Abstract
The endothelin receptor antagonist atrasentan improved kidney outcomes in the SONAR trial for type 2 diabetes and chronic kidney disease (NCT01858532), though individual responses varied. To identify molecular biomarkers of atrasentan response and outcome, we conducted a nested case-control proteomics study (N = 180) within the SONAR trial population and identified urinary clusterin (uCLU) as the top candidate. Transcriptomic analyses of human kidney biopsies at tissue and single cell level from independent cohorts revealed higher CLU mRNA levels associated with worse kidney function and outcomes. An endothelin signaling activation score derived from pathway genes was reduced by atrasentan in mice with diabetic kidney disease. In the SONAR trial (N = 3,060) population, higher uCLU predicted worse outcomes, while atrasentan reduced uCLU by 42.6% over six weeks. Early uCLU changes independently predict improved kidney outcomes. In summary, uCLU is associated with kidney disease progression and response to atrasentan treatment, supporting its potential as a pharmacodynamic biomarker to target therapy.
Similar content being viewed by others
Introduction
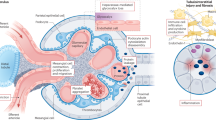
Endothelin-1 (ET-1) is a potent vasoactive peptide that is often elevated in the context of hyperglycemia, hyperinsulinemia, inflammation, and increased renin-angiotensin-system activity1. Binding of ET-1 to the endothelin receptor type A (ETA) in the kidney causes sustained vasoconstriction of the afferent arterioles, hyperfiltration, podocyte damage, endothelial glycocalyx dysfunction, and proteinuria, leading to progressive kidney injury and decline in estimated glomerular filtration rate (eGFR), a measure of kidney function2,3,4. Selective endothelin receptor A antagonists (ERAs) have been shown to ameliorate kidney injury in animal models of chronic kidney disease (CKD) and slow the progression of CKD in clinical trials5,6,7. Specifically, the ERA, atrasentan, decreased albuminuria and improved kidney outcomes in patients with type 2 diabetes and CKD (T2D-CKD)8,9. However, there were variations in the response to atrasentan among participants, indicating that not everyone benefited from the treatment to the same extent10,11.
A precision medicine approach could help identify those most responsive to ERA treatment. In the phase 3 SONAR trial of atrasentan in patients with T2D-CKD (Clinicaltrials.gov: NCT01858532), the change in urinary albumin to creatinine ratio (uACR), a validated surrogate of kidney function decline and failure, was used to select responders to atrasentan treatment9,12. Trial completion, however, revealed that this strategy had not worked as anticipated6. Many aspects of the trial design could have contributed to the poor performance of uACR as a treatment-response biomarker, including the large fluctuations in uACR and difficulty in separating random variations in uACR from pharmacological atrasentan-specific treatment effects9. Another limitation was that uACR, as a generic downstream biomarker of CKD progression, lacked specificity for atrasentan response since it did not reflect the activity of the endothelin signaling system. A molecular biomarker or a biomarker panel associated with both the pathogenesis of T2D-CKD and the activity of the endothelin system could be a more promising pharmacodynamic biomarker candidate to stratify individual treatment responses to atrasentan.
Recent advances in multi-omics technologies have enabled mechanistic biomarker identification for prediction of CKD progression and/or response to treatment9,13,14,15,16. The ideal mechanistic treatment response biomarker or biomarker panel should be supported by preclinical model systems with mechanistic insight anchored in human kidney tissue, which can then be captured by a routine, non-invasive diagnostic procedure.
This study aims to identify and validate mechanistic pharmacodynamic biomarkers significantly associated with the pathophysiology of T2D-CKD and response to atrasentan. In a nested case-control study of SONAR trial participants, sequential urinary proteomic profiles are used to identify candidate urinary protein biomarkers associated with disease progression and treatment response. Here, we show that kidney tissue transcriptomics in atrasentan treated preclinical CKD animal models and human CKD cohorts provide mechanistic support for the kidney-specific link between the biomarker, endothelin pathway activation and kidney outcomes. Finally, the pharmacodynamic performance of the biomarker is validated across the entire SONAR trial population. The analytical approach used in this study is schematically summarized in Fig. 1.
Schematic of the overall approach to identifying, prioritizing, and validating a non-invasive urinary biomarker of atrasentan responsiveness. Responder: uACR reduction > 45% during 6-week treatment and eGFR change between − 1.0 and + 1.0 mL/min/1.73 m² per year over a median 2.2-year period. Non-responder: uACR reduction < 15% during 6-week treatment and GFR decline > 3.0 mL/min/1.73 m² per year over the same period. Conditional responder: uACR reduction > 45% during 6-week treatment, but eGFR decline > 3.0 mL/min/1.73 m² per year after transitioning to placebo over the same period. Abbreviations: uACR – urinary albumin to creatinine ratio; eGFR – estimated glomerular filtration rate.
Results
Dynamic biomarkers associated with atrasentan response
To identify biomarkers associated with response to atrasentan treatment, we conducted a nested case-control biomarker discovery study from the SONAR trial including three groups: responders, non-responders and conditional responders to treatment with atrasentan, based on changes in uACR during the 6 weeks treatment with atrasentan and on annual changes in eGFR during a median 2.2 year atrasentan treatment (Table 1 and Supplementary Fig. S1). Using these two parameters we defined a “responder” group, (R), as participants with an uACR reduction of at least 45% and with a stable eGFR changes between − 1.0 to + 1.0 mL/min/1.73m2 per year; a “non-responder” group, (NR), defined as participants with an uACR reduction of less than 15% and with an eGFR decline of more than 3 mL/min/1.73 m2 per year; a “conditional responder” group (CR), defined as participants with an uACR reduction of at least 45% and who transitioned to placebo at randomization and with an eGFR decline of more than 3.0 mL/min/1.73 m2 per year after transition to placebo. The three groups were well balanced with respect to risk factors of eGFR decline, with the exception that the baseline eGFR in responders (42.6 compared to 44.5 mL/min/1.73m2 overall) was slightly lower compared to the other 2 groups. The standardized mean differences for the variables used to assess group matching are provided in Supplementary Table S1.
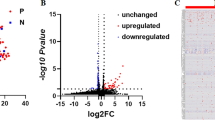
The urinary proteome was profiled using aptamer-based protein quantification (SOMAscan technology) for baseline and at the end of the run-in period of the 180 patients selected for biomarker discovery. An additional urine sample 4 weeks after conditional responders were transitioned to placebo was also included in the analysis to assess off-drug effects. Among 1275 proteins assessed, the concentration of 16 urinary protein biomarkers significantly changed during 4 weeks of treatment with atrasentan (univariate logistic model) (p < 0.01). In conditional responders, the concentrations of these 16 biomarkers reversed when participants were transitioned from atrasentan to placebo (Supplementary Fig. S2). For 4 of the 16 biomarkers, independent assays (ELISA) established passing feasibility and validity thresholds for urine protein quantification (details in “Methods” Section). Significant correlations were observed between the concentrations of urinary kallikrein B1 (uKLKB1), alpha-2-macroglobulin (uA2M), plasminogen (uPLG), and clusterin (uCLU) when measured by ELISA in the same set of samples used for the aptamer profiling, indicating consistency across the measurement platforms derived from these two technologies (Supplementary Fig. S3A). A one-unit increase in the concentration changes of these four markers was associated with a reduced likelihood of being a responder by 34% (95% confidence interval (CI) = 14–56%), 45% (95%CI = 22–62%), 36% (95%CI = 16–51%), and 33% (95%CI = 10–51%), for uKLKB1, uA2M, uPLG and uCLU, respectively (Fig. 2). This association was technically replicated by analyzing the same set of samples with ELISA assays (Supplementary Fig. S3B).
This forest plot shows the mean odds ratios (ORs) and 95% confidence intervals (CIs) for the associations between concentration changes in four urine biomarkers, kallikrein B1 (KLKB1), α2-macroglobulin (A2M), plasminogen (PLG) and clusterin (CLU), and patient treatment response. Increased biomarker concentrations from baseline to 6 weeks post-treatment is associated with reduced likelihood of being a responder (binary classified as “R” for responder or “NR” for non-responder, n = 60 for each group). Each biomarker and its OR range are depicted by horizontal lines. A dashed line at OR = 1.0 indicates no effect. ORs to the right suggest a tendency towards non-response; to the left towards response. The ORs are computed using logistic regression adjusted for age, sex, systolic and diastolic blood pressures, estimated glomerular filtration rate (eGFR), urine albumin-to-creatinine ratio (uACR), and hemoglobin A1c (HbA1C). Statistical significance, p < 0.05, was assessed using the Wald test. Mean changes for responders (R), non-responders (NR), and the overall mean change of all patients are included.
Atrasentan reversible signature in T2D-CKD mouse model
To test for a mechanistic link between disease processes reversed by atrasentan and the urinary biomarkers identified in the SONAR proteomic study, a preclinical study of atrasentan response in a mouse model for T2D-CKD was utilized. The leptin-deficient BTBR mouse strain with the ob/ob mutation (BTBR ob/ob) develops progressive weight gain, type 2 diabetes, and diabetic kidney disease that has many features of advanced human diabetic nephropathy and responds to endothelin blockade with amelioration of nephropathy5.
To establish the gene regulatory landscape of atrasentan response in BTBR ob/ob mice, differential gene expression analysis was done across RNAseq data from kidney glomerular tissues from wild type (WT, n = 5), 6 wk of saline (T2D-CKD vehicle, n = 5) or atrasentan (T2D-CKD Atra, n = 7) treated BTBR ob/ob mice. The two comparisons for this analysis were between the T2D-CKD vehicle and WT, and the T2D-CKD Atra and vehicle. A total of 1353 genes were identified as differentially expressed in both sets of comparisons after adjustment for multiple testing (q < 0.1). A total of 1068 dysregulated genes (78.9%; chi-square, p = 1.5e-100) in T2D-CKD vehicle: WT were reversed by atrasentan treatment in T2D-CKD Atra:vehicle (q < 0.1), including 818 genes up-regulated in T2D-CKD vehicle that showing down-regulation in T2D-CKD Atra, and 250 genes showed the inverse pattern (Fig. 3a). Of the 1068 reversed genes, 731 mapped to human orthologs. Kidney biopsy derived gene expression profiles from the T2D-CKD datasets from the European Renal cDNA biobank (ERCB)17,18,19 were evaluated for overlap with the expression profile reversed by atrasentan in the BTBR/obob mice. In the ERCB biopsies, 312 of the 731 genes that were reversed in the mouse model were differentially expressed between T2D-CKD and living donors, and identified as the atrasentan-reversible gene signature in humans.
a Identification of atrasentan reversed T2D-CKD-associated molecular signature(q < 0.1). b Endothelin 1 pathway is activated in T2D-CKD Vehicle and reversed by atrasentan treatment (T2D-CKD Atra). Pathway maps were derived from Ingenuity pathway analysis (IPA). The purple edge of the shapes indicates “group” or “complex” of molecules. c Atrasentan Reversible-Endothelin Activation Signature (AR-EAS) score was significantly increased in T2D-CKD Vehicle (n = 5) compared to wild type (WT, n = 5, p = 0.0317). AR-EAS score in T2D-CKD mice with atrasentan treatment (T2D-CKD Atra) (n = 7) was significantly reduced compared to T2D-CKD Vehicle, (p = 0.0025), to a level comparable to WT mice. The center line depicts the median, the box extends from the lower quartile to the upper quartile, and the whiskers indicate the minimum and maximum values. A two-sided Student’s t-test was used to compare each pair of groups. d CLU mRNA levels were significantly and highly correlated with AR-EAS score (Spearman correlation, ρ = 0.67, p = 0.0016) in experimental mice.
To define the regulatory landscape of endothelin receptor blockade conserved between murine and human CKD, we identified canonical pathways among the 312 atrasentan-reversible genes that are associated with T2D-CKD (Supplementary Table S2). Endothelin-1 signaling is among the top three significantly activated pathways along with glutathione-mediated detoxification, and TCA cycle II. Figure 3b shows the activation (T2D-CKD Vehicle) and inhibition (T2D-CKD Atra) of the endothelin-1 pathway in the mouse model.
Next, an atrasentan reversible-endothelin activation signature (AR-EAS) was established. For the AR-EAS, genes were selected if they were part of the enriched endothelin-1 pathway and were cross-validated in patients with T2D-CKD studies (Supplementary Table S3). The expression values of these selected genes were used to generate a Z-score as described previously15. The AR-EAS was generated for each mouse, and average Z-scores were compared among different treatment groups. As shown in Fig. 3c, the AR-EAS score was significantly higher in T2D-CKD Vehicle compared to WT, whereas significantly reduced in T2D-CKD Atra compared to vehicle-treated mice.
The kidney mRNA levels of the four candidate biomarkers from the human biomarker discovery proteomic study were then evaluated for their association with AR-EAS score in experimental mice. Murine Clu mRNA levels were significantly positively correlated with the AR-EAS score (rρ = 0.67, p = 0.0016) (Fig. 3d). The mRNA expression levels of Klkb1, Plg and A2m in experimental mice were low (0 or close to 0 counts for A2m and Plg, and a mean of the raw counts was less than 5 for Klkb1) and did not pass quality control, thus no correlation data with AR-EAS score is available for those three genes.
CLU in human CKD kidneys at tissue and single cell level
A significant correlation was observed only between Clu mRNA levels and AR-EAS scores in mice with T2D-CKD, coupled with the observation that the additional three biomarkers did not significantly enhance the predictive capacity provided by CLU in the SONAR proteomic discovery study, we prioritized CLU as the marker for further evaluation.
The first step in the evaluation determined the relationship between intra-kidney expression of CLU mRNA and CLU protein in urine and plasma samples from patients with CKD. uCLU/Cr and plasma CLU (pCLU) levels were quantified using a validated ELISA assay. No significant correlation between pCLU and uCLU/Cr (r = 0.19, p = 0.07) was observed (Supplementary Fig. S4a), suggesting that the increased uCLU is unlikely to be from passive glomerular filtration of pCLU. In contrast, uCLU/Cr exhibited significant correlations with CLU tissue mRNA levels in both the glomerular and tubulointerstitial compartment (r = 0.35, p = 0.008, and r = 0.44, p = 0.0001), respectively, consistent with an intrarenal source of the urinary CLU/Cr (Supplementary Fig. S4b).
In the next step, intrarenal CLU expression sources and regulation in human kidney tissue were assessed in healthy living donor biopsies and T2D-CKD. In micro-dissected glomerular and tubulointerstitial compartments of ERCB biopsies, CLU mRNA was expressed at a significantly higher level in patients with T2D-CKD, compared to healthy living donors (Fig. 4a). Furthermore, CLU mRNA was significantly correlated with eGFR (r = − 0.64, p = 0.0014, and r = − 0.81, p < 0.0001, for glomerular and tubulointerstitial CLU mRNA, respectively) (Fig. 4b), indicating higher CLU mRNA level to be associated with worse kidney function, represented by lower eGFR. Furthermore, at time of biopsy, a higher CLU mRNA level, from both glomerular and tubulointerstitial compartments of micro-dissected kidney biopsies was significantly associated with higher risk of progression to composite endpoint of end stage kidney disease (ESKD) or 40% reduction of baseline eGFR in an independent CKD biopsy cohort from the Nephrotic Syndrome Study Network (NEPTUNE, Fig. 4c, d), a longitudinal study involving people with primary proteinuric glomerular diseases20,21.
a CLU mRNA is markedly upregulated in the glomerular and tubulointerstitial compartments among T2D-CKD participants (n = 12 and 17 for glomerular and tubulointerstitial compartments, respectively) when compared to living donors (LD) (n = 21 and 31 for glomerular and tubulointerstitial compartments, respectively). A two-sided Student’s t-test was used for two-group comparison (p < 0.005). b Increased CLU mRNA in both glomerular and tubulointerstitial compartments is significantly correlated with reduced eGFR in patients with T2D-CKD (n = 22, p < 0.005). c, d Kaplan-Meier survival analyses with 5 years of follow-up data indicate that higher CLU mRNA, in either compartment, is significantly associated (p < 0.005, log-rank test) with increased risk of advancing to the composite end point of ESKD or a 40% decrease from baseline eGFR in the NEPTUNE cohort. Participants were grouped into tertiles based on CLU mRNA levels, with ranges of expression for each tertile in the glomerular and tubular compartments included in each figure legend, respectively. Abbreviations: eGFR estimated glomerular filtration rate; ESKD end stage kidney disease or kidney failure.
To examine whether there is an association between CLU and endothelin/receptor signaling, we conducted a correlation analysis between CLU and EDN1 (the gene encoding endothelin1), as well as AR-EAS, using transcriptomic data from the tubulointerstitial compartment of kidney biopsies (n = 371) from the NEPTUNE cohort. We identified a significant positive correlation between CLU and EDN1 (r = 0.70, p = 8.7e-17; Supplementary Fig. S5A). In addition, CLU mRNA showed a significant correlation with AR-EAS (r = 0.35, p < 0.0001; Supplementary Fig. S5B). To determine the cellular source of CLU mRNA expression in the kidney, single cell RNASeq (scRNAseq) data from the Kidney Precision Medicine Project (KPMP)22 were assessed for CLU expression and showed abundant CLU mRNA in epithelial, endothelial and mesenchymal cell populations, including but not limited to proximal tubular cells (PT), principal cells (PC), ascending thin limb (ATL), thick sending limb (TAL), intercalated cells (IC), endothelial cells (EC), PT/distal convoluted tubular cells (DTL), and interstitial cells (Fig. 5a). Immunohistochemistry (IHC) staining confirmed protein expression in glomerular capillary cells and distal tubular cells (Supplementary Fig. S6). We further compared the CLU mRNA expression in these cell clusters between health and disease. As illustrated by dot plot analysis in Fig. 5b, CLU is highly expressed in about 30% of ECs and over 70% of tubular cells in T2D-CKD, compared to living donors with markedly lower average intensity and detected in less than 10% of ECs and less than 50% of tubular cells. To evaluate if CLU in tubular cells expressing the CLU gene (CLU + ) is associated with clinical or histopathological phenotypes in CKD, we performed a pseudo bulk analysis in the single cell data from KPMP and examined the correlation between CLU mRNA levels in tubular cells with eGFR and tubular interstitial fibrosis (IF). We found that CLU is significantly correlated with eGFR (r = − 0.64, p = 0.03, Supplementary Fig. S7A) and %IF (r = 0.83, p = 0.001, Supplementary Fig. S7B). These results support the involvement of tubular CLU in disease-associated clinical and histopathological phenotypes.
a UMAP of single-cell RNAseq data reveals the expression of CLU across various kidney cell-type clusters from living donors (LD) and patients with T2D-CKD. b Dot plot analysis demonstrating CLU mRNA is expressed in a greater number of endothelial and tubular cells (dot size represents the percentage of cells expressing CLU) and at a higher expression level (dot intensity reflects mean expression), in patients with T2D-CKD versus LD. Statistical significance was assessed by multiple testing-adjusted P value (p < 0.05) using the FindMarkers function in Seurat. c CLU co-expressing genes in endothelial and tubular cells were identified by comparing differentially expressed genes (DEG) between the cells expressing CLU (CLU + ) versus those that did not (CLU-). The significance threshold was set at an absolute Log2 Fold Change |Log2FC| > 0.263 with FDR < 0.001. Significantly enriched functional modules were identified in CLU co-expressing genes using the Humanbase platform in (d). EC and (e) combined tubular cells. Circles represent genes within the module. Red star denotes CLU.
CLU co-expressing gene programs in kidney cells
EC cells expressing CLU greater than 0 (CLU > 0), normalized gene expression, in T2D-CKD were referred to as EC-CLU +. Differential expression analysis identified 314 genes significantly differently expressed between EC-CLU + versus ECs not expressing CLU mRNA (EC-CLU -) with a false discovery rate (FDR) < 0.001, and log2-transformed fold change either above + 0.26 or less than − 0.26 (Fig. 5c). The 314 genes were clustered into six functional modules in a kidney-relevant network analysis using Humanbase23. Using the same cut-off value, 1270 differentially expressed genes were identified between tubular cells expressing CLU (TUB-CLU + ) and those not expressing CLU (TUB-CLU-). Eight modules were identified enriched in functions that are associated with various kidney functions (Fig. 5d). It is worth noting that in both EC cells and tubular cells, CLU is mapped into the functional module that is involved in cardiovascular system development, vasculature development, and regulation of cell/endothelial cell migration (M2 in EC cell modules and M6 in tubular cell modules, red star denotes CLU) (Fig. 5e).
Validation of uCLU – association with kidney outcomes
To validate the association of change in uCLU/Cr with atrasentan treatment in a larger patient group, we used urine samples from baseline and the end of the active run-in period from SONAR trial participants, outside of the discovery nested case-control study. Baseline clinical characteristics of the 3060 included participants in the validation study were similar to the overall SONAR population (Table 1). Mean eGFR was 43.3 (Standard deviation [SD] 14) mL/min/1.73 m2, Median uACR was 810 (25th to 75th Percentile 450, 1519) mg/g, and median uCLU/Cr was 495 [25th 75th P 215, 1394] ng/mg.
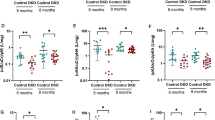
Participants were followed for a median duration of 2.1 (25th to 75th percentile: 1.2 to 2.9) years, during which 270 (8.8%) participants experienced a composite kidney endpoint. In multivariable analyses, adjusting for participant demographic, physical, and biochemical characteristics, including history of cardiovascular disease and antihypertensive medications, demonstrated that each doubling in uCLU/Cr was statistically significantly associated with an increased risk of the composite kidney endpoint with an adjusted hazard ratio (HR) of 1.10 (95% CI 1.03, 1.17, p = 0.004) (Fig. 6a/d). When uCLU/Cr was modeled as a categorical variable, the relative risk of the kidney outcome was significantly higher in the highest vs lowest quartile of (Fig. 6a).
This analysis included 3060 patients from the SONAR trial with available samples at the start and end of the 6-weeks active run-in period. a Association between baseline uCLU/Cr and clinical kidney outcomes divided in quintiles and analyzed on a continuous scale. which shows the hazard ratio associated with each increment in uCLU/Cr. Model 1 was adjusted for age, sex and race/ethnicity. Model 2 was additionally adjusted for systolic blood pressure, GFR, (log)uACR, HbA1c, body weight, hemoglobin, history of cardiovascular disease, antihypertensive medications, and treatment allocation. b Change in uCLU/Cr during 6 weeks treatment with atrasentan. Geometric means are shown, with error bars representing the 95% confidence interval. Shown are the baseline and week 6 uCLU/Cr concentrations and the percentage change from baseline. c Association between the change from baseline in urinary uCLU/Cr during 6 weeks atrasentan treatment and subsequent clinical kidney outcomes, divided in quintiles and analyzed on a continuous scale. Model 1 was adjusted for age, sex and race/ethnicity. Model 2 was additionally adjusted for baseline log transformed uCLU/Cr, systolic blood pressure, estimated glomerular filtration rate, (log)uACR, HbA1c, body weight, hemoglobin, history of cardiovascular disease, antihypertensive medications, and treatment allocation. d, e Associations of baseline uCLU/Cr (d) and change in uCLU/Cr (e) with clinical kidney outcomes analyzed on a continuous scale. The red line represents the hazard ratio at different levels of clusterin creatinine ratio at baseline, and the light pink area represents the corresponding 95% confidence interval. f Hazard ratio of the clinical outcomes by changes in both uCLU/Cr and uACR.
Six weeks treatment with atrasentan 0.75 mg/day was associated with a reduction in geometric mean of uCLU/Cr from 556 (SD 5.6) ng/mg to 320 (SD 5.7) ng/mg, corresponding to a geometric mean percentage change of -42.6% (95%CI 39.4, 45.4; Fig. 6b). After 6 weeks atrasentan treatment, 2260 (73.9%) participants experienced a reduction in uCLU/Cr from baseline. The change in uCLU/Cr during six weeks treatment with atrasentan was associated with the composite endpoint during subsequent follow-up. In multivariable analysis after adjustment for all covariates, each halving of uCLU/Cr was independently associated with a 10% lower risk of the composite kidney endpoint (HR 0.90 (95% CI 0.84, 0.98; p = 0.009) (Fig. 6c/e). When delta uCLU/Cr was modeled as a categorical variable, relative to the third quintile, the first and second quintiles were associated with a lower risk of the composite kidney endpoint in multivariable adjusted analyses, while the fourth and fifth quintiles were associated with a higher risk (Fig. 6c). Since atrasentan reduces uACR and early changes in uACR during treatment with atrasentan are associated with the composite kidney endpoint, we finally analyzed the correlations between changes in uACR and uCLU/Cr. Pearson correlation coefficient showed a weak correlation between change in (log2)uACR and (log2)uCLU/Cr (r = 0.19; p = < 0.001). The risk of attaining the composite kidney endpoint showed consistency across all uACR and uCLU/Cr categories (Fig. 6f). Specifically, early reduction in uCLU/Cr was associated with a lower risk of the composite kidney endpoint regardless of the uACR change, with the lowest risk - observed in participants with a reduction in both uACR and uCLU/Cr, and the highest risk observed in those with an increase in both parameters.
Discussion
The selective endothelin receptor type antagonist atrasentan reduces the relative risk of clinical kidney outcomes in patients with T2D-CKD. However, like many other nephroprotective drugs, a large between-individual variation in the response to atrasentan exists. Currently, there is a lack of quantitative, evidence-based methods to distinguish between patients who will or will not benefit from atrasentan. To address this unmet need, we employed an integrative approach that identified mechanism-based pharmacodynamic biomarker candidates whose short-term changes can predict patient’s longitudinal outcome during atrasentan treatment. uCLU emerged as a top candidate biomarker, demonstrating a strong association with kidney disease progression and value for identifying patients in whom the benefit of atrasentan may be more pronounced.
Our integrative approach, by leveraging kidney tissue transcriptomic data from both humans and atrasentan-treated mouse models of T2D-CKD, emphasizes biomarkers linked to underlying endothelin biology and response to atrasentan. This connection to the targeted disease mechanism improves on traditional approaches used to identify biomarkers where, typically, the focus is primarily on associations between variables and outcomes.
Using this strategy, we identified and validated uCLU as a biomarker associated with the risk of kidney failure independent of other clinical characteristics, suggesting that uCLU is involved in the pathophysiology of CKD in T2D-CKD. Moreover, early changes in uCLU level after initiation of atrasentan treatment was also associated with the risk of the composite kidney endpoint. This association was independent of other patient characteristics and changes in albuminuria. These findings, together with the association we show between intra-kidney CLU mRNA levels with and endothelin 1 pathway activation, suggest that early changes in uCLU during treatment with atrasentan may serve as pharmacodynamic response marker to monitor individual’s response to atrasentan treatment.
Clusterin, also known as apolipoprotein J, is a glycosylated protein involved in both apoptotic and antiapoptotic pathways and is found across tissues including the kidney24. It is thought to be involved in cardiovascular- related diseases, including dyslipidemia, atherosclerosis, obesity, and type 2 diabetes25,26,27. Increased clusterin expression in the kidney tubular compartment is associated with tissue injury and considered to be a stress-activated injury-response protein. Clusterin also serves a protective function through suppressing inflammation or scavenging toxic lipid byproducts and denatured proteins, or restoring mitochondrial function via CLUlow density lipoprotein receptor-related protein 2 (LRP2)- activation calcium/calmodulin dependent protein kinase 1D signaling in proximal tubular cells28,29,30 The urinary excretion of clusterin is believed to be derived from kidney tissues rather than filtration of plasma clusterin31. Clusterin is thus thought to be a specific marker of damage to tubular cells32, supported by prior studies demonstrating association of uCLU with nephrotoxicity in animals33,34 and in humans35,36,37. Specifically, uCLU was better at detecting proximal tubular injury compared to other nephrotoxicity markers including CysC, B2M and total protein38. uCLU has also been suggested for use as an early tissue damage marker with a similar profile to KIM-134,39. It is therefore perhaps not surprising that regulatory agencies qualified uCLU, along with five other biomarkers, to be used in non-clinical and clinical drug development to aid in the detection of drug-induced kidney tubular injury in phase 1 trials in healthy volunteers (https://www.fda.gov/media/115635/download).
Although uCLU is used in the setting of acute kidney damage, little is known about its association with kidney disease progression in CKD. One study in 159 patients with type 2 diabetes and GFR over 60 mL/min/1.73 m2 reported that increased uCLU was associated with progression to CKD stage 3 and above27. The limited sample size and lack of clinical kidney endpoints, warrants validation in a contemporary well-phenotyped population. Our study offers evidence for association of uCLU with CKD progression, more importantly, adds further insight that the relationship between uCLU and clinical outcomes are likely primarily mediated through tissue clusterin: firstly, we demonstrated that CLU mRNA is abundantly expressed in kidney and it’s expression is up-regulated in CKD compared to healthy individuals, at both tissue compartment and single cell level; secondly, uCLU is significantly correlated with kidney tissue CLU mRNA but not with pCLU, suggesting that uCLU is not just a passive filtrate from the systemic circulation, but rather is kidney tissue derived and as a result may reflect clusterin-associated CKD pathology; thirdly, the significant association between intrarenal tissue CLU mRNA at the time of biopsy and composite kidney endpoints in the NEPTUNE cohort supports the observed associations between baseline uCLU and risk of the composite kidney endpoint in the SONAR trial.
The mechanism by which early changes in uCLU concentration during atrasentan treatment are associated with long-term outcomes remains incompletely understood. Although endothelin-1 is known for its anti-apoptotic role40,41,42 and clusterin can exert context-dependent pro- or anti-apoptotic effects30,43,44,45, our study did not find significant enrichment of apoptosis-related pathways in the canonical pathway analysis of differentially expressed genes between CLU⁺ and CLU⁻ cells. Our findings suggest that, in the context of CKD, the primary functional impact of clusterin may involve processes such as cell adhesion, epithelial/endothelial-mesenchymal transition, fibrosis, and atherosclerosis. The significant correlation between CLU mRNA levels in pseudobulk tubular cells and patients’ interstitial fibrosis scores supports the aforementioned biological process. While apoptosis remains an important cellular process that clusterin may be associated with, the evidence from our study supporting this association is currently limited.
However, it is worth noting that our animal studies data and single-cell and bulk RNASeq analysis of human kidney biopsies provide further insight into the potential link between CLU and EDN1 signaling pathway. In atrasentan-treated BTBR ob/ob mice, clusterin mRNA levels are significantly correlated with the AR-EAS score, a transcriptomic measure based on involving endothelin pathway genes reversed by atrasentan (Fig. 3d), indicating a link between CLU RNA levels and the status of endothelin signaling. In human CKD kidney biopsies, we also observed a significant strong positive correlation between CLU and EDN1 mRNAs and AR-EAS score. Furthermore, integrating single-cell analysis with kidney-specific molecular network analysis revealed that clusterin is a component of a functional module involved in cardiovascular system development, vasculature development, and regulation of cell/endothelial cell migration. These pathophysiological processes are well known to be significantly impacted by endothelin 1 signaling pathways46,47,48,49. These findings support an association between clusterin and endothelin 1 signaling in the context of pathogenesis of cardio-kidney damage, including vasculature development and damage as well as regulation of endothelial cell migration. While these results are promising, further bioinformatic analysis, such as ligand-receptor analysis, and in vitro and in vivo studies are required to further elucidate the mechanisms underlying these strong associations. Investigation into whether atrasentan reduces clusterin expression in human kidneys and whether this reduction is associated with the alleviation of kidney injury will also help to gain better insight into this question. Ongoing or future biopsy studies involving endothelin receptor antagonists, such as the SPARTAN study (NCT04663204), could contribute a more comprehensive understanding of the mechanistic link.
Although the focus of this paper is on uCLU, concentrating on the kidney-expressed clusterin and its associated processes and functions, clusterin is ubiquitously expressed in human tissues and organs, such as adipocytes, liver, lung, whole brain and heart (data from BioGPS). Dysregulation of clusterin, in particular plasma- and adipocyte-derived circulating clusterin, have been implicated in various diseases, including cardiometabolic and cardiovascular disease26,50,51,52. In our hands, pCLU does not correlate with uCLU, supporting the hypothesis that uCLU is not simply the result of passive filtration of pCLU; instead, a marker likely reflecting intrarenal biological processes.
In addition to uCLU, urinary A2M, PLG, and KLKB1 also demonstrated significant associations with atrasentan response in our initial biomarker discovery phase. Due to the lack of significant additive value of these markers when added, either one-by-one or together, to the model including clinical variables and clusterin, and their absence in cross-species transcriptomic support in the mouse model, these other three markers were not included in the validation in the large SONAR cohort. However, their association with response to atrasentan in the discovery study suggests potential for future investigation. A2M as a nonspecific protease inhibitor, is likely involved in increased glomerular matrix accumulation in kidney disease through inhibiting matrix-degrading proteases53. A2M mRNA expression in the kidney demonstrated an endothelial cell-specific and disease outcome-associated pattern in patients with glomerular disease54. uA2M has been considered as a predictor of severe loss of glomerular permselectivity55. Atrasentan treatment significantly reduced uA2M maybe an early indication of improved glomerular permselectivity and/or the reduced matrix accumulation. The glycoprotein zymogen PLG contributes to renal interstitial fibrosis via promoting epithelial-to-mesenchymal transition56. Baseline uPLG was reported to be independently associated with increased risk of ESKD in a cohort of 1010 patients with glomerular disease57. Future studies are needed to evaluate the utility of these individual markers - or their various combinations as a pharmacodynamic biomarker panel - in the SONAR trial or other independent cohorts, taking into account model performance, practicality, and cost-effectiveness.
The results of this study are clinically relevant in view of the development and registration of new ERAs for the treatment of CKD. Sparsentan, a dual endothelin angiotensin receptor blocker, reduced eGFR decline in patients with immunoglobulin A (IgA) nephropathy58 and is registered for the treatment of IgA nephropathy. Similarly, atrasentan reduced proteinuria in 270 patients with IgA nephropathy in the Phase III ALIGN study (NCT04573478)59. Another selective ERA, zibotentan, is currently in development as a fixed-dose combination with the SGLT2 inhibitor dapagliflozin for high-risk patients with CKD60. Validation of uCLU as a biomarker of risk and treatment response will contribute to the use and clinical implementation of clusterin.
While the strengths of this study are the multi-omics translational approach to discover and validate a pharmacodynamic biomarker to monitor treatment response to atrasentan, and the use of samples from a large randomized controlled clinical trial with clinical endpoints, this study has a few limitations. Firstly, kidney tissue samples from patients treated with atrasentan are not yet available. This precluded direct clinical confirmation of the association between changes of CLU mRNA and change of uCLU during atrasentan treatment, as well as their association with longitudinal outcomes. Consequently, no causal or mediating role can be inferred from our current study. Furthermore, we used transcriptomic data of patients from a CKD cohort (NEPTUNE), to assess the association between tissue CLU mRNA and kidney longitudinal outcomes. It is important to note that few participants in the NEPTUNE cohort have diabetes. We selected the NEPTUNE dataset in part because the available T2D-CKD cohorts lacked sufficient sample size and transcriptomic data to support robust survival analyses for evaluating long-term outcomes. Importantly, analyzing the NEPTUNE cohort allowed us to examine whether the association between clusterin and disease progression extends beyond T2D-CKD to a broader spectrum of CKD. This is particularly relevant, as these diseases often share underlying mechanisms of progression, thereby supporting the potential generalizability of our findings. Finally, participants of the SONAR trial were carefully selected patients with T2D-CKD who did not have heart failure and peripheral edema, which may limit the generalizability of our findings to the broader CKD population. In our future work we intend to include additional analyses and clinical studies utilizing kidney biopsies and matching urine samples from patients treated with atrasentan to determine whether clusterin mediates disease progression and the response to ERA treatment
In conclusion, using an integrative approach we identified and validated uCLU as a promising pharmacodynamic biomarker of atrasentan treatment response and kidney disease progression, independent of uACR and other baseline clinical variables. This finding warrants further clinical validation in well-designed, independent studies with larger sample sizes and greater event numbers, and assessment for future clinical implementation. Collectively these efforts may pave the way for precision medicine in kidney diseases.
Methods
Ethics approvals
The protocol of the SONAR trial was approved by all relevant ethics committees and regulatory bodies. All SONAR participants provided informed consent before any study-specific procedure commenced. The biomarker studies described here were covered by the original ethics committee approvals and consent process.
The mouse experimental protocol was reviewed and approved by the Animal Care Committee of the University of Washington in Seattle.
Overall design
The analytical workflow of this study (Fig. 1) initiated with an advanced proteomic analysis of a nested case-control study to identify candidate biomarkers of a patient’s response to atrasentan. Following the biomarker discovery phase, selected biomarkers were subjected to technical validation via ELISA assays applied to the same bio-samples used for biomarker discovery. To explore the biological significance of the biomarkers and verify that the candidate biomarkers matched disease pathophysiology with atrasentan’s mechanism of action, transcriptomic analyses were carried out using an atrasentan treated diabetic mouse model for progressive CKD and using biopsy samples from patients with T2D-CKD, to demonstrate the association between gene encoding the candidate biomarker and kidney disease severity and progression. Additionally, these studies also aimed to establish a quantitative link between the candidate biomarker and a gene signature representing atrasentan reversed endothelin activation signature. This multifaceted approach identified a premier biomarker candidate, which subsequently underwent further validation in a large prospective phase 3 clinical trial with atrasentan (SONAR Clinicaltrials.gov ID: NCT01858532).
Data sources
Animal studies
Kidney tissues from previously described experiments with BTBR ob/ob mice were used5. Briefly, female BTBR WT, BTBR/ob heterozygotes [BTBRob+/−; BTBR.V(B6)-Lepob/WiscJ; stock no. 004824]) mice from Jackson Laboratories (Bar Harbor, ME) maintained under 12 h light cycle with free access to standard diet (standard chow dietPicolab Rodent Diet 20, Brentwood, MO) and water were either treated with saline (n = 5) or atrasentan 5 mg/kg body weight per day (n = 7) in drinking water. Wild type mice (n = 5) were included as control. Female BTBR ob/ob mice are more resilient than males and are less prone to urine scalds and poorly healing skin abrasions or wounds, leading to risk of premature sacrifice, therefore, only female mice were included in this study. Atrasentan or saline (vehicle) treatment was initiated when animals were 18 weeks of age and continued for 6 weeks until 24 weeks of age. The changes in phenotypic characteristics across treatment groups were published previously5.
Clinical studies
The SONAR trial recruited 5,117 patients aged 18 to 85 years with a diagnosis of type 2 diabetes, eGFR between 25 and 75 mL/min/1.73 m2, and uACR between 300 and 5,000 mg/g (> 33.9-565.6 mg/mmol). All eligible participants were using an ACE-inhibitor or ARB and received atrasentan 0.75 mg during a 6-week open-label active run-in period to select patients likely to respond to atrasentan, defined as reduction from baseline in uACR of 30% or more. All responder patients who tolerated atrasentan (N = 2648) and a selection of non-responder patients (N = 1020) proceeded to the randomization visit and were assigned in a 1:1 ratio to continue atrasentan or to transition to placebo. The composite kidney endpoint of the trial was a composite of doubling of serum creatinine, kidney failure or death due to kidney failure.
To identify biomarkers whose change from baseline to 6 weeks atrasentan treatment associated with patients’ long-term response, we conducted a nested case-control study of the SONAR trial (Supplementary Fig. S1). The remaining 3060 patients (83.4% of total cohort) from the SONAR trial with available samples at start and end of the 6-weeks active run-in period were used for biomarker validation purposes.
Transcriptomic data from healthy controls and people with kidney diseases
Transcriptomic data from micro-dissected glomerular and tubulointerstitial compartment of patients with kidney disease were obtained from European Renal cDNA Biobank18, Nephrotic Syndrome Study Network (NEPTUNE)14 and Nephroseq (https://www.nephroseq.org). Single cell RNASeq data was derived from Kidney Precision Medicine project (KPMP)22 (https://www.kpmp.org/). Details are provided in “Data Availability Section”.
Measurements
Animal studies
Total RNA was isolated from mouse glomeruli enriched cortex tissue using RNeasy isolation kits (Qiagen). RNA quantity and integrity was confirmed on an Agilent 2100 bioanalyzer. RNA sequencing was performed on oligo(dT) purified poly(A)+ mRNA. A standard TruSeq mRNA library was constructed using TruSeq RNA Sample Prep Kit v2 (Illumina, Cat.# RS-122-2001). The library was sequenced by Illumina HiSeq 4000 using a single-end run (SE50 bases). cDNA, NGS library prep and sequencing prep for an Illumina platform were generated following standard protocols. QC on the NGS library prep was done using the Agilent TapeStation. More details can be found in the supplementary information.
High throughput biomarker profiling
Urine samples at baseline and after 6 weeks atrasentan treatment from 180 patients participating in the SONAR trial were used. Of these, in 60 patients, who were randomized to placebo after 6 weeks atrasentan treatment, additional urine samples were used after 4 weeks placebo treatment to assess off-drug effects. The SOMAscan 1.3 K was for biomarker identification. The SOMAscan protein array platform renders 1317 protein analytes for human protein profiling. Urine samples were processed according to the SomaLogic’s recommended standard protocols61,62 for the SOMAscan Assay Human Cell & Tissue kit (CLTH_4.2_20161012_1.5k). Details of measurement and normalization were included in the supplementary Information. SOMAscan log2 transformed expression values were derived from quantile-normalized data files. The expression value for each individual analyte was normalized by the urine creatinine value of the same sample to correct for hydration status.
ELISA protein assay
Commercially available ELISA assays were used for biomarker technical validation. ELISA assays went through assay feasibility evaluation (standard curve parallelism, dynamic concentration range, endogenous levels in healthy versus disease, dilution linearity, selection of QCs, and optimized condition for different biofluid matrices) and assay validation (upper & lower limit of quantification, precision & accuracy, matrix effects, analyte stability, and selectivity & specificity). Assays passed feasibility and assay validation evaluation were selected for further analysis. Concentrations of alpha-2-macroglobulin (A2M) (R&D DY1938, Minneapolis, MN, USA), clusterin (CLU) (Abcam,ab174447, Waltham, MA, USA), plasminogen (PLG) (Abcam, ab108893, Waltham, MA, USA) and kallikrein (KLKB1) (Abcam, ab202405, Waltham, MA, USA) were measured in urine samples as per standard or modified standard protocol at 1:10, 1:32, 1:32, and 1:6 dilutions respectively. Quality control samples with high, medium and low concentrations of the respective analytes were included in each ELISA plate to monitor batch effect. The inter-assay coefficient of variation of the quality controls for the above-mentioned biomarkers was 11%, 3.0%, 3.4%, and 2.6%, respectively. The concentrations of biomarkers were normalized by urine creatinine to correct for hydration status, and the log2 transformed ratio was used for subsequent analysis.
Data Analysis
Animal and human transcriptomic data analysis
Determination of reversal gene signature by atrasentan
Normalization and log-transformation of the raw counts was performed in R using the calcNormFactors function in the edgeR package63 and voom64 in the limma package. Differential expression was subsequently fit to linear models using the lmFit and eBayes functions in limma. To identify differentially expressed (DE) genes in the diabetic mouse model (with respect to WT mice) that were subject to reversal upon treatment with atrasentan, differential expression (DE) was identified using R with the limma package for two contrasts: the vehicle-treated BTBR ob/ob mice vs. wild-type (T2D-CKD vehicle: WT), and the atrasentan-treated vs. vehicle-treated ob/ob mice (T2D-CKD atra:vehicle). With the T2D-CKD vehicle group in common between the two contrasts, it was possible to directly query the effect of atrasentan treatment on the diabetic phenotype at the level of gene expression.
Molecular feature annotation and mapping strategy
The Ensembl human Gene ID was defined as a common denominator for molecular feature annotation for Omics experiments65. Version 92 of the Ensembl reference was used to map Ensembl Gene IDs to official human Gene Symbols. Pairwise human-mouse orthologs were retrieved from the OMA database (https://omabrowser.org)66. Only genes having a 1:1 match were kept. Mouse genes without respective human orthologs were excluded from further analyses.
Cross -species integration with transcriptomics data for patient with T2D-CKD
Dysregulated genes in BTBR ob/ob mice (T2D-CKD Vehicle) versus BTBR WT mice and reversed by atrasentan treatment (T2D-CKD Atra, q < 0.1, Benjamini-Hochberg method) were identified. They were then mapped to the human ortholog. The expression of the human ortholog genes in human ERCB T2D-CKD datasets17,67 were evaluated by comparing living donor and patients with T2D-CKD. Genes concordantly regulated in patients with T2D-CKD and the BTBR ob/ob mouse model were used for further analysis.
Pathway enrichment analysis and kidney relevant functional network analysis
Ingenuity Pathway Analysis software (IPA, QIAGEN Inc.) was used to identify enriched canonical pathways68. To determine the significantly enriched functional gene modules in the differentially expressed gene sets, we projected gene signatures into the HumanBase functional network (https://humanbase.io/module/), in which gene signatures were clustered using community clustering in higher-order tissue-specific functional network, to identify tightly connected sets and pathways in the context of kidney69.
Generation of the atrasentan reversible-endothelin activation signature
Genes mapped to enriched Endothelin-1 pathways by Ingenuity Pathway Analysis (IPA) software was extracted. The expression values of cross-validated genes were used to generate a Z-score, using a previously described method15, referred as Atrasentan reversible-endothelin activation signature (AR-EAS) for each mouse and average Z-scores were compared among different groups of treatment.
Identification of CLU co-expressing gene signatures
ScRNA-seq analysis was performed using the cortex region of the kidney biopsy and processed according to the Kidney Precision Medicine Project single-cell protocol22,54. CLU co-regulated genes were identified using similar method previously published13. In brief, genes that were differentially expressed in CLU-expressing (CLU + ) versus CLU-no expressing (CLU–) cells from patients with T2D-CKD were identified using the FindMarkers Seurat function70,71,72,73. CLU + versus CLU– cells were based on greater than 0 (CLU > 0) normalized gene expression. To identify the differential gene signature, comparisons between groups were made using the Welch’s t test. Only genes that exhibited an FDR (false discovery rate) of less than 0.001 and also demonstrated an absolute log2 fold change greater than 0.26 were selected for subsequent analysis. We performed psuedobulk analysis first on the tubular cells expressing CLU mRNA (aggregating the single-cell expression by sample at the cell type level to mimic the bulk RNASeq profiles) and then correlated with the phenotype74,75.
Discovery of biomarkers
The case-control design included three groups of patients who were selected based on the uACR change during the 6-weeks atrasentan treatment period and the rate of eGFR decline during median 2.2 year double blind treatment with atrasentan: We defined a “responder” group, (R), defined as patients whose uACR reduction was reduced at least 45% during 6 weeks treatment with atrasentan and whose eGFR slope during a median 2.2 year treatment with atrasentan was stable between − 1.0 to + 1.0 mL/min/1.73 m2 per year; A “non-responder” group, (NR), defined as patients whose uACR reduction was less than 15% during 6-weeks atrasentan and whose eGFR decline during median 2.2 year atrasentan treatment was more than 3 mL/min/1.73 m2 per year; A “conditional responder” group (CR),defined as patients whose reduction in uACR was at least 45% during 6-weeks atrasentan treatment and who transitioned to placebo at randomization and whose eGFR decline after transition to placebo was more than 3.0 mL/min/1.73 m2 per year. Differences in biomarker changes from baseline to week 6 among the three groups were analyzed for marker identification. Three groups of patients were selected by propensity matching using the following baseline demographic and clinical characteristics: age, gender, systolic blood pressure, hemoglobin, eGFR and uACR. Standardized differences among groups were computed to determine that the groups were adequately matched. Differences in biomarker concentrations were compared between responders and non-responders. Univariable logistic regression analyses were subsequently performed to identify biomarkers whose change from baseline to week 6 were significantly associated with atrasentan response comparing responders and non-responders. Urinary protein markers with logistic regression associations of p < 0.01, a threshold selected as a trade-off between sensitivity and specificity, ensuring the inclusion of a sufficient number of candidate proteins (for downstream multi-step validation) while reducing the likelihood of false positives, were then further restricted to those showing a reversibility in the direction of the effect during the off-treatment phase in the conditional responders. This selection was performed to ensure that the selected biomarkers reflected a true pharmacological treatment response. Candidate biomarkers were then further selected based on commercial ELISA assay availability, and assay performance evaluation based on feasibility and validity in measuring urine samples, as described above.
Validation of biomarker
In the validation cohort, we assessed the association between baseline log-transformed urinary CLU:creatinine ratio (uCLU/Cr) with the primary kidney outcome using Cox proportional hazard regression. uCLU/Cr was log-transformed to take into account its skewed distribution. uCLU/Cr was entered in the Cox models as quintiles, using the third quartile as common reference for the other quartiles and as a continuous variable. To examine the association, two models were constructed. Model 1 was adjusted for age, sex and race/ethnicity. In Model 2, we additionally adjusted for systolic blood pressure, eGFR, (log)uACR, HbA1c, body weight, hemoglobin, history of cardiovascular disease, antihypertensive medications, and treatment allocation. We calculated the geometric mean change from baseline in uCLU/Cr during 6 weeks treatment with atrasentan and tested the statistical significance using a one sample t test. uCLU/Cr was subsequently entered in a Cox proportional hazard model to assess association between the change in uCLU/Cr and primary kidney outcome. Change from baseline in uCLU/Cr was categorized in quintiles with the third quintile being used as a common reference for the other quintiles. We also entered change from baseline in uCLU/Cr as a continuous variable in the Cox proportional hazard model. All aforementioned models additionally adjusted for (log)baseline uCLU/Cr were used for assessing the association between change in uCLU/Cr and primary kidney outcome. A sample size of 3060 provided 91.2% power to detect a multivariable adjusted HR for the association between changes in uCLU/Cr and the primary kidney outcome of 0.90 assuming standard deviation of the difference in uCLU/Cr between the two visits of 2.15, square of the correlation (i.e., R2) between the covariate of interest and other covariates in the model of 0.3, probability of event of interest of 10%, and p-value of 0.0576. Power analysis was conducted using command “power cox” in Stata 18.0. Cox proportional hazard regression analyses were finally performed to assess the effect of atrasentan compared to placebo in reducing the risk of the composite kidney endpoint in subgroups stratified by the median change in uCLU/Cr during 6-weeks open label atrasentan treatment. Consistent with the primary analysis model of the SONAR trial, the treatment effect in the Cox model was adjusted for log-transformed uACR values, serum albumin, age, and eGFR at randomization. A p-value for interaction was calculated by adding an interaction term between treatment and uCLU/Cr strata to the Cox model.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The transcriptomic data supporting the findings of this study are available in the public domain in Nephroseq (https://www.nephroseq.org) and the Gene Expression Omnibus (GEO) repository under the following: GSE104954, GSE104948, GSE108112, GSE182380, GSE104066, GSE133288, GSE197307, and GSE30122, and can be downloaded. The transcriptomic data generated from the mouse model experiments are available under GSE307122. The clinical datasets used for the SONAR patients in the current study are not publicly available owing to patient confidentiality considerations and compliance with institutional privacy regulations. This clinical trial data can be requested by any qualified researchers who engage in rigorous, independent scientific research, and will be provided following review and approval of a research proposal and Statistical Analysis Plan (SAP) and execution of a Data Sharing Agreement (DSA) by the corresponding author, Dr. Hiddo J.L. Heerspink. Data requests can be submitted at any time and will be processed as soon as possible. The data will be accessible for 12 months, with possible extensions considered. Restrictions imposed on data use will be described in the applicable data use agreement. All other data resulting from this study are available in the main text or in the Supplementary Materials. Source data are provided as a Source Data file. Source data are provided in this paper.
References
Kohan, D. E. & Barton, M. Endothelin and endothelin antagonists in chronic kidney disease. Kidney Int. 86, 896–904 (2014).
Helal, I., Fick-Brosnahan, G. M., Reed-Gitomer, B. & Schrier, R. W. Glomerular hyperfiltration: definitions, mechanisms and clinical implications. Nat. Rev. Nephrol. 8, 293–300 (2012).
Yang, Y. & Xu, G. Update on Pathogenesis of Glomerular Hyperfiltration in Early Diabetic Kidney Disease. Front. Endocrinol. 13, 872918 (2022).
Martínez-Díaz, I. et al. Endothelin Receptor Antagonists in Kidney Disease. Int. J. Mol. Sci. 24, https://doi.org/10.3390/ijms24043427 (2023).
Hudkins, K. L., Wietecha, T. A., Steegh, F. & Alpers, C. E. Beneficial effect on podocyte number in experimental diabetic nephropathy resulting from combined atrasentan and RAAS inhibition therapy. Am. J. Physiol. Renal Physiol. 318, F1295–f1305 (2020).
Egido, J. et al. Atrasentan for the treatment of diabetic nephropathy. Expert Opin. Investig. Drugs 26, 741–750 (2017).
Andress, D. L. et al. Clinical efficacy of the selective endothelin A receptor antagonist, atrasentan, in patients with diabetes and chronic kidney disease (CKD). Life Sci. 91, 739–742 (2012).
de Zeeuw, D. et al. The endothelin antagonist atrasentan lowers residual albuminuria in patients with type 2 diabetic nephropathy. J. Am. Soc. Nephrol. 25, 1083–1093 (2014).
Heerspink, H. J. L. et al. Atrasentan and renal events in patients with type 2 diabetes and chronic kidney disease (SONAR): a double-blind, randomised, placebo-controlled trial. Lancet 393, 1937–1947 (2019).
Waijer, S. W. et al. The effect of atrasentan on kidney and heart failure outcomes by baseline Albuminuria and kidney function: a post Hoc analysis of the SONAR randomized trial. Clin. J. Am. Soc. Nephrol. 16, 1824–1832 (2021).
Koomen, J. V. et al. Individual Atrasentan Exposure is Associated With Long-term Kidney and Heart Failure Outcomes in Patients With Type 2 Diabetes and Chronic Kidney Disease. Clin. Pharmacol. Ther. 109, 1631–1638 (2021).
Heerspink, H. J. L. et al. Rationale and protocol of the Study Of diabetic Nephropathy with AtRasentan (SONAR) trial: A clinical trial design novel to diabetic nephropathy. Diabetes Obes. Metab. 20, 1369–1376 (2018).
Sen, T. et al. Sodium glucose co-transporter 2 inhibition increases epidermal growth factor expression and improves outcomes in patients with type 2 diabetes. Kidney Int. 104, 828–839 (2023).
Mariani, L. H. et al. Precision nephrology identified tumor necrosis factor activation variability in minimal change disease and focal segmental glomerulosclerosis. Kidney In. 103, 565–579 (2023).
Liu, J. et al. Multi-Scalar data integration links glomerular angiopoietin-tie signaling pathway activation with progression of diabetic kidney Disease. Diabetes 71, 2664–2676 (2022).
Limonte, C. P. et al. Urinary proteomics identifies cathepsin D as a biomarker of rapid eGFR decline in type 1 diabetes. Diabetes Care 45, 1416–1427 (2022).
Ju, W. et al. Defining cell-type specificity at the transcriptional level in human disease. Genome Res. 23, 1862–1873 (2013).
Ju, W. et al. Tissue transcriptome-driven identification of epidermal growth factor as a chronic kidney disease biomarker. Sci. Transl. Med. 7, 316ra193 (2015).
Yasuda, Y., Cohen, C. D., Henger, A. & Kretzler, M. Gene expression profiling analysis in nephrology: towards molecular definition of renal disease. Clin. Exp. Nephrol. 10, 91–98 (2006).
Gadegbeku, C. A. et al. Design of the Nephrotic Syndrome Study Network (NEPTUNE) to evaluate primary glomerular nephropathy by a multidisciplinary approach. Kidney Int. 83, 749–756 (2013).
Hodgin, J. B. et al. Quantification of glomerular structural lesions: associations with clinical outcomes and transcriptomic profiles in nephrotic syndrome. Am. J. Kidney Dis. 79, 807–819 (2022).
Lake, B. B. et al. An atlas of healthy and injured cell states and niches in the human kidney. Nature 619, 585–594 (2023).
Greene, C. S. et al. Understanding multicellular function and disease with human tissue-specific networks. Nat. Genet. 47, 569–576 (2015).
Park, J. W. et al. Clusterin is highly expressed in tubular complexes during spontaneous pancreatitis of spontaneous hypertensive rats. J. Vet. Med. Sci. 80, 1553–1557 (2018).
Park, S., Mathis, K. W. & Lee, I. K. The physiological roles of apolipoprotein J/clusterin in metabolic and cardiovascular diseases. Rev. Endocr. Metab. Disord. 15, 45–53 (2014).
Won, J. C. et al. Plasma clusterin (ApoJ) levels are associated with adiposity and systemic inflammation. PLoS ONE 9, e103351 (2014).
Kim, S. S. et al. Urine clusterin/apolipoprotein J is linked to tubular damage and renal outcomes in patients with type 2 diabetes mellitus. Clin. Endocrinol. 87, 156–164 (2017).
Weng, X. et al. Clusterin regulates macrophage expansion, polarization and phagocytic activity in response to inflammation in the kidneys. Immunol. Cell Biol. 99, 274–287 (2021).
Girton, R. A., Sundin, D. P. & Rosenberg, M. E. Clusterin protects renal tubular epithelial cells from gentamicin-mediated cytotoxicity. Am. J. Physiol. Renal Physiol. 282, F703–F709 (2002).
Gujarati, N. A. et al. Podocyte-specific KLF6 primes proximal tubule CaMK1D signaling to attenuate diabetic kidney disease. Nat. Commun. 15, 8038 (2024).
van Meer, L., Moerland, M., Cohen, A. F. & Burggraaf, J. Urinary kidney biomarkers for early detection of nephrotoxicity in clinical drug development. Br. J. Clin. Pharmacol. 77, 947–957 (2014).
Rosenberg, M. E. & Silkensen, J. Clusterin: Physiologic and pathophysiologic considerations. Int. J. Biochem. Cell Biol. 27, 633–645 (1995).
Vlasakova, K. et al. Evaluation of the relative performance of 12 urinary biomarkers for renal safety across 22 rat sensitivity and specificity studies. Toxicol. Sci. 138, 3–20 (2014).
Vinken, P. et al. Tissue Kim-1 and urinary clusterin as early indicators of cisplatin-induced acute kidney injury in rats. Toxicol. Pathol. 40, 1049–1062 (2012).
Lane, B. R. et al. Evaluation of urinary renal biomarkers for early prediction of acute kidney injury following partial nephrectomy: a feasibility study. Eur. Urol Focus 6, 1240–1247 (2020).
Mohamed, F. et al. Nephrotoxicity-induced proteinuria increases biomarker diagnostic thresholds in acute kidney injury. BMC Nephrol. 18, 122 (2017).
Da, Y. et al. Serial quantification of urinary protein biomarkers to predict drug-induced acute kidney injury. Curr. Drug Metab. 20, 656–664 (2019).
Dieterle, F. et al. Urinary clusterin, cystatin C, beta2-microglobulin and total protein as markers to detect drug-induced kidney injury. Nat. Biotechnol. 28, 463–469 (2010).
Voshaar, T., Schweisfurth, H., Schottes, C., Prinsen, J. & Koch, H. P. Serum theophylline concentrations in patients with chronic obstructive lung diseases: a comparison of 2 methods]. Pneumologie 44, 283–284 (1990).
Raymond, M. N., Bole-Feysot, C., Banno, Y., Tanfin, Z. & Robin, P. Endothelin-1 inhibits apoptosis through a sphingosine kinase 1-dependent mechanism in uterine leiomyoma ELT3 cells. Endocrinology 147, 5873–5882 (2006).
Shichiri, M., Kato, H., Marumo, F. & Hirata, Y. Endothelin-1 as an autocrine/paracrine apoptosis survival factor for endothelial cells. Hypertension 30, 1198–1203 (1997).
Vanholder, R., Hoefliger, N., De Smet, R. & Ringoir, S. Extraction of protein bound ligands from azotemic sera: comparison of 12 deproteinization methods. Kidney Int. 41, 1707–1712 (1992).
Tuncdemir, M. & Ozturk, M. The effects of ACE inhibitor and angiotensin receptor blocker on clusterin and apoptosis in the kidney tissue of streptozotocin-diabetic rats. J. Mol. Histol. 39, 605–616 (2008).
Kim, N. et al. Human nuclear clusterin mediates apoptosis by interacting with Bcl-XL through C-terminal coiled coil domain. J. Cell Physiol. 227, 1157–1167 (2012).
Takase, O. et al. Inhibition of NF-kappaB-dependent Bcl-xL expression by clusterin promotes albumin-induced tubular cell apoptosis. Kidney Int. 73, 567–577 (2008).
Shohet, R. V. et al. Mice with cardiomyocyte-specific disruption of the endothelin-1 gene are resistant to hyperthyroid cardiac hypertrophy. Proc. Natl. Acad. Sci. USA 101, 2088–2093 (2004).
Zollmann, F. S. & Paul, M. Transgenic models for the study of endothelin function in the cardiovascular system. J Cardiovasc. Pharmacol. 35, S13–S16 (2000).
Masaki, T., Kimura, S., Yanagisawa, M. & Goto, K. Molecular and cellular mechanism of endothelin regulation. Implications for vascular function. Circulation 84, 1457–1468 (1991).
Parris, R. J. & Webb, D. J. The endothelin system in cardiovascular physiology and pathophysiology. Vasc. Med. 2, 31–43 (1997).
Koller, L. et al. Clusterin/apolipoprotein J is independently associated with survival in patients with chronic heart failure. J. Clin. Lipidol. 11, 178–184 (2017).
Bradley, D. et al. Clusterin impairs hepatic insulin sensitivity and adipocyte clusterin associates with cardiometabolic risk. Diabetes Care 42, 466–475 (2019).
Wittwer, J. & Bradley, D. Clusterin and Its Role in Insulin Resistance and the Cardiometabolic Syndrome. Front. Immunol. 12, 612496 (2021).
Barrett, A. J. & Starkey, P. M. The interaction of alpha 2-macroglobulin with proteinases. Characteristics and specificity of the reaction, and a hypothesis concerning its molecular mechanism. Biochem. J. 133, 709–724 (1973).
Menon, R. et al. Single cell transcriptomics identifies focal segmental glomerulosclerosis remission endothelial biomarker. JCI Insight 5, e133267 (2020).
Joachim, G. R., Cameron, J. S., Schwartz, M. & Becker, E. L. Selectivity of protein excretion in patients with the nephrotic syndrome. J. Clin. Invest. 43, 2332–2346 (1964).
Zhang, G. et al. Plasmin(ogen) promotes renal interstitial fibrosis by promoting epithelial-to-mesenchymal transition: role of plasmin-activated signals. J. Am. Soc. Nephrol. 18, 846–859 (2007).
de Cos, M. et al. Urinary plasminogen as a marker of disease progression in human glomerular disease. Am. J. Kidney Dis. 84, 205–214 (2024).
Heerspink, H. J. L. et al. Sparsentan in patients with IgA nephropathy: a prespecified interim analysis from a randomised, double-blind, active-controlled clinical trial. Lancet 401, 1584–1594 (2023).
Lambers Heerspink, H. et al. Wcn23-1085 A Phase 3, Randomized, double-blind, placebo-controlled study of atrasentan in patients with Iga nephropaty- the align study. Kidney Int. Rep. 8, S279–S280 (2023).
Heerspink, H. J. L. et al. Zibotentan in combination with dapagliflozin compared with dapagliflozin in patients with chronic kidney disease (ZENITH-CKD): a multicentre, randomised, active-controlled, phase 2b, clinical trial. Lancet 402, 2004–2017 (2023).
Candia, J. et al. Assessment of variability in the SOMAscan assay. Sci. Rep. 7, 14248 (2017).
Kim, C. H. et al. Stability and reproducibility of proteomic profiles measured with an aptamer-based platform. Sci. Rep. 8, 8382 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Cunningham, F. et al. Ensembl 2019. Nucleic Acids Res. 47, D745–D751 (2019).
Altenhoff, A. M. et al. The OMA orthology database in 2018: retrieving evolutionary relationships among all domains of life through richer web and programmatic interfaces. Nucleic Acids Res. 46, D477–D485 (2018).
Berthier, C. C. et al. Enhanced expression of Janus kinase-signal transducer and activator of transcription pathway members in human diabetic nephropathy. Diabetes 58, 469–477 (2009).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
Krishnan, A. et al. Genome-wide prediction and functional characterization of the genetic basis of autism spectrum disorder. Nat. Neurosci. 19, 1454–1462 (2016).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Stuart, T. et al. Comprehensive Integration of Single-Cell Data. Cell 177, 1888–1902 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Badia, I. M. P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinform. Adv. 2, vbac016 (2022).
Murphy, A. E. & Skene, N. G. A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis. Nat. Commun. 13, 7851 (2022).
Hsieh, F. Y. & Lavori, P. W. Sample-size calculations for the Cox proportional hazards regression model with nonbinary covariates. Control Clin. Trials 21, 552–560 (2000).
Acknowledgements
The authors thank Emily Tanner, Yuee Wang, and Virginia Vega-Warner for their technical support. The list of NEPTUNE study contributors are included in Supplementary Acknowledgments. This project has received funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement No. 115974 (BEAt-DKD) to HJLH. This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation program and EFPIA and JDRF. This project also received funding from the European Union’s HORIZON Research and Innovation Actions (HORIZON-HLTH-2022-TOOL-11-01 - Tools and technologies for a healthy society- PRIME-CKD) undertaking under grant agreement No. 101095146 to HJLH. This project is funded by the European Union. Views and opinions expressed are however, those of the author(s) only and do not necessarily reflect those of the European Union. Neither the European Union nor the granting authority can be held responsible for them. MFG received support from the Swedish Research Council (grant # 2018-02837), the Swedish Heart and Lung Foundation (grant # 2019-0470), the Swedish Strategic Research Area Exodiab (grant # 2009-1039) and the Swedish Foundation for Strategic Research (LUDC-IRC, grant # IRC15-0067). Additional support was from the George M. O’Brien Michigan Kidney Translational Resource Center (MKTC), funded by NIH/NIDDK U54DK137314 (MK, WJ, VN). The KPMP (MK, co-PI) is funded by the following grants from the NIDDK: U01DK133081, U01DK133091, U01DK133092, U01DK133093, U01DK133095, U01DK133097, U01DK114866, U01DK114908, U01DK133090, U01DK133113, U01DK133766, U01DK133768, U01DK114907, U01DK114920, U01DK114923, U01DK114933, U24DK114886, UH3DK114926, UH3DK114861, UH3DK114915, UH3DK114937. The Nephrotic Syndrome Study Network Consortium (NEPTUNE), U54-DK-083912 (MK co-PI), is a part of the NIH Rare Disease Clinical Research Network (RDCRN), supported through a collaboration between the Office of Rare Diseases Research, National Center for Advancing Translational Sciences, and the NIDDK. Additional funding and/or programmatic support for this project has also been provided by the University of Michigan, NephCure Kidney International, and the Halpin Foundation to NEPTUNE.
Author information
Authors and Affiliations
Contributions
All coauthors have contributed to the manuscript. Specifically, H.J.L.H., M.K., and W.J. conceived the project, were involved in the study design, data generation, data collection, analysis and interpretation. P.V., J.D.S., V.N., M.L., E.M., and K.S. performed or contributed to statistical analyses. W.J. and V.N. designed and performed pathway analyses, single-cell RNA sequence analyses and bioinformatic analysis. K.L.H. and C.E.A. conducted a mouse model experiment. M.F.G. and K.L.D. were involved in data interpretation. W.J. and E.L. were involved in ELISA assay selection, feasibility analysis and validation, biomarker measurement, and data generation. M.H., M.B. and C.O.’C were involved in IHC tissue staining analysis. J.H., F.B., M.I., V.N., P.P., and W.J. were involved in transcriptomic data analysis of the mouse model. L.S. contributed to the figures, edited and formatted the text. H.J.L.H. was involved in the study design and data collection of the SONAR trial. W.J. and H.J.L.H. wrote the first draft of the manuscript. All authors reviewed the manuscript drafts, provided critical feedback that helped shape the research, analysis, and article. All authors approved the final version for submission and take responsibility for the accuracy and integrity of the data. H.J.L.H. had full access to the data in the study and final responsibility for the decision to submit for publication.
Corresponding authors
Ethics declarations
Competing interests
M.K. reports grants and contracts through the University of Michigan unrelated of this work from Chan Zuckerberg Initiative, AstraZeneca, NovoNordisk, Eli Lilly, Boehringer-Ingelheim, European Union Innovative Medicine Initiative, Certa Therapeutics, RenalytixAI, Regeneron, Novo Nordisk, Sanofi, Dimerix, Travere and Vera Therapeutics. Also outside of this work, M.K. has received consulting fees through the University of Michigan from NovoNordisk, Alexion, Novartis, Roche Diagnostics and Vera Therapeutics. W.J., V.N., and M.K. hold a patent PCT/EP2014/073413 “Biomarkers and methods for progression prediction for chronic kidney disease” licensed. P.P. is an employee at Delta4 GmbH. Unrelated to this work, W.J. received consultancy fees from Travere Therapeutics. M.F.G. has received financial and non-financial (in kind) support from Boehringer Ingelheim Pharma GmbH, JDRF International, Eli Lilly, AbbVie, Sanofi-Aventis, Astellas, Novo Nordisk A/S, Bayer AG, within EU grant H2020-JTI-lMl2-2015-05 (Grant agreement number 115974 - BEAt-DKD): She has also received financial and in kind support from Novo Nordisk, Pfizer, Follicum, Coegin Pharma, Abcentra, Probi, Johnson & Johnson, within a project funded by the Swedish Foundation for Strategic Research on precision medicine in diabetes (LUDC-IRC #15-0067). MFG has received personal consultancy fees from Lilly and Tribune Therapeutics AB. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Rhian Touyz and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ju, W., Nair, V., Vart, P. et al. Urinary clusterin as a biomarker of human kidney disease progression and response to the endothelin receptor antagonist atrasentan: An exploratory analysis from the SONAR trial. Nat Commun 17, 2482 (2026). https://doi.org/10.1038/s41467-026-68973-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68973-5