line represents a 2-fold change in expression levels between the peptidergic cluster with the highest expression and the late cnidocyte cluster. Neuropeptide GPCRs and neuropeptide precursors from ref. 16. C WGCNA gene modules eigengene value across cell clusters. Gene modules (rows) are annotated as neural-specific, broadly neural or secretory-specific. Cell clusters (columns) colors as in Fig. 3G. D Transcription factor membership across WGCNA modules (same rows as in C) for bHLH, bZIP, homeodomain, zinc finger and other transcription factor families. Each transcription factor is a column, with low (semi-opaque, membership ≥ 0.35) or high (opaque, membership ≥ 0.5) module membership grades to highlight TF associated to multiple modules. Horizontal lines demarcate WGCNA modules associated with broad or restricted neural identities, mature cnidocytes, developing cnidocytes or secretory cells (respectively, bottom to top, as in C). E Gene modules top co-expression relationships with the gene module identifiers, same module colors as in (C). Broadly neural gene modules are in light blue. F Neural modules overview. Co-expression networks for broad neuronal and subtype- specific neural gene modules. (Above) Transcription factors in each module, colored by family. (Below) Effector genes colored as in B, with neural effectors expressed across both late cnidocytes and peptidergic neurons (in black in B) in white. Pan-peptidergic module 6, broadly neural module 5 and two neural- specific modules (7 and 8) are highlighted, with all neural-associated modules in Supplementary Fig. 18. Data is provided as Source Data File 6.
Neural cells arising from different lineages deploy distinct TF-effector gene modules
Since mature cnidocytes and peptidergic neurons acquire their neural identities through distinct processes, we further examined whether they converge on a shared neural transcriptional signature (Fig. 6A). We find that the expression profiles of neural effector genes are distinct (Fig. 6B). Neuropeptide precursors, processing enzymes and neuropeptide GPCRs are expressed in peptidergic neurons but not in late cnidocytes, suggesting that peptidergic transmission is not utilized by cnidocytes. Other neural-expressed GPCRs and ion channels are specific to either late cnidocytes or peptidergic neurons, with only a limited overlap between the two. Overall, these results demonstrate that mature cnidocytes and peptidergic neurons have separate neural effector expression profiles.
Despite the distinct differentiation trajectories and effector profiles of these neural cells, we examined whether they shared pan-neural transcription factors. Using weighted gene co-expression network analysis (WGCNA)50 to generate modules of genes whose expression is highly correlated, we identify 48 cell-type specific and broader class-specific modules across all cell types (Fig. 6C). Each neural identity combines a cell-type-specific module (modules 7–21 and 25) with shared neural module(s) (modules 3–6) (Fig. 6C and Supplementary Fig. 18). This is specific to neural cells, and contrasts with the lack of a shared pan-secretory module for the dozen molecularly distinct secretory subtypes. Among the shared neural modules, we first identify a pan-peptidergic module (Module 6) for peptidergic neural identity. This module encompasses pan-peptidergic effectors and transcription factors including a GATA factor, a ETS factor, and bHLH factors such as NvAshA/Nem17, an achaete-scute-related gene involved in embryonic peptidergic specification12,51 (Fig. 6D and Supplementary Fig. 18). Thus, we propose that this set of transcription factors underlies early peptidergic maturation, consistent with our model of Elav-positive peptidergic neurons arising from a shared cellular lineage. These genes are also specifically enriched in putative peptidergic precursors in the progenitor-specific subclustering (Supplementary Fig. 12C). Top co-expression relationships between modules further highlight this pan-peptidergic module 6 as a “hub” for peptidergic subtype-specific gene modules 7–21 (Fig. 6E). Secondly, we find a smaller pan-neural module partially shared between late cnidocytes and peptidergic neurons (Module 5). It includes few neural effectors and transcription factors, such as the SNAIL family C2H2 zinc finger transcriptional repressor Scratch, also used in vertebrate neural maturation52,53. While this module spans both late cnidocytes (neural phase) and peptidergic neurons, it is not utilized in the first stage of cnidocyte maturation (capsule phase) and thus does not correspond to a common lineage intermediate. Furthermore, as our early peptidergic neural fate specification model suggests, progenitor Module 1 includes the atonal-related bHLH transcription factor Nem10 (NvAth-like/NvArp3), an embryonic neurogenesis regulator6,8,12 (Fig. 6D). In contrast, the developing cnidocyte module (Module 26) encompasses all known embryonic regulators of early cnidocyte development (PaxA, the bZIP cnidoJun, NR12, and zinc finger factors like ZNF84538,39,48,49). These genes are also specifically enriched in putative cnidocyte precursors in the progenitor-specific subclustering (Supplementary Fig. 12C). While cnidocyte and peptidergic neural programs exhibit limited transcriptional convergence and rely on distinct maturation strategies, there are a few shared factors which suggest some pan-neural-shared regulation.
Overall, we find classes of transcription factors systematically associated with pan- (bHLH, bZIP classes) and subtype-specific (Homeodomain, zinc fingers classes) neural modules. bHLH transcription factors play proneural roles in bilaterians, and cnidarian orthologs of the achaete/scute and atonal/NeuroD6,8,12,51 families are associated here with shared pan-neural modules and broad peptidergic and cnidocyte neural fate, along with Myc family genes (Fig. 6D and Supplementary Fig. 18). Sets of bZIP factors are enriched in pan-peptidergic, pan-neural, developing, and mature cnidocyte modules (respectively, modules 6, 3–5, 26, and 25). In addition to class-specific neural bHLH and bZIP factors, each neural subtype is defined by a unique homeodomain and zinc finger code (Fig. 6D and Supplementary Fig. 19). Distinct homeodomain factors, with minimal overlap, specify peptidergic neurons, late cnidocytes, and sensory cells; a strategy exclusive to neural cells and absent in secretory cells. This homeodomain code is associated with subtype-specific effectors, including ion channels, neuropeptide precursors, and receptors (Fig. 6D and F). Developing and late cnidocytes also exhibit unique homeodomain and ZF transcription factors, with only PouIV, a BTB (Broad complex, Tramtrack, Bric-à-brac family), and an H2C2 ZF transcription factor shared between the two phases.
Together, these analyses highlight that adult neural identity integrates three transcriptional modules: (1) a pan-peptidergic module defining peptidergic neural identity, (2) a small, broadly neural module partially shared between late cnidocytes and peptidergic neurons, and (3) subtype-specific programs with a homeodomain code. Cnidocytes undergo a two-step differentiation process, each phase employing distinct transcription factors separate from those in peptidergic neurons. While peptidergic neurons and cnidocytes follow independent cellular trajectories and transcriptional programs to establish their mature neural identities, they utilize similar transcription factor classes—bZIP and bHLH factors act broadly within neural classes, whereas homeodomain and zinc finger factors are subtype-specific. These modular neural renewal strategies are unique to neural cells and are not employed in digestive secretory cells, despite their shared cellular mode of renewal with peptidergic neurons.
Discussion
A diversity of animals, including cnidarians, planarians and acoels, display indeterminate growth, necessitating lifelong (re)generation of multiple neuron types1,24,25. Here, we elucidate the cellular and transcriptional trajectories underlying adult neurogenesis in the sea anemone Nematostella vectensis. Using reporter tracing, nerve net imaging and single-cell transcriptomics, we show that active neurogenesis expands the nerve net along the primary body axis through two distinct pathways: (1) peptidergic neurons, the major neural cell class, arise directly from multipotent progenitors, with subtype identities specified in proportion to existing peptidergic populations and (2) cnidocytes undergo a two-stage maturation process characterized by a transcriptionally distinct intermediate phase, marked by phase-specific transcription factors. In the context of indeterminate growth, our observations show that neurogenesis occurs across the primary body axis, without a restricted growth zone, reminiscent of the distributed processes described in planarians and acoels1,25. The neurogenic rate uncovered—more than one new cell for two existing ones in the body per one-week period, across both peptidergic neurons and cnidocytes, is consistent with the growth rates observed in fed adults26. Importantly, the concomitant generation of neural subtypes in fixed proportions occurs independent of injury. This suggests that regenerative capacities may be rooted in continuous cell renewal rather than exclusively as a damage response. The occurrence of homeostatic neural renewal is tightly associated with extensive injury-induced regeneration and asexual reproduction abilities across metazoans1. The reoccurrence of the known peptidergic, cnidocyte, and pan-neural embryonic regulators in the corresponding adult gene modules further suggests that the molecular programs governing neural and cnidocyte specification may be deeply conserved across developmental stages.
We show the continuous differentiation of cells from the adult stem-like progenitor compartment31,54,55 into multiple cell types, including neural and secretory cells. Our results suggest potential self-renewal within a subset of the progenitor pool; however, this has not yet been directly demonstrated. Thus, whether some of these cells are bona fide stem cells and whether they might correspond to the dispersed slow-cycling cells reported in the literature46,55 remains unresolved. Such definitive identification of a self-renewing stem cell pool awaits future advances in genetic-lineage tracing in this species. The transcriptional signature of this progenitor pool resembles that of adult multi- to pluripotent stem cells found in planarians, acoels and in Hydra and Hydractinia29,30,56,57, pointing to shared molecular mechanisms of stem cell potency in animals. Notably, our study shows that the Elav>KikGR+ expression dynamics reveal that Piwi1+ progenitor cells, despite retaining undifferentiated transcriptional profiles, are already committed to differentiation. This indicates that the decision to differentiate (and the fate specification) occurs early, in cells with stem-cell-like molecular identity, rather than being limited to progenitors that have begun expressing cell-type-specific genes (putative precursors). This further suggests that fate decisions can be made alongside the cell cycle, as proposed for planarian neoblasts58. This contrasts with neural development in fixed-growth animals, where transit-amplifying intermediates, such as Drosophila neuroblasts, or mammalian apical radial glia, sequentially generate neurons during embryonic neurogenesis59. In Nematostella, direct neurogenesis may enable rapid neuron production to sustain indeterminate growth, with all body wall peptidergic neuron subtypes arising from undifferentiated progenitors per 4-day period. However, this could constrain molecular diversity, producing fewer neuron types compared to fixed-growth bilaterians.
Our study further uncovers lineage biases within the stem-like progenitor pool. Although all three drivers label the progenitor compartment, their resulting progeny show only partial overlap, with differential neural and secretory cell class recovery. This suggests two possible models for neural cell fate determination. One model proposes a shared “neuroglandular” lineage, where cells exiting the progenitor state are committed to peptidergic and digestive secretory cell fates36. Alternatively, a second model proposes that commitment to these fates occurs earlier within the progenitor compartment, with distinct cellular lineages forming upon exit. Our data, along with the reanalysis of a Prdm14d driver14, strongly support the latter scenario. This is consistent with insm1 reporter data60, expressed in progenitors, secretory cells and neurons in adults. Furthermore, the concurrent generation of peptidergic neurons and digestive secretory cells, in proportions that reflect their relative subtype abundance, suggests that subtype identity is pre-determined at exit from the progenitor state, consistent with the distinct putative peptidergic and secretory-specific progenitor subsets.
In contrast to the direct generation of peptidergic neurons, cnidocytes, the hallmark stinging cell of the phylum, undergo a distinct two-step transcriptional maturation. They transition from a cnidarian-specific capsule-forming program to a bilaterian-like neural program, each associated to distinct sets of transcription factors. The secondary neural phase of cnidocyte development shares some transcription factors with peptidergic neurons, suggesting limited transcriptional convergence across neural classes. Cnidocytes are conserved across all cnidarians41,61,62, but their renewal mechanisms vary. In hydrozoans, which diverged from anthozoans over 500 million years ago, such as the jellyfish Clytia hemisphaerica, they arise from stem cell niches in tentacle bulbs, while Hydra produces migratory nests of 4, 8, 16, and 32 cells from multipotent interstitial stem cells63,64. In contrast, the sea anemone Nematostella vectensis (an anthozoan) displays dispersed early-stage cnidocytes in the epidermis and mesenteries, even as it remains unclear whether they arise from the stem-like progenitor pool (also in refs. 32,56). This highlights the evolutionary plasticity of cnidocyte development, despite their conserved capsule genes. Thus, investigating whether orthologous transcription factors and neural effector genes are used in cnidocytes and peptidergic neurons across cnidarians could shed light on neural cell evolution and diversification, particularly considering their lineage-specific neural effector expansion (e.g., voltage-gated potassium channels, GPCRs16,65).
Although cnidocytes and peptidergic neurons use distinct renewal modes, their early development is marked by transient SoxC expression, followed by downregulation during maturation. This parallels vertebrate neurogenesis, where SoxC factors (SOX4, SOX11, and SOX12) play a critical role in neural progenitor differentiation66, underscoring the evolutionary conservation of neurodevelopmental pathways across eumetazoans, despite lineage-specific diversification. Overall, we propose a model where SoxC, SoxB(2), and distinct bHLH and bZIP transcription factors are associated with neural class specification, while homeodomain and zinc finger genes specify subtype identities. Intriguingly, the homeodomain code delineating neural identities in Nematostella is unique to neural cells and absent in digestive secretory cells. This is not clearly associated with axial patterning, as body wall neurons do not appear clearly spatially segregated along the primary (oral–aboral) or secondary (directive) body axes, across currently characterized populations14,15,18,35,49,67. Furthermore, the neural-subtype-specific homeodomain proteins are distinct from the ANTP-class Hox genes specifying the secondary (directive) body axis68,69, expressed in cnidarian-specific gastrodermal cells in adults. In bilaterians, homeodomain transcription factors are essential for primary body axis (antero-posterior) patterning (ANTP-class Hox genes), and act as terminal selectors of neural identity70,71,72,73. Thus, given the early diversification of homeodomain family genes before the cnidarian–bilaterian split, their neural subtype-specific expression in Nematostella suggests an ancestral role in neural subtype specification, separate from spatial patterning11,74. Supporting this hypothesis, the POU-family homeodomain gene PouIV/Brn3 is critical for cnidocyte maturation49,75, as its removal results in blocked immature cnidocytes that fail to initiate their secondary neural program. Systematic functional studies of homeodomain-class genes during adult neurogenesis would help shed light on this evolutionary link.
Our identification of the populations, lineages, and transcriptional programs underlying adult neural cell generation in a sea anemone reveals how indeterminate growth organisms scale their nervous system. We uncover two distinct modes of renewal across the main neural classes, shedding light on the strategies underlying lifelong neurogenesis. The shared molecular features with bilaterians point to deeply conserved neural specification mechanisms, tracing back over 540 million years to the Precambrian. These findings provide new insights into the molecular basis of adult neurogenesis and the evolutionary diversification of neurons across animal lineages.
Methods
Transgenic line generation
For the FoxL2>KikGR, SoxC>KikGR, and Elav>KikGR reporter line generation, plasmid constructs were generated from a pNvT-MHC::mCh vector backbone (a gift from Ulrich Technau76, Addgene plasmid #67943) with the KikGR fluorophore (from the pCAG::KikGR vector, a gift from Anna-Katerina Hadjantonakis, Addgene plasmid #3260833). KikGR reporter expression was driven by the published 2.4 kb upstream region of Elav from ref. 20 or the upstream regions of either SoxC (2.2 kb) or FoxL2 (2.4 kb), with primers in Supplementary Data 1. Stable transgenic animals were generated using I-SceI-mediated transgenesis76, with the following modified injection mixture: 50 ng/μL plasmid concentration, 1X I-SceI buffer (10 mM Tris–HCl, 10 mM MgCl2, 1 mM DTT, pH8.8), 0.5 mM Patent Blue VF (Sigma, #198218), 0.2 U/μL I-SceI (adjusted to 0.4 U/μL after 30 min at 37 °C). All experiments used stably integrated SoxC>KikGR, FoxL2>KikGR or Elav>KikGR 5–8-week-old heterozygote animals (sexually immature adults) raised at 18 °C in 12.5 ppt artificial seawater in the dark and fed 4 times a week with freshly hatched Artemia nauplii.
Whole animal photoconversion nerve net in situ imaging and quantification
Individual SoxC>KikGR, FoxL2>KikGR or Elav>KikGR immature adults were relaxed in 12.5 ppt artificial seawater and immobilized by a 1:1 dilution with 70 mg/mL MgCl2, then transferred between two coverslips, with tape layers to adjust depth to ca. 0.5 mm for photoconversion and live confocal imaging. Whole animal KikGR photoconversion was performed with a ZEISS Axio Imager 2 (ZEISS), with a X-Cite 120LED Boost (EXCELITAS) light source at 80% power, with manually tiled regions with 8 sec exposure (emission BP 445/50 nm) at ×20 magnification. Photoconversion efficiency was confirmed by imaging and flow cytometry (Supplementary Fig. 3B) following animal dissociation into single cells. Photoconverted animals were kept in 12.5 ppt artificial seawater, with either 1% DMSO or without. For confocal imaging of the body wall nerve net, individual SoxC>KikGR, FoxL2>KikGR, or Elav>KikGR adults at 2-, 4-, or 7-day post-KikGR photoconversion were immobilized (as above) and imaged at ×40 on a Zeiss LSM800 inverted confocal microscope, with 8–10 optical sections per z-stack with 1 µm between optical slices, with the enhanced green fluorescent protein (eGFP) spectrum used for detecting non-photoconverted KikGR and mCherry spectrum used for the photoconverted KikGR protein, adjusted to remove overlap. For quantification purposes, all images from the same reporter line at 2, 4, or 7 days post-photoconversion were acquired with the same settings. Figures 1B and 3A, C composite extended depth of focus and single slice images and Supplementary Movies 1–9 are for illustration only. All cell segmentations and quantifications were performed on raw z-stack data. For cell quantification, a cellpose 2.0 neural net model77,78 was manually trained on randomly sampled individual image slices from SoxC>KikGR, Elav>KikGR, and FoxL2>KikGR live body wall confocal z-stacks, then used to segment all raw z-stacks at all timepoints post-photoconversion separately on either the red (photoconverted KikGR signal) or green (new KikGR signal) channel for all drivers with the following parameters: diameter: 0, chan2: 0, cellprob_threshold: −0.5,flow_threshold: 0.8, stitch_threshold: 0.25 after optimization. The two sets of cell masks per image (red+ set and green+ set) were then merged into a single set for each z-stack for quantification. Representative videos of the z-stacks with the segmentation are provided in Supplementary Movies 1–9.
Single cell dissociation and KikGR+ cell sorting
For scRNA-seq sampling, individual SoxC>KikGR, FoxL2>KikGR or Elav>KikGR adults at 2-, 4-, or 7-day post-KikGR photoconversion (with initial experiments containing 1% DMSO added to their medium during the timecourse, and not in latter experiments, see Supplementary Data 2 and 3) were immobilized with MgCl2 as above and sampled below their pharynxes (body without pharynx region or tentacles) to be consistent with our body wall imaging data, then dissociated into single cells according to79, with 1–2 individuals per dissociation. In brief, the sample medium was replaced with calcium magnesium-free seawater (495 mM NaCl, 9.7 mM KCl, 27.6 mM NaHCO3, 50 mM Tris–HCl pH 8) with collagenases (0.05 mg/mL Liberase TM, Roche), and dissociated by gentle pipetting at room temperature. The cell suspension was filtered (Ø35 µm) and stained with live cell dye Calcein Violet AM (ThermoFisher Scientific, 1:2000 dilution of 1 µg/µL DMSO stock) or a combination of Calcein Violet and SytoxRed dead cell dye (Thermo Fisher Scientific, 1:1000 dilution of a DMSO 5 µM stock).
Individual live KikGR+ cells were sorted using a Fluorescence-Assisted Cell Sorter (FACS Aria IIu and III, BD Biosciences) into 384-well plates containing 2 µL lysis buffer and barcoded polyT capture oligonucleotides (MARS-Seq protocol4,80,81), then frozen at −80 °C until processing. FACS gating strategy is shown in Supplementary Figs. 3 and 4; in brief, to sort live KikGR+ cells, cell-sized particles were selected using their FSC-A (logicle-scale) vs. SSC-A (logicle-scale) signal (ca. ≥3 µm in diameter, using sizing beads (SpheroTech) as an absolute size reference). Multiplets were excluded with FSC-W and FSC-H, then Calcein Violet AM-positive live singlets were selected (ex.: 405 nm, em.: BP 450/40 nm). Initial sorts included a SytoxRed signal: APC channel (ex.: 633 nm, em.: 660/20 nm) and mildly depleted the Calcein Violetlow/SytoxRedmid population. From live singlets, KikGR+ cells were selected based on their new KikGR signal (FITC-A, ex.:488 nm, em. filters: LP 502 nm and BP 530 nm/30) and photoconverted KikGR (PE-A, ex.: 561 nm, em. Filters: BP 582/15 nm) signal, with the FITC-Alow/PE-Alow autofluorescent cells excluded, with a negative wild-type (no KikGR) control. A shared KikGR+ gate was used for all Elav>KikGR+ and SoxC>KikGR+ cells, and for 17/41 of FoxL2>KikGR+ libraries (Supplementary Fig. 3). For 16/41 FoxL2>KikGR+ libraries, we depleted their large epithelial cell fraction, as enabled by its specific photoconverted KikGR-positive/low new KikGR signal profile (“Epithelial-depleted KikGR+” sorting). To enrich for green and orange cells, we also sampled only New KikGRmid/high cells for 4 FoxL2>KikGR+ libraries (“New KikGR+” gate). All quantifications of reporter status (green, orange, and red cell proportions) from FACS-sorted cells are from cells sorted with the shared KikGR+ gate throughout the paper, with ternary plots (R package ggtern82) or barplots (R package ggplot2) visualizations, filtering out clusters with <10 sampled cells. For each KikGR+-sorted cell, cell reporter status (i.e. green, orange or red) for all drivers (Elav>KikGR, FoxL2>KikGR, SoxC>KikGR) was assigned using red, orange, or green gates defined across all sorted cells (Supplementary Fig. 4). Cell identifiers were corrected for two sorted 384-well plates (4 libraries: RP0885_E/O, RP0886_E/O), as the flow cytometer mechanical arm moved by one column prior to the sort. For each library sorted with the KikGR+ gate and used for color reporter analysis, matched flow cytometry data were validated through their correlation between FSC-A signal (a cell volume proxy) and total UMI per cell.
Single-cell RNA-Seq analysis
We generated 84 scRNA-Seq libraries according to published MARS-Seq protocols4,80,81. In brief, all plates were processed with the same protocol, and mRNA was reverse transcribed into cDNA with a barcoded oligodT capture oligonucleotide in the lysis buffer containing the cell barcode and unique molecular identifier. Exonuclease treatment (ExoI, NEB) was used to remove unused oligodT nucleotides. Pools of cDNA were then amplified by in vitro transcription (T7 polymerase), then fragmented. Each RNA fragment was ligated to an oligonucleotide containing a library pooling barcode with a T4 ssDNA:RNA ligase before reverse transcription. Final libraries were obtained from PCR amplification with 17 cycles. Library fragment distributions and concentrations were assessed by Tapestation 4150 (Agilent) and Qubit3.0 (Invitrogen). scRNA-Sseq libraries were pooled equimolarly and sequenced in paired-end mode on a NextSeq550 (Illumina) using a high-output 75-cycle kit (up to 32 libraries per run with 190 cells per half-plate library). Median sequencing depth per library was 13.4M reads (for 190 sorted cells). Fastq files were reformatted to the following read structure for compatibility with STAR Solo: Read1: mRNA fragment; Read2: 4 nucleotides (nt) library barcode, 7 nt well barcode (11nt total for the cell barcode), 8 nt UMI barcode. Reads were mapped and demultiplexed using STARSolo (2.7.10b), with the following parameters --soloType CB_UMI_Simple --soloUMIlen 8 --soloCBlen 11 --soloUMIstart 12 --soloCBstart 1 --soloCBmatchWLtype 1MM --soloUMIdedup Exact; with the cell barcode list for each sequencing run as the soloCBwhitelist. Paired-end fastq files per half-plate library are provided in the GEO archive (GSE288441), along with cell barcode lists for each pool of demultiplexing-compatible libraries.
Reads were mapped to the jaNemVect1.1 Nematostella vectensis genome assembly (from the Darwin Tree of Life (DToL) project—Wellcome Open Research83), with the KikGR sequence added33, using the gene models from the RefSeq Nematostella vectensis Annotation Release 101 from NCBI (GCF_932526225.1, NCBI) (in GEO archive GSE288441). Of the 84 libraries, from 27 independent biological dissociations (see Supplementary Fig. 10A, C, E, and G), a total of 19, 24, and 41 libraries correspond to the Elav>KikGR, SoxC>KikGR, and FoxL2>KikGR drivers, respectively, with an average of 2131 median RNA molecules recovered per library. We removed the 164/16,012 control empty wells, and the remaining 15,848 cells were filtered based on RNA molecules recovery (excluding cells with ≤300 or ≥13,000 recovered RNA molecules, which removed 468/15,848 (2.9%) low-quality cells), and low quality clusters (lower UMIs, no specific marker gene, a large fraction of stress-related genes and ribosomal proteins) were filtered out, as well as clusters with cells from one biological dissociation, only from DMSO+ samples or from a single sequencing run for a total of 3030/15,848 (19%) filtered out cells. After filtering, 12,352/15,848 (78%) cells were retained. Filtered and unfiltered UMI tables are available in the GEO archive (GSE288441). Libraries’ descriptions with biological samples, experimental conditions, and gene recovery statistics are in Supplementary Data 2. We then used Seurat 5.2.1 with SCTransform v2 (sctransform 0.4.1) normalization for clustering, using the top 4000 variable genes (excluding the KikGR gene)84,85, with the top 65 PCA components used and a 2.4 clustering resolution. Clusters were annotated based on literature-curated markers (Supplementary Data 15,14,16,18,31,35,38,40,41,42,42,44,47,49,49,51,86,89). Elav>KikGR+ cells (Fig. 2 and Supplementary Figs. 5 and 11), SoxC>KikGR+ (Supplementary Fig. 11), and FoxL2>KikGR+ (Supplementary Fig. 11) cells were also individually clustered as described above, with a resolution of 2, 2.5, and 2, respectively.
Due to the multiple genome assemblies and gene models used in the literature, gene annotations were transferred to the GCF_932526225.1 gene models (jaNemVect1.1 genome assembly, gene identifiers starting with LOC) using either BLASTP of protein sequences, primerBLAST of in situ hybridization probes from the original publications or best reciprocal BLASTP hits; as listed in Supplementary Data 1, with the GCF_932526225.1 gene annotation used by default for all other genes. Transcription factors were predicted with AnimalTFDB87, with bZIP transcription factors named from their families from ref. 87, and bHLH transcription factors annotated from refs. 12,52. Cell cycle module genes (22 G2/M and 29 S-phase genes) were identified in Nematostella by mapping Seurat’s mouse cell cycle markers84 using eggNOG mapper annotations. The distribution of cell identities across libraries was consistent across biological samples for each driver and fluorescence-activated cell sorting (FACS) gating strategy (Supplementary Fig. 10). Gene expression across clusters is visualized using DotPlot from SCTransform data for the global clustering and from the RNA data for the line-specific reclustering, with expression values normalized to the maximum expression observed across clusters, and dot size representing the percentage of cells expressing the gene in each cluster.
To investigate differential neural effector gene usage between peptidergic neuron clusters (18 clusters) and late cnidocytes (1 cluster), we annotated genes coding for Class A GPCRs, mGluR-like receptors, and various ion channels based on PFAM domains using eggNOG mapper v2. The PFAM domains used included 7tm_1 and 7tm_3 for GPCRs (respectively, Class A and mGluR-like), as well as ASC, Anoctamin, NT_gated_channel, iGluR, PKD_channel, Lig_chan-Glu_bd, and Neur_chan_LBD for ion channels. De-orphanized neuropeptide GPCR receptors were further annotated from ref. 16. To assess the differential usage of neural-specific genes, we excluded lowly expressed genes (defined as those with fewer than 50 UMI across neural clusters) and genes whose highest expression occurred in non-neural clusters. Given the presence of multiple peptidergic clusters but only a single late cnidocyte cluster, we systematically compared the late cnidocyte cluster to the peptidergic cluster with the highest expression of each effector gene. The expression matrix was aggregated using Seurat’s AggregateExpression function, with a scale factor of 1000. We calculated the log2-fold change in expression between the late cnidocytes and the peptidergic cluster with maximal effector expression, plotting this against the average expression between the two clusters for 499 genes across six categories (Source Data File 6).
For the cnidocytes developmental trajectory inference, cnidocytes (clusters 6 and 14) were subsetted from the global scRNA-seq clustering and processed in Seurat with standard normalization to generate a UMAP embedding. Pseudotime analysis was performed using slingshot88 (2.12.0) on the UMAP coordinates. Gene expression along pseudotime was then estimated using a generalized additive model with the gam function from the mgcv R package (1.9-1).
For the progenitor-specific reclustering, progenitors were subsetted from the global scRNA-seq clustering and processed in Seurat with standard normalization, using 30 PCs and a resolution of 1.5, and annotated based on the expression of progenitor-enriched genes and cell-type-specific marker genes.
Weighted gene co-expression network analysis (as implemented in the WGCNA R package, version 1.73) was used to identify 48 transcriptional modules in the global clustering50. Gene expression data from the Seurat object were filtered to retain genes with at least 50 UMIs, and aggregated expression was computed per cluster with log normalization and a scale factor of 1000. A weighted gene co-expression network was then constructed using a soft-threshold power of 6. The adjacency matrix was transformed into a topological overlap matrix (TOM), and hierarchical clustering was performed. Modules were identified via dynamic tree cutting (deepSplit = 3, minClusterSize = 20), then merged based on eigengene correlations, using a dissimilarity threshold of 0.25. Module eigengenes were computed, and gene-module membership was determined by correlating gene expression with module eigengenes across clusters. A low (0.35) and a high (0.5) threshold were used to visualize soft module membership for transcription factors. Gene module expression across clusters (Supplementary Fig. 18) was computed using Seurat’s AddModuleScore function with each module composed of genes with module membership ≥ 0.5 (stringent threshold), and each gene only associated with its top-scoring module (Source Data File 6). These thresholded gene modules were then used to visualize the network of module top co-expression relationships, derived from the average pairwise TOM values, by filtering out module-module average scores below 0.002.
Reanalysis of RNA-Seq data
Upregulated and downregulated genes from the PouIV-KO and Prdm14d>GFP+ RNA-Seq datasets, as reported in the original publications14,49, were converted to GCF_932526225.1 gene identifiers using best reciprocal BLASTp hits (Supplementary Data 1). Gene set enrichment analysis was then performed using the GSEA R package (fgsea 1.30.0), with cluster gene markers from the FindAllMarkers in Seurat function on the global clustering (with an adjusted p-value threshold of 0.01) used for the ‘pathways’ lists (26–1265 genes per cluster-associated set) (Source Data File 7).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The SoxC>KikGR, Elav>KikGR, and FoxL2>KikGR transgenic Nematostella vectensis animals and associated plasmids are available upon request to the corresponding author. Sequencing data is available in GEO (accession: GSE288441), with paired-end fastq files for each sequenced library, alongside the reference genome and associated gtf used for mapping (jaNemVect1.1 Nematostella vectensis genome assembly with the KikGR sequence added), cell metadata information, and raw and filtered global count tables. Source data are provided with this paper as Source Data Files, as well as Supplementary Movies of the nerve net imaging. Source data are provided with this paper.
Code availability
Data analysis code (R version 4.4.1) and processed annotated Seurat sc RNA-Seq objects are available on Zenodo ([https://doi.org/10.5281/zenodo.14977132]).
References
Srivastava, M. Beyond casual resemblance: rigorous frameworks for comparing regeneration across species. Annu. Rev. Cell Dev. Biol. 37, 415–440 (2021).
Rentzsch, F., Juliano, C. & Galliot, B. Modern genomic tools reveal the structural and cellular diversity of cnidarian nervous systems. Curr. Opin. Neurobiol. 56, 87–96 (2019).
Bosch, T. C. G. et al. Back to the basics: cnidarians start to fire. Trends Neurosci. 40, 92–105 (2017).
Sebé-Pedrós, A. et al. Cnidarian cell type diversity and regulation revealed by whole-organism single-cell RNA-Seq. Cell 173, 1520–1534.e20 (2018).
Richards, G. S. & Rentzsch, F. Transgenic analysis of a SoxB gene reveals neural progenitor cells in the cnidarian Nematostella vectensis. Development 141, 4681–4689 (2014).
Richards, G. S. & Rentzsch, F. Regulation of Nematostella neural progenitors by SoxB, Notch and bHLH genes. Development 142, 3332–3342 (2015).
Levy, S. et al. A stony coral cell atlas illuminates the molecular and cellular basis of coral symbiosis, calcification, and immunity. Cell 184, 2973–2987.e18 (2021).
Watanabe, H. et al. Sequential actions of β-catenin and Bmp pattern the oral nerve net in Nematostella vectensis. Nat. Commun. 5, 5536 (2014).
Ryan, T. J. & Grant, S. G. N. The origin and evolution of synapses. Nat. Rev. Neurosci. 10, 701–712 (2009).
Flici, H. et al. An evolutionarily conserved SoxB-Hdac2 crosstalk regulates neurogenesis in a cnidarian. Cell Rep. 18, 1395–1409 (2017).
Larroux, C. et al. Genesis and expansion of metazoan transcription factor gene classes. Mol. Biol. Evol. 25, 980–996 (2008).
Simionato, E. et al. Origin and diversification of the basic helix-loop-helix gene family in metazoans: insights from comparative genomics. BMC Evol. Biol. 7, 33 (2007).
Magie, C. R., Pang, K. & Martindale, M. Q. Genomic inventory and expression of Sox and Fox genes in the cnidarian Nematostella vectensis. Dev. Genes Evol. 215, 618–630 (2005).
Lemaître, Q. I. B. et al. NvPrdm14d-expressing neural progenitor cells contribute to non-ectodermal neurogenesis in Nematostella vectensis. Nat. Commun. 14, 4854 (2023).
Havrilak, J. A. et al. Characterization of NvLWamide-like neurons reveals stereotypy in Nematostella nerve net development. Dev. Biol. 431, 336–346 (2017).
Thiel, D. et al. Large-scale deorphanization of Nematostella vectensis neuropeptide GPCRs supports the independent expansion of bilaterian and cnidarian peptidergic systems. eLife 12, RP90674 (2024).
Takahashi, T. Comparative aspects of structure and function of cnidarian neuropeptides. Front. Endocrinol. 11, 339 (2020).
Sachkova, M. Y. et al. Toxin-like neuropeptides in the sea anemone Nematostella unravel recruitment from the nervous system to venom. Proc. Natl. Acad. Sci. USA 117, 27481–27492 (2020).
Hauser, F., Koch, T. L. & Grimmelikhuijzen, C. J. P. Review: the evolution of peptidergic signaling in Cnidaria and Placozoa, including a comparison with Bilateria. Front. Endocrinol. 13, 973862 (2022).
Nakanishi, N., Renfer, E., Technau, U. & Rentzsch, F. Nervous systems of the sea anemone Nematostella vectensis are generated by ectoderm and endoderm and shaped by distinct mechanisms. Development 139, 347–357 (2012).
Levy, S. et al. Ectopic activation of GABAB receptors inhibits neurogenesis and metamorphosis in the cnidarian Nematostella vectensis. Nat. Ecol. Evol. 5, 111–121 (2021).
Ramos-Vicente, D. et al. Metazoan evolution of glutamate receptors reveals unreported phylogenetic groups and divergent lineage-specific events. eLife 7, e35774 (2018).
Goulty, M., Botton-Amiot, G., Rosato, E., Sprecher, S. G. & Feuda, R. The monoaminergic system is a bilaterian innovation. Nat. Commun. 14, 3284 (2023).
Sebens, K. P. The ecology of indeterminate growth in animals. Annu. Rev. Ecol. Evol. Syst. 18, 371–407 (1987).
Baguñà, J. et al. Growth, degrowth and regeneration as developmental phenomena in adult freshwater planarians. In Experimental Embryology in Aquatic Plants and Animals (ed. Marthy, H.-J.) 129–162 (Springer US, 1990).
Garschall, K. et al. The cellular basis of feeding-dependent body size plasticity in sea anemones. Development 151, dev202926 (2024).
Röttinger, E. Nematostella vectensis, an emerging model for deciphering the molecular and cellular mechanisms underlying whole-body regeneration. Cells 10, 2692 (2021).
Labusch, M., Mancini, L., Morizet, D. & Bally-Cuif, L. Conserved and divergent features of adult neurogenesis in zebrafish. Front. Cell Dev. Biol. 8, 525 (2020).
Wyss, L. S., Bray, S. R. & Wang, B. Cellular diversity and developmental hierarchy in the planarian nervous system. Curr. Opin. Genet. Dev. 76, 101960 (2022).
van Wolfswinkel, J. C. Piwi and potency: PIWI proteins in animal stem cells and regeneration. Integr. Comp. Biol. 54, 700–713 (2014).
Miramón-Puértolas, P., Pascual-Carreras, E. & Steinmetz, P. R. H. A population of Vasa2 and Piwi1 expressing cells generates germ cells and neurons in a sea anemone. Nat. Commun. 15, 8765 (2024).
Tritschler, S. et al. Concepts and limitations for learning developmental trajectories from single cell genomics. Development 146, dev170506 (2019).
Nowotschin, S. & Hadjantonakis, A.-K. Use of KikGR a photoconvertible green-to-red fluorescent protein for cell labeling and lineage analysis in ES cells and mouse embryos. BMC Dev. Biol. 9, 49 (2009).
Fritzsche, S. & Hunnekuhl, V. S. Cell-specific expression and individual function of prohormone convertase PC1/3 in Tribolium larval growth highlights major evolutionary changes between beetle and fly neuroendocrine systems. EvoDevo 12, 9 (2021).
Busengdal, H. & Rentzsch, F. Unipotent progenitors contribute to the generation of sensory cell types in the nervous system of the cnidarian Nematostella vectensis. Dev. Biol. 431, 59–68 (2017).
Steger, J. et al. Single-cell transcriptomics identifies conserved regulators of neuroglandular lineages. Cell Rep. 40, 111370 (2022).
Fritz, A. E., Ikmi, A., Seidel, C., Paulson, A. & Gibson, M. C. Mechanisms of tentacle morphogenesis in the sea anemone Nematostella vectensis. Development 140, 2212–2223 (2013).
Babonis, L. S. & Martindale, M. Q. PaxA, but not PaxC, is required for cnidocyte development in the sea anemone Nematostella vectensis. EvoDevo 8, 14 (2017).
Babonis, L. S., Enjolras, C., Ryan, J. F. & Martindale, M. Q. A novel regulatory gene promotes novel cell fate by suppressing ancestral fate in the sea anemone Nematostella vectensis. Proc. Natl. Acad. Sci. USA 119, e2113701119 (2022).
Steinmetz, P. R. H., Aman, A., Kraus, J. E. M. & Technau, U. Gut-like ectodermal tissue in a sea anemone challenges germ layer homology. Nat. Ecol. Evol. 1, 1535–1542 (2017).
Zenkert, C., Takahashi, T., Diesner, M.-O. & Özbek, S. Morphological and molecular analysis of the Nematostella vectensis cnidom. PLoS ONE 6, e22725 (2011).
Moran, Y. et al. Analysis of Soluble Protein Contents from the Nematocysts of a Model Sea Anemone Sheds Light on Venom Evolution. Mar. Biotechnol. 15, 329–339 (2013).
Li, X. et al. Ether-à-go-go family voltage-gated K+ channels evolved in an ancestral metazoan and functionally diversified in a cnidarian–bilaterian ancestor. J. Exp. Biol. 218, 526–536 (2015).
Cole, A. G. et al. Muscle cell-type diversification is driven by bHLH transcription factor expansion and extensive effector gene duplications. Nat. Commun. 14, 1747 (2023).
Amiel, A. R. et al. Characterization of morphological and cellular events underlying oral regeneration in the sea anemone, Nematostella vectensis. Int. J. Mol. Sci. 16, 28449–28471 (2015).
Amiel, A. R., Foucher, K., Ferreira, S. & Röttinger, E. Synergic coordination of stem cells is required to induce a regenerative response in anthozoan cnidarians. Preprint at bioRxiv https://doi.org/10.1101/2019.12.31.891804 (2019).
Thorington, G. U. & Hessinger, D. A. Activated L-type calcium channels inhibit chemosensitized nematocyst discharge from sea anemone tentacles. Biol. Bull. 238, 180–192 (2020).
Sunagar, K. et al. Cell type-specific expression profiling unravels the development and evolution of stinging cells in sea anemone. BMC Biol. 16, 108 (2018).
Tournière, O. et al. NvPOU4/Brain3 functions as a terminal selector gene in the nervous system of the cnidarian Nematostella vectensis. Cell Rep. 30, 4473–4489.e5 (2020).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Layden, M. J., Boekhout, M. & Martindale, M. Q. Nematostella vectensis achaete-scute homolog NvashA regulates embryonic ectodermal neurogenesis and represents an ancient component of the metazoan neural specification pathway. Development 139, 1013–1022 (2012).
Paul, V. et al. Scratch2 modulates neurogenesis and cell migration through antagonism of bHLH proteins in the developing neocortex. Cereb. Cortex (New York, NY, 1991) 24, 754–772 (2014).
Dattoli, A. A. et al. Domain analysis of the Nematostella vectensis SNAIL ortholog reveals unique nucleolar localization that depends on the zinc-finger domains. Sci. Rep. 5, 12147 (2015).
Talice, S. et al. Candidate stem cell isolation and transplantation in Hexacorallia. Cell Rep. 43, 114944 (2024).
Denner, A. et al. Nanos2 marks precursors of somatic lineages and is required for germline formation in the sea anemone Nematostella vectensis. Sci. Adv. 10, eado0424 (2024).
Varley, Á, Horkan, H. R., McMahon, E. T., Krasovec, G. & Frank, U. Pluripotent, germ cell competent adult stem cells underlie cnidarian regenerative ability and clonal growth. Curr. Biol. 33, 1883–1892.e3 (2023).
Hobmayer, B. et al. Stemness in Hydra—a current perspective. Int. J. Dev. Biol. 56, 509–517 (2012).
Raz, A. A., Wurtzel, O. & Reddien, P. W. Planarian stem cells specify fate yet retain potency during the cell cycle. Cell Stem Cell 28, 1307–1322.e5 (2021).
Holguera, I. & Desplan, C. Neuronal specification in space and time. Science 362, 176–180 (2018).
Tournière, O., Gahan, J. M., Busengdal, H., Bartsch, N. & Rentzsch, F. Insm1-expressing neurons and secretory cells develop from a common pool of progenitors in the sea anemone Nematostella vectensis. Sci. Adv. 8, eabi7109 (2022).
Engel, U. et al. A switch in disulfide linkage during minicollagen assembly in Hydra nematocysts. EMBO J. 20, 3063–3073 (2001).
Tursch, A., Mercadante, D., Tennigkeit, J., Gräter, F. & Özbek, S. Minicollagen cysteine-rich domains encode distinct modes of polymerization to form stable nematocyst capsules. Sci. Rep. 6, 25709 (2016).
Denker, E., Manuel, M., Leclère, L., Le Guyader, H. & Rabet, N. Ordered progression of nematogenesis from stem cells through differentiation stages in the tentacle bulb of Clytia hemisphaerica (Hydrozoa, Cnidaria). Dev. Biol. 315, 99–113 (2008).
David, C. N. & Gierer, A. Cell cycle kinetics and development of Hydra attenuata: III. Nerve and nematocyte differentiation. J. Cell Sci. 16, 359–375 (1974).
Lara, A., Simonson, B. T., Ryan, J. F. & Jegla, T. Genome-scale analysis reveals extensive diversification of voltage-gated K+ channels in stem cnidarians. Genome Biol. Evol. 15, evad009 (2023).
Kavyanifar, A., Turan, S. & Lie, D. C. SoxC transcription factors: multifunctional regulators of neurodevelopment. Cell Tissue Res. 371, 91–103 (2018).
Havrilak, J. A., Al-Shaer, L., Baban, N., Akinci, N. & Layden, M. J. Characterization of the dynamics and variability of neuronal subtype responses during growth, degrowth, and regeneration of Nematostella vectensis. BMC Biol. 19, 104 (2021).
Finnerty, J. R. Cnidarians reveal intermediate stages in the evolution of Hox clusters and axial complexity1. Am. Zool. 41, 608–620 (2001).
DuBuc, T. Q., Stephenson, T. B., Rock, A. Q. & Martindale, M. Q. Hox and Wnt pattern the primary body axis of an anthozoan cnidarian before gastrulation. Nat. Commun. 9, 2007 (2018).
Reilly, M. B., Cros, C., Varol, E., Yemini, E. & Hobert, O. Unique homeobox codes delineate all the neuron classes of C. elegans. Nature 584, 595–601 (2020).
Smith, J. J. et al. A molecular atlas of adult C. elegans motor neurons reveals ancient diversity delineated by conserved transcription factor codes. Cell Rep. 43, 113857 (2024).
Hobert, O. Homeobox genes and the specification of neuronal identity. Nat. Rev. Neurosci. 22, 627–636 (2021).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Ryan, J. F. et al. The cnidarian-bilaterian ancestor possessed at least 56 homeoboxes: evidence from the starlet sea anemone, Nematostella vectensis. Genome Biol. 7, R64 (2006).
Gahan, J. M., Kouzel, I. U., Jansen, K. O., Burkhardt, P. & Rentzsch, F. Histone demethylase Lsd1 is required for the differentiation of neural cells in Nematostella vectensis. Nat. Commun. 13, 465 (2022).
Renfer, E., Amon-Hassenzahl, A., Steinmetz, P. R. H. & Technau, U. A muscle-specific transgenic reporter line of the sea anemone, Nematostella vectensis. Proc. Natl. Acad. Sci. USA 107, 104–108 (2010).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Pachitariu, M. & Stringer, C. Cellpose 2.0: how to train your own model. Nat. Methods 19, 1634–1641 (2022).
Plessier, F., Schmutz, S., Novault, S. & Marlow, H. Single-cell transcriptomic analysis in the regenerating cnidarian Nematostella vectensis. Methods Mol. Biol. 2450, 565–581 (2022).
Jaitin, D. A. et al. Massively parallel single cell RNA-Seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Keren-Shaul, H. et al. MARS-seq2.0: an experimental and analytical pipeline for indexed sorting combined with single-cell RNA sequencing. Nat. Protoc. 14, 1841–1862 (2019).
Hamilton, N. E. & Ferry, M. ggtern: ternary diagrams using ggplot2. J. Stat. Softw. 87, 1–17 (2018).
Fletcher, C. & da Conceicoa, L. P. The Genome Sequence of the Starlet Sea Anemone, Nematostella vectensis (Stephenson, 1935) (Wellcome Open Research, Open Access Publishing Platform, 2023).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Reinke, A. W., Baek, J., Ashenberg, O. & Keating, A. E. Networks of bZIP protein–protein interactions diversified over a billion years of evolution. Science 340, 730–734 (2013).
Shen, W.-K. et al. AnimalTFDB 4.0: a comprehensive animal transcription factor database updated with variation and expression annotations. Nucleic Acids Res. 51, D39–D45 (2022).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genom. 19, 477 (2018).
Columbus-Shenkar, Y. Y. et al. Dynamics of venom composition across a complex life cycle. eLife 7, e35014 (2018).
Acknowledgements
We thank Sébastien Bastide for data analysis discussions and comments on the manuscript. We thank Caroline Kaiser, Ilaria Rebay, Paschalis Kratsios, and Urs Schmidt-Ott, and members of the Marlow lab, for discussions and comments on the manuscript. We thank Colette Plessier for the Nematostella silhouettes. We thank Baptiste Saudemont for MARS-Seq training, Sophie Novault and Sandrine Schmutz for FACS training, and expert advice. We thank the Flow Cytometry Core Facility in Institut Pasteur (Paris, France) for BD FACS AriaIII access. We thank David Leclerc, Mike Olson, and Bert Ladd of the University of Chicago Cytometry and Antibody Technology Facility for BD FACS AriaII training and advice (Facility RRID: SCR_017760). We thank Marie-Pierre Mailhé for assistance with wild-type animal care. We thank the Developmental and Stem Cell Biology Department of Institut Pasteur (Paris, France) for Zeiss LSM800 confocal microscope access. We thank Matt Kaufman for image processing advice. We thank the University of Chicago’s Research Computing Center for HPC access for image processing. We thank the École Normale Supérieure de Lyon and the Ministère de l’Enseignement Supérieur et de la Recherche (France) for funding (to F.P.). This study was funded in part by NIH NIGMS grant 5R35GM155159 to H.M.
Author information
Authors and Affiliations
Contributions
H.M. conceived the study. H.M. and F.P. designed experiments and interpreted the data. F.P. performed experiments, developed and implemented data analyses, generated figures, and wrote the initial draft. H.M. and F.P. wrote and edited the manuscript. H.M. supervised the project and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks José Martín-Durán and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Plessier, F., Marlow, H. Cellular and transcriptional trajectories of neural fate specification in sea anemone uncover two modes of adult neurogenesis. Nat Commun 17, 2611 (2026). https://doi.org/10.1038/s41467-026-68802-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68802-9