Abstract
Pyroptosis plays a crucial role in immune defense against infections and endogenous threats by eliminating harmful cells and modulating the immune response through inflammation. However, the natural activation of pyroptosis involves intricate signaling pathways, posing significant challenges for its artificial manipulation in research and therapies. Here, we present DAMAGE (Death Manipulation Gene), an innovative system that integrates gasdermins within the type III-E CRISPR framework, enabling the specific recognition of target RNA (tgRNA) and triggering pyroptosis. This mechanism allows DAMAGE to selectively identify and eliminate virus-infected, cancerous, and senescent cells, all of which exhibit altered RNA transcriptomes. Additionally, DAMAGE exhibits considerable promise as a platform for mRNA-LNP therapy. Our study highlights the potential of this CRISPR-based system in the controllable induction of pyroptosis, offering an innovative therapeutic strategy for treating RNA-heterogeneous diseases.
Similar content being viewed by others
Introduction
Pyroptosis is a lytic cell death induced by pathogen infection or endogenous challenge, playing a pivotal role in immune defense1,2,3. This process is primarily executed by members of the gasdermin family proteins (GSDMs), which include GSDMA, GSDMB, GSDMC, GSDMD, GSDME, and DFNB594. All GSDMs except DFNB59 possess a conserved two-domain structure: the N-terminal domain (NTD) interacts with the cell membrane to oligomerize and form pores, leading to membrane lysis and pyroptosis, while the C-terminal domain (CTD) inhibits NTD activity5,6,7,8. Pyroptosis is characterized as a “hot” reaction in immunology, eliciting a robust inflammatory response by releasing inflammatory cytokines and danger-associated molecular patterns (DAMPs), effectively activating the immune system4,9,10,11. Consequently, several studies have explored the therapeutic potential of pyroptosis in cancer treatment, yielding encouraging results11,12,13. However, the intricate signaling pathways involved in pyroptosis complicate efforts to induce this process through natural signaling interventions14,15,16. To tackle this challenge, we strive to develop a synthetic biology system that can respond to specific signals and controllably induce pyroptosis.
To counteract exogenous genetic elements, prokaryotes possess a wide range of defense mechanisms, among which the CRISPR-Cas system is especially prominent17,18. Although the CRISPR-Cas system primarily functions in adaptive immunity through RNA-guided cleavage of nucleic acids, growing research attention has shifted toward auxiliary proteins encoded adjacent to CRISPR loci—termed the CRISPR-associated protease (CASP) system19,20. Recent studies have uncovered complex interactions among diverse proteins within the type III-E CASP system, establishing a foundation for its practical application. The type III-E effector Cas7-11 (also known as gRAMP) is a single-protein effector composed of four Cas7 domains and one Cas11 domain21,22. Cas7-11 specifically recognizes tgRNA that is complementary to CRISPR RNA (crRNA; CR), subsequently activating the activity of a protease from the CHAT family of tetrapeptide repeats (Csx29, or TPR-CHAT)23,24,25,26,27. The activated Csx29 specifically cleaves the protein Csx30 at amino acids 427 to 429, facilitating the signal transduction from RNA to protease23,25,26.
In view of this signal conversion mechanism, we hypothesize that Csx30 can serve as a linker between the NTD of GSDMs (GSDMs-N) and the CTD of GSDMs (GSDMs-C) to construct a fusion protein, named GSDMs-Csx30. The Cas7-11–crRNA complex identifies tgRNA and subsequently activates Csx29. This activation leads to the cleavage of GSDMs-Csx30 by Csx29, resulting in the release of GSDMs-N and ultimately triggering pyroptosis. Herein, we introduce an innovative system designed to detect tgRNA and actively induce pyroptosis, referred to as DAMAGE (Death Manipulation Gene). DAMAGE specifically targets and eliminates various transcriptome-altered cells, such as virus-infected, cancerous, and senescent cells. Furthermore, DAMAGE demonstrates the potential to be transcribed into mRNA and delivered in vivo via Lipid Nanoparticles (LNPs) for therapeutic applications. Overall, our study establishes an ingenious method for targeting RNA-heterogeneous diseases, which holds great promise as an innovative strategy for their treatment.
Results
The construction and functional validation of DAMAGE
Type III-E CASP system contains a variety of proteins, including Cas7-11, Csx29, Csx30, Csx31, and RpoE21,22,23,24,25,26,27. We constructed expression plasmids derived from the type III-E CASP loci of Desulfonema ishimotonii, including Cas7-11, Csx29, and Csx30. Additionally, a nuclease-inactive Cas7-11 variant (dCas7-11; D429A/D654A) and a protease-inactive Csx29 variant (dCsx29; H615A/C658A) (Supplementary Fig. 1a) were also generated. The interactions among Cas7-11, Csx29, and Csx30 indicated a significant functional interrelationship between these proteins (Supplementary Fig. 1b). The predicted structure generated by AlphaFold2 reveals that Csx30 comprises an NTD (amino acids 1 to 377) and a CTD (amino acids 419 to 565), which are connected by a long, flexible linker region (amino acids 378 to 418) (Supplementary Fig. 1c). Based on these findings, we performed a series of truncation assays and found that the CTD of Csx30 alone is sufficient for cleavage by activated Csx29 (Supplementary Fig. 1d–h). These results support a role for Csx30 as a linker within an “X-Csx30-Y” architecture, enabling the fusion of two distinct protein domains and their integration into the type III-E CRISPR system.
Consequently, we developed the DAMAGE (Death Manipulation Gene) system (Fig. 1a), which is designed to respond to specific tgRNA and induce pyroptosis. It consists of five essential components: Cas7-11, Csx29, GSDMs-Csx30, crRNA, and tgRNA, each of which plays a critical role in the functionality of the system. The core of DAMAGE lies in the GSDMs-Csx30 effector, wherein Csx30 functions as a linker bridging GSDMs-N and GSDMs-C. As GSDMD is one of the most extensively studied members of the GSDMs, we aligned amino acid sequences of murine GSDMD (mGSDMD) and human GSDMD (hGSDMD), and created a series of truncated GSDMD-Csx30 variants (Supplementary Fig. 2a) to determine the optimal length of the Csx30 linker. Through a series of screenings, D-X demonstrated tightly regulated pyroptotic activity upon tgRNA recognition and was identified as the optimal fusion protein (Supplementary Fig. 2b–f). Furthermore, shorter truncated forms of GSDMD-Csx30 induced irregular and unpredictable nonspecific pyroptosis (Supplementary Fig. 2g, h). These findings indicate that Csx30 407-565 (amino acids 407 to 565) may serve as the most suitable linker.
a Schematic of the DAMAGE (Death Manipulation Gene) system. All GSDMs-Csx30 effectors, except D-Y (Csx30 397-565), use Csx30 407-565 as the linker. b LDH release assay of all GSDMs-Csx30 effectors. OFF: dCas7-11, Csx29, GSDMs-Csx30, EGFP, and CR-NT; ON: replace CR-NT with EGFP-CR-Mix; N+: positive control, replace GSDMs-Csx30 with GSDMs-Csx30-N in the ON groups. Heat map of LDH release, labeled with the mean of biological replicates (n = 6). c Flow cytometry (FCM) analysis of all GSDMs-Csx30 effectors. Left Y axis (EGFP+ (%)): EGFP-positive non-pyroptotic live cells; Right Y axis (PI+ (%)): PI-stained pyroptotic dead cells. PI: propidium iodide. d, ATP-based cell viability assay of all GSDMs-Csx30 effectors. OFF: dark green; ON: medium green; N+: light green; ctrl (-): negative control, treated only with transfection reagent; ctrl (+): positive control, transfected with GSDMD-N. e Cell imaging to assess the pyroptotic activity of C-FL in HEK293T cells. White arrows indicate pyroptotic cells. Scale bar, 200 μm, zoom inset, 50 μm. f Immunofluorescence imaging to evaluate the pyroptotic activity of A-FL in HeLa cells. Cells were stained with DAPI for nuclei and incubated with anti-Flag antibody for the detection of Flag-tagged A-FL effector. Scale bar, 10 μm. g Co-culture experiment validating the specificity of DAMAGE in eliminating target cells. Cells expressing EGFP and mCherry were co-transfected with DAMAGE targeting EGFP mRNA (DAMAGE-EGFP). Scale bar, 200 μm. All experiments were repeated at least three times (b–g). Data are means ± s.d. (c, d: n = 3). Source data are provided as a Source Data file.
To develop additional GSDMs-Csx30 effectors, we performed a comparative analysis of amino acid sequences among various members of the GSDMs family. The results showed significant homology between GSDMs-N and GSDMs-C, while the linker region exhibited lower similarity (Supplementary Fig. 4a). We therefore inserted the Csx30 407-565 segment into the natural cleavage sites of GSDMs to create the full-length GSDMs-Csx30, termed GSDMs-Csx30-FL (A-FL to E-FL). GSDMs were further truncated at their conserved amino acid sites, and the linker region was replaced with Csx30 407-565 to generate the short-length GSDMs-Csx30, designated as GSDMs-Csx30-SL (A-SL to E-SL). Furthermore, the previously designed constructs D-X and D-Y were also included, incorporating an additional nuclear export signal (NES) sequence upstream of GSDMD-N, thereby distinguishing them from D-SL and D-FL. Meanwhile, we constructed a plasmid expressing GSDMs-Csx30-N (N) as the positive control to mimic the cleavage product of GSDMs-Csx30 generated by activated Csx29 (Fig. 1b–e). We found that the loss of nuclease activity in Cas7-11 had no significant effect on the activation of the DAMAGE system (Supplementary Fig. 1e, 2b). Therefore, to eliminate the possible interference caused by nuclease activity, this study mainly used dCas7-11 for subsequent experiments.
We initially evaluated the functionality of the DAMAGE system by employing Enhanced Green Fluorescent Protein (EGFP) as the target. The control groups (OFF) were transfected with a non-targeting crRNA (CR-NT), whereas the trigger groups (ON) were treated with a crRNA mixture (CR-Mix) specifically targeting EGFP mRNA (Supplementary Fig. 3a–c and Supplementary Table 1). Compared to the OFF groups, GSDMs-Csx30 in the ON groups exhibited significant cleavage and subsequent induction of pyroptosis. In these ON groups, cell membrane lysis released intracellular contents into the supernatant, leading to decreased levels of Cas7-11, Csx29, and GSDMs-Csx30 in the cell lysate, whereas the cleavage-generated GSDMs-Csx30-N could be detected (Supplementary Fig. 4b, c). Following the release of cellular contents, the full-length C-terminal Flag-tagged GSDMs-Csx30 and the cleaved GSDMs-Csx30-C, were pulled down from the culture medium via immunoprecipitation (IP) by Flag-tagged beads (Supplementary Fig. 4c). Compared to the OFF group, the two forms above showed a significantly increase in the supernatant of the ON group, especially pronounced in A-SL and A-FL. The results demonstrate that the ON group displayed significantly distinct trends compared to the OFF group, with performance closely aligning with the N+ positive control. This was reflected in elevated lactate dehydrogenase (LDH) release (Fig. 1b), a reduction in the EGFP+ live cell population (Fig. 1c) accompanied by a rise in Propidium Iodide (PI) positive dead cells (PI+), and diminished cell viability (Fig. 1d and Supplementary Fig. 12a). The ON group also displayed distinct morphological features of pyroptosis, such as cellular swelling, plasma membrane blebbing, rupture, and the release of intracellular contents (Fig. 1e, f and Supplementary Fig. 5a, b). All these results demonstrated the occurrence of pyroptosis upon triggering of DAMAGE. To further evaluate the specificity of DAMAGE, we co-cultured EGFP- and mCherry-expressing cells and transfected them with DAMAGE targeting EGFP mRNA (DAMAGE-EGFP). The results demonstrated a significant reduction in green fluorescence, while red fluorescence remained unchanged (Fig. 1g). All these findings indicated that DAMAGE can selectively eliminate target cells without affecting non-target cells.
In conclusion, the DAMAGE system demonstrates a robust capacity to respond to tgRNA and induce highly specific pyroptosis in target cells. These findings indicate that the DAMAGE system drives tgRNA-dependent pyroptosis activation for achieving clearance of targeted cell populations.
DAMAGE responds to RSV infection and induces pyroptosis
When viruses invade host cells, they produce exogenous RNA that is markedly distinct from the host transcriptome. This discrepancy offers a promising opportunity for DAMAGE to selectively target and eliminate virus-infected cells. In this study, we utilized human respiratory syncytial virus (RSV) as a model virus, which is a negative-sense, single-stranded RNA virus that belongs to the Pneumoviridae family28. The RSV genome contains 10 genes that encode 11 proteins. The mRNAs encoding these proteins are sequentially transcribed from the non-segmented genome, resulting in a progressive decline in transcriptional efficiency.
Consequently, we selected the RSV nucleocapsid protein (RSV-N) for its high transcription level as the tgRNA to detect RSV, and developed DAMAGE-RSV to specifically target and eliminate RSV-infected cells (Fig. 2a). Through a preliminary and relatively comprehensive screening, the effectors GSDMB/C-Csx30 (B-FL, B-SL, C-FL, C-SL) demonstrated superior lethality, consistent with the previous findings, and B-SL was selected for subsequent assay due to its low nonspecific pyroptotic activity (Fig. 2b and Supplementary Fig. 6a). In both plasmid transfection and viral infection, DAMAGE-RSV was activated upon detection of RSV-N mRNA, resulting in efficient cleavage of GSDMs-Csx30 effectors (Supplementary Fig. 6c) and the occurrence of pyroptosis (Fig. 2c) along with obvious morphological changes (Fig. 2e). The tagged proteins, including dCas7-11-HA, Csx29-Myc, RSV-N-Myc, GSDMs-Csx30-Flag, and the cleaved GSDMs-Csx30-C-Flag, were successfully pulled down from the culture medium (Fig. 2d and Supplementary Fig. 6d, e). Furthermore, a reduction in cell viability was observed in the ON groups (Supplementary Fig. 6b and 12b), which provides additional evidence supporting the efficacy of DAMAGE. These results indicate that in the presence of RSV-N mRNA, whether transcribed from either exogenous transfection or viral infection, can be detected by DAMAGE-RSV to induce efficient pyroptosis, leading to cellular membrane lysis and the release of intracellular components.
a Schematic of DAMAGE-RSV. b The pyroptotic activity of DAMAGE-RSV. In HEK293T cells, RSV-N as tgRNA. OFF: dCas7-11, Csx29, GSDMs-Csx30, RSV-N CR-Mix and Vector; ON: replace Vector with RSV-N; N+: positive control, replace GSDMs-Csx30 with GSDMs-Csx30-N in the ON groups. The p values between the ON group and the OFF group of each GSDMs-Csx30 are labeled in the figure. c LDH release assay of DAMAGE-RSV in HeLa cells. d: dCas7-11 or dCsx29. d Immunoblotting and immunoprecipitation (IP) of supernatants to validate the pyroptosis-targeting activity of DAMAGE-RSV. Left bands: cell-lysate; right bands: supernatant-IP. e Immunofluorescence analysis of DAMAGE-RSV in HeLa cells. OFF: uninfected; ON: RSV-infected. RSV-N antibody indicates RSV infection; Flag-tag antibody indicates B-SL. Scale bar, 10 μm. f, g Gradient analysis of tgRNA in DAMAGE-RSV. Left: FCM assay, “-” represents PI-negative cells; “+” represents PI-positive cells; right: LDH release assay. RSV-N plasmid gradient (f). RSV multiplicity of infection (MOI) gradient (g). h LDH release assay of DAMAGE-RSV in A549 cells. Left: RSV-N plasmid transfection; right: RSV infection. i Cell imaging of DAMAGE-RSV in A549 cells. White arrows indicate pyroptotic cells. Scale bar, 50 μm. B-SL as effector, RSV-N as tgRNA (c–i). All experiments were repeated at least three times (b–i). Data are means ± s.e.m. (b: n = 3) or means ± s.d. (c, f–h: n = 3). p values were calculated by two-way ANOVA with Tukey’s multiple comparisons test (b), two-way ANOVA with Dunnett’s multiple comparisons test (c) and two-tailed unpaired t-test (h). Source data are provided as a Source Data file.
To further assess the sensitivity of DAMAGE-RSV, we conducted a series of gradient assays using negative quadratic dilution under both exogenous transfection and viral infection conditions. We observed that as the dilution factor increased, the pyroptotic activity of GSDMs-Csx30 progressively diminished. The cleavage efficiency of GSDMs-Csx30 decreased in response to reduced levels of tgRNA (Supplementary Fig. 7a–d), leading to a gradual decline in the PI positivity (Fig. 2f, g and Supplementary Fig. 6f, g) and a corresponding reduction in LDH release (Fig. 2f, g and Supplementary Fig. 6h, i). These findings highlight the remarkable sensitivity of DAMAGE in detecting minimal levels of tgRNA and its efficacy in triggering pyroptosis.
Since RSV predominantly infects respiratory epithelial cells, A549 cells, a human lung adenocarcinoma epithelial cell line, are extensively employed as a well-established in vitro model for the study of RSV infection. We performed transfection and infection experiments in A549 cells and observed consistent experimental outcomes (Fig. 2h). At the morphological level, the DAMAGE system successfully converts RSV-induced apoptosis into pyroptosis, thereby achieving efficient clearance of infected cells (Fig. 2i).
These findings demonstrate that DAMAGE can accurately identify “non-self” tgRNA originating from viruses or other pathogens and eliminate infected cells without adversely impacting normal cells. Moreover, DAMAGE exhibited high sensitivity in identifying these RNA signatures. This discovery provides pioneering strategies for combating viral infections by distinguishing the unique RNA profiles of viruses.
DAMAGE selectively targets and eliminates HPV-infected cervical cancer cells
Persistent infection with high-risk human papillomavirus (HPV) is recognized as the primary causative factor in the development of cervical cancer29,30. The integration of HPV DNA into the host genome results in the persistent expression of viral oncoproteins, particularly E6 and E7, making them pivotal targets for therapeutic vaccine development against HPV-associated malignancies31. Notably, the HeLa cell line, one of the most extensively utilized models in scientific research, originates from cervical cancer cells caused by HPV18 infection and accompanied by subsequent genomic integration. Therefore, we selected HeLa cells as a model system to examine the specific cytotoxic effects of DAMAGE on cervical cancer.
The validation experiments were initially conducted in HEK293T cells to evaluate the ability of DAMAGE to specifically target and recognize the E6 and E7 mRNA transcripts of HPV16/HPV18. The results indicated that the DAMAGE system can be specifically activated upon recognition of HPV mRNA (Supplementary Fig. 8a–c), leading to the induction of pyroptosis and a significant decrease in cell viability (Supplementary Fig. 12e-h). Consequently, we have developed DAMAGE-HPV, a system specifically designed to target and eliminate HPV infected cells. Through initial validation using individual crRNAs (CR1 to CR5) in HeLa cells, we found that endogenous HPV18-E6/E7 mRNA, transcribed from the HeLa genome, is sufficient to activate DAMAGE-HPV and induce pyroptosis (Fig. 3b). To further enhance the targeting capability of crRNA for HPV RNAs, we adopted a previously established strategy (Supplementary Fig. 3c) by fusing five individual target sites for each protein, constructing CR-E6, CR-E7, as well as a CR-E6+E7 combining the effective target sites from both E6 and E7. These constructs were used for subsequent experiments. We next evaluated the endogenous triggering-induced lethality of all effectors in HeLa cells (Supplementary Fig. 8d–g). The fluorescence intensity of EGFP and cell viability were assessed following co-transfection of cells with EGFP and DAMAGE-HPV to evaluate the extent of pyroptosis (Fig. 3c, d and Supplementary Fig. 8i). Consistent with previous findings in HEK293T cells, B-FL and C-FL exhibited the highest cytotoxicity in HeLa cells and were therefore selected as the primary effectors for further investigation.
a Schematic of DAMAGE-HPV. b Screening of crRNAs targeting HPV18-E6/E7 mRNA from the HeLa genome. Mock: CR-NT. The Flag-tag antibody detects D-X; endogenous GSDMD antibody detects D-X-N. c, d Assessment of pyroptotic activity mediated by various GSDMs-Csx30 effectors in DAMAGE-HPV. OFF: dCas7-11, Csx29, GSDMs-Csx30 and CR-NT; ON: replace CR-NT with HPV18 CR-Mix. Quantitative assays of EGFP green fluorescence intensity (c) and ATP-based cell viability assays (d) to evaluate cell viability. The p values for each group under ON and OFF conditions are shown in the figure. e–g Time gradient analysis of DAMAGE-HPV. FCM analysis (e). LDH release assay. ctrl (-): negative control; ctrl (+): positive control, transfected with GSDMD-Csx30-N. p values indicate the statistical differences between each group and the control group (ctrl (-)) at the 48 h time point (f). Immunoblot analysis (g). h Cell imaging of DAMAGE-HPV in HeLa cells. Pyroptotic cells are indicated by white arrows. Scale bar, 100 μm, zoom inset, 25 μm. i Cell imaging of DAMAGE-HPV in HeLa and C33-A cells. Scale bar, 100 μm. All experiments were repeated at least three times, in HeLa cells or C33-A cells, endogenous HPV18-E6/E7 mRNA as tgRNA (b–i), B-FL as the effector (e–i). Data are means ± s.d. (c, d: n = 3) and means ± s.e.m. (f: n = 3). p values were calculated by two-way ANOVA with Sidak’s multiple comparisons test (c, d) and two-way ANOVA with Dunnett’s multiple comparisons test (f). Source data are provided as a Source Data file.
With increasing transfection duration, both the PI-positive rate and LDH release levels gradually increased, peaking at 36 h post-transfection of DAMAGE-HPV (Fig. 3e–g). To visualize the effects of DAMAGE-HPV, we transfected HeLa cells with DAMAGE-HPV and co-transfected EGFP as a fluorescent marker to observe the morphological changes. Compared to the OFF groups (CR-NT), all three ON groups (CR-E6, CR-E7, and CR-E6+E7) significantly induced pyroptosis, along with a notable decrease in green fluorescence (Fig. 3h). To further investigate the recognition specificity of DAMAGE-HPV under endogenous activation conditions, we selected the HPV18-negative cervical cancer cell line C33-A as a negative control. In the absence of endogenous HPV18 mRNA, DAMAGE-HPV remains inactive in C33-A cells, preserving their viability and structural integrity. In contrast, HPV18-positive HeLa cells display pronounced pyroptotic morphological changes, accompanied by a significant decrease in fluorescence intensity (Fig. 3i).
These findings demonstrate that DAMAGE can specifically sense endogenous viral RNA transcribed by integrated viral genomic and induce pyroptosis in infected cells. This suggests that DAMAGE holds promise as a therapeutic strategy not only for acute infections like RSV, but also for chronic viral infections, such as HPV, HBV, and HIV.
DAMAGE identifies single nucleotide mutations and specifically triggers pyroptosis in KRAS-mutant cancer cells
KRAS is one of the most prevalent oncogenes, with mutations present in ~10% of all cancers32. Several mutations have been identified at the 12th amino acid Gly in KRAS, including G12C, G12D, G12R, G12A, G12S, and G12V33,34,35. Despite being identified more than four decades ago, it is only recently that inhibitors like sotorasib, which is a covalent binder of the KRAS-G12C mutation, have been approved for clinical application36.
Notably, significant base mutations exist in the mRNA between KRAS wild-type (KRAS-WT) and KRAS mutants (KRAS-mut), which facilitates the development of specifically targeted therapies. To evaluate the capability of DAMAGE in identifying single nucleotide mutations, we aligned the sequences of KRAS-WT and KRAS-mut. Two crRNAs were initially designed for each KRAS-mut variant, featuring spacers that perfectly match the KRAS-mut sequences while containing a single nucleotide mismatch relative to KRAS-WT (Supplementary Fig. 9a). We observed that the ON group transfected with KRAS-mut exhibited significant D-X cleavage, whereas the OFF group transfected with KRAS-WT showed negligible cleavage (Supplementary Fig. 9b), indicating that DAMAGE can discriminate single-nucleotide variants. Based on these findings, we have developed DAMAGE-KRAS, a targeted system for the detection of KRAS single nucleotide mutations.
To further enhance recognition accuracy, we focused on KRAS-G12C and designed 24 distinct crRNAs targeting KRAS-G12C by varying the position of the mutant nucleotide within the 24-bp spacer region of the crRNA (Fig. 4a). We systematically evaluated the capacity of these 24 crRNAs to detect single nucleotide mutations in KRAS-G12C. The results revealed that crRNA-16, crRNA-23, and crRNA-24 exhibited superior performance, with crRNA-23 displaying the highest specificity in discriminating KRAS-G12C from KRAS-WT (Fig. 4b–e and Supplementary Fig. 9c–g). To investigate the impact of endogenous KRAS mutations, HEK293T and HeLa cells stably expressing mCherry-tagged KRAS-WT or EGFP-tagged KRAS-G12C were subsequently constructed. In the presence of crRNA-23, DAMAGE-KRAS markedly induced pyroptosis in EGFP-KRAS-G12C stable cell lines, whereas no significant effect was observed in mCherry-KRAS-WT stable cell lines (Fig. 4f–h and Supplementary Fig. 9h). Time-dependent ATP-based cell viability assays further demonstrated that crRNA-16 and crRNA-23 significantly reduced cell viability in the KRAS-G12C groups compared to the KRAS-WT controls (Supplementary Fig. 12i, j). In addition, we systematically screened the six KRAS-mut subtypes and identified the optimal crRNAs to specifically target the mRNAs of these KRAS-mut (Fig. 4i and Supplementary Fig. 9i). Moreover, utilizing the NCI-H23 cells that endogenously harbor the KRAS-G12C mutation (Fig. 4j) and the A549 cells carrying the KRAS-G12S mutation (Fig. 4k), we demonstrated that endogenous KRAS-mut mRNA can effectively activate the DAMAGE-KRAS system, thereby indicating its potential applicability in clinical therapy.
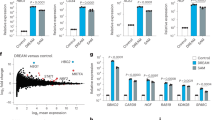
a Schematic of the design of KRAS-G12C crRNAs. Red lowercase indicates the mutated base. b, c Characterization of all KRAS-G12C crRNAs in DAMAGE-KRAS. OFF: KRAS-WT; ON: KRAS-G12C. p values for OFF vs. ON comparisons of CR-16, CR-23, and CR-24 were all less than 0.0001. PI staining and FCM analysis (b). LDH release assay, data are means with s.d., n = 6 (c). d FCM analysis of KRAS-G12C crRNA-23. PI+ pyroptotic cells were shown by contour plots in each group. e Immunoblotting assay of all KRAS-G12C crRNAs. ON groups showed significant GSDMs-Csx30 cleavage. f Cell imaging of DAMAGE-KRAS. In HEK293T stable cell lines, crRNA-23 to target KRAS-G12C mRNA. White arrows indicate pyroptotic cells. Scale bar, 25 μm. g LDH release assay of DAMAGE-KRAS in HEK293T stable cell lines. Mock: only treated with transfection reagent. h LDH release assay of DAMAGE-KRAS in HeLa stable cell lines. Mock: only treated with transfection reagent. i PI-positive cell statistics by FCM analysis (left) and LDH release rate (right) of the optimal crRNA for all KRAS-G12 mutant isoforms. j, k Characterization of DAMAGE-KRAS in cancer cell lines harboring primary KRAS mutations. NCI-H23: KRAS-G12C mutant (j). A549: KRAS-G12S mutant (k). All experiments were repeated at least three times, B-FL as the effector (b–k). Data are means ± s.d. (b, g–k: n = 3). p values were calculated by one-way ANOVA with Dunnett’s multiple comparisons test (g, h) or two-way ANOVA with Sidak’s multiple comparisons test (b, c, i) and two-tailed unpaired t-test (j, k). Heatmap data are presented as the mean of three independent biological replicates (b: n = 3). Source data are provided as a Source Data file.
Our findings demonstrate that the position of single nucleotide mutations within the crRNA spacer significantly influences the pyroptotic activity of DAMAGE. Following systematic optimization, DAMAGE achieves effective and precise detection of single nucleotide variants, underscoring its potential for therapeutic applications in a wide range of cancers driven by genetic alterations, including those involving BRAF, NRAS, and TP53. This category encompasses single- or multi-base substitutions, insertions or deletions, as well as chromosomal rearrangements.
DAMAGE targets p16/p21 mRNA and selectively eliminates senescent cells
Cellular senescence is a stress response that prevents the proliferation of aged or damaged cells. However, when these senescent cells accumulate in tissues over time, they may contribute to age-related decline and disease progression37. This process is triggered by the activation of retinoblastoma (RB) and p53 tumor suppressor networks, resulting in the transcription upregulation of cyclin-dependent kinase inhibitors (CDKIs), such as CDKN1A (p21CIP1; p21) and CDKN2A (p16INK4a; p16), which are commonly used as senescence markers38.
To selectively target senescent cells, we designed crRNAs for p16 and p21, including five individual crRNAs and a five-in-one crRNA-Mix, thereby establishing DAMAGE-Aging (Fig. 5a and Supplementary Table 1). Compared to the OFF groups (CR-NT), transient transfection of DAMAGE-Aging into HEK293T cells resulted in significant pyroptosis in the ON groups (p16/p21 CR-Mix), as evidenced by marked GSDMs-Csx30 cleavage, elevated LDH release, and an increased number of PI-labeled dead cells (Fig. 5b, c and Supplementary Fig. 10a, b). As the duration of transfection increased, the proportion of PI-positive dead cells rose progressively and cell viability declined in the ON groups, whereas no significant changes were observed in the OFF groups, even after 48 h (Supplementary Fig. 10c and 12c, d).
a Schematic of DAMAGE-Aging. b, c Assessment of pyroptotic activity mediated by various GSDMs-Csx30 effectors in DAMAGE-Aging. OFF: dCas7-11, Csx29, GSDMs-Csx30, p16/p21 plasmid and CR-NT; ON: replace CR-NT with p16/p21-CR-Mix; ctrl (-): negative control; ctrl (+): positive control, transfected with GSDMD-Csx30-N. Heatmap of PI positive rate (b). LDH release assay (c). d, e Activation of DAMAGE-Aging by endogenous p16/p21 mRNA. Co-transfected EGFP as a fluorescent marker. Cell imaging, scale bar, 200 μm (d). LDH release assay (e). f–i p16-KO blocks the endogenous activation of DAMAGE-Aging. Co-transfected EGFP as a fluorescent marker, p16-CR5 to detect endogenous p16 mRNA. Cell imaging, white arrows indicate pyroptotic cells, scale bar, 100 μm; zoom inset, 25 μm (f). FCM analysis (g). PI staining of pyroptotic cells (h). LDH release assay (i). j MS37452 treatment inhibits the endogenous activation of DAMAGE-Aging. Co-transfected EGFP as a fluorescent marker, EGFP fluorescence in the control group (DMSO, CR-NT) was defined as 100%, live cell proportion is indicated by EGFP+ (%). k Etoposide and doxorubicin treatment enhanced DAMAGE-Aging-induced pyroptosis. All experiments were repeated at least three times (b–k). In HEK293T cells, C-SL as effector, endogenous p16/p21 mRNA as tgRNA (d–k). Data are means ± s.d. (c, e, h–k: n = 3). p values were calculated by one-way ANOVA with Dunnett’s multiple comparisons test (e, j), two-way ANOVA with Tukey’s multiple comparisons test (h, i) or two-way ANOVA with Sidak’s multiple comparisons test (k). Heatmap data are presented as the mean of three independent biological replicates (b: n = 3). Source data are provided as a Source Data file.
However, it was unexpected to observe that the abundant endogenous mRNAs of p16 and p21 in HEK293T cells (Fig. 5d, e and Supplementary Fig. 10d) and HeLa cells (Supplementary Fig. 11a–c) were capable of activating DAMAGE-Aging and inducing pyroptosis, especially p16. Therefore, we established a HEK293T p16 knockout (HEK293T-p16-KO) cell line and successfully isolated a monoclonal cell line in which p16 mRNA was no longer detectable by p16 crRNA-5 (p16-CR5) (Supplementary Fig. 3a, b). Next, we introduced DAMAGE-Aging into HEK293T wild-type (HEK293T-WT) and HEK293T-p16-KO cells, along with EGFP as the fluorescent marker. In the ON groups (p16-CR5), HEK293T-WT cells exhibited a significant pyroptosis phenotype, marked by increased PI-positive dead cells and reduced green fluorescence. In contrast, HEK293T-p16-KO cells showed no significant pyroptotic response and resembled the OFF groups (CR-NT) (Fig. 5f–i). Consistent results were also observed in the HeLa-p16-KO cell line (Supplementary Fig. 11d, e). These results demonstrate that p16-KO effectively suppresses endogenous p16 mRNA-mediated DAMAGE-Aging activation, further confirming its target specificity.
To further assess the ability of DAMAGE-Aging to identify changes in RNA transcript levels, we modulated p16 and p21 mRNA translation using pharmacological agents. We found that MS37452, a small-molecule inhibitor of p16 mRNA translation, partially attenuated DAMAGE-Aging-induced pyroptosis, resulting in an increased proportion of EGFP-positive viable cells (Fig. 5j and Supplementary Fig. 10e). Furthermore, to inversely validate whether upregulation of p16/p21 mRNA could enhance the pyroptotic activity of DAMAGE-Aging, cells were treated with etoposide (2 μM) or doxorubicin (100 nM), both function as transcriptional activators of p16 and p21. These treatments significantly enhanced DAMAGE-Aging-induced pyroptosis, as indicated by increased PI-positive cells, elevated LDH release, and characteristic pyroptotic morphology (Fig. 5k and Supplementary Fig. 10f, 11g).
Based on the experimental results above and the gradient experiments illustrated in Fig. 2, we have confirmed that DAMAGE can effectively detect differences at the mRNA level. By specifically targeting highly expressed mRNAs in senescent cells, such as p16 and p21, DAMAGE-Aging shows a great potential for selectively eliminating senescent cells and offers an innovative strategy for targeted senotherapies.
The potential of DAMAGE in mRNA-LNP therapeutic applications
Given the proven efficacy of mRNA vaccines in combating the coronavirus disease 2019 (COVID-19) pandemic, this innovative platform has garnered considerable attention for its potential application in developing effective therapeutics for a range of infectious diseases and oncological conditions30,39,40. In this study, we attempt to transcribe DAMAGE into mRNA in vitro and deliver it utilizing Lipid Nanoparticles (LNPs) to explore its potential for mRNA-LNP therapeutic applications.
Originally, DAMAGE consisted of five components: Cas7-11, Csx29, GSDMs-Csx30, crRNA, and tgRNA, each integrated into separate plasmids. To simplify the DAMAGE system, we cloned Cas7-11, Csx29, and GSDMs-Csx30 into a single plasmid and renamed it DAMAGE-Plus (Fig. 6a). We initially transfected all the DAMAGE-Plus plasmid into HEK293T and HeLa cells. The results demonstrated that DAMAGE-Plus functioned in a similar trend to the original DAMAGE and even showed a lower level of nonspecific pyroptosis. Among them, DAMAGE-Plus B-FL performed the best and was chosen for subsequent experiments (Fig. 6b and Supplementary Fig. 13a). Consistent with prior findings, DAMAGE-Plus effectively targeted all previously investigated tgRNAs and induced significant pyroptosis (Supplementary Fig. 13b–d). Furthermore, DAMAGE-Plus can accurately detect tgRNAs, such as EGFP mRNA and HPV18-E7 mRNA, thereby specifically inducing pyroptosis in targeted cells without affecting non-target cells. This functional specificity has been confirmed through cell co-culture assays (Fig. 6c, d and Supplementary Fig. 13e, f) and live-cell imaging (Supplementary Fig. 15 and Supplementary Movie 1–6). We also verified the recognition capability for single-nucleotide mutation of DAMAGE-Plus in HEK293T-KRAS-G12C stable cell lines, which was consistent with the results presented in Fig. 4 (Supplementary Fig. 13g, h).
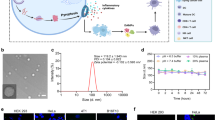
a Schematic of DAMAGE-Plus. DAMAGE-Plus mRNA encodes a large fusion protein that is self-cleaved into three independent proteins by T2A and P2A. IRES: Internal Ribosome Entry Site; P2A: Type 1 Porcine teschovirus 2A; T2A: Thosea asigna virus 2A; LNP: Lipid Nanoparticles. b FCM analysis of the DAMAGE-Plus for all GSDMs-Csx30. In HEK293T cells, EGFP as tgRNA, the gray dotted line shows the OFF group, the colored solid line shows the ON group. c, d Co-culture assay of HeLa-EGFP and HeLa-mCherry stable cell line. Endogenous EGFP mRNA or HPV18-E7 mRNA as tgRNA, GSDMD-Csx30-N as a positive control. Cell imaging, white arrows indicate pyroptotic cells, scale bar, 50 μm (c). LDH release assay (d). e, f Assessment of pyroptotic activity of DAMAGE-Plus in mRNA form in HEK293T cells; ctrl (-): negative control; ctrl (+): positive control, transfected with GSDMD-Csx30-N mRNA. p values between the ON group and the OFF group of each GSDMs-Csx30 were all less than 0.0001. FCM analysis (e); LDH release assay (f). g, h The pyroptotic activity of DAMAGE-Plus in mRNA form. In HeLa cells, endogenous HPV18-E7 mRNA as tgRNA. Cell imaging, scale bar, 25 µm (g). LDH release assay (h). i LDH release assay of DAMAGE-Plus in mRNA-LNP form. In HeLa cells, endogenous HPV18-E7 mRNA as tgRNA. All experiments were repeated at least three times (b–i), DAMAGE-Plus B-FL as effector (c–i). Data are means ± s.d. (d, f, h, i: n = 3) or violin plots with all individual data points (e: n = 3). p values were calculated by one-way ANOVA with Dunnett’s multiple comparisons test (d), two-way ANOVA with Sidak’s multiple comparisons test (e, f) and two-tailed unpaired t-test (h, i). Source data are provided as a Source Data file.
Next, we performed in vitro transcription of all components of DAMAGE-Plus into RNA, and then co-transfected these RNAs into HEK293T cells. In the ON groups, DAMAGE-Plus in RNA form demonstrated the ability to effectively recognize all tgRNAs and induce pyroptosis (Fig. 6e, f and Supplementary Fig. 14a, b). Furthermore, we validated whether DAMAGE could be delivered in the form of LNP-encapsulated mRNA (mRNA-LNP). Considering that an excessively large size of DAMAGE-Plus would reduce the efficiency of in vitro transcription and complicate LNP encapsulation. We divided DAMAGE-Plus into two components, Cas7-11 and Csx29-B-FL, which were transcribed separately in vitro (Supplementary Fig. 14c–e). Meanwhile, mRNA-LNP was synthesized to evaluate the therapeutic potential of DAMAGE (Supplementary Fig. 14f, g). In HEK293T cells, whether in the form of mRNA (Supplementary Fig. 14h) or mRNA-LNP (Supplementary Fig. 14i), DAMAGE was found to effectively recognize tgRNA and trigger pyroptosis upon activation. Furthermore, both mRNA transfection (Fig. 6g, h and Supplementary Fig. 14j) and mRNA-LNP delivery (Fig. 6i and Supplementary Fig. 14k) of the DAMAGE system in HeLa cells could also lead to specific recognition of endogenous HPV18-E7 mRNA and efficient induction of pyroptosis.
These findings demonstrate that DAMAGE can be transcribed into mRNA in vitro and delivered in the form of mRNA-LNP, underscoring its potential as an mRNA therapeutic for in vivo delivery and treatment, with promising prospects for translation into clinical application.
Discussion
Pyroptosis is a crucial mechanism in immune defense against infections and endogenous threats. This inflammatory form of programmed cell death can enhance the anti-tumor immune response by recruiting immune cells, including cytotoxic T lymphocytes. Recent studies have illuminated the potential applications of pyroptosis in cancer therapy. For instance, Wang et al. developed a bio-orthogonal chemical system for the selective delivery of GSDMD to tumor cells12. Zhang et al. demonstrated that tumor-derived GSDME functions as a tumor suppressor by activating pyroptosis and enhancing anti-tumor immunity13. Additionally, Xia et al. constructed a synthetic circuit by fusing the cleavage site of tobacco etch virus protease (TEVP) between GSDMs-N and GSDMs-C, enabling programmable control of mammalian cell death11. However, these methods pose certain challenges in clinical applications, encompassing opportunities for improvement in efficiency, specificity, and practicality, as well as aspects related to natural controllability that could benefit from further refinement.
To address these challenges, we have developed the DAMAGE system, which can precisely trigger pyroptosis in harmful target cells without affecting normal cells, featuring high specificity and controllability. The activation of DAMAGE occurs upon recognition of tgRNA, rendering it particularly suitable for the treatment of RNA-heterogeneous diseases, including those involving diverse RNA sequence types, base mutations, and transcriptional-level variations. In terms of RNA sequence types, upon viral infection of a host cell, the virus generates viral RNA that is distinct from the host transcriptome. The DAMAGE system can specifically induce pyroptosis in virus-infected cells by recognizing these exogenous “non-self” RNA. Owing to this high recognition specificity, the DAMAGE system holds significant potential for application in the treatment of a broad spectrum of viral infectious diseases, including acute viral infections (RSV, SARS-CoV-2 and DENV) and chronic viral infections (human papillomavirus, hepatitis B virus and HIV). In terms of RNA base mutations, the DAMAGE system demonstrates single-base recognition accuracy. Thus, we hypothesize that, given sufficient abundance of mutant RNA, DAMAGE can effectively target all disease types related to gene mutations, including single or multiple base mutations, gene insertions or deletions, and chromosomal alterations. Furthermore, the pyroptotic activity of DAMAGE can be activated to varying degrees depending on the levels of tgRNA, allowing for the specific targeting and elimination of pathological cells exhibiting significant transcriptomic alterations, such as senescent cells. In this study, we attempted to transcribe the DAMAGE system into mRNA in vitro and deliver it to cells via mRNA-LNP, thereby successfully inducing specific pyroptosis in target cells. However, the DAMAGE system consists of three proteins and two RNA components, and its multi-component architecture along with high molecular weight poses significant challenges to in vitro mRNA transcription efficiency and the packaging capacity of LNPs. As a result, the delivery efficiency of this system requires further validation in practical applications. Future efforts should focus on optimizing the expression design of key components or utilizing delivery carriers with enhanced loading capacity, which will be critical for advancing the clinical translation of this system.
With regard to the future development of the DAMAGE system, the primary focus will be on the Csx29 and Csx30 effectors. In this study, a total of 12 GSDMs-Csx30 effector constructs were designed. It is noteworthy that the GSDM-Csx30 effectors exhibit varying degrees of pyroptotic activity. Compared to the full-length GSDMs-Csx30-FL, the truncated GSDMs-Csx30-SL exhibits a reduced capacity to induce pyroptosis after cleavage, but also shows reduced nonspecific pyroptosis, particularly when comparing B-SL and B-FL. Among the 12 effectors tested, GSDMB-Csx30 and GSDMC-Csx30 exhibited the most potent lethal effects. The rank order of potency was B-FL > C-FL > C-SL > B-SL. However, increased lethality comes with higher nonspecific pyroptotic activity. Therefore, it is difficult to define the best effector in this study. Interestingly, D-SL and D-FL can induce a relatively strong nonspecific pyroptotic response, whereas D-X and D-Y, which were engineered with N-terminal NES sequences, exhibit markedly attenuated nonspecific pyroptosis, particularly D-X. The mechanism by which NES attenuates GSDMD toxicity may involve its effect on the GSDMD-N structure, thereby impairing its pore-forming activity. These phenomena suggest that appropriately weakening the ability of GSDMs-Csx30 to induce pyroptosis can significantly decrease the occurrence of nonspecific pyroptosis. Furthermore, our findings indicate that even in its inactive state, Csx29 has the potential to directly cleave GSDMs-Csx30, a phenomenon particularly evident with D-Y. Therefore, stringent regulation of Csx29 protease activity and further structural optimization of GSDMs-Csx30 may represent promising directions for future improvements to the DAMAGE system.
In broader applications, DAMAGE serves as a powerful research tool for the artificial regulation of pyroptosis. By regulating RNA transcription or the expression of proteins associated with any of the five components (for example, by integrating a tetracycline-inducible eukaryotic operon system), pyroptosis can be precisely initiated at specific time points and within particular cell types. In addition to the GSDMs family, the DAMAGE system holds potential for application in regulating other zymogens or signaling pathway proteins, such as members of the caspase family or transcription factors, enabling the construction of artificial RNA-responsive signaling pathways.
Although the DAMAGE system has demonstrated a specific cytotoxic effect on target cells across multiple cell lines, its efficacy and functional behavior have not yet been evaluated in more complex biological systems, such as in organoid or murine models. Therefore, the system is still in an early, exploratory phase of development. Additional in vivo studies are necessary to evaluate its potential and feasibility for future clinical application. In summary, DAMAGE represents a sophisticated circuit that transduces signals by converting RNA information to proteolytic activity, facilitating the cascading amplification of these signals. We maintain high expectations regarding the potential applications of DAMAGE, which may offer an innovative therapeutic strategy for a wide range of RNA-heterogeneous diseases.
Methods
Cells
The HEK293T (CRL-1573, ATCC), A549 (CCL-185, ATCC), HeLa and C33-A (obtained from Dr. Zheng Hu, Wuhan University) cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, 11995065, Gibco) supplemented with 10% fetal bovine serum (FBS, 086150, WISENT) and 1% Penicillin-Streptomycin (15140-122, Gibco). The NCI-H23 cells (GDC0651, CCTCC) were cultured in RPMI Medium 1640 basic (C11875500BT, Gibco) supplemented with 10% FBS and 1% Penicillin-Streptomycin. Cells used in experiments were from passages fewer than 30. All cell cultures were maintained at 37 °C in a humidified incubator with 5% CO2.
Plasmid construction
The Cas7-11, Csx29, and Csx30 genes were cloned from the DiCas7-11 full locus plasmid (172501, Addgene), and the fragments were ligated into restriction enzyme-digested expression vectors using T4 DNA ligase (RK21501, ABclonal). The crRNAs were designed using the Cas13design online platform, selecting the five top-ranked, non-overlapping complementary spacers (https://cas13design.nygenome.org/)41. For the construction of GSDMs-Csx30, gene fragments of all human GSDMs were obtained through gene synthesis (Tsingke Biotech). The GSDMs-Csx30 gene fragments were constructed by bridge PCR and cloned into the vector via homologous recombination by Hieff Clone Universal II One Step Cloning Kit (10923ES20, Yeasen). All inserted fragments have been validated through DNA sequencing to ensure their accuracy and reliability.
Plasmid transfection
HEK293T or HeLa cells were seeded in a 12-well plate, a 24-well plate or a 96-well plate and then incubated overnight to reach a density of approximately 70% before transfection. The five components of DAMAGE: dCas7-11, Csx29, GSDMs-Csx30, crRNA, and tgRNA were maintained at a mass ratio of 1:1:1:1:1. A total of 2 μg of plasmid was transfected into each well of a 12-well plate, 1 μg into each well of a 24-well plate and 0.5 μg into each well of a 96-well plate, with the same ratio maintained. When DAMAGE was constructed in a single plasmid, 1.2 μg DAMAGE-Plus plasmid, 0.4 μg tgRNA plasmid and 0.4 μg crRNA plasmid were transfected into each well of a 12-well plate. For endogenous trigger experiments, the tgRNA plasmid was omitted. For transfection of HEK293T or HeLa cells in a 12-well plate, all the plasmids were diluted in 50 μL Opti-MEM (31985-070, Gibco) for every 1 μg of plasmid transfection added with 1 μL of Neofect™ DNA transfection reagent (TF20121201, NEOFECT). For transfection of A549, NCI-H23 and C33-A cells, Lipofectamine™ 3,000 Transfection Reagent (L3000075, Invitrogen) was employed in accordance with the manufacturer’s instructions. Equal amounts of DNA were added to each well.
Virus infection
HEK293T, HeLa or A549 cells were seeded in a 12-well plate and incubated overnight until they reached a density of ~70% before viral infection. In the infection procedure, the culture medium was removed, and the cells were washed with PBS (SH30256.01, Cytiva) and then exposed to DMEM (11995065, Gibco) containing RSV (strain A2) for 2 h with different MOI values. Subsequently, the medium was replaced with fresh DMEM supplemented with 4% FBS, followed by transfection.
Cell co-culture assay
In the co-culture assay of transient transfection, HEK293T or HeLa cells were divided into two groups and seeded in a 12-well plate at the same density. After overnight culture, one group was transfected with mCherry, while the other was transfected with EGFP. Simultaneously, both groups were transfected with DAMAGE targeting EGFP mRNA. 8 h post transfection, the two groups of cells were mixed at a 1:1 ratio. The OFF groups transfected with CR-NT were employed as a negative control, as they did not trigger DAMAGE activation. For the stable cell lines, the same amount of HEK293T-mCherry-KRAS-WT and HEK293T-EGFP-KRAS-G12C, or HeLa-mCherry-KRAS-WT and HeLa-EGFP-KRAS-G12C, or HeLa-EGFP and HeLa-mCherry were collected and seeded in a 12-well plate. The cells were transfected with the corresponding plasmids after overnight culture.
mRNA production and transfection
DAMAGE-Plus was cloned into a DNA backbone featuring a T7 promoter, and the plasmid was linearized through restriction endonucleases after purification for mRNA production. Then, the mMESSAGE mMACHINE™ T7 ULTRA Transcription Kit (AM1345, Invitrogen) was employed to transcribe DAMAGE-Plus mRNA with a 5’ cap structure and a 3’ polyA tail in vitro. To generate RNA for tgRNA and crRNA, PCR was utilized to amplify them from plasmids, with a T7 promoter incorporated at the 5’ end. The resulting DNA amplicons were then subjected to in vitro transcription using the TranscriptAid T7 High Yield Transcription Kit (K0441, Invitrogen). Finally, all these RNA products were purified with the MEGAclear™ Transcription Clean-Up Kit (AM1908, Invitrogen), followed by transfection executed using Lipofectamine™ MessengerMAX™ Transfection Reagent (LMRNA008, Invitrogen) in accordance with the manufacturer’s instructions. The components: DAMAGE-Plus, crRNA and tgRNA were transfected at molar ratio of 1:2:1.
The preparation and delivery of mRNA-LNP
The two plasmids (Cas7-11 and Csx29-B-FL) were cloned into a DNA backbone featuring a T7 promoter respectively, serving as the template for in vitro RNA transcription. The mRNA-LNP synthesis was performed by GenScript Biotech Corporation with the molar ratio of Cas7-11: Csx29-B-FL: crRNA = 1:1:3, and the product was stored at −80 °C until used. For the delivery experiment in cells, HEK293T or HeLa cells were seeded in a 12-well plate and cultured overnight to the density of 60%–70%. Next, the frozen mRNA-LNP was allowed to thaw on ice for 10 min and mixed by gentle inversion 3–5 times (vortexing was avoided). The thawed LNPs was then diluted in Opti-MEM to a final concentration of 1 μg mRNA per well and added to the culture medium. The plate was rocked gently for even distribution of the LNPs, and then incubated for 24 h prior to analysis.
Cell line engineering
For the construction of stable expression or CRISPR/Cas9 knockout cell lines, lentiviral plasmids carrying the desired gene or sgRNA were initially transfected into HEK293T cells along with the packaging plasmids pSPAX2 (12260, Addgene) and pMD2G (12259, Addgene) at a ratio of 3:2:1 for lentiviral packaging. The supernatants were collected 48 h post-transfection and filtered through a 0.45-μm filter to remove cellular debris, and the filtered supernatant was used to infect the indicated cells for another 48 h. The stable expression cells were selected with blasticidin (3513-03-9, Yeasen), while p16/p21-KO cells were generated under puromycin selection (540411, Sigma-Aldrich). Fluorescence-tagged stable lines were subsequently validated by fluorescence microscopy, while knockout efficiency was confirmed by immunoblotting34.
Reagent treatment
For MS37452 (HY-119344, MCE) treatment, concentration gradients of 0 (DMSO), 25, 50, and 100 μM were established. After cells adhesion, the compound was added for a 12 h treatment period, after which transfection was performed. The experiment included the following treatment groups: a DMSO control, and groups treated with 25, 50, or 100 μM MS37452. Cells in each group were transfected with the DAMAGE system, using either the non‑targeting CR‑NT (OFF groups) or the p16‑targeting crRNAs (p16‑CR5 or p16‑CR‑Mix; ON groups) constructs. For flow cytometry data analysis, the EGFP+ population in the CR-NT (OFF) in each treatment was set as 100%. The percentage of live cells was then calculated as the ratio of the EGFP+ cells in the p16-CR5 or p16-CR-Mix (ON) group to that in its corresponding CR-NT (OFF) group.
In the Etoposide (HY-13629, MCE) and Doxorubicin (HY-15142, MCE) treatment, the cells were seeded in a 12-well plate, and transfection was performed after the overnight culture. 8 h post transfection, the supernatant was replaced with fresh medium containing Etoposide (2 μM) or Doxorubicin (100 nM), and an equal amount of DMSO was added as a control. The samples were collected for assay after 36 h of treatment.
Antibodies
The antibodies used in the study are listed as follows: DYKDDDDK Tag (D6W5B) Rabbit mAb (14793S, Cell Signaling Technology), DYKDDDDK Tag (9A3) Mouse mAb (8146S, Cell Signaling Technology), Anti-HA-tag Mouse mAb (M180-3, MBL), Anti-Myc-tag Mouse mAb (M192-3, MBL) were utilized to detect labeled proteins. Gasdermin A (49307S), Gasdermin B (76439S), Gasdermin C (61921S), Gasdermin D (39754S), Gasdermin E (19453S) were used for the direct detection of expressed GSDMs (all from Cell Signaling Technology). Mouse monoclonal anti-RSV Nucleoprotein (N) (ab94806, Abcam), Mouse monoclonal anti-RSV Phosphoprotein (P) (ab94965, Abcam) and Mouse monoclonal anti-RSV M2-1 (ab94805, Abcam) were used to detect viral proteins. p16 INK4A (E6N8P) Rabbit mAb (18769S, Cell Signaling Technology) and p21 Waf1/Cip1 (12D1) Rabbit mAb (2947S, Cell Signaling Technology) were utilized to detect endogenous p16/p21. α-Tubulin Mouse mAb (AC012, ABclonal) and GAPDH Mouse mAb (AC033, ABclonal) were employed as loading controls. Goat anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 488 (A-11029, Invitrogen); Goat anti-Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 594 (A-11012, Invitrogen); and Goat anti-Mouse IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 594 (A-11005, Invitrogen) were used for immunofluorescence assay in confocal microscopy.
Microscopy
Cells were seeded in a 12-well plate for transfection. After 24 h for HEK293T or 36 h for HeLa post-transfection, each well was stained with appropriate dye according to the recommended dilution ratio, conditions and time: PI (P3566, Invitrogen) at a ratio of 1:1000; Hoechst (C1029, Beyotime) at a ratio of 1:1000; or 2 drops of SYTOX Green (R37168, Invitrogen). Subsequently, cell morphology was observed and images were captured using a Leica DMi8 microscope. For confocal microscopy, cells were seeded in a 24-well plate with aseptic coverslips. 20 h post-transfection, the supernatant was removed and the cells were washed with PBS three times. The samples then fixed with 4% paraformaldehyde at room temperature for 20 min and permeabilized with 0.2% Triton X-100 (T109028, Aladdin) in PBS for 10 min. Subsequently, cells were blocked using a buffer containing 3% BSA (BioFrox, 4240GR500) for 30 min. They were then incubated with the primary antibody at 4 °C overnight and followed by a fluorescent labeled secondary antibody at room temperature for 1 h. The nuclei were stained for 10 min using DAPI (D9542, Sigma-Aldrich) at 50 μg/mL. After three 5-min PBS washes, the coverslips were inverted onto glass slides with ProLong Diamond Antifade Mountant (P36970, Invitrogen) and stayed overnight in the dark before observation. The images were captured using a Zeiss LSM 980 confocal microscope and analyzed by ZEN2.3, and the scale bar was added using ImageJ software (Experimental Methodology Reference Zhou, B. et al.)42.
Live cell imaging
Live cell imaging was performed on a Zeiss LSM 980 equipped with a stage-top incubator. The cells were cultured in a Φ20 mm TC-treated confocal dish (BS-20-GJM, Biosharp) for transfection. Live-cell imaging was initiated 12 h post-transfection, acquiring one frame every 5 min for a total of 100 frames. Each group was imaged under identical laser intensity settings. The movies were processed and laid out by ZEN2.3, and the scale bar was added using ImageJ43.
Cell stain and flow cytometry
Cells seeded in a 12-well plate and transfected with plasmid or mRNA for 24–36 h were stained and harvested for flow cytometry. For PI (P3566, Invitrogen) staining, the dye was added to the cell supernatant at 1:1000 dilution and incubated at 37 °C for 30 min. All cells were collected for analysis. Floating cells in supernatant were first transferred to a 1.5 mL Eppendorf tube and pelleted by centrifugation (200 × g, 5 min; Eppendorf 5424 R). The remaining adherent cells were trypsinized and combined with the pelleted floating cells. Then, all cells were harvested by centrifugation at 200 × g for 5 min. The pellet was resuspended in 1 mL PBS and analyzed using CytoFLEX flow cytometer (Beckman Coulter). Forward and side scatter were set as follows: 500/500 for HEK293T, 200/200 for HeLa, and 300/300 for A549 cells. Cellular fluorescence was detected using two main channels: FITC (for EGFP; excitation 488 nm, emission 525/40 nm; gain 5–10), PE (for PI, excitation 488 nm, emission 585/42 nm; gain 50–100) or PC5.5 channel (for PI, excitation 488 nm, emission 690/50; gain 50). For experiments involving two colors, single-color controls were used to establish the compensation matrix with the CytExpert software (Version 2.0.4.8). All the results were analyzed by FlowJo 10.10, using negative controls (unstained) to define positivity thresholds. The gray dashed line was employed to mark the fluorescence intensity threshold separating PI-negative viable cells (left) from PI-positive dead cells (right) in this study.
Cell viability and cytotoxicity experiments
LDH release was measured using the CytoTox 96® Non-Radioactive Cytotoxicity Assay (G1781, Promega) to evaluate the process of cell pyroptosis according to the manufacturer’s instructions. Cell culture supernatant was collected 24–48 h post transfection and centrifuged at 16000 × g for 5 min to remove cell debris. For the assay, 50 μL supernatant of each sample was added to the wells of a 96-well plate, followed by the addition of an equal volume of reaction mix. The plate was gently shaken to mix the components and incubated in the dark for 20–30 min. The reaction was stopped by adding 50 μL stop solution, and the absorbance at 490 nm was measured using a Multi-function microplate reader (Molecular Devices, SpectraMax iD3). Each sample did 3 parallels. The LDH release rate was calculated according to the manufacturer’s instructions. To detect the cell viability, the ATP-based viability assements were conducted using the CellTiter-Glo® Luminescent Cell Viability Assay (G7571, Promega) according to the manufacturer’s instructions. 13,000 HEK293T cells were seeded per well of a 96-well plate and transfected at approximately 60% confluency. Following 24 h of incubation, 100 μL assay reagent was added to an equal volume of culture medium in each well. Plates were mixed on the orbital shaker (80 cycles per minute, 10 min), incubated for 10 min at room temperature, and then the luminescence was recorded through a Multi-function microplate reader. ATP-based cell viability was calculated as the luminescence of the transfected wells normalized to that of un-transfected controls (Opti-MEM only). Each sample did 3 parallels.
RNA extraction, reverse transcription PCR, and real-time PCR
Total RNA was extracted using Trizol reagent (15596026CN, Invitrogen) in accordance with the manufacturer’s protocol. 1 μg mRNA was used for reverse transcription with the ABScript II cDNA First-Strand Synthesis Kit (RK20400, ABclonal) to obtain cDNA. Subsequently, the expression level of RSV-N mRNA after gradient RSV-N transfection or RSV infection was quantified by real-time PCR using 2× Universal SYBR Green Fast qPCR Mix (RK21203, ABclonal). Moreover, the expression level of the target gene was normalized to that of GAPDH as the housekeeping gene.
Immunoblotting and supernatant immunoprecipitation
To investigate the effect of DAMAGE, cells were lysed in lysis buffer (150 mM NaCl, 50 mM Tris-HCl pH 7.4, 1% Triton X-100, 1 mM EDTA pH 8.0, 0.1% SDS) containing protease inhibitor cocktail. Proteins in the cell lysates were separated via SDS-PAGE and analyzed through immunoblotting. Given that the Flag tag is fused to the C-terminus of effectors, the detection of N-terminal cleavage products requires the usage of endogenous antibodies. For comprehensive detection of all effectors and their N-terminal fragments, the endogenous antibody anti-GSDMA/B/C/D/E were diluted to the recommended dilution ratio. To ascertain whether the cellular contents were released into the supernatant upon cell rupture, culture supernatant was collected and incubated with tag-coupled beads overnight at 4 °C. The supernatant was subjected to three rounds of immunoprecipitation (IP). In the first round, the supernatant was incubated with Flag-tagged agarose beads (651503, Biolegend) overnight at 4 °C, followed by centrifugation at 6000 × g for 2 min. The supernatant was then transferred to a new tube, while the beads were washed three times with lysis buffer and subsequently denatured in 1× SDS-PAGE loading buffer for the immunoblotting assay. The supernatant underwent a second round of IP and was incubated with Myc-tagged magnetic beads (HY-K0206, MCE), following the same procedure as above. The third round of IP was performed with HA-tagged magnetic beads (HY-K0207, MCE), using identical steps.
Co-immunoprecipitation
The interaction between Cas7-11-HA, Csx29-Myc, and Csx30-Flag were investigated for three pairs: Cas7-11-HA and Csx29-Myc; Csx29-Myc and Csx30-Flag; Cas7-11-HA and Csx30-Flag. For the experiments of Cas7-11-HA and Csx29-Myc, cells in a 6-well plate were harvested and lysed 24 h post transfection. Lysates were incubated with HA-tagged beads at 4 °C overnight to immunoprecipitate HA-tagged Cas7-11. Interaction was assessed by detecting co-precipitated Csx29-Myc in the denatured beads via immunoblotting. To further verify the interaction, reciprocal co-immunoprecipitation was performed using Myc-tagged beads to pull down Csx29-Myc and detect associated Cas7-11-HA. Given the nuclease activity of Cas7-11 and the proteolytic activity of Csx29, corresponding inactive variants (dCas7-11 and dCsx29) were also included in the assays. The other two interaction pairs were assayed in the same manner.
Protein structure analysis
The structures of Csx30 was predicted by AlphaFold2. UCSF ChimeraX and PyMOL were employed to visualize the predicted outcomes44,45,46.
EGFP fluorescence intensity detection
To assess the pyroptotic capacity of various DAMAGE effectors for endogenous activation in HeLa cells, EGFP fluorescence intensity was quantitatively measured as a proxy for cell viability. Cells were seeded in a 96-well plate at 13,000 cells per well and cultured overnight to allow attachment. Upon reaching ~70% confluency, cells were co-transfected with DAMAGE-HPV and EGFP, using endogenous HPV18-E7 mRNA as the tgRNA. As cell death occurs, the fluorescence diminishes, accompanied by a decrease in EGFP intensity and cell viability. Therefore, EGFP fluorescence intensity can serve as an indicator of cell viability. The transfected cells were incubated for 24–36 h, and EGFP fluorescence intensity was detected using a Multi-function microplate reader. Wells transfected with EGFP alone served as the positive control (set as 100%), while wells treated with Opti-MEM only were used as the negative control. The relative EGFP intensity of each well was calculated by the formula: Y% = (X − negative control) / (positive control − negative control).
Statistics and reproducibility
The sample sizes and analytical measures are indicated in the figure legends and method details. In all cases, data from at least three independent experiments were used. Excel (Microsoft), ImageJ, ZEN 2.3, LAS X (Version 3.7.4, Leica), FlowJo (Version 10.10, BD Biosciences), and GraphPad Prism (Version 10, Dotmatics) were used. To visually and statistically represent data, each flow cytometry plot displays 10,000 or 20,000 randomly selected events.
Statistical analyses were conducted using GraphPad Prism 10. Different types of t-test, one-way ANOVA test and two-way ANOVA test were utilized as appropriate for different experimental conditions, and details of the statistical analyses performed and p values were described in the figure legends. Data sets were considered significantly different if the p value was less than 0.05. All Western blotting experiments were repeated three times. All micrographs were repeated three times, with at least three random fields of view captured per experiment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this study are available within the paper and its supplementary information files. Source data are provided with this paper.
References
Broz, P. & Dixit, V. M. Inflammasomes: mechanism of assembly, regulation and signalling. Nat. Rev. Immunol 16, 407–420 (2016).
Jorgensen, I. & Miao, E. A. Pyroptotic cell death defends against intracellular pathogens. Immunol. Rev 265, 130–142 (2015).
Rathinam, V. A. K., Zhao, Y. & Shao, F. Innate immunity to intracellular LPS. Nat. Immunol 20, 527–533 (2019).
Broz, P., Pelegrı´N, P. & Shao, F. The gasdermins, a protein family executing cell death and inflammation. Nat. Rev. Immunol 20, 143–157 (2020).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Ding, J. et al. Pore-forming activity and structural autoinhibition of the gasdermin family. Nature 535, 111–116 (2016).
Aglietti, R. A. et al. GsdmD p30 elicited by caspase-11 during pyroptosis forms pores in membranes. Proc. Natl. Acad. Sci. USA 113, 7858–7863 (2016).
Kuang, S. et al. Structure insight of GSDMD reveals the basis of GSDMD autoinhibition in cell pyroptosis. Proc. Natl. Acad. Sci. USA 114, 10642–10647 (2017).
Liu, X. et al. Inflammasome-activated gasdermin D causes pyroptosis by forming membrane pores. Nature 535, 153–158 (2016).
Liu, X., Xia, S., Zhang, Z., Wu, H. & Lieberman, J. Channelling inflammation: gasdermins in physiology and disease. Nat. Rev. Drug Discov 20, 384–405 (2021).
Xia, S. et al. Synthetic protein circuits for programmable control of mammalian cell death. Cell 187, 2785–2800.e16 (2024).
Wang, Q. et al. A bioorthogonal system reveals antitumour immune function of pyroptosis. Nature 579, 421–426 (2020).
Zhang, Z. et al. Gasdermin E suppresses tumour growth by activating anti-tumour immunity. Nature 579, 415–420 (2020).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Kayagaki, N. et al. Noncanonical inflammasome activation by intracellular LPS independent of TLR4. Science 341, 1246–1249 (2013).
Shi, J. et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature 514, 187–192 (2014).
Bernheim, A. & Sorek, R. The pan-immune system of bacteria: antiviral defence as a community resource. Nat. Rev. Microbiol 18, 113–119 (2020).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Shmakov, S. A. et al. Systematic prediction of genes functionally linked to CRISPR-Cas systems by gene neighborhood analysis. Proc. Natl. Acad. Sci. USA 115, E5307–E5316 (2018).
Shah, S. A. et al. Comprehensive search for accessory proteins encoded with archaeal and bacterial type III CRISPR-cas gene cassettes reveals 39 new cas gene families. RNA Biol 16, 530–542 (2019).
Özcan, A. et al. Programmable RNA targeting with the single-protein CRISPR effector Cas7-11. Nature 597, 720–725 (2021).
Sam, P. B., van Beljouw et al. The gRAMP CRISPR-Cas effector is an RNA endonuclease complexed with a caspase-like peptidase. Science 373, 1349–1353 (2021).
Strecker, J. et al. RNA-activated protein cleavage with a CRISPR-associated endopeptidase. Science 378, 874–881 (2022).
Kato, K. et al. Structure and engineering of the type III-E CRISPR-Cas7-11 effector complex. Cell 185, 2324–2337.e16 (2022).
Hu, C. et al. Craspase is a CRISPR RNA-guided, RNA-activated protease. Science 377, 1278–1285 (2022).
Kato, K. et al. RNA-triggered protein cleavage and cell growth arrest by the type III-E CRISPR nuclease-protease. Science 378, 882–889 (2022).
Stella, G. & Marraffini, L. Type III CRISPR-Cas: beyond the Cas10 effector complex. Trends. Biochem. Sci 49, 28–37 (2024).
Langedijk, A. C. & Bont, L. J. Respiratory syncytial virus infection and novel interventions. Nat. Rev. Microbiol 21, 734–749 (2023).
Schiffman, M. et al. Human papillomavirus and cervical cancer. Lancet 370, 890–907 (2007).
Ramos da Silva, J. et al. Single immunizations of self-amplifying or non-replicating mRNA-LNP vaccines control HPV-associated tumors in mice. Sci. Transl. Med 15, eabn3464 (2023).
Doorbar, J. Molecular biology of human papillomavirus infection and cervical cancer. Clin. Sci 110, 525–541 (2006).
Weng, C., Faure, A. J., Escobedo, A. & Lehner, B. The energetic and allosteric landscape for KRAS inhibition. Nature 626, 643–652 (2024).
Hofmann, M. H. et al. Expanding the reach of precision oncology by drugging all KRAS mutants. Cancer Discov 12, 924–937 (2022).
Bond, M. J. et al. Targeted degradation of oncogenic KRASG12C by VHL-recruiting PROTACs. ACS Cent Sci 6, 1367–1375 (2020).
Aqil, F. et al. Milk exosomes - natural nanoparticles for siRNA delivery. Cancer Lett 449, 186–195 (2019).
Skoulidis, F. et al. Sotorasib for lung cancers with KRAS p.G12C mutation. N. Engl. J. Med 384, 2371–2381 (2021).
Baker, D. J. et al. Naturally occurring p16(Ink4a)-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
McHugh, D., Durán, I. & Gil, J. Senescence as a therapeutic target in cancer and age-related diseases. Nat. Rev. Drug. Discov 24, 57–71 (2024).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med 383, 2603–2615 (2020).
Baden, L. R. et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N. Engl. J. Med 384, 403–416 (2021).
Wessels, H. H. et al. Massively parallel Cas13 screens reveal principles for guide RNA design. Nat. Biotechnol 38, 722–727 (2020).
Zhou, B. et al. Full-length GSDME mediates pyroptosis independent from cleavage. Nat. Cell. Biol 26, 1545–1557 (2024).
Becker, M. E. et al. Live imaging of airway epithelium reveals that mucociliary clearance modulates SARS-CoV-2 spread. Nat. Commun 15, 9480 (2024).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein. Sci 32, e4792 (2023).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein. Sci 30, 70–82 (2021).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein. Sci 27, 14–25 (2018).
Acknowledgements
We gratefully acknowledge Professor Zheng Hu from Wuhan University School of Medicine, Wuhan, China for the generous gift of HeLa and C33-A cells, and the original plasmids of HPV16-E6/E7 and HPV18-E6/E7, which were used for further modification. This work was supported by grants from the National Key R&D Program of China (2023YFC2307800), the National Natural Science Foundation of China (82550107, 82130064, 32470177, U24A20348 and U22A20337), and the Natural Science Foundation of Wuhan (2024040701010047).
Author information
Authors and Affiliations
Contributions
M.H. and W.W. co-performed the experiments. M.H., W.W., and M.C. designed the study and wrote the paper. M.H. mainly contributed to plasmid design and construction, flow cytometry, Western blotting and data analysis. W.W. mainly contributed to the detection of pyroptosis-related biochemical indicators, flow cytometry, cell imaging and data analysis. Y.Q. and M.C. supervised the study. H.Z., C.L., C.Z., J.L. and Y.H. provided experimental assistance. All authors discussed the results and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, M., Wang, W., Zhou, H. et al. A synthetic system for RNA-responsive pyroptosis based on type III-E CRISPR nuclease-protease. Nat Commun 17, 2565 (2026). https://doi.org/10.1038/s41467-026-69179-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69179-5