Abstract
Alcoholic liver disease (ALD) remains a significant clinical challenge with limited therapeutics. It is strongly associated with sarcopenia, which further worsens the prognosis in liver cirrhosis. Zinc-deficiency is a hallmark of ALD, impairing the liver’s antioxidant defenses and alcohol metabolism. Here we identify a chaperone role for MG53/TRIM72 in facilitating crosstalk between zinc and alcohol dehydrogenase (ADH) and aldehyde dehydrogenase (ALDH2), acting as a metabolic regulator of muscle-liver communication. In ALD, muscle-derived MG53 is transported to liver and mitigates liver damage. MG53 deficiency exacerbates hepatic zinc-deficiency and impairs ADH and ALDH2 activity, which are reversed by systemic administration of recombinant human MG53 (rhMG53). MG53’s B-box motif coordinates zinc interaction with ADH and ALDH2, enhancing their enzymatic activity to clear toxic alcohol byproducts. We developed a bio-derived nanoparticle for co-delivery of rhMG53 and zinc, offering a promising therapeutic approach for ALD. In vivo findings are applicable to the male sex only.
Similar content being viewed by others
Introduction
Alcoholic liver disease (ALD) is a progressive condition caused by chronic alcohol consumption, leading to liver pathologies such as steatosis, hepatitis, and cirrhosis1,2,3. A significant complication of ALD is sarcopenia, which is characterized by the loss of muscle mass and strength4. Sarcopenia greatly exacerbates hepatic encephalopathy and mortality in patients with liver cirrhosis5,6. However, the effect of sarcopenia on ALD has not been clarified. Myokines, signaling molecules produced by skeletal muscle, play a key role in regulating metabolic processes and tissue regeneration, supporting the health of the liver and other vital organs7,8. Understanding the role of myokines in tissue protection and regeneration is essential for developing effective treatments for ALD.
About 15 years ago, our group identified MG53, a muscle-derived zinc-binding protein, as a critical component of the cell membrane repair machinery9. Dubbed a “molecular bandage”, MG53 has been shown to protect a variety of vital organs, including muscle, heart, liver, lung, kidney, eye, and brain, under injurious conditions10,11,12,13,14. Notably, hepatocytes do not contain endogenous MG53 protein, relying instead on muscle-derived MG53 delivered through circulation to aid in liver protection under stress conditions13. While extensive studies have demonstrated MG53’s therapeutic potential as a tissue repair protein, its role in protecting against chronic liver injury and metabolic dysfunction associated with ALD has yet to be explored.
Zinc is a key element in various metabolic processes, including enzyme function, antioxidant defense, and cell membrane repair15,16,17,18,19,20. Chronic alcohol consumption is known to deplete hepatic zinc levels, leading to impaired liver and muscle health4,21. Zinc deficiency worsens liver damage in ALD by weakening the liver’s antioxidant defenses and reducing its ability to metabolize toxic alcohol byproducts, while also compromising the liver’s regenerative capacity17,22.
In this study, we identify a chaperone role for MG53 in facilitating the crosstalk between zinc and alcohol dehydrogenase (ADH) and aldehyde dehydrogenase 2 (ALDH2), acting as a metabolic regulator of muscle–liver communication. By modulating zinc metabolism in ALD, MG53 helps preserve ADH and ALDH2 activity, thereby enhancing the liver’s metabolic capacity under chronic-plus-binge ethanol exposure. To harness the synergistic effects of MG53 and zinc in combating ALD, we developed a bio-derived nanoparticle for the co-delivery of recombinant human MG53 (rhMG53) and zinc. We demonstrated its safety and efficacy in preserving liver metabolic function in rodent models, offering a promising therapeutic strategy for ALD treatment.
Results
ALD causes MG53 transfer from the circulation to the liver tissues
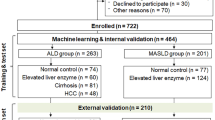
To investigate the role of sarcopenia in ALD, we first analyzed data from the National Health and Nutrition Examination Survey (NHANES). A total of 1858 participants with complete information were included, of whom 498 were classified as ALD (Fig. S1A, B). Both crude and adjusted analyses showed that sarcopenia was significantly associated with an approximately 2-fold higher risk of ALD (Fig. 1A). Sarcopenia was defined using appendicular skeletal muscle mass normalized to body mass index (ASM/BMI) as an index23. Restricted cubic spline analysis revealed a nonlinear relationship between ASM/BMI and ALD probability (p < 0.001), with a higher risk in male participants (Fig. 1B).
A Association of sarcopenia and ALD risk based on the National Health and Nutrition Examination Survey (NHANES) dataset. Odds ratios in Model 1 were crude, and in Model 2 were adjusted by age, sex, race, marital status, education, smoking, and alcohol drinking. B Association of skeletal muscle mass/body mass index (ASMBMI) with ALD probability among female (red) and male (blue) participants. C Schematic representation of the workflow identifying candidate proteins involved in skeletal muscle–liver crosstalk. D Survival analysis of alcohol abuse patients with liver cancer, stratified by high and low MG53 expression levels (79 patients in the high-expression group, 36 in the low-expression group). E H/E staining and MG53 immunohistochemistry staining in liver tissues from ALD patients. Para-carcinoma liver tissues from patients without an alcohol history served as normal controls (Scale bars, 200 μm. n = 6 per group). F Immunofluorescence staining of MG53 in liver tissues from mice after pair-fed or EtOH-fed for 4 weeks (Scale bars, 100 μm. n = 10 per group). Cytokeratin 18 (CK18), highly concentrated in hepatocytes, serves as a marker to identify hepatocytes in liver tissue. G Serum and hepatic MG53 expressions in ALD patients (left) and ALD mice (right) determined by immunoblotting. ALD patients were compared with healthy donors and normal controls (para-carcinoma liver tissues from patients without an alcohol history), showing on the left (n = 6 per group). Mice after an EtOH-fed diet for 4 weeks were compared with a pair-fed diet, showing on the right (n = 10 per group). Ponceau S or GAPDH was used as a loading control for serum samples or liver tissue. H Correlation analysis between serum MG53 and AST in ALD patients (left, n = 10) and ALD mice (right, n = 15). Data were presented as odds ratios with 95% confidence intervals in (A) and as mean ± SEM in (B). Logistic regression was performed for (A) restricted cubic splines with a likelihood ratio test were performed for (B) and a simple linear regression model was performed for (H). Source data are provided as a Source Data file.
Further to examine the crosstalk between skeletal muscle and liver in ALD, we performed a combined analysis using data from the Human Liver Proteome Database24, The Human Protein Atlas25,26, and liver proteomics from ALD patients. Proteins typically present in normal liver tissue were excluded, leaving a list of 455 exogenous proteins in ALD liver, suggesting these were secreted from other organs and taken up by the liver. Among these, two potential secreted myokines - MG53/TRIM72 and Ankyrin 1 (ANK1) were identified (Fig. 1C). A literature search revealed no published studies linking ANK1 to liver function or the pathogenesis of ALD. Consistently, a 5-year clinical follow-up data (Fig. S1C) showed no significant impact of ANK1 expression on patient survival. In contrast, higher expressions of MG53 were positively correlated with increased survival in patients with alcohol abuse and liver cancer, as demonstrated using the Kaplan–Meier Plotter database (Fig. 1D)27. These findings highlight that MG53 emerged as a primary candidate myokine mediating skeletal muscle–liver communication in ALD.
MG53, a skeletal muscle-derived protein involved in membrane repair, has been shown to protect against acute liver injury and suppress hepatocellular carcinoma13,18,28,29. Since our discovery of MG53 in 20099, extensive studies from our group and others have established that MG53 is predominantly expressed in skeletal and cardiac muscle (see Fig. S1D, taken from the FANTOM5 database). Consistent with previous reports13,29, MG53 is absent in normal liver tissue; however, we detected MG53 in the liver of ALD patients with chronic alcohol abuse and in mice fed with ethanol (EtOH, 2.8% v/v) for 4 weeks (Fig. 1E, F and Fig. S1E, F). The ALD mouse model was successfully established, as confirmed by hematoxylin-eosin (H&E) staining showing hepatic lesions and increased ALT, AST, triglyceride, and cholesterol levels (Fig. S2A–D). This suggests a translocation of MG53 from circulation to the liver, as its concentration decreased in the blood but increased in the liver of both ALD patients and mice (Fig. 1G). Moreover, we found an inverse correlation between serum MG53 levels and liver function markers (serum ALT, AST, and LDH levels) (Fig. 1H and Fig. S2E). Using AAV-mediated muscle-specific expression of MG53-EGFP (AAV9-MHCK7-MG53-EGFP) in mice30,31, we tracked the dynamic transfer of MG53-EGFP from skeletal muscle to other tissues during chronic ethanol exposure. After 4 weeks of ethanol feeding, immunofluorescence revealed MG53-EGFP translocation from skeletal muscle into injured hepatocytes (Fig. S2F). Together, these findings suggest that MG53 acts as a key mediator of skeletal muscle–liver communication, influencing hepatic function and contributing to ALD progression.
MG53 deficiency exacerbates ALD severity, and exogenous rhMG53 ameliorates ALD in mice
To investigate MG53’s role in ALD, MG53 knockout (MG53−/−) and littermate wild-type (WT) mice were fed with ethanol (EtOH) for 4–8 weeks (Figs. S3 and S4). Under physiological conditions, no difference was observed between MG53−/− and WT mice in terms of liver weight/body weight ratio and liver function (Fig. S3A–C), along with increased liver MG53 in ethanol-stimulated WT mice but no significant difference in ethanol intake between the WT and MG53−/− mice (Fig. S3D-E). MG53 deficiency significantly reduced the survival rates over the 8-week observation period (Fig. 2A). Due to the high mortality at 8 weeks, liver function was assessed at the 4-week time-point (Fig. S4A).
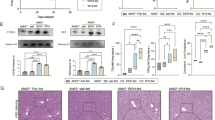
A Effect of MG53 deficiency on the survival rate in mice subjected to an 8-week pair-fed or EtOH-fed diet, with replenishment of rhMG53 via micro-osmotic pump (1 mg/kg with 1 μg/h) (n = 20 per group). B Liver-body weight ratio in mice after a 4-week pair-fed or EtOH-fed diet. C Impact of MG53 deficiency on liver function, assessed by serum AST levels in mice after pair-fed or EtOH-fed diets for 4 weeks. rhMG53 replenishment reduces AST levels in MG53−/− mice subjected to EtOH treatment. D Liver structural changes due to MG53 deficiency with or without rhMG53 replenishment, as shown in (C) evaluated by H/E staining (Scale bars, 100 μm) in mice after pair-fed or EtOH-fed diets for 4 weeks. E Liver steatosis in the different groups of mice, as shown in (C) determined by Oil Red O staining (Scale bars, 50 μm) after pair-fed or EtOH-fed diets for 4 weeks. F Hepatic triglyceride level in mice subjected to a 4-week EtOH-fed diet treatment was more elevated in MG53−/− mice compared with WT mice, and was reduced with rhMG53 treatment. G Reactive oxygen species (ROS) levels in the different mice, detected by DHE staining (Scale bars, 50 μm). H Different liver ultrastructure, determined by TEM (300× and 1000×) in mice after pair-fed or EtOH-fed diets for 4 weeks. LD lipid droplet, M mitochondria, N nucleus. I Inflammatory response in mice, indicated by serum TNF-α levels after pair-fed or EtOH-fed diets for 4 weeks. J Liver fibrosis in mice, assessed by Sirius red staining (Scale bars, 50 μm). K Collagen I expression in mice liver. From 2B to 2 K, n = 6 for MG53−/− mice with EtOH fed, n = 8 for rhMG53 treatment group, n = 8 for other groups. Data were presented as mean ± SD. Survival curve with Log-rank (Mantel–Cox) test was performed for (A); two-way ANOVA with Sidak’s test was performed for (B, C, F, I and K). Source data are provided as a Source Data file.
In EtOH-fed mice, MG53 deficiency increased liver weight/body weight ratio (Fig. 2B) and muscle loss (Fig. S4B), along with worsened liver function, as indicated by elevated serum ALT, AST, and LDH levels (Fig. 2C and Fig. S4C). H&E staining revealed that EtOH-fed MG53−/− mice displayed severe hepatic lesions, with higher pathological scores and pronounced hepatocyte ballooning, compared with WT mice (Fig. 2D and Fig. S4D). These MG53−/− mice also showed aggravated hepatic steatosis, as evidenced by Oil Red O staining (Fig. 2E and Fig. S4E) and increased liver triglyceride and cholesterol levels (Fig. 2F and Fig. S4F).
Additionally, MG53 deficiency led to heightened oxidative stress, mitochondrial injury, and inflammation injury, as measured by DHE fluorescence (Fig. 2G and Fig. S4G), MPO activities (Fig. S4H), and transmission electron microscopy (TEM) (Fig. 2H and Fig. S4I), alongside elevated levels of inflammation markers with TNF-α, IL-6, and IL-1β levels (Fig. 2I and Fig. S4J). The EtOH diet also significantly exacerbated fibrosis in MG53−/− mice at 4-weeks post EtOH treatment, as shown by Sirius red staining (Fig. 2J and Fig. S4K) and increased collagen I and IV expression (Fig. 2K and Fig. S4L).
To further explore MG53’s protective role in ALD, a rescue study was conducted by administering recombinant human MG53 protein (rhMG53, 1 mg/kg with 1 µg/h) via micro-osmotic pump to MG53−/− mice (Fig. S4A). Replenishment of rhMG53 improved survival rate (Fig. 2A), reduced the skeletal muscle loss (Fig. S4B), mitigated hepatic injury (Fig. 2B, C), steatosis, inflammation, and fibrosis (Fig. 2D–K and Fig. S4), supporting the important function of MG53 in protection against ALD in mice.
ADH and ALDH2 are MG53-interacting proteins that control alcohol metabolism in ALD
We performed mass spectrometry to identify potential MG53-binding proteins in the liver of ALD mice after immunoprecipitation with MG53 antibody (Fig. 3A). A total of 417 potential MG53-binding proteins were detected, and after merging with 9749 ALD-related genes, 205 candidate targets were selected for further analysis.
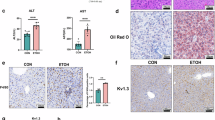
A Screening process of MG53-binding key targets. Venn diagram showing 205 overlapping proteins between MG53-potential proteins (417; identified via mass spectrometry using MG53 antibody in the liver tissue from EtOH-fed mice) and ALD-related genes (9747, retrieved from the GeneCards and CTD databases). B Gene Ontology (GO) enrichment analysis of 205 overlapping proteins with the top 10 terms in each enrichment. C Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of 205 overlapping proteins. The top 10 representative pathways, including “alcoholic liver diseases”, are presented with gene counts. D Schematic representation of the ALD pathway in the KEGG pathway map with MG53-binding proteins (red) (https://www.kegg.jp/entry/map04936). E Co-immunoprecipitation of ADH/ALDH2 and MG53 in liver tissues from pair-fed or EtOH-fed mice after 4 weeks. F Hepatic zinc concentration in mice following pair-fed or EtOH-fed diets with or without rhMG53 treatment. G, H Serum ethanol and aldehyde levels in MG53−/− mice after 4-week pair-fed or EtOH-fed diets with or without rhMG53 treatment. I, J ADH and ALDH2 enzyme activities in AML12 cells treated with EtOH (400 mM) and rhMG53 (10 μg/ml) for 24 h (n = 8 per group). K Immunofluorescence staining of hepatic ADH/ALDH2 and MG53 in pair-fed or EtOH-fed mice (Scale bars, 20 μm). Mice after 4-week exposure to pair-fed or EtOH-fed were administered with Cy5.5-labeled rhMG53 (1 mg/kg) 2 h prior to sacrifice (n = 6 per group). L Immunofluorescence staining of ADH/ALDH2 and MG53 in AML12 cells treated with EtOH (400 mM) and rhMG53 (10 μg/ml) for 24 h (Scale bars, 20 μm. n = 6 per group). From (F–H), n = 6 for MG53−/− mice with EtOH fed, n = 8 for rhMG53 treatment group, n = 8 for other groups. Data were presented as mean ± SD. A two-sided hypergeometric test was performed for (C), a two-way ANOVA with Sidak’s test was performed for (F–H), and a one-way ANOVA with Tukey’s test was performed for (I, J). Source data are provided as a Source Data file.
GO enrichment analysis of molecular functions, cell components, and biological processes revealed that these proteins are predominantly involved in alcohol metabolism, mitochondrial function, and membrane-related pathways (Fig. 3B). Of the top 10 KEGG pathways, “alcoholic liver disease” was highlighted as potentially playing a critical role in MG53’s protective effects against ALD (Fig. 3C).
In the “alcoholic liver disease” pathway, ADH and ALDH are two key enzymes responsible for alcohol metabolism (Fig. 3D). Our data demonstrate that ADH and ALDH2 are direct binding partners of MG53, a finding confirmed through co-immunoprecipitation (Fig. 3E and Fig. S5A, B) in liver and extra-liver, including gut (Fig. S5C, D). These enzymes, essential for antioxidant defense and liver protection under stress, are regulated by cellular zinc levels17,22. Extensive research shows that hepatic zinc deficiency, a hallmark of ALD, impairs the function of both enzymes32,33. Notably, the ability of ADH and ALDH2 to metabolize toxic alcohol byproducts is zinc-dependent17,22.
Spectroscopic analysis revealed a progressive decline in hepatic zinc levels associated with chronic EtOH exposure34,35. Interestingly, MG53−/− mice exhibited more severe zinc deficiency in the liver compared with WT mice after 4 weeks of EtOH exposure. However, this zinc deficiency was ameliorated in MG53−/− mice following rhMG53 administration (Fig. 3F).
Given the pivotal role of ADH and ALDH2 in alcohol metabolism, we measured serum ethanol and acetaldehyde levels. In MG53−/− mice, ethanol and acetaldehyde concentrations were significantly higher compared to controls, but these levels were markedly reduced by rhMG53 replenishment (Fig. 3G, H). This suggests that MG53 exerts a protective effect against ALD by binding with ADH and ALDH, enhancing their activities and improving alcohol metabolism, thus reducing EtOH-induced liver injury. We also treated cultured AML12 liver cells with rhMG53 in the presence of EtOH for 24 h (Fig. S5E–G). As the concentration of EtOH escalates, there is a corresponding exacerbation of cellular lipid droplet accumulation and injury in AML12 cells, as indicated by the increased fluorescent intensity of Nile red (Fig. S5E) and the marked reduction in cell viability (Fig. S5F). EtOH treatment (400 mM) for 24 h also caused significant ROS elevation, evidenced by increased fluorescent intensity of DHE (Fig. S5H) and increased levels of LDH in the culture medium (Fig. S5I), indicating cellular damage. However, the addition of rhMG53 (10 µg/mL) significantly mitigated the harmful effects of EtOH on AML12 cells. This was reflected in the substantially improved cell viability (Fig. S5G), reduction of DHE fluorescence (Fig. S5H), and decreased LDH release (Fig. S5I). Further immunofluorescence analysis confirmed that rhMG53 treatment increases intracellular zinc levels and enhances its co-localization with ADH and ALDH2 in EtOH-stimulated AML12 cells (Fig. S5J). Indeed, in AML12 cells subjected to EtOH treatment, rhMG53 significantly increased the enzymatic activity of both ADH and ALDH2 (Fig. 3I, J).
We systemically administered Cy5.5-labeled rhMG53 to mice and observed markedly elevated levels of Cy5.5-rhMG53 in the livers of ALD mice (after 4-week exposure to EtOH), but not in control mice on a normal diet (Fig. 3K). Immunofluorescence staining revealed clear co-localization of rhMG53 with both ADH and ALDH2 (Fig. 3K) in livers from ALD mice. Furthermore, consistent with previous reports of MG53 localizing to injured cell membrane13,18, we observed that rhMG53 could enter the cytosol and co-localize with ADH and ALDH2 in AML12 cells treated with EtOH (Fig. 3L), which also co-precipitated in rhMG53-treated primary mouse hepatic parenchymal cells (Fig. S5K). These findings confirm the intracellular action of rhMG53 to enhance the enzymatic activities of ADH and ALDH2 under ALD conditions.
MG53 coordinates with zinc recruitment to enhance ADH/ALDH2 function in ALD
To investigate the mechanisms underlying MG53’s regulation of ADH/ALDH2 functions, we obtained the structure coordinates of MG53, ADH, and ALDH2 from the AlphaFold Protein Structure Database and performed molecular docking analysis to model their interactions. These studies revealed that three key residues in MG53-Glu87, Asp100, and Arg101 are critical for its interaction with ADH, while Leu103 is essential for MG53’s interaction with ALDH2 (Fig. 4A).
A Molecular docking analysis of MG53 binding to ADH and ALDH2. MG53 binds to ADH at residues 87, 100, and 101 (left), and binds ALDH2 at residue 103 (right). B Molecular dynamic simulation of MG53-ADH and MG53-ALDH2 complexes for 100 ns. Left: Gibbs free energy landscape of the MG53-ADH complex, with principal component 1 (PC1) ranging from 0.0 to 1.13, and PC2 between 2.97 and 3.23. Right: Gibbs free energy landscape of the MG53-ALDH2 complex, with PC1 values from 0.0 to 1.05, and PC2 values between 2.71 and 3.03. C Root Mean Square Deviation (RMSD) trends of MG53-ADH (blue) and MG53-ALDH2 (red) complexes, showing a rapid increase from 0 to 20 ns, followed by stabilization from 20 to 100 ns. D Binding affinity of rhMG53 to ADH (left) and ALDH2 (right) measured at various concentrations of MG53. ADH/ALDH2 were immobilized on streptavidin (SA) biosensors, and binding was detected by bio-layer interferometry. E Binding process of rhMG53 protein (10 μg/ml) and rhMG53-ADH-binding mutant (rhMG53-3A) to ADH, analyzed by bio-layer interferometry. ADH was immobilized on SA biosensors. rhMG53-3A protein represents MG53 mutations at ADH-binding residues 87, 100, and 101. (F) ADH enzyme activity in AML12 cells co-cultured with 400 mM EtOH and treated with 10 μg/m lrhMG53 or rhMG53-3A (n = 8 per group). G Binding process of rhMG53 and rhMG53-ALDH2-binding mutant (rhMG53-1A) to ALDH2, analyzed by bio-layer interferometry. ALDH2 was immobilized on SA biosensors. rhMG53-1A represents MG53 mutation at the ALDH2-binding residue 103. H ALDH2 enzyme activity in AML12 cells co-cultured with 400 mM EtOH and treated with 10 μg/ml rhMG53 or rhMG53-1A (n = 8 per group). ADH enzyme activities (I) ALDH2 enzyme activities (J) cell viability (K) and intracellular LDH release (L) in AML12 cells co-cultured with 400 mM EtOH and treated with 10 μg/ml rhMG53 or rhMG53-zinc-binding mutant (rhMG53-C29L/C105S), representing mutations in MG53-zinc-binding domains at residues 29 and 105 (n = 8 per group). Data were presented as mean ± SD. P-value was determined by one-way ANOVA with Tukey’s test. Source data are provided as a Source Data file.
To further assess the stability of the MG53/ADH or MG53/ALDH2 complexes, we conducted molecular dynamics simulations (Fig. 4B and Fig. S6A–D). The Gibbs free energy landscape demonstrated stable complex formation for both MG53/ADH and MG53/ALDH2 (Fig. 4B and Fig. S6E). During the 100 ns simulations, rapid binding of ADH and ALDH2 to MG53 occurred within the initial 20 ns, with complex stabilization observed throughout the remainder of the simulation (Fig. 4C). The specific interactions between rhMG53 and ADH, and rhMG53 and ALDH2, were confirmed by Bio-Layer Interferometry (BLI) assay (Fig. 4D). These data confirm the stable combination between rhMG53 and ADH/ALDH2.
Guided by the molecular docking studies (Fig. 4A), we performed site-directed mutagenesis to mutate the three critical residues in MG53 (Glu87, Asp100, and Arg101) to alanines, generating the mutant rhMG53-3A protein using our established E. coli fermentation protocol (Fig. S7A, B)10. BLI assays confirmed that the rhMG53-3A mutant lost its specific binding with ADH (Fig. 4E). While the wild-type rhMG53 protein enhanced the enzymatic activity of ADH in an in vitro biochemical assay, the rhMG53-3A mutant was ineffective in stimulating ADH function (Fig. 4F). Similarly, when Leu103 was mutated to alanine, the rhMG53-1A mutant lost its interaction with ALDH2 in the BLI assay (Fig. 4G), and was unable to enhance ALDH2 enzymatic activity in the in vitro conditions (Fig. 4H). These results indicate that the stimulating effects of rhMG53 on ADH and ALDH2 are dependent on its interaction with these enzymes.
Domain homology analysis identified two zinc-binding domains in MG53, located in the RING finger and B-box motifs (Fig. S7C)18,36,37. Mutations of Cys29 to Leucine (C29L) in the RING finger and Cys105 to Serine (C105S) in the B-box motif are known to disrupt MG53’s zinc-binding capacity18. We generated the rhMG53-C29L/C105S mutant protein (Fig. S7). Interestingly, the mutant rhMG53-C29L/C105S protein, which lost its zinc-binding ability18, showed significantly reduced capacity to activate both ADH and ALDH2, compared to the wild-type rhMG53 protein (Fig. 4I, J). Moreover, rhMG53-C29L/C105S was ineffective in protecting AML12 cells from EtOH-induced injury, as shown by the minimum impact on cell viability (Fig. 4K) and LDH release (Fig.4L). We performed additional experiments to evaluate the rhMG53-C29L/C105S mutant in the ALD mouse model. MG53−/− mice were subjected to a 4-week chronic-plus-binge ethanol feeding and treated with rhMG53-C29L/C105S via micro-osmotic pump (1 mg/kg with 1 µg/h). Unlike wild-type rhMG53, the mutant protein had no significant effect on alcohol-induced mortality or liver injury in MG53−/− mice (Fig. S7D–F).
Collectively, these findings reveal that MG53 acts as a chaperone by coordinating the transfer of zinc to ADH and ALDH2, thereby boosting their enzymatic function in metabolizing toxic alcohol byproducts under ALD conditions.
Developing a bio-nanoparticle for co-delivery of rhMG53 and zinc to treat ALD
Given the crucial role of MG53 and its interaction with zinc in combating ALD, we developed bio-derived MG53@Zn nanoparticles (MG53@Zn-NPs) as a delivery platform for the co-delivery of rhMG53 and zinc (Fig. 5A). We reasoned that this targeted co-delivery system could not only enhance liver regeneration through MG53’s tissue-protective properties but also improve the liver’s metabolic capacity to clear toxic alcohol byproducts by boosting the MG53-zinc-ADH/ALDH2 axis.
A Schematic illustration of the synthesis of MG53@Zn nanoparticles via desolvation. B Scanning electron microscopy image of MG53@Zn nanoparticles (Scale bar, 200 nm). Upper right: High-angle annular dark-field image (Scar bar, 100 nm). Element mapping shows the distribution of zinc (Zn, blue) and oxygen (O, red) in MG53@Zn nanoparticles. n = 5 independent experiments. C Size distribution of MG53@Zn nanoparticles. D Time-dependent uptake of Cy5.5-labeled MG53@Zn nanoparticles (10 μg/ml with rhMG53 dose equivalents) in AML12 cells. E, F Effect of MG53@Zn nanoparticles on cell viability and LDH release in AML12 cells. Cells were co-cultured with 400 mM EtOH and MG53@Zn for 24 h (n = 8 per group). G Effect of MG53@Zn on ROS levels in AML12 cells co-cultured with 400 mM EtOH and treated with MG53@Zn for 24 h, detected by DHE staining (Scale bars, 20 μm. n = 8 per group). H Effect of MG53@Zn on lipid accumulation in AML12 cells co-cultured with 400 mM EtOH and MG53@Zn treatment (10 μg/ml with rhMG53 dose equivalents) for 24 h, determined by Nile red staining and intensity analysis (Scale bars, 20 μm. n = 8 per group). I In vivo distribution of Cy5.5-labelled MG53@Zn (1 mg/kg with rhMG53 dose equivalents) in C57BL/6J mice and major organs. Whole-body fluorescence and ex vivo fluorescence of major organs were detected 24 h after injection. J Immunofluorescence analysis of MG53 enrichment in the liver of C57BL/6J mice 24 h post-injection of MG53@Zn (Scar bar, 20 μm. n = 6 per group). K Time course of MG53@Zn accumulation in the liver, showing retention from 0 to 72 h and a significant decrease at 120 h post-injection (n = 6 per group). L Bio-safety evaluation of MG53@Zn in mice using H&E staining of major organs (Scar bars, 50 μm). Mice were administered MG53@Zn (1 mg/kg with rhMG53 equivalents, every 5 days, intramuscularly) for 30 days and observed (n = 6 per group). Data were presented as mean ± SD. P-value was determined by one-way ANOVA with Tukey’s test. Source data are provided as a Source Data file.
The MG53@Zn-NPs were synthesized using a desolvation protocol, involving the sequential mixing of the rhMG53/Zn complex with human serum albumin (HSA) (Fig. 5A). The resulting nanoparticles had a multi-leaf shape and an average size of ~250 nm (Fig. 5B, C). When tested in cultured AML12 cells, Cy5.5-labeled rhMG53@Zn-NPs was taken by AML12 cells in a time-dependent manner (Fig. 5D) and provided superior protective effects against EtOH-induced injury (400 mM, 24 h) compared to rhMG53 alone, as evidenced by improved cell viability and reduced LDH release (Fig. 5E, F). Additionally, MG53@Zn-NP enhanced oxidative stress resistance in AML12 cells, as demonstrated by reduced DHE fluorescence (Fig. 5G) and MDA levels, along with increased SOD and GSH levels (Fig. S8A). These nanoparticles also lowered lipid oxidation, as indicated by decreased Nile red fluorescence (Fig. 5H).
Systemic administration of MG53@Zn-NP facilitated targeted delivery of the MG53/Zn complex to the liver. Following tail vein injection (1 mg/kg with rhMG53 equivalent) in normal mice, we observed prominent accumulation of Cy5.5-labeled rhMG53@Zn-NPs in the liver (Fig. 5I), confirmed by immunofluorescence staining and immunoblotting of Zn2+ and rhMG53 of hepatocytes (Fig. 5J and Fig. S8B). Liver frozen sections incubated with a Zn2+ probe further revealed a marked increase in intracellular Zn2+ fluorescence, with clear co-localization of the Zn2+ signal and Cy5.5-labeled nanoparticles by confocal microscopy (Fig. 5J). The half-life of rhMG53 in the liver was approximately 5 days, after intramuscular injection of MG53@Zn-NPs (Fig. 5K). We also conducted repetitive dosing of MG53@Zn-NPs in WT mice (1 mg/kg every 5 days for 30 days) and observed no adverse effects on vital organs (Fig. 5L and Fig. S8C). Before in vivo studies, we tested MG53@Zn-NPs in EtOH-stressed AML12 cells, where they restored ADH and ALDH2 activities that were diminished by EtOH exposure, surpassing endogenous ADH/ALDH2 activity levels (Fig. 6A). In EtOH-fed WT mice, more than 60% of the animals died within 8 weeks (Fig. 6B, red; Fig. S9A), but MG53@Zn-NPs notably improved the survival of the mice (Fig. 6B, purple).
C57BL/6J mice were fed an EtOH diet for 8 weeks. A pair-fed diet was used as a control. MG53@Zn nanoparticles (1 mg/kg with rhMG53 dose equivalents) were administered intramuscularly every 5 days, while rhMG53 (1 mg/kg) was administered by micro-osmotic pump. A ADH and ALDH2 enzyme activities in AML12 cells co-cultured with 400 mM EtOH and MG53@Zn (10 μg/ml with rhMG53 dose equivalents) for 24 h (n = 8 per group). B Survival rate of mice with pair-fed or EtOH-fed diets over 8 weeks with rhMG53 or MG53@Zn treatment (n = 20 per group). C Liver-body weight ratio in mice treated with MG53@Zn on pair-fed or EtOH-fed diets (n = 8). D Serum AST levels in treated mice after pair-fed or EtOH-fed diets (n = 8 per group). E H&E staining in liver tissues (Scale bars, 100 μm. n = 8 per group). F Liver steatosis evaluated by Oil Red O staining (Scale bars, 50 μm. n = 8 per group). G Hepatic triglyceride levels after MG52@Zn or rhMG53 treatment in mice with pair-fed or EtOH-fed diets (n = 8). H Hepatic zinc content in treated mice after 8 weeks on pair-fed or EtOH-fed diets (n = 6). I ROS levels in liver tissue measured by DHE staining and intensity analysis (Scale bars, 50 μm. n = 8 per group). J, K Serum ethanol and aldehyde levels in treated mice after 8 weeks on pair-fed or EtOH-fed diets (n = 8 per group). L Serum TNF-α levels in treated mice after 8 weeks on pair-fed or EtOH-fed diets (n = 8 per group). M Liver fibrosis evaluated by Sirius red staining (Scale bars, 50 μm. n = 8 per group). N Liver collagen Ⅰ expressions (n = 8 per group). Data were presented as mean ± SD. One-way ANOVA with Tukey’s test was performed for (A) survival curve with Log-rank (Mantel–Cox) test was performed for (B) two-way ANOVA with Sidak’s test was performed for (C, D, G–L and N). Source data are provided as a Source Data file.
To show the long-term effect of MG53@Zn-NPs on liver structure and function, we took 8-week treatment as the observation time-point, which is unlike that in MG53−/− mice. MG53@Zn-NPs reduced liver weight/body weight ratio (Fig. 6C), and serum AST, ALT, and LDH levels (Fig. 6D and Fig. S9B), indicating a reduction in EtOH-induced liver injury. Histological analysis showed improved hepatocyte integrity and steatosis in treated mice (Fig. 6E, F and Fig. S9C, D), with a mitigation of EtOH-induced triglyceride and cholesterol elevation (Fig. 6G and Fig. S9E).
Spectroscopic analysis of liver tissue revealed that MG53@Zn-NPs significantly restored zinc levels in EtOH-fed mice, bringing them close to normal (Fig. 6H). MG53@Zn-NPs also reduced oxidative stress (measured by DHE fluorescence) (Fig. 6I), improved the liver’s antioxidant function as evidenced by decreased serum levels of EtOH and aldehyde (Fig. 6J-K). Additionally, TNF-α, IL-6, and IL-1β, elevated by EtOH-induced inflammation, were markedly reduced by MG53@Zn-NPs (Fig. 6L and Fig. S9F), contributing to the long-term benefits of MG53@Zn-NPs in alleviating EtOH-induced liver fibrosis (Fig. 6M and Fig. S9G), as quantified by the changes in collagen I and Ⅳ as biomarkers of fibrosis (Fig. 6N and Fig. S9H). Importantly, MG53@Zn-NPs showed superior therapeutic efficacy for ALD compared to treatment with rhMG53 alone (1 mg/kg with 1 μg/h via micro-osmotic pump) (Fig. 6 and Fig. S9).
Discussion
Skeletal muscle, traditionally regarded as an organ for motion and energy metabolism, has been increasingly recognized for its secretory function, playing a vital role in inter-organ communication. In conditions of liver dysfunction, skeletal muscle can act as an auxiliary detoxification organ5,38, secreting myokines that modulate liver metabolism, inflammation, and tissue repair39,40. Our study identifies MG53 as a myogenic zinc metallochaperone that bridges muscle–liver crosstalk, highlighting its capacity to regulate zinc metabolism and protect the liver under chronic alcohol exposure.
We took an integrative approach combining bioinformatics and big data analysis to identify exogenous myokines, like MG53, that are enriched in the liver during ALD. The use of public databases for high-throughput screening of potential therapeutic targets is a relatively new approach in medical research, providing a cost-effective and time-efficient method for identifying novel disease-related factors24,25,26,27,41. As bioinformatics continues to evolve, it is likely to play an increasingly important role in accelerating the discovery and development of targeted therapies for a wide range of diseases. It is worth mentioning that we used Kaplan–Meier Survival curves for alcohol users with liver cancer as a rationale for studying MG53, but the dataset is based on transcriptomics41. The RNA expression of MG53 in liver tissue dictating survival would be through a different mechanism compared to the uptake of serum-based protein, such as exosome-based mRNA delivery, which can be further explored in future studies.
MG53’s involvement in liver health adds an important layer to our understanding of myokines as mediators of inter-organ communication. Previous studies have shown that other muscle-derived proteins, such as meteorin-like protein, offer protection against hepatic inflammation by modulating immune cell infiltration42, while irisin, another exercise-induced myokine, mitigates mitochondrial dysfunction and oxidative stress in the liver43,44. MG53 now joins this growing list of myokines, with our data revealing its crucial role in maintaining liver homeostasis through zinc-dependent regulation of alcohol metabolism enzymes, ADH and ALDH2. Our previous studies have demonstrated that loss of MG53 increases oxidative stress in various disease models (e.g., myocardial ischemia-reperfusion and chemical-induced lung injury), whereas rhMG53 protein administration attenuates such damage12,45. Chronic alcohol consumption shifts alcohol metabolism from the ADH-dependent pathway to the cytochrome P450 system via CYP2E1, increasing ROS production and causing Zn2+ deficiency2,21,46,47. This underscores a novel therapeutic mechanism by which muscle-derived factors can actively contribute to liver detoxification and protection.
MG53’s newly identified role as a zinc metallochaperone broadens our understanding of how proteins maintain intracellular zinc homeostasis, especially in the context of chronic diseases like ALD. Zinc plays a vital role in numerous biological processes, including enzyme function, gene expression, and cell signaling. Zinc metallochaperones, such as ZNG1 and METAP148, have been identified as critical for ensuring zinc is properly allocated to proteins or organelles. In the context of ALD, hepatic zinc deficiency is a hallmark that exacerbates liver dysfunction, compromising the organ’s antioxidant defenses and impairing its ability to metabolize alcohol effectively. Our data demonstrate that MG53 helps to preserve zinc homeostasis by facilitating zinc transfer to ADH and ALDH2, enhancing their activities. This zinc-mediated interaction enables more efficient detoxification of ethanol and its toxic byproducts, illustrating MG53’s dual role in muscle–liver communication and zinc regulation.
To further elucidate the molecular mechanisms underlying MG53’s regulation of ADH and ALDH2 functions, we employed molecular docking analysis to pinpoint key residues in MG53 essential for its interaction with these enzymes. Our findings revealed that MG53’s RING and B-box motifs play a critical role in coordinating zinc interactions with ADH and ALDH2. This zinc-chaperone ability enhances the enzymatic activity of ADH and ALDH2, thereby reducing oxidative stress and facilitating the clearance of toxic alcohol byproducts. Moreover, the role of extrahepatic ALDH2, particularly in the gut, is crucial in the overall systemic metabolism of alcohol and acetaldehyde, and a gut–liver axis is a key concept in ALD49. Delineating the specific role and mechanism of MG53 in the gut will be the subject of a dedicated future investigation in ALD. These findings not only expand MG53’s known role in membrane repair but also establish it as a central regulator of metabolic processes critical to liver health, especially under chronic-plus-binge ethanol exposure.
The therapeutic potential of MG53/Zn in ALD is especially significant, given the limitations of current treatments. ALD remains a major public health burden, with alcohol consumption contributing to approximately 3 million deaths annually worldwide46. Despite decades of research, specific therapies for alcoholic hepatitis and advanced ALD have remained elusive, in part due to the complex and multifactorial nature of the disease. Our study offers a promising new avenue for ALD treatment by leveraging MG53’s dual role in membrane repair and zinc homeostasis to restore ADH and ALDH2 activity and protect against ethanol-induced liver injury.
Our development of MG53@Zn-NPs represents a novel advancement in the targeted delivery of therapeutics. Nanoparticle-based drug delivery systems are increasingly being explored for their ability to improve bioavailability, reduce off-target effects, and enhance therapeutic outcomes50. By encapsulating MG53 and zinc within a bio-derived nanoparticle, we were able to overcome the limitations associated with direct zinc supplementation, such as poor bioavailability, potential toxicity, and inefficient liver targeting51. The hepatic targeting ability of MG53@Zn-NPs is primarily attributed to the intrinsic biological properties of its albumin shell, which facilitates both passive accumulation and active uptake in the liver52,53,54. Our MG53@Zn-NPs demonstrated remarkable stability, biocompatibility, and efficacy in vivo, offering a novel therapeutic approach with potential applications beyond ALD, in conditions involving oxidative stress and zinc dysregulation.
Extrahepatic ALDH2, particularly in the intestine, plays a critical role in systemic ethanol/acetaldehyde clearance, and the gut–liver axis is central to ALD pathogenesis49. In line with this concept, we observed MG53 accumulation in the gut of ethanol-stimulated mice. ALDH2 is well established as a mitochondrial enzyme, and our previous work demonstrated that MG53 localizes to mitochondria through binding to cardiolipin, thereby preserving mitochondrial integrity12,55. We have also shown that MG53 traverses the sinusoidal endothelium and enters hepatocytes to protect against liver injury13,29. Extending these findings, our biochemical analyses revealed a direct interaction between MG53 and ALDH2 in intestinal tissues from chronically ethanol-fed mice.
Despite the promising findings, our study has several limitations. The small sample size of human patients restricts the generalizability of our results, and future studies should aim to include larger, more diverse cohorts to confirm our findings. Additionally, while rodent models offer valuable insights, they have inherent limitations in accurately replicating human physiology. Future research should consider proof-of-concept studies in large animal models, which may provide more reliable data for translational applications. Moreover, the rhMG53 protein used in our study is produced via E. coli fermentation, and there is a need to explore alternative production methods, such as mammalian cell lines, to ensure the scalability and safety of rhMG53 for clinical trials.
Overall, our study identifies MG53 as a key regulator of muscle–liver communication in the context of ALD, acting as a zinc metallochaperone to enhance ADH and ALDH2 activity and promote alcohol metabolism. The development of MG53@Zn-NPs offers a novel, targeted therapeutic strategy for ALD, with potential broader applications in the treatment of other metabolic and oxidative stress-related diseases. Our findings provide a foundation for future studies aimed at translating these insights into clinical interventions that can benefit patients with ALD and related conditions.
Methods
Patients
Human samples were collected from Daping Hospital (Chongqing, China), with the study receiving approval from the Daping Hospital Institutional Ethics Review Board (ID223). Blood samples from ALD patients were obtained from individuals with a history of alcohol abuse spanning more than 5 years, who continued alcohol consumption at the time of sampling. Control blood samples were obtained from inpatients with no history of alcohol abuse and normal liver function. Liver biopsy tissues were obtained from patients with alcoholic hepatitis or cirrhosis, who had a similar history of alcohol abuse. Normal liver samples were obtained from non-pathological para-carcinoma tissues from patients undergoing surgical resection for hepatic carcinoma. All participants were fully informed and provided consent before sample collection. Liver tissues were either immediately submerged in liquid nitrogen for protein extraction and stored at −80 °C, or preserved in 4% paraformaldehyde for a minimum of 24 h before being flash-frozen or embedded in paraffin. Blood samples were processed by centrifugation at 16,000 × g for 15 min at room temperature to collect serum, which was subsequently stored at −80 °C. This study strictly adhered to the ethical guidelines outlined in the World Medical Association’s Declaration of Helsinki for research involving human participants.
Animals
All animal study protocols were approved by the Institutional Animal Care and Use Committee of the Third Military Medical University (Chongqing, China) (Certification No: AMUWEC20191177). Eight-week-old C57BL/6 mice were purchased from Vital River Laboratories (Beijing, China). MG53 knockout (MG53−/−) mice and their wild-type (WT) littermates were generated as described previously9,56. Mice were housed in individually ventilated cages, maintained under a 12:12 h light–dark cycle at 22–24 °C and 40–60% humidity, with free access to food and water. The animal facility maintains a thermoneutral micro-environment, eliminating hypothermia as a confounder. All procedures were performed under anesthesia, and every effort was made to minimize suffering. Mice were euthanized by cervical dislocation, and blood, liver, skeletal muscle, heart, spleen, lung, and kidney were harvested for further analysis.
To establish the ALD mouse model, mice were fed a Lieber–DeCarli ethanol-containing diet (purchased from XiaoShuYouTai Biotechnology Co., Ltd., Beijing, China), which involved increasing ethanol concentrations over time based on the NIAAA model57. The diet began with 0.75% ethanol for 3 days, followed by 1.5% for 4 days, and with 2.8% EtOH for either 4 or 8 weeks, finally followed by gavage of a single dose of ethanol (28.5%, 20 μL/g body weight) 9 h before being euthanized. Mice in the control group were given an isocaloric Lieber–DeCarli pair-fed diet. The rhMG53 protein from Novoprotein (Suzhou, China) was administered via micro-osmotic pump (ALZET, 1004, Beijing, China) at a dose of 1 mg/kg body weight. Mutated rhMG53 plasmids were constructed in the MiaoLing Plasmid Platform (MiaoLing Biology, Wuhan, China), and the mutant proteins (rhMG53-1A, rhMG53-3A, and rhMG53-C29L/C105S) were produced by Novoprotein.
Recombinant viruses and infection
Muscle-restricted gene transfer was accomplished with AAV9 vectors in which the MHCK7 promoter drives either MG53-EGFP or a control sequence (AAV9-MHCK7-Control and AAV9-MHCK7-Trim72-EGFP; HanBio, Shanghai)30,31. After isoflurane induction, the lower-limb fur of 8-week-old male C57BL/6J mice was clipped, and the skin was wiped with 70% ethanol. The same viral dose (1 × 1012 vg/ml, 10 µl per site) was slowly introduced into 4 sites—quadriceps femoris and gastrocnemius of both legs.
Histological analysis and transmission electron microscopy
Tissues were fixed in 4% paraformaldehyde for 24 h and embedded in paraffin. Thin sections (4 μm) were prepared using a microtome. For pathological observation, paraffin sections of liver, heart, spleen, lung, and kidney samples were stained with hematoxylin-eosin (H&E). To evaluate hepatic fibrosis, liver sections were stained with Sirius red. For analyzing hepatic lipid accumulation, fresh liver tissues were embedded in a Tissue-Tek optimal cutting temperature (OCT) compound (SAKURA, Torrance, California) and rapidly frozen. These samples were then sectioned to 8-μm using a cryotome (Sakura Finetek) and stained with Oil Red O. All histological staining kits were purchased from Solarbio Life Science (Beijing, China), and staining was performed according to the manufacturer’s instructions. The pathological features of liver tissues were assessed and scored as described previously58,59.
For transmission electron microscopy (TEM), fresh liver tissues were trimmed into 2 mm × 2 mm samples and fixed in 2% glutaraldehyde at 4 °C overnight. Regions were excised randomly using a glass scribe and mounted for ultramicrotomy. After negative staining, the sections were observed, and images were taken using a transmission electron microscope (Fischione Instruments).
Biochemical assays
The levels and activities of ALT, AST, LDH, cholesterol, triglyceride, MPO, SOD, GSH, MDA, and ethanol were quantified using specific assay kits from Solarbio Life Science, in accordance with the manufacturer’s protocols. These include Glutamic-pyruvic Transaminase Activity Assay Kit, Glutamic-oxalacetic Transaminase Activity Assay Kit, Lacate Dehydrogenase (LDH) Activity Assay Kit, Triglyceride Content Assay Kit, Total Cholesterol Content Assay Kit, Myeloperoxidase (MPO) Activity Assay Kit, Superoxide Dismutase (SOD) Activity Assay Kit, Glutathione Peroxidase (GPX) Activity Assay Kit, and Malondialdehyde (MDA) Content Assay Kit. Serum acetaldehyde was measured using the Acetaldehyde Assay Kit from Sigma-Aldrich, following the manufacturer’s instructions. Absorbance readings were measured at the corresponding wavelength using a spectrophotometer (Thermo Scientific).
Hepatic zinc detection
Liver tissues (about 100 mg wet weight per sample) were placed in 15 mL clean centrifuge tubes and dissolved with aqua regia. The treated samples were then introduced into the injector of an Inductively Coupled Plasma Mass Spectrometry (Perkin Elmer Nexion 1000, Waltham, MA). Following plasma ignition, the ion source was optimized to ensure full ionization of the sample elements, allowing them to enter the mass spectrometer. In the mass spectrometer, zinc ions were selectively screened and quantified based on their respective nucleus-mass ratio (m/z) for spectroscopic analysis. The ion signals detected by the mass spectrometer were subsequently transmitted to a computer for data collection and storage.
Mass spectrometry (MS)
Liver tissues were obtained from C57BL/6 mice after an EtOH-fed diet for 4 weeks. To detect liver proteins, the total protein of liver tissues was prepared and quantified using the BCA Protein Assay Kit (Beyotime Biotechnology), then diluted to 1 μg/μL with 200 μg of proteins per sample. Proteins were reduced with DTT and alkylated with IAA, then precipitated with acetone and redissolved in ammonium bicarbonate. Trypsin digestion was performed overnight at 37 °C. The peptides were desalted, resuspended in 0.1% FA, and separated by UltiMate3000 RSLCnano LC. Elution was performed with a gradient of solvents A and B. The eluate was analyzed by Orbitrap Exploris 480 MS in data-dependent mode. MS and MS/MS spectra were acquired with specific resolutions and settings. Data were processed using Proteome Discoverer software with the Mouse UniProt Protein Database. Peptides were identified using the SEQUEST algorithm with specified parameters, including trypsin as the protease, mass tolerances, and a false discovery rate of 1%.
According to the instructions of the Protein A/G Immunoprecipitation Kit (Beaverbio, Suzhou, China), the total proteins of liver tissues were extracted and incubated with BeaverBeads™ MG53 antibody (Proteintech, Wuhan, China) immunomagnetic beads at 4 °C overnight to form the MG53 antibody-antigen complex. Following magnetic separation, the beads containing the complex were washed and stored in immunoprecipitation washing buffer. Samples of MG53 antibody-antigen complex were analyzed using a Q Exactive mass spectrometer (Thermo Finnigan). Proteome Discoverer 1.4 software (Thermo Scientific) was used to analyze raw MS files and generate a data matrix that consisted of the retention time, mass-to-charge ratio (m/z) values, and peak intensity. Specific peptides were used to recognize protein. Bioinformatic analyses were performed by Bioinformatics online (https://www.bioinformatics.com.cn/).
Co-immunoprecipitation
Protein was extracted from mouse liver tissues as described above, and incubated with specific antibodies, gently rotating at 4 °C overnight. The immunocomplex was captured by adding BeaverBeads™ immunomagnetic beads and rotating gently at 4 °C for 3 h. The mixture was then centrifuged at 2500 × g for 5 min at 4 °C to obtain the precipitate. The precipitate was washed 3 times with ice-cold RIPA buffer, and the immunocomplex was separated from the beads by resuspending and boiling, followed by centrifugation to collect the supernatant. The supernatant was subsequently analyzed via SDS-PAGE, followed by western blotting with the appropriate antibodies. Rabbit anti-MG53, rabbit anti-ADH, and rabbit anti-ALDH2 antibodies were purchased from Proteintech.
Molecular docking
Protein structures of MG53, ADH, and ALDH2 were obtained from AlphaFold Protein Structure Database. These structures were introduced to the Molecular Operating Environment (MOE) 2019.01software to predict the binding interactions of MG53 and ADH, as well as MG53 and ALDH2. At the end of the docking procedure, 3D structural maps and planar representations were created, with binding sites marked. To assess the interaction and stability of protein complexes, a molecular dynamics simulation was performed using Gromacs 2022.3 software over a period of 100 ns. Stability was evaluated using Gibbs energy landscapes, Root Mean Square Deviation (RMSD), Root Mean Square Fluctuation (RMSF), Radius of Gyration, the number of hydrogen bonds, and Solution Accessible Surface Area (SASA), and MM/GBSA of contributions (Nine Chapters Calculation, Sichuan, China).
Protein–protein interaction assay
Biotin labeling
Initially, ADH and ALDH2 proteins were dissolved in phosphate-buffered saline with 0.02% Tween 20 (TBST) to achieve the appropriate concentrations. Streptavidin (SA) biotin was then added to 100 μl protein solutions at a molar ratio of 3:1 relative to the protein and incubated at room temperature for 30 min to ensure adequate conjugation. Labeled protein solutions were purified using a desalting column to remove unbound biotin.
Bio-layer interferometry assay
SA sensors were immersed in solutions containing biotin-labeled ADH or ALDH2 proteins to immobilize the proteins onto the sensor surface. The sensors were then incubated in solutions with varying concentrations of rhMG53 (or mutated) proteins to conduct binding assays. Real-time monitoring of interference spectral changes induced by the binding of rhMG53 (or mutated) proteins to the sensors was performed. Experimental procedures were carried out on the Octet RED96 instrument, utilizing consumables recommended by SARTORIUS (Beijing, China). A biotinylation kit was purchased from Genemore (Suzhou, China). Purified ADH protein and ALDH2 protein were purchased from CUSABIO (Wuhan, China), and purified rhMG53 (including mutated) proteins were purchased from Novoprotein.
Cell culture and viability assay
AML12 cells, mouse hepatic parenchymal cells, and the corresponding media were purchased from Pricella Biotechnology (Wuhan, China). Cells were maintained at 37 °C in a humidified cell incubator with 5% CO2, and the culture medium was replaced every two days. Following the fourth passage, cells at 60–70% confluency were used for experiments. To establish the ALD cell model, AML12 cells were treated with 50-, 100-, 200-, 400-, and 800-mM EtOH for 24 h to determine the appropriate ethanol concentration. For the therapeutic observation, cells cultured with ethanol were supplemented with a gradient concentration of rhMG53 (2.5–20 μg/mL) or rhMG53@Zn-NPs for 24 h.
For cell viability assessment, cells were plated in 96-well plates at a density of 5 × 103 cells/well and cultured overnight to allow for adherence. Cells were then pretreated with various interventions for 24 h at 37 °C. Cell viability was estimated using the cell counting kit-8 (CCK-8) colorimetric assay (Solarbio Life Science), and LDH release was detected by LDH Cytotoxicity Assay Kit (Beyotime), according to the manufacturer’s instructions.
Immunohistochemistry and immunofluorescence
Primary antibodies included mouse anti-MG53, rabbit anti-MG53, rabbit anti-CK18 (HUABIO, Zhejiang, China), rabbit anti-ADH, rabbit anti-ALDH2, and rabbit anti-Alb (all from Proteintech). Secondary antibodies included goat anti-rabbit and goat anti-mouse (Proteintech). For tissue staining, samples were fixed in 4% paraformaldehyde for 24 h, embedded in paraffin, and sectioned to 4-μm thickness. Paraffin-embedded sections were deparaffinized with deparaffinization buffer (Solarbio Life Science).
For immunohistochemical (IHC) staining, liver sections were immersed in 10 mM citric acid buffer (pH 6.0) for 30 min and treated with 3% H2O2 for 30 min to quench endogenous peroxidase activity. Sections were blocked in QuickBlock ™ Blocking Buffer for Immune Staining (Beyotime Biotechnology) for 1 h to prevent nonspecific antibody binding. Primary antibodies (1:100) were applied in a humidified chamber at 4 °C overnight, followed by secondary antibodies (1:100) for 1 h at room temperature. In each step, liver sections were rinsed with phosphate buffer saline (PBS) three times for 5 min each. Images were acquired using an SLIDEVIEW VS200 microscope (Olympus, Japan).
For tissue immunofluorescence (IF) staining, liver sections were immersed in 50× Citrate Antigen Retrieval Solution (pH 6.0) (Beyotime Biotechnology) for 30 min and cooled naturally to room temperature to retrieval antigen, and then blocked in QuickBlock ™ Blocking Buffer for Immune Staining for 1 h to prevent nonspecific antibody binding. MG53, CK18, ADH, ALDH2, and Alb staining were performed by incubating with the corresponding primary antibodies (1:100) in a humidified chamber at 4 °C overnight, followed by incubation with secondary antibodies (1:100) for 1 h at room temperature. In each step, liver sections were cleaned with PBS for 3 times × 5 min. Finally, sections were sealed with Antifade Mounting Medium with DAPI (Beyotime Biotechnology). Images were acquired using a Confocal Laser Scanning Microscope (Olympus).
For cellular immunofluorescence staining, AML12 cells were cultured in 24-well plates with cell slides at a density of 1 × 104 cells/mL. After treatment, the culture medium was removed, and cells were fixed with 4% paraformaldehyde for 30 min. Blocking and antibody incubation were performed with the above method, with PBS rinses performed three times for 5 min each. Finally, cell slides were sealed on the object slide with Antifade Mounting Medium with DAPI. Images were acquired using a Confocal Laser Scanning Microscope.
Cellular zinc level measurement
AML12 cells were cultured in 24-well plates covered with cell slides at a density of 1 × 104 cells/mL. After treatment, the culture medium was removed, and 4% paraformaldehyde was added to fix cells for 30 min, and washed with PBS for 3 times × 5 min. Cell slides were incubated with 25 μM zinquin ethyl ester (MedChemExpress, Shanghai, China) in PBS for 30 min at 37 °C, and washed with PBS for 3 times × 5 min. Finally, cell slides were taken out and sealed on the object slide with Antifade Mounting Medium with DAPI. Images were acquired using a Confocal Laser Scanning Microscope at 368 nm excitation.
ROS level measurement in liver tissue and AML12 cells
For ROS measurement, liver tissues were embedded in OCT compound, rapidly frozen, and sectioned into 8-μm-thick slides. The sections were cleaned with PBS for 3 times × 5 min. AML12 cells were also cultured in 24-well plates covered with cell slides and treated as necessary. Both cleaned frozen sections and cell slides were incubated with Dihydroethidium (DHE) solution (Beyotime Biotechnology) in a dark chamber at 37 °C for 30 min. After washing with PBS for 3 times × 5 min, samples were mounted with Antifade Mounting Medium with DAPI. Images were acquired using a confocal laser scanning microscope.
Enzyme activity assay
The enzyme activity of hepatic ADH and ALDH2 was evaluated using Athe lcohol Dehydrogenase Activity Assay Kit and Acetaldehyde Dehydrogenase Activity Assay Kit from Solarbio Life Science.
Enzyme-linked immunosorbent assay (ELISA)
Serum levels of IL-6, TNF-α, and IL-1β were determined by the mouse IL-6 ELISA kit, the mouse TNF-α ELISA kit, and the mouse IL-1β ELISA kit, separately, purchased from Solarbio Life Science. Measurement processes followed the manufacturer’s protocols.
Western blot
The total protein of liver tissues, serum, and AML12 cells was prepared and quantified using the BCA Protein Assay Kit (Beyotime Biotechnology). Equal amounts of protein samples were subjected to 4–20% SDS-PAGE and transferred to polyvinylidene fluoride membranes. The membranes were blocked with 5% nonfat milk-TBST at room temperature for 30 min and were washed with TBST for 3 times 5 min, and then were incubated with primary antibodies at 4 °C overnight, followed by incubating with fluorescent secondary antibodies in the dark at room temperature for 1 h, washed with TBST for 3 times × 5 min. The loading serum protein was determined by staining with ponceau S (MeilunBio, Shanghai, China) for photography and washed with TBST 3 times × 5 min. The primary antibodies include rabbit anti-MG53 (1:1000, Proteintech), rabbit anti-ADH (1:1000, Proteintech), rabbit anti-ALDH2 (1:1000, Proteintech), and mouse anti-GAPDH (1:1000, Proteintech). The secondary antibodies include goat anti-rabbit IR Dye 800 and goat anti-mouse IR Dye 800 (1:10,000, Li-Cor Biosciences, Beijing, China). The protein-bound complexes were detected by the Odyssey Infrared Imaging System (LI-Cor Biosciences), and the band intensities were measured by ImageJ software.
Quantitative real-time PCR
Total RNA samples of liver tissues were extracted using TRIzol reagent. Two μg of total RNA was used to synthesize the first-strand cDNA (TAKARA, Kusatsu, Japan) for reverse transcription, and the fluorescence was detected using the CFX96 Real-Time PCR Detection System (Bio-Rad, San Diego, CA) according to the manufacturer’s instructions. The expression of the target gene was normalized by GAPDH mRNA as an internal control.
Synthesis and characterization of MG53@Zn nanoparticles
To synthesize MG53@Zn-NPs, 1 mL of rhMG53 (2 mg/mL) was mixed with 0.2 mL zinc acetate solution (14 mg/mL), followed by stirring at 4 °C for 4 h. The mixture was then dialyzed with 8000–14,000 molecular weight dialysis bags (Solarbio Life Science) in deionized water at 4 °C for 12 h, resulting in MG53@Zn solution. Next, 400 μL human serum albumin (HSA, 10 mg/mL) was combined with 100 μL MG53@Zn solution. The pH was adjusted to 9 by adding 2–8 μL NaOH (1 M), followed by the addition of 4 times the volume of anhydrous ethanol (2 mL), at a drop rate of about 1 mL/min, while stirring at 4 °C for 10 min. Subsequently, 25 μL of NHS-TK-NHS linker solution (10 mg/ml, xarxbio, Shanxi, China) solution (10 mg/mL) was added, and the reaction was allowed to continue under stirring at 4 °C for 2 h. After dialysis in deionized water at 4 °C for 12 h, the final MG53@Zn-NPs were obtained.
The particle size and distribution of MG53@Zn-NPs were measured using Dynamic Light Scattering in Zeta Size Nano series (Malvern Panalytical, UK). To visualize the nanoparticles, the solution was dropped onto the surface of the 4 mm × 4 mm silicon wafer, dried naturally, and then imaged using a scanning electron microscope (SEM) (Fischione Instruments). For additional characterization, the NPs solution was applied to copper meshes (200-mesh), air-dried, and analyzed using TEM (FEI TECNAI G2 F20, Fischione Instruments), while the elemental composition was examined by XFlash 6T-30 detector (Bruker), generating specific elementpeak maps.
In vivo distribution and in vitro uptake of MG53@Zn-NPs
For in vivo distribution analysis, C57BL/6J mice were intravenously injected with MG53@Zn-NPs labeled with Cy5.5, a near-infrared dye. The bio-distribution of these NPs within the mice was detected and visualized by Small Animal In Vivo Imaging (SAIVI™). After imaging, the mice were euthanized with anesthesia, and their heart, livers, spleens, lungs, and kidneys were harvested to assess organ-specific distribution. The livers were further imaged to evaluate the localization of the MG53@Zn-NPs. For the in vitro experiments, AML12 cells were co-cultured with Cy5.5-labelled MG53@Zn-NPs for 0, 1, 3, 6, 12, and 24 h. After each time-point, cells were fixed and stained, then observed and imaged by a Confocal Laser Scanning Microscope to assess NP uptake over time.
Available data
Potential crosstalk proteins between skeletal muscle and alcohol-injured liver were revealed by combining ALD-related proteins, muscle-specific proteins, and secreted proteins, and then eliminating proteins detected in a normal liver. ALD-related proteins were obtained from the Human Liver Proteome Database (http://www.liverproteome.org/) and liver samples of ALD patients from our center (https://wwwn.cdc.gov/nchs/nhanes/Default.aspx)24. Muscle-specific proteins, secreted proteins, and liver-detected proteins were obtained from The Human Protein Atlas (https://www.proteinatlas.org/)25,26. The associations between expression levels of potential proteins and survival rates of liver cancer patients with alcohol abuse were evaluated on the Kaplan–Meier Plotter database (http://kmplot.com)41. Liver cancer patients with alcohol abuse were split by auto select best cutoff in 60 months follow-up threshold to produce the overall survival analysis27. The two patient cohorts with different expression levels of potential proteins are compared by a Kaplan–Meier survival plot, and the hazard ratio with 95% confidence intervals and log-rank P-value are calculated.
Epidemiological analysis of NHANES data
Study population: The data used in this study were derived from the National Health and Nutrition Examination Survey (NHANES) 2017–2018, a cross-sectional survey conducted in the United States. NHANES collects comprehensive data on the health and nutritional status of non-institutionalized U.S. children and adults through household interviews, mobile examination centers (MEC), blood samples, and physical examinations60. Inclusion criteria: All participants from the NHANES 2017–2018 survey. Exclusion criteria: (1) Pregnant individuals; (2) Individuals under 18 years of age; (3) Missing diagnosis of sarcopenia; (4) Missing records of alcohol consumption; (5) Missing data on liver damage diagnosis. The NHANES protocol received ethical approval from the National Centre for Health Statistics (NCHS) review board, with written informed consent provided by all participants.
Definition of ALD
The NHANES 2017–2018 survey incorporated vibration-controlled transient elastography (VCTE) to measure liver stiffness and the controlled attenuation parameter (CAP) to quantify hepatic fat content, providing an estimate of liver fibrosis. VCTE has been widely validated for assessing liver steatosis and fibrosis. CAP values range from 100 to 400 dB/m, with higher values indicating increased hepatic fat content61.
In this study, alcohol consumption was categorized into four levels: never drinking, light drinking, moderate drinking, and heavy drinking. “Never drinking” was defined as consuming alcohol fewer than 12 times in lifetime; “Former drinking” was defined as consuming alcohol more than 12 times a year and did not drink last year, or did not drink last year but drank more than 12 times in lifetime; “Mild drinking” included women consuming up to one drink per day and men consuming up to two drinks per day; “Moderate drinking” included women who drank at least two times per day and men who drank at least three times per day, or those who consumed alcohol at least two days per month in binge patterns; “Heavy drinking” was defined as women consuming at least three drinks per day and men consuming at least four drinks per day, or those who binge drank five or more days per month. Binge drinking was defined as consuming four or more drinks per day in women and five or more drinks per day in men62.
For the definition of ALD in this study, participants were required to meet all three of the following criteria: (1) Alcohol consumption of at least mild drinking; (2) CAP score ≥248 dB/m (to indicate hepatic steatosis)63; (3) Elevated alanine aminotransferase (ALT) or aspartate aminotransferase (AST) levels (women >19 U/L or men >29 U/L62). Participants who did not meet these diagnostic criteria were classified as non-ALD.
Definition of sarcopenia
The NHANES 2017–2018 survey utilized dual-energy X-ray absorptiometry (DXA) to measure body composition, a standard method for diagnosing sarcopenia. DXA-derived appendicular skeletal muscle mass (ASM), the sum of lean mass in the arms and legs, was used to define sarcopenia. In this study, the skeletal muscle mass index (ASMBMI = ASM/BMI), as proposed by the National Institutes of Health (NIH) Foundation, was used for sarcopenia diagnosis. Specifically, an ASMBMI < 0.789 for men and <0.512 for women was considered indicative of sarcopenia23.
Covariates
Body mass index (BMI) was calculated as weight (kg) divided by height squared (m²), and categorized into three groups: <25.0 kg/m², 25.0–30.0 kg/m², and ≥30.0 kg/m². Ethnicity was classified based on self-reported data from NHANES and included Mexican American, non-Hispanic Black, non-Hispanic White, other Hispanic, and other races (including non-Hispanic and multi-racial individuals). Marital status was categorized as married, divorced, widowed, or never married. Education was categorized into three levels: less than high school, high school graduate, and college graduate or higher. Household income was divided into three categories based on the poverty income ratio (PIR): <1.0, 1.0–3.0, and ≥3.0. Smoking status was categorized into three groups: never smoker (smoked fewer than 100 cigarettes in a lifetime), former smoker (smoked ≥100 cigarettes but currently not smoking), and current smoker (smoked ≥100 cigarettes and currently smoking either daily or occasionally).
Statistical analysis
For NHANES data
Given the complex multi-stage (stratified and clustered) sampling design of NHANES, all analyses were weighted using the “Survey” package in R, applying the 2017–2018 “WTMEC2YR - Full Sample 2 Year MEC Examination Weights” to adjust for survey design. For covariates with missing values, such as marital status, education, and smoking, multiple imputation was performed using the “Mice” package. Comparisons of baseline characteristics were conducted using independent-samples t-tests for continuous variables and χ2 tests for categorical variables. The relationship between sarcopenia and ALD was analyzed using two-sided logistic regression. Both unadjusted and adjusted models were used, adjusting for age, sex, race, marital status, smoking status, and alcohol consumption. Odds ratios (OR) and 95% confidence intervals (CI) were estimated. Nonlinear associations between ASMBMI (as a continuous variable) and the prevalence of ALD were analyzed using two-sided restricted cubic splines (RCS) with a likelihood ratio test. All statistical analyses were performed using R software (version 4.4.1), which is available at http://www.R-project.org. A p-value of <0.05 was considered statistically significant.
For experimental data
Data were expressed as mean ± standard deviation (SD). An unpaired Student’s t-test was used to compare two groups. For three or more group comparisons, one-way ANOVA with Tukey’s test or two-way ANOVA with Sidak’s test was performed. Survival curve with Log-rank (Mantel–Cox) test was performed for Kaplan–Meier survival analysis. Two-tailed linear regression with Pearson’s Correlation Coefficient was used to determine the correlation between two different variables. All data are representative of more than 3 independent experiments. GraphPad Prism 9 was used for statistical analyses. Results were considered to be statistically significant when p < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
National Health and Nutrition Examination Survey (NHANES) data were obtained from the available dataset at https://wwwn.cdc.gov/nchs/nhanes/Default.aspx. Original data for creating all graphs in the paper are provided in Source data 1. A public dataset of liver proteomics for ALD patients that supports the findings of this study is available in the HUMAN LIVER PROTEOME DATABASE (http://www.liverproteome.org/). Liver mass spectrometry data for ALD patients from our hospital center have been deposited in ProteomeXchange with the primary accession code PXD058395 (https://www.iprox.cn/page/home.html). Human skeletal muscle-specific proteins, secreted proteins, and normal liver-detected mRNA lists that support the findings of this study are available in the Human Protein Atlas (https://www.proteinatlas.org/). Original data for creating all graphs in the paper are provided in Source data 2 and 3. All the source data are provided with this paper. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request. Source data are provided with this paper.
References
Huang, D. Q. et al. Global epidemiology of cirrhosis - aetiology, trends and predictions. Nat. Rev. Gastroenterol. Hepatol. 20, 388–398 (2023).
Thursz, M. & Lingford-Hughes, A. Advances in the understanding and management of alcohol-related liver disease. BMJ 383, e077090 (2023).
Wu, X. et al. Recent Advances in understanding of pathogenesis of alcohol-associated liver disease. Annu. Rev. Pathol. 18, 411–438 (2023).
Thapaliya, S. et al. Alcohol-induced autophagy contributes to loss in skeletal muscle mass. Autophagy 10, 677–690 (2014).
Chang, K. V. et al. Is sarcopenia associated with hepatic encephalopathy in liver cirrhosis? A systematic review and meta-analysis. J. Formos. Med. Assoc. 118, 833–842 (2019).
Tantai, X. et al. Effect of sarcopenia on survival in patients with cirrhosis: a meta-analysis. J. Hepatol. 76, 588–599 (2022).
Pedersen, B. K. & Febbraio, M. A. Muscles, exercise and obesity: skeletal muscle as a secretory organ. Nat. Rev. Endocrinol. 8, 457–465 (2012).
Severinsen, M. C. K. & Pedersen, B. K. Muscle-organ crosstalk: the emerging roles of myokines. Endocr. Rev. 41, 594–609 (2020).
Cai, C. et al. MG53 nucleates assembly of cell membrane repair machinery. Nat. Cell Biol. 11, 56–64 (2009).
Weisleder, N. et al. Recombinant MG53 protein modulates therapeutic cell membrane repair in treatment of muscular dystrophy. Sci. Transl. Med. 4, 139–185 (2012).
Cao, C. M. et al. MG53 constitutes a primary determinant of cardiac ischemic preconditioning. Circulation 121, 2565–2574 (2010).
Gumpper-Fedus, K. et al. MG53 preserves mitochondrial integrity of cardiomyocytes during ischemia reperfusion-induced oxidative stress. Redox Biol. 54, 102357 (2022).
Han, Y. et al. Membrane-delimited signaling and cytosolic action of MG53 preserve hepatocyte integrity during drug-induced liver injury. J. Hepatol. 76, 558–567 (2022).
Duann, P. et al. MG53-mediated cell membrane repair protects against acute kidney injury. Sci. Transl. Med. 7, 279–236 (2015).
Coleman, J. E. Zinc proteins: enzymes, storage proteins, transcription factors, and replication proteins. Annu. Rev. Biochem. 61, 897–946 (1992).
Zago, M. P. & Oteiza, P. I. The antioxidant properties of zinc: interactions with iron and antioxidants. Free Radic. Biol. Med. 31, 266–274 (2001).
Zhou, Z. et al. Zinc supplementation prevents alcoholic liver injury in mice through attenuation of oxidative stress. Am. J. Pathol. 166, 1681–1690 (2005).
Cai, C. et al. Zinc binding to MG53 protein facilitates repair of injury to cell membranes. J. Biol. Chem. 290, 13830–13839 (2015).
Maret, W. Zinc biochemistry: from a single zinc enzyme to a key element of life. Adv. Nutr. 4, 82–91 (2013).
Lin, P. H. et al. Zinc in wound healing modulation. Nutrients 10, 16 (2017).
Bode, J. C. et al. Hepatic zinc content in patients with various stages of alcoholic liver disease and in patients with chronic active and chronic persistent hepatitis. Hepatology 8, 1605–1609 (1988).
Kang, X. et al. Zinc supplementation reverses alcohol-induced steatosis in mice through reactivating hepatocyte nuclear factor-4alpha and peroxisome proliferator-activated receptor-alpha. Hepatology 50, 1241–1250 (2009).
Studenski, S. A. et al. The FNIH sarcopenia project: rationale, study description, conference recommendations, and final estimates. J. Gerontol. A Biol. Sci. Med. Sci. 69, 547–558 (2014).
Niu, L. et al. Dynamic human liver proteome atlas reveals functional insights into disease pathways. Mol. Syst. Biol. 18, e10947 (2022).
Digre, A. & Lindskog, C. The Human Protein Atlas-spatial localization of the human proteome in health and disease. Protein Sci. 30, 218–233 (2021).
Ren, Z. et al. Single-cell reconstruction of progression trajectory reveals intervention principles in pathological cardiac hypertrophy. Circulation 141, 1704–1719 (2020).
Menyhárt, O., Nagy, Á & Győrffy, B. Determining consistent prognostic biomarkers of overall survival and vascular invasion in hepatocellular carcinoma. R. Soc. Open Sci. 5, 181006 (2018).
Ma, X. et al. The E3 ubiquitin ligase MG53 inhibits hepatocellular carcinoma by targeting RAC1 signaling. Oncogenesis 11, 40 (2022).
Yao, W. et al. MG53 anchored by dysferlin to cell membrane reduces hepatocyte apoptosis which induced by ischaemia/reperfusion injury in vivo and in vitro. J. Cell. Mol. Med. 21, 2503–2513 (2017).
Liu, Y. et al. Muscle-derived small extracellular vesicles induce liver fibrosis during overtraining. Cell Metab. 37, 824–841 (2025).
Ma, S. et al. Skeletal muscle-derived extracellular vesicles transport glycolytic enzymes to mediate muscle-to-bone crosstalk. Cell Metab. 35, 2028–2043 (2023).
Grüngreiff, K., Reinhold, D. & Wedemeyer, H. The role of zinc in liver cirrhosis. Ann. Hepatol. 15, 7–16 (2016).
Vatsalya, V. et al. Association of serum zinc with markers of liver injury in very heavy drinking alcohol-dependent patients. J. Nutr. Biochem. 59, 49–55 (2018).
Aydemir, T. B., Sitren, H. S. & Cousins, R. J. The zinc transporter Zip14 influences c-Met phosphorylation and hepatocyte proliferation during liver regeneration in mice. Gastroenterology 142, 1536–1546.e1535 (2012).
Wang, Z. et al. Effect of orally administered hydroxypropyl chitosan on the levels of iron, copper, zinc and calcium in mice. Int. J. Biol. Macromol. 64, 25–29 (2014).
Ma, Y., Ding, L., Li, Z. & Zhou, C. Structural basis for TRIM72 oligomerization during membrane damage repair. Nat. Commun. 14, 1555 (2023).
Park, S. H. et al. Structure and activation of the RING E3 ubiquitin ligase TRIM72 on the membrane. Nat. Struct. Mol. Biol. 30, 1695–1706 (2023).
Johnson, D. J. et al. Branched chain amino acid uptake and muscle free amino acid concentrations predict postoperative muscle nitrogen balance. Ann. Surg. 204, 513–523 (1986).
Bhanji, R. A. et al. Sarcopenia in hiding: the risk and consequence of underestimating muscle dysfunction in nonalcoholic steatohepatitis. Hepatology 66, 2055–2065 (2017).
Kuchay, M. S. et al. Non-alcoholic fatty liver disease-related fibrosis and sarcopenia: an altered liver-muscle crosstalk leading to increased mortality risk. Ageing Res. Rev. 80, 101696 (2022).
Győrffy, B. Discovery and ranking of the most robust prognostic biomarkers in serous ovarian cancer. GeroScience 45, 1889–1898 (2023).
Du, Y. et al. Meteorin-like protein overexpression ameliorates fulminant hepatitis in mice by inhibiting chemokine-dependent immune cell infiltration. Acta Pharmacol. Sin. 44, 1404–1415 (2023).
Guo, M. et al. Irisin ameliorates age-associated sarcopenia and metabolic dysfunction. J. Cachexia Sarcopenia Muscle 14, 391–405 (2023).
Bi, J. et al. Irisin alleviates liver ischemia-reperfusion injury by inhibiting excessive mitochondrial fission, promoting mitochondrial biogenesis and decreasing oxidative stress. Redox Biol. 20, 296–306 (2019).
Li, H. et al. MG53 attenuates nitrogen mustard-induced acute lung injury. J. Cell. Mol. Med. 26, 1886–1895 (2022).
Seitz, H. K. et al. Alcoholic liver disease. Nat. Rev. Dis. Primers. 4, 16 (2018).
Szabo, G. et al. IL-1 receptor antagonist plus pentoxifylline and zinc for severe alcohol-associated hepatitis. Hepatology 76, 1058–1068 (2022).
Weiss, A. et al. Zn-regulated GTPase metalloprotein activator 1 modulates vertebrate zinc homeostasis. Cell 185, 2148–2163.e2127 (2022).
Fu, Y. et al. Coordinated action of a gut-liver pathway drives alcohol detoxification and consumption. Nat. Metab. 6, 1380–1396 (2024).
Xue, C. et al. Bio-inspired engineered ferritin-albumin nanocomplexes for targeted ferroptosis therapy. J. Control. Release 351, 581–59 (2022).
Sun, Q. et al. Dysregulation of hepatic zinc transporters in a mouse model of alcoholic liver disease. Am. J. Physiol. Gastrointest. Liver Physiol. 307, G313–322 (2014).
Lian, B. et al. Folate-conjugated human serum albumin-encapsulated resveratrol nanoparticles: preparation, characterization, bioavailability and targeting of liver tumors. Artif. Cells Nanomed. Biotechnol. 47, 154–165 (2019).
Wang, Y. et al. Self-sulfhydrated, nitro-fixed albumin nanoparticles as a potent therapeutic agent for the treatment of acute liver injury. ACS Nano 18, 20772–20791 (2024).
Wang, W. et al. Fabrication and evaluation of nanoparticle-assembled BSA microparticles for enhanced liver delivery of glycyrrhetinic acid. Artif. Cells Nanomed. Biotechnol. 45, 740–747 (2017).
Liao, L. et al. MG53 deficiency mediated skeletal muscle dysfunction in chronic obstructive pulmonary disease via impairing mitochondrial fission. Redox Biol. 83, 103663 (2025).
Bian, Z. et al. Sustained elevation of MG53 in the bloodstream increases tissue regenerative capacity without compromising metabolic function. Nat. Commun. 10, 4659 (2019).
Bertola, A. et al. Mouse model of chronic and binge ethanol feeding (the NIAAA model). Nat. Protoc. 8, 627–637 (2013).
Altamirano, J. et al. A histologic scoring system for prognosis of patients with alcoholic hepatitis. Gastroenterology 146, 1231–1239 (2014).
Lackner, C. et al. Development and prognostic relevance of a histologic grading and staging system for alcohol-related liver disease. J. Hepatol. 75, 810–819 (2021).
Yetley, E. & Johnson, C. Nutritional applications of the Health and Nutrition Examination Surveys (HANES). Annu. Rev. Nutr. 7, 441–463 (1987).
Tang, A., Cloutier, G., Szeverenyi, N. M. & Sirlin, C. B. Ultrasound elastography and MR elastography for assessing liver fibrosis: part 1, principles and techniques. AJR Am. J. Roentgenol. 205, 22–32 (2015).
Rattan, P. et al. Inverse association of telomere length with liver disease and mortality in the US population. Hepatol. Commun. 6, 399–410 (2022).
Karlas, T. et al. Individual patient data meta-analysis of controlled attenuation parameter (CAP) technology for assessing steatosis. J. Hepatol. 66, 1022–1030 (2017).
Acknowledgements
This work was supported in part by the Major Science and Technology Projects in Yunnan Province (202302AA310041, 202403AC100021) to C.Z. and J.W., the National Natural Science Foundation of China (82470299, 82460021) to Y.H. and Y.Y., US National Institutes of Health (R01 AG072430, R01EY036243) to J.M., and Open Project of the Key Laboratory of Geriatric Cardiovascular and Cerebrovascular Disease Research Ministry of Education of China (KFKT202501) and Postdoctoral Innovative Talent Support Program (ZXBSH032) to T.S. We are deeply indebted to Xiaoyang Wang from the Army Medical University for her expert technical support in the realm of proteomics, and we would like to thank all the participants and professionals contributing to the NHANES, HUMAN LIVER PROTEOME DATABASE, and Human Protein Atlas.
Author information
Authors and Affiliations
Contributions
C.Z., Y.H., and J.M. conceptualized and designed the study, supervised the overall project, and participated in text revision. T.S. and X.Z. performed experiments, analyzed data, designed figures, and drafted the manuscript. J.W. participated in statistical analyses, text writing, and revising. H.J. constructed a nanoparticle, participated in animal imaging, and performed fluorescence labeling and big data analyses. Y.W., M.Z., P.C., J.X., and N.L contributed to clinical samples and information gathering from patients. B.H. and D.F. established the animal model system and performed pathological staining. L.J., Y.J., and L.Z. participated in animal model establishment, mass spectrometry analysis, and cell culture. L.Y. performed mass spectrometry analysis of human samples. M.T. and Y.Y. participated in animal observation and data gathering. K.L., Y.C., J.K., and R.L. participated in data analyses and manuscript writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xingxing Kong, Mingqian Tan, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shu, T., Zeng, X., Wang, J. et al. MG53 mediates skeletal muscle-liver cross-talk and enhances alcohol metabolism in alcoholic liver disease. Nat Commun 17, 2722 (2026). https://doi.org/10.1038/s41467-026-69132-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69132-6