Abstract
Cells regulate processes through protein interaction networks. Most chemically induced dimerization (CID) systems respond to exogenous molecules, limiting integration with endogenous signaling. Here, we repurpose nuclear receptor (NR) ligand-binding domains (LBDs) and coactivators to develop hormone- or clinically approved drug-responsive CIDs. Using the LBDs of TRβ, VDR, RARγ, ERβ, and GR2 with a TIF2 coactivator peptide, we constructed CIDs responsive to triiodothyronine, vitamin D, retinoic acid, estrogen, cortisol, and their antagonists. These CIDs enable two-input transcriptional switches for gene regulation. Furthermore, we design hormone-responsive liquid-liquid phase-separated (LLPS) condensates that strongly amplify transcription when exceeding a critical interaction threshold. These functional LLPS condensates provide a tunable platform for transcriptional control with up to several hundred-fold activation. Our findings offer an approach for integrating synthetic biology with physiological signaling, advancing applications in gene circuits, biosensing, and therapeutics through ligand-controlled LLPS formation.
Similar content being viewed by others
Introduction
Cells use networks of protein interactions to integrate and process external and internal signals into appropriate cellular responses. Advances in synthetic biology enabled the manipulation of these networks, paving the way for innovative tools for time-resolved control of cellular processes. Particularly, the development of a small molecule-responsive molecular switches, termed chemically induced dimerization (CID) systems, advanced this field by enabling chemical control of protein interactions1,2. These systems typically consist of heterodimerization domains, which reassemble only in the presence of a target small molecule3,4,5. Most CID systems are designed to respond to small molecules, with recent designs incorporating clinically-approved drugs to enhance their therapeutic applicability5,6,7,8,9,10,11,12,13,14. The development of actuators that respond to endogenous ligands such as hormones offers the construction of synthetic circuits that integrate with the body’s physiological signaling network. Therefore, the construction of CIDs that respond to endogenous small molecules could be used as biosensors and theragnostic devices by leveraging the existing signaling pathways.
Human nuclear receptors (NRs) are a superfamily of 48 ligand-activated transcription factors that regulate a wide range of physiological processes, including inflammation, immunity, hormonal signaling, and metabolic homeostasis15,16. This ligand-dependent activation is highly specific, as NRs bind structurally diverse natural ligands, such as hormones, vitamins, and fatty acids, enabling them to generate distinct biological outputs in response to different input signals17. NRs have modular composition, comprising a DNA- (DBD) and a ligand-binding domain (LBD). The LBD contains the activation function-2 (AF-2) region, whose function is regulated by ligand binding and the conformational state of helix 12 (H12)18. Ligand binding induces allosteric changes in H12, determining whether the receptor recruits coactivators or corepressors. An agonist binding stabilizes H12 in an active conformation, promoting interactions with coactivator proteins such as steroid receptor coactivator-2 (SRC-2) and transcriptional intermediary factor 2 (TIF2)19,20. Coactivators interact with NRs via an α-helix containing an LXXLL motif (L = leucine, X = any amino acid), which fits into a hydrophobic groove formed by the rearranged H12 of LBD21,22. NRs are also major drug targets, leading to the development of clinically-approved synthetic agonists and antagonists that function as molecular switches to modulate NR activity for therapeutic purposes23. Several reports previously utilized NRs and their LBDs to construct ligand-controlled gene expression systems in mammalian cells24,25,26.
Further, NRs and their LBDs have been repurposed to create synthetic transcription factors by fusing LBDs to alternative DNA-binding domains, enabling ligand-dependent regulation of synthetic gene circuits27,28,29. These synthetic transcription factors rely on the endogenous recruitment of coactivators and corepressors to modulate transcription. Beyond transcriptional control, LBDs have also been integrated into synthetic sensor-actuator devices. For instance, a chemically induced dimerization (CID) system was designed using two LBDs that dimerize upon binding a synthetic hybrid or bifunctional ligand28. In another approach, ligand-induced homodimerization of LBDs was employed to construct biosensors based on bioluminescence resonance energy transfer29,30,31.
Here, we repurposed the endogenous interaction between the nuclear receptor (NR) ligand-binding domains (LBDs) and coactivator proteins to develop chemically induced dimerization (CID) systems that respond to endogenous signals such as hormones. Specifically, we demonstrate that the interaction between the LBDs of TRβ, VDR, RARγ, ERβ, and GR2 and an LXXLL motif-containing peptide derived from the TIF2 coactivator can be leveraged to construct CIDs responsive to triiodothyronine (T3), vitamin D, retinoic acid, estrogen, and cortisol, respectively. Using these CIDs, we designed two-input sensors capable of either activating or repressing gene expression in response to specific agonist/antagonist signals. Additionally, we engineered synthetic, NR ligand-responsive liquid-liquid phase-separated condensates based on NRs above a certain threshold of peptide repeats, which enhanced the selected gene expression by several hundred-fold. The results show the potential of LBD-based CIDs to regulate cellular processes in response to physiological signals, enabling the development of precisely controlled, hormone-responsive gene circuits for mammalian cell engineering and autonomously regulated synthetic systems.
Results
Design and validation of LBDs-based CIDs
To engineer a ligand-responsive chemically induced dimerization (CID) system, we hypothesized that the natural interaction between the ligand-binding domain (LBD) of nuclear receptors (NRs) and their coactivators could be used for constructing CIDs responsive to hormones and clinically approved drugs in mammalian cells. As a proof of concept, we initially selected the LBD of the thyroid receptor beta (TRβ), which exhibits high-affinity binding to the triiodothyronine (T3) hormone and a coactivator peptide derived from the TIF2, containing a canonical two-turn α-helix LXXLL motif that interacts with LBD32. To evaluate the interaction between TRβ-LBD and the TIF2 peptide in mammalian cells, a split luciferase assay was employed. Specifically, the LBD of TRβ and TIF2 peptide were fused to the N- and C-terminal fragments of split luciferase (nLuc and cLuc), respectively (Fig. 1a). Upon T3 binding, the LBD undergoes a conformational change, exposing the interaction surface for binding the TIF2 peptide and reconstituting luciferase activity. In HEK293T cells, T3 stimulation (1 µM) led to a significant increase in a luciferase activity compared to unstimulated controls, demonstrating efficient dimerization and reconstitution of a split luciferase (Fig. 1b). Additionally, the response kinetics was rapid, with luciferase activity plateauing within 10 min of T3 addition to cells (Fig. 1c). Next, we evaluated the system’s ability to regulate protein subcellular localization. To this end, we targeted a TagBFP-tagged TRβ-LBD to the plasma membrane by fusing it with the Lyn11 localization tag (Lyn11:TagBFP:TRβ), while TIF2 fused to YFP was localized in the cytosol (TIF2:YFP) (Fig. 1d). Upon T3 addition, the cytosolic TIF2-YFP translocated to the plasma membrane (Fig. 1e), confirming ligand-induced recruitment.
a Schematic representation of the LBD-TIF2 CID system. Ligand binding induces a conformational change of H12 in the LBD, exposing the TIF2 binding site. Ligand-dependent dimerization of LBD and TIF2 peptide results in split luciferase reporter reconstitution (nLuc and cLuc) and a measurable output signal. b Ligand-induced luciferase reconstitution. HEK293T cells were transfected with TRβ:cLuc (50 ng) and nLuc:TIF2 (50 ng) and stimulated with triiodothyronine (T3, 1 µM) for 24 h. Cells expressing only Renilla luciferase (10 ng) served as a negative control. Fold induction values (stimulated vs. non-stimulated) are shown above the graphs. Data represent means ± s.d. from n = 8 biological replicates pooled from two independent experiments. c Kinetics of TRβ-LBD dimerization in response to T3 (1 µM). Data represent means ± s.d. from n = 4 biological replicates, representative of two independent experiments. d Schematic of ligand-induced recruitment of TIF2 to the plasma membrane. TRβ-LBD is membrane-targeted via a Lyn11 tag (Lyn11-TagBFP-TRβ), while mCitrine-tagged TIF2 remains cytosolic. Cytosol to PM translocation of TIF2 by dimerization of LBDs indicates successful recruitment. e Confocal microscopy images showing mCitrine:TIF2 (50 ng) colocalization with Lyn11-TagBFP-TRβ (100 ng) 24 h after T3 (1 µM) or DMSO treatment. Corresponding fluorescence intensity graphs for mCitrine and TagBFP are shown. Images are representative of two independent experiments. Scale bar = 20 µm. f Scheme of ligand-induced recruitment of TIF2-5ptaseOCRL to its membrane-bound dimerization partner, Lyn11-TagBFP-TRβ, leading to PIP₂ hydrolysis and GFP-C1-PLCδ-PH dissociation from the plasma membrane. g Representative images show HEK293T cells co-transfected with TIF2-5ptaseOCRL (50 ng), Lyn11-TagBFP-TRβ (100 ng), and GFP-C1-PLCδ-PH (50 ng) before and 60 s after T3 (1 µM) stimulation. Corresponding GFP-C1-PLCδ-PH cytoplasmic intensity data (n = 6) are plotted. Images are representative of two independent experiments. Scale bar = 20 µm. Statistical comparisons in (b) were performed using a two-sided unpaired t-test with Welch’s correction (****P < 0.0001; **P < 0.01). All schematics were created using Inkscape (ver. 1.2.1).
To demonstrate the functional utility of the LBD-CID system, we applied it to regulate polyphosphoinositide (PIP) metabolism (Fig. 1f). Plasma membrane-resident secondary lipid messengers, such as PI(4,5)P2 (PIP2) and its precursor PI(4)P, are in rapid flux during signal transduction33. To achieve membrane-localized PIP2 depletion, we co-expressed a plasma membrane-targeted TRβ-LBD and a cytosolic phosphoinositide 5-phosphatase domain of OCRL (5-ptaseOCRL) fused to the TIF2 peptide in HEK293T cells. In the absence of T3, PIP2 levels remained stable, as monitored by the membrane localization of the PIP2 fluorescent biosensor GFP-C1-PLCδ-PH at the (Fig. 1g, Supplementary Fig. 1, Supplementary Movies 1 and 2). Upon T3 stimulation, 5-ptaseOCRL was recruited to the plasma membrane, resulting in a rapid decrease in PIP2 levels, evidenced by the redistribution of the PIP2 fluorescent sensor from the plasma membrane to the cytosol. To monitor enzyme translocation, we generated a fluorescent fusion protein, mCherry:TIF2:5-ptaseOCRL. While membrane localization of this construct was not clearly detectable on the same timescale as biosensor redistribution (Supplementary Fig. 1b), likely due to limited expression, it became apparent at later timepoints (Supplementary Fig. 1c). To further validate functional recruitment, we performed a control time-lapse imaging using a cytosolic TIF2-YFP fusion protein (Supplementary Figure 1d). Upon T3 stimulation, we observed rapid recruitment to the plasma membrane, with visible enrichment occurring within 60 s (Supplementary Figure 1d). Additionally, to exclude the possibility that PH domain redistribution is driven by T3 treatment alone, we transfected cells with only the PH domain biosensor (without 5ptaseOCRL constructs) and stimulated with T3 for 24 h, where no translocation of the biosensor was observed (Supplementary Figure 1e). These results highlight that the LBD-CID system can be used for modulating intracellular localization.
Design of ON/OFF LBD-CID systems
Next, we wanted to determine if our LBD-CID systems could integrate two ligand inputs with opposing effects to achieve a more precise dynamic control over cellular processes. Dual-input systems allow for ligand-mediated switching between the “on” and “off” states, enabling versatility in modulating protein-protein interactions. LBDs, in combination with their interacting coactivator TIF2 peptide, are ideal for this purpose, as many LBDs can bind both an agonist ligands (inducing the “on” state via TIF2 association) and competitive antagonist ligands (inducing the “off” state via dissociation of the TIF2 peptide) (Fig. 1a). We first implemented the GR2-LBD, which has a well-characterized agonist cortisol (COR) and an antagonist mifepristone (MIF) ligand. To assess ligand-induced assembly and disassembly of the dimerization domains, we utilized a split luciferase reporter assay in HEK293T cells. As designed, the addition of cortisol led to a substantial increase in luciferase activity compared to non-stimulated controls, demonstrating efficient cortisol-mediated interaction between GR2-LBD and TIF2 peptide (Fig. 2b). To evaluate antagonist-mediated inhibition, we added MIF in the presence of cortisol. Mifepristone effectively inhibited cortisol-induced luciferase activity, as indicated by a marked reduction in luminescence (Fig. 2c). Time-course experiments further demonstrated rapid and reversible control of the system: COR stimulation induced the luciferase activity, which was subsequently inhibited by the addition of MIF (Fig. 2d). A second, higher-dose of cortisol restored the activity and demonstrated the reversibility of the system. These results validate the potential of GR2-LBD to function as a dual-input CID system with tight control over interaction dynamics. We next applied the same design principles to the ERβ-LBD, which binds an agonist estradiol (EST) and an antagonist 4-hydroxytamoxifen (4-OHT). Stimulation with estradiol triggered luciferase reconstitution, resulting in increased luminescence (Fig. 2e). In the case of this NR, however, we observed high constitutive interaction between the ERβ-LBD and TIF2 peptide already in the absence of EST. This constitutive interaction was efficiently suppressed upon the addition of 4-OHT, leading to a decline in the luciferase activity, demonstrating an antagonist-driven dissociation of the complex. Importantly, subsequent re-stimulation with EST reversed the inhibitory effect of 4-OHT, demonstrating reversibility also in this system (Fig. 2f). Additionally, we combined ERβ and GR2 to localize TIF2-YFP to different subcellular locations (Supplementary Fig. 2). COR recruited TIF2-YFP to membrane-localized GR2, while EST induced colocalization with mitochondrially targeted ERβ. These results highlight the versatility and precision of dual-input LBD-CID systems utilizing the same TIF2 peptide in dynamically regulating protein-protein interactions.
a Schematic representation of an LBD-CID system responding to two types of input signals. The system can be switched to the ON and OFF states depending on the presence of an agonist or antagonist ligand. b Ligand-induced split luciferase reconstitution. HEK293T cells were transfected with GR2:cLuc (50 ng) and nLuc:TIF2 (50 ng) and stimulated with cortisol (COR, 1 µM) for 24 h. Cells transfected with only Renilla luciferase (10 ng) served as a negative control. c Luciferase activity of the GR2-CID system after treatment of transfected HEK293T cells with cortisol (COR, 1 µM), mifepristone (MIF, 10 µM), or both for 24 h. d Fast kinetics of the sequential luciferase activation and suppression in the GR2-LBD system in response to COR (1 µM), sequential addition of MIF (10 µM), and finally the addition of excess COR (100 µM). d Kinetics of luciferase activation and suppression in the GR2-LBD system following stimulation with COR (1 µM), sequential addition of MIF (10 µM), and final addition of excess COR (100 µM). e Ligand-induced split luciferase reconstitution in HEK293T cells transfected with ERβ:cLuc (50 ng) and nLuc:TIF2 (50 ng), followed by stimulation with estradiol (EST, 1 µM) (left) or 4-hydroxitamoxifen (4-OHT, 1 µM) (right) for 24 h. Cells expressing only Renilla luciferase (10 ng) served as a negative control. f Kinetics of ERβ-LBD dimerization in response to 4-OHT (1 µM) and sequential addition of EST (10 µM). Data in (d) and (f) represent means ± s.d. from n = 4 biological replicates, as a representative of two independent experiments. Data in (b), (c) and (e) represent means ± s.d. from n = 8 biological replicates pooled from two independent experiments. The firefly luciferase units (nFluc) were calculated by normalizing the luciferase activity of each time-point to the value/signal of non-stimulated conditions. Statistical comparisons in were performed using a two-sided unpaired t-test with Welch’s correction (****P < 0.0001; **P < 0.01). All schematics were created using Inkscape (ver. 1.2.1).
LBD-CID enables two-input transcriptional control
The ability of LBD-CID systems to toggle between the ON and OFF states through agonists and antagonists represents a versatile platform for precise control of cellular processes. This dynamic modulation is could be particularly beneficial for applications requiring fine-tuned transcriptional regulation. To evaluate the potential of LBD-CID systems in regulating gene expression, we employed a split CRISPR/dCas9-based transcriptional activation9,34. Initially, we designed a transcriptional activator based on GR2-TIF2 interaction that is responsive to the agonist cortisol (COR) and antagonist mifepristone (MIF) to regulate transcriptional activity of a reporter gene in HEK293T cells. The GR2-LBD domain was genetically fused to a strong chimeric VPR activation domain (VP64-p65-Rta)35 while TIF2 peptide was fused to dCas9 DNA-binding domain. Ligand-induced dimerization between GR2 and TIF2 reconstituted a functional transcriptional activator, enabling transcription of the luciferase reporter gene with ten binding sites for the dCas9:gRNA complex (Fig. 3a). Stimulation with cortisol failed to significantly activate the reporter gene when only a single TIF2 peptide was fused to dCas9 (Fig. 3b).
a Schematic representation of the split transcriptional activation system. Agonist-induced recruitment of LBD:VPR to dCas9:TIF2 activates transcription of a reporter gene, while antagonist treatment displaces LBD:VPR, suppressing activation. b Transcriptional activation of a reporter with 10 binding sites (10 × bs) was tested in HEK293T cells transfected with GR2-VPR and dCas9:TIF2 and stimulated with cortisol (COR, 1 µM) for 24 h. c Increasing the number of bound GR2-VPR molecules by concatenating TIF2 repeats to the dCas9 DNA-binding domain significantly enhances transcriptional activation of a reporter plasmid containing 10 × bs for the dCas9:gRNA complex. Cells were stimulated with COR (50 nM) for 24 h. d Dose-response curve of GR2-VPR in combination with dCas9:6 × TIF2 upon stimulation with increasing concentrations of COR. e Transcriptional activation of a reporter with 10 × bs was tested using ERβ-VPR and dCas9 with varying numbers of TIF2 repeats in HEK293T cells stimulated with estradiol (EST, 100 nM) for 24 h. f Dose-response curve for the dCas9:6 × TIF2 ERβ-VPR system. g Evaluation of LBD-transcriptional activation systems orthogonality by stimulating cells transfected with individual LBD-CID with different indicated ligands. Individual data points are presented in Supplementary Fig. 3. h, i Dynamic control of transcriptional activation using agonist and antagonist ligands. Simultaneous treatment with cortisol and mifepristone (MIF) inhibits activation in the GR2 system (g), while estradiol with 4-hydroxytamoxifen (4-OHT) suppresses activation in the ERβ system (h). Agonist and antagonist ligand additions are indicated by filled circles beneath the graphs, with the corresponding expected outcomes marked by arrows. j, k Transcriptional activation of a reporter with a single binding site (1 × bs) using dCas9:6×TIF2 and either GR2-LBD (i) or ERβ-LBD (j) in response to their respective agonist ligands. Results are shown as fold activation relative to non-stimulated mock cells transfected with an empty plasmid (pcDNA3) or reporter-only control. Data represent means ± s.d. from n = 8 biological replicates pooled from two independent experiments. Conditions were compared using a two-sided unpaired t-test with Welch’s correction (****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns > 0.05). All schematics were created using Inkscape (ver. 1.2.1).
Several strategies have been developed to enhance transcriptional activation by recruiting multiple activation domains to the DNA-binding domain through dimerization motif concatenation36,37. Such designs are intended to enhance transcriptional strength via multivalent recruitment of activators at specific genomic loci. Therefore, we explored TIF2 peptide concatemerization by engineering multimeric TIF2 repeats (2 × TIF2, 4 × TIF2, and 6 × TIF2) fused to the dCas9 DNA-binding domain. This design recruited multiple GR2-VPR transcriptional activators upon ligand binding, aiming to enhance transcriptional activation. Indeed, the addition of cortisol resulted in an increase in transcriptional activity with an increasing number of TIF2 repeats (Fig. 3c). Dose-response experiments revealed that the system using dCas9:6×TIF2 achieved nanomolar sensitivity to cortisol, a concentration within the physiologically relevant range38 (Fig. 3d). We extended this strategy to an ERβ-based LBD-CID system using the agonist estradiol (EST) and the antagonist 4-hydroxytamoxifen (4-OHT). Similar to the GR2-based system, a single TIF2 peptide fused to dCas9 induced weak transcriptional activation upon estradiol stimulation (Fig. 3e). However, concatenated 6×TIF2 peptides dramatically enhanced transcriptional activation of the reporter gene, achieving robust activation within physiologically relevant estradiol concentrations (Fig. 3f)39.
To further evaluate the generalizability of the LBD-CID strategy, we also tested LBDs derived from the vitamin D receptor (VDR) and retinoic acid receptor gamma (RARγ). When fused to VPR and combined with dCas9:6×TIF2 on the reporter with ten binding sites for gRNA:dCas9 complex, both VDR and RARγ LBDs supported specific ligand-dependent transcriptional activation in response to estradiol (EST), cortisol (COR), all-trans retinoic acid (ATRA), and calcitriol (CAL) (active form of vitamin D) (Fig. 3g, Supplementary Figure 3). Importantly, for both GR2- and ERβ-based transcriptional systems, which have well-characterized agonist/antagonist pairs, the simultaneous addition of antagonist ligands (MIF for GR2 and 4-OHT for ERβ) effectively inhibited reporter gene expression, even in the presence of the corresponding agonists (Fig. 3h, i). LBD-CID systems can therefore dynamically toggle between activation and repression states in response to dual ligand inputs. Finally, we assessed whether transcriptional activation could be achieved using a reporter containing only a single binding site for the dCas9:gRNA complex. Despite using dCas9 fused to 6 × TIF2 to maximize recruitment of LBD–VPR activators, agonist stimulation resulted in significantly weaker transcriptional activation compared to reporters containing multiple binding sites in both GR2- and ERβ-based systems (Fig. 3j, k).
Engineering multivalent liquid-liquid phase-separating LBD-CIDs
Given the steep increase in transcriptional activation observed when four or more TIF2 peptides were concatenated, compared to constructs containing only one or two copies, we suspected that a threshold in multivalency may trigger a cooperative state transition, possibly via liquid-liquid phase separation (LLPS). Many natural transcription factors utilize phase separation with coactivators to form biomolecular condensates, which enhance transcriptional activation by concentrating transcriptional machinery at specific genomic loci40. Inspired by this, we wanted to test if LBD-CID systems are capable of liquid-liquid phase separation (LLPS) upon the addition of the ligand.
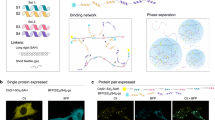
We first tested whether interactions between the GR2- and ERβ-LBDs and concatenated TIF2 peptides were able to drive condensate formation. Constructs containing 1×, 2×, or 6× TIF2 repeats fused to TagBFP were co-expressed with YFP-tagged LBDs in the presence of their respective agonists. Neither 1×TIF2 nor 2×TIF2 showed any visible condensate formation, while 6×TIF2 supported weak formation of condensates (Supplementary Fig. 4a, b). To enhance multivalency to robustly drive phase separation, we decided to use a designed coiled-coil (CC) peptide scaffold composed of five repeats of P3 (5×P3). P3 peptide forms a CC heterodimerdimer with a complementary P4 peptide, which was fused to six tandem TIF2 repeats (6×TIF2). We hypothesized that this design could provide higher multivalency concentrating TIF2 interaction motifs, necessary for robust LLPS formation, while the homodimerization of GR2- and ERβ-LBDs upon agonist binding supports condensate formation (Fig. 4a)41,42. To confirm LLPS formation, we coexpressed YFP-tagged GR2-LBD (YFP-GR2), TagBFP-tagged P4:6×TIF2 (P4:BFP:6×TIF2), and miRFPnano3-tagged 5×P3 (mRFPnano3:5×P3) in the cytoplasm of HEK293T cells. Upon cortisol stimulation, the GR2-based constructs formed distinct, cytoplasmic puncta, indicative of LLPS (Fig. 4b, Supplementary Figure 4c). These condensates were absent in unstimulated GR2-LLPS cells and were also absent when the cells were treated with cortisol and its antagonist mifepristone (MIF). In contrast, the ERβ-LLPS system exhibited constitutive protein droplet formation even in the absence of the agonist ligand (Fig. 4c). This observation aligns with the above mentioned results showing an intrinsic interaction between ERβ and TIF2, which can promote phase separation independent of ligand binding (Fig. 2e). Notably, LLPS was observed even when cells were cultured in medium supplemented with 10% charcoal-stripped FBS to remove possible estrogens in the media (Supplementary Figure 5). The addition of the antagonist 4-hydroxytamoxifen (4-OHT) disrupted these preformed condensates, confirming ligand-dependent modulation of phase separation dynamics (Fig. 4d). Additionally, when we tested a TagBFP-tagged P4 construct with only a single TIF2 motif (P4:BFP:1×TIF2), we did not observe LLPS formation, indicating that multivalency is essential for the formation of phase-separated droplets (Supplementary Fig. 6).
a Schematic representation of constructs used for ligand-induced liquid-liquid phase separation (LLPS). b Confocal microscopy images of HEK293T cells expressing GR2:YFP (50 ng), P4:TagBFP:6×TIF2 (50 ng), and miRFPnano3:5 × P3 (50 ng), taken 24 h after treatment with cortisol (COR, 100 nM), mifepristone (MIF, 10 μM), or DMSO (non-stimulated). Scale bar = 10 µm. Confocal microscopy images of HEK293T cells expressing ERβ:YFP (50 ng), P4:TagBFP:6×TIF2 (50 ng), and miRFPnano3:5 × P3 (50 ng), taken 24 h after treatment with (c) estradiol (EST, 1 µM), d 4-hydroxytamoxifen (4-OHT, 100 nM), or DMSO (non-stimulated). Scale bars: 10 µm (c, d). e Fluorescence recovery after photobleaching (FRAP) analysis of GR2- and ERβ-based LLPS. The plots show normalized YFP fluorescence recovery of GR2-YFP or ERβ-YFP (n = 4). Representative confocal images of bleached condensates are shown (right), along with acquisition times. Images also illustrate fusion events between condensates. Scale bar = 5 µm. Data represent means ± s.d. f Time-lapse confocal microscopy images showing droplet disassembly following the addition of 4-OHT (100 nM) and subsequent addition of EST (1 µM). Images are representative of two independent experiments. All schematics were created using Inkscape (ver. 1.2.1).
To evaluate the fluidity of the condensates within cells, we performed fluorescence recovery after photobleaching (FRAP) analysis on both GR2- and ERβ-based systems. The fluorescence recovery rates demonstrated the dynamic liquid nature of the droplets, with a rapid recovery time for both systems (Fig. 4e). Notably, ERβ droplets exhibited higher recovery compared to GR2 droplets, suggesting potential differences in their internal organization and fluidity. Furthermore, droplet fusion events were observed, an additional hallmark of LLPS. Treatment with 10 % 1,6-hexanediol (1,6-HD), a compound known to disrupt weak hydrophobic protein-protein interactions required for droplet formation, completely dissolved the GR2 and ERβ-based droplets (Supplementary Fig. 7)40,43. Additionally, we demonstrated the reversible control of ERβ-based LLPS (Fig. 4f). Pre-existing condensates rapidly dissolved upon treatment with an antagonist 4-OHT, and subsequent addition of an excess estradiol triggered the re-formation of droplets. Together, these results establish that LBD-CID systems can be engineered to form artificial chemically regulated biomolecular LLPS condensates.
Chemically-induced LLPS boosts transcriptional activation
To investigate whether GR2- or ERβ-LLPS condensates could enhance transcription from a reporter containing a single DNA-binding site. We utilized a previously characterized CRISPR/dCas9 system fused with multiple P3 repeats (termed cccTag) to introduce multivalency44. GR2 was fused to the trimeric VPR activation domain, and 6×TIF2 was fused to BFP (Fig. 5a). All constructs included a nuclear localization signal (NLS) to direct the proteins to the nucleus. First, we tested transcriptional activation using cccCas9 constructs fused to increasing numbers of P3 repeats (1×, 2×, 4×, and 10×) in combination with GR2-LLPS system. A single P3 segment failed to enhance transcriptional activation, likely due to insufficient multivalent interactions required for efficient transcriptional activation (Fig. 5b). However, increasing the number of P3 repeats significantly boosted transcriptional activation, with activity plateauing at four concatenated peptide repeats, leading to >200 fold activation. Next, we tested the ERβ-based LLPS system. Upon stimulation with estradiol, we observed a robust increase in transcriptional activation that also plateaued at four P3 peptide repeats (Fig. 5c). Notably, leakage of the ERβ-based system was observed at a higher number of P3 repeats, consistent with ligand-independent ERβ-TIF2 interaction. To benchmark the performance of the GR2- and ERβ-based LLPS transcriptional activators, we compared them to previously described hormone-responsive INSPIRE-CID systems45, as well as their CC-enhanced versions (cccINSPIRE) that allow multivalent recruitment of effector domains via cccCas9 (Supplementary Fig. 8). In both cortisol- and estrogen-responsive contexts, the LLPS-based systems consistently outperformed their CID-based counterparts and even surpassed the transcriptional activity of the constitutively active dCas9:VPR fusion, confirming enhanced transcriptional output of the LLPS-based gene activation strategy. As expected, both GR2- and ERβ-based LLPS gene activation systems formed distinct phase-separated LBD-YFP a P4:NLS:BFP:6xTIF2 condensates in the nuclei of transfected HEK293T cells upon stimulation with their respective agonist ligands (Fig. 5d). We found that treatment of cells with 1,6-HD caused a disassembly of nuclear puncta formed (Supplementary Fig. 9). To additionally confirm that the observed activation enhancement was due to the multivalent recruitment of activation domains, we performed control experiments using a single repeat of the TIF2 peptide. In this setup, we observed minimal transcriptional activation upon cortisol stimulation even when using multiple repeats of a P3 peptide (Supplementary Fig. 10), highlighting the necessity of multivalent interactions for robust transcriptional activation.
a Schematic representation of the engineered LLPS-based transcriptional activation system. P4:NLS:6 × TIF2 binds to cccTag-dCas9:10×P3, and agonist-mediated recruitment of LBD-VPR activation domains undergoes LLPS, leading to gene activation of a reporter with a single DNA binding site. b Reporter transcriptional activation using cccTag-dCas9 fused to increasing P3 peptide repeats (1 × P3, 2 × P3, 4 × P3, and 10 × P3) measured 24 h after cortisol (COR, 50 nM) stimulation. c Estradiol (EST, 100 nM)-induced transcriptional activation in the ERβ-based system, demonstrating increased activation with higher dCas9-P3 peptide repeats. d Representative confocal image of HEK293T cells co-expressing dCas9:10P3, P4:NLS:BFP:6xTIF2, and GR2- or ERβ:NLS:VPR:YFP, along with [ab]nt sgRNA. The image shows nuclear localization of GR2- and ERβ-based LLPS puncta after 24 h stimulation with the ligands EST (100 nM) and COR (100 nM). Scale bar = 10 µm. Corresponding fluorescence intensity graphs for mCitrine and TagBFP are shown. Images are representative of two independent experiments. Arrowheads point to the representative condensates. e, f Dynamic regulation of GR2- and ERβ-based LLPS transcriptional activation through agonist and antagonist treatment. Cortisol-induced activation is inhibited by mifepristone (MIF) (e), while estradiol-induced activation is suppressed by 4-hydroxytamoxifen (4-OHT) (f). Agonist and antagonist ligand additions are indicated by filled circles beneath the graphs, with the corresponding expected outcomes marked by arrows. g Transcriptional activation of ASCL1 endogenous gene by GR2- (left) and ERβ- (right) based LLPS systems after stimulation of HEK293T cells with indicated ligands. In all experiments, HEK293T cells were co-transfected with 50 ng of the reporter plasmid [ab]:pMin:Fluc, 25 ng of the corresponding gRNA, 25 ng of the dCas9:n×P3, 50 ng of the P4:NLS:6(1)×TIF2, and 50 ng of either ERβ:NLS:VPR or GR2:NLS:VPR encoding plasmids. Results are presented as fold activation relative to non-stimulated mock cells transfected with an empty plasmid (pcDNA3) or reporter-only control. Data represent means ± s.d. from n = 8 biological replicates pooled from two independent experiments. Conditions were compared using a two-sided unpaired t-test with Welch’s correction (****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; ns > 0.05). All schematics were created using Inkscape (ver. 1.2.1).
To investigate whether transcriptional activation correlates with nuclear LLPS condensate formation driven by the multivalency of P3 (1×, 2×, 4×, and 10×) and TIF2 (1× and 6×) repeats, we performed imaging experiments. Surprisingly, both ERβ- and GR2-LBDs, when fused to YFP and the VPR activation domain, formed nuclear puncta regardless of the number of P3 or TIF2 repeats (Supplementary Fig. 11). Notably, in the case of the 6×TIF2 construct, we observed colocalization with nuclear LBD:YFP:VPR condensates, whereas 1×TIF2 did not show such colocalization. This is consistent with multivalency-driven LLPS in the cytoplasm (Supplementary Figures 3 and 6) and demonstrates the differences in condensate formation in the cytosol versus the nucleus. We speculate that the nuclear puncta may be supported by the contribution of VPR, consistent with previous reports showing that dCas9-VPR independently forms nuclear condensates, possibly through interactions with nucleic acids or nuclear proteins46, or from potential recruitment of the LBD fusion proteins into endogenous nuclear condensates. Supporting this, transfection of cells with LBD-VPR fusion constructs alone was sufficient to induce nuclear puncta (Supplementary Fig. 12). The observation of nuclear puncta independent of P3 or TIF2 repeat number complicates the direct correlation between condensate formation and transcriptional activity. Multivalency mediated through P3 scaffolds and TIF2 repeats is important for achieving high transcriptional activation, as evidenced by the cytoplasmic LBD-LLPS data. Next, we evaluated the dynamic regulation of LLPS-based transcriptional activators upon the addition of antagonist ligands. The simultaneous presence of cortisol and the GR2 antagonist mifepristone, or estradiol and the ERβ antagonist 4-hydroxytamoxifen (4-OHT), effectively inhibited transcriptional activation (Fig. 5e, f), demonstrating that these systems can be dynamically regulated. Finally, we demonstrated that the designed multivalent transcriptional activators could be leveraged for custom transcriptional induction of an endogenous ASCL1 gene (Fig. 5g). These results highlight the potential of multivalent LLPS-based synthetic transcriptional activators to achieve robust gene activation, even on a single DNA-binding site.
Discussion
Designed sensor and actuator tools to manipulate mammalian cellular processes are desired for potential applications in synthetic biology-based control systems, with distinct advantages of human derived protein-based modules in terms of biocompatibility and functional integration with endogenous signaling. Here we leveraged the natural interaction between ligand-binding domains (LBDs) of nuclear receptors and their coactivators17 to create ligand-responsive molecular switches and leverages the formation of LLPS to potently amplify the response and decrease the concentration range of chemical regulators. As a proof-of concept this strategy was demonstrated using the LBDs of TRβ, VDR, RARγ, GR2, and ERβ, which interact with peptides derived from the TIF2 coactivator and respond to physiologically important ligands—triiodothyronine, vitamin D, retinoic acid, cortisol, and estrogen, respectively (Figs. 1 and 2). While our strategy focused on these receptors, the concept could be extendable to other NRs that bind well-characterized ligands such as vitamins, fatty acids, and additional hormones18. However, we recognize that many human NRs, selective ligands are not known or they exhibit complex endogenous regulation, restricting the generalizability of the platform. Unlike many CID platforms that rely on fully orthogonal protein components (e.g., plant, virus, or microbial proteins), this system is based on native human receptors and ligands9,11,12,47,48,49,50. On the other hand, CID systems based on human protein domains and responsive to FDA-approved drugs, such as thalidomide analogs51, Bcl-xL/Bcl-2 inhibitors (e.g., venetoclax)52,53, or the INSPIRE platform45, inevitably target not only engineered CID constructs but also their endogenous counterparts, which may compromise orthogonality and specificity. The advantage of LBD-CIDs is their potential integration with endogenous signaling pathways, without immune rejection of engineered cells, which could be advantageous in physiological or therapeutic contexts. At the same time, this integration raises the risk of off-target effects, such as activation of endogenous genes or crosstalk with native NR functions. Therefore, future applications should carefully assess such context-dependent effects. To mitigate such issues, NRs engineered for reduced sensitivity to endogenous ligands (e.g., mutant ER variants insensitive to endogenous hormones) could be used, with mutants that respond to small-molecule agonists or antagonists that do not trigger natural receptors54. Moreover, the constitutive or engineered interactions between LBDs and coactivator or corepressor peptides can be exploited to expand the control logic of these systems. For example, corepressor motifs derived from N-CoR or SMRT, which preferentially bind to LBDs in the absence of ligand, could be harnessed to create ligand-gated “off-switches” that dissociate upon ligand addition55,56.
Functional outputs can integrate multiple cooperating or competing signals that regulate a shared or different outputs57,58,59. For example, a first signal could activate a desirable function while a competing ligand could suppress it. Switchable behavior was recently demonstrated by de novo designed multi-input molecular switch based on the receiver viral NS3a protease and designed reader proteins which recognize the receiver-ligand complex9. We demonstrated that LBD-CIDs can reversibly toggle between “on” and “off” state of cellular output dependent on the presence of agonist/antagonist ligands, which may represent an additional safety latch in applications (Fig. 2). This switchable LBD-CIDs was applied to control gene expression using CRISPR/dCas9, which can be used to design therapeutic circuits and modulate the expression of in principle any gene of interest.
To enhance the transcriptional output, we concatenated multiple TIF2 peptides, which potently increased transcriptional activation upon ligand stimulation (Fig. 3). While this strategy robustly enhanced transcription in reporters with multiple binding sites, activation of a single binding site gene was achieved by providing coiled-coil dimer-mediated multimerization. Transcription factors and their coactivators, often regulate gene expression through the formation of transcriptional condensates via multivalent interactions60,61. Liquid-liquid phase separation (LLPS) at super-enhancers has been shown to concentrate activation domains to drive the transcription of highly active genes40. Additionally, fusing intrinsically disordered domains of condensate forming protein (e.g. FUS, DDX4 etc.). to synthetic transcription factors has been demonstrated to induce phase separation and enhance transcriptional activation62,63. Recently, CID tools have also been used to trigger droplet assembly in living cells by fusing a chemically responsive domain to intrinsically disordered domains of condensate forming proteins or multimerization domains64,65,66,67,68. However, many of the previously described systems used to trigger phase separation suffer from irreversibility as most of them used rapamycin-inducible CIDs or compounds to dissolve the condensates. We have previously designed artificial condensates composed of weak coiled-coil dimer-forming peptides54. Inspired by this, we engineered multivalency by introducing strong coiled-coil dimer-forming peptides to promote LLPS44,69. This combination successfully enhanced the formation of small-molecule-induced liquid condensates in the cytoplasm of mammalian cells (Fig. 4). These condensates could be dissolved upon the antagonist addition and reformed upon agonist stimulation. Furthermore, ligand-induced phase separation of inducible transcription factors robustly amplified the transcriptional activation (Fig. 5). The results demonstrate that potent amplification of transcription could be generated either by the multimerization of DNA binding sites or CC-mediated multimerization that provided sufficient valency for the formation and or recruitment of liquid condensates. However, a direct link between multivalency and nuclear condensates is difficult to establish, as LBD-VPR constructs form distinct nuclear condensates even in the absence of multivalent scaffolds. Recently Zhen et al. introduced testosterone-responsive synthetic condensates based on cytoplasmic androgen receptor (AR) constructs, which required deletion of the nuclear localization signal and DNA-binding domain hinge region to uncouple AR’s natural transcriptional activity70. In contrast, our LBD-CID system offers a simpler and more generalizable approach, without the need for extensive domain engineering or prior knowledge of LLPS-driving elements within the receptor. This approach complements and expands the growing synthetic condensate toolkit by enabling transcriptionally active, hormone-responsive condensates.
In summary, LBD-TIF2-based switches can be used for transducing sensed endogenous signals into functional cellular responses. These systems hold significant potential for advancing synthetic biology, particularly in unraveling the design and mechanism of LLPS and for therapeutic interventions.
Methods
Plasmid construction
All plasmids were constructed using the Gibson assembly method71 and are listed in Supplementary Table 1. Amino acid sequences of protein-coding genes and individual protein domains are listed in Supplementary Table 2. Nucleotide sequences of gRNAs and promoter sequences with DNA target sites are listed in Supplementary Table 3.
Mammalian cell culture and transfection
Human embryonic kidney (HEK) 293 T cells (ATCC, HEK293T, Catalog No. CRL-3216) were cultured in DMEM medium (Thermo Fisher Scientific) supplemented with 10% v/v FBS (Thermo Fisher Scientific) in a humidified incubator at 37 °C with 5% CO2. For luciferase experiments, 2-2.5 × 104 cells per well were seeded in CoStar White 96-well plates (Corning). For confocal microscopy experiments, 5 × 104 HEK293T cells per well were seeded in 8-well tissue-culture chambers (m-Slide 8 well, Ibidi). At 30–90% confluence, HEK293T cells were transfected with a mixture of DNA and PEI (6 µl/500 ng DNA, stock concentration 0.324 mg/ml, pH 7.5). The amounts of transfected plasmids are indicated in the figure captions. HEK293T cells were stimulated one day after transfection by replacing the media with fresh media containing the appropriate concentration of the ligand by 1000x dilution of the stock concentration (final DMSO concentration 0.1%).
1,6-Hexanediol treatment
For 1,6-hexanediol (1,6-HD) treatment, HEK293T cells were seeded in 8-well tissue culture chambers and transfected and/or stimulated with the appropriate plasmids and ligands. A 1,6-HD stock solution prepared in PBS was diluted in culture medium to a final concentration of 10 %. Cells were incubated with this working solution for 1 min, after which they were immediately imaged using confocal microscopy.
Luciferase assays
All luminescence measurements were made using a Centro LB 963 microplate reader (Berthold Technologies) with LightCompass® software. For endpoint luminescence measurements, the HEK293T cells were harvested at the indicated timepoints after stimulation and lysed in 25 ul of 1 × Passive Lysis buffer (Promega). Firefly luciferase and Renilla luciferase activity were measured in cell-lysates using the dual luciferase assay (Promega). Relative luciferase units (RLU) were calculated by normalizing each sample’s firefly luciferase activity to the constitutive Renilla luciferase activity determined in the same sample. For in situ kinetics, the cell culture medium was removed 48 h after transfection and replaced with 100 uL of assay medium (DMEM supplemented with 10% FBS, 2 mM ATP, 0.54 mM D-luciferin; with or without ligand). Continuous measurements of firefly luciferase activity were obtained every 15 s. The firefly luciferase units (nFluc) were calculated by normalizing the luciferase activity of each time-point to the value/signal of non-stimulated conditions.
Confocal microscopy
A Leica TCS SP5 inverted laser-scanning microscope on a Leica DMI 6000 CS module, using x63 oil-immersion objective and equipped with numerical aperture 0.4 (Leica Microsystems) was used for confocal microscopy. One day after transfection, HEK293T cells were stimulated with the indicated ligands and imaged 24 h after stimulation with the indicated ligands. A 514-nm laser line of a 100-mW argon laser was used for YFP excitation, and the emitted light was detected between 520 and 560 nm with a PMT (photomultiplier tube). A 50-mW 405-nm diode laser was used for TagBFP excitation, and the emitted light was detected between 420 nm and 460 nm with PMT. A 10-mW 633-nm HeNe laser was used for miRFP670nano3 excitation, and the emitted light was detected between 650 nm and 690 nm. Leica LAS AF Lite software version 1.0 was used for image acquisition.
Fluorescence recovery after photobleaching (FRAP)
FRAP experiments were performed on transfected HEK293T cells using a Leica TCS SP5 confocal microscope equipped with a 63× oil-immersion objective. A region of interest (ROI) within the condensate was selectively bleached using a 488-nm laser line at 100% power. Fluorescence intensity was recorded over time, capturing four pre-bleaching frames followed by post-bleaching frames at a rate of one frame every 2 s. To account for photobleaching effects, fluorescence intensity was also measured in an adjacent, unbleached droplet of similar size. Raw data were background-subtracted and normalized to the maximum pre-bleach intensity.
RNA extraction, reverse transcription, and quantitative PCR
HEK293T cells were harvested at the indicated timepoints after transfection and/or stimulation with 250 µl trypsin, centrifuged at 1000 x g for 5 min, resuspended in 200 µl PBS; thereafter, RNA was extracted using the High-pure RNA isolation kit (Roche). Reverse transcription was performed with the High-capacity cDNA reverse transcription kit (Applied Biosystems) with a mixture of random oligonucleotides. Quantitative PCR was performed with the LightCycler 480 SYBR Green I master mix (Roche) on the LightCycler 480 microplate reader. Oligonucleotides used for quantitative PCR: GAPDHf1: GGAGCGAGATCCCTCCAAAAT, GAPDHr1: GGCTGTTGTCATACTTCTCATGG, hASCL1f1: CGCGGCCAACAAGAAGATG and hASCL1r1: CGACGAGTAGGATGAGACCG. Results are shown as fold change compared to non-stimulated mock cells transfected with an empty vector plasmid (pcDNA3.1) after normalization to GAPDH expression using the ΔΔCt method72.
Statistics and reproducibility
Biological replicates represent parallel measurements of firefly luciferase reporter activity or in the HEK239T of distinct wells cultured in a multi-well plates under the same conditions and transfected with the same mixture of plasmids. At selected time points after cell transfection and/or stimulation, each well (biological replicate) was measured individually. Technical replicates (i.e. repeated measurements of the same sample) were performed only in the case of qPCR, where each biological replicate was measured in duplicate for each PCR reaction. Independent experiments indicate the repetition of the whole experiment. The dose reponse curves were fit to four parameter logistic (4-PL) curve using GraphPad Prism 8 (log[agonist] vs. response – variable slope; Equation: Y = Bottom + (Top-Bottom)/(1 + 10^((LogEC50-X)*HillSlope)). Microscopic images are representative of two independent experiments and at least four separate observations within the same experiment. All statistical analysis was calculated with GraphPad Prism 8. Exact P-values of Welch T-test and the number of replicates (n) are provided in figure captions. All figure schematics were created using the free and open-source software Inkscape (ver. 1.2.1; www.inkscape.org).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the article or Supplementary Information. Source data are provided with this paper.
References
Bashor, C. J., Hilton, I. B., Bandukwala, H., Smith, D. M. & Veiseh, O. Engineering the next generation of cell-based therapeutics. Nat. Rev. Drug Discov. 21, 655–675 (2022).
Brayshaw, L. L. et al. The role of small molecules in cell and gene therapy. RSC Med. Chem. 12, 330 (2021).
Fegan, A., White, B., Carlson, J. C. T. & Wagner, C. R. Chemically controlled protein assembly: Techniques and applications. Chem. Rev. 110, 3315–3336 (2010).
Stanton, B. Z., Chory, E. J. & Crabtree, G. R. Chemically induced proximity in biology and medicine. Science 359. https://doi.org/10.1126/science.aao5902 (2018).
Derose, R., Miyamoto, T. & Inoue, T. Manipulating signaling at will: Chemically-inducible dimerization (CID) techniques resolve problems in cell biology. Pflugers Archiv Eur. J. Physiol. 465, 409–417 (2013).
Rihtar, E. et al. Chemically inducible split protein regulators for mammalian cells. Nat. Chem. Biol. 1–8 https://doi.org/10.1038/s41589-022-01136-x (2022).
Kang, S. et al. COMBINES-CID: An Efficient Method for de Novo Engineering of Highly Specific Chemically Induced Protein Dimerization Systems. J. Am. Chem. Soc. 141, 10948–10952 (2019).
Glasgow, A. A. et al. Computational design of a modular protein sense-response system. Science 366, 1024–1028 (2019).
Foight, G. W. et al. Multi-input chemical control of protein dimerization for programming graded cellular responses. Nat. Biotechnol. 37, 1209–1216 (2019).
Martinko, A. J. et al. Switchable assembly and function of antibody complexes in vivo using a small molecule. Proc. Natl. Acad. Sci. Usa. 119, e2117402119 (2022).
Miyamoto, T. et al. Rapid and orthogonal logic gating with a gibberellin-induced dimerization system. Nat. Chem. Biol. 8, 465–470 (2012).
Liang, F.-S., Ho, W. Q. & Crabtree, G. R. Engineering the ABA Plant Stress Pathway for Regulation of Induced Proximity. Sci. Signal. 4, rs2 (2011).
Spencer, D., Wandless, T., Schreiber, S. & Crabtree, G. Controlling signal transduction with synthetic ligands. Science 262, 1019–1024 (1993).
Putyrski, M. & Schultz, C. Protein translocation as a tool: The current rapamycin story. FEBS Lett. 586, 2097–2105 (2012).
Scholtes, C. & Giguère, V. Transcriptional control of energy metabolism by nuclear receptors. Nat. Rev. Mol. Cell Biol. 23, 750–770 (2022).
Evans, R. M. & Mangelsdorf, D. J. Nuclear Receptors, RXR, and the Big Bang. Cell 157, 255–266 (2014).
Sladek, F. M. What are nuclear receptor ligands?. Mol. Cell. Endocrinol. 334, 3–13 (2011).
Rastinejad, F., Ollendorff, V. & Polikarpov, I. Nuclear receptor full-length architectures: confronting myth and illusion with high resolution. Trends Biochem. Sci. 40, 16–24 (2015).
Weikum, E. R., Knuesel, M. T., Ortlund, E. A. & Yamamoto, K. R. Glucocorticoid receptor control of transcription: precision and plasticity via allostery. Nat. Rev. Mol. Cell Biol. 18, 159–174 (2017).
Yu, X. et al. Structural Insights of Transcriptionally Active, Full-Length Androgen Receptor Coactivator Complexes. Mol. Cell 79, 812–823.e4 (2020).
Fondell, J. D., Ge, H. & Roeder, R. G. Ligand induction of a transcriptionally active thyroid hormone receptor coactivator complex. Proc. Natl. Acad. Sci. USA. 93, 8329–8333 (1996).
Lin, B. C., Hong, S. H., Krig, S., Yoh, S. M. & Privalsky, M. L. A conformational switch in nuclear hormone receptors is involved in coupling hormone binding to corepressor release. Mol. Cell. Biol. 17, 6131–6138 (1997).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Ko, M. S., Takahashi, N., Sugiyama, N. & Takano, T. An auto-inducible vector conferring high glucocorticoid inducibility upon stable transformant cells. Gene 84, 383–389 (1989).
Friedman, H. M. et al. Use of a glucocorticoid-inducible promoter for expression of herpes simplex virus type 1 glycoprotein gC1, a cytotoxic protein in mammalian cells. Mol. Cell. Biol. 9, 2303–2314 (1989).
Gallinari, P. et al. A Functionally Orthogonal Estrogen Receptor-Based Transcription Switch Specifically Induced by a Nonsteroid Synthetic Ligand. Chem. Biol. 12, 883–893 (2005).
Rössger, K., Charpin-El-Hamri, G. & Fussenegger, M. A closed-loop synthetic gene circuit for the treatment of diet-induced obesity in mice. Nat. Commun. 4, 2825 (2013).
Braselmann, S., Graninger, P. & Busslinger, M. A selective transcriptional induction system for mammalian cells based on Gal4-estrogen receptor fusion proteins. Proc. Natl. Acad. Sci. 90, 1657–1661 (1993).
Mata de Urquiza, A., Solomin, L. & Perlmann, T. Feedback-inducible nuclear-receptor-driven reporter gene expression in transgenic mice. Proc. Natl. Acad. Sci. 96, 13270–13275 (1999).
Dirnberger, D., Unsin, G., Schlenker, S. & Reichel, C. A Small-Molecule–Protein Interaction System with Split-Ubiquitin as Sensor. ChemBioChem 7, 936–942 (2006).
Choi, G. et al. Novel Estrogen Receptor Dimerization BRET-Based Biosensors for Screening Estrogenic Endocrine-Disrupting Chemicals. Biomater. Res. 28, 0010 (2024).
Bledsoe, R. K. et al. Crystal Structure of the Glucocorticoid Receptor Ligand Binding Domain Reveals a Novel Mode of Receptor Dimerization and Coactivator Recognition. Cell 110, 93–105 (2002).
Idevall-Hagren, O., Dickson, E. J., Hille, B., Toomre, D. K. & De Camilli, P. Optogenetic control of phosphoinositide metabolism. Proc. Natl. Acad. Sci. 109, E2316–E2323 (2012).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049 (2016).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A Protein-Tagging System for Signal Amplification in Gene Expression and Fluorescence Imaging. Cell 159, 635–646 (2014).
Scarsi, A., Pedone, D. & Paolo Pompa, P. A dual-color plasmonic immunosensor for salivary cortisol measurement. Nanoscale Adv. 5, 329–336 (2023).
Watson, C. S., Jeng, Y.-J. & Kochukov, M. Y. Nongenomic actions of estradiol compared with estrone and estriol in pituitary tumor cell signaling and proliferation. FASEB J. 22, 3328–3336 (2008).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Song, D. et al. ERα and ERβ Homodimers in the Same Cellular Context Regulate Distinct Transcriptomes and Functions. Front. Endocrinol. 13, 930227 (2022).
Presman, D. M. et al. DNA binding triggers tetramerization of the glucocorticoid receptor in live cells. Proc. Natl. Acad. Sci. 113, 8236–8241 (2016).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, eaar2555 (2018).
Lebar, T., Lainšček, D., Merljak, E., Aupič, J. & Jerala, R. A tunable orthogonal coiled-coil interaction toolbox for engineering mammalian cells. Nat. Chem. Biol. 16, 513–519 (2020).
Rihtar, E. et al. Chemically inducible split protein regulators for mammalian cells. Nat. Chem. Biol. 19, 64–71 (2023).
Fu, Y. et al. Dynamic properties of transcriptional condensates modulate CRISPRa-mediated gene activation. Nat. Commun. 16, 1640 (2025).
Ziegler, M. J. et al. Mandipropamid as a chemical inducer of proximity for in vivo applications. Nat. Chem. Biol. 18, 64–69 (2022).
Wang, T. et al. Repurposing salicylic acid as a versatile inducer of proximity. Nat. Chem. Biol. 1–13 https://doi.org/10.1038/s41589-025-01918-z (2025).
Chin, S. E. et al. A simeprevir-inducible molecular switch for the control of cell and gene therapies. Nat. Commun. 14, 7753 (2023).
Beltrán, J. et al. Rapid biosensor development using plant hormone receptors as reprogrammable scaffolds. Nat. Biotechnol. 40, 1855–1861 (2022).
Jan, M. et al. Reversible ON- and OFF-switch chimeric antigen receptors controlled by lenalidomide. Sci. Transl. Med. 13, eabb6295 (2021).
Shui, S. et al. A rational blueprint for the design of chemically-controlled protein switches. Nat. Commun. 12, 5754 (2021).
Giordano Attianese, G. M. P. et al. Dual ON/OFF-switch chimeric antigen receptor controlled by two clinically approved drugs. Proc. Natl. Acad. Sci. USA 121, e2405085121.
Feil, R., Wagner, J., Metzger, D. & Chambon, P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem. Biophys. Res. Commun. 237, 752–757 (1997).
Lee, J. W., Cheong, J., Lee, Y. C., Na, S.-Y. & Lee, S.-K. Dissecting the molecular mechanism of nuclear receptor action: transcription coactivators and corepressors. Exp. Mol. Med. 32, 53–60 (2000).
Watson, P. J., Fairall, L. & Schwabe, J. W. R. Nuclear hormone receptor co-repressors: Structure and function. Mol. Cell. Endocrinol. 348–135, 440–449 (2012).
Bertschi, A., Wang, P., Galvan, S., Teixeira, A. P. & Fussenegger, M. Combinatorial protein dimerization enables precise multi-input synthetic computations. Nat. Chem. Biol. 19, 767–777 (2023).
Gao, Y., Wang, L. & Wang, B. Customizing cellular signal processing by synthetic multi-level regulatory circuits. Nat. Commun. 14, 8415 (2023).
Daniel, R., Rubens, J. R., Sarpeshkar, R. & Lu, T. K. Synthetic analog computation in living cells. Nature 497, 619–623 (2013).
Wei, M.-T. et al. Nucleated transcriptional condensates amplify gene expression. Nat. Cell Biol. 22, 1187–1196 (2020).
Chong, S. et al. Tuning levels of low-complexity domain interactions to modulate endogenous oncogenic transcription. Mol. Cell 82, 2084–2097.e5 (2022).
Chen, R. et al. Specific multivalent molecules boost CRISPR-mediated transcriptional activation. Nat. Commun. 15, 7222 (2024).
Schneider, N. et al. Liquid-liquid phase separation of light-inducible transcription factors increases transcription activation in mammalian cells and mice. Sci. Adv. 7, eabd3568 (2021).
Yoshikawa, M., Yoshii, T., Ikuta, M. & Tsukiji, S. Synthetic Protein Condensates That Inducibly Recruit and Release Protein Activity in Living Cells. J. Am. Chem. Soc. 143, 6434–6446 (2021).
Garabedian, M. V. et al. Designer membraneless organelles sequester native factors for control of cell behavior. Nat. Chem. Biol. 17, 998–1007 (2021).
Wu, J. et al. Modulating gene regulation function by chemically controlled transcription factor clustering. Nat. Commun. 13, 2663 (2022).
Qian, Z.-G., Huang, S.-C. & Xia, X.-X. Synthetic protein condensates for cellular and metabolic engineering. Nat. Chem. Biol. 18, 1330–1340 (2022).
Wang, Y. et al. Programmable solid-state condensates for spatiotemporal control of mammalian gene expression. Nat. Chem. Biol. 1–10 https://doi.org/10.1038/s41589-025-01860-0 (2025).
Ramšak, M. et al. Programmable de novo designed coiled coil-mediated phase separation in mammalian cells. Nat. Commun. 14, 7973 (2023).
Zhen, N. et al. Engineering bi-directional chemically-modulated synthetic condensates for cellular control. Nat. Commun. 16, 6587 (2025).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, (2009).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
This research was supported by grants from the Slovenian Research Agency (J3-60061, P4-0176, J7-4640, J7-4493, R.J.) and project CTGCT, which received Teaming for Excellence funding under the European Union’s Horizon research and innovation program Grant agreement ID: 101059842 (R.J.).
Author information
Authors and Affiliations
Contributions
E.R. and R.J. conceptualized the research; E.R. designed the experiments. E.R., T.F., F.I., and E.K. performed the experiments on cell culture. E.R. analyzed the data and interpreted the results. E.R. drafted the initial manuscript. E.R., T.F., and R.J. critically reviewed, edited, and provided feedback on the manuscript. R.J. supervised the work and provided funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rihtar, E., Fink, T., Ivanovski, F. et al. Repurposing nuclear receptors for ligand-responsive liquid condensate formation and gene regulation. Nat Commun 17, 2218 (2026). https://doi.org/10.1038/s41467-026-69099-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69099-4