Abstract
Heterotopic ossification (HO), the abnormal formation of bone in soft tissues, is a debilitating complication that occurs after severe burns, trauma, and joint replacement surgery. Current diagnostic methods detect HO only after substantial progression, limiting opportunities for early intervention. In this study, we show that circulating mesenchymal progenitor cells (cMPCs), isolated using a microfluidic iChip from patients undergoing hip replacement and a mouse model of traumatic HO, exhibit HO-associated gene expression as early as 6 hours post-injury—41 days before radiographic detection. RNA sequencing of cMPCs enabled development of a liquid biopsy-based HO risk prediction model, achieving up to 90% sensitivity and 100% specificity in human subjects. Furthermore, the model detected significant reductions in HO risk following prophylactic treatment, correlating with decreased HO volume. These findings establish a noninvasive platform for early detection and monitoring of HO and suggest broader utility for diseases involving aberrant mesenchymal cell fate. This approach enables high-throughput screening of at-risk patients and real-time assessment of therapeutic efficacy.
Similar content being viewed by others
Introduction
Heterotopic ossification (HO), the abnormal growth of bone in soft tissues, is a common yet debilitating condition. Although genetic forms of HO remain rare, the incidence of trauma-induced HO is common. In adults with traumatic tendon injuries, fractures and sports injuries, the incidence of HO ranges from ~10–20%, whereas the incidence for HO ranges from 60–70% for blast injuries, 20–80% for hip replacements and more than 60% for major burns1,2,3,4,5,6. HO severely diminishes a patient’s quality of life due to pain, limited mobility, nerve damage, and poor wound healing. The gold-standard techniques for HO diagnosis include X-ray and computed tomography (CT); however, these technologies only detect mature bone 6–8 weeks after injury, when HO prevention or reversal is no longer possible. Currently, there are no technologies that can diagnose HO prior to irreversible symptoms such as joint contracture and pain.
While prophylactic strategies have been the focus of many clinical studies, the only currently approved modalities are radiation therapy and non-steroidal anti-inflammatory medications. However, these treatments are non-specific and have off-target effects, including fracture non-union, tissue fibrosis, malignancy, and wound healing complications7,8, making the need for precision medicine (only treating those at risk) a high priority. Additionally, the timing of initiation and termination of these prophylactic strategies remains unknown, and it is necessary to optimize treatment duration. Thus, a large percentage of patients (up to 80%) who will not go on to form HO end up receiving these potentially harmful prophylactic interventions. In addition to those treated unnecessarily, many patients are also treated at the wrong time and for the incorrect duration, given a lack of treatment guiding technologies. Thus, a point-of-care decision support tool to predict which patients will develop HO and assess the impact of treatment efficacy will provide a more “precision medicine” approach.
Rare cells in circulation have been shown as potential precursors of neoplastic disease and allow early detection and monitoring of cancer as a “liquid biopsy” approach. Circulating mesenchymal/skeletal cells also have been found in patients following trauma9,10,11. Blood samples from people living with a rare genetic form of HO, fibrodysplasia ossificans progressiva (FOP), have been shown to have increased circulatory osteogenic precursors during active episodes of HO formation compared to stable disease. Indeed, previous studies detected HO-associated circulating mesenchymal progenitor cells (cMPCs) following an HO-inducing burn/tenotomy (B/T) in our proven mouse model12. While several studies suggest that circulating mesenchymal cell types contribute to HO formation, their contribution is likely limited13.
In this work, we investigate if cMPCs can be used as an early marker of trauma-induced HO. First, we collect and analyze blood samples from mice after trauma with and without HO to determine differences in cMPC abundance. Next, we identify an HO-specific cMPC-based molecular signature using blood samples from hip replacement patients and a mouse model of HO. Then, using machine learning, we train a predictive model achieving up to 100% specificity and 90% sensitivity in identifying early-stage HOpos samples. These findings provide compelling evidence supporting the feasibility of a liquid biopsy approach in early detection and surveillance of traumatic HO. Finally, to assess therapeutic efficacy, we treat mice after trauma with a proven inhibitor of HO. Measuring the molecular phenotype and concentration of cMPCs – a “liquid biopsy” – can serve as a sensitive, minimally invasive diagnostic marker, or “footprint” of HO, enabling screening of high-risk patients and evaluating therapeutic efficacy to more precisely guide early treatment. Just as liquid biopsies have emerged as critical tools to diagnose and monitor certain malignancies14,15,16,17,18,19,20,21,22, we propose that this technology holds great promise for musculoskeletal conditions involving aberrant mesenchymal cell fate and repair, such as HO.
Results
HO-inducing injury leads to the early release of PDGFRa+ cMPCs
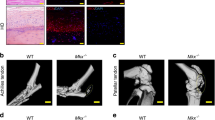
To investigate if peripheral blood contains cMPCs that can be used as biomarkers to enable early detection of HO, we performed either 1) a combination of 30% total body surface area (TBSA) burn injury followed by transection at the midpoint dorsal hind limb tendon (Achilles tenotomy) which results in HO (HOpos) or 2) a combination of 30% TBSA burn and a skin incision (B/S) which does not cause HO (HOneg) according to protocols previously published by our collaborative team and collected blood samples at days 1, 3 or 7 after injury12 (Fig. 1a). The injuries were performed at UT Southwestern and samples were shipped to Massachusetts General Hospital utilizing normothermic stabilization of whole blood for cMPC isolation using our established protocols23. We then isolated live cMPC populations using iChip and quantified CD45- cells expressing PDGFRa and CD90, a marker combination containing MPCs24, using flow cytometry (Fig. 1b, Supplementary Fig. 1).
a Experimental design to enrich cMPCs. Whole blood is drawn from mice with burn/tenotomy (B/T) and burn/sham injury controls (B/S), incubated with antibodies targeting leukocytes and magnetic beads; blood is then processed via iChip to enrich cMPCs. Created in BioRender. Karabacak, M. (2026) https://BioRender.com/u7f26d4. b Flow cytometry gating to analyze the marker combination CD45-PDGFRa+CD90+ that contains cMPCs. c CD45-PDGFRa+CD90+ cell numbers following B/T are significantly increased compared to B/S days 1 and 3 post-injury (Day1: n = 4 B/S, n = 9 B/T; Day 3: n = 4 B/S, n = 9 B/T, data are presented as mean values + /-SD). d We repeated the B/T model in transgenic Gli1- and Adipoq-lineage tracing mice and quantified tdTomato cells in enriched blood samples. Left panel shows total number of tdTomato+ cells observed (p = 0.0629), right panel shows %tdTomato labeled cells within PDGFRa/CD90 cMPC gate. e Immunofluorescent histology of uninjured mouse hindlimb and 12 weeks post-injury hindlimb in Gli1CreERT2:ROSA-LSL-TdTomato (shown in purple) with chevrons marking double-positive cells (Col1a1(2.3 kB)-GFP is shown in green, DAPI shown in blue, scale bars = 100 µm, data are presented as mean values ± SEM). f Quantification of the %tdTomato+ cells within Col1a1(2.3)-GFP+ osteoblasts+ (n = 4) and Aggrecan+ chondrocytes (n = 3) in HO anlagen isolated from Gli1creERT2; tdTomato mice (error bars showing mean ± SD). Source data are provided with this paper.
In day 1 post-B/S control mice, we detected a mean of 4 PDGFRa+/CD90+ per 1 mL blood sample (Fig. 1c, HO-associated MPCs; mean of n = 5 mice), whereas day 1 post-B/T MPC numbers increased precipitously to 16 PDGFRa+/CD90+ (n = 9) and 30 PDGFRa+/CD90+ at day 3-post B/T (n = 9). This dramatic increase was completely reversed by day 7 post-B/T, as the number of MPCs decreased to 3 PDGFRa+/CD90+(n = 5).
Next, we asked whether the cells identified in the circulation were derived from the HO site. To answer this question, we used two lineage tracing systems previously validated by our team25. Specifically, we have shown that Gli1-lineage cells mark bone progenitor cells in the periosteum, whereas Adipoq-lineage cells (adipoCAR cells26,27) mark progenitor cells in the bone marrow. To query if the cMPCs we isolated were from the periosteum or bone marrow, we activated the Cre system with tamoxifen using our previously validated models, followed by a burn tenotomy. We then collected blood and isolated cMPCs as above (Fig. 1a). Interestingly, we were able to detect Gli1CreERT2 cells that were also PDGFRa and CD90 positive, whereas we could not detect adipoCAR cells indicating that the circulating cells most likely derived from Gli1-CreER lineage cells (Fig. 1d). While we previously validated this lineage tracing system in fracture models, we now wanted to confirm that the Gli1 lineage cells marked the same cells at the HO site. Thus, we next employed our Gli1CreERT2;TdTom;Col1a1(2.3)GFP mice to lineage trace the Gli1CreERT2 cells to validate that they are the bone-forming Col1a1 cells. Indeed, prior to injury Gli1CreERT2;TdTom cells were located in the periosteum and peritenon, and co-stained for Col1a1(2.3)GFP (Fig. 1e). Next, we performed our burn/tenotomy model in these mice and analyzed the HO site 12 weeks after injury at which point mature heterotopic bone was noted to stain for both Gli1CreERT2;TdTom and Col1a1(2.3)GFP. This is consistent with previous studies that identified Gli1CreERT2;TdTom cells to mark heterotopic bone28. Importantly, these Gli1CreERT2 cells were not identified in the bone marrow (Fig. 1e), confirming that the cMPCs were not derived from the bone marrow. When we quantified Col1a1(2.3)-GFP+ osteoblasts and Aggrecan+ chondrocytes, we found that %24.5 ± 3.5 of chondrocytes and %33.4 ± 12.6 of osteoblasts within the Gli1CreERT2;TdTom+ population in HO anlagen (Fig. 1f). Thus, for the first time, to the best of our knowledge, we have demonstrated that rare circulating MPCs are present in the blood after an HO-inducing injury and that these derive from Gli1-lineage cells. Given that the Gli1+ lineage cells are not present in the circulation in uninjured mice, and they become detectable in the blood after injury to the Achilles tendon, they are more likely to derive from tissues adjacent to the injury site. Although this is compelling supporting evidence that the circulating cells originate from the HO site, direct evidence to conclusively show this has not yet been obtained. In future experiments, we will narrow down the source of the cells through the use of different Cre models and different injury models.
cMPCs post-B/T express distinct HO-associated transcripts
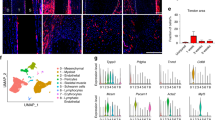
Previous RNA sequencing analyses of MPCs described detailed transcriptional changes that are responsible for HO in mice, including during the critical early time points12,29,30,31. To validate the detection of circulating cells expressing cMPC markers in HOpos conditions, we again leveraged the polytrauma musculoskeletal HO model, where mice receive an Achilles tenotomy at the same time as a 30% total body burn, which reliably forms HO at the site of the tenotomy. We performed single-cell RNAseq on cMPC-Chip-enriched blood from mice after burn/tenotomy (injury leading to HO). We were able to identify cMPCs (Fig. 2a, 13 cMPCs across 2 mice, day 1 post-B/T), expressing mesenchymal markers Dcn, Col1a2, and Fbln2 (Supplementary Fig. 2a).
a UMAP representation of cells detected in iChip-enriched blood product from mouse B/T samples (n = 2). b Experimental design to enrich cMPCs and create a cMPC-based HO RNA signature. Whole blood is drawn from patients that went through hip arthroplasty and mice with B/T or B/S. Blood is then processed via iChip to enrich cMPCs, and RNA is analyzed to identify genes with higher expression in HOpos patients. Stepwise filtering of differentially expressed genes identifies cMPC-based markers highly specific for HO. The gene selection was performed as following: selected genes 1) enriched in HOpos vs HOneg POD1 post-hip arthroplasty, 2) enriched in mouse HOpos B/T when compared to HOneg B/S, and 3) detected in single mouse HO anlagen MPCs by scRNAseq in HOpos B/T model, then 4) removed PBMC genes (expressed >0.5 TPM HPA dataset). Created in BioRender. Karabacak, M. (2026) https://BioRender.com/jgy9pht. c Expression levels of all HO signature genes across HOneg and HOpos hip arthroplasty patients. Exhibit shows log(CPM + 1) normalized data collected from blood samples taken at POD 1 (n = 12 HOneg, n = 10 HOpos). d Expression levels of selected HO signature genes in HOneg and HOpos hip replacement patients across different sampling times (n = 12 POD1 HOneg, n = 11 POD1 HOpos, n = 9 POD14 HOneg, n = 5 POD14 HOpos). e Expression levels of selected HO signature genes in HOneg B/S and HOpos B/T mice across different sampling times (B/S n = 12, B/T n = 27). f Expression levels of HO-associated cMPC genes in MPCs of the injury site following B/T, across different times post-B/T. These data are from previously published work (Clusters 4–7 and 11–12 of the UMAP in Fig. 10a from Sorkin et. al. GEO: GSE126060 dataset), where we referred to these cells as the MPC cluster. Source data are provided with this paper.
We then hypothesized that transcriptional signatures of cMPCs have a specific and sensitive molecular footprint that can predict HO much earlier than current radiographic approaches. To identify a HO-predictive molecular profile of cMPCs, we collected cMPCs from human patients that underwent hip replacement surgery, as well as mice with HOpos B/T and HOpos B/S. We then performed bulk RNAseq and identified genes that are i) increased in HOpos patients on post-operative day 1 (POD1) and Day 1 and 3 post-B/T in mice compared to HOneg samples, ii) expressed in HO-site MPCs based on previously published data and iii) not expressed by PBMCs (Fig. 2b). We identified 32 such cMPC genes (Fig. 2c). As examples, Fig. 2d, e exhibits 6 genes that were particularly enriched: Amotl2, Bace1, Cdh11, Cryab, Lox and Matn2 expression levels in patient and mouse blood samples respectively. These genes were also upregulated in MPCs resident in HO sites in our previously published scRNAseq datasets (Fig. 2f). Thus, we have identified a panel of distinct HO-associated cMPC transcripts that provide a window into the transcriptional changes at the site of musculoskeletal injury and HO formation.
A ML-trained HO prediction model shows strong accuracy for HO
Next, we set to leverage the key genes expressed in our cMPCs to create a prediction model. From the HO gene signature, we created a prediction model trained by 3 commonly used ML approaches using the concentrations of 32 markers as input. We used linear regression (LR), random forest (RF) and support vector machines (SVM). SVM-trained model could achieve >90% sensitivity for human HO with an AUC-ROC of 0.961 (Fig. 3a). We also tested if the SVM model trained on patient data could predict mouse HO and found that it achieved an AUC-ROC of 0.8 at D1 and D3 post-B/T. These models detected HO on average 4 weeks earlier when compared to the current gold standard radiography (Fig. 3). Taken together, these datasets confirm the presence of circulating immature MPC subsets specific to an HO-inducing injury with strong accuracy.
a We used three machine learning approaches (LR: logistic regression, RF: random forest, SVM: support vector machine) to build a model for predicting HO using cMPC HO RNA signature in patients at risk of HO. Data from randomly selected patients (70%) were used to train the model, and the rest (30%) was used to validate and generate an ROC curve. Since the pilot data is relatively small, we repeated the model building and validation 10 times and averaged the model performance, interpolating TPR values from fixed FPRs. Confidence intervals represent 95% of the TPR distribution for a given FPR. The best performing model for predicting HO on POD 1, SVM, achieved an AUC of 0.90, implying a strong predictive power. b We applied an SVM model trained on patient data to validate predictive power in the mouse cMPC bulkRNAseq dataset. Predictions of HO in HOneg B/S to HOpos B/T yielded ROC-AUC of 0.78. Source data are provided with this paper.
Expression of key cMPC genes decreases due to HO-reducing prophylactic treatment
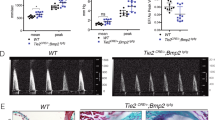
One future goal is to develop our liquid biopsy approach for monitoring patients during prophylactic treatments. This approach could assist in predicting early therapeutic response and guide the therapeutic window for each patient, allowing for more precise treatment timing. We tested the feasibility of detecting a therapy response using cMPC-based liquid biopsy (Fig. 4).
a Micro-CT 3-dimensional reconstruction of mice after B/T treated with BAPN or vehicle control (CTL). b Quantification of HO volume in mice treated with BAPN vs. vehicle control (n = 14 control, n = 14 BAPN). c cMPC genes following BAPN treatment. Left panel exhibits significantly decreased cMPC HO signature genes (all genes q < 0.10 FDR, n = 4, vehicle and BAPN treated mice 3 days post B/T), right panel exhibits the average of all cMPC HO signature genes (p < 0.05). d other cMPC genes that are significantly decreased upon BAPN treatment (all genes q < 0.05 FDR, p-values range from 0.001 to 0.11, n = 4, vehicle and BAPN treated mice 3 days post B/T). These genes were not sensitive and specific for the early detection of human HO, but were detected in mouse B/T. Source data are provided with this paper.
To do this, we enriched mouse cMPCs and performed bulk RNAseq following B/T with or without lysyl oxidase inhibition via β-aminopropionitrile (BAPN) treatment, which we have shown to significantly decrease HO formation32 (Fig. 4a, b). Indeed, bulk RNAseq of enriched cMPCs data showed that mice treated with BAPN had a statistically significant decrease in expression of 8 genes in BAPN-treated mice compared to control (Fig. 4c). Furthermore, we detected several MPC-specific genes and ECM-associated genes33 to be significantly reduced due to BAPN treatment (Fig. 4d, Supplementary Fig. 3). In summary, we have shown that the HO prediction model can also help predict therapeutic efficacy.
Non-HO inducing musculoskeletal injury can be distinguished from HO-inducing injury using cMPC concentration or cMPC HO RNA signature
To test if cMPC release is specific to injury that leads to HO bone formation, we obtained blood samples from mice that went through other modes of injury: 1) a severe closed head concussive injury via weight drop that mimics human traumatic brain injury34 and 2) muscle fibrosis induced by intramuscular injection of cardiotoxin35. We have not observed a significant increase in cMPC numbers in these non-HO inducing models (Supplementary Fig. 4).
Circulating cells with osteogenic progenitor character have been observed in patients following fracture10,11. Therefore, we performed a fracture model36 in mice to ask whether cMPCs post-HO inducing B/T were distinct from the circulating osteogenic cells following non-HO inducing fracture. We performed iChip enrichment followed by bulk RNA sequencing and compared the cMPC HO signature between HOneg fracture and HOpos B/T. While some of the cMPC genes were observed in both conditions with statistically indistinguishable levels, we found that Lrig, Gpx8, P3h3, Hmcn1, Cdh11 and Cryab were significantly higher in HOpos B/T at POD1, when compared to fracture (Supplementary Fig. 5), and the mean of the cMPC HO signature was higher post-B/T when compared to fracture at day 1.
Overall, this set of data supports the approach to use cMPC concentration and RNA signature at an early time point to predict HO following surgery and trauma.
Discussion
Changes associated with HO occur much earlier than standard radiographic detection; however, these modalities require special imaging and are not regularly available37,38,39. Currently, clinical detection of HO is performed with CT (takes an average of 23 days after symptom development to detect HO)40; bone scans with 99mTc-MDP (low specificity, which leads to difficulties in discriminating HO from other inflammatory, traumatic, or degenerative skeletal processes) and serum alkaline phosphatase measurement (sensitive, not specific, with alterations dependent on hepatic and renal function)41. Thus, one of the major knowledge gaps in HO treatment is a lack of accurate early diagnostic modalities. Precision medicine approaches to guide therapy for those patients who are developing HO, as well as precise timing of treatment, would allow for shorter treatment duration with minimization of adverse consequences, decreases in cost, and improvement in treatment adherence. Additionally, an in vivo blood-based biomarker would allow for targeted treatment on the basis of early identification of changes in MPC fate. Demonstration of therapeutic efficacy based on patient selection upon cMPC evidence of HO would dramatically enhance current treatment paradigms.
When taken together, the data presented here confirm the presence of cMPCs that are sensitive and specific to an HO-inducing injury, both in patients and in a trauma-based mouse model of HO. The data presented suggest that MPCs are mobilized from the HO site into the bloodstream after an HO-inducing injury and express HO-specific genes, long before HO can be detected radiographically. Beyond early diagnosis, our findings also validate cMPCs as a liquid biopsy to accurately assess treatment efficacy.
Previous studies focused on near-infrared (NIR) tetracycline markers that are deposited within ossifying tissues and can be observed through non-invasive in vivo imaging38. Near infrared tetracycline imaging can detect mineral deposition as early as one week after trauma and at least 5 weeks before radiographic evidence38. Similarly, Raman Spectroscopy and spectral ultrasound technologies have been used for HO detection in the B/T mouse models used here42,43,44. These technologies, however, have yet to be translated into human use. Each of these imaging systems are also large and require complex equipment and would be difficult to deploy in the setting of prolonged field care, thus making our microfluidic technology an important advance for portable, point of care HO detection. The refinement of a sensitive, non-invasive technique will identify candidates with the greatest risk for HO and allow us to implement inhibitor therapies before soft tissues have undergone irreversible osteo-chondrogenic remodeling in patients with cMPC evidence of early HO.
In addition to leveraging cMPC transcriptional data for HO detection, we think that the identified transcripts from cMPCs may also provide novel pathways that can be targeted therapeutically since these genes have functional relevance for endochonral ossification. While HO site MPCs could also provide such data, it is currently contraindicated to perform additional tissue biopsies after surgery in patients at risk for HO. Thus, cMPCs provide the only tissue that can be isolated and analyzed early after surgery during the time frame in which the key aberrant transcriptional programs are active. The presented human cMPC data identified 24 genes related to biological processes associated with bone formation (Supplementary Table 1).
These genes encompass signaling pathway modulators (Amotl2 in Wnt/Hippo signaling45, Evc2 in hedgehog signaling46 Lrig3 in BMP signaling47, Tspan10 in Notch signaling), ECM components and processors (Bmper as a regulator of bone formation48, Hmcn1 as ECM component49,50, Matn2 in matrix assembly51, Lox and P3h3 in collagen crosslinking and processing52, Pcolce2 in collagen synthesis53), metabolic and cellular regulation (Gfpt2 in glucose metabolism, Igfbp5 in IGF signaling54), bone formation (Cryab in chondrogenic and osteogenic differentiation55,56, Postn in skeletal response to parathyroid hormone57,58, Scara3 in mesenchymal progenitor cell fate59), tissue modeling (Mmp2). Overall, these genes have functional relevance to tHO, and are not detected in the bloodstream for surgery patients that do not develop HO and mice without HO. Therefore, screening these markers in circulation may benefit future studies of traumatic HO and other musculoskeletal pathologies, and for monitoring of prophylactic strategies.
This study has several limitations. First, our microfluidic technology, iChip, requires freshly drawn whole blood samples analyzed within 24 h; therefore, preserved and cryobanked samples currently cannot be used to enrich cMPCs in circulation. We also only focused on hip arthroplasty patients, but have not explored other clinical conditions such as burn and spinal cord injury. Analysis of blood samples from these patient populations is needed for a broader validation and further refinement of our method.
Our study used RNA sequencing to provide a comprehensive transcriptomic analysis for the identification of HO-associated cMPC transcriptional signatures. While RNAseq offers broad molecular profiling capabilities, droplet digital PCR (ddPCR) represents a more scalable and highly sensitive method for targeted RNA analysis. ddPCR provides absolute quantification and demonstrates outstanding sensitivity for detecting low-abundance, rare cell-derived transcripts, which are critical advantages for clinical diagnostic applications.
For future clinical validation studies, we plan to develop a ddPCR assay targeting the HO-associated transcriptional signatures identified in this work. The standardized protocols and faster turnaround times of ddPCR will facilitate the prospective clinical validation of our liquid biopsy approach, ultimately enabling translation of our findings into clinical practice.
Liquid biopsy approach using cMPCs could address in HO prevention which patients are at risk, the duration of treatment and the timing of treatment initiation. Treatment timing is important for several reasons, including: 1. Reduced treatment duration—by identifying the optimal treatment timing, we can minimize treatment length and monitor for recurrence. This targeted timing will minimize adverse consequences of therapeutics, improve patient adherence, and reduce costs; and 2. Precision medicine-based patient selection—portable, point-of-care, non-invasive diagnostic modalities that can risk-stratify patients will allow for targeted treatments so that only patients at risk receive treatment. The potential impact of cMPC liquid biopsy is not limited to the patients at risk of HO. Aberrant cell fate and repair associated with mesenchymal progenitor cells are also observed in osteoarthritis60 and fibrosis61. Early blood-based detection strategies that can be routinely applied to patients at risk are an active area of research for these diseases. When further developed and validated, cMPC liquid biopsy could provide direct and easy access to cMPCs for molecular, physiological, and pharmacological characterization in a vast array of acute or chronic diseases.
Methods
Ethical statement
All human research was reviewed and approved by UTSW IRB (STU-2020-0552). Informed consent was obtained from all human research participants. All mice studies were reviewed and approved by the University of Texas Southwestern Medical School Institutional Animal Care and Use Committee (Protocol 2020-102949).
Mouse injury models
Gli1creERT mice (JAX:007913), Adiponectin-cre mice (JAX:028020), Rosa-LSL-tdTomato mice (JAX:007909), and Col1a1(2.3kB)-GFP mice (JAX:013134) were purchased from Jackson Laboratory. B/T injuries were performed as previously described12. Briefly, 8–10-week-old C57B6 mice (Jackson Laboratory) were provided buprenorphine sustained release for analgesia (1 mg/kg subcutaneous injection) and isoflurane in oxygen (2.5%) for anesthesia. Mice were prepared for surgery by sharing the back and left hindlimb with clippers. Burn and tenotomy sites were cleaned by three alternating scrubs of ethanol and betadine. Anesthetic depth was assessed by hindlimb pinch prior to injury. The tenotomy was performed by making a small skin incision to expose the Achilles’ tenotomy, isolating the tendon using surgical scissors, and fully transecting the tendon at its midpoint. For B/S procedures, the tendon was isolated but not transected. Skin was closed using absorbable 5–0 Vicryl suture. Metal blocks were warmed to 60 °C in a water bath and applied to the mouse's back for 18 s. Mice were recovered from anesthesia in a cage warmed by a heating pad before returning to their original cage. Mice were assessed for complications and pain daily for three days following injury. The surgeon was not blinded to the surgical procedure. For bicortical fractures, the tibia was fractured mid-diaphysis by 3-point bending using a Zondervan apparatus36. The location of the pin was verified by X-ray radiography on an AMI HTX Optical Imaging System (Spectral Instruments Imaging). Buprenorphine SR was injected immediately after the surgery, while the mice were still anesthetized.
Patient enrollment
We established a clinical research protocol to collect blood from patients undergoing hip arthroplasty as well as to follow their eventual HO formation by x-ray and CT. To determine the molecular profile of cMPCs, we sampled peripheral blood from 22 patients undergoing hip arthroplasty pre-operatively and on post-operative day (POD)1 and 14 (Sex was self-reported (11 male and 11 female patients/donors). Age range was 20–94 and average age was 68. Sex-based analyses were not performed due to the limited sample size). We then followed these patients by x-ray at days 21 and 42 for determining HOpos patients (we included two representative x-rays from the enrolled patients in Supplementary Fig. 6).
BAPN treatment
Following B/T or B/S, 8 mg/ml BAPN in 5% sucrose water, or 5% sucrose water control was provided to mice in bottles for ad libitum consumption. Bottles were shaken daily and replaced twice weekly. Mice were treated until tissue harvest.
Microfluidic enrichment
All experiments used an iChip equipped with a non-equilibrium inertial separation array (NISA)62. Blood samples were drawn in the University of Texas Southwestern Medical Center, into ACD tubes, transported overnight under normothermic conditions and processed within 24 h of blood draw. Before enrichment, cells were incubated with antibodies against biotinylated CD45, CD16 and CD66b for a minimum of 30 min, and then with 1 µm streptavidin-coated superparamagnetic beads for 20 min17,63. Then, whole blood is passed through iChip, and an enriched product containing cMPCs is collected. We then performed a second step of enrichment by repeating the reagent incubation and microfluidic enrichment steps.
Bulk RNA sequencing
RNA was purified from lysed cells using the Qiagen Micro RNeasy kit. We prepared libraries for bulk RNAseq using the Plexwell Rapid Library Preparation kit (Seqwell)64 following manufacturer instructions. Sequencing was performed at MGH Nextgen Core Facility using Illumina NextSeq 2000. Following debarcoding, the data was aligned and converted to reads files in Illumina Basespace RNA-Seq Alignment App (v.2.0.2, Illumina, CA).
Gene selection
Gene-level RNA quantification from Salmon files was read into a count matrix of genes and samples. edgeR65 and DESeq66 analysis pipelines were applied to identify differentially expressed genes between HOpos and HOneg populations. A database of 20,162 genes and their expression levels (TPM) in human PBMCs was obtained from the Human Protein Atlas67. 6,021 genes expressed at >0.5 TPM in human PBMCs were filtered from the differential expression analysis. Genes with ratio of geometric means for (CPM + 1) expression level above 7 were considered differentially expressed (GM(HOpos(CPM + 1)) / GM(HOneg(CPM + 1)) > 7). CPM of 13,262 genes from i) enriched mouse cMPC bulk RNAseq datasets (B/T and B/S conditions) and ii) 1049 single mouse MPCs from GEO: GSE12606012 were log-transformed, and mouse genes were mapped to their human orthologs using the R package orthogene. 11,285 genes (85%) where human orthologs existed were retained after mapping. An average CPM across all 1049 cells was computed for each gene. HO resident MPC genes were then ranked in order of average single-cell expression, and edgeR65 and DESeq66 analysis pipelines were applied to identify differentially expressed genes between HOpos B/T and HOneg B/S mice. HO signature gene candidates were then selected from the union of differential expression among HOpos patients and mice when compared to HOneg counterparts, and the highest-ranked genes expressed in resident MPCs.
Model training
A gene signature defined as the sum of the log(CPM + 1) expression of selected genes among the 22 patient samples was constructed, and a binary response column was added to indicate HO status as either a 1 or a 0. The data was partitioned into random training (70%) and validation (30%) sets. A linear Support Vector Machine (SVM) classifier was then trained on these data using the svm function in the R package e1071 and validated using the predict function in the stats package. Similarly, random forest (RF) and linear regression models were trained and validated by the randomforest and stats packages. The receiver-operator characteristic (ROC) curve for the reported calibrated probabilities by these classifiers was constructed, and the area under the curve (AUC) was computed.
Single-cell RNA sequencing
iChip-enriched cells were captured and barcoded via the 10X instrument at MGH Nextgen Core Facility. 10x Genomics Cell Ranger 7.1.0 (via 10x Genomics Cloud Analysis) was used to perform sample de-multiplexing, barcode processing, and single-cell gene counting (alignment, barcoding, and unique molecular identifier [UMI] count)68. Reads were aligned to the mm10-2020-A reference genome as appropriate for the input dataset. Downstream analysis steps were performed using Seurat v569. Cells identified as poor libraries or resulting from doublet cells were filtered by eliminating cells on both ends of the distribution, as well as cells with unusually high percent mitochondrial transcripts. Normalization, scaling, and dimensional reductions using principal component analysis (PCA) and uniform manifold approximation and projection (UMAP) were performed using Seurat v5. Integrative analysis was performed using Harmony70. Cell type labels were assigned based on characteristic relative marker gene expression levels between cell clusters using the FindAllMarkers function. Module scores were generated using the AddModuleScore function of Seurat to identify cell populations of interest. Module scores were calculated as the level of gene expression enrichment of a set gene list relative to a random control list, with higher module score values representing positive enrichment beyond background. Dimensional feature plots, violin plots, and dot plots were used to visualize gene expression within the population of interest.
Flow cytometry
iChip isolated cells were labeled with a cocktail of antibodies in 0.5% BSA, 0.1% F68-PBS at 4 °C. The antibody cocktail consisted of PDGFRa-SB600 (APA5, Invitrogen), CD90-eFluor450 (Thy-1.1, Invitrogen), CD45-BUV395 (30-F11, BD), Ter119-BUV395 (Ter-119, BD). Then, cells were washed and suspended in PBS 0.5% BSA before being analyzed on a FACS Aria Fusion (BD Biosciences). For compensation, all antibodies were conjugated to OneComp eBeads (Thermo Fisher Scientific). Unstained control cells and compensation beads were used for performing compensation. We used PDGFRa + CD90 + MPCs isolated from bone tissue using previously established protocols71,72 and PBMCs for determining the gating strategy. For transgenic mouse experiments, we also used Gli1CreERT2-tdTomato+ control cells isolated from periosteum as compensation controls and for determining gates.
Microscopy
Freshly dissected tissues were fixed overnight in 4% paraformaldehyde in PBS at 4 °C, then washed several times in PBS. Tissues were then decalcified in 10% EDTA (w/v) in PBS for 4 weeks, then paraffin-embedded. 5 μm thick tissue sections were cut and adhered to glass slides. Sections were then deparaffinized and stained using the following procedure. Sections were permeabilized for 10 min with 0.05% NP40 and 2.5% DMSO in Hank’s balanced salt solution (HBSS), then washed in PBS and blocked in PBS with 5% normal donkey serum (Jackson Immunoresearch) for 1 h. Slides were then stained overnight with combinations of the following antibodies: chicken anti-GFP (1–200, Aves Labs, GFP-1020), goat anti-dsRed (1–200, LifeSpan Biosciences, LS-C340696), rabbit anti-Aggrecan (1–35, EMD Millipore, AB1031). Secondary antibodies included donkey anti-chicken Alexa Fluor 488 (1–250, Jackson Immunoresearch), donkey anti-rabbit Alexa Fluor 647 (1–250, Jackson Immunoresearch), and donkey anti-goat CF-555 (1–500, Biotium). Sections were counterstained with DAPI to stain nuclei before mounting. Slides were mounted with Prolong Gold anti-fade reagent (Invitrogen). Images were acquired with a Zeiss LSM880 confocal microscope. Confocal images were processed and analyzed using Fiji (Image J) and Photoshop (Adobe Systems).
Statistics and reproducibility
Graphpad Prism 10 and BioRender were used to perform statistical analyses and prepare illustrations. All statistical analyses are two-sided. Gene expression is illustrated using scattered dot plots, with bars indicating the mean value and error bars representing standard deviations. No data was excluded from analysis. No statistical method was used to predetermine sample size. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Flow cytometry data was deposited to the Flow Repository and is available under Repository ID FR-FCM-Z7CJ. RNAseq data have been deposited to GEO and is available under accession codes GSE267142, GSE267143GSE267144, and GSE316836. The previously published sequencing data [13] used in this work is available via accession number GSE126060. Source data are provided with this paper.
References
Zeckey, C. et al. Risk of symptomatic heterotopic ossification following plate osteosynthesis in multiple trauma patients: an analysis in a level-1 trauma centre. Scand. J. Trauma Resusc. Emerg. Med 17, 55 (2009).
Mital, M. A., Garber, J. E. & Stinson, J. T. Ectopic bone formation in children and adolescents with head injuries: its management. J. Pediatr. orthopedics 7, 83–90 (1987).
Levi, B. et al. Risk factors for the development of heterotopic ossification in seriously burned adults: a National Institute on Disability, Independent Living and Rehabilitation Research burn model system database analysis. J. Trauma Acute Care Surg 79, 870–876 (2015).
Nauth, A. et al. Heterotopic ossification in orthopaedic trauma. J. Orthop. Trauma 26, 684–688 (2012).
Dey, D. et al. The traumatic bone: trauma-induced heterotopic ossification. Transl. Res 186, 95–111 (2017).
Kluger, G., Kochs, A. & Holthausen, H. Heterotopic ossification in childhood and adolescence. J. Child Neurol. 15, 406–413 (2016).
Merkely, G., Chisari, E., Lola Rosso, C. & Lattermann, C. Do nonsteroidal anti-inflammatory drugs have a deleterious effect on cartilage repair? a systematic review. Cartilage 13, 326S–341S (2021).
Oh, J. H. et al. Do selective COX-2 inhibitors affect pain control and healing after arthroscopic rotator cuff repair? A preliminary study. Am. J. Sports Med 46, 679–686 (2018).
Suda, R. K. et al. Circulating osteogenic precursor cells in heterotopic bone formation. Stem Cells 27, 2209–2219 (2009).
Alm, J. J. et al. Circulating plastic adherent mesenchymal stem cells in aged hip fracture patients. J. Orthop. Res 28, 1634–1642 (2010).
Eghbali-Fatourechi, G. Z. et al. Circulating osteoblast-lineage cells in humans. N. Engl. J. Med 352, 1959–1966 (2005).
Sorkin, M. et al. Regulation of heterotopic ossification by monocytes in a mouse model of aberrant wound healing. Nat. Commun. 11, 722 (2020).
Loder, S. J. et al. Characterizing the Circulating Cell Populations in Traumatic Heterotopic Ossification. Am. J. Pathol. 188, 2464–2473 (2018).
Sullivan, J. P. et al. Brain tumor cells in circulation are enriched for mesenchymal gene expression. Cancer Discov. 4, 1299–1309 (2014).
Ting, D. T. et al. Single-cell RNA sequencing identifies extracellular matrix gene expression by pancreatic circulating tumor cells. Cell Rep. 8, 1905–1918 (2014).
Miyamoto, D. T. et al. RNA-Seq of single prostate CTCs implicates noncanonical Wnt signaling in antiandrogen resistance. Science 349, 1351–1356 (2015).
Fachin, F. et al. Monolithic Chip for High-throughput Blood Cell Depletion to Sort Rare Circulating Tumor Cells. Sci. Rep. 7, 10936 (2017).
Kalinich, M. et al. An RNA-based signature enables high specificity detection of circulating tumor cells in hepatocellular carcinoma. Proc. Natl. Acad. Sci. USA 114, 1123–1128 (2017).
Hong, X. et al. Molecular signatures of circulating melanoma cells for monitoring early response to immune checkpoint therapy. Proc. Natl. Acad. Sci. USA 115, 2467–2472 (2018).
Kwan, T. T. et al. A Digital RNA Signature of Circulating Tumor Cells Predicting Early Therapeutic Response in Localized and Metastatic Breast Cancer. Cancer Discov. 8, 1286–1299 (2018).
Miyamoto, D. T. et al. An RNA-Based Digital Circulating Tumor Cell Signature Is Predictive of Drug Response and Early Dissemination in Prostate Cancer. Cancer Discov. 8, 288–303 (2018).
Ozkumur, E. et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci. Transl. Med 5, 179ra47 (2013).
Wong, K. H. K. et al. Whole blood stabilization for the microfluidic isolation and molecular characterization of circulating tumor cells. Nat. Commun. 8, 1733 (2017).
Agarwal, S. et al. Surgical excision of heterotopic ossification leads to re-emergence of mesenchymal stem cell populations responsible for recurrence. STEM CELLS Transl. Med. 6, 799–806 (2017).
Jeffery, E. C., Mann, T. L. A., Pool, J. A., Zhao, Z. & Morrison, S. J. Bone marrow and periosteal skeletal stem/progenitor cells make distinct contributions to bone maintenance and repair. Cell Stem Cell 29, 1547–1561.e6 (2022).
Zhong, L. et al. Single cell transcriptomics identifies a unique adipose lineage cell population that regulates bone marrow environment. eLife 9, e54695 (2020).
Baccin, C. et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat. Cell Biol. 22, 38–48 (2020).
Kan, C. et al. Gli1-labeled adult mesenchymal stem/progenitor cells and hedgehog signaling contribute to endochondral heterotopic ossification. Bone 109, 71–79 (2018).
Huber, A. K. et al. Immobilization after injury alters extracellular matrix and stem cell fate. J. Clin. Invest. 130, 5444–5460 (2020).
Lee, S. et al. NGF-TrkA signaling dictates neural ingrowth and aberrant osteochondral differentiation after soft tissue trauma. Nat. Commun. 12, 4939 (2021).
Vishlaghi, N. et al. Vegfc-expressing cells form heterotopic bone after musculoskeletal injury. Cell Rep. 43, 114049 (2024).
Kang, H. et al. The HIF-1α/PLOD2 axis integrates extracellular matrix organization and cell metabolism, leading to aberrant musculoskeletal repair. Bone Res. 12, 17 (2024).
Feuermann, M. et al. A compendium of human gene functions derived from evolutionary modelling. Nature 640, 146–154 (2025).
Khuman, J. et al. Tumor necrosis factor alpha and Fas receptor contribute to cognitive deficits independent of cell death after concussive traumatic brain injury in mice. J. Cereb. Blood Flow. Metab. 31, 778–789 (2011).
Guardiola, O. et al. Induction of acute skeletal muscle regeneration by cardiotoxin injection. J. Vis. Exp. 54515 https://doi.org/10.3791/54515 (2017).
Zondervan, R. L., Vorce, M., Servadio, N. & Hankenson, K. D. Fracture apparatus design and protocol optimization for closed-stabilized fractures in rodents. J. Vis. Exp. 58186 https://doi.org/10.3791/58186 (2018).
Peterson, J. R. et al. Early detection of burn-induced heterotopic ossification using transcutaneous Raman spectroscopy. Bone 54, 28–34 (2013).
Perosky, J. E. et al. Early detection of heterotopic ossification using near-infrared optical imaging reveals dynamic turnover and progression of mineralization following Achilles tenotomy and burn injury. J. Orthop. Res 32, 1416–1423 (2014).
Ranganathan, K. et al. High-frequency spectral ultrasound imaging (SUSI) visualizes early post-traumatic heterotopic ossification (HO) in a mouse model. Bone 109, 49–55 (2018).
Brownley, R. C. et al. Characterization of heterotopic ossification using radiographic imaging: evidence for a paradigm shift. PloS one 10, e0141432 (2015).
Citak, M. et al. The roles of serum alkaline and bone alkaline phosphatase levels in predicting heterotopic ossification following spinal cord injury. Spinal Cord. 54, 368–370 (2016).
Lin, Y., Lin, W. Y., Kao, C. H. & Tsai, S. C. Easy interpretation of heterotopic ossification demonstrated on bone SPECT/CT. Clin. Nucl. Med 39, 62–63 (2014).
Eekhoff, E. M. W. et al. 18F]NaF PET/CT scan as an early marker of heterotopic ossification in fibrodysplasia ossificans progressiva. Bone 109, 143–146 (2018).
Ghanem, M. A., Dannoon, S. & Elgazzar, A. H. The added value of SPECT-CT in the detection of heterotopic ossification on bone scintigraphy. Skelet. Radio. 49, 291–298 (2020).
Dai, X. et al. Phosphorylation of angiomotin by Lats1/2 kinases inhibits F-actin binding, cell migration, and angiogenesis. J. Biol. Chem. 288, 34041–34051 (2013).
Kwon, E. K. et al. The role of ellis-van creveld 2(EVC2) in mice during cranial bone development. Anat. Rec. (Hoboken) 301, 46–55 (2018).
Herdenberg, C. et al. LRIG proteins regulate lipid metabolism via BMP signaling and affect the risk of type 2 diabetes. Commun. Biol. 4, 90 (2021).
Cao, H. et al. The Pitx2:miR-200c/141:noggin pathway regulates Bmp signaling and ameloblast differentiation. Development 140, 3348–3359 (2013).
Cui, Y. et al. Hemicentin-1 is an essential extracellular matrix component during tooth root formation by promoting mesenchymal cell differentiation. Front Cell Dev. Biol. 12, 1435241 (2024).
Wischmann, J. et al. Matrix mineralization controls gene expression in osteoblastic cells. Exp. Cell Res. 372, 25–34 (2018).
Li, P. et al. Mice lacking the matrilin family of extracellular matrix proteins develop mild skeletal abnormalities and are susceptible to age-associated osteoarthritis. Int. J. Mol. Sci. 21, 666 (2020).
Vallet, S. D. & Ricard-Blum, S. Lysyl oxidases: from enzyme activity to extracellular matrix cross-links. Essays Biochem 63, 349–364 (2019).
Vadon-Le Goff, S. et al. Identification of PCPE-2 as the endogenous specific inhibitor of human BMP-1/tolloid-like proteinases. Nat. Commun. 14, 8020 (2023).
Mukherjee, A. & Rotwein, P. Insulin-like growth factor binding protein-5 in osteogenesis: facilitator or inhibitor? Growth Horm. IGF Res 17, 179–185 (2007).
Tian, B. et al. CRYAB suppresses ferroptosis and promotes osteogenic differentiation of human bone marrow stem cells via binding and stabilizing FTH1. Aging (Albany NY) 16, 8965–8979 (2024).
Granéli, C. et al. Novel markers of osteogenic and adipogenic differentiation of human bone marrow stromal cells identified using a quantitative proteomics approach. Stem Cell Res 12, 153–165 (2014).
Bonnet, N., Conway, S. J. & Ferrari, S. L. Regulation of beta-catenin signaling and parathyroid hormone anabolic effects in bone by the matricellular protein periostin. Proc. Natl. Acad. Sci. 109, 15048–15053 (2012).
Duchamp de Lageneste, O. et al. Periosteum contains skeletal stem cells with high bone regenerative potential controlled by Periostin. Nat. Commun. 9, 773 (2018).
Chen, P. et al. Scara3 regulates bone marrow mesenchymal stem cell fate switch between osteoblasts and adipocytes by promoting Foxo1. Cell Prolif. 54, e13095 (2021).
Sono, T. et al. Perivascular Fibro-Adipogenic Progenitor Tracing during Post-Traumatic Osteoarthritis. Am. J. Pathol. 190, 1909–1920 (2020).
Harvey, T., Flamenco, S. & Fan, C.-M. A Tppp3+Pdgfra+ tendon stem cell population contributes to regeneration and reveals a shared role for PDGF signalling in regeneration and fibrosis. Nat. Cell Biol. 21, 1490–1503 (2019).
Mishra, A. et al. Ultrahigh-throughput magnetic sorting of large blood volumes for epitope-agnostic isolation of circulating tumor cells. Proc. Natl. Acad. Sci. USA 117, 16839–16847 (2020).
Karabacak, N. M. et al. Microfluidic, marker-free isolation of circulating tumor cells from blood samples. Nat. Protoc. 9, 694–710 (2014).
Hahaut, V. et al. Fast and highly sensitive full-length single-cell RNA sequencing using FLASH-seq. Nat. Biotechnol. 40, 1447–1451 (2022).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Uhlen, M. et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science. 366, eaax9198 (2019).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Severe, N. et al. Stress-induced changes in bone marrow stromal cell populations revealed through single-cell protein expression mapping. Cell Stem Cell 25, 570–583.e7 (2019).
Baryawno, N. et al. A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell 177, 1915–1932.e16 (2019).
Acknowledgments
We are grateful to the MGH Nextgen Sequencing, MGH Flow and Mass Cytometry Cores for technical assistance with Flow Cytometry and RNA sequencing. We thank the following orthopedic surgeons at UTSW that contributed to this work: Drs. Robert Weinschenk, Varatharaj Mounasamy, Alexandra Callan, Michael Huo, Garen Collett, Ishvinder Grewal, Antonia Chen. We thank the technical assistance from Drs. Avanish Mishra, Jon Edd, Ezgi Antmen, Qin Tao, and Sweta Munagapati. This work was supported by funding from Shriners Hospital for Children, MGH ECOR, Orphan Disease Center (to N.M.K.). Dr. Levi is grateful for support from the NIH (R61AR078072/R33AR078072 and R01AR078324). We also appreciate support from Surgical Partners in Research (SPAR) at UTSW, including Jarrett Hubbard and Katie Naumann.
Author information
Authors and Affiliations
Contributions
J.H.N. performed animal experiments; M.H. performed iChip enrichment, data analysis, machine learning; S.K. performed bioinformatic analysis; H.K., F.L., and H.S. performed iChip enrichment and RNA-seq experiments; C.P. developed experimental design and performed animal experiments; A.S. performed bioinformatic analysis; E.C.J. developed experimental design and animal experiments; M.E., A.C., and A.F.C developed clinical design and supported patient enrollment; S.N. performed flow cytometry; R.T. designed bioinformatic approach; J.C. performed animal experiments and microCT analysis, H.K. performed animal experiments; C.A. provided mouse model expertise; M.T. contributed methodology; B.L. and N.M.K. conceived the study, directed the study, contributed methodology, and coordinated the collaborative work. N.M.K. and B.L. wrote the paper. All the authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Massachusetts General Hospital has filed a patent application for the use of rare cell isolation technology in heterotopic ossification described in this publication (PCT Application No. PCT/US2024/055005). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Matthew Greenblatt and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nunez, J., Holtz, M., Korlakunta, S. et al. Early detection of aberrant cell fate and repair using circulating progenitor cells in patients with heterotopic ossification. Nat Commun 17, 2231 (2026). https://doi.org/10.1038/s41467-026-68857-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68857-8