Abstract
Staphylococcus aureus is a leading cause of severe infections in neonatal intensive care units (NICUs). We present results of precision surveillance of S. aureus carriage and invasive infection in a cohort study of high-risk infants in the NICU with the goal of identifying novel strategies for prevention and control. Using whole genome sequence guided epidemiology over a 3-year timeframe, we identify spatial and temporal links for transmission between babies, suggesting shared spaces, caregivers, and physical proximity as major risks for transmission in the NICU. Moreover, environmental surveillance reveals potential environmental reservoirs of S. aureus. Remarkably, specific clusters of S. aureus strains that are associated with invasive infection are also detected in more infants over time during routine surveillance, suggesting a strong link between the rate of dissemination and disease in this vulnerable population. Overall, our findings demonstrate a strong association amongst colonization, transmission, persistence, and the development of invasive infections, underscoring the importance of targeted measures to prevent S. aureus infections in the NICU setting.
Similar content being viewed by others
Introduction
Infants hospitalized in Neonatal Intensive Care Units (NICUs) are at increased risk for healthcare-associated infections (HAIs), which are a major cause of morbidity, mortality, adverse long-term neurodevelopmental outcomes, and a significant contributor to the escalating cost of neonatal care1,2,3. Staphylococcus aureus colonizes and persists on the skin and mucous membranes of infants early in life and is a major cause of invasive infection4,5,6. A recent study of 100,000 very preterm infants born across the US between 2018 and 2020 revealed that one-quarter of HAIs were attributed to S. aureus7. Single and multicenter investigations of S. aureus infections in NICUs have identified significant rates of infection-specific morbidity and mortality, with one study reporting an ~10% case fatality rate7,8. There is an urgent need for improved strategies and effective interventions to combat S. aureus colonization, transmission, and infection.
Currently, there are no standard approaches for NICU surveillance of S. aureus colonization, and there is little evidence for when to initiate or stop surveillance, how frequently to test, and which populations to target9. Guidelines suggest that surveillance is likely to be most useful in outbreaks or ongoing healthcare-associated transmission, especially when results will be used to direct infection prevention strategies such as cohorting, isolation, and decolonization10. However, the true impact of surveillance-guided prevention strategies for S. aureus on HAI outcomes remains unclear11.
At the heart of addressing these gaps in knowledge is the ability to measure the persistence and transmission of specific strains of S. aureus in the NICU setting over time, especially while interventions are being implemented. Traditional molecular epidemiology methods, such as multi-locus sequence typing (MLST), spa-typing, or SCCmec-typing, lack the necessary granularity as genetically distinct S. aureus clones may differ by as few as ~100 single-nucleotide polymorphisms (SNPs)12,13,14,15,16. Whole-genome sequencing (WGS) offers an advantage by enabling direct assessment of the clonality of bacterial isolates. While prior WGS studies have primarily focused on outbreak investigations of methicillin-resistant S. aureus (MRSA), few have monitored the colonization dynamics of both MRSA and methicillin-susceptible S. aureus (MSSA) over extended periods of time17,18,19,20.
Here we report the results of a 3-year, unit-wide surveillance screening for S. aureus across the Children’s Hospital of Philadelphia (CHOP) NICU, encompassing WGS of surveillance isolates of NICU patients, blood cultures from both NICU and non-NICU patients at CHOP, and NICU environmental samples. We aimed to measure the introduction, transmission, persistence, and the risk of invasive infection from S. aureus (MSSA and MRSA) using WGS of samples from serial surveillance screening, bloodstream infections, and environmental sampling in the NICU. Our findings provide evidence of S. aureus transmission and persistence within the NICU and suggest that clusters with documented bloodstream infection display higher rates of dissemination, highlighting new opportunities to prevent serious invasive infections through targeted interventions.
Results
High prevalence of S. aureus colonization and infections in the NICU
S. aureus is a critical threat to the health of high-risk infants, and novel surveillance approaches are needed to track, prevent, and control invasive S. aureus infections in the NICU setting. Here, we designed a precision surveillance approach tracking S. aureus carriage and invasive infection across a 3-year time period in the NICU. Throughout the study surveillance period (October 2021–June 2024), S. aureus was consistently detected with recurrent positive swabs from patients across all sections of the NICU (Fig. 1a). S. aureus was also isolated in 43 blood cultures from 41 infants across 38 distinct bed locations, spanning all seven NICU sections (Fig. 1d). Stable MSSA and MRSA colonization rates were observed over time across all NICU locations. MSSA colonization rates (mean rate, 25.97% colonized patients/sampled patients; [95% CI, 19.41–34.62%]; median rate, 25% [Q1–Q3, 23.79–28.07%]) significantly exceeded those of MRSA (mean rate, 4.98% colonized patients/sampled patients [95% CI, median rate, 4.58% [Q1–Q3, 1.27–9.43%]) (Fig. 1d). This pattern was mirrored in blood stream infection (BSI) rates, where MSSA BSIs (mean rate, 0.37 BSIs/1000 patient-days [95% CI, 0.27–0.47]; median rate, 0.33 [Q1–Q3, 0.32–0.36]) occurred more frequently than MRSA BSIs (mean rate, 0.06 BSIs/1000 patient-days [95% CI, 0.014–0.105]; median rate, 0 [Q1–Q3, 0–0]) (Fig. 1d).
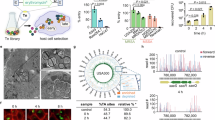
a NICU floorplan visualization of S. aureus methicillin-resistant (MRSA)/S. aureus methicillin-susceptible (MSSA) surveillance results: positive S. aureus colonizing samples (blue dots, size proportional to frequency) and S. aureus bacteremia samples (red crosses). b Core-genome maximum likelihood phylogeny annotated with sample source (outer ring, red: invasive, blue: colonizing, gray: environmental) and clonal complexes (CC) assignment (bars and text labels). Genomes assigned to CC 1, 5, and 8 were paraphyletic in the phylogenetic analysis. Branch colors indicate bootstrap support, with values ≥ 70 supporting 66.4% of branches. c Pair-wise SNP analysis reveals clonal relationships among S. aureus genomes by comparing genomes across three datasets: (1) within NICU genomes (NICU), (2) against a local Philadelphia pediatric database (Local), and (3) against a curated global publicly available database (Global). d Monthly MSSA/MRSA colonization and infection rates. Colonization rates are calculated as a percentage of colonized patients among sampled patients, and infection rates are calculated as BSIs per 1000 patient-days. Both colonization and infection rates were significantly higher for MSSA than for MRSA. e Workflow schematic for identifying strains, clones, and transmission clusters. Red boxes represent decision points, while blue boxes indicate process outputs, serving as inputs for subsequent analytical steps. Source data are provided as a Source Data file.
Genomic diversity and clonality among NICU S. aureus isolates
We performed WGS on 1670 isolates, retaining 1446 high-quality genomes after assembly (982 surveillance, 449 blood cultures, and 15 environmental isolates). These genomes spanned 10 clonal complexes (CCs) and 66 sequence types (STs), with CC398, CC8, and CC5 predominating. A core-genome ML tree (Fig. 1b) indicated substantial genomic diversity. Notably, genomes from surveillance swabs (colonizing), positive blood cultures (invasive), and environmental samples were distributed across all major phylogenetic clades, showing that the ability to cause invasive infections was not restricted to specific lineages.
We identified 328 patients with two or more genomes, a total of 1142 genomes. While most genomes from the same patient formed monophyletic groups (median, 3 genomes/patient [Q1–Q3, 2–4]), 158 genomes from 36 patients (median, 4 genomes/patient [Q1–Q3, 2–5]) were in distinct phylogenetic clades. Twenty-four distinct phylogenetic clades of highly related genomes were identified, encompassing 1442 genomes (median, 26 genomes/clade [Q1–Q3, 4–73]) from 657 patients (median, 11 patients/clade [Q1–Q3, 2–36]). Clonal relationships were further determined through comprehensive SNP analysis using three datasets. We constructed pairwise SNP distance matrices comparing NICU genomes against (1) themselves (NICU matrix), (2) a local database of 349 pediatric S. aureus genomes collected in Philadelphia (Local matrix)21, and (3) a curated global database of 68,298 high-quality publicly available S. aureus genomes22. The NICU matrix uniquely demonstrated a significant cluster of closely related genomes (SNP distance < 100) (Fig. 1c).
Based on the phylogenetic pattern, clonality, and high rates of S. aureus colonization across the NICU, we hypothesized that at least some infants may have acquired S. aureus through transmission within the NICU. To rigorously test for transmission within the NICU, we used a dynamic, phylogeny-corrected SNP threshold (phylothreshold), which allowed for flexible SNP thresholds across different groups (Extended Data Table S1). Briefly, this approach first groups genomes using hierarchical clustering based on average silhouette scores calculated from branch lengths on an inclusive genome tree. Within each group, we determine the number and composition of strains by single-linkage clustering, imposing a threshold value for SNP distance. To determine the optimal threshold, we sequentially increase the SNP distance threshold until reaching a plateau where further increases no longer change the number or composition of strains within the group. This approach revealed that single-SNP thresholds occasionally did not capture clonal relationships within phylogenetic trees, as putatively linked genomes were not always monophyletic (Extended Data Fig. S1). Thus, we incorporated a phylogenetic correction step that checks for monophyly and bootstrap support within each strain. Specifically, genomes are added to plateau-defined groups through this phylogenetic correction step if they belong to the same clade and the clade has bootstrap support ≥70. By employing “phylothresholds”, our approach better reflects phylogenetic inference, and it avoids the limitations of using a single threshold across all genomes from different genetic backgrounds. While phylothresholds are tailored to fit specific groups, they are also consistent with strict thresholds defined across numerous other studies14. For example, when we compared the SNP distances within our phylothreshold-defined strains against 40 published S. aureus transmission studies reporting SNP thresholds or SNP distance ranges within transmission clusters, there was strong concordance (Extended Data Table S3; Extended Data Fig. S2).
Widespread transmission linked to infections and source
We defined a “strain” as either a group of genomes defined by a phylothreshold or singletons with no SNP distance matches. A “clone” was designated as a strain detected in at least two distinct samples on more than one date or patient, while a “transmission cluster” refers to a clone found in more than one patient. Using the above phylothresholds, we identified 490 strains (360 NICU, 130 non-NICU), 209 clones (150 NICU, 59 non-NICU), and 69 transmission clusters (69 NICU) (Fig. 1d, Extended Data Table S2). The 355 non-NICU blood culture genomes from 140 patients comprised 134 non-cluster strains, with only 19 genomes belonging to 7 transmission clusters (Clusters 1, 2, 10, 29, 37, 51, and 58). All cluster-associated non-NICU genomes originated from patients with prior NICU admission, and 97% of non-cluster strains detected outside NICU (130/134) were genomically distinct from NICU strains. This differentiation validated NICU-specific transmission clusters and supported our hypothesis that most infants who acquired S. aureus did so within the NICU rather than from hospital-wide circulation. We quantified colonization events by noting the first time that a strain of S. aureus was detected in a patient’s surveillance sample. We observed 528 NICU colonization events, and 176 (33.33%) were identified as transmission-acquired after excluding 69 index patient cases (the first patient in which the cluster was observed) (Extended Data Fig. S3). CC398 MSSA was most prevalent, with 62/490 strains and 14/69 clusters. This observation aligns with many studies reporting the recent global spread of the CC398 MSSA lineage23, as well as findings from other NICUs in the US17. Of 69 transmission clusters, 19 “invasive clusters” were identified based on the detection of the strain from any BSI (Fig. 2a). Among these, CC398 MSSA accounted for 5/19 (26%), highlighting the association between CC398 MSSA and HAIs in the NICU setting.
a Transmission clusters as determined by phylothresholds (n = 69) represented as boxes, colored according to invasive status (red: invasive clusters; blue: colonizing clusters) and annotated with methicillin resistance status. R indicates methicillin-resistant S. aureus (MRSA) cluster and S indicates methicillin-susceptible S. aureus (MSSA) cluster. Within each cluster, patient status is indicated by colored dots (blue: colonization only; red: infection only; yellow: both colonization and infection), with individual patient IDs shown as numbers. Environmental isolates are marked by gray diamonds. Two bars below each cluster box indicate: (1) the specific NICU section where the cluster was detected, and (2) the assigned treatment team at the time of detection. Among all transmission clusters identified, 19 were classified as invasive. b Temporal comparison of genomic emergence and transmission index cases versus index patient admission dates (n = 48 transmission clusters, each row representing a distinct transmission cluster with ≥4 genomes). Green bars represent 95% confidence intervals for transmission index case dates (bounds: weighted 2.5th and 97.5th percentiles of inferred transmission index dates across posterior samples), and dark green lines indicate medoid dates. Blue bars represent 95% highest posterior density intervals for tMRCA (time to most recent common ancestor) from BEAST2 molecular clock analysis (bounds: lower and upper 95% HPD boundaries from the posterior distribution), and dark blue lines indicate the mean. Purple triangles indicate index patient admission dates, and purple circles indicate collection dates of the earliest cluster genomes. The 95% confidence intervals of transmission index dates overlap with the 95% HPD of tMRCA for all clusters, indicating that tMRCA serves as both a genomic emergence indicator and a reliable proxy for transmission index cases. Index patient admission dates fell within the 95% HPD of tMRCA for 50% (24/48) of clusters, suggesting that admission of pre-colonized index patients represents a major introduction route. Source data are provided as a Source Data file.
We captured the progression of 19 patients (19/618) from colonization to invasive infection with the same strain. The median interval was 19 days (Q1–Q3, 9–138, mean, 66.95). At the strain level, for the 19 strains associated with both colonization and invasive infection (10 of which were transmission clusters), the median interval from earliest colonization to earliest invasive infection was 147 days (Q1–Q3, 14–245, mean, 156.47). Among the 69 transmission clusters identified, the median length from earliest detection to detection in a new patient was 91 days (Q1–Q3, 18–210, mean: 131.10).
A total of 44 BSIs were associated with strains detected in the NICU, including 26 from 19 invasive clusters. After excluding 11 index patient cases, 15 infections (34.10%) were attributed to transmission (Extended Data Fig. S3). Cluster 1 was the largest transmission cluster, involving 30 patients, and four patients were diagnosed with Cluster 1 BSI. To investigate whether transmission clusters originated from putative index patients pre-colonized by the respective strains upon admission, we compared admission dates of the index patients in the clusters with genomic emergence dates estimated through (i) time to the most recent common ancestor (tMRCA) and (ii) transmission index dates from TransPhylo analysis24. The medoid of dates of transmission index inferred by TransPhylo v1.4.5 fell within the 95% highest posterior density (HPD) intervals of the tMRCA, and the 95% confidence intervals of transmission index dates overlapped with the 95% HPD for all 48 clusters (Fig. 2b). This concordance indicates that tMRCA serves as both an indicator of genomic emergence and a reliable proxy for transmission index cases. Among the 48 evaluable transmission clusters, 50% (24/48) had putative index patient admission dates that fell within the 95% HPD of tMRCA, suggesting that these pre-colonized patients likely introduced the clusters upon admission rather than acquiring them within the NICU. In the remaining 50% (24/48) of clusters, transmission index dates either predated or postdated the putative index patients’ admission dates, indicating alternative introduction routes. We also employed TransPhylo and a Hidden Markov Model (HMM; msm v1.8.225; Extended Data Fig. S10) to estimate rates of unsampled genomes in each transmission cluster.
These findings suggest that at least half of transmission clusters were introduced through the admission of pre-colonized patients captured in our NICU. Alternative introduction routes for the remaining clusters may include introduction by unsampled patients, healthcare workers, visitors, or environmental reservoirs. Together, these results illuminate the primary mechanisms of cluster introduction while underscoring the complexity of S. aureus transmission dynamics in the NICU setting.
Spatial and temporal proximity between patients is associated with transmission
Only eight of the 69 transmission clusters were MRSA, with three being invasive clusters and five colonizing clusters, suggesting that antibiotic resistance alone does not fully explain transmission and persistence (Fig. 2a). We hypothesized that spatial and temporal proximity between patients would be a strong determinant of transmission. As exemplified in Cluster 1 (Fig. 3a, c), patients shared overlapping lengths of stay and locations across all seven NICU sections, generating a strong epidemiological link in the cluster.
Temporal analysis of Cluster 1 by location and treatment team assignments. a Patient timeline showing NICU sections (colored rectangles), with admission (triangles) and discharge dates (squares). b Corresponding treatment team assignments over time (colored rectangles). In both panels, sampling events are indicated by dots (red: invasive isolates; blue: colonizing isolates). c NICU floorplan visualization of patient movements. Spatial and temporal proximity serve as the primary drivers of transmission, as visualized in the floor plan. Arrow and dot colors represent different patients. The legend for patient dot and arrow colors is omitted in this panel. For complete panels with legends, see Extended Data Fig. S3.
Complementing the epidemiological link in the cluster and serving as a potential proxy for unsampled healthcare workers, shared treatment team assignments revealed potential healthcare worker-mediated transmission routes. For instance, despite physical separation in distinct NICU sections, patients 229 and 235, as well as patients 414 and 399, were linked through concurrent care by the same NICU treatment team (Fig. 3b). This pattern observed in Cluster 1, where strong epidemiological links emerged through both spatial proximity and overlapping treatment team assignments, was seen in 60.9% (42/69) of transmission clusters, as exemplified by Cluster 1 (Supplementary Data S1–S3). An additional 27.5% (19/69) of clusters demonstrated links only through shared NICU sections, while one cluster (1.4%) showed overlapping treatment teams without a shared NICU section (Supplementary Data S1–S3).
Notably, 10.1% of clusters (7/69) exhibited no epidemiological links in our analysis. The SNP distances within these clusters confirmed accurate strain determination by our pipeline, suggesting that analytical artifacts are unlikely (Extended Data Fig. S2d). These transmission clusters lacking epidemiological links suggest transmission may have occurred through unsampled intermediaries, shared equipment, environmental reservoirs, or transmission from parents not observed in this study. Collectively, these findings suggest that shared NICU spaces and treatment teams represent the primary drivers of transmission, but also highlight a role of additional cryptic transmission that was not detected by our study.
Isolates recovered from environmental sampling represented three transmission clusters
To further detect potential reservoirs of transmission, we collected 210 environmental samples from the surfaces in all seven NICU sections using swabs and broad-based culture methods. Of the 19 S. aureus isolated from these surfaces, 15 produced high-quality genomes (sampling locations in Extended Data Fig. S4). Remarkably, we recovered environmental isolates matching three of the previously defined transmission clusters (Clusters 22, 58, 67) (Fig. 4). There was high spatial proximity between environmental isolate collection sites and patient beds for all three (Fig. 4b). The time intervals between the most recent cluster detections in patients and the corresponding environmental recovery (30–42 days) suggest the possibility of persistent environmental reservoirs despite routine NICU sterilization procedures. This also could be explained by high abundance or prevalence in the NICU, and shedding from colonized patients, healthcare workers, or visitors. Regardless of the possible scenarios, our findings demonstrate that environmental surfaces in NICU settings could serve as critical intermediary reservoirs facilitating transmission dynamics between patients, healthcare workers, and the NICU environment.
aTemporal distribution of isolate sampling (red dot: invasive, blue dot: colonizing, gray diamond: environmental, numbers within circles indicate de-identified patient IDs, and text within diamonds indicates environmental sampling location), demonstrating the prolonged persistence of environmental reservoirs, demonstrating the detection of environmental isolates after the most recent detection of clusters. b Spatial mapping of environmental reservoirs (gray squares) relative to patient movement within the clusters, showing high spatial proximity between environmental isolate collection sites and patient bed locations. Environmental detection may represent persistent reservoirs or shedding from colonized patients, healthcare workers, or visitors.
Persistence in the NICU is associated with increased invasive infection
We examined the length of cluster persistence in the NICU by measuring the time between the first isolation date of a transmission cluster and the most recent date of detection. Invasive clusters, defined by at least one BSI isolate in the cluster, demonstrated significantly higher NICU persistence (median persistence, 421 days [Q1–Q3, 212–570]) than the 50 colonizing clusters (median persistence, 128 days [Q1–Q3, 35.25–394]) (Fig. 6a). To further assess persistence of transmission clusters, we calculated the tMRCA for 48 transmission clusters with more than four genomes (18 invasive, 30 colonizing). The molecular clock persistence times of 18 invasive clusters (median persistence, 576 days [Q1–Q3, 304.5–907.25]) were higher than those for the 30 colonizing clusters (median persistence, 383 days [Q1–Q3, 195.5–655.25]) (Extended Data Table S2). These results suggest that persistence in the NICU is highly associated with subsequent invasive infection. Additionally, our analyses revealed transitions from colonizing to invasive states in several invasive clusters. For instance, Clusters 1, 2, and 4, which were the top three most persistent invasive clusters, initially manifested as colonizing clusters for a few months before causing invasive disease (Fig. 5). These findings suggest that strains with enhanced persistence and transmission capabilities pose a significantly higher risk for invasive infections in the NICU setting. This increased risk may be mediated by a variety of bacterial factors that promote both persistence, spread, or invasive disease, unmeasured host characteristics that render the infant more vulnerable to persistent colonization and infection, or the overall abundance or prevalence of specific clones.
Temporal distribution of isolate sampling of invasive clusters (n = 19), with invasive (red), colonizing (blue), and environmental (gray diamonds) sources. Numbers within circles indicate de-identified patient IDs, and text within diamonds indicates environmental sampling location. The temporal patterns demonstrate transitions from colonizing to invasive status after transmission and prolonged persistence in the NICU setting. Source data are provided as a Source Data file.
Enhanced dissemination is associated with the risk of invasive infections
Invasive clusters persisted in the NICU longer than colonizing clusters (median difference, 293 days; P = 0.01) and, on average, were detected in more patients (median difference, 2 patients; P = 0.01) (Fig. 6a, b). To test if this difference applies across equivalent time periods, we calculated the correlation between the number of patients and the duration of persistence. Invasive clusters exhibited a slope that was approximately five-fold higher than colonizing clusters (Fig. 6c). Next, we did a temporal analysis of the accumulated number of new patients in whom the strain was detected throughout the time of NICU persistence (Fig. 6d, e). Invasive clusters were detected in new patients more rapidly than colonizing clusters, with this difference becoming particularly obvious from day 200 to day 500 after initial detection. Linear Mixed-Effects Models also confirmed that invasive clusters were detected in more patients over time (interaction term P < 2e−16). Although Cluster 1 exhibited exceptional dissemination across all metrics and could potentially cause bias, analyses excluding this cluster confirmed that invasive clusters remained significantly higher in accumulated patient numbers and persistence duration (Extended Data Fig. S5). To account for bias introduced by combining genomes from blood cultures with genomes collected during surveillance, we also recalculated NICU persistence and accumulated patient detection, limiting our analysis to genomes collected during surveillance (Extended Data Fig. S8). In this analysis, transmission clusters that had been independently seen in blood cultures were still more likely to be found in more patients and demonstrated higher inferred rates of dissemination. Furthermore, when clusters were stratified by the number of patients with bloodstream infections, we observed a graded relationship, in which dissemination rates were highest in clusters with two or more patients with bloodstream infections, intermediate in those with a single patient, and lowest in those with none (Extended Data Fig. S9).
a Persistence times were significantly longer in invasive clusters (n = 19) compared to colonizing clusters (n = 50). b Invasive clusters (n = 19) involved significantly more patients than colonizing clusters (n = 50). For a and b, box plots display the median as the center line, box bounds represent the 25th and 75th percentiles, and whiskers extend to the smallest and largest values within 1.5 × IQR within each group. Statistical significance was determined using two-sided Wilcoxon rank-sum tests with continuity correction and without adjustment for multiple comparisons; *P < 0.05 (a: P = 0.012; b: P = 0.010). c Linear regression analysis revealed that invasive clusters were associated with a higher patient count over the same persistence. Separate regression lines are shown for colonizing (blue; n = 50) and invasive (red; n = 19) clusters. Shaded bands represent 95% confidence intervals for each regression line. For a–c, each dot represents a distinct transmission cluster. d Accumulated patient numbers over days post-initial detection are plotted for each cluster, with colors indicating invasive (red, n = 19) versus colonizing (blue, n = 50) status. e Mean accumulated patients per day post-initial detection by invasive status (n = 19 invasive clusters; n = 50 colonizing clusters). Lines show mean values; shaded ribbons indicate ± standard error. Blue represents colonizing clusters, red represents invasive clusters, and dotted lines indicate a single remaining cluster (no standard error). A linear mixed-effects model with cluster as a random effect was fitted using restricted maximum likelihood (REML), and models with and without the interaction term (invasive status × time) were compared using a likelihood ratio test with maximum-likelihood estimation. Invasive clusters accumulated patients significantly faster than colonizing clusters (interaction term coefficient = 0.011 patients/day, χ² = 5585.9, df = 1, P < 2.2 × 10⁻¹⁶. This P-value represents the minimum reportable P-value in R). Source data are provided as a Source Data file.
These striking findings raise the possibility that certain clones pose a significantly greater risk by being more transmissible and potentially more invasive, providing a clear rationale for prioritizing surveillance and prevention targeting these particular strains. Together, this comprehensive study of S. aureus in the NICU setting creates a new paradigm for our understanding of colonization, transmission, and invasive infection of S. aureus in high-risk neonates and provides a roadmap for future surveillance and intervention strategies.
Discussion
Infections due to S. aureus remain a major cause of morbidity and mortality in the NICU setting7. Colonization is associated with increased risk of infection, and exposure to the NICU environment and equipment, healthcare workers, and parents are likely the major factors that impact transmission and colonization26,27,28. Despite decades of infection prevention efforts and collaborative implementation of bundles of care, rates of invasive infections, such as central line-associated bloodstream infections, have plateaued, especially in complex and high-risk populations in children’s hospitals28. While bloodstream infections due to coagulase-negative staphylococci have declined in our NICU, S. aureus (mostly MSSA) remains the predominant pathogen associated with bloodstream infections. Point prevalence testing in our NICU indicates high S. aureus colonization rates among NICU infants, primarily MSSA with a few MRSA cases. New strategies are needed to mitigate S. aureus transmission and infection in the NICU. In this study, we employed WGS of samples from surveillance screening and prospective collection of invasive infections to measure the dynamics of dissemination of S. aureus in our NICU.
For determining the dissemination of clones, most studies usually rely on SNP thresholds set at a single value across groups of genomes, which can lead to false negative and positive assignment of genomes to clonal groups29. In addition, the length of time a strain has been persistent and variations in mutation rate could make single cutoffs problematic30. We show that even conservative cutoffs (≤25 SNPs) can produce transmission clusters incongruent with clonal groups on a phylogenetic tree. Thus, we used a strategy with variable cutoffs across groups that also corrected for phylogenetic discrepancies and support values, which we call phylothresholds. Our method infers transmission clusters at multiple SNP distances, enabling assessment of both conservative and broader thresholds (0–500 SNPs) that adhere to phylogenetic monophyly (Extended Data Table S1). This new method captures clear clonal relationships amongst NICU genomes, strengthening the inference of isolate “sameness” and enhancing our understanding of transmission dynamics.
WGS has been used previously for tracking S. aureus dissemination in NICU settings17,18,19. However, previous studies have lacked analyses combining both MRSA and MSSA, and only a small number have identified links between dissemination and invasive infection19,31,32,33,34,35. Our study leveraged a comprehensive collection of isolates from multiple sources in the CHOP NICU (routine surveillance samples, infant blood culture, and environmental samples) with both MRSA and MSSA. The hospital-wide non-NICU blood culture genomes served as controls that corroborated the transmission within the NICU and also enabled tracking of colonization persistence in patients transferred out of the NICU. The prolonged length of surveillance and the large number of samples sequenced ensure the robustness of the inference of the dynamics of dissemination, persistence, and introduction.
Data from this study were used to inform various infection prevention efforts in the NICU, including improving the quality of hand hygiene and mupirocin decolonization to aimed at decreasing S. aureus prevalence. Spatial localization of clusters helped uncover that transmission was occurring unit-wide and not limited to specific sections or staffing groups. Environmental sampling suggested that the patient environment is likely a reservoir for S. aureus and has led to an increased focus on environmental cleaning efforts, such as UV cleaning, to reduce bacterial burden in the NICU. Ongoing genomic investigation will help measure the impact of each of these sequentially implemented interventions on rates of colonization and transmission of various strains of S. aureus, as well as invasive infections. Additionally, WGS is helping us define S. aureus strains at the highest risk of dissemination and infection in the specific context of the NICU. As next steps, we are designing new approaches to rapid detection of high-risk S. aureus strains, to facilitate deployment of targeted infection prevention approaches against the high-risk strains.
Our findings suggest that targeting the elimination of specific strains may be a highly effective approach to reducing invasive disease. In our analyses, strains that were detected in more patients over time were more likely to be invasive. It remains to be investigated whether this association is due to an inherent biological link between transmissibility and virulence or simply that highly prevalent strains are more likely to encounter a vulnerable host. We also evaluated spatial spread patterns as a potential proxy of transmissibility by rescaling spatial distance based on phylogenetic topology (detailed in Spatial Distance Rescaled Phylogenetic Analysis in Methods, Visualization in Supplementary Data 4). This analysis mirrored the patient transmission outcome, suggesting that invasive clusters spread more rapidly in space than do colonizing clusters during equivalent time periods (Extended Data Fig. S6). However, we acknowledge that additional studies are needed to validate the epidemiological significance of these spatial patterns. Despite these uncertainties and the need for further validation, targeting the most prevalent strains may offer a high-impact strategy to reduce invasive disease. If high-risk strains can be identified early and reliably, then targeted strategies such as patient care cohorting, contact precautions, intranasal mupirocin, and topical chlorhexidine decolonization could reduce the risk of invasive S. aureus infection. We also identified links between environmental sources of S. aureus and invasive clusters, highlighting that enhanced cleaning efforts may be needed to eliminate high-risk strains from the environment. The precision surveillance approach presented in this study will provide us with the ability to accurately measure the impact of targeted interventions on decreasing the dissemination of high-risk strains. Identification of the biological and genomic properties of high-risk strains will be a major focus of future work.
This study has several potential limitations. As a single-center investigation, our findings may not be fully generalizable, particularly to lower-acuity settings. Additionally, our phylothreshold technique might underestimate instances of strain reintroduction to the NICU, and future sampling of locally circulating S. aureus clones outside the hospital will strengthen this work. Previous studies have reported that healthcare workers can be sources of transmission to patients in clinical settings12, and the epidemiological characteristics of some of our transmission clusters suggest this possibility. However, the absence of samples from healthcare workers limits our ability to comprehensively track transmission dynamics, as healthcare workers likely serve as key intermediaries for transmission during routine care activities. Similarly, we did not collect samples from parents, which precludes investigation of parent-to-infant transmission. Parents could theoretically transfer S. aureus between their own infant, the environment, and others during visits, representing an understudied transmission route. These sampling limitations highlight the need for future studies incorporating surveillance of all potential human reservoirs to fully elucidate transmission dynamics.
In addition to this host sampling limitation, the design of the monthly surveillance sampling imposes constraints on genomic resolution. Although we collected isolates from a large cohort of patients (n = 617), the number of genomes recovered per patient was limited (mean, 2.32 genomes; median, 2 [Q1–Q3, 1–3]). This sampling density is insufficient to resolve within-host S. aureus population diversity or evolution within the NICU setting. The reliance on a single-isolate-per-timepoint strategy may therefore underestimate transmission events, particularly those involving multiple co-colonizing lineages. Additionally, the intermittent sampling intervals create gaps that likely miss some patients resident in the NICU for short periods and transient periods of colonization. Overall, any sampling strategy will miss some colonization, transmission, and infection events. Putative donors and recipients, and unsampled intermediate hosts or environmental sources, can only be inferred by techniques such as transmission trees, phylogenetic trees, or models that incorporate missing data, such as the HMM models used here (Extended Data Fig. S10). While these techniques can help ameliorate problems with missing data or sampling limitations, they are limited in their ability to definitively identify transmission sources and true colonization rates. Also, while a detailed analysis of host characteristics was beyond the scope of this project, future studies will incorporate that information to improve understanding of the interaction of high-risk strains and vulnerable hosts.
Importantly, the correlation between the dissemination rate of specific strains and invasive infections could arise from multiple factors, including increased transmissibility rates, increased ability to persist once transmitted, higher prevalence or durability in the environment, or multiple other intrinsic biological factors. It is also possible that other opportunistic factors, rather than intrinsic biological characteristics, are at play, such as introduction from a patient with a high bacterial burden, repeated introductions from a staff member, or proximity to immunocompromised infants. While our current data cannot definitively distinguish between these factors, it is likely that multiple processes contribute jointly to the dynamics of dissemination.
Our study shows extensive transmission, persistence, and introduction of both MRSA and MSSA in the NICU, with MSSA posing a greater problem for colonization and invasive disease. To our knowledge, this is the first report linking high transmission rates to invasive S. aureus clusters in the NICU setting. Our findings underscore the importance of detecting colonization and transmission for infection prevention and suggest future directions for infection prevention with real-time interventions targeting highly transmissible strains.
Methods
Study setting and surveillance
This study was approved by the CHOP Institutional Review Board (IRB 022889 and 17-014648). Informed consent was waived, and no participant compensation was provided. Sex and gender were not considered in the study design, as dissemination of S. aureus is expected to be independent of patient sex and gender. The CHOP NICU system comprises two facilities: a 102-bed quaternary care unit (main campus) and an 18-bed Level III NICU at the King of Prussia (KOP) campus. These units manage approximately 2900 patient-days monthly and 12,000 central line-days annually.
A comprehensive S. aureus colonization surveillance program was implemented across both facilities. At the main campus NICU, we initiated quarterly MSSA nares screening in addition to monthly MRSA surveillance between October 2021 and July 2022, followed by monthly MRSA/MSSA nares surveillance, collecting 5466 samples from 2092 patients across seven distinct NICU sections. The KOP NICU initiated MRSA/MSSA monthly surveillance in December 2023, with 133 samples collected from 95 patients by June 2024. Given the monthly point prevalence sampling approach and the prolonged hospitalization experienced by many preterm and term infants with complex medical conditions, patients were re-sampled if they remained admitted in the NICU for additional point prevalence sampling dates. As long as samples met quality control parameters, repeat samples (surveillance or positive blood culture) from the same patients were included in the analysis.
During the timeframe of the study, the NICU and Infection Prevention and Control teams were actively engaged in quality improvement and infection prevention efforts to decrease S. aureus colonization and infection rates, in addition to ongoing efforts to prevent healthcare-associated infections. All infants identified to be positive with either MSSA or MRSA based on surveillance or results of diagnostic testing for clinical indication, underwent decolonization with a 5-day course of mupirocin, starting in July 2022. All infants with central venous catheters (term corrected age or older) received daily chlorhexidine bathing. The NICU also embarked on a hand hygiene and ‘bare below the elbows’ campaign in July 2022 to re-engage staff in infection prevention efforts. Finally, the NICU added initiatives to enhance cleaning procedures for NICU patient rooms through the course of the study.
Concurrent with NICU colonization surveillance, S. aureus-positive blood culture isolates were collected hospital-wide from patients with bloodstream infections as part of routine sampling at CHOP. Between October 2021 and June 2024, 94 isolates from 34 patients within the NICU and 355 isolates from 140 patients outside the NICU were obtained. These isolates served as geographically proximate controls to assess NICU-specific transmission and enabled longitudinal tracking of patients transferred from the NICU to other locations.
Study definitions
Bloodstream infections (BSI) and central line-associated bloodstream infections (CLABSI) were defined as per the Centers for Disease Control National Healthcare Safety Network (NHSN) definitions for lab-confirmed bloodstream infections36. BSI was defined as primary when no secondary source was identified, or secondary when identified to be seeded from a localized infection. CLABSI was defined as a BSI in the setting of a central line in place for more than two consecutive calendar days, with no secondary source, in concordance with NHSN definitions. For S. aureus, a single positive blood culture is sufficient to diagnose BSI.
Clinical sampling and isolate processing
Blood culture: During cases of suspected bacteremia, Bactec PedsPlus/F (BD, New Jersey, USA) bottles were loaded onto a Bactec FX automated analyzer for up to 5 days of incubation at the CHOP Infectious Disease Diagnostics Laboratory (IDDL). Positive blood samples are then plated to 5% sheep blood Tryptic Soy Agar (BD), Chocolate agar (BD), and MacConkey (BD) agar for aerobic growth. All plates are grown at 5% CO2 and 35-37 °C. Plates are analyzed daily for growth. Species-level identification is confirmed using a Biotyper CA (Bruker, MA, USA), matrix-assisted LASER desorption/ionization time-of-flight mass spectrometer (MALDI-TOF MS) platform. Single colonies were subcultured and archived for long-term storage and downstream sequencing. From subcultures, a sterile half-loop (10 µL inoculating loop) of growth from the plate was transferred into the Qiagen PowerSoil 0.5 mm glass bead tube (cat #13116-50) containing 500 µL of molecular-grade water for the Illumina direct from colony DNA extraction. Tubes were briefly vortexed to mix and either processed immediately or stored at −80 °C. A fresh loop is then used to collect remaining growth and transfer it into 1 mL of 20% LB-glycerol solution for cryobanking.
S. aureus nasal surveillance: BBL™ CultureSwab™ EZ collection (BD) from the anterior nares were sent to the CHOP IDDL and plated onto Chromagar™ MRSA II (Chromagar, St-Denis, France) and Mannitol salt agar (BD). Each surveillance collection involved all infants in the NICU and was performed over 2–3 days. Plates were incubated in 5% CO2 at 35 °C and analyzed following overnight growth. Identification as S. aureus was confirmed using a Staphaurex (ThermoFisher Scientific, MA, USA) latex agglutination assay and as MRSA by the PBP2a SA Culture Colony Test. Subcultured single colonies of confirmed isolates were then processed for cryobanking and direct from colony DNA extraction following the same methods described above.
Environmental sampling
A pilot environmental sampling was conducted across all seven sections of the neonatal intensive care unit (NICU) to identify potential environmental reservoirs of transmission. A total of 210 surfaces representing high-touch and high-traffic areas (e.g., cribs, monitors, sinks, chairs, toys, closets) were systematically sampled using a standardized protocol. A Neogen® Sponge-Stick with Neutralizing Buffer (Neogen cat. # SSL10NB) for culturing, ensuring consistent order of sampling from the crib area outward to the general room environment. Samples were transported immediately to the laboratory and processed the same day. Sponge samples were homogenized using a BagMixer 400 to extract the liquid fraction, which was then aliquoted into sterile conical tubes. Aliquots (10 µL) were plated onto a panel of these selective CHROMagar media (cat. # TA672) for the detection of S. aureus amongst other media for organisms outside the scope of this study. In parallel, 100 µL of each homogenized sample was inoculated into BHI broth for enrichment to facilitate the recovery of low-abundance organisms. All plates and enrichment cultures were incubated at 37 °C for 18–24 h. Plates were examined for colony growth and morphology after incubation. Distinct colonies were subcultured using sterile 10 µL loops onto LB agar plates for isolation. Representative colonies were archived in 20% LB–glycerol solution and stored at −80 °C. Downstream whole-genome sequencing was performed on single colonies from all S. aureus isolates.
Whole genome sequencing
Single colony isolates from the CHOP IDDL were stored at −80 °C at the CHOP microbial ARchive and Cryo-collection (microbialARC) strain bank and were sequenced at the CHOP Microbiome Center. DNA from all isolates was extracted using the Illumina Direct from Colony protocol. Briefly, for lysis, bead tubes were shaken at 20 Hz for 10 min, then centrifuged at 4500×g for 2 min to pellet the beads. Twenty microliters of the supernatant were transferred into a fully skirted PCR plate. Crude DNA was purified using the AMPure XP SPRI bead cleanup, with two washes in 80% ethanol and final elution in 80 µL of elution buffer. The resulting DNA was stored at −20 °C until sequencing. Extracted DNA was quantified with the Quant-iT PicoGreen® dsDNA assay kit (ThermoFisher Scientific) before library generation. Libraries were generated from 7.5 ng DNA using the Illumina DNA Prep kit and IDT for Illumina unique dual indexes at 1:4 scale reaction volumes. Library success was assessed by Quant-iT PicoGreen® dsDNA assay kit, and samples with library yields <1 ng/µl were re-prepped as needed. After all samples for a given pool were prepped, an equal volume of library was pooled from every sample, and then the pool was sequenced using a 300-cycle Nano kit on the Illumina MiSeq. This low-cost pilot run provided preliminary read counts and indexing performance, allowing us to assess library balance across samples. Libraries were subsequently re-normalized and repooled based on the demultiplexing statistics of the MiSeq Nano run to ensure more even representation in the final sequencing pool. Final libraries were sequenced on an Illumina NovaSeq 6000 v1.5 flow cell, producing 2 × 150 bp paired-end reads. Extraction blanks and nucleic acid-free water were processed along with experimental samples to empirically assess environmental and reagent contamination. A laboratory-generated mock community consisting of DNA from Vibrio campbellii and Lambda phage was included as a positive sequencing control.
Genome assembly and quality control
Raw reads underwent quality control using Sunbeam 4.3.7, including adapter trimming and host decontamination37. Isolate genomes were assembled using SPAdes v3.15.538. To obtain clean genomes, 102 isolate genomes were processed through the Anvi’o v8 pipeline using MEGAHIT v1.2.9 for contig filtering and removal of contaminating sequences39,40. Assembly quality was assessed using CheckM v1.2.2 with the following criteria: ≥95% completeness, ≤5% contamination, and “Staphylococcus (UID301)” lineage classification41. Species-level contamination was evaluated using Mash version 2.342. Assemblies were filtered to retain genomes between 2.55 and 3.15 Mb. Of 1670 assembled genomes, 1446 passed quality criteria and were used for downstream analyses. Genomes rejected from quality control appeared to be a random subset of the data; we found no temporal, patient-based, location, sequence type, or anatomical collection site associations (https://github.com/qianxuans/CHOP-NICU-Scripts/blob/main/Genome_assembly_and_quality_control/failed_genomes.xlsx). All passed assemblies are available on NCBI (accession numbers available in https://github.com/qianxuans/CHOP-NICU-Scripts/blob/main/Genome_assembly_and_quality_control/passed_genomes.xlsx).
The genome assemblies demonstrated robust sequencing depth with a mean coverage of 90.22X (median, 76.79X [Q1–Q3, 42.78−115.92X]) and assembly contiguity as reflected by contig length averaging 18,541.07 base pairs (median, 18.361 bp [Q1–Q3, 13,357.5–22,566.5 bp]). This comprehensive genomic dataset allowed high-resolution investigation of clonality and transmission dynamics. The genome assemblies deposited in the NCBI GenBank database may exhibit minor variations from those employed in our analyses, due to GenBank’s standard processing pipeline, which includes sequence trimming and the exclusion of contigs below 200 base pairs. Accession numbers are available in the Data Availability section.
Core-genome phylogeny inference
The 1446 high-quality genome assemblies were annotated using Bakta v1.9.2 with Database v5.143. The resulting gff3 files were utilized for core-genome alignment, which was carried out using panaroo 1.5.044. This alignment was then used to infer the maximum-likelihood phylogeny in IQ-TREE version 2.3.045, employing the general time-reversible (GTR) substitution model46 and accounting for among-site rate heterogeneity using the Gamma distribution and four rate categories47.
The phylogenetic inference was performed with 100 initial parsimony trees. Unbiased branch support values were provided by UFBoot using 10,000 bootstrap replicates48. SH-like approximate-likelihood ratio test was implemented with 10,000 replicates to assess branch support49. The resulting phylogeny was midpoint-rooted and visualized using a customized R script with packages ggplot2 and ggtree50,51.
Clonality determination
To test for and determine clonal strain composition, we first examined pairwise SNP distances calculated using the wrapper dnadiff in MUMmer 4.0.0rc52, using the gSNP in the dnadiff output as the number for SNP distance. We constructed pairwise SNP distance matrices comparing NICU genomes against: (1) themselves (NICU matrix), (2) a local database of 349 pediatric S. aureus genomes collected in Philadelphia (Local matrix)21, and (3) a curated global database of 68,298 high-quality publicly available S. aureus genomes53.
Next, we used a hierarchical clustering approach using silhouette scores to partition all genomes into 28 groups using the phylogenetic tree described previously in core-genome phylogeny inference. For the hierarchical clustering process, we started by computing the pairwise distances between any pair of genomes using branch lengths in the phylogenetic tree using the cophenetic.phylo() function in R package ape54. The generated distance matrix was used to test every possible grouping pattern using the function hclust() from stats, a built-in R package. To evaluate the quality of the grouping pattern, we calculated the average silhouette scores using the function silhouette from the R package cluster55, and the grouping pattern with the highest average silhouette scores was used to partition the genomes. Every group with two or more genomes was supported with 100 ultrafast bootstrap support48 and 100 SH-like approximate-likelihood ratio test support (SH-aLRT)49, expect for Group2 with 97 SH-aLRT support (Extended Data Table S1).
In total, 28 groups were identified by hierarchical clustering. If there was only one genome in a group, that genome was considered a singleton. For groups with 2 or more genomes, we also filtered out any genomes that were more than 1000 SNPs away from every member of the group. These were treated as singletons, which minimizes the possibility of disruptive long branches in subsequent phylogenetic inference.
Twenty-four out of 28 groups consisted of two or more genomes, averaging 60 genomes (median, 26 [Q1–Q3, 4–73]). After excluding three singletons from three groups, we implemented a fine-grained phylogenetic analysis for each of the 24 groups. We used the curated database of 68,298 high-quality S. aureus genomes mentioned previously to identify the five most similar publicly available genomes for every study genome in the group. Those publicly available genomes limited the thresholds to focus on potential transmission clusters within the NICU. We then produced multi-sequence alignments containing study genomes and the most similar publicly available genomes using Snippy 4.6.056, with the study genome with the highest N50 in each group used as the reference genome. The resulting alignments were used as the input for inferring the reference-based maximum-likelihood phylogenetic trees with IQ-TREE version 2.3.045, employing the general time-reversible (GTR) substitution model46 and accounting for among-site rate heterogeneity using the Gamma distribution and four rate categories47, and tested for robustness with standard bootstrap supports57. We implemented a single linkage clustering (SLC) algorithm to determine closely related strains at every possible potential SNP threshold, from the minimal distance in the group to a threshold of 500. We then selected the SNP threshold at which the number and composition of strains plateaued. The plateaus of 24 groups averaged 21.58 SNPs (median, 18.5 [Q1–Q3, 10.75−27.25]), and 486 strains were determined (223 singletons and 263 clones). At each SNP threshold tested, the strain compositions determined by pairwise SNP distances were mapped to the corresponding reference-based phylogenetic tree of the group. Interestingly, even at conservative SNP thresholds (<25), the strains determined by SNP thresholds were sometimes not monophyletic on the phylogenetic tree (multi-SNP-threshold plot, Extended Data Fig. S1). To correct for this problem, we expanded each group to include the smallest clade that included all SLC members as well as any non-SLC members in that clade, when the bootstrap support for the clade ≥ 70. We compared our SNP thresholds to those used in 40 other studies in NICU (Extended Data Table S3; Extended Data Fig. S2), in ongoing work, we are formally validating our pipeline with simulated and other real-world genomic datasets.
Cluster analysis and visualizations
If any bacteremia isolate was present in a transmission cluster, it was defined as an invasive cluster; otherwise, it was referred to as a colonizing cluster.
Methicillin resistance in transmission clusters was confirmed through clinical laboratory testing and detection of the mecA gene using BLASTx 2.14.0+ against a customized protein database containing 13 PBP2a family β-lactam-resistant peptidoglycan transpeptidase amino acid sequences from the NCBI protein database58,59. The accession numbers of the sequences were WP_057521704.1, WP_063852670.1, WP_063852677.1, WP_063852683.1, WP_000721309.1, WP_000721306.1, WP_063852626.1, WP_063852617.1, WP_000721310.1, WP_063851348.1, WP_012655867.1, WP_063852710.1, and WP_000725529.1.
We used customized R scripts to create visualizations. These included plots depicting patient metadata, invasive status, methicillin resistance, and cluster persistence. Additionally, we generated swimmer plots and annotated NICU floor plans using separate customized R scripts.
Transmission cluster source analysis
We conducted two complementary analyses to infer the sources and timing of transmission clusters. Molecular clock analysis was performed to estimate the genomic emergence of transmission clusters. For genomes identified as belonging to the same strain, we generated whole-genome alignments and masked putative recombination regions using ClonalFrameML v1.1360. Time to the most recent common ancestor (tMRCA) was estimated using BEAST v2.7.661, employing an HKY substitution model and a strict molecular clock. The analysis incorporated a coalescent constant population model with clock rates following log-normal distributions (1.0E−6 to 1.0E−7) as Bayesian priors. Markov Chain Monte Carlo (MCMC) chains were run up to 500 million steps to ensure effective sample sizes exceeded 200 for all parameters. The resulting phylogenetic trees were generated in NEXUS format and can be accessed at https://github.com/qianxuans/CHOP-NICU-Scripts/tree/main/Molecular_Clock_Analysis/BEAST2_Tree_Visualization/nexus.
To complement the inference of cluster source, we used TransPhylo24 to infer the transmission trees and to estimate the medoids using the timed phylogenies inferred with BEAST v2.7.661. The 95% confidence intervals were calculated as the 2.5th and 97.5th percentiles of inferred transmission index dates, weighted by sampling frequency in the posterior distribution. We estimated the shape and scale parameters of the gamma-distributed generation time by fitting observed time intervals between primary colonization events and subsequent secondary colonization events in previously uncolonized patients within identified transmission clusters (Extended Data Fig. S7). MCMC chains were run for 1,000,000 iterations to ensure effective sample sizes exceeded 100 for all parameters. Transmission index date confidence intervals and medoids were calculated from inferTTree() results after discarding the first 10% as burn-in. Colored phylogenetic trees, transmission trees, and outbreak plots showing sampled versus unsampled cases are provided in Supplementary Data 5–7.
Spatial distance rescaled phylogenetic analysis
For each cluster containing at least four genomes with a genomic maximum-likelihood (ML) phylogeny, pairwise spatial distances between all genomes were calculated using the Pythagorean theorem based on collection site coordinates from floor plans, generating a Naïve spatial distance matrix measured in meters.
A spatial-distance-rescaled phylogeny was constructed using the nnls.tree() function from the phangorn package62, with the Naïve spatial distance matrix and genomic ML phylogeny as inputs. This function preserves the ML phylogeny topology while rescaling branch lengths with spatial distances using ordinary least squares (OLS) regression to minimize the residual sum of squares (RSS).
The phylogenetic rescaled spatial distance matrix for each cluster was computed using the cophenetic.phylo() function in R package ape54. To calculate the accumulated distance through days post-initial detection, samples collected each day were compared to all previously collected samples using phylogenetic rescaled spatial distances. Single linkage clustering was applied to determine new distances added daily, where the minimum distance between each new sample and all previously collected samples represented the new distance contribution for that day. This approach corrects for genomes in a cluster that were isolated close to each other in the NICU but are not the closest relatives in the phylogeny.
Statistical analysis
We examined the persistence in the NICU, patient numbers, and travel distances of transmission clusters within the NICU. To evaluate statistical significance for patient numbers, persistence, and the total spatial distance-rescaled branch length between colonizing and invasive clusters, we applied the two-sided nonparametric Mann–Whitney–Wilcoxon test with continuity correction using R’s built-in functions, without adjustment for multiple comparisons. Linear regression models were fitted using the lm() function from the R stats package to assess the relationships between total spatial distance rescaled branch length, persistence in the NICU, and patient count. Additionally, we employed linear mixed-effects models via the lmer() function from the lme4 package to model the relationships between accumulated distance, accumulated patient numbers, and days post-initial detection, with cluster as a random effect63. Models were fitted using restricted maximum likelihood (REML), and likelihood ratio tests with maximum-likelihood estimation were used to compare models with and without interaction terms.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing reads and genome assemblies generated in this study have been deposited in the NCBI database under BioProjects PRJNA1218714, PRJNA1390285, and PRJNA1219099 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1218714; https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1390285; https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1219099). The patient-level clinical data (including patient movement, treatment team assignment, admission, and discharge dates) underlying Figs. 3, 4b, and Supplementary Data S1–S3 are available under restricted access due to patient privacy. Access can be obtained by contacting L.S. or the corresponding authors (J.P.Z., A.M.M., or P.J.P.), subject to institutional review board approval and a data use agreement. Data will be made available to qualified researchers for legitimate research purposes. Requests will receive a response within 30 days. Access will be granted for a period to be determined based on the study's needs, following approval. The processed source data generated in this study are provided in the Source Data file. The processed source data generated in this study are provided in the Source Data file. Source data are provided with this paper.
Code availability
Custom analytical code used for data analysis and visualization is available at https://github.com/qianxuans/CHOP-NICU-Scripts and archived in Zenodo (https://doi.org/10.5281/zenodo.18177031)64. The ready-to-use pipeline for clonality determination is publicly available for download at https://github.com/microbialARC/THRESHER.
References
Lake, J. G. et al. Pathogen distribution and antimicrobial resistance among pediatric healthcare-associated infections reported to the National Healthcare Safety Network, 2011–2014. Infect. Control Hosp. Epidemiol. 39, 1–11 (2018).
Hsu, H. E. et al. Health care-associated infections among critically ill children in the US, 2013–2018. JAMA Pediatr. 174, 1176–1183 (2020).
Stoll, B. J. Neurodevelopmental and growth impairment among extremely low-birth-weight infants with neonatal infection. JAMA 292, 2357 (2004).
Ericson, J. E. et al. Burden of invasive Staphylococcus aureus infections in hospitalized infants. JAMA Pediatr.169, 1–8 (2015).
Shane, A. L. et al. Methicillin-resistant and susceptible Staphylococcus aureus Bacteremia and meningitis in preterm infants. Pediatrics 129, 914 (2012).
Zervou, F. N., Zacharioudakis, I. M., Ziakas, P. D. & Mylonakis, E. MRSA colonization and risk of infection in the neonatal and pediatric ICU: a meta-analysis. Pediatrics 133, 1015 (2014).
Flannery, D. D., Edwards, E. M., Coggins, S. A., Horbar, J. D. & Puopolo, K. M. Late-onset sepsis among very preterm infants. Pediatrics (Evanston) 150, 1 (2022).
Horbar, J. D. et al. Mortality and neonatal morbidity among infants 501 to 1500 grams from 2000 to 2009. Pediatrics 129, 1019–1026 (2012).
Akinboyo, I. C. et al. SHEA neonatal intensive care unit (NICU) white paper series: practical approaches to Staphylococcus aureus disease prevention. Infect. Control Hosp. Epidemiol. 41, 1251–1257 (2020).
Yokoe, D. S. et al. A compendium of strategies to prevent healthcare-associated infections in acute care hospitals: 2014 updates. Infect. Control Hosp. Epidemiol. 35, 967–977 (2014).
Petersen, R. Y. et al. Effects of discontinuation of weekly surveillance testing on methicillin-resistant Staphylococcus aureus in the NICU. Am. J. Perinatol. 41, e1337–e1343 (2024).
Price, J. R. et al. Transmission of Staphylococcus aureus between health-care workers, the environment, and patients in an intensive care unit: a longitudinal cohort study based on whole-genome sequencing. Lancet Infect. Dis. 17, 207–214 (2017).
Coll, F. et al. Longitudinal genomic surveillance of MRSA in the UK reveals transmission patterns in hospitals and the community. Sci. Transl. Med. 9, 413 (2017).
Coll, F. et al. Definition of a genetic relatedness cutoff to exclude recent transmission of meticillin-resistant Staphylococcus aureus: a genomic epidemiology analysis. Lancet Microbe 1, e328–e335 (2020).
Tong, S. Y. C. et al. Genome sequencing defines phylogeny and spread of methicillin-resistant Staphylococcus aureus in a high transmission setting. Genome Res. 25, 111–118 (2015).
Price, J. R. et al. Whole-genome sequencing shows that patient-to-patient transmission rarely accounts for acquisition of Staphylococcus aureus in an Intensive Care Unit. Clin. Infect. Dis. 58, 609–618 (2014).
Annavajhala, M. K. et al. Genomic and epidemiological features of two dominant methicillin-susceptible Staphylococcus aureus clones from a neonatal intensive care unit surveillance effort. mSphere 7, e0040922 (2022).
Harris, S. R. et al. Whole-genome sequencing for analysis of an outbreak of meticillin-resistant Staphylococcus aureus: a descriptive study. Lancet Infect. Dis. 13, 130–136 (2013).
Nurjadi, D. et al. Surveillance for colonization, transmission, and infection with methicillin-susceptible Staphylococcus aureus in a Neonatal Intensive Care Unit. JAMA Netw. Open 4, e2 (2021).
Knox, J., Uhlemann, A. & Lowy, F. D. Staphylococcus aureus infections: transmission within households and the community. Trends Microbiol. 23, 437–444 (2015).
She, Q. et al. Local Database of 349 Pediatric S. aureus Genomes. Zenodo, 3 Feb. 2025, https://doi.org/10.5281/zenodo.14795788 (2025).
Theiller, E. & A. Moustafa. S.aureus - WhatsGNU Ortholog Database. Zenodo, 28 Aug. 2024, https://doi.org/10.5281/zenodo.13388052 (2024).
Bouiller, K., Bertrand, X., Hocquet, D. & Chirouze, C. Human infection of methicillin-susceptible Staphylococcus aureus CC398: a review. Microorganisms (Basel) 8, 1737 (2020).
Didelot, X., Fraser, C., Gardy, J. & Colijn, C. Genomic infectious disease epidemiology in partially sampled and ongoing outbreaks. Mol. Biol. Evol. 34, 997–1007 (2017).
Jackson, C. Multi-state models for panel data: the msm package for R. J. Stat. Softw. 38, 1–28 (2011).
Zervou, F. N., Zacharioudakis, I. M., Ziakas, P. D. & Mylonakis, E. MRSA colonization and risk of infection in the neonatal and pediatric ICU: a meta-analysis. Pediatrics 133, e1015–e1023 (2014).
Shane, A. L. et al. Methicillin-resistant and susceptible Staphylococcus aureus bacteremia and meningitis in preterm infants. Pediatrics 129, e914–e922 (2012).
Achermann, Y. et al. Epidemiology of methicillin-susceptible Staphylococcus aureus in a neonatology ward. Infect. Control Hosp. Epidemiol. 36, 1305–1312 (2015).
Hawken, S. E. et al. Threshold-free genomic cluster detection to track transmission pathways in health-care settings: a genomic epidemiology analysis. Lancet Microbe 3, e652–e662 (2022).
Köser, C. U. et al. Rapid whole-genome sequencing for investigation of a neonatal MRSA outbreak. N. Engl. J. Med. 366, 2267–2275 (2012).
Layer, F. et al. Molecular typing of toxic shock syndrome toxin-1-and Enterotoxin A-producing methicillin-sensitive Staphylococcus aureus isolates from an outbreak in a neonatal intensive care unit. Int. J. Med. Microbiol. 305, 790–798 (2015).
Cremers, A. J. et al. Surveillance-embedded genomic outbreak resolution of methicillin-susceptible Staphylococcus aureus in a neonatal intensive care unit. Sci. Rep. 10, 2619 (2020).
Hatcher, J. et al. Healthcare-worker-associated outbreak of Panton–Valentine-leukocidin-producing meticillin-sensitive Staphylococcus aureus in a large neonatal unit in London: successful targeted suppression therapy following failure of mass suppression therapy. J. Hosp. Infect. 122, 148–156 (2022).
Madera, S. et al. Prolonged silent carriage, genomic virulence potential and transmission between staff and patients characterize a neonatal intensive care unit (NICU) outbreak of methicillin-resistant Staphylococcus aureus (MRSA). Infect. Control Hosp. Epidemiol. 44, 40–46 (2023).
Slingerland, B. C. et al. Neonatal Staphylococcus aureus acquisition at a tertiary intensive care unit. Am. J. Infect. Control 48, 1023–1027 (2020).
CDC NHSN. Bloodstream Infection (BSI) Event (CDC NHSN, 2025).
Clarke, E. L. et al. Sunbeam: an extensible pipeline for analyzing metagenomic sequencing experiments. Microbiome 7, 46 (2019).
Prjibelski, A., Antipov, D., Meleshko, D., Lapidus, A. & Korobeynikov, A. Using SPAdes de novo assembler. Current Protocols Bioinform. 70, e102 (2020).
Li, D., Liu, C., Luo, R., Sadakane, K. & Lam, T. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Eren, A. M. et al. Community-led, integrated, reproducible multi-omics with anvi’o. Nat. Microbiol. 6, 3–6 (2020).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Schwengers, O. et al. Bakta: rapid and standardized annotation of bacterial genomes via alignment-free sequence identification. Microb. Genom. 7, 11 (2021).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 1–21 (2020).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Lanave, C., Preparata, G., Sacone, C. & Serio, G. A new method for calculating evolutionary substitution rates. J. Mol. Evol. 20, 86–93 (1984).
Yang, Z. Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J. Mol. Evol. 39, 306–314 (1994).
Minh, B. Q., Nguyen, M. A. T. & Haeseler, vonA. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 30, 1188–1195 (2013).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Wickham, H., Chang, W. & Wickham, M. H. Package ‘ggplot2’. Create elegant data visualisations using the grammar of graphics. Version 2, 1–189 (2016).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Moustafa, A. M. & Planet, P. J. WhatsGNU: a tool for identifying proteomic novelty. Genome Biol. 21, 58 (2020).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Struyf, A., Hubert, M. & Rousseeuw, P. Clustering in an object-oriented environment. J. Stat. Softw. 1, 1–30 (1997).
Seemann, T. snippy: Fast Bacterial Variant Calling from NGS Reads https://github.com/tseemann/snippy (2015).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 49, 10 (2021).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Didelot, X. & Wilson, D. J. ClonalFrameML: efficient inference of recombination in whole bacterial genomes. PLoS Comput. Biol. 11, e1 (2015).
Bouckaert, R. et al. BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 15, e1–e1006650 (2019).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Bates, D. et al. Package ‘lme4’. Convergence 12, 2 (2015).
She, Q. CHOP NICU S. aureus Transmission Analysis Scripts. Zenodo, 7 Jan. 2026, https://doi.org/10.5281/zenodo.18177031 (2026).
Acknowledgements
We thank the NICU staff, the NICU Safety Quality Specialist nursing team (Theresa O’Connor, AmyRose Neppach, Ariana Lowe, Kayla Mason, and Nakisha Barr), Lindsay Brim, Infection Prevention and Control, and the Infectious Disease Diagnostics Laboratory at the Children’s Hospital of Philadelphia for their assistance with this study. This work was funded and supported by The Center for Microbial Medicine at the Children’s Hospital of Philadelphia. L.S., P.J.P., A.M.M., and J.P.Z. were supported by 1R01AI185544. Q.S. was supported by the Chappell Culpeper Family Foundation Fellowship in the Center for Microbial Medicine at the Children's Hospital of Philadelphia.
Author information
Authors and Affiliations
Contributions
Q.S. performed formal analysis, developed analytical methodology (code and pipelines), created visualizations, and wrote the original draft. L.S. initiated the comprehensive S. aureus colonization surveillance program, coordinated routine sampling of S. aureus-positive blood culture isolates, contributed to project administration and conceptualization, and wrote the original draft. E.T., B.E.G., T.N., S.M.J., A.A., and A.M.M. performed whole-genome sequencing. A.F. performed data analysis. R.F.P. coordinated clinical sampling and isolate processing. E.H., K.M.P., R.N.B., S.E.C., K.A.G., K.P.S., R.M.H., and M.Z.D. provided critical input on methodology and contributed to manuscript review and editing. L.S., J.P.Z., A.M.M., and P.J.P. contributed to project administration, conceptualization, methodology, formal analysis, visualization, supervision, funding acquisition, validation, and writing (original draft and review & editing). All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.P.Z. has consulted in the past for Vedanta Biosciences, Inc. and AstraZeneca. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Talima Pearson, Jason Kwong, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
She, Q., Srinivasan, L., Theiller, E. et al. Rapid dissemination of Staphylococcus aureus in the neonatal intensive care unit is associated with invasive infection. Nat Commun 17, 2550 (2026). https://doi.org/10.1038/s41467-026-69074-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69074-z