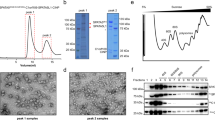
Abstract
R2SP belongs to the R2TP-like quaternary chaperone family and consists of RUVBL1/RUVBL2 AAA+ ATPases, which powers the machinery, and SPAG1 and PIH1D2 adapter proteins that engage specific clients to promote their quaternary assembly. However, little is known about the structure of R2SP and the precise mode of action of these R2TP-like complexes. Here, we combined biochemical (ATPase and fluorescence polarization assays) and structural approaches (NMR, structural mass spectrometry, cryo-EM) to investigate the 3D organization of the R2SP complex, its mode of assembly and ATPase activity. Together with our binding, mutational and kinetic studies, these results led us to propose a model in which SPAG1 and PIH1D2 bind and cooperatively engage RUVBL1/RUVBL2 to produce R2SP. This reveals a 3D structure close to the canonical R2TP complex but also highlights differences in RUVBL1/RUVBL2 ATPase activity, as well as the cooperative binding of SPAG1 and PIH1D2 to this catalytic core that may explain functional difference between the two systems.
Similar content being viewed by others
Introduction
Many proteins across all domains of life function as part of multimeric complexes that require precise and regulated quaternary assembly. In mammals, the assembly of these quaternary structures is supported by a multisubunit chaperone complex composed of an R2TP core, which includes two AAA+ ATPases RUVBL1 and RUVBL2, RNA polymerase II-associated protein 3 (RPAP3), and Protein Interacting with HSP90 1 domain-containing protein 1 (PIH1D1)1,2,3,4,5,6. RUVBL1/RUVBL2 are the catalytic core of the R2TP complex, while RPAP3 and PIH1D1 act as flexible scaffolding and regulatory units, respectively. R2TP complexes interact with the HSP90 chaperone as well as a Prefoldin-like (PFDL) module to form the so-called PAQosome7,8,9,10,11, which ensures proper assembly of both protein (RNA Polymerases, PIKKs, …)1,8 or ribonucleoprotein (snoRNPs, U4/U5 snRNPs, …)2,3,4,5 complexes. The existence of an endogenous R2T complex composed of RUVBL1, RUVBL2, RPAP3 and devoid of PIH1D1 has been observed both in humans, owing to RPAP3 isoform 2 that does not bind PIH1D112,13 and very recently in Arabidopsis thaliana, whose genome does not include PIH1D114.
In addition to R2TP, human cells also express a variant of the canonical R2TP complex termed R2SP, where two protein adapters, SPAG1 (sperm-associated antigen 1), and PIH1D2 (Protein Interacting with HSP90 1 domain-containing protein 2) replace RPAP3 and PIH1D1 respectively. R2SP is responsible for the quaternary assembly of distinct set of clients involved in motile cilia biogenesis, cell adhesion, cell migration, and the organization of the synaptic zone15. The null mutants of SPAG1 causing Primary Cilia Dyskinesia16. R2TP and R2SP target different clients. Furthermore, whereas R2TP is ubiquitous, R2SP exhibit tissue-specific expression such as in the biogenesis of motile cilia, for the quaternary assembly of axonemal dynein arms complexes. In this process, the R2SP functions with the help of dynein axonemal assembly factors16,17,18, whose mutations result in ciliary mobility defects associated with various diseases such as respiratory infections, sinusitis, otitis, or male fertility loss19,20. Interaction probing also suggests that the R2SP is involved in a variety of other nuclear and cytoplasmic functions15. Therefore, given that R2TP and R2SP share the same RUVBL1/RUVBL2 catalytic core, the difference in client preference and function are likely due to the differences in the protein adapters (RPAP3 and PIH1D1 vs SPAG1 and PIH1D2, respectively), and how these adapters affect R2SP complex, its mode of assembly and ATPase activity. However, many of these mechanistic details are currently missing due to the lack of structural understanding of R2SP.
Previous studies showed that within R2TP-like complexes, RUVBL1/RUVBL2 exists in an equilibrium between single and double hetero-hexameric rings (i.e. hexamers and dodecamers) depending on the nucleotide load and the nature of their partners21. Each protomer comprises three domains. Domain I (DI) and domain III (DIII) form the ATPase-core ring of RUVBL1/RUVBL2 responsible for the ring-like oligomerization, whereas flexible domain II (DII) protrudes from the ATPase ring and: (i) recruits cofactors/clients via an external region (DIIext)22,23,24,25; (ii) establishes a structural and conformational link between clients and the ATPase core via its internal region (DIIint)24,26; (iii) interacts with nucleotides through the DIIext region27; and (iv) interacts with DII’s from other hexameric rings, forming DII-stacked hexameric rings (dodecamers), which may represent an inhibited/storage form of the complex28. RUVBL1/RUVBL2 hetero-hexamers bind and hydrolyze nucleotides, which is the basis of their chaperoning activity, the N-terminal parts of RUVBL1/RUVBL2 interact with both their own DIIext domains and the nucleotide, and has been proposed to block the exchange of ADP to ATP24,26. Furthermore, these N-terminal regions become unstructured in the absence of a nucleotide, suggesting that nucleotide binding exerts a major influence over RUVBL1/RUVBL2 and how they recruit their partners; however, these questions remain a matter of debate13,24,29. In this context, although available 3D structures of human24,30 and yeast31,32 R2TP, and plant14 R2T complexes helped depict the spatial organization of the complex, the high-resolution structure for the full-length R2TP complex is still missing, most likely because of its high degree of flexibility and dynamic nature.
Here, we show a combination of biochemical, biophysical and structural approaches to characterize the recently identified human R2SP complex. Our results reveal that the C-terminal domain of SPAG1 binds the RUVBL1/RUVBL2 ring in the same manner as RPAP3. However, we also show significant differences in the binding mode and activity of R2SP compared to the canonical R2TP complex which hints towards its specific mode of assembly and action.
Results
SPAG1:PIH1D2 binds cooperatively to RUVBL1/RUVBL2 to form R2SP
To investigate structure and function of the human R2SP complex, we expressed all its components in E. coli, separately or together, and used truncated versions when full-length (FL) proteins proved insoluble. In those cases, specific truncations were informed based on the existing literature as follows. The C-terminal amino-acid sequences of RPAP3 and SPAG1 share 25% identity and 35% similarity scores, suggesting a conserved 3D fold for this domain and thus, putative conserved functions33. We designed a soluble fragment of SPAG1 comprising residues 796 to 926 (SPAG1_C), corresponding to RPAP3 C-terminal domain (RPAP3_C) used for NMR structure determination15, and a longer C-terminal soluble fragment encompassing residues 622 to 926, which additionally included the TPR3 and SPAG1_C domains (SPAG1622-926) (Fig. 1). In our hands, FL SPAG1 can be stably over-expressed and was shown to bind the chaperones HSP70 and 90 in vitro33, however its ability to bind RUVBL1/RUVBL2 proteins alone or in complex with PIH1D2 was not yet described.
A Domain organization of R2TP and R2SP components and their comparison: RUVBL1 and RUVBL2 – AAA + ATPases present in both R2TP and R2SP; RPAP3 (the hatched area corresponds to isoform 1) and SPAG1 – two homologous adapters featuring N-terminal tetratricopeptide repeat (TPR) domains (TPR1 and 2 in RPAP3) and (TPR1-3 in SPAG1), PIH1Dx-interacting domain (PRD), and the C-terminal RUVBL1/RUVBL2 binding domain (RBD); and PIH1D1 and PIH1D2 – two homologous adapter proteins featuring N-terminal Protein Interacting with HSP90 (PIH) domain, and C-terminal CS (fCHORD-containing proteins and Sgt1 domain) domain. B AlphaFold3 (AF3) predicted structures of PIH1D2 (left panel) and SPAG1622-926 (right panel) protein structures.
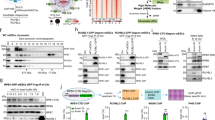
We first investigated interactions between different components of the R2SP complex in vitro using co-expression and co-immunoprecipitation assays, and compared it with R2TP. We observed that RUVBL1/RUVBL2 complex co-purified with all SPAG1 constructs (i.e., FL SPAG1, SPAG1622-926, and SPAG1_C; Supplementary Fig. 1A–C). This indicates that SPAG1_C was sufficient to bind the RUVBL1/RUVBL2 complex, similarly to the RPAP3_C domain of RPAP3 (Supplementary Fig. 1A, C; controls in Supplementary Fig. 2A). In addition, NMR assays demonstrated that the presence of RUVBL1/RUVBL2 affected the 13C-METHYL-SOFAST spectrum of SPAG1796-926 regardless of the presence of a nucleotide (Apo, ADP, or ATP) (Supplementary Fig. 3), indicating that this interaction was nucleotide-independent.
Co-expression of PIH1D1 or PIH1D2 with RUVBL1/RUVBL2 led to moderate co-elution with different band intensity profiles (Supplementary Fig. 1D) with the majority of PIH1D2 observed to be insoluble (Supplementary Fig. 1D). However, the remaining soluble PIH1D2 (found in the supernatant and elution fractions) appeared to be intact and able to interact with RUVBL1/RUVBL2, similar to PIH1D1 (Supplementary Fig. 1D). PIH1D1 and PIH1D2 interactions with RUVBL1/RUVBL2 became more intense with the addition of RPAP3 and SPAG1, respectively, with a more pronounced effect seen for R2SP (Supplementary Fig. 1E). This suggests that PIH1D2 might require SPAG1 for efficient integration into the R2SP complex. Finally, we showed that the SPAG1 fragment 742-791 (downstream of the SPAG1’s TPR3 domain) could form an R2SP complex (Supplementary Fig. 1F), possibly by interacting with the CS domain of PIH1D2, similar to the binding of RPAP3396-455 to PIH1D112.
To measure interaction affinities (KD) and the kinetics (kon and koff) of binding within the R2SP complex, we performed Surface Plasmon Resonance (SPR) assays between different SPAG1:PIH1D2 sub-complexes (i.e. SPAG1622-926:PIH1D2 and SPAG1622-926:PIH1D2231-315; Fig.1), and different RUVBL1/RUVBL2 variants (i.e. wild type ADP loaded RUVBL1/RUVBL2; catalytically dead mutant ATP loaded RUVBL1D302N/RUVBL2D299N34,35; and DIIext domain deletion mutants ADP loaded \({\rm{RUVBL1}}^{\Delta {\rm{D}}_{{\rm{IIext}}}}/{\rm{RUVBL2}}^{\Delta {\rm{D}}_{{\rm{IIext}}}}\)) (Fig. 2 and Supplementary Table 1). The experiments demonstrated that the RUVBL1/RUVBL2 complex had nanomolar affinity for SPAG1622-926:FL-PIH1D2 (KD = 5.3 ± 0.4 nM) and SPAG1622-926:PIH1D2231-315 (KD = 22 ± 1.1 nM), with resolved kinetics that allowed the determination of kon and koff (Fig. 2 and Supplementary Table 1). The catalytically dead RUVBL1D302N/RUVBL2D299N complex, which binds but doesn’t hydrolyze ATP, exhibited similar kinetic profile of binding to SPAG1622-926:PIH1D2231-315 as the wild type (Supplementary Fig. 4A); however, in the presence of FL PIH1D2 the KD increased to the micromolar range for the RUVBL1D302N/RUVBL2D299N complex (Supplementary Fig. 4C). A similar effect was observed with the dodecameric RUVBL1/RUVBL2 complex with truncated DIIext, which displayed only low-affinity and transient interactions with SPAG1622-926:FL-PIH1D2 (Supplementary Fig. 4B and 4D, Supplementary Table 1). Note that our SPAG1622-926 construct features an 8-residue C-terminal tag that does not appear to prevent complex formation, although we cannot exclude the possibility that it affects the kinetics. To further deconvolute interactions within R2SP and unveil its assembly mechanism, we tested individual SPAG1 and PIH1D2 proteins against the RUVBL1/RUVBL2 complex. By SPR, SPAG1622-926 alone interacted very transiently with the RUVBL1/RUVBL2 complex, with KDss of 1.6 ± 0.2 µM (Supplementary Fig. 4E, F and Supplementary Table 1), supporting previous IP LUMIER assays performed in vivo15. However, we could not measure KD values for SPAG1, SPAG1796-926, and PIH1D2 alone as they failed to form functional active surfaces during the SPR assay development, even when using the Extract2Chip method36.
Colored curves represent recorded data and are overlaid with a 1:1 Langmuir binding model kinetic fit (black). The Top concentration and following injections are related by a 2-fold dilution. The KD is taken as the ratio of the dissociation rate constant (kd) divided by the association rate constant (ka). Experiment was performed in absence of any nucleotide in the running buffer.
Taken together, our data suggest that both SPAG1 and RPAP3 interact directly with the RUVBL1/RUVBL2 complex but with different affinities (milli- vs nano-molar, respectively). Moreover, SPAG1 interaction with the RUVBL1/RUVBL2 complex is stronger in the presence of PIH1D2, suggesting distinct cooperativity models for the assembly of R2SP and R2TP. Our results also suggest that DII domains of RUVBL1 and RUVBL2 play a key role in binding SPAG1:PIH1D2, which is further modulated by the nucleotide molecule present in the active pocket. These results are in agreement with previous structural studies of RUVBL1/RUVBL2 showing that ATP binding induces “closed conformation” where DII domains moves towards the active site (closed conformation)24,26, resulting in a conformation that is likely incompatible with FL PIH1D2 binding but not PIH1D2231-315 (i.e. the CS domain).
R2TP and R2SP exhibit different ATP binding and ATPase activity
Next, we compared the effect of SPAG1:PIH1D2 and RPAP3:PIH1D1 on the ATPase activity of RUVBL1/RUVBL2. Using steady state kinetics, we demonstrated that both RPAP3:PIH1D1 and SPAG1:PIH1D2 stimulate RUVBL1/RUVBL2 ATPase activity in a concentration-dependent and saturable manner, allowing the measurement of AC50 (concentration required for half-maximum activation; Fig. 3A and B). RPAP3:PIH1D1 stimulated the ATPase activity of RUVBL1/RUVBL2 by 1.2-fold with an AC50 of 20 nM, in agreement with previous reports13, while SPAG1:PIH1D2 stimulated RUVBL1/RUVBL2 by 2-fold with an AC50 of 700 nM, indicating some differences between the two systems.
RPAP3:PIH1D1 (A) and SPAG1:PIH1D2 (B) concentration-dependent steady state ATPase activity of RUVBL1/RUVBL2 was measured by a coupled assay. Data points (black dots) represent the means ± SD of three measurements at 37 °C. The solid lines through data points represent the best rectangular hyperbola fit (orange for RPAP3:PIH1D1 and purple for SPAG1:PIH1D2). C–H Kinetics of association and dissociation of fluorescent ATPγS on RUVBL1/RUVBL2 (blue), R2TP (orange) and R2SP (purple) was measured by a fluorescence polarization assay. Both showed a bi-exponential behavior (Supplementary Fig. 5). Bars represent mean values and error bars indicate the standard deviation (SD) calculated from n = 3 or 5 independent technical replicates. Statistical significance was determined using a two-sided Student’s t test (ns, p ≥ 0.05; *, p< 0.05; **, p< 0.01; ***, p< 0.001). Apparent association constants kobs1 (C) and kobs2 (D) were extracted from best bi-exponential fits determined for RUVBL1/RUVBL2, R2TP and R2SP (n = 3). For panel (C, D) comparisons yielded the following exact p-values: RUVBL1/RUVBL2 vs R2TP (p = 5.93 × 10−4 and p = 0.0185), RUVBL1/RUVBL2 vs R2SP (p = 2.48 × 10−4 and p = 8.14 × 10−3), and R2TP vs R2SP (p = 6.98 × 10−4 and p = 0.0367). A similar experiment was repeated on samples where bound ADP nucleotides were removed prior by alkaline phosphatase treatment and apparent association constants kobs1 (E) and kobs2 (F) with the same color code were determined (n = 5). For panel E-F, comparisons yielded the following exact p values: RUVBL1/RUVBL2 vs R2TP (p = 0.0101 and p = 0.468), RUVBL1/RUVBL2 vs R2SP (p = 0.0101 and p = 0.444), and R2TP vs R2SP (p = 0.955 and p = 0.967). Finally, the nucleotide dissociation constants koff1 (G) and koff2 (H) were determined from best bi-exponential decay fits for RUVBL1/RUVBL2, R2TP and R2SP (n = 5). For panel G-H, comparisons yielded the following exact p values: RUVBL1/RUVBL2 vs R2TP (p = 6.73 × 10−4 and p = 0.0234), RUVBL1/RUVBL2 vs R2SP (p = 0.0138 and p = 2.70 × 10−2), and R2TP vs R2SP (p = 0.0133 and p = 0.0113).
We also examined how SPAG1:PIH1D2 and RPAP3:PIH1D1 affect nucleotide binding. Using BODIPY-ATPγS (BDP-ATP), a fluorescent-labeled non-hydrolysable ATP analog, and fluorescent polarization measurement, we followed its binding and dissociation kinetics from RUVBL1/RUVBL2. We observed that nucleotide binding by RUVBL1/RUVBL2 followed a bi-exponential regime suggesting that the RUVBL1/RUVBL2 hexamer possesses two nucleotide binding sites with distinct association and dissociation rates, both differing by about 10-fold in magnitude and likely corresponding to the active sites of RUVBL1 and RUVBL2 (Supplementary Fig. 5). We named the two apparent association rate constants kobs1 and kobs2, and the two dissociation rate constants koff1 and koff2, respectively. To characterize the effect of two co-chaperones on nucleotide binding, we first evaluated nucleotide association after the addition of either RPAP3:PIH1D1 or SPAG1:PIH1D2 (Fig. 3C and D). We observed that both apparent association rates increase upon the addition of the co-chaperones, however RPAP3:PIH1D1 was 2 to 3-times more efficient in accelerating the association rates than SPAG1:PIH1D2. Because our recombinant RUVBL1/RUVBL2 preparation co-purified with endogenous E. coli ADP, we checked if the kobs would change if we first depleted nucleotides from RUVBL1/RUVBL2. Indeed, we observed that the apparent nucleotide binding rates of RUVBL1/RUVBL2 were greatly accelerated in the absence of pre-bound ADP (Fig. 3E, F). Interestingly, adding SPAG1:PIH1D2 had no effect on the apparent binding rates but did accelerate nucleotide release (Fig. 3G, H).
Together, these observations suggest that RPAP3:PIH1D1 and SPAG1:PIH1D2 are more likely to facilitate nucleotide dissociation rather than nucleotide association. Although our experiments did not allow us to pinpoint which RUVBL has the slow or the rapid dissociating site, we observed that RPAP3:PIH1D1 was more potent in increasing koff1 of the rapidly dissociating site, while SPAG1:PIH1D2 was a better activator of the slow dissociation site and increased koff2. These differences could be related to the different binding mechanisms of RPAP3:PIH1D1 and SPAG1:PIH1D2 to the RUVBL1/RUVBL2 AAA + ring, as already gleaned from the results of the SPR experiments (this work and ref. 15).
The C-terminal domain of SPAG1 and RPAP3 display similar 3D folds
To get further insight into the R2SP 3D organization and structurally characterize the putative RUVBL1/RUVBL2 binding domain (RBD) of SPAG1 (Fig. 1), we determined the NMR structure of SPAG1796-926 in solution. We obtained a well-resolved ensemble for which statistics are described in Supplementary Table 2. Briefly, SPAG1796-926 is composed of 8 α-helices that pack together to form a globular domain (Fig. 4A). When compared to the NMR structure of RPAP3’s RBD (RPAP3535-665, Fig. 1), SPAG1796-926 displays a similar overall fold and features, including a break at the end of helix α4, which seems to represent a strong signature of RBD 3D fold. The main distinguishing feature is the presence of an extended loop between helices α7-α8 that covers helices α4 and α5 in SPAG1796-926 (covering loop, Fig. 4B). Further comparison between RPAP3535-665 and SPAG1796-926 showed an RMSD value of ~ 2.0 Å for Cα atoms in secondary structures. This suggests that SPAG1796-926 may not be able to perfectly fit the binding site of RPAP3535-665 on RUVBL1/RUVBL2, even though the positioning of the interfacing residues in RPAP3535-665 appeared conserved in SPAG1796-926. This could explain why the affinity of RUVBL1/RUVBL2 for SPAG1622-926 which contains the SPAG1796-926 region (Supplementary Fig. 4), differ from the one of RPAP3535-66515.
A Backbone representation of the 20 best NMR structures (α-helices indicated in green) of SPAG1_C. B Cartoon representation of the superimposition of the NMR structures of the C-terminal domain of human SPAG1 (SPAG1_C, green) and RPAP3 (RPAP3_C, blue) proteins. The break in the helix α4, common to SPAG1 and RPAP3 is indicated, as well as the covering loop specific of SPAG1. C Slices extracted from the 1H-15N NOESY-HSQC spectrum recorded on a 2H/13C/15N-labeled PIH1D2_CS:SPAG1622-762 complex. NOE cross-peak linking amide protons belonging to individual spin systems are highlighted using dash blue lines. They reveal a spatial proximity between residues whose label is indicated on the top of the slice (green and orange labels for SPAG1622-762, and PIH1D2’s CS domain (P2_CS), respectively). D Superimposition of the five AF3 predicted models of the PIH1D2_CS:SPAG1622-762 complex structure. In the upper panel, five predicted structures were aligned on the one PIH1D2_CS (orange). The TPR3 domains of five SPAG1 predicted structures were shown in different colors (TPR3_1 to TPR3_5), indicating that relative position of SPAG1 TPR3 with respect to PIH1D2_CS was poorly predicted by AF3. The region of the complex with the highest convergence rate is highlighted in a circle and includes PIH1D2_CS (orange) and SPAG1752-762 region of all five SPAG1622-762 predicted structures. The lower panels shown a zoom in of the high convergence region, highlighting residues at the interface, with dashed lines indicating experimentally observed NOE data from Fig. 4C. The fragment 752-762 of SPAG1 is shown in green, PIH1D2_CS is shown in orange. E Structures of the PIH1D2_CS, PIH1D1_CS, and yeast Pih1_CS bound to SPAG1, RPAP3 and yeast Tah1, respectively. The structures come from AF3 (this work), X-ray (6GXZ.pdb) and NMR (2MNJ.pdb), respectively. The limits of each protein fragment are indicated.
Validating the SPAG1:PIH1D2 interface using NMR and XL-MS constraints
Next, we used NMR and cross-linking mass spectrometry (XL-MS) to characterize interactions between PIH1D2 and SPAG1 at the atomic level. We successfully produced a 2H/13C/15N-labeled complex between the putative CS domain of PIH1D2 (PIH1D2231-315 or PIH1D2_CS) and an extended version of SPAG1’s TPR3 domain (SPAG1622-762) (Fig. 1), and used NMR to analyze the SPAG1622-762:PIH1D2_CS structure. Although we were not able to obtain assignments for all backbone resonances, which prevents the full calculation of the 3D structure (Supplementary Fig. 6A), we did observe experimental NOE cross-peaks between the HN nuclei of R754, I756, Q759, and V761 in SPAG1 and the HN nuclei of V269, L267, A299/K300/F301, and F301/K303 in PIH1D2_CS respectively (Fig. 4C). The strength of the recorded NOEs and their unambiguous assignments indicate that SPAG1 makes extensive contacts with PIH1D2-CS. To visualize the possible protein-protein interface, we used AlphaFold3 (AF3) to build a model of PIH1D2_CS:SPAG1622-762. The predicted structures produced an interaction mode between the region 752-762 in SPAG1 and PIH1D2_CS (Fig. 4D), in very good agreement with intermolecular NMR distances (Fig. 4C). Furthermore, AF3 predicted structures revealed the characteristic intermolecular β-sheet, which was previously observed in structures of RPAP3:PIH1D1 and Tah1:Pih1 (yeast ortholog) complexes (Fig. 4E). Interestingly, the positioning of the TPR core in SPAG1622-762 did not converge toward a stable position on PIH1D2_CS in AF3 predicted structures (Fig. 4D). This is in line with the computed chemical shift variations between free SPAG1622-742 and PIH1D2_CS-bound SPAG1622-742 which do not reveal significant perturbations all over the region 622-738 in SPAG1 (Supplementary Fig. 6B, C), suggesting that SPAG1 TPR core does not contribute to PIH1D2_CS binding. Therefore, our NMR data and AF3 structure predictions allow us to define residues 752-762 as the minimal PIH1D2-binding sequence in SPAG1.
In parallel, we performed XL-MS analysis of SPAG1622-926:FL-PIH1D2 heterodimer using DSBU (DiSuccinimidyl diButyric Urea), a cross-linking agent that creates bonds between lysine residues with Cα-Cα distances between 5 and 32 Å (Supplementary Fig. 7). Overall, we identified 58 unique cross-links, 27 intra-SPAG1622_926, 10 intra-PIH1D2 and 21 inter-SPAG1622-926:PIH1D2 (Supplementary Table 3 and Supplementary Fig. 7). Among those, 7 inter-SPAG1622-926:PIH1D2 cross-links comprised the SPAG1622_926_TPR3 domain involved in HSP70/90 binding and 11 inter-SPAG1622-926:PIH1D2 cross-links were in the phospho-peptide binding pocket of the PIH domain (Supplementary Fig. 7). The data suggest that, although not engaging PIH1D2_CS (see above), SPAG1622_926 TPR core binds the PIH domain in PIH1D2. This was also previously seen for the TPR1 domain of RPAP3 and the PIH domain of PIH1D112. In further agreement with the NMR and AF3 predictions, no cross-links were noticed between the TPR core in SPAG1622-926 and PIH1D2’s CS domain; nonetheless, several inter-SPAG1622-926:PIH1D2 cross-links involved residues K741 from SPAG1622-926 (close to the minimal PIH1D2 binding region) and K290 from PIH1D2 (in its CS domain) were observed, thus corroborating the SPAG1622-926:PIH1D2 binding interface observed by NMR.
Up to three SPAG1622-926:PIH1D2 heterodimers interact with one RUVBL1/RUVBL2 heterohexamer
To extend our structural analysis to the entire R2SP complex, we assembled a truncated version of the R2SP complex featuring SPAG1622-926:PIH1D2 bound to RUVBL1/RUVBL2 in the presence of ADP because the FL SPAG1 suffers proteolysis (Supplementary Fig. 8); we refer to this as the R2S622-926P complex. To assess the stoichiometry of the R2S622-926P complex, we used native mass photometry (nMP) and native mass spectrometry (nMS). While nMP profiles obtained for RUVBL1/RUVBL2 alone revealed the coexistence of hexameric and dodecameric assemblies (Supplementary Fig. 9), the R2S622-926P nMP histograms revealed a much more heterogeneous distribution, with more hexameric species compared to dodecameric ones (73% vs. 27% respectively) (Fig. 5A, B). Similarly, comparison of nMS spectra obtained for RUVBL1/RUVBL2 and R2S622-926P also demonstrated that SPAG1622-926:PIH1D2 binding disrupted the RUVBL1/RUVBL2 dodecamer to form a stable hexameric R2S622-926P complex (Supplementary Fig. 9B and Fig. 5C), similar to what was seen upon binding of other RUVBL1/RUVBL2 partners (e.g., DPCD or RPAP323,24). Regarding the hexameric species, both approaches revealed the coexistence of undecorated RUVBL1/RUVBL2 hexamers (313,503 ± 8 Da, 20%), or carrying one (384,692 ± 13 Da, 35%), or two (456,039 ± 22 Da, 18%) SPAG1622-926:PIH1D2 units, each containing an average of 6 ADP molecules (Fig. 5C). We also observed a fourth very low-abundance population (< 5%) that corresponds to a RUVBL1/RUVBL2 hexamer with three SPAG1622-926:PIH1D2, however, due to the low abundance we were not able to assign an accurate mass to this assembly.
Overall, our data indicate that SPAG1622-926:PIH1D2 binding shifts RUVBL1/RUVBL2 hexamer/dodecamer equilibrium towards the hexameric assembly, resulting in formation of R2SP complexes where RUVBL1/RUVBL2 hexamers engage one, two or three SPAG1622-926:PIH1D2 heterodimers in that order of preference. These stoichiometries are similar to those seen within the R2TP complex24.
A Mass Photometry measurements in triplicates, shown in shades of grey, at a droplet concentration of 50 nM. MP shows that RUVBL1/RUVBL2 hexamers co-exist with RUVBL1/RUVBL2:SPAG1(S1)622-926:PIH1D2(P2) and RUVBL1/RUVBL2:(S1622-926:P2)2, and that dodecameric RUVBL1/RUVBL2 is also formed. Standard deviations come from measurement replicates (n = 3). The low mass range (30–100 kDa) exhibits a peak that may correspond to free RUVBL1 or RUVBL2 monomers, with close molecular weights of 51.7 and 51.9 kDa, respectively. In the 100–550 kDa mass range, three populations are identified and quantified: undecorated RUVBL1/RUVBL2 hexamers (species a, 311.0 ± 2.0 kDa, 20 ± 1%), or RUVBL1/RUVBL2 with one (species b, 373.0 ± 2.6 kDa, 35 ± 1%) and two (species c, 438.3 ± 7.6 kDa, 18 ± 3%) S1622-926:P2 bound. Above 550 kDa, populations of RUVBL1/RUVBL2 dodecamers (species d, 620.6 ± 7.8 kDa, 10 ± 1%) with one (species e, 677.0 ± 4.4 kDa, 13 ± 4%), or two (species f, 739.0 ± 14.1 kDa, 5 ± 4%) S1622-926:P2 bound are detected. * represents low abundance RUVBL1/RUVBL2:(S1622-926:P2)3 species (< 5%). Results are shown as an average ± SD (n = 3, measurement replicates). B Relative abundances of the different stoichiometry, in % of total counts, measured in MP. Standard deviations come from measurements replicates (n = 3). C native MS confirmed the presence of R2S622-926P complexes with different stoichiometries ranging from RUVBL1/RUVBL2 hexamers bound to one (384,692 ± 13 Da) or two S1622-926:P2 (456,039 ± 22 Da), each containing in average 6 ADP molecules.
R2S622-926P and R2TP complexes share similar 3D architecture
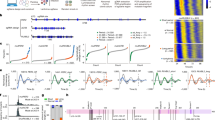
We next investigated the 3D structures of R2S622-926P using cryo-electron microscopy (cryo-EM) and single particle analysis (SPA). Initial 2D and 3D classification rounds using RELION37, revealed the coexistence of dodecameric and hexameric assemblies (Supplementary Fig. 10), in agreement with MS characterization. The image processing scheme allowed us to reconstruct a 3.5 Å map of the dodecameric assembly formed by two toe-to-toe RUVBL1/RUVBL2 hexamers interacting via their DII domains (Supplementary Fig. 11) in a skewed manner, as already observed by other groups for full-length RUVBL1/RUVBL2 assemblies23,30. Conformational heterogeneity studies of the RUVBL1/RUVBL2 dodecamer using cryoDRGN38 showed movements of one hexamer relative to the other (Supplementary Movie 1), which likely reflects the structural dynamics between the two hexamers as previously suggested39. In line with this observation, local resolution of the consensus cryo-EM map ranges from ~ 3 Å for the inner AAA + core ring, to 6-7 Å for the DII domains (Supplementary Fig. 10), suggesting that the latter is very flexible. Of note, 2D classes as well as low-pass filtered cryo-EM maps of dodecameric assemblies reveal horn-like densities crowning the AAA + core of RUVBL1/RUVBL2 (Supplementary Fig. 12). Although these densities were not reconstructed in the high-resolution cryo-EM map, we hypothesize that they correspond to the dynamic binding of SPAG1622-926, as hinted by our nMP and nMS data, as well as further cryo-EM analyzes of hexameric particles described below.
The global resolution of the R2S622-926P hexameric particle reconstruction was 3.6 Å (Supplementary Fig. 13). The map revealed an AAA+ hexameric ring corresponding to the DI-DIII domains of RUVBL1/RUVBL2, with local resolutions ranging between 3 and 4 Å from the central pore outward, as assessed with ResMap40 (Supplementary Fig. 13). This ring was topped by at least one “horn” feature, with local resolutions between 3.5 and 5 Å from its base to its apex, while the laterally positioned, structurally mobile DIIext domains of RUVBL1/RUVBL2 were less well resolved (5–7 Å). We used this cryo-EM map (Fig. 6A) together with the NMR structure of the C-terminal domain of SPAG1 (SPAG1_C) to build an atomic model of the RUVBL1/RUVBL2:SPAG1_C complex (Fig. 6B). The “horn” atop the DI-DIII hexameric ring could be assigned to the SPAG1 region spanning amino acids 800–926. As in its RPAP3 paralog, this domain bridges one RUVBL1 and one RUVBL2 subunit (Fig. 6A, B). Because the DIIext domains of RUVBL1 were poorly resolved, their X-ray structures were fitted into the corresponding densities by rigid-body docking; nevertheless, they could be unambiguously placed in an open conformation. Notably, the RUVBL2 DIIext domain engaged by SPAG1_C on the DI-DIII ring appeared even more flexible than the others.
A Top (left panel) and side (right panel) surface views of the cryo-EM map of hexameric complexes (composed of RUVBL1/RUVBL2, SPAG1622-926 and PIH1D2) determined at a global resolution of 3.6 Å. Local resolutions are shown in Supplementary Fig. 13. B Corresponding views of the atomic model derived from the cryo-EM map shown in A. RUVBL1 (R1) and RUVBL2 (R2) are shown in pink and blue, respectively. SPAG1’s C-terminal domain (SPAG1_C) is shown in lime green. C, D Details of the cryo-EM map (grey mesh) and reconstructed atomic model of the contact zone between SPAG1_C and RUVBL2 (C) and of the ADP moieties in the RUVBL1/RUVBL2 hexamer (D). Color coding is the same as panels A, B.
Inspection of the upper ring/horn region of the RUVBL1/RUVBL2:SPAG1_C cryo-EM map shows that SPAG1 residue R881 lies within < 4 Å of RUVBL2 residues R392, T397, Y430, D435, and R438, while SPAG1 residue M884 contacts at least RUVBL2 residues E396 and I393 (Fig. 6C). These interactions were supported by point-mutation analyzes and XL-MS, as described below. Moreover, and consistent with nMS data, six ADP molecules could be unambiguously fitted into the cryo-EM density (Fig. 6D). A density located above the DI-DIII domains of the RUVBL1/RUVBL2 hexameric ring was attributed to the partial presence of a second SPAG1 molecule (Fig. 6A, B), again in agreement with nMS results. To further analyze the stoichiometry of SPAG1 on RUVBL1/RUVBL2 particles, we performed focused classification of the hexameric particle set (Supplementary Supplementary Fig. 13). This analysis showed that the RUVBL1/RUVBL2 heterohexamer can be crowned by either one or two SPAG1_C domains (Supplementary Supplementary Fig. 14), with particle proportions of 63% and 37%, respectively. In parallel, we used cryoDRGN38 to sort out structural heterogeneity in the hexameric dataset using AI-based approaches. This analysis revealed RUVBL1/RUVBL2 heterohexamers capped by one, two, or even three SPAG1 C-terminal domains (Supplementary Movie 2, PC1 analysis), consistent with nMP and nMS observations.
Low-pass filtered 3D classes (Supplementary Fig. 14B) as well as 2D classes (Supplementary Fig. 14C) of hexameric R2S622-926P particles always revealed fuzzy densities beneath the RUVBL1/RUVBL2 DII domains, at the center of the inner cavity of the RUVBL1/RUVBL2 ring (Supplementary Fig. 14B and Supplementary Movie 2, PC1 analysis). Despite extensive attempts, these densities could not be refined to high resolution. However, they were clearly detected in a cryoDRGN analysis of the binned cryo-EM images (Supplementary Movie 2, PC1 analysis). By homology with the R2TP complex24 and supported by our XL data described below, we propose that these densities correspond to PIH1D2.
Based on these results, we next tested different surface interaction mutants for the association of RUVBL2 with SPAG1622-926, focusing on RUVBL2 residue D435 and SPAG1 residues R881A/M884A. These SPAG1 mutations correspond to residues R623 and M626 in RPAP3, which are essential for RPAP3-RUVBL1/RUVBL2 binding15. Using both co-expression in E. coli (Supplementary Fig. 15) and LUMIER co-IPs in mammalian cells (Supplementary Fig. 16), we found that the D435 of RUVBL2 is essential for interaction with both SPAG1 and RPAP3. Co-expression assays showed that the RUVBL1/RUVBL2 D435A mutant still interacted with RPAP3_C or full-length RPAP3 (Supplementary Fig. 15B, D), whereas the SPAG1 mutants lost binding to RUVBL1/RUVBL2 (Supplementary Fig. 15E). Notably, both approaches revealed that addition of PIH1D2 restored the interaction between mutant RUVBL2 and SPAG1, an effect not fully reproduced by PIH1D1 (Supplementary Figs. 15F, G, 16B). This finding is consistent with our SPR results, where FL PIH1D2 ameliorated the binding between SPAG1 and RUVBL1/RUVBL2 (Supplementary Table 1). Together, these analyzes underscore a key interplay between SPAG1 and PIH1D2 during assembly of the R2SP complex, which is different from what is seen for the R2TP complex.
XL-MS reveals the importance of RUVBL1/RUVBL2’s DII domains in SPAG1 and PIH1D2 interaction
To complement our cryo-EM data, we used XL-MS to study in-solution interactions formed within the R2S622-926P complex. Using DSBU as a cross-linker (5-32 Å range for Cα-Cα) allowed us to identify a total of 208 unique cross-links (99 intra- and 109 inter-protein interactions, Fig. 7A and Supplementary Table 4). We detected direct interactions between SPAG1 and both RUVBL1 and RUVBL2, as well as interactions between PIH1D2 and both RUVBL1 and RUVBL2. Almost 37% (40 peptide pairs) of the unique inter-XLs involved PIH1D2 and either RUVBL1 (26) or RUVBL2 (14). Half of them interacted via the DII domains of RUVBL1/RUVBL2, and within these, 14 involved the PIH domain of PIH1D2. Thus, the XL-MS data supports that PIH1D2 interacts with RUVBL1/RUVBL2 via their DII domains, as previously suggested.
A Circular plot showing identified intra- and inter-XLs in R2S622-926P dataset. All intra-SPAG1622-926 XLs of the TPR3 domain could be mapped and distance-validated on NMR TPR3 SPAG1622-926 structure (6I57.pdb). B Interaction network between RUVBL1, RUVBL2, SPAG1622-926 and PIH1D2 in R2S622-926P as identified by XL-MS. C XL-mapping on R2S622-926P cryo-EM structure, with XL distance-validated in green and violated in red. Overall, 84% of mapped XLs are in accordance with the structure. The SPAG1 positioning and conformation are confirmed by 4 inter-XLs and 4 intra-XLs, all distance-validated.
On the SPAG1622-926 side, the network plot revealed many interactions between SPAG1622-926 and RUVBL1/RUVBL2 (27 cross-link pairs, 25% of the total inter-XL pairs), involving the DIII domain of RUVBL2 (residues K368, K417 and K427) and all RUVBL1 domains (DI, DII and DIII) (Fig. 7B and Supplementary Table 4). In addition, the XL-MS dataset also revealed 17 inter SPAG1622-926:PIH1D2 cross-links, mainly between the CS domain of PIH1D2 (K252, K293 and K306) and the TPR3 of SPAG1622-926 (K641, K670, K706 and K710), in agreement with the NMR results. Finally, the PIH domain of PIH1D2 was also in close proximity of SPAG1622-926_TPR3, as depicted by the cross-links between K42 of PIH1D2 and K628/K744 of SPAG1622-926 (Supplementary Fig. 17). Overall, XL-MS data suggest that although firmly anchored via its SPAG1_C to RUVBL2, the remaining of the SPAG1622-926 construct is relatively flexible and engages RUVBL1, as well as PIH1D2.
We next mapped the identified cross-links onto the cryo-EM maps (Fig. 7C). As the high-resolution cryo-EM reconstruction lacks PIH1D2 and is poorly resolved in the RUVBLs DIIext regions, only 51 XL pairs (24%) could be mapped on the EM structure (30 intra-links and 21 inter-links). Among these inter-protein XLs, 10 involved RUVBL1 and RUVBL2, while 5 accounted for SPAG1622-926:RUVBL1/RUVBL2 pairs. Remarkably, 84% of measurable Cα-Cα distances obtained from the EM structure agreed with DSBU distance constraints (5-32 Å). The violated distances (8 XL-pairs) all included a highly dynamic DIIext (Supplementary Fig. 18). The XL-MS dataset thus cross-validated the EM results and in particular the position of SPAG1622-926 within the cryo-EM structure.
An integrative 3D model of the R2S622-926P complex
To assess the overall 3D architecture of the full R2S622-926P complex, we integrated our NMR, cryo-EM and XL-MS data to model the structure of the full R2S622-926P complex. Our cryo-EM map allowed a high-resolution reconstruction of the hexameric RUVBL1/RUVBL2 domains DI-DIII crowned by one SPAG1_C domain. Although less well resolved, densities located under this hexameric ring could nonetheless unambiguously be attributed to the DIIext of RUVBL1/RUVBL2 in an open conformation. Cryo-EM maps also revealed a hazy density that could not be definitively interpreted, but which we hypothesized corresponded to the presence of PIH1D2 (see above). As there is currently no experimental 3D structure of PIH1D2’s PIH domain, we used AF3 structure predictions of PIH1D2 and/or SPAG1622-926:PIH1D2 complex and combined them with our NMR, cryo-EM and XL-MS data to generate a 3D model of the “full” R2S622-926P complex (Fig. 8, Supplementary Fig. 20). Positioning of SPAG1622-926 was done using 7 XLs with RUVBL1 and 4 with RUVBL2, while PIH1D2 positioning was done using 5 XLs with RUVBL1 and 2 XLs with RUVBL2 (Supplementary Fig. 20). The resulting model was in good agreement with the XL-MS dataset, as it accommodated 148 out of the identified 208 XL-pairs, a 3-fold increase compared to the cryo-EM structure of R2SP with the missing densities (51/208). Out of these 148 XL-pairs, 71% were distance-validated, supporting overall positioning of proteins in the integrative model (Fig. 8B). SPAG1622-926 and PIH1D2 conformations were distance-validated with 100% and 64% of intra-protein XLs, respectively. Altogether, this integrative architectural model suggests that the SPAG1_C and PIH domains of SPAG1 and PIH1D2 clamp the RUVBL1/RUVBL2’s hexameric ring at their top and bottom sides, respectively, while SPAG1’s TPR3 and PIH1D2’s CS domains interact with the DIIext of RUVBL1/RUVBL2 (Fig. 8A). Interestingly, using AF3 structure prediction exclusively, generated a structure of the “full” R2S622-926P complex that placed DIIext of RUVBL1 and RUVBL2 in a closed position (Supplementary Fig. 19A). This is in contrast to our cryo-EM data, where DIIext seemed to be in open conformation and moving along a plane that is perpendicular to the RUVBL1/RUVBL2 central tunnel (see Supplementary Movie 2, PC2 analysis). Additionally, this AF3 predicted structure of full R2S622-926P complex validates 58% of XL-pairs, with no agreement with predicted inter SPAG1622-926:PIH1D2 interface (11/12 violated) and less than half (11/27) distance-validated XLs of SPAG1622-926:RUVBL1/RUVBL2 (Supplementary Fig. 19B). Therefore, AF3 alone was not sufficient to yield a R2S622-926P complex structure compatible with experimental data, highlighting the benefits of our integrative approach.
A Proposed 3D model of the full R2S622-926P assembly based on NMR, XL-MS, cryo-EM and AF3 predictions. RUVBL1 and RUVBL2 are represented in pink and blue, respectively, the SPAG1_C and TPR3 domains of SPAG1 (SPAG1622-926) are shown in lime and forest green, respectively. PIH1D2 is represented in dark red (PIH domain) and orange (CS domain). B Representation of cross-link pairs validated on the integrative model (71% of the total XL pairs, black dashed lines).
Discussion
In this study we used biochemical, biophysical and integrative structural biology approaches to unveil mechanistic and architectural aspects of R2SP complex, a R2TP-like quaternary chaperone with distinct client preference, tissue expression and function. Although SPAG1 and PIH1D2, two adapter proteins found in R2SP, share several homologous domains with RPAP3 and PIH1D1, adapters that belong to the archetypal R2TP complex12, our results indicate some key differences between how these adapters engage and modulate structure and activity of RUVBL1/RUVBL2 ATPase core. Within R2TP, RPAP3 is the sole binder of RUVBL1/RUVBL213,15. W. Houry’s group suggested that PIH1D1 does not directly interact with RUVBL1/RUVBL2 in the presence of RPAP3, but is rather positioned near the ring due to its direct interaction with RPAP313. This is further supported by the existence of sub-complexes such as R2T13 with yet unknown functions, and the existence of an endogenous version of RPAP3 (isoform2) unable to interact with PIH1D112. Such stable sub-complexes for R2SP and splicing isoforms for SPAG1 have not yet been identified in vivo, and our data indicates that they might not form at all. Our co-expression and SPR analysis hint towards a cooperative mechanism of SPAG1 and PIH1D2 binding to the RUVBL1/RUVBL2 ring, as SPAG1 seems to require PIH1D2 for high affinity complex formation. This is reinforced by our integrative structural model (informed by the results of our NMR, cryo-EM and XL-MS studies, and point mutations) that places PIH1D2 CS domain in contact with SPAG1, and both PIH1D2 CS and PIH domains in contact with DII domains of RUVBL1/RUVBL2 (Fig. 8). It will be interesting to further elucidate the details of the binding of PIH1D2 to RUVBL1/RUVBL2 (here this was technically not achievable due to PIH1D2 alone being unstable in the SPR assays). Therefore, our data clearly indicate that the behavior of PIH1D2 (from R2SP) differs from PIH1D1 (from R2TP) by playing a crucial role in the assembly of the R2SP complex.
ATP binding and/or hydrolysis are required for most of the RUVBL1/RUVBL2-related quaternary chaperones41. Our activity results show that SPAG1:PIH1D2 stimulates the RUVBL1/RUVBL2’s ATPase activity almost twice as much as RPAP3:PIH1D1. This might be correlated with our SPR analysis, in which the use of ATP-loaded dead mutant versions of RUVBL1/RUVBL2 negatively impacts the interaction with SPAG1622-926:PIH1D2 when compared to WT, i.e. ADP-containing RUVBL1/RUVBL2. These data indicate that SPAG1:PIH1D2 binding to RUVBL1/RUVBL2 is nucleotide-dependent; however, how the presence of ATP in the active site influences SPAG1622-926:PIH1D2 binding kinetics and vice versa remain elusive. The nucleotide binding could either lead to changes in the DI and DIII ring’s apical region, interfering with SPAG1 binding, or influence DIIext positioning as previously proposed26, thus affecting PIH1D2 binding. Furthermore, our kinetic assays show that both RPAP3:PIH1D1 and SPAG1:PIH1D2 facilitate nucleotide dissociation rather than association to RUVBL1/RUVBL2 binding sites. Our experiments with RUVBL1/RUVBL2 alone further reveal the presence of two types of nucleotide binding sites with distinct association and dissociation rates (one faster than the other). RPAP3:PIH1D1 appears more potent in dissociating non-hydrolysable ATP from the fast site, while SPAG1:PIH1D2 is more efficient on the slower, rate-limiting site. We hypothesize that those two ATP binding sites correspond to RUVBL1 and RUVBL2, but further experiments are needed to identify which one is which. Of note, in our cryo-EM structure, all active sites are fully occupied by ADP, regardless of being bound to SPAG1 or PIH1D2. This suggests a different molecular mechanism from R2TP where PIH1D1 seems to promote nucleotide release from the active site of an RUVBL2 protomer, while the other ring’s active sites remain ADP bound24. Here, it’s worth pointing out that another group13 used ATPase assays to show no significant effect of RPAP3:PIH1D1 on the activity of RUVBL1/RUVBL213. To summarize our data, we built two schematic models describing the cooperativity of SPAG1 and PIH1D2 binding to RUVBL1/RUVBL2 in R2SP (Fig. 9A) versus the independent binding of RPAP3 and PIH1D1 in R2TP (Fig. 9B). These mechanistic differences may be in part due to key structural differences between the two quaternary chaperones.
A R2SP assembly is proposed to involve cooperativity between SPAG1 and PIH1D2 as they engage RUVBL1/RUVBL2. B R2TP assembly is proposed to involve independent binding of RPAP3 and PIH1D1 to RUVBL1/RUVBL2. In both panels, the position of the pointer on the counter represents the rate of ADP release.
Although similar, structural models of R2SP and R2TP exhibit key differences in terms of how adapter proteins engage with the RUVBL1/RUVBL2 core. Our three-dimensional model of the R2SP chaperone shows that PIH1D2’s PIH domain, known to recruit hypothetical phosphorylated clients42, is located below the ring formed by the DIIs of RUVBL1/RUVBL2. In contrast, its CS domain binds closely to SPAG1’s TPR3 domain, close to the RUVBL1/RUVBL2’s DII external parts, most probably on the side of RUVBL1/RUVBL2 hexameric assembly. This is distinct from what is observed in R2TP, and likely caused by different lengths in the unstructured regions linking the structured domains of PIH1D2 and SPAG1, compared to PIH1D1 and RPAP3. Indeed, whereas RPAP3 has a long linker connecting PRD to RBD (Fig. 1), proposed to play a role in the competition between RPAP3 and PIH1D1 for RUVBL1/RUVBL2 DIIext binding13, SPAG1 has a short linker between PRD and RBD. This short length might impose PIH1D2 protein to adopt bilobal conformation with the CS domain near the RUVBL1/RUVBL2 ring’s vicinity, and the PIH domain in the basal region. We hypothesize that the bilobal conformation adopted by PIH1D2 could compensate for the shorter SPAG1 linker region and thus facilitate SPAG1 binding to RUVBL1/RUVBL2. We further propose that these differences could help explain why the R2SP assembly mechanism differs from the R2TP’s one.
An open set of questions in the field revolves around how clients engage with quaternary chaperones. Although we did not experimentally address client engagement in this study, our R2SP 3D model suggests that clients most likely engage the DII side of the ring. SPAG1, which is anchored via its C-terminal RBD to the opposite side of the RUVBL1/RUVBL2 ring, would thus keep HSP70/90 chaperones known to bind SPAG1 N-terminal TPR domains (Fig. 1)33,43 hanging close by to assist the assembly process. The clients would be recruited through PIH1D215 or the HSPs. To date, no clients have been shown to bind the upper side of the RUVBL1/RUVBL2 ring (SPAG1_C side), this binding polarity might likely be extended to other quaternary folding processes involving RUVBL1/RUVBL2. Importantly, we observed that RUVBL1/RUVBL2 ring engages up to three SPAG1/PIH1D2 complexes. Our extensive cryo-EM image analysis also hints towards high flexibility of RUVBL1/RUVBL2 DIIext, PIH1D2 and SPAG1’s TPR3 in R2SP complex. As such, we propose that R2SP is an octopus-like dynamic platform for quaternary client assembly whereby its spatial organization and flexibility allows R2SP to synchronize interactions with different clients and their transfer from HSP70/90. In summary, our study of the human R2SP complex adds another biophysical and structural piece to the RUVBL1/RUVBL2-based quaternary chaperone puzzle, shading light into chaperoning molecular mechanisms deeply dependent on fine-tuned protein-protein interactions.
Methods
NMR
NMR protein samples were prepared with a classic procedure involving, (i) production in E. coli BL21(DE3) pRARE2 cells grown in a minimal M9 media containing 13C-D6-glucose and 15NH4Cl, and (ii) a two-steps purification process based on immobilized metal ion affinity chromatography followed by gel filtration.
More in details, SPAG1796-926 (SPAG1_C) was produced with a N-terminal 6xHIS tag that allowed its purification. The tag was cleaved by 3C protease and the protein was isolated on a S75 Superdex column. The 13C/15N-labeled NMR sample was concentrated at 1 mM in 10 mM NaPi, 150 mM NaCl, pH 6.4, 0.5 mM TCEP (5%/95% D2O/H2O). 1H-15N HSQC, 1H-13C (aliphatic and aromatic), HNCO, HNCACO, HNCA, HNCOCA, HNCACB, CBCACONH, HBHACONH, HCCCONH, CCCONH, HCCH-TOCSY, HCCH-COSY, HNHA, 1H-1H NOESY, 1H-15N-NOESY-HSQC, and 1H 13C-NOESY-HSQC (NOESY mixing time, 120 ms) spectra were recorded at 298 K using a 600 MHz AVANCE III spectrometer equipped with a cryoprobe, enabling the nearly complete 1H, 13C, and 15N resonance assignments. The assignment was performed with CARA1.8.444.
For structure calculation of SPAG1_C, backbone chemical shifts were converted into dihedral angle restraints using TALOS-N. The CYANA 3.97 automated procedure was then employed to extract distance restraints from 2D 1H-1H NOESY, 3D 1H 15N-NOESY-HSQC, and 3D 1H 13C-NOESY-HSQC spectra, all recorded with a mixing time of 120 ms. The resultant sets of dihedral angles and inter-proton restraints underwent thorough verification and were utilized to generate 100 CYANA structures, subsequently refined in explicit water using the AMBER-based Portal Server for NMR structures (AMPS-NMR). The 20 structures with the lowest constraint energies were chosen as the most representative. Visualization of all 3D structures was performed using Pymol software45. Chemical shifts were deposited in the Biological Magnetic Resonance Bank under reference 34968. The 3D structure of SPAG1_C was deposited in the Protein Data Bank under reference 9HKR.
For the PIH1D2_CS:SPAG1622-762 complex, the bacterial cultures were performed as for SPAG1_C, but in D2O. The two proteins fragments were co-expressed due to the insolubility of the free PIH1D2_CS. The complex was purified thanks to the 6xHIS tag at the N-terminal end of SPAG1622-762, PIH1D2_CS being untagged. The tag was cleaved by the 3C protease and the complex was isolated on a S200 Superdex column. The 2H/13C/15N NMR sample was concentrated at 0.5 mM in 10 mM NaPi, 150 mM NaCl, pH 6.4, 0.5 mM TCEP (5%/95% D2O/H2O). This permitted to exchange amide and all labile 2H with the 1H coming from the buffer. The assignment was based on the analysis of HNCA, CBCACONH, HNCACB, HNCO, HNCACO and 1H-15N NOESY-HSQC (120 ms mixing time) spectra recorded at 293 K using a 600 MHz AVANCE III spectrometer equipped with a cryoprobe. The assignment of the spectra was guided by the assignment of the free SPAG1622-74233. This allowed a partial assignment of backbone nuclei of the PIH1D2_CS:SPAG1622-762 complex (Cα,Cβ, N, HN and CO). HN-HN cross-peaks on the 1H-15N NOESY-HSQC spectrum were detected using CARA1.8.4.
To monitor the binding of unlabeled RUVBL proteins to 13C-labeled SPAG1_C, 1D METHYL-SOFAST-HMQC spectra were recorded. Interaction experiments were conducted at 298 K and 600 MHz in the RUVBL1/RUVBL2 buffer (20 mM Tris-HCl pH 8.0, 250 mM NaCl, 5% glycerol, 2 mM MgCl2, 0.5 mM TCEP). RUVBL1 was tagged with a 6xHistidine at the N-terminus, and RUVBL2 was tagged with a 3xFLAG. The concentration of 13C-labeled proteins was approximately 10 μM. The 1H dimension was edited to selectively detect 1H nuclei attached to 13C nuclei. Protons attached to 13C nuclei in the range of 5 to 35 ppm were selectively excited with a bandwidth of 3 ppm centered at 0 ppm. The relaxation delay was set to 150 ms, and the number of scans was 2048. The final concentration ratio between unlabeled and labeled proteins was maintained at 1:1, considering one monomer of SPAG1_C and one heterodimer of RUVBL1/RUVBL2.
LUMIER IP
HEK293T cells from ATCC were seeded in 24-well plates and transfected with 450 ng of RL fusion and 50 ng of the 3xFLAG-FFL fusion, with 1 μL of JetPrime (PolyPlus), as recommended by the manufacturer. 48 h later, cells were extracted for 15 min at 4 °C in 450 μl of HNTG containing protease inhibitor cocktail (Roche) and spun down at 4 °C and at 20,000 x g for 10 min. The IP was performed duplicated, by putting 100 μL of the extract in each of four wells in a 96-well plate, with two wells being coated with anti-FLAG antibody (see below), and two control wells without antibodies. Plates were incubated for 3 h at 4 °C and then washed 5 times with 300 μL of ice-cold HNTG, for 10 min at 4 °C for each wash. After the last wash, 10 μL of PBL buffer (Promega) was added in each well. To measure the input, 2 μl of extract and 8 μL of 1 x PBL buffer were put in empty remaining wells. Plates were then incubated 5 min at room temperature, and FFL and RL luciferase activities were measured in IP and input wells, using the dual luciferase kit (Promega). Every transfection was performed three times as independent replicates.
To coat the wells of the 96-well plates with M2 anti-FLAG antibodies, High-binding plates were used (Lumitrac, Greiner), and 70 μL of M2 antibody (10 μg/mL in 1 x PBS; F1804 Sigma-Aldrich) were put in each well and incubated overnight at room temperature in the dark. The next day, wells were blocked with 300 μL of blocking buffer, for 1 h at room temperature. IP control wells were treated the same way except that no antibody was put in the well. Blocking buffer was 3% Bovine Serum Albumin (BSA), 5% sucrose, 0.5% Tween 20, 1xPBS). HNTG buffer was 20 mM HEPES-KOH pH 7.9, 150 mM NaCl, 1% Triton X-100, 10% glycerol, 1 mM MgCl2, 1 mM EGTA.
Co-expression assays and cryo-EM samples preparation
The sequences coding the proteins or protein fragments of interest, wild type and mutants, are subcloned between NdeI and BamHI rectriction sites into 3 pET-based vectors: pnEA which is ampicillin resistant and encodes a protein carrying an N-terminal 6-Histidine tag followed by a 3C protease site, pnCS which is spectinomycin resistant and encodes a native protein, and pnYK which is kanamycin resistant and encodes also a native protein.
For co-expression assays, the vectors encoding the proteins or protein fragments are co-transformed into E. coli BL21(DE3)pRARE2 expression strain. The clones selected on LB-agar plates supplemented with the adequate antibiotics are grown in 100 mL LB medium at 37 °C; when OD600 reaches 0.7, the cultures are induced with 0.2 mM IPTG and placed O/N at 20 °C under agitation.
Then, the cells are harvested, and the pellets resuspended into 4 mL lysis buffer (HEPES 25 mM pH 7.5; NaCl 300 mM; imidazole 10 mM; TCEP 0.5 mM), sonicated, and the lysate is cleared by centrifugation. 200 µL TALON resin (50% slurry) are added to the supernatant (soluble fraction) and after a 15 to 30 min binding step on a rotary shaker, the resin is spun down, washed 3 times with 500 µL lysis buffer, and 250 µL elution buffer (HEPES 25 mM pH 7.5; NaCl 300 mM; Imidazole 300 mM; TCEP 0.5 mM) are added to recover the complexes.
Purification steps are followed on SDS-PAGE where insoluble fraction (P), Soluble fraction (So), proteins Bound to the resin (B) and Elution (E) are loaded and colored by Coomassie blue staining.
For reconstituted R2SP and R2S622-926P cryo-EM samples preparation, RUVBL1/RUVBL2 and SPAG1:PIH1D2 or SPAG1622-926:PIH1D2 are produced in large cultures and purified separately, then mixed.
For R2SP, transformation of pETDuet:His-RUVBL1/RUVBL2-FH8 on one hand, and co-transformation of pnEA::PIH1D2 and pnYK::SPAG1 (pnYK::SPAG1622-926 for R2S622-926P) on the other hand are performed into E. coli BL21(DE3)pRARE2 expression strain. The clones selected on LB-agar plates supplemented with the adequate antibiotics are grown in 1 L flasks in LB medium at 37 °C and when OD600 reaches 0.7, the cultures are induced with 0.2 mM IPTG. After O/N growth at 20 °C under agitation, the cells are harvested and pellets resuspended into lysis buffer (HEPES 25 mM pH 7.5; NaCl 300 mM; Imidazole 10 mM; TCEP 0.5 mM; Glycerol 5% for RUVBL1/RUVBL2; ADP 0.5 mM for R2S622-926P) (40 mL/L culture pellet), and sonicated. The lysate is cleared by centrifugation and the complexes are purified on TALON resin as followed: 2 mL resin (50% slurry) are added to the 40 mL supernatant (soluble fraction) and after a 15 to 30 min binding step on a rotary shaker, the resin is spun down, washed 3 times with 5 mL lysis buffer and the complexes are eluted with 3 x 5 mL elution buffer (HEPES 25 mM pH 7.5; NaCl 300 mM; Imidazole 300 mM; TCEP 0.5 mM; Glycerol 5% for RUVBL1/RUVBL2; ADP 0.5 mM for R2S622-926P). Human Rhino Virus 3C (HRV-3C) protease is added to the elution for O/N cleavage at 4 °C of the FH8 C-terminal tag for RUVBL2 and the Hisx6 N-terminal tag for SPAG1:PIH1D2 (or SPAG1622-926:PIH1D2). After size exclusion chromatography (SEC) on Superose6 column, RUVBL1/RUVBL2 is mixed with excess SPAG1:PIH1D2 (or SPAG1622-926:PIH1D2), and the reconstituted R2SP (or R2S622-926P) complex is purified again on Superose6 column in final buffer (HEPES 20 mM pH 7.5; NaCl 150 mM; TCEP 0.5 mM; ADP 0.5 mM for R2S622-926P) to eliminate excess of SPAG1:PIH1D2 (or SPAG1622-926:PIH1D2) and generate a cryo-EM sample at 1.1 mg/mL.
Protein co-expression and purification for the SPR experiment
RUVBL1 and RUVBL2 were produced as described in ref. 15 with the following modifications: (i) RUVBL2 has an N-terminal FH8-tag preceded by an HRV-3C cleavage site; (ii) Peak fractions from the HisTrap were incubated with 5 mM CaCl2 and loaded onto a HiPrepTM Octyl FF 16/10 column (Cytiva). A Superose 6 Increase 10/300 GL equilibrated with 20 mM Tris-HCl pH 8.0, 150 mM NaCl, 5% glycerol, 2 mM MgCl2 and 0.5 mM TCEP, was used to separate oligomeric species. RUVBL1D302N/RUVBL2D299N active site mutant variant was purified in the presence of 4 mM ATP, and RUVBL1ΔDII/RUVBL2ΔDII truncated variant complex was purified in the absence of nucleotides.
SPAG1622-926 carries a C-terminal Flag-tag without a cleavage site, while PIH1D2_CS or PIH1D2 have a C-terminal StrepTag II preceded by an HRV-3C cleavage site. SPAG1622-926:PIH1D2_CS or SPAG1622-926:PIH1D2 were co-expressed in E. coli (DE3*) (Novagen, 71400), with 50 µM IPTG overnight at 18 °C in a New Brunswick™ (Innova®) 44 R Shaker at 150 rpm. The complexes were immobilized in a 5 mL StrepTactin XT HC (IBA life sciences), previously equilibrated in 20 mM HEPES pH 8, 300 mM NaCl, 0.5 mM TCEP, and eluted with 50 mM Biotin. Peak fractions collected from the StrepTactin XT were injected in a Superdex 200 16/60 XK, allowing the isolation of a heterodimer. Collected fractions from the main peak were diluted to reduce the concentration of NaCl to 50 mM, further polished in Resource TM Q (Cytiva), and eluted with a linear gradient, allowing the separation of a major peak corresponding to the intact complex (w/o degradation) at approximately 170 mM NaCl. Tag removal from collected peak fractions was performed by incubating 18 h at 4 °C with 1% (w/w) HRV-3C protease (Thermo Fisher Scientific, TFS). Digested sample was injected into a 5 mL StrepTactin XT in tandem with 1 mL HisTrap. All purification steps were carried out at room temperature and were monitored by NuPage Bis-Tris gels (Invitrogen, NP0302).
Surface plasmon resonance
SPAG1622-926:PIH1D2_CS, SPAG1622-926:PIH1D2, or SPAG1622-926 protein constructs were immobilized onto CM5 (Series S) sensor chips using standard amine coupling. 10 mM HEPES, pH 7.4, 150 mM NaCl (HSB-N) was used as the background buffer during immobilization. The carboxymethyl dextran surface of all flow cells was activated with 20 mM EDC and 5 mM NHS for 1.5 min. Both SPAG1/PIH1D2 constructs were injected on the chip surface at a final concentration of 0.5 µg/mL in 10 mM Sodium Acetate buffer pH 5.5, while SPAG1622-926 was injected in 10 mM Bis-Tris pH 6.5. The enzymes were coupled to the surface with a 1 to 2 min injection time at a flow rate of 10 μL min−1. The remaining activated groups were blocked with a 5 min injection of 1.0 M ethanolamine, pH 8.5. Typically, 100 ± 10 response units (RU) were obtained for the immobilization of all protein constructs. Negative controls were performed by immobilizing either BSA (TFS) or human Cyclophilin D43-207 (CypD) with the same RU levels as the proteins of interest. CypD43-207 is a 22 kDa in-house purified protein with similar size to SPAG1_C and confirmed to be active through binding to Cyclosporin A. All RUVBL1/RUVBL2 complexes were directly dissolved in running buffer (20 mM NaKPi pH 7.5, 150 mM NaCl, 5 mM MgCl2, 1 mM DTT, 0.05% P20), in the absence or presence of nucleotides and analyzed with a Biacore 4000 (Biacore AB, Cytiva, Uppsala, Sweden) at 25 °C. RUVBL complex was tested at 10 different concentrations using a 2-fold dilution series. Interaction analysis cycles consisted of a 300 s sample injection (30 µL min−1) followed by 600 s of buffer flow (dissociation phase), with a data collection rate of 10 Hz. All sensorgrams were processed by subtracting the binding response recorded from the control surface (reference spot), followed by subtracting of the blank buffer injection from the reaction spot (double referencing). All datasets were fitted to a simple 1:1 Langmuir interaction model to determine kinetic parameters (ka, kd, KD) and affinity constants at steady state (KDss). A local Rmax was applied for all interaction with resolved binding kinetics.
ATPase assay
The steady-state ATPase activity of RUVBL1/RUVBL2 complex was evaluated using a coupled enzymatic assay (PK/LDH) that monitors the oxidation of NADH in the presence of excess ATP12, with a modified reaction buffer (50 mM HEPES/KOH pH=7.5, 100 mM KCl, 5 mM ATP; 5 mM MgCl2) in 96-well plates on a CLARIOstar reader (BMG Labtech) at 37 °C. The RUVBL1/RUVBL2 concentration was 1 µM of hexamer. The concentration required for half-maximum activation between RUVBL1/RUVBL2 and RPAP3:PIH1D2 or SPAG1:PIH1D2 was calculated by relative specific ATPase activity (SA) of RUVBL1/RUVBL2 and fitted with the following equation using R software46: \({SA}={{SA}}_{R1R2}+\frac{{{SA}}_{\max }\times \left[R3P1{orS}1P2\right]}{{{AC}}_{50}+\left[R3P1{orS}1P2\right]}\)
Fluorescence polarization assay
The fluorescence polarization of BODIPY® FL ATPγS (Jena Bioscience) was measured by Clariostar (filters: excitation: 482-16; dichroic: LP 504; emission: 530-40) on 384 wells-plate at 25 °C. All fluorescence polarization assays were performed into FP buffer (25 mM HEPES/KOH pH = 7.5, 150 mM KCl, 5 mM MgCl2) with a final concentration of BDP-ATPγS at 10 nM. The nucleotides of RUVBL1/RUVBL2 complex were removed by incubation with Agarose-phosphatase alkaline (P0762, Sigma) at 25 °C overnight. A fraction of the protein sample was then denatured at 100 °C for 20 min, followed by centrifugation at 15,000 x g for 20 min. The level of remaining unlabeled nucleotides per monomer of RUVBL1/RUVBL2 in the supernatant was measured by absorbance at 260 nm using an extinction coefficient of ε = 15,400 M⁻¹ cm⁻¹47 (6.6% for the sample treated by alkaline phosphatase). For binding and equilibrium assays, the proteins were deposited into the wells using a pipette, and the BDP-ATPγS were added via the injector of Clariostar. For exit assays, proteins and BDP-ATPγS were deposited together with a pipette on the wells and unlabeled nucleotides were added via the injector of Clariostar. The data were fitted using R with these equations: (binding assay) \({FP}={{FP}}_{t=0}+({a}_{1}\times (1-{e}^{-{k}_{{obs}1}\times t}))+({a}_{2}\times (1-{e}^{-{k}_{{obs}2}\times t}))\); (exit assay) \({FP}={{FP}}_{t=0}+\left({a}_{1}\times {e}^{-{k}_{{off}1}\times t}\right)+\left({a}_{2}\times {e}^{-{k}_{{off}2}\times t}\right)\)
Native mass spectrometry
Samples were buffer exchanged against ammonium acetate (200 mM, pH 7.5) with Zeba Spin desalting columns (7 kDa cutoff, TFS, Rockford, IL, USA). Sample concentrations were determined by UV absorbance using a NanoDrop 2000 spectrophotometer (TFS, France). Prior to injection, full length RUVBL1/RUVBL2 and SPAG1622-926:PIH1D2 were diluted and mixed in ratio 1:2 (detected species were the same even with higher proportions of SPAG1622-926:PIH1D2). nMS experiments were carried out on a hybrid electrospray quadrupole time-of-flight mass spectrometer (Synapt G2 HDMS, Waters, Manchester, UK) coupled to an automated chip-based nanoelectrospray source (Triversa Nanomate, Advion, Ithaca, USA) operating in the positive ion mode. Mass spectrometer calibration was performed from 1000 to 20,000 m/z using singly charged ions produced by a 2 mg/mL solution of cesium iodide in 2-propanol/water (v/v). Instrumental parameters were optimized to ensure the transmission of high masses while maintaining non-covalent interactions. The backing pressure was set to 6 mbar and the cone voltage to 200 V. Data interpretation was performed using MassLynx v4.1 (Waters, Manchester, UK).
Mass photometry
Measurements of were done on a TWO MP (Refeyn Ltd, Oxford, UK) at room temperature. We used microscope slides (24 x 50 mm, 170 ± 5 µm, No. 1.5H, Paul Marienfeld GmbH & Co. KG, Germany) that were cleaned (milli-Q water, isopropanol, milli-Q water) and dried (clean nitrogen stream) and further assembled with six-well silicone gaskets (cut from CultureWellTM, 50-3 mm DIA x 1 mm Depth, 3-10 µL, Grace Bio-Labs, Inc., Oregon, USA). Focus was made each time with one 18 µL PBS droplet before mixing with 2 µL of sample to analyze. A contrast-to-mass calibration was performed twice a day with a mix of BSA, Bevacizumab and Glutamate Dehydrogenase in PBS. For RUVBL1/RUVBL2 and R2SP analysis, samples were first diluted in their native sample to reach 500 nM and 2 µL of these stock solutions were then quickly diluted to 50 nM in the 18 µL droplet of PBS. Three movies of 60 s (3000 frames) were recorded for each sample using AcquireMP sotware (Refeyn Ltd, Oxford, UK), data were processed with DiscoverMP software (Refeyn Ltd, Oxford, UK).
XL-MS
SPAG1:PIH1D2 heterodimer and R2SP complex (20 mM HEPES; 150 mM NaCl; 0.5 mM TCEP; pH 7.5) were both incubated in biological triplicate with DSBU cross-linker (CF plus Chemicals) at 100 molar excesses for 45 min (RT) and quenched with 20 mM ammonium bicarbonate for 20 min. XL reaction was controlled using home-made SDS-PAGE (12%). Samples were then reduced for 30 min at 37 °C (5 mM DTT), alkylated for 1 h in the dark (15 mM Iodoacetamide), and finally digested overnight at 37 °C with Trypsin/Lys-C (Promega, Madison, USA) at a 50:1 Protein:Enzyme ratio. Digestion was quenched with 1% v/v TriFluoroacetic Acid (TFA), and peptides were submitted to a SPE cleanup (Sep-Pak C18 1cc, 50 mg). After evaporation and dilution in 2% ACN/0.1% (Formic Acid) FA, samples were analyzed in nanoLC-MS/MS using a nanoAcquity UPLC (Waters, Milford, USA) coupled to a Q-Exactive HF-X Orbitrap mass spectrometer (TFS) controlled by XCalibur software (v4.0.27.9). Trapping was done on a nanoEase M/Z Symmetry C18 precolumn (180 µm x 20 mm, 5 µm particle size, Waters) and separation on a nanoEase M/Z Peptides BEH C18 (75 µm x 250 mm, 1.7 µm particle size, Waters). The following 105 min gradient was applied using mobile phase A (water/0.1 % FA) and B (ACN/0.1% FA): 3% for 5 min, 3–40% B in 90 min, 40–99% B in 2 min, 99% B for 5 min, 99-3% B in 2 min. Flow rate was set to 400 nL/min and column temperature to 60 °C. Full scan MS spectra were acquired in positive mode with a resolution of 120,000, with a 300–1800 m/z range. Fragmentation of Top 10 most intense ions (charge states 3–8 +) were done using HCD stepped collision energy (27, 30, 33% NCE). MS/MS spectra were acquired with a resolution of 30,000 and a 2 m/z isolation window. Raw data were converted into.mgf files and processed using MeroX software v2.0.1.4 with a mass tolerance of 5 ppm for precursor ions and 10 ppm for product ions. A 5% FDR cut-off and a signal-to-noise of at least 2 were also applied. Proteases cleavage sites were defined as Lys and Arg with maximum 3 missed cleavages, only peptides of at least 5 residues were considered. Cysteine carbamidomethylation was set as fixed modification and Methionine Oxidation as variable modification (2 max. mod.). Lys and N-ter as well as Ser, Thr and Tyr were considered as cross-linking sites. Database containing interest proteins and > 50 common contaminants was used for identification with RISEUP mode and 1 max. missing ion. Cross-links composed of consecutive amino acid sequences were not used and all cross-links identified were manually validated. Only cross-links identified in at least 2/3 XL replicates were validated. The SPAG1:PIH1D2 and R2SP XL-MS datasets have been deposited to ProteomeExchange via the PRIDE partner repository, with dataset identifier PXD056494. For mapping on sequences and distance validation of identified XLs on the different structures (< 32 Å cutoff for Cα-Cα), we used the xiVIEW webserver (https://xiview.org)48. XLs were further visualized using PyMol version 2.5.4 (Schrödinger, LLC)45 and UCSF ChimeraX version 1.849.
Grid preparation and cryo-EM images acquisition
Cryo-EM grids were prepared and systematically checked at METI, Toulouse. Immediately after glow discharge, 3.5 µL of purified R2S622-926P particles (at 0.45 mg/mL as estimated by Bradford) were deposited onto QUANTIFOIL holey carbon grids (R2/1, 300 Mesh). Grids were plunge-frozen using a Leica EM-GP automat; the temperature and humidity level of the loading chamber were maintained at 20 °C and 95%, respectively. Excess solution was blotted with a Whatman filter paper no. 1 for 1.7–1.9 s, and grids were immediately plunged into liquid ethane (− 183°C).
Images were recorded on a Titan Krios electron microscope (FEI, TFS) located at EMBL, Heidelberg, Germany. The cryo-electron microscope operated at 300 kV and was equipped with a Gatan K2 summit direct electron detector using counting mode. Automatic image acquisition was performed with SerialEM, at a magnification corresponding to a calibrated pixel size of 0.645 Å and a total electron dose of 51.92 e-/Å2 over 40 frames. Nominal defocus values ranged from − 0.8 μm to − 2.8 μm.
Single-particle analysis
Image analysis was performed using Relion 3.150, unless otherwise stated. An overview of the image processing scheme is presented in Supplementary Figs. 10 and 13. Sixteen thousand five hundred and sixty-six stacks of frames were collected at EMBL Heidelberg. Frame stacks were aligned to correct for beam-induced motion using MotionCor251. Contrast transfer function (CTF) and defocus estimation were performed on the realigned stacks using GCTF52. After selection upon CTF estimation quality, maximum resolution on their power spectra, and visual checking, ‘good’ micrographs were retained for further analysis. A total of 2,158,039 particles were initially picked and then extracted in boxes of 304 × 304 pixels, using the LogPicker Autopick option from Relion 3.150. A first 2D classification was performed (on particle images binned by a factor of 4) to sort out bad particles, yielding a total of 1,870,648 remaining particles. Of note, on the 1,870,648 picked particles, at least 1,303,376 were classified as top views. These preferential orientations thus represent at least 70% of the imaged particles, suggesting a strong air-water interface bias during plunge freezing. A subset of 2D classes were first used for an ab initio 3D reconstruction, and the resulting dodecameric 3D map was used as a reference for a 3D classification in five classes of the particle dataset. One class harbored a dodecameric form of the R2SP with a good level of detail. The 253,429 particles from this class were re-extracted without imposing any binning factor, and a consensus 3D structure was obtained using RELION’s 3D auto-refine option, that reached a resolution of 3.5 Å according to gold-standard Fourier shell correlation (FSC) at 0.143 after CTF Refinement and particle polishing. Local resolution estimated by ResMap40 and particles angular distribution are shown on Supplementary Fig. 10. A post-processed map of the dodecameric assembly was calculated using DeepEMhancer51. The remaining classes were merged and subjected to a 3D classification (K = 5), using an hexameric RUVBL1/RUVBL2 atomic model derived from (PDB-ID: 2XSZ)53 as reference. One of the five 3D classes resulted in a clearly defined hexameric shape typical to the RUVBL1/RUVBL2 assembly. The four others were either featureless or exhibited a strong anisotropy of features corresponding to most top views, which were subsequently discarded (Supplementary Fig. 10, lower right panel). The 302,119 particles from the “good” 3D class were re-extracted without imposing any binning factor, and a consensus 3D structure was obtained at 3.6 Å resolution according to gold-standard FSC procedure. Local resolution ranged from 3.2 to 7.0 Å according to ResMap estimation40. Particles angular distribution and local resolution are shown on Supplementary Fig. 13 (lower left panel). To try to sort out structural heterogeneity within these hexameric particles, two different strategies were used (Supplementary Fig. 13). The first was to perform a global, unmasked 3D classification (K = 5) while skipping orientation searches, and the second used a mask around a monomer of RUVBL1 + RUVBL2 + SPAG1_C for a masked and focused 3D classification while skipping orientation searches. From the first strategy, a “cleaned” 3D class of 167,033 particles was obtained. These particles were subjected to 3D auto refinement, and the maps were subsequently improved by particle-based motion correction, B-Factor weighting, and CTF refinement, resulting in a final resolution of 3.6 Å that displayed a hexameric ring crowned by at least one “horn” (corresponding to SPAG1_C).
To assess how many SPAG1_C could be associated with RUVBL1/RUVBL2 hexameric ring, the second strategy (right panel on Supplementary Fig. 11) was to apply a mask around a monomer of RUVBL1, RUVBL2, SPAG1_C, and perform a focused 3D classification while skipping orientation searches. This resulted in 5 classes with either one or two horn-like structures of SPAG1_C which were refined to resolutions ranging from 5.0 to 3.8 Å (Supplementary Fig. 13).
Structural heterogeneity assessment within the cryo-EM datasets
To characterize their nature and level of structural heterogeneity, 302,119 particles corresponding to hexameric, or 252,248 particles corresponding to dodecameric assemblies (dodecamer before polishing) were also subjected to AI-based sorting by cryoDRGN version 2.3.038. All models were trained for 50 epocs, parallelized through several GPU’s. Both hexameric and dodecameric particles were downsampled to a box size of 128 pixels with a pixel size of 1.52 Å/pixel for evaluation. We trained a cryoDRGN model on these downsampled particles with three layers and 256 nodes per layer for both the encoder and decoder network architectures (termed 256 × 3) and an 8-dimensional latent variable Z (dodecameric assemblies were trained on layers with 512 nodes). This resulted in a smooth distribution of Z values, indicating that heterogeneity in the dataset originates purely from continuous conformational changes and not from populations of particles with distinct compositions, in agreement with the 3D classification results using RELION. We then represented the maps (Supplementary movies 1 and 2) using the traversal of the first, second, and third principal components of variability in the Z values, as implemented in cryoDRGN. For the hexameric particles, the resulting maps show a variety of states, including states with different number of SPAG1_C’s decorating the “top” region of the ring, the presence of density “beneath” the RUVBL1/RUVBL2 ring that might correspond to PIH1D2, DII from RUVBL1 moving laterally, and lack of density for DII of RUVBL2 that are bound to SPAG1_C. For the dodecameric particles, the resulting maps show different ways that both hexamers might move relative to each other, including twisting, compaction, and tilting, but also DII density variations, specifically from RUVBL2.
Cryo-EM Maps Interpretation
Atomic models of the RUVBL1/RUVBL2 hexamer (PDB-ID 6H7X, https://doi.org/10.2210/pdb6H7X/pdb, 2C9O, https://doi.org/10.2210/pdb2C9O/pdb; and 6FO1, https://doi.org/10.2210/pdb6FO1/pdb) as well as of the NMR structure of SPAG1_C herein presented were first fitted in the cryo-EM maps of interest as rigid body using the “fit” command in UCSF Chimera54. Manual refinements and adjustments, as well as flexible and jiggle fittings were then realized on various chains of the models in Coot55. Final atomic model was refined using REFMAC556. Atomic coordinates of domains DIIext of RUVBL1 (PDB 2C9O) and RUVBL2 (PDB 6H7X) were rigid-body docked in the cryo-EM map. Because of the low resolution of the map in this region, these atomic coordinates were not further refined. Final model evaluation was done with MolProbity57. Overfitting statistics were calculated by a random displacement of atoms in the model, followed by a refinement against one of the half-maps in REFMAC5, and Fourier shell correlation curves were calculated between the volume from the atomic model and each of the half-maps in REFMAC5 (Supplementary Table 5) Maps and models visualization was done with Coot, UCSF Chimera and ChimeraX49. Figures and movies were created using ChimeraX and Kdenlive (https://kdenlive.org/en/).
R2S622-926P integrative model establishment
The 3D structures of PIH1D2 and SPAG1, either in their full-length form or cut into smaller domains, were predicted using AlphaFold358. Namely, the following amino acid sequences of SPAG1622-926 and SPAG1622-926_TPR3_plus (622-780); or PIH1D21-315, PIH1D2_PIH_plus (1-163), PIH1D2_CS_plus (160-315) were submitted to structure prediction by AlphaFold3 (AF3) (Fig. 1). AF3 structural prediction of PIH1D2’s PIH domain was fitted on a segmented low-resolution cryo-EM map presented in Supplementary Movie 2 using Chimera’s Segger option. Then, combinations of SPAG1622-926:PIH1D2 (full length or fragments above described) were systematically submitted to AF3. Finally, different sequences enclosing RUVBL1 and RUVBL2 DIIs (namely: RUVBL1DII: 120-296; RUVBL1DIIext: 120-240; RUVBL2DII: 126-295; RUVBL2DIIext: 126-232) were also added to SPAG1622-926:PIH1D2 complexes, yielding SPAG1622-926:PIH1D2:RUVBL1/RUVBL2 assemblies for AF3 prediction. Assays using different copy numbers of these proteins/domains were also systematically performed (SPAG1622-926 and PIH1D2: 1 or 2 copies; RUVBL1 and RUVBL2: 1, 2 or 3 copies of each protein segment). For each of these AF3 predictions, all 5 output structures were examined using Chimera. The final structures of PIH1D2’s CS and SPAG1622-926 TPR3 domains that were retained for building the integrative R2S622-926P model corresponded to the 4th best AF3 prediction of the following construct, 2xSPAG1622-926_TPR3_plus:2xPIH1D2:1xRUVBL1DIIext:1xRUVBL2DIIext, because it was in best agreement with the experimental data from cryo-EM, NMR and XL-MS. Atomic coordinates of the PIH1D2’s CS and SPAG1622-926 TPR3 domains were extracted from this AF3 prediction and added to the R2S cryo-EM atomic model supplemented with PIH1D2’s PIH domain using Coot and Chimera.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data needed are present in the paper and/or the Supplementary Materials. Additional data related to this paper may be requested from the corresponding authors. The cryo-EM maps and the models have been deposited in the Electron Microscopy Data Bank (EMDB) with accession codes EMD-52013 and EMD-52336 for the hexameric R2S622-926P and dodecameric RUVBL1/RUVBL2 cryo-EM maps, respectively, and PDB ID 9HB4 and 9HPO for the RUVBL1/RUVBL2:SPAG1_C and dodecameric RUVBL1/RUVBL2 models, respectively. The chemical shifts and NMR structure of the human SPAG1_C are deposited in BMRB (ID: 34968) and PDB (ID: 9HKR), respectively. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE59 partner repository with the dataset identifier PXD056494. Source data are provided in this paper.
References
Horejsi, Z. et al. CK2 phospho-dependent binding of R2TP complex to TEL2 is essential for mTOR and SMG1 stability. Mol. Cell 39, 839–850 (2010).
Boulon, S. et al. The Hsp90 chaperone controls the biogenesis of L7Ae RNPs through conserved machinery. J. Cell Biol. 180, 579–595 (2008).
Machado-Pinilla, R., Liger, D., Leulliot, N. & Meier, U. T. Mechanism of the AAA+ ATPases pontin and reptin in the biogenesis of H/ACA RNPs. RNA 18, 1833–1845 (2012).
Schlotter, F. et al. Proteomic analyses reveal new features of the box H/ACA RNP biogenesis. Nucleic Acids Res. 51, 3357–3374 (2023).
Bizarro, J. et al. Proteomic and 3D structure analyses highlight the C/D box snoRNP assembly mechanism and its control. J Cell Biol. 207, 463–480 (2014).
Bizarro, J. et al. NUFIP and the HSP90/R2TP chaperone bind the SMN complex and facilitate assembly of U4-specific proteins. Nucleic Acids Res. 43, 8973–8989 (2015).
Houry, W. A., Bertrand, E. & Coulombe, B. The PAQosome, an R2TP-Based Chaperone for Quaternary Structure Formation. Trends Biochem. Sci. 43, 4–9 (2018).
Boulon, S. et al. HSP90 and its R2TP/Prefoldin-like cochaperone are involved in the cytoplasmic assembly of RNA polymerase II. Mol. Cell 39, 912–924 (2010).
Cloutier, P. et al. R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein. Nat. Commun. 8, 15615 (2017).
Coulombe, B., Cloutier, P. & Gauthier, M. S. How do our cells build their protein interactome? Nat. Commun. 9, 2955 (2018).
Cloutier, P. et al. Upstream ORF-Encoded ASDURF Is a Novel Prefoldin-like Subunit of the PAQosome. J. Proteome Res. 19, 18–27 (2020).
Henri, J. et al. Deep Structural Analysis of RPAP3 and PIH1D1, Two Components of the HSP90 Co-chaperone R2TP Complex. Structure https://doi.org/10.1016/j.str.2018.06.002 (2018).
Seraphim, T. V. et al. Assembly principles of the human R2TP chaperone complex reveal the presence of R2T and R2P complexes. Structure 30, 156–171 (2022).
Palacios-Abella, A. et al. The structure of the R2T complex reveals a different architecture from the related HSP90 cochaperone R2TP. Structure 33, 740–752.e8 (2025).
Maurizy, C. et al. The RPAP3-Cterminal domain identifies R2TP-like quaternary chaperones. Nat. Commun. 9, 2093 (2018).
Knowles, M. R. et al. Mutations in SPAG1 cause primary ciliary dyskinesia associated with defective outer and inner dynein arms. Am. J. Hum. Genet. 93, 711–720 (2013).
Yamamoto, R., Hirono, M. & Kamiya, R. Discrete PIH proteins function in the cytoplasmic preassembly of different subsets of axonemal dyneins. The J. Cell Biol. 190, 65–71 (2010).
Li, Y., Zhao, L., Yuan, S., Zhang, J. & Sun, Z. Axonemal dynein assembly requires the R2TP complex component Pontin. Development 144, 4684–4693 (2017).
Leigh, M. W. et al. Primary Ciliary Dyskinesia (PCD): A genetic disorder of motile cilia. Transl. Sci. Rare Dis. 4, 51–75 (2019).
Zariwala, M. A. et al. Primary Ciliary Dyskinesia (University of Washington, Seattle, Seattle (WA), 2007).
Dauden, M. I., López-Perrote, A. & Llorca, O. RUVBL1-RUVBL2 AAA-ATPase: a versatile scaffold for multiple complexes and functions. Curr. Opin. Struct. Biol. 67, 78–85 (2021).
Serna, M. et al. CryoEM of RUVBL1–RUVBL2–ZNHIT2, a complex that interacts with pre-mRNA-processing-splicing factor 8. Nucleic Acids Res. 50, 1128–1146 (2022).
Dos Santos Morais, R. et al. Deciphering cellular and molecular determinants of human DPCD protein in complex with RUVBL1/RUVBL2 AAA-ATPases. J. Mol. Biol. 434, 167760 (2022).
Munoz-Hernandez, H. et al. Structural mechanism for regulation of the AAA-ATPases RUVBL1-RUVBL2 in the R2TP co-chaperone revealed by cryo-EM. Sci. Adv. 5, eaaw1616 (2019).
Ayala, R. et al. Structure and regulation of the human INO80–nucleosome complex. Nature 556, 391–395 (2018).
Silva, S. T. N. et al. X-ray structure of full-length human RuvB-Like 2 – mechanistic insights into coupling between ATP binding and mechanical action. Sci. Rep. 8, 13726 (2018).
Theobald, D. L., Mitton-Fry, R. M. & Wuttke, D. S. Nucleic acid recognition by OB-fold proteins. Annu. Rev. Biophys. Biomol. Struct. 32, 115–133 (2003).
Ewens, C. A. et al. Architecture and nucleotide-dependent conformational changes of the Rvb1-Rvb2 AAA+ complex revealed by cryoelectron microscopy. Structure 24, 657–666 (2016).
López-Perrote, A. et al. Regulation of RUVBL1-RUVBL2 AAA-ATPases by the nonsense-mediated mRNA decay factor DHX34, as evidenced by Cryo-EM. ELife 9, e63042 (2020).
Martino, F. et al. RPAP3 provides a flexible scaffold for coupling HSP90 to the human R2TP co-chaperone complex. Nat. Commun. 9, 1501 (2018).
Tian, S. et al. Pih1p-Tah1p Puts a Lid on Hexameric AAA+ ATPases Rvb1/2p. Structure 25, 1519–1529 (2017).
Rivera-Calzada, A. et al. The structure of the R2TP complex defines a platform for recruiting diverse client proteins to the HSP90 molecular chaperone system. Structure https://doi.org/10.1016/j.str.2017.05.016 (2017).
Chagot, M. E. et al. Binding properties of the quaternary assembly protein SPAG1. Biochem J. 476, 1679–1694 (2019).
Mézard, C., Davies, A. A., Stasiak, A. & West, S. C. Biochemical properties of RuvBD113N: a mutation in helicase motif II of the RuvB hexamer affects DNA binding and ATPase activities1. J. Mol. Biol. 271, 704–717 (1997).
Lakomek, K., Stoehr, G., Tosi, A., Schmailzl, M. & Hopfner, K. P. Structural basis for dodecameric assembly states and conformational plasticity of the full-length AAA+ ATPases Rvb1. Rvb2. Structure 23, 483–495 (2015).
Paiva, A. C. F. et al. Extract2Chip—bypassing protein purification in drug discovery using surface plasmon resonance. Biosensors 13, 913 (2023).
Scheres, S. H. W. A Bayesian view on cryo-EM structure determination. J Mol. Biol. 415, 406–418 (2012).
Zhong, E. D., Bepler, T., Berger, B. & Davis, J. H. CryoDRGN: reconstruction of heterogeneous cryo-EM structures using neural networks. Nat. Methods 18, 176–185 (2021).
Lopez-Perrote, A., Munoz-Hernandez, H., Gil, D. & Llorca, O. Conformational transitions regulate the exposure of a DNA-binding domain in the RuvBL1-RuvBL2 complex. Nucleic Acids Res. 40, 11086–11099 (2012).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Yenerall, P. et al. RUVBL1/RUVBL2 ATPase Activity drives PAQosome maturation, DNA replication and radioresistance in lung cancer. Cell Chem. Biol. 27, 105–121.e14 (2020).
Horejsi, Z. et al. Phosphorylation-dependent PIH1D1 interactions define substrate specificity of the R2TP cochaperone complex. Cell Rep. 7, 19–26 (2014).
Dermouche, S., Chagot, M.-E., Manival, X. & Quinternet, M. Optimizing the first TPR domain of the human SPAG1 protein provides insight into the HSP70 and HSP90 binding properties. Biochemistry https://doi.org/10.1021/acs.biochem.1c00052 (2021).
Keller, R. L. J.Optimizing the Process of Nuclear Magnetic Resonance Spectrum Analysis and Computer-Aided Resonance Assignment. (ETH Zurich, 2005).
Schrodinger, L.The PyMOL Molecular Graphics System, Version 1 (2015).
R. Core Team. R: A Language and Environment for Statistical Computing.R Foundation for Statistical Computing, Vienna, Austria. (2025).
Briggs, L. C. et al. Analysis of nucleotide binding to P97 reveals the properties of a tandem AAA hexameric ATPase *. J. Biol. Chem. 283, 13745–13752 (2008).
Combe, C. W., Graham, M., Kolbowski, L., Fischer, L. & Rappsilber, J. xiVIEW: Visualisation of crosslinking mass spectrometry data. J. Mol. Biol. 436, 168656 (2024).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. ELife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Gorynia, S. et al. Structural and functional insights into a dodecameric molecular machine - the RuvBL1/RuvBL2 complex. J. Struct. Biol. 176, 279–291 (2011).
Pettersen, E. F. et al. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Cryst. D 71, 136–153 (2015).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Cryst. D 67, 355–367 (2011).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Cryst. D 66, 12–21 (2010).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Acknowledgements
The authors would like to thank S. Balor and V. Soldan (METI platform, Toulouse, member of the national infrastructure France-BioImaging supported by the French National Research Agency (ANR-10-INBS-04)), for technical support in electron microscopy, and the engineers from the CALMIP HPC facility. We thank the Biophysics and Structural Biology (B2S) platform of SMP (“Service Mutualisé de Plateformes, Université de Lorraine”, https://umsibslor.univ-lorraine.fr/en/facility/biophysics-structural-biology-b2s) for access to the NMR and SLS core facilities. We also acknowledge editing services from Dr. M. Kostic (Life Science Editors). This work was supported by the CNRS, the University of Toulouse, the University of Strasbourg, “l’Université de Lorraine”, the “Agence Nationale de la Recherche” (ANR-23-CE12-0022 YMCR; ANR-23-CE44-0035 NAProt-XLMS) and the French Proteomics Infrastructure (ProFI; ANR-10-INBS-08-03 and ANR-24-INBS-0015). H.G.F., E.D., and M.L. acknowledge the French Ministry for Education and Research for funding of their PhD. This work was also funded by “Fundação para a Ciência e Tecnologia/Ministério da Ciência, Tecnologia e Ensino Superior” (FCT/MCTES, Portugal) through national funds to iNOVA4Health (UIDB/04462/2020 and UIDP/04462/2020) and the Associate Laboratory LS4FUTURE (LA/P/0087/2020). This work was partly performed using HPC resources from CALMIP (Grant 2014-P1406).
Author information
Authors and Affiliations
Contributions
P.E.S., M.E.C., H.G.F., M.L., T.C., E.D., L.P., and A.C.F.P. performed the experiments. P.E.S., M.E.C., H.G.F., P.M.F.S., C.V., T.M.B., P.M., S.C., X.M., C.P.C., and M.Q. conceived the study. P.M.F.S., E.B., B.C., C.V., T.M.B., P.M., S.C., X.M., C.P.C., and M.Q. supervised the work. P.E.S., M.E.C., H.G.F., A.C.F.P., P.M.F.S., C.V., M.Q., T.M.B., P.M., S.C., C.P.C., and X.M. wrote the manuscript. All co-authors proofread the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Santo, P.E., Chagot, ME., Gizardin-Fredon, H. et al. An integrative structural biology approach reveals the dynamic organization of the R2SP quaternary chaperone complex. Nat Commun 17, 2605 (2026). https://doi.org/10.1038/s41467-026-69157-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69157-x