Abstract
Inorganic phosphate (Pi) is essential for all living organisms. PstSCAB, a bacterial high-affinity ABC transporter, imports Pi under limiting conditions via five subunits: PstA and PstC forming the transmembrane domain (TMD), periplasmic PstS that switches between free and TMD-docked forms for Pi capture and delivery, and two cytosolic PstB subunits for ATP binding and hydrolysis. Its malfunction affects the virulence of pathogenic bacteria, making it pharmaceutically attractive. However, complete structural pictures of PstSCAB in different states remain lacking. Here, we determine cryo-EM structures of PstSCAB in resting, pretranslocation, and catalytic intermediate states, which reveal that conformational changes in PstS and ATP binding/unbinding in PstB collectively induce rigid-body movements of TMD, generating inward- or outward-facing conformations. In TMD, Pi specificity is determined by positively charged Arg220 (PstA) and Arg237 (PstC). This study advances understanding of bacterial Pi import and supports drug development targeting PstSCAB.
Similar content being viewed by others
Introduction
Phosphorus, the fifth most abundant element in cells, is essential for all living organisms. At the cellular level, phosphorus plays a role in the inheritance of genetic materials, energy metabolism, membrane integrity, and intracellular signaling. At the macroscopic level, misregulation of phosphorus can result in human disorders1, decreased agricultural productivity2, and lethality in microorganisms3, etc., therefore, it is of medical, agricultural, and pharmaceutical significance.
Inorganic phosphate (Pi) is the main form of phosphorus available in the environment, and different organisms have evolved distinct Pi translocating systems for the exchange of Pi across cell membranes. In Escherichia coli (E. coli), two Pi transporter systems are involved in Pi import: the low-affinity inorganic phosphate transporter (PiT) system, and the high-affinity phosphate-specific transport (Pst) system4. Under an excessive Pi environment, Pit is responsible for Pi uptake, and Pst is repressed, whereas under Pi-limiting conditions, Pst is activated. Conditional activation of PstSCAB is possible because it is part of the Pho regulon controlled by a two-component regulatory system comprised of a membrane-spanning histidine kinase sensor protein (PhoR) and a cytosolic response regulator (PhoB)5. Under Pi limitation, PhoR functions as a kinase that phosphorylates PhoB, allowing PhoB to bind to promoters with a consensus DNA sequence called the PHO box and regulate the expression of genes in the Pho regulon. When Pi is abundant, PhoR functions as a phosphatase that dephosphorylates PhoB, causing the dissociation of PhoB from downstream promoters and the reversal of gene expression regulation. Although the exact signal sensed by PhoR is not clear, extracellular Pi concentration has been ruled out, as PhoR does not seem to contain a periplasmic sensory domain; instead, alternative conformations in PstSCAB have been proposed as a possible signal6, probably involving PhoU that is encoded in the same operon (PstSCAB-phoU operon)7.
In addition to the genes involved in maintaining phosphate homeostasis, the Pho regulon contains numerous virulence genes associated with quorum sensing, chemotaxis, toxin secretion, and regulation, which contribute to changes in virulence-related phenotypes, such as biofilm formation, intestinal colonization, motility, and cytotoxicity. Therefore, pathological or experimental disruption of PstSCAB function has been shown to compromise its control on the Pho regulon and thereby the virulence of pathogenic bacteria8,9,10,11,12. For example, in urinary tract infections (UTI) causing Proteus mirabilis, PstS and PstC homologs were identified as virulence factor required for colonization of the urinary tract11. Some studies showed that mutations of PstA/C affected host cell adherence by atypical enteropathogenic E. coli (EPEC) and virulence of EPEC-like murine pathogen Citrobacter rodentium8. In another study adopting a functional genomics approach, PstS and PstC were among the identified virulence factors involved in Edwardsiella tarda pathogenesis12. Inactivation of the Pst system attenuated the virulence of Avian pathogenic E. coli (APEC) strain chi7122 in a chicken infection model10. In addition, PstSCAB is directly or indirectly involved in the pathogenicity of the septicemia-inducing E. coli strain in calves and piglets, and disruption of PstC by transposon abolished its pathogenicity9. More recently, it was shown that PstS from Mycobacterium tuberculosis (Mtb) PstSCAB is targeted by antibodies isolated from Mtb-infected patients13, implying its role in Mtb pathogenesis.
In PstSCAB transporter, PstS, a phosphate-binding protein with a two-lobed structure, is the periplasmic subunit that captures Pi from the periplasm and then delivers them to the substrate binding site in the transmembrane domain (TMD) formed by PstA and PstC. Therefore, PstS has to undergo conformational changes between the open, Pi-free state and closed, Pi-bound state. Two cytosolic PstB subunits form the nucleotide-binding domains (NBDs) that bind and hydrolyze ATP, providing the energy needed to release TMD-bound Pi into the cytosol. Although the periplasmic subunit PstS has been subjected to extensive structural studies13,14,15,16,17,18,19,20,21, the architecture of the whole PstSCAB complex and its structures in different functional states remain elusive, leaving the molecular mechanism of Pi specificity and Pi import in TMD unaddressed.
Here, we determine the cryo-electron microscopy (cryo-EM) structures of PstSCAB in resting, inward-facing/pretranslocation, and outward-facing/catalytic intermediate states. Along with ATPase assay, MST assay, Pi import assay, and molecular dynamics (MD) simulations, our study sheds light on the molecular mechanism of Pi specificity and import in PstSCAB.
Results
Biochemical characterization and structure determination
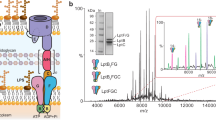
In E. Coli, all 4 subunits in PstSCAB are encoded in an operon named PstSCAB-PhoU. We initially tried to purify E. coli PstSCAB transporter from BL21(DE3) cells overexpressing the whole PstSCAB operon, utilizing an affinity tag on PstB, which only yielded a complex without PstS (referred to as PstCAB hereafter) (Fig. 1a). We then changed strategy and reconstituted PstSCAB complex using PstCAB and PstS that were expressed and purified separately, with an E179Q mutation in PstB that prevents ATP hydrolysis. Using ATPase assays, we showed that while the wild-type PstCAB displayed robust ATPase activity, the ATPase activity for PstCABE179Q was largely abolished (Fig. 1b, c). Substrate inhibition effect was observed at higher ATP concentrations (Fig. 1c), which was also observed in “Bce-type” ABC transporter BceAB from Bacillus subtilis22. Such inhibition is likely due to accumulation of Pi from ATP hydrolysis, as higher concentrations of Pi also showed inhibitory effect (Fig. 1d). Surprisingly, addition of PstS did not have notable effects on ATPase activity of PstCAB (Fig. 1e), unlike the stimulatory effect of substrate-binding proteins (SBPs) observed in other bacterial ABC importers such as maltose transporter23, and E. coli cysteine/cystine ABC transporter YecSC-FliY24. Additionally, using MST assays, we demonstrated that PstCAB could bind Pi, with a Kd of 7.76 mM (Fig. 1f).
a Purification of PstCAB. Size exclusion chromatography (SEC) trace and SDS-PAGE gel image were shown. SEC was repeated twice with similar results. b ATPase activity of wild-type PstCAB and PstCABE179Q. c Concentration-dependent ATPase activity of wild-type PstCAB. Data were fit with a model of substrate inhibition. d, e Effect of Pi (d) and PstS (e) on ATPase activity of wild-type PstCAB. f Binding curves for Pi with purified PstCAB, measured with MST assays. In b–f data represent mean ± SD (biological replicates, n = 3), except for the control in (f) (biological replicates, n = 2). Source data are provided as a Source data file.
By incubating PstCABE179Q and PstS with ATP, we managed to obtain the PstSCAB complex in an ATP-bound, catalytic intermediate state (referred to as PstSCABE179Q_ATP hereafter) (Supplementary Fig. 1a). However, we did not see binding between PstCAB and PstS in the absence of ATP. To capture PstSCAB in pretranslocation state (ATP unbound form), we inspected the interfaces between PstS and PstA/C in its ATP-bound outward-facing structure (described in detail later) and introduced two pairs of cysteine residues in N-lobe and C-lobe of PstS, and periplasmic loops of PstA and PstC, respectively (PstSN48C-PstAE149C and PstSK167C-PstCD63C, referred to as “4Cys” hereafter), followed by cross-linking with Dithio-bismaleimidoethane, with wild-type PstB subunits and in the absence of ATP, which yielded PstSCAB4cys representing pretranslocation state (Supplementary Fig. 1b).
Purified PstCAB, PstSCAB4Cys and PstSCABE179Q_ATP were reconstituted into nanodiscs and then subjected to single-particle cryo-EM analysis. 3D reconstructions yielded 4 high-resolution maps in 3 distinct conformations, namely resting, pre-translocation, and catalytic intermediate states, at overall resolutions of 3.0/3.3, 2.9, and 2.7 Å respectively (Supplementary Figs. 2–4 and Table 1). For the PstSCABE179Q_ATP dataset, two 3D reconstructions corresponding to inward-facing PstCAB and outward-facing PstSCAB conformations were obtained during 3D classification. Overall, the quality of the electron densities is superb for most part of the complex, allowing accurate model building (Supplementary Fig. 5). The only exception is PstB, which can be reliably modeled only in PstSCABE179Q_ATP, but not in PstCAB and PstSCAB4Cys, owing to its poor density. Therefore, PstB in PstCAB and PstSCAB4Cys was built by rigid-body fitting of its coordinates from PstSCABE179Q_ATP. The ATP-free, inward-facing PstCAB maps obtained from PstCAB and PstSCABE179Q_ATP are indistinguishable from each other (Supplementary Fig. 6a). Therefore, the one from PstCAB with higher resolution was deposited and used for discussion.
Structure of the resting state PstCAB
In the resting state, the transporter presumably has just gone through conformational changes that opened its Pi-binding site in the membrane-spanning subunits to the cytosol and released the bound Pi into the cell, and is ready for another Pi import cycle. In agreement with our biochemical data, the inward-facing structure consists of one copy of PstA and PstC, and two copies of PstB, but not PstS (Fig. 2a, b). This is similar to the resting state structure of MBP-MalFGK2 and Methanosarcina acetivorans ModBC25, whose SBPs are not docked, but different from those of trehalose importer LpqY-SugABC26, oligopeptide permease OppABCD27 and heme/tri- or tetra-peptides importer DppABCD28 from Mycobacterium tuberculosis (Mtb), whose SBPs are already associated with TMD in the resting state (Supplementary Fig. 7).
a, b Cryo-EM density map (a), and cartoon representations (b) of PstSCAB structure in the resting state. c Schematic representations of the topology of PstA and PstC. d Superimposition of PstA and PstC subunit, highlighting their high structural homology. e Pseudo symmetry observed between PstA and PstC in the PstCAB structure. f Residues in PstA and PstC involved in forming the gate facing the periplasm, as viewed from the angle indicated by the eye symbol in (b). g, h The coupling apparatus between PstB and PstA (g) or PstC (h), corresponding to the regions indicated in (b). Different subunits are colored differently (as labeled in a, b) throughout this figure. In b, the cytosol-facing cavity is shown as surface. In g, h, hydrogen bonds and salt bridges are shown as green and yellow dashed lines, respectively.
Both PstA and PstC contain six transmembrane helices (TMs) (Fig. 2c, d). Similar to ModB2C2 and MBP-MalFGK2, two subunits in the membrane (PstA and PstC) are intertwined, with TM1 of one subunit contacts with TM2-TM6 of the other subunit (Fig. 2e). Although of limited sequence homology (~19% sequence identity), the structure of PstA and PstC are similar to each other, with an root mean square deviation (RMSD) of 1.78 Å over 231 residues when superimposed, and are therefore related to each other via a 2-fold pseudo symmetry (Fig. 2e). PstA and PstC form a transmembrane heterodimer in the membrane, with a cavity that faces the cytosol and is inaccessible from the periplasm, owing to a closed gate formed by several patches of conserved residues (“gating regions”) (Fig. 2f). Residues in these “gating regions” are located at the periplasmic tip of TM5 from both PstA (Ala 226, Leu 229, and Phe 230) and PstC (Thr 242, Met 243, and Phe 247), and the loop between TM5 and TM6 in PstA (Phe 254) and PstC (Ala 272), right above the Pi binding site in the membrane (see later sections) (Fig. 2f). This gating regions ensure that the Pi binding site, which is located in the interface between PstA and PstC, is accessible only from the cytosol.
Two PstBs bind to PstA and PstC, one for each, through a coupling apparatus conserved among type I ABC transporters (e.g., a groove formed in the ATP-binding subunits and a coupling helix within the EAA loops of membrane-spanning subunits) (Fig. 2g, h). Like the counterparts in other bacterial ABC transporters, PstB forms a NBD, which can be further divided into RecA-like subdomain containing Walker A and B motifs and a helical subdomain containing the LSGGQ signature motif (Fig. 2b), essential for ATP binding. For some ABC transporters, e.g., MBP-MalFGK2 and Methanosarcina acetivorans ModBC, there is a regulatory domain in addition to NBD, which forms contact with that of the neighboring subunit (Supplementary Fig. 7). In PstSCAB, since no such regulatory domain is present, its resting state structure is held together solely by interactions between PstA and PstC (with a buried surface area of ~3468 Å2), which explains that the poor density at the distal end of PstBs is due to their flexibility (Fig. 2a).
Structure of PstSCAB in pretranslocation state
To capture the structure of PstSCAB in the pretranslocation state, we had to introduce cysteines into the interface between PstS and PstA/PstC, followed by crosslinking, implying that the pretranslocation (pre-T in short hereafter) state is probably transient (ATP binding and hydrolysis likely happen shortly after Pi-loaded PstS docks onto PstA/C). TMD of the pre-T state structure is in inward-facing conformation, similar to that of the resting state structure (Fig. 3a, b). One copy of closed, Pi-bound PstS docks on TMD from the periplasmic side (Fig. 3a, b).
a, b Cryo-EM density map (a) and cartoon representations (b) of PstSCAB structure in the pretranslocation state. c The Pi binding pocket in PstS and the bound Pi, with its electron density shown as mesh (contoured at rmsd threshold of 3 Å). d–f Interactions between the N-lobe of PstS and PstA (d), between the C-lobe of PstS and PstA (e), and between the P1 and P3 loop of PstC and PstS (f), corresponding to the regions boxed in (b). In b, the position of introduced cysteins is shown as yellow spheres. In c, d, hydrogen bonds are shown as green dashed lines.
As has been shown in previous structural characterization, PstS is structurally similar to other cluster D-IIIa SBPs and adopts a two-lobed structural fold that is shared among the periplasmic components of bacterial ABC type importers29 (Fig. 3b and Supplementary Fig. 7). Structure of PstS in the pre-T state is similar to the isolated crystal structure of PstS in Pi-bound closed conformation (pdb # 1ixh), with an RMSDs of 0.69 Å over 319 residues when superimposed (Supplementary Fig. 8a). In PstS structure of pre-T state, an extra non-protein density corresponding to Pi is observed in the cavity between the two lobes, which overlaps with bound Pi in the previously reported PstS crystal structure in Pi-bound form (Fig. 3c and Supplementary Fig. 8a). This observation consolidates our claim that this structure is in the pre-T state, in which PstS is loaded with Pi and is ready to release its bound Pi into the Pi binding site in the membrane (in the outward-facing or catalytic intermediate state, see below).
PstS docks onto PstA and PstC in a manner that its N-lobe sits above PstA, whereas its C-lobe sits above PstC (with a buried surface area of ~749 and ~606 Å2 respectively) (Fig. 3b). Between PstA and the N-lobe of PstS, Asp 21, Lys 30 and Asn 32 in PstS form hydrogen bonds with Asn 235, Gln 236 and Trp 238 in PstA respectively (Fig. 3d). Between PstC and the C-lobe of PstS, Lys 175 in PstS forms hydrogen bond with Asn 273 in PstC (Fig. 3f). In addition, Val 31, and Thr 49 in PstS form interactions with Cys 149, and Phe 237 in PstA (Fig. 3d). While the interactions between PstS and PstA/C seem sparse and occurred only at one side of each interface, two long loops between TM1 and TM2 (P1 loop), and TM5 and TM6 (P3 loop) in both PstA and PstC, extend upward into the cleft between N-lobe and C-lobe of PstS, and form additional interactions with them, which further stabilize PstSCAB complex (Fig. 3d–f). In PstA, while Pro 73 in P1 loop form hydrophobic interactions with the C-lobe of PstS (e.g., Gln 201), Pro 260 and Phe 261 in P3 loop form hydrophobic interaction with the C-lobe of PstS (e.g., Gln 201 and Asn 202) (Fig. 3e). Similarly, in PstC, Asn 273, Glu277 and Ser 280 in P3 loop form interactions with the N-lobe of PstS (e.g., Lys 43 and Lys 175) (Fig. 3f).
Surprisingly, the structures of PstA, PstC and PstB are almost identical between the resting and pre-T state, with an RMSD of 1.25 Å over 923 residues (Fig. 3a, b and Supplementary Fig. 6b). This is similar to LpqY-SugABC, but different from MBP-MalFGK2, as MalF and MalG were brought closer in the pre-T state upon binding of maltose-bound MBP, which is an intermediate between resting and outward-facing states (Supplementary Fig. 7)30. Other transient pre-T intermediate states may exist, where PstS adopts intermediate conformations between closed and open states. Transition to catalytic intermediate states is probably initiated by conformational changes in PstS, since two PstBs are too far away to jointly bind ATPs and initiate such transition. Potentially, in these pre-T intermediate states not captured in this study, two PstB monomers will be close enough to allow joint ATP binding by two PstBs, further accelerating the conformational transition of PstSCAB into an outward-facing conformation.
Structure of PstSCAB in ATP-bound, outward-facing state
In the ATP-bound state of PstSCAB, the binding of ATPs tightly glues two copies of PstB together, resulting in the outward-facing conformation of the whole transporter complex that allows the release of Pi from PstS into the Pi binding site in TMD (Fig. 4a, b). PstB binds ATP in a manner that is conserved in ABC transporters, involving residues in the Walker A and B motifs from one monomer and the LSGGQ motif of the opposite monomer (Fig. 4c). The docked PstS is in open, ligand-free conformation, highly similar to the previously reported crystal structure of PstS in the ligand-free state (PDB # 1oib), with a RMSD of 1.13 Å over 317 residues when superimposed (Supplementary Fig. 8b). The interfaces between PstS and TMD are dramatically different from that of pre-T state, involving completely different set of residues (Fig. 4d, e). For example, interactions are formed between Pro 71, Tyr 141, Ser 234 in PstA and Asp 21, Asn 32, Gln 34, Lys 43, Thr 44 in the N-lobe of PstS (Fig. 4d). On the other hand, Leu 132, Asn 251, Gln 254, Asn 273, in PstC are involved in forming interactions with residues in the C-lobe of PstS (Arg 134, Gly 138, Pro 169, Lys 175 and Arg 186) (Fig. 4e). In addition, Met 258 & Pro 260 in PstA, and Ala 276 and Glu 277 in PstC form interactions with residues in C-lobe (Tyr 198 and Gln 201) and N-lobe (Ser 38 and Ser 39) of PstS (Fig. 4d, e), further sealing the cavity between PstS and TMD. Consistent with the stable binding of PstS to TMD in the catalytic intermediate state, the buried surface areas are ~964 and ~909 Å2 between N-lobePstS and PstA, and C-lobePstS and PstC, respectively, as compared to ~749 and ~606 Å2 in the pre-T state. The increase in buried surface area seems to be mainly contributed by PstS (Supplementary Fig. 9). Moreover, dramatic conformational changes occurred in transmembrane subunits (PstA and PstC) and their interacting ATP-binding modules (PstBs), as will be discussed in detail later.
a, b Cryo-EM density map (a) and cartoon representation (b) of PstSCAB structure in the outward-facing, catalytic intermediate state. c Binding of ATP at the interface between two PstBs. Interactions of the N-lobe of PstS with PstA (d), and the C-lobe of PstS with PstC (e). Hydrogen bonds are shown as green dashed lines. f Pi binding site in the TMD, with modeled Pi shown as sticks, and its density shown as mesh (contoured at rmsd threshold of 3 Å). g Distance distribution histograms between the Pi molecule and the proximal residues from all MD runs. The distances were measured as the minimum distance between any atom pairs of two residue groups, including side chains. h Binding curves for Pi with purified PstCAB and its Pi-coordinating residue mutants, measured with MST assays. i Pi import activity of Pi binding residue mutants. In g data represent mean ± SD (for all data points in each run, n = 2000). In data represent mean ± SD (biological replicates, n = 3 for h, n = 3 for i), except for the control (biological replicates, n = 2) and PstCE241QAB (biological replicates, n = 4) in (h). Source data are provided as a Source data file.
A density is observed in the middle of TMD in the cryo-EM map of the outward-facing conformation of PstSCAB, inside the helical bundle formed by TM4 and TM5 of both PstA and PstC, but not in the inward-facing state and pre-T state, likely representing Pi (Fig. 4f). Although the density does not fit perfectly with a Pi molecule, several pieces of evidence support the identity of this density as being Pi. First, location of this density in PstSCAB is similar to the bound maltose in MBP-MalFGK2, as well as trehalose in LpqY-SugABC, all positioned in a cavity in the interface between two TMD subunits (Supplementary Fig. 7). Second, residues that are in coordination distance with this density are mostly positively charged, with their sidechains pointing toward the abovementioned density, which agrees well with the favored chemical environment for Pi binding (see below for details), similar to the Pi binding environment in XPR131. Moreover, as reported previously, mutations of some of these residues were shown to abolish the Pi transport activity, whereas the ATPase activity remained unaffected32,33,34. Third, the modeled Pi binds stably in the pocket throughout a 1 μs MD simulation (Fig. 4g and Supplementary Fig. 10). To rigorously assess Pi-binding stability, we analyzed the distances between Pi and its coordinating residues. Pi forms stable contacts with PstASer130, PstAArg220, and PstCArg237. The time-dependent distance profiles further confirm this claim: the distances from Pi to PstAArg220 and PstCR237 are exceptionally stable, showing minimal fluctuation over the entire simulation (Supplementary Fig. 11). The distance to PstASer133 exhibits more variation but remains within a bound state. In addition, we further calculated the root-mean-square fluctuation (RMSF) for both Cα atoms and side chains (averaged per residue) of the Pi-neighboring residues (Supplementary Fig. 12). The results confirm that the Cα-RMSF for all neighboring residues is very low, while the sidechain RMSF shows a certain mobility.
Pi is coordinated by residues from both PstA and PstC, e.g., the side chain of Arg 220 and the main chain of TM5 from PstA, as well as the side chain of Arg 237 and the main chain of TM5 from PstC (Fig. 4f). Earlier studies have shown that mutation of PstAArg220 and PstCArg237 largely compromised the Pi transport activity32,33,34. Moreover, mutation of PstCGlu241 also abolished Pi transport activity. According to our structure, PstCGlu241 forms salt bridge with PstCArg237, thereby stabilizing the Pi binding site (Fig. 4f). Pseudo-symmetrically, PstAGlu224 also forms salt bridge with PstAArg220 (Fig. 3f), although its mutation did not have any effects on Pi transport activity32. Using MST assays, we show that mutations of its coordinating residues largely reduced the Pi binding affinity of PstCAB (Fig. 4h and Supplementary Table 2). Moreover, in our Pi import assays, mutation of these residues by Crispr-cas9 editing in E. coli resulted in largely impaired Pi import activity when compared with that of the wild-type PstSCAB (Fig. 4i).
Structural transitions among different states
Dramatic structural changes were observed among PstSCAB structures in different states. While the main structural difference between PstCAB and PstSCAB is the absence or presence of PstS in PstCAB, structural changes occur broadly within the complex when comparing PstSCABE179Q_ATP and PstSCAB4cys (Fig. 5a).
a Structure superimposition of PstSCAB structures in the inward-facing and outward-facing state. b Comparison of PstS structures in inward-facing and outward-facing states. Their N-lobe was superimposed to highlight the movement in the C-lobe. c, d Comparison of the dimeric PstB structures in inward-facing (c) and outward-facing (d) states, shown as surfaces. The distance between a pair of Tyr152 in PstBs is indicated by double arrows and labeled. e, f Structure comparison of TM4-TM5 of PstSCAB in inward-facing (e) and outward-facing state (f). Side chains of the gating residues are shown as sticks. The distance between Tyr181 in PstA and Val198 in PstC (both near the coupling helix) is indicated by double arrows and labeled. Superimposition of PstA (g) and PstC (h) in inward-facing and outward-facing states. The view in the insets highlights the rigid-body movement in TM2-5. Cartoon representations were used throughout the figure except (c, d).
Specifically, PstS switches from a compact conformation in PstSCAB4cys (the cleft between two lobes of PstS is closed, with Pi engulfed inside it) to a loose conformation in PstSCABE179Q_ATP (the cleft between two lobes of PstS is open, without bound Pi), around the inter-domain hinge (Fig. 5b). When the N-lobes of PstS from these two states were aligned, the C-lobe of PstS in PstSCABE179Q_ATP swings ~15° away from the central cleft (Fig. 5b). Two PstBs undergo large conformational changes, involving both translational and rotational movements (Fig. 5c, d). As a result, two PstBs are brought closer to each other in PstSCABE179Q_ATP, allowing joint binding of two ATPs in the resulting dimer interface (Fig. 5c, d). In TMD, the gate at periplasmic side is closed and the gate at cytoplasmic side is open in the structure of PstSCAB4cys, therefore it represents the inward-facing, pre-T state (Fig. 5e). By contrast, in TMD of PstSCABE179Q_ATP, periplasmic half of PstA and PstC, which are directly coupled with two lobes of PstS, move away from the center of two-fold pseudo-symmetric axis, whereas their cytoplasmic half move towards the center of two-fold pseudo-symmetric axis (Fig. 5f). Consequently, the gate at periplasmic side becomes dilated and the gate at cytoplasmic side is constricted, thereby exposing the binding site in the membrane to the periplasm instead of the cytoplasm, which is the basis of alternative access model of most transporters (Fig. 5e, f and Supplementary Fig. 13). Cytoplasmic gate is mainly formed by TM4 and TM5 of PstA and PstC, which can move conveniently along with PstB, as the connecting helix between TM4 and TM5 (the coupling helix) in both PstA and PstC is responsible for forming direct interaction with PstB (Figs. 2g, h and 5e, f). Similar to other ABC transporters, conformational changes in TMD are achieved by rigid-body movements in PstA and PstC, particularly their TM2-TM5 (Fig. 5g, h).
Discussion
In this study, we presented the overall structure and Pi transporting mechanism of PstSCAB, the high-affinity Pi transporter broadly existed in bacteria, using single particle cryo-EM, complemented with biochemical (Pi transport assays and ATPase assays) and computational (MD simulations) approaches.
In short, our study provides several snapshots of the key steps in the Pi import cycle of PstSCAB (Fig. 6 and Supplementary Movie 1). In the resting state, PstCAB is in inward-facing conformation and is ready for the docking of PstS loaded with Pi from the periplasm side. Once Pi-loaded PstS is docked onto PstCAB, the pre-T PstSCAB state is formed, which is probably transient and will rapidly turn into the outward-facing, catalytic intermediate state (Fig. 6). This process is likely initiated by the opening of the cleft between N and C-lobe of PstS, which causes conformational changes in TMD (Fig. 6). As a result, two PstBs are brought closer to each other, allowing joint binding of ATP molecules, which further accelerate the formation of the outward-facing, catalytic intermediate state. During this process, Pi bound in PstS is released into the binding site in TMD (Fig. 6). Finally, ATP molecules are hydrolyzed into ADP and Pi, which breaks the PstB dimer and results in the formation of the inward-facing state of the Pi import complex (Fig. 6). During this step, Pi binding site in TMD becomes accessible from the cytosol, allowing Pi bound in TMD to be released into the cytosol. Meanwhile, PstS disassociates with other components of PstSCAB, resetting the complex into a resting state (Fig. 6). From this point, the Pi import complex is ready for another Pi import cycle. Although not captured in this study, a few other states must also exist, including a pre-T intermediate state, in which two PstBs are closer to each other and allow joint ATP binding, and an occluded state, in which Pi is sealed in the binding site in TMD and is inaccessible from either side of the membrane. It is also unclear after the catalytic intermediate state, during which state PstS becomes disassociated with TMD. Ultimately, integrating these structural insights with multi-scale simulations across all functional states will be crucial for a complete dynamical understanding of the transport cycle.
The states with or without experimental structures are highlighted by solid and dashed boxes, respectively. ATP and ADP molecules are represented by stars and rhombs, respectively. Pi molecules are shown as spheres. Question marks were used to indicate that it is unknown during which state PstS became disassociated with PstCAB.
Among the Gram-negative bacterial type I ABC importers with structural information, MBP-MalFGK2 is the most well-characterized30,35,36,37,38,39,40. Although PstSCAB shows similarity to MBP-MalFGK2 regarding overall architecture and translocation mechanism, there are a few striking differences. First, architecture-wise, while the two subunits (PstA and PstC) in PstSCAB contain 6-TMs commonly seen in bacterial ABC transporters (e.g., ModB2C241, DppABCD28 and LpqY-SugABC)26, the counterparts in MBP-MalFGK2 (MalF and MalG) possess extra TMs. Second, in the pre-T state, where both SBPs (PstS and MBP) are in substrate-bound closed conformation, while binding of MBP to MalFGK2 induces conformational changes in TMD into an intermediate conformation, binding of PstS does not have the same effect in PstCAB. Third, a scoop loop in TMD facing the periplasm has been observed in MBP-MalFGK2 and a few other bacterial ABC transporters, which can insert into the substrate binding site in SBPs and function to dislodge the bound substrate in SBPs26,27,28,40. However, the counterpart in PstSCAB does not seem to have the same function, as it is not inserted into the Pi binding site in PstS in the catalytic intermediate state (Fig. 4). Instead, these loops likely only function to seal the cavity formed between PstS and TMD.
Two Pi binding sites, one in PstS and the other in TMD, are observed in different structures of PstSCAB. The density for bound Pi in PstS of PstSCAB structure in the pre-T state is much stronger than that of the bound Pi in TMD (in the interface between PstA and PstC) of PstSCAB structure in the catalytic intermediate state, which correlates with a much more extensive interaction network (up to 13 hydrogen bonds) involved in Pi binding in PstS. Consequently, PstS has high substrate specificity towards monobasic/dibasic Pi, and against other inorganic oxyanions, including some close analogues of Pi, such as arsenate and sulfate14. By contrast, fewer interactions are involved in the binding of Pi in the TMD (Fig. 4f). Further studies will be required to see which site has more contributions in in achieving high Pi specificity in PstSCAB.
As introduced earlier, PstSCAB has been found to be essential for the virulence of various pathogenic bacteria. Strikingly, antibodies have been suggested to play a protective role in individuals exposed to Mtb. PstS1, the homolog of PstS in Mtb, was recently found to be the antigen targeted by at least two antibodies isolated from one Mtb-infected patient13. Superposition of our structure of holo PstSCAB complex with crystal structures of PstS1 in complex with the above-mentioned antibodies came to a conclusion that is in line with the one drawn from the superimposition of Mtb PstS1 with MetQ of MetNIQ methionine ABC transporter complex: these antibodies do not work by disrupting the interaction between PstS and TMD13 (Supplementary Fig. 14). Our accurate atomic model provides a better template for generating a structure model for Mtb PstSCAB, which can be used for screening or designing antibodies with protective effects against Mtb infection. In addition, the same strategy could be applied to screen or design protective antibodies or bioactive peptides (an important source for new drug development42,43,44,45,46,47,48) against bacteria that are infectious to humans or animals, e.g., those involved in urinary tract infection, pneumonia, or peritonitis.
Methods
Cloning and overexpression of PstS and PstSCAB
All primers used in this study have been listed in Supplementary Table 3. The protein-coding sequence of pstS and pstSCAB, which are contained in the pstSCAB-phoU operon, were amplified from E. coli DH5α genomic DNA by PCR, and cloned into pET28a(+), using either conventional digestion and ligation (pstS, digested with Nco I/Xho I) or Gibson assembly (pstSCAB). These constructs allowed the expression of target proteins fused with a 6×His tag at the C terminus. For preparing the inward-facing, pre-translocation state and outward-facing, catalytic intermediate state samples, constructs for expressing PstSN48C/K167C and PstSCD63CAE149CB or PstSCABE179Q were prepared following the standard protocol of site-directed mutagenesis, and verified by sequencing (Sangon Biotech, China).
The resulting plasmids were transformed into E. coli BL21 (DE3). Single colonies were cultured in Luria broth (LB) with 50 μg/mL kanamycin for 12 h at 37 °C. The resulting starter culture was transferred into 1 L LB medium containing kanamycin (50 μg/ml), and incubated at 37 °C until the OD600 reached 0.8. Then, 0.5 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) was added, and the induction continued at 16 °C for 20 h. Cells were harvested by centrifugation, flash-frozen in liquid nitrogen, and stored at −80 °C.
Protein purification
To purify the PstCAB/PstCABE179Q complex, frozen cell pellets were re-suspended in lysis buffer containing 50 mM Tris, pH 8.0, 200 mM NaCl and 1 mM PMSF (buffer A), and lysed by using a high-pressure homogenizer (JNBIO, Guangzhou) at 1000 bar for 4 cycles (same procedure for cell lysis throughout the method section). Unbroken cells and large debris were removed by centrifugation at 10,000 × g for 30 min at 4 °C, and 1 % (w/v) Lauryl maltose neopentyl glycol (LMNG, Anatrace) were added into the supernatant for membrane protein extraction (gentle agitation for 2 h at 4 °C). After centrifuging at 4 °C, 20,000 × g for 30 min, the membrane protein-containing supernatant was applied to a disposable column with 1 mL Nickel-nitrilotriacetic acid (Ni-NTA) agarose beads (GE Healthcare). After extensive wash steps, the protein was eluted with 20 mM Tris, pH 8.0, 150 mM NaCl (buffer B) containing 0.01 % (w/v) LMNG and 200 mM imidazole, concentrated, and purified using a Superose 6 Increase column (GE Healthcare) in buffer B containing 0.006 % (w/v) LMNG. Protein fractions with the highest homogeneity were collected and concentrated to ~4 mg ml−1 and stored at −80 °C.
To obtain the PstSCAB4Cys complex, PstSN48C/K167C and PstCD63CAE149CB protein were purified separately. The cell pellets of PstSN48C/K167C were re-suspended in phosphate-buffered saline (PBS) (pH 7.2) containing 1 mM PMSF and lysed. Unbroken cells and large debris were removed by centrifugation at 10,000 × g for 30 min at 4 °C. The supernatant was loaded on Ni-NTA agarose beads (GE Healthcare) affinity column and incubated for 30 min at 4 °C followed by elution with PBS buffer containing 200 mM imidazole. The protein was then loaded into a Superdex 75 Increase column equilibrated with PBS, and peak fractions were pooled. PstCD63CAE149CB was purified following a similar protocol to PstCAB/PstCABE179Q except for using PBS buffer containing 1% (w/v) LMNG for membrane protein extraction. PstSN48C/K167C and PstCD63CAE149CB were then incubated in PBS buffer containing 0.01% LMNG (w/v) at a molar ratio of 1:1 with the presence of 1 mM dithio-bismaleimidoethane (DTME, Thermo Scientific) for 1 h at room temperature. Successful crosslinking was confirmed by treatment with 100 mM dithiothreitol (DTT, Thermo Scientific) followed by SDS-PAGE electrophoresis (Supplementary Fig. 1b). The crosslinked products were then purified using Superose 6 Increase column equilibrated with PBS, pH 7.2 containing 0.006 % (w/v) LMNG.
To obtain the PstSCABE179Q_ATP complex, PstS was purified following a similar protocol to PstSN48C/K167C, except for using lysis buffer containing buffer A. The purification procedure for the PstCABE179Q complex was described above. The protein samples (PstS: PstCABE179Q at a molar ratio of 2:1) were incubated with 1 mM ATP, 1 mM MgCl2, and 2 mM HPO42− for 30 min at room temperature, followed by size-exclusion chromatography on a Superose 6 Increase column (GE Healthcare) in buffer B containing 0.006 % (w/v) LMNG to yield a stable complex.
For Nanodisc reconstitution, PstCAB, PstSCAB4Cys or PstSCABE179Q_ATP, MSP1D1 membrane scaffold protein, and POPG were mixed at a molar ratio of 1:1:30 in buffer B containing 0.006 % (w/v) LMNG and incubated for 1 h at 4 °C. Detergents were removed by incubation with 50 mg ml−1 Bio-Beads SM2 (Bio-Rad) for 1 h at 4 °C, followed by centrifugation. For the resulting supernatant, the same procedure was repeated 3 more times (incubation period of 1 h, 1 h, and overnight, respectively). Nanodisc-embedded PstCAB, PstSCAB4Cys or PstSCABE179Q_ATP were further purified using a Superose 6 Increase column in buffer B.
Cryo-EM sample preparation and data acquisition
For the PstCAB, PstSCABE179Q_ATP and PstSCAB4cys samples, Quantifoil R1.2/1.3(300 mesh) holey carbon grids were first glow discharged for 50 s using a Pelco easiGlow glow discharge unit, and 4 μL sample was applied to the surface of the grid at a temperature of 5 °C and humidity level of 100%. Grids were then blotted for 2.0 s before being plunge-frozen in liquid ethane using Vitrobot Mark IV. The cryo-specimens were loaded onto a Titan Krios transmission electron microscope (Thermo Fisher) operated at 300 kV for data collection. The microscope is equipped with a GIF-Quantum energy filter (Gatan), which was used with a slit width of 20 eV. Automatic data collection was performed using EPU software. Images were recorded with Gatan K3 direct electron detectors operating in super-resolution counting mode at pixel sizes of 0.856, 0.856 and 0.67 Å, respectively. The exposure was performed with a dose rate of 15 e-/pixel/s and an accumulative dose of ~50 e-/Å2 for each image, fractionated into 36 movie-frames. The final defocus ranges of the datasets were approximately −(1.2–2.5) μm.
Image processing and 3D reconstruction
For the PstCAB data, the dose-fractionated image stacks were subjected to beam-induced motion correction using MotionCor249. Initial contrast transfer function (CTF) values for each micrograph were calculated with CTFFIND450. Micrographs with an estimated resolution limit worse than 6 Å were discarded in the initial screening. About 10,000 particles were blob-picked and subjected to 2D classification to generate templates for auto-picking against the entire dataset. The subsequent image processing and reconstruction were performed using cryoSPARC51. 3,228,106 particles were picked from 6777 micrographs. Then the picked particles were extracted and subjected to three rounds of reference-free 2D classification in cryoSPARC, which yielded 1,194,741 particle projections. This subset was subjected to one round of ab-initio reconstruction followed by Heterogeneous Refinement. The predominant class containing one subset of 579,652 best particles shows the clear features of secondary structural elements. These particles were subjected to a non-uniform refinement, which yielded a reconstruction at 3.0 Å resolution (Supplementary Fig. 2).
The PstSCABE179Q_ATP and PstSCAB4cys datasets were processed similarly to the above PstCAB data. The detailed image processing and reconstruction are shown in Supplementary Figs. 3 and 4. Local resolution estimate was performed with cryoSPARC.
Model building
Model building was manually conducted in Coot52. Accurate residue registration was achieved based mainly on the clearly defined densities for bulky residues (Phe, Trp, Tyr, and Arg). Models were refined against cryo-EM maps using real-space refinement in PHENIX53, with secondary structure and non-crystallography symmetry restraints applied. The statistics of the models’ geometries were generated using MolProbity54. Ion translocating pathways were calculated using Caver 3.055. All the figures were prepared using PyMol56 or ChimeraX57.
ATPase assay
All ATPase assays were performed using the ATPase/GTPase Activity Assay Kit (Sigma-Aldrich). Purified PstCAB and PstCABE179Q (1 μg) were incubated in a 20-μl reaction volume in buffer B containing 1 mM ATP and 1 mM MgCl2 (Sangon Biotech) for 10 min at 37 °C, which represents the standard reaction system. For measurement of concentration-dependent ATPase activity, varying concentrations of ATP were used. For the effect of Pi and PstS, varying concentrations of Pi and PstS were added in the standard reaction system. After the reaction was stopped and the colorimetric product was generated, absorbance at 620 nm was measured at room temperature using a SpectraMax iD3 multifunction reader (Molecular Devices). ATPase activity was represented as phosphate (nanomoles) produced by 1 mg of protein per minute.
Microscale thermophoresis (MST) assays
The PstCAB complex and its relevant mutants were prepared following a similar protocol to PstCAB/PstCABE179Q except for using the Hepes buffer (pH 7.4). Protein labeling was performed using the Monolith His-Tag Labeling Kit RED-tris-NTA 2nd Generation (Nanotemper Technologies, Cat# MO-L018). Specifically, 90 μL of His-tagged proteins (200 nM) and 90 μL of dye (100 nM) were mixed and incubated for 30 min at room temperature under no-light conditions. Following centrifugation at 15,000 × g for 10 min at 4 °C, the labeled protein was immediately subjected to phosphate-binding assays using the Monolith NT.115 (NanoTemper Technologies). The assay mixture containing labeled proteins and varying concentrations of Pi was loaded into the Monolith NT.115 capillaries (Nanotemper Technologies, Cat# MO-K022). The MST measurements were performed with 40% LED/excitation power and high MST power. All experiments were performed in triplicate. Data analysis was carried out using the manufacturer’s analysis software (NanoTemper software).
Gene editing
A modified CRISPR/Cas9 system (pEcCas/pEcgRNA) was used for gene knock-out and knock-in in E. coli strain TOP10, following the procedure described previously58. In brief, the pEcCas plasmid, encoding Cas9 and λ-Red recombinase, was used to facilitate homologous recombination, with λ-Red expression induced by 10 mM L-arabinose. The pEcgRNA plasmid, carrying the sgRNA and an N20-specific sequence, was employed to target the gene for editing. Target-specific 20 bp spacers were predicted using the CHOPCHOP website (https://chopchop.cbu.uib.no). Homologous arms, 500 bp up and downstream of the target gene, were linked via overlapping extension PCR to create the knockout donor template. The strain carrying pEcCas was cultured at 37 °C until the OD600 reached 0.2, after which 10 mM arabinose was added to induce λ-Red recombinase expression. The culture was kept at 37 °C until the OD600 reached 0.8, after which competent cells were prepared. These cells were electroporated with 200 ng of pTarget plasmid and 800 ng of donor DNA, and transformants were plated on LB agar containing 50 μg/mL kanamycin and 50 μg/mL spectinomycin after 1 h of incubation at 37 °C. Positive transformants were confirmed by colony PCR and DNA sequencing. The pTarget plasmid was cured using 10 mM rhamnose, and the pEcCas plasmid was eliminated by adding 10 mM sucrose.
Phosphate uptake measurements
Procedure for Pi uptake measurement is adapted from a previous study6. Two types of medium were used: MOPS (morpholinepropanesulfonic acid), MOPS with 0.4% glucose and 0.1 mM Pi (referred to as MOPS LoPi). Cells were grown overnight in 5 ml MOPS LoPi. After washing twice with 5 ml MOPS medium, the cells were resuspended to an OD600 of ~0.45 in MOPS medium containing 0.4% glucose, but no Pi. The Pi-starved cells were incubated at 37 °C on a roller drum for 2 h. For transport assays, 750 μl of cells at an OD600 of 0.45 were added to 750 μl of a 10.5 μM solution of K2HPO4. After incubating for 45 s at room temperature, 1000 μl of cells were removed and filtered through pre-wet 0.2 μm nitrocellulose filters. Using the Malachite Green assay (Sigma), Pi concentrations were measured for each filtrate sample. The net amount of Pi (in nmoles) in the filtrate (the amount in a blank reaction containing no cells was subtracted) was divided by the dry weight of cells in each sample and the time, which yielded the Pi uptake rate. Each strain was assayed using three biological replicates in triplicate.
Molecular dynamics simulations
Systems for the PstSCAB structure in the catalytic intermediate state were built using the CHARMM-GUI server59, with each structure embedded in a palmitoyl-oleyl-phosphatidylcholine bilayer and hydrated with 50,662 TIP3P water molecules60. The final systems were neutralized with the sodium ion, and extra NaCl was added to model the experimental salt concentration of 0.15 M, resulting in a simulation box approximately 110 Å × 110 Å × 173 Å in dimension, encompassing a total of 216,586 atoms. All simulations were carried out using GROMACS-2025.2 software package61 with the CHARMM36m force field62. The van der Waals cutoff was set to 12 Å and PME was used for long-range electrostatic energy calculations. These systems were first minimized for 50,000 steps with the steepest descent algorithm, followed by an equilibration process at 300 K for 1125 ps in NPT ensemble. The final production simulations were performed at 300 K under the NPT ensemble with the integration step of 2 fs. The stochastic velocity rescaling method63 and the Parrinello–Rahman method64 were used for temperature and anisotropic pressure coupling, respectively. Three independent runs with the identical starting structures and randomized initial velocities were carried out to improve samplings, with each trajectory lasting for 1 μs. All data were analyzed using the facilities from GROMACS-2025.2 software package and all figures were prepared using gnuplot 5.4. The RMSD values were computed relative to the initial structure with Cα atoms. Supplementary Fig. 10 shows that the systems converged rapidly. Accordingly, data from the 400–1000 ns time interval were selected for all subsequent analyses to ensure converged calculations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Structure coordinates and cryo-EM density maps have been deposited at the Protein Data Bank and Electron Microscopy Data Bank under accession numbers 9K3Y and EMD-62032 for PstSCAB in the resting state; 9K3X and EMD-62031 for PstSCAB in pretranslocation state; and 9K3S and EMD-62027 for PstSCAB in catalytic intermediate state. Other structure coordinates analyzed in this paper are indicated in the text. Final states of the MD simulation trajectory are available as Supplementary Data 1. The source data underlying Figs. 1a–f; 4h, i and Supplementary Fig. 1a–c are provided as a Source data file. Source data are provided with this paper.
References
Legati, A. et al. Mutations in XPR1 cause primary familial brain calcification associated with altered phosphate export. Nat. Genet. 47, 579–581 (2015).
Liu, J. et al. A vacuolar phosphate transporter essential for phosphate homeostasis in Arabidopsis. Proc. Natl. Acad. Sci. USA 112, E6571–E6578 (2015).
Sethuraman, A., Rao, N. N. & Kornberg, A. The endopolyphosphatase gene: essential in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 98, 8542–8547 (2001).
Jansson, M. Phosphate uptake and utilization by bacteria and algae. Hydrobiologia 170, 177–189 (1988).
Santos-Beneit, F. The Pho regulon: a huge regulatory network in bacteria. Front. Microbiol. 6, 402 (2015).
Vuppada, R. K., Hansen, C. R., Strickland, K. A. P., Kelly, K. M. & McCleary, W. R. Phosphate signaling through alternate conformations of the PstSCAB phosphate transporter. BMC Microbiol. 18, 8 (2018).
Gardner, S. G., Johns, K. D., Tanner, R. & McCleary, W. R. The PhoU protein from Escherichia coli interacts with PhoR, PstB, and metals to form a phosphate-signaling complex at the membrane. J. Bacteriol. 196, 1741–1752 (2014).
Cheng, C. et al. Contribution of the pst-phoU operon to cell adherence by atypical enteropathogenic Escherichia coli and virulence of Citrobacter rodentium. Infect. Immun. 77, 1936–1944 (2009).
Daigle, F., Fairbrother, J. M. & Harel, J. Identification of a mutation in the pst-phoU operon that reduces pathogenicity of an Escherichia coli strain causing septicemia in pigs. Infect. Immun. 63, 4924–4927 (1995).
Lamarche, M. G. et al. Inactivation of the pst system reduces the virulence of an avian pathogenic Escherichia coli O78 strain. Infect. Immun. 73, 4138–4145 (2005).
Burall, L. S. et al. Proteus mirabilis genes that contribute to pathogenesis of urinary tract infection: identification of 25 signature-tagged mutants attenuated at least 100-fold. Infect. Immun. 72, 2922–2938 (2004).
Srinivasa Rao, P. S., Lim, T. M. & Leung, K. Y. Functional genomics approach to the identification of virulence genes involved in Edwardsiella tarda pathogenesis. Infect. Immun. 71, 1343–1351 (2003).
Watson, A. et al. Human antibodies targeting a Mycobacterium transporter protein mediate protection against tuberculosis. Nat. Commun. 12, 602 (2021).
Elias, M. et al. The molecular basis of phosphate discrimination in arsenate-rich environments. Nature 491, 134–137 (2012).
Luecke, H. & Quiocho, F. A. High specificity of a phosphate transport protein determined by hydrogen bonds. Nature 347, 402–406 (1990).
Vyas, N. K., Vyas, M. N. & Quiocho, F. A. Crystal structure of M tuberculosis ABC phosphate transport receptor: specificity and charge compensation dominated by ion-dipole interactions. Structure 11, 765–774 (2003).
Wang, Z., Luecke, H., Yao, N. & Quiocho, F. A. A low energy short hydrogen bond in very high resolution structures of protein receptor-phosphate complexes. Nat. Struct. Biol. 4, 519–522 (1997).
Ledvina, P. S., Tsai, A. L., Wang, Z., Koehl, E. & Quiocho, F. A. Dominant role of local dipolar interactions in phosphate binding to a receptor cleft with an electronegative charge surface: equilibrium, kinetic, and crystallographic studies. Protein Sci. 7, 2550–2559 (1998).
Hirshberg, M. et al. Crystal structure of phosphate binding protein labeled with a coumarin fluorophore, a probe for inorganic phosphate. Biochemistry 37, 10381–10385 (1998).
Yao, N., Ledvina, P. S., Choudhary, A. & Quiocho, F. A. Modulation of a salt link does not affect binding of phosphate to its specific active transport receptor. Biochemistry 35, 2079–2085 (1996).
Wang, Z., Choudhary, A., Ledvina, P. S. & Quiocho, F. A. Fine tuning the specificity of the periplasmic phosphate transport receptor. Site-directed mutagenesis, ligand binding, and crystallographic studies. J. Biol. Chem. 269, 25091–25094 (1994).
George, N. L., Schilmiller, A. L. & Orlando, B. J. Conformational snapshots of the bacitracin sensing and resistance transporter BceAB. Proc. Natl. Acad. Sci. USA 119, e2123268119 (2022).
Gould, A. D., Telmer, P. G. & Shilton, B. H. Stimulation of the maltose transporter ATPase by unliganded maltose binding protein. Biochemistry 48, 8051–8061 (2009).
Sabrialabed, S., Yang, J. G., Yariv, E., Ben-Tal, N. & Lewinson, O. Substrate recognition and ATPase activity of the E. coli cysteine/cystine ABC transporter YecSC-FliY. J. Biol. Chem. 295, 5245–5256 (2020).
Gerber, S., Comellas-Bigler, M., Goetz, B. A. & Locher, K. P. Structural basis of trans-inhibition in a molybdate/tungstate ABC transporter. Science 321, 246–250 (2008).
Liu, F. et al. Structural basis of trehalose recycling by the ABC transporter LpqY-SugABC. Sci. Adv. 6, https://doi.org/10.1126/sciadv.abb9833 (2020).
Yang, X. et al. An oligopeptide permease, OppABCD, requires an iron-sulfur cluster domain for functionality. Nat. Struct. Mol. Biol. 31, 1072–1082 (2024).
Hu, T. et al. Molecular basis for substrate transport of Mycobacterium tuberculosis ABC importer DppABCD. Sci. Adv. 10, eadk8521 (2024).
Scheepers, G. H., Lycklama, A. N. J. A. & Poolman, B. An updated structural classification of substrate-binding proteins. FEBS Lett. 590, 4393–4401 (2016).
Oldham, M. L. & Chen, J. Crystal structure of the maltose transporter in a pretranslocation intermediate state. Science 332, 1202–1205 (2011).
He, Q. et al. Structural basis of phosphate export by human XPR1. Nat. Commun. 16, 683 (2025).
Webb, D. C., Rosenberg, H. & Cox, G. B. Mutational analysis of the Escherichia coli phosphate-specific transport system, a member of the traffic ATPase (or ABC) family of membrane transporters. A role for proline residues in transmembrane helices. J. Biol. Chem. 267, 24661–24668 (1992).
Cox, G. B., Webb, D. & Rosenberg, H. Specific amino acid residues in both the PstB and PstC proteins are required for phosphate transport by the Escherichia coli Pst system. J. Bacteriol. 171, 1531–1534 (1989).
Cox, G. B., Webb, D., Godovac-Zimmermann, J. & Rosenberg, H. Arg-220 of the PstA protein is required for phosphate transport through the phosphate-specific transport system in Escherichia coli but not for alkaline phosphatase repression. J. Bacteriol. 170, 2283–2286 (1988).
Chen, J. Molecular mechanism of the Escherichia coli maltose transporter. Curr. Opin. Struct. Biol. 23, 492–498 (2013).
Oldham, M. L., Chen, S. & Chen, J. Structural basis for substrate specificity in the Escherichia coli maltose transport system. Proc. Natl. Acad. Sci. USA 110, 18132–18137 (2013).
Chen, S., Oldham, M. L., Davidson, A. L. & Chen, J. Carbon catabolite repression of the maltose transporter revealed by X-ray crystallography. Nature 499, 364–368 (2013).
Oldham, M. L. & Chen, J. Snapshots of the maltose transporter during ATP hydrolysis. Proc. Natl. Acad. Sci. USA 108, 15152–15156 (2011).
Khare, D., Oldham, M. L., Orelle, C., Davidson, A. L. & Chen, J. Alternating access in maltose transporter mediated by rigid-body rotations. Mol. Cell 33, 528–536 (2009).
Oldham, M. L., Khare, D., Quiocho, F. A., Davidson, A. L. & Chen, J. Crystal structure of a catalytic intermediate of the maltose transporter. Nature 450, 515–521 (2007).
Hollenstein, K., Frei, D. C. & Locher, K. P. Structure of an ABC transporter in complex with its binding protein. Nature 446, 213–216 (2007).
Wei, Z. et al. Inhibition of miRNA-365-2-5p targeting SIRT1 regulates functions of keratinocytes to enhance wound healing. FASEB J. 39, e70560 (2025).
Wu, Y. T. et al. Peptide Cy (RL-QN15) accelerates hair regeneration in diabetic mice by binding to the frizzled-7 receptor. Zool. Res. 45, 1287–1299 (2024).
Li, C. et al. The direct binding of bioactive peptide andersonin-W1 to TLR4 expedites the healing of diabetic skin wounds. Cell. Mol. Biol. Lett. 29, 24 (2024).
Jia, Q. et al. Hydrogel loaded with peptide-containing nanocomplexes: symphonic cooperation of photothermal antimicrobial nanoparticles and prohealing peptides for the treatment of infected wounds. ACS Appl. Mater. Interfaces 16, 13422–13438 (2024).
Li, Y. et al. Peptide RL-QN15 regulates functions of epidermal stem cells to accelerate skin wound regeneration via the FZD8/β-catenin axis. Exploration https://doi.org/10.1002/EXP.20240090.
Wang, L. et al. Cyclic heptapeptide FZ1 acts as an integrin alphavbeta3 agonist to facilitate diabetic skin wound healing by enhancing angiogenesis. J. Med. Chem. 68, 19503–19520 (2025).
Ru, Z. Q. et al. Ultra-short cyclic peptide Cy (RL-QN15) acts as a TLR4 antagonist to expedite oral ulcer healing. Zool. Res. 46, 1187–1202 (2025).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: phenix.real_space_refine. Comput. Crystallogr. Newsl. 4, 43–44 (2013).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput. Biol. 8, e1002708 (2012).
Schrodinger, L. L. C. The PyMOL Molecular Graphics System, Version 1.8 (2015).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Li, Q. et al. A modified pCas/pTargetF system for CRISPR-Cas9-assisted genome editing in Escherichia coli. Acta Biochim. Biophys. Sin. 53, 620–627 (2021).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Pall, S. et al. Heterogeneous parallelization and acceleration of molecular dynamics simulations in GROMACS. J. Chem. Phys. 153, 134110 (2020).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101–014101 (2007).
Aoki, K. M. & Yonezawa, F. Constant-pressure molecular-dynamics simulations of the crystal-smectic transition in systems of soft parallel spherocylinders. Phys. Rev. A 46, 6541–6549 (1992).
Acknowledgements
Cryo-EM data were collected at the Cryo-EM Center of Southern University of Science and Technology. This work was supported in part by grants from the National Natural Science Foundation of China (32271012 and 32571403 to Q.C.; 32560219 to H.X.; 32100129 to S.L.; 32360138 to X.Y.; 32200998 to J.W.), Tianjin Fund for Distinguished Young Scholars (20JCJQJC00080 to Q.C.), Yunnan Fund for Distinguished Young Scholars (202401AV070004 to Q.C.; 202301AY070001-301 to X.Y.), the Xingdian Scholar Fund of Yunnan to Q.C., the Applied Basic Research Foundation of Yunnan Province (202401CF070179 to H.X.; 202301AS070036 to X.Y.), High-level Health Technology Talent in Yunnan Province (L-2025003 to X. Y.), the First-Class Discipline Team of Skin and Mucosal Regenerative Medicine of Kunming Medical University (2024XKTDTS10 to X.Y.), and Shenzhen Science and Technology Program (JCYJ20240813112508011 to S.L.).
Author information
Authors and Affiliations
Contributions
S.Liu, Q.C., X.Y., and D.L. conceived and supervised the project. H.X., X.J., and J.L. prepared the samples; S.Liu, S.Li, S.X., J.W., L.Z., and Q.C. performed data acquisition, image processing and structure determination; H.X. and Y.H. performed gene editing, ATPase assays and Pi import assays; R.Q., Q.C., and X.Y. performed molecular dynamic simulations; all authors participated in research design, data analysis, and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Türkan Haliloğlu and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiao, H., Li, S., Qi, R. et al. Molecular mechanism of phosphate import by the bacterial PstSCAB transporter. Nat Commun 17, 2294 (2026). https://doi.org/10.1038/s41467-026-69153-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69153-1