Abstract
Condensin organizes eukaryotic genomes into three-dimensional (3D) chromosome architectures that support accurate chromosome segregation during mitosis. However, the molecular mechanisms underlying this organization remain unclear. Here, we identify a previously unrecognized interaction between the condensin subunit Cnd1 and the mediator subunit Pmc4 in fission yeast, Schizosaccharomyces pombe. We characterize a condensin mutation, cnd1-K658E, which disrupts this interaction and observe that it impairs the formation of condensin-mediated chromatin domains during mitosis, resulting in chromosome segregation defects. This condensin-mediator interaction facilitates condensin recruitment to highly transcribed genes and mitotically activated genes, the latter of which demarcate condensin-mediated domains. Moreover, 1,6-hexanediol treatment and Pmc4 mediator depletion impair expression of mitotically activated genes, diminish condensin enrichment at those boundary genes, and disrupt domain boundaries, suggesting that mediator contributes to mitotic gene expression and chromosome architecture via phase separation. Together, these results reveal a mechanism by which mitotic gene expression patterns shape condensin-mediated chromosome architecture to ensure faithful chromosome segregation.
Similar content being viewed by others
Introduction
The cohesin and condensin complexes consist of Structural Maintenance of Chromosomes (SMC) factors, and they are collectively referred to as SMC complexes1,2,3,4. These complexes are evolutionarily conserved among eukaryotes5,6 and are primarily known for their roles in sister chromatid cohesion and mitotic chromosome condensation, respectively7,8,9,10,11,12,13. In addition to these canonical functions, cohesin and condensin have also been shown to participate in 3D genome organization14,15,16,17,18.
Cohesin forms chromatin contacts and topologically associating domains (TADs), also referred to as cohesin loops and domains19,20,21. It is important to note that genomic contacts and chromatin domains are related concepts; genomic contacts comprising multiple smaller interactions can collectively define chromatin domains. Cohesin loops are formed through loop extrusion and phase separation mechanisms22,23,24,25,26. TADs are associated with replication timing, and genes within the same TADs on the X chromosome tend to be co-regulated during early differentiation of mouse embryonic stem cells, although the precise role of TADs in gene regulation remains incompletely understood27,28,29. In this context, it has been shown that loss of cohesin, which eliminates TADs and cohesin loops, has only a modest impact on global gene expression and that transcriptional elongation disrupts cohesin loops, suggesting that TADs and loops may primarily reflect underlying transcriptional activity19,30. Nonetheless, some evidence indicates that TADs can functionally constrain enhancer activity within the domains, thereby preventing ectopic expression of developmental genes and oncogenes under certain conditions31,32. Mechanistically, CTCF has been proposed to define TAD boundaries by stalling cohesin-mediated loop extrusion17,20,21. In fission yeast, however, TAD boundaries are instead formed at convergently transcribed genes33,34.
In addition to cohesin, condensin also forms chromatin contacts and domains, also referred to as condensin domains in this study34,35,36,37,38,39. Condensin-mediated genomic contacts are thought to arise through loop extrusion and diffusion capture mechanisms40,41,42. In addition to its architectural role, condensin has also been implicated in gene regulation36,43,44,45,46,47,48. While CTCF has been established as a key factor in forming TADs and cohesin loops, the molecular factors that define condensin-mediated genomic contacts and domains remain largely unknown.
Our previous study has demonstrated that fission yeast cohesin and condensin form genomic contacts primarily within and beyond 100 kb, respectively34. Furthermore, cohesin and condensin establish chromatin domains of approximately 30–40 kb and 300 kb–1 Mb, respectively, which are inversely regulated during the cell cycle38. Mechanistically, the Cnd2 subunit of condensin interacts with the TBP TATA box-binding protein Tbp1 in fission yeast, and disruption of this interaction between Cnd2 and Tbp1 leads to reduced condensin localization across the genome49. These findings suggest that the interaction between condensin and the general transcription factor TBP contributes to condensin loading to gene loci16. Nevertheless, additional mechanisms are likely to contribute to condensin recruitment and the formation of condensin-mediated genomic contacts and domains.
This study demonstrates that the Pmc4 subunit of the mediator complex interacts with the Cnd1 subunit of condensin and that this interaction is required for accurate mitotic chromosome segregation. We developed a condensin mutation that disrupts the condensin-mediator interaction and found that the interaction is necessary for the formation of condensin domains. Mechanistically, the condensin-mediator interaction promotes the recruitment of condensin to highly transcribed genes and mitotically activated genes located at domain boundaries. Mediator is known as a transcriptional regulator/initiator complex50,51. This study demonstrates that mitotic chromosome architecture is shaped by the interplay between the transcriptional machinery, mediator, and the chromosome compaction machinery, condensin.
Results
Protein interaction between the Cnd1 condensin subunit and the Pmc4 mediator subunit
To elucidate how condensin cooperates with other factors to organize mitotic chromosome architecture, we searched for proteins that interact with condensin in fission yeast. For this purpose, we employed a yeast two-hybrid (Y2H) screening approach to identify interactions between condensin subunits and proteins encoded by a cDNA library (Yeast Genetic Resource Center, Osaka City University). The Y2H screen identified the mediator subunit Pmc4 as a potential interactor of the condensin subunit Cnd1 (Fig. 1a). Mediator has previously been shown to interact with cohesin in murine ES cells52, suggesting that mediator might also interact with condensin to organize mitotic chromosome structure. To test this possibility, we examined the Cnd1-Pmc4 interaction in fission yeast by co-immunoprecipitation (co-IP) analysis and detected an interaction (Fig. 1b). Furthermore, fission yeast Cnd1 and Pmc4 proteins were expressed in Sf9 insect cells and purified using GST pull-down. Analysis of size exclusion chromatography fractions using a Superose 6 column revealed a stable and stoichiometric complex between Cnd1 and Pmc4 (Fig. 1c). These complementary assays consistently supported a stable Cnd1-Pmc4 interaction.
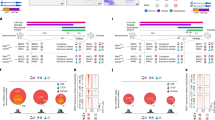
a Yeast two-hybrid (Y2H) analysis of the interaction between the condensin subunit Cnd1 and the mediator subunit Pmc4. The pmc4 cDNA fragment encoding 10-239 amino acid (aa) residues of the full-length Pmc4 (239 aa) was initially isolated by Y2H screening. The interaction between Cnd1 and Cnd2 condensin subunits serves as a positive control (bottom). b Co-IP result showing the interaction between Cnd1-Myc condensin and Pmc4-Pk mediator subunits in fission yeast cells. Cnd1-Myc and Pmc4-Pk were expressed from their endogenous loci. Three independent experiments showed similar results. c GST pull-down analysis on GST-Pmc4 with His-Cnd1. GST-Pmc4 and His-Cnd1 were co-expressed in sf9 insect cells and subjected to GST pull-down, size exclusion chromatography, SDS-PAGE, and Coomassie Brilliant Blue staining. Two biological replicates showed similar results. d Genome-wide distributions of Cnd1-Pk, Cut14-Pk, Pmc4-Pk, Tbp1-Pk, and Rad21-Myc. The no-tag control distribution was included to represent the background signal in ChIP-seq analysis. e Summary of binding sites co-occupied by Cnd1-Pk and Pmc4-Pk (left) and by Cut14-Pk and Pmc4-Pk (right). The genetic features are explained in Supplementary Fig. 1a. f Venn diagrams showing the overlaps between Cnd1-Pk and Pmc4-Pk binding sites (upper left) and their associated genes (upper right), as well as between Cut14-Pk and Pmc4-Pk binding sites (bottom left) and their associated genes (bottom right). Results are summarized as in Supplementary Fig. 1b. g Heatmaps showing the distributions of Cnd1-Pk (left), Cut14-Pk (middle), and Pmc4-Pk (right) at their common target genes (n = 407). Pol II genes commonly bound by Cnd1-Pk, Cut14-Pk, and Pmc4-Pk were ordered from top to bottom based on Cnd1-Pk ChIP-seq enrichment. Results are represented as described in Supplementary Fig. 1g, h. h Co-localization of Cnd1-Pk, Cut14-Pk, and Pmc4-Pk at a 130 kb genomic region containing Pol III genes, such as tRNA and 5S rRNA, and the mitotically activated eng1 gene. Binding patterns of Cnd1-Pk, Cut14-Pk, and Pmc4-Pk at the eng1 (i) and rps23 (j) gene loci. The rps23 gene is highly transcribed and encodes a ribosomal protein. Source data for (b, c) are provided as a Source Data file.
Co-localization of condensin and mediator at gene regions
If condensin and mediator interact, they may co-localize across the fission yeast genome. To test this possibility, we performed ChIP-seq analyses to map the condensin subunits Cnd1 and Cut14 and the mediator subunit Pmc4 (Fig. 1d). We found that the condensin subunits co-localized with Pmc4 at RNA polymerase II-transcribed (Pol II) genes, RNA polymerase III-transcribed (Pol III) genes, such as tRNA and 5S rRNA genes, long terminal repeats (LTRs), and non-coding RNA (ncRNA) genes (Fig. 1e). The condensin subunits were significantly co-localized with Pmc4 at their binding sites (P < 7.63 × 10–334 for Cnd1 and P < 1.54 × 10–760 for Cut14, hypergeometric distribution test) and at Pol II genes (P < 8.04 × 10- 313 for Cnd1 and P < 1.90 × 10−690 for Cut14, hypergeometric distribution test; Fig. 1f). Among the 407 Pol II genes co-occupied by the condensin subunits and Pmc4, Cnd1 and Cut14 were preferentially enriched at the 3’ ends, whereas Pmc4 was enriched at the 5’ ends of genes (Fig. 1g). Moreover, the condensin subunits were clearly co-localized with Pmc4 at Pol II and Pol III genes (Fig. 1h). Interestingly, strong enrichment of the condensin subunits and Pmc4 was observed at the eng1 gene and other mitotically activated genes (Fig. 1g–i). The Ace2 transcription factor induces expression of these mitotically activated genes53,54. The condensin subunits and Pmc4 were also enriched at highly transcribed housekeeping genes, including the rps23 ribosomal protein gene (Fig. 1j). These results suggest that condensin and mediator tend to co-localize at highly transcribed genes and mitotically activated Ace2 target genes.
Additionally, we analyzed genome-wide distributions of Tbp1(TATA-box binding protein, TBP) and the Rad21 cohesin subunit because we previously found that Tbp1 interacts with the Cnd2 condensin subunit and that cohesin and condensin both tend to localize at the 3’ ends of Pol II genes34,49. We observed that Cnd1, Cut14, Pmc4, Tbp1, and Rad21 were enriched at Pol II genes, Pol III genes, LTRs, and ncRNA genes (Supplementary Fig. 1a). The condensin subunits were significantly co-localized with Pmc4 and Tbp1 at their binding sites (P < 5.58 × 10–299 for Cnd1 and P < 7.48 × 10−304 for Cut14, hypergeometric distribution test) and at Pol II genes (P < 1.32 × 10–265 for Cnd1 and P < 2.76 × 10−230 for Cut14, hypergeometric distribution test), supporting interactions between condensin and both TBP and mediator (Supplementary Fig. 1b). Consistent with previous studies, the condensin subunits and the Rad21 cohesin subunit were localized at the kinetochore-forming and heterochromatic centromere regions, respectively (Supplementary Fig. 1c)9,49,55. In contrast, Tbp1 and Pmc4 were enriched at centromeric tRNA genes, implying that the co-localization between condensin and the transcription-related factors, TBP and mediator, is largely restricted to gene regions (Supplementary Fig. 1c). To further support this observation, the condensin subunits, Pmc4, and Tbp1 were enriched at Ace2 target genes, the top 10% highest transcribed Pol II genes, and Pol III genes (Supplementary Fig. 1d–f).
Moreover, we observed a clear difference in condensin and cohesin localizations. Although Rad21 was also localized at the top 10% highest transcribed Pol II genes and Pol III genes, where the condensin subunits, Pmc4, and Tbp1 were also enriched (Supplementary Fig. 1e, f), the heatmap illustrating their distributions across all Pol II genes indicated that the condensin subunits, Pmc4, and Tbp1 were tightly co-localized at Pol II genes (n = 286), whereas Rad21 was more broadly distributed at the 3’ ends of many additional Pol II genes (Supplementary Fig. 1g, h). The condensin subunits, Pmc4, and Tbp1 were frequently co-localized at highly transcribed Pol II genes and mitotically activated Ace2 target genes (Supplementary Fig. 1h). These ChIP-seq and biochemical results collectively suggest that condensin co-localizes with mediator and Tbp1 at gene regions through protein-protein interactions.
Impairment of Cnd1-Pmc4 interaction by cnd1-K658E mutation
To investigate the function of the Cnd1-Pmc4 interaction, we set out to generate cnd1 mutations that specifically inhibit the interaction without disrupting the condensin complex. We employed a Y2H screening approach combined with PCR-based random mutagenesis and identified the cnd1-m1 mutant allele (Fig. 2a and Supplementary Fig. 2a, b). The cnd1-m1 allele consisted of five point mutations: T584A, K658E, R905H, M1057K, and a nonsense mutation at the Y1136 residue. We dissected these point mutations and found that the K658E substitution alone was sufficient to disrupt the Cnd1-Pmc4 interaction (Fig. 2a). Notably, the Cnd1-Cnd2 interaction is known to be the only interaction required for Cnd1 to integrate into the condensin complex56. The Y2H data revealed that the Cnd1-Cnd2 interaction was unaffected by the cnd1-K658E mutation, implying that this mutation impairs the Cnd1-Pmc4 interaction without interfering with condensin complex formation (Fig. 2a). This finding was further supported by co-IP analyses using the endogenous fission yeast system, again showing that the cnd1-K658E mutation disrupts the Cnd1-Pmc4 interaction but not the Cnd1-Cnd2 interaction (Fig. 2b, c).
a Y2H analysis examining the interaction of wild-type (WT) and mutant Cnd1 with the mediator subunit Pmc4 and the condensin subunit Cnd2. Wild-type or mutant Cnd1 fused to GAL4-BD was co-expressed with Pmc4 or Cnd2 fused to GAL4-AD. The control plate (SD-Trp-Leu) was used to maintain the GAL4-BD and GAL4-AD plasmids, whereas the +Aur plate (SD-Trp-Lue-His-Ade containing 125 ng/ml aureobasidin A) was used to assess Y2H interactions. b Co-IP analysis investigating the interaction of wild-type and mutant Cnd1-Myc with Pmc4-Pk. Three independent experiments showed similar results. c Co-IP analysis showing the interactions of wild-type and mutant Cnd1-Myc with Cnd2-Pk. Three independent experiments showed similar results. d Chromosome segregation defects in the cnd1-K658E mutant strain. Cells carrying the cnd1-WT and cnd1-K658E genes at the endogenous locus were subjected to immunofluorescence analysis to visualize spindle microtubules (mitotic marker) and DAPI-stained DNA. Mitotic cells displaying lagging chromosomes (arrowheads) were quantified. The experiments were conducted in three independent replicates. e Chromosome segregation defects in cells depleted of Pmc4. Cells were cultured in EMM liquid medium containing auxin to deplete the endogenous Pmc4, while exogenous Pmc4 was expressed from a plasmid. The vector control represents cells lacking exogenous Pmc4 expression (Pmc4 depletion). Mitotic defects were analyzed as described in (d). Data in (d, e) are represented as mean ± SD and P-values were calculated by two-sided Student’s t test. Source data for (b-e) are provided as a Source Data file.
Involvement of Cnd1-Pmc4 interaction in chromosome segregation
To elucidate the role of the Cnd1-Pmc4 interaction in chromosome segregation, we analyzed mitotic chromosomes in the cnd1-K658E mutant strain. We found that the frequency of chromosome segregation defects was significantly increased in cnd1-K658E cells compared to wild-type cells (Fig. 2d). This result suggests that the cnd1-K658E mutation causes segregation defects by disrupting the Cnd1-Pmc4 interaction.
Moreover, to investigate the role of condensin and mediator in chromosome segregation, we employed the auxin-inducible degron (AID) system to deplete endogenous proteins. For this purpose, we first optimized the AID system by testing different combinations of degrons (IAA7 and IAA17) and F-box proteins (atTIR1, atAFB2, and osTIR1). To this end, the Pmc4 mediator subunit was fused to a degron, and we found that IAA17 in combination with the two F-box proteins atAFB2 and osTIR1 was most effective (Supplementary Fig. 2c). Specifically, cells grew well on control plates without auxin, but growth was impaired on auxin-containing plates, consistent with the essential role of Pmc4 in cell viability. Moreover, expression of Pmc4 proteins from plasmids rescued the growth defects of this strain, supporting the conclusion that the growth defects resulted from Pmc4 depletion (Supplementary Fig. 2d).
To investigate the function of the Cnd1-Pmc4 interaction, we depleted endogenous Cnd1 expression by inducing targeted protein degradation and expressed wild-type and mutant Cnd1 proteins from plasmids. We observed that cells lacking exogenous Cnd1 (vector; Cnd1 depletion) did not grow on the EMM plates supplemented with Auxin, indicating effective depletion of Cnd1 (Supplementary Fig. 2e). Supporting this, western blot analysis showed that endogenous Cnd1 fused to AID tag was reduced following activation of the AID system (Supplementary Fig. 2f). In contrast, cells grew on the same plates when wild-type or mutant Cnd1 proteins were provided from plasmids, suggesting that exogenous Cnd1 can compensate for the loss of endogenous Cnd1. Interestingly, the frequency of chromosome segregation defects was significantly higher in cells expressing Cnd1-K658E compared to those expressing wild-type Cnd1 (Supplementary Fig. 2g). We also observed that cells depleted of endogenous Cnd1 and expressing exogenous Cnd1-K658E exhibited similar growth to those expressing wild-type Cnd1 (Supplementary Fig. 2e), implying that the cnd1-K658E mutation is sufficient to cause chromosome segregation defects without affecting cell viability at a detectable level. Therefore, the cnd1-K658E mutation likely serves as a valuable genetic tool for investigating the function of the condensin-mediator interaction in chromosome segregation. Moreover, cnd1-K658E mutant cells did not show increased sensitivity to thiabendazole (TBZ), a microtubule-destabilizing agent, compared to wild-type cells (Supplementary Fig. 2h). These results collectively suggest that the Cnd1-Pmc4 interaction is required for faithful chromosome segregation. Furthermore, we detected chromosome segregation defects in cells depleted of Pmc4, reinforcing the idea that mediator contributes to chromosome segregation during mitosis through its interaction with condensin (Fig. 2e).
Roles of Cnd1-Pmc4 interaction in the formation of condensin-mediated genomic contacts and chromatin domains
We examined the effects of the cnd1-K658E mutation on 3D chromosome architecture during mitosis. To obtain mitotic cells, we used the cdc25-22 G2/M block-release method and confirmed proper cell cycle synchronization in the strains used for our analysis (Supplementary Fig. 3a). We performed in situ Hi-C experiments using wild-type and cnd1-K658E mutant mitotic cells (Fig. 3a). The difference map and distance curves indicated that genomic contacts at distances of approximately 75 to 800 kb were reduced in the cnd1-K658E mutant compared to wild-type cells (Fig. 3b, c). Genomic contacts within this range are known to be mediated by condensin38. The enlarged difference map consistently revealed that genomic contacts within condensin domains were impaired in the cnd1-K658E mutant relative to wild-type cells (Fig. 3d). In this study, we frequently used a 1.9 Mb region as a model locus to examine chromatin domains and boundaries, as we previously found that the eng1 and SPAC343.20 genes are mitotically activated by the Ace2 transcription factor and serve as condensin domain boundaries34. Contact scores within condensin domains were significantly reduced in the cnd1-K658E mutant (P = 2.10 × 10−10, two-sided paired Student’s t test; Fig. 3e).
a Genome-wide contact maps of mitotic cells from the wild-type strain (cnd1-WT; left) and the cnd1-K658E mutant strain (cnd1-K658E; right). ICE-normalized matrices at 20 kb resolution were used for data representation. Mitotic cells were prepared as described in Supplementary Fig. 3a. b Difference map showing changes in contact scores between cnd1-K658E and cnd1-WT cells. The Hi-C data shown in (a) were used to calculate the difference scores. c Relationships between contact scores and genomic distances in cnd1-WT and cnd1-K658E cells. Contact matrices at 10-kb resolution were used for this analysis. d Enlarged difference map of a 1.9 Mb region of chromosome I (dotted box in (b)). Data representation is explained in Supplementary Fig. 3e. e Averages and log2 ratios of contact scores between cnd1-K658E and the cnd1-WT cells, and between biological replicates from mitotic cell cultures (inset). Analysis was performed as described in Supplementary Fig. 3f. f Contact maps of mitotic cells expressing wild-type Pmc4 from a plasmid (Pmc4 WT; left) and cells lacking exogenous Pmc4 expression (Pmc4 depletion; right). Endogenous Pmc4 proteins were degraded using the AID system. Mitotic cells were prepared as in Supplementary Fig. 3a. g Genome-wide difference map showing changes in contact scores between Pmc4-depleted and Pmc4 WT cells. h Relations between contact scores and genomic distances in Pmc4 WT and Pmc4-depleted cells. i Enlarged difference map of the 1.9 Mb region on chromosome I (dotted box in (g)) comparing Pmc4-depleted and Pmc4 WT cells. Border strength in each sample was calculated as described38. The difference in the border strength scores between the two samples is shown on the right. Arrows mark domain boundaries impaired by Pmc4 depletion. j Averages and log2 ratios of contact scores between Pmc4-depleted and Pmc4 WT cells. The analysis was performed as described in Supplementary Fig. 3f. Insets show biological replicates from mitotic cell cultures. Across the entire study, the same conditions were used for representing Hi-C matrices, generating difference maps, plotting distance curves, and performing statistical analyses of condensin domains.
We also performed in situ Hi-C experiments with mitotic cells depleted of endogenous Cnd1 and expressing exogenous wild-type or mutant Cnd1 (Supplementary Fig. 3b). The difference maps and distance curves indicated that both Cnd1-K658E expression and Cnd1 depletion reduced genomic contacts at distances of approximately 75 to 800 kb (Supplementary Fig. 3c, d). We also observed that a cross-shaped pattern was enhanced around centromeres in Cnd1-depleted cells, suggesting increased contacts between centromere-proximal regions in the absence of condensin. The enlarged difference maps revealed that both conditions impaired genomic contacts within condensin domains (Supplementary Fig. 3e). Contact scores within condensin domains were significantly reduced by both Cnd1-K658E expression and Cnd1 depletion (P = 2.46 × 10−14 and 1.85 × 10−26 for Cnd1-K658E expression and Cnd1 depletion, respectively, two-sided paired Student’s t-test; Supplementary Fig. 3f). These results collectively indicate that the interaction between Cnd1 and Pmc4 contributes to the formation of condensin-mediated genomic contacts and chromatin domains.
It has previously been shown that the Cnd2 condensin subunit interacts with Tbp149. Therefore, we also examined the 3D chromosome architecture in cells depleted of endogenous Cnd2 and expressing either wild-type or mutant Cnd2 (Supplementary Fig. 4a). The cnd2-C703R mutation has been shown to impair the Cnd2-Tbp1 interaction without affecting condensin complex formation. We found that Cnd2-C703R expression and Cnd2 depletion significantly diminished condensin domains (Supplementary Fig. 4a–d). Notably, the effects of Cnd2-C703R expression and Cnd2 depletion on domain formation were relatively weaker than those of Cnd1-K658E expression and Cnd1 depletion, likely because asynchronous cell populations were subjected to Hi-C analysis. Nevertheless, these results indicate that condensin interactions with mediator and TBP contribute to the formation of condensin-mediated genomic contacts and chromatin domains.
Mediator contributes to the formation of condensin-mediated genomic contacts and chromatin domains
To investigate the involvement of mediator in shaping 3D chromosome architecture during mitosis, we performed in situ Hi-C using mitotic cells in which Pmc4 proteins were degraded through the AID system (Fig. 3f). The difference map between Pmc4 WT and Pmc4-depleted cells, along with the distance curves, revealed that genomic contacts between 75 and 800 kb were impaired by Pmc4 depletion, although the effect was relatively milder than Cnd1 depletion or Cnd1-K658E expression (Fig. 3g, h). The enlarged difference map and statistical analysis also indicated that condensin-mediated domains and genomic contacts were significantly disrupted by Pmc4 depletion (Fig. 3i, j; P = 3.73 × 10−4, two-sided paired Student’s t-test). In addition to the defects in condensin domain formation, we observed that Pmc4 depletion enhanced genomic contacts between two adjacent condensin domains and that border strength scores at domain boundaries were reduced by Pmc4 depletion (Fig. 3i). These results suggest that mediator contributes not only to the formation of condensin-mediated genomic contacts but also to the assembly of domain boundaries.
Mediator is a large protein complex composed of approximately 23 subunits in fission yeast57. These subunits are organized into two modules: the core complex and the CDK8 regulatory subcomplex, which have distinct roles in specific biological pathways51,58. We therefore extended our analysis to additional mediator subunits and subjected nine non-essential mediator deletion mutants to Hi-C analysis. We found that condensin-mediated genomic contacts and chromatin domains were impaired in med10∆, med20∆, and pmc3∆ (also known as med27∆) mutants (Supplementary Fig. 5). However, the effects of these deletions were weaker than those observed with Pmc4 depletion, likely because asynchronous cell populations were used for the Hi-C analysis of mediator mutants. We also observed that border strength scores were decreased around the eng1 gene locus in the mediator mutants (med10∆ and med20∆), suggesting that the chromatin boundary at the eng1 locus was compromised. These results suggest that mediator as a whole, and not Pmc4 alone, contributes to the formation of condensin-mediated genomic contacts and chromatin domains.
Effects of condensin-mediator double depletion on 3D genome organization during mitosis
Previous studies have shown that growth defects caused by condensin mutations can be partially suppressed by mutations in mediator components59,60, suggesting that mediator may play an inhibitory role in condensin-mediated chromosome organization. To investigate the underlying mechanisms, we combined Cnd1 condensin and Pmc4 mediator depletion and performed in situ Hi-C analysis using mitotic cells (Supplementary Fig. 6a). The difference maps and distance curves indicated that, compared to wild-type cells, Cnd1-Pmc4 double depletion reduced genomic contacts within condensin domains (Supplementary Fig. 6b–d). Additionally, we calculated the log2 ratio of contact scores between wild-type and Cnd1-Pmc4 double-depleted cells and found that condensin domains were significantly impaired in Cnd1-Pmc4 double-depleted cells compared with wild-type cells (Supplementary Fig. 6e).
Furthermore, when Cnd1-Pmc4 double depletion was compared to Cnd1 single depletion, we observed that genomic contacts within approximately 800 kb were increased, suggesting that mediator can inhibit genomic contacts within this range (Supplementary Fig. 6b–e). Notably, these genomic contacts were enhanced in the absence of condensin. In contrast, when Cnd1-Pmc4 double depletion was compared to Pmc4 single depletion, genomic contacts within approximately 800 kb were decreased, suggesting that condensin promotes genomic contacts within this range (Supplementary Fig. 6b–e). Taken together, these observations support the idea that mediator plays an inhibitory role in genomic contacts up to ~800 kb during mitosis, which may explain why mediator mutations can partially suppress condensin mutations that impair contacts within a similar range. Our results suggest that condensin and mediator cooperate to establish the functional chromosome architecture required for faithful chromosome segregation (see “Discussion”).
Detection of condensin domain disruption caused by cnd1-K658E mutation using FISH microscopy
Since Hi-C analysis indicated that condensin domains were impaired in the cnd1-K658E mutant, we sought to examine domain organization in the mutant using a FISH-based microscopic approach. For this analysis, we designed FISH probes to visualize the two paired genomic loci either within the same domain or across different domains (Fig. 4a). Interphase and mitotic cells were distinguished by spindle staining. The visualized loci are consistently 110 kb apart for both intra- and inter-domain FISH probes. In wild-type cells, the intra-domain probes were positioned significantly closer during mitosis than during interphase, whereas the inter-domain probes were also located closer during mitosis than during interphase, albeit to a lesser extent than the intra-domain probes (Fig. 4b). This observed difference between intra- and inter-domain probes reflects domain-based chromosome compaction during mitosis. We next visualized the same paired loci in the cnd1-K658E mutant and found that the shortening of the intra-domain probe distances observed in wild-type cells during mitosis was significantly compromised in the mutant, indicating that domain-based chromosome compaction during mitosis is impaired in the cnd1-K658E mutant (Fig. 4b).
a Positions of FISH probes. Probes #1 and #2 span adjacent condensin domains (dotted lines), while probes #2 and #3 are positioned within a single domain. The genomic regions targeted by these probes are consistently separated by 110 kb for both inter-domain and intra-domain FISH probes. An enlarged difference map of the 1.9 Mb region on chromosome I, comparing Cnd1-depleted and Cnd1-WT cells, is shown to represent condensin domains; experimental details are provided in Supplementary Fig. 3. b Quantification of distances between inter-domain or intra-domain FISH probes in interphase and mitotic cells of the wild-type (cnd1-WT) and cnd1-K658E mutant (cnd1-K658E) strains. Representative FISH-IF images were shown below, displaying two loci (red and green) and spindle microtubules (blue; mitotic marker). c Quantification of distances between inter-domain or intra-domain FISH probes in interphase and mitotic cells expressing Pmc4 from a plasmid (Pmc4 WT) or lacking plasmid-derived Pmc4 (Pmc4 depletion). Endogenous Pmc4 was removed by the AID system. d Schematic representations of mitotic condensin domains under the indicated conditions. Green and red dots indicate the schematic positions of genomic regions visualized by FISH probes (#1, #2, and #3). In wild-type cells, two adjacent condensin domains are formed during mitosis. In mitotic cells of the cnd1-K658E mutant, these domains are impaired and occupy larger areas, leading to increased intra-domain distances compared to wild-type cells. In mitotic cells depleted of Pmc4, the distances between the two domains are shortened, suggesting a disruption of domain boundary formation. The central bar represents the median, with the boxes indicating the upper and lower quartiles. The whiskers extend to data points within 1.5× the interquartile range from the box. P-values were calculated by two-sided Mann–Whitney U test. Source data for (b, c) are provided as a Source Data file.
Moreover, we designed other sets of FISH probes to visualize the two paired genomic loci located either within the same domain or across different domains (Supplementary Fig. 7a). The visualized loci are consistently 410 kb apart for both intra- and inter-domain FISH probes. We analyzed these loci in cells depleted of endogenous Cnd1 and expressing exogenous wild-type or mutant Cnd1, and found that when wild-type Cnd1 was expressed, the intra-domain probes were positioned significantly closer during mitosis than during interphase, whereas the inter-domain probes were also closer during mitosis than during interphase, although to a lesser extent than the intra-domain probes (Supplementary Fig. 7b). In contrast, the shortening of the intra-domain probe distances observed in mitotic cells expressing wild-type Cnd1 was significantly compromised in cells expressing Cnd1-K658E (Supplementary Fig. 7b). These results consistently demonstrate that the Cnd1-Pmc4 interaction promotes domain-based chromosome compaction during mitosis.
Furthermore, we visualized condensin domains using multiple FISH probes (Supplementary Fig. 7c). We observed that the areas occupied by FISH signals varied depending on fluorescent dyes and imaging conditions. Therefore, we measured the distance between the centers of FISH signals as a reproducible parameter to assess the organization of the two condensin domains. We examined condensin domains in cells depleted of endogenous Cnd1 and expressing exogenous Cnd1 from plasmids. We found that when wild-type Cnd1 was expressed, the centers of the two condensin domains were positioned significantly closer during mitosis than during interphase, reflecting domain-based chromosomal compaction during mitosis (Supplementary Fig. 7d). In contrast, the distances between the two domains were not shortened during mitosis relative to interphase in cells depleted of Cnd1 or expressing Cnd1-K658E (Supplementary Fig. 7d). These results suggest that the mitosis-specific shortening in distance between the two condensin domains depends on condensin and the Cnd1-Pmc4 interaction. The distances between the two condensin domains in mitotic cells expressing the Cnd1-K658E were similar to those of Cnd1-depleted mitotic cells, suggesting that the Cnd1-Pmc4 interaction plays a major role in condensin domain organization, although it also remains possible that the cnd1-K658E mutation impairs condensin function independently of the Cnd1-Pmc4 interaction.
Detection of condensin domain disruption caused by Pmc4 depletion using FISH microscopy
We visualized two paired genomic loci located either within the same domain or across different domains in cells depleted of endogenous Pmc4, with or without exogenous Pmc4 expression (Fig. 4a, c). We found that the distances between the intra-domain probes were significantly shorter during mitosis than during interphase in cells expressing Pmc4, whereas the inter-domain probes were also positioned closer during mitosis than during interphase, albeit to a lesser extent than the intra-domain probes (Fig. 4c). Interestingly, we observed that the shortening of the intra-domain probe distances observed in mitotic cells expressing Pmc4 was modestly impaired in Pmc4-depleted cells, suggesting weakened intra-domain compaction. In addition, the distances between the inter-domain probes were slightly but significantly shorter during mitosis in Pmc4-depleted cells compared with Pmc4-expressing cells, implying a compromised domain boundary. Together, these results indicate that Pmc4 depletion slightly impairs domain-based chromosome compaction and diminishes boundary formation.
We next visualized the two condensin domains using two groups of multiple probes and found that, when the wild-type Pmc4 was expressed, the centers of the two condensin domains were positioned significantly closer during mitosis than during interphase (Supplementary Fig. 7e). This mitosis-specific shortening of distance between the condensin domains was partly compromised in Pmc4-depleted cells, presumably reflecting impaired condensin domain formation and weakened boundaries. Based on overall FISH and Hi-C results, our model is that domain-based chromosomal compaction is diminished in the cnd1-K658E mutant, whereas domain compaction is partially compromised in Pmc4-depleted cells, coinciding with impaired boundary formation (Fig. 4d).
Cnd1-Pmc4 interaction is involved in condensin recruitment
Our results suggested that the Pmc4 mediator and Cnd1 condensin subunits interact and co-localize at highly transcribed genes and mitotically activated genes (Fig. 1). Disruption of the Cnd1-Pmc4 interaction impaired condensin-mediated genomic contacts and domain organization (Figs. 3 and 4). These results led us to hypothesize that the mediator recruits condensin to gene loci. To test this hypothesis, we performed ChIP-seq experiments to map wild-type and mutant Cnd1 expressed from the endogenous locus (Fig. 5a). We found that Cnd1-K658E enrichment was impaired across the genome, including at mitotically activated Ace2-target genes, highly transcribed genes, and Pol III genes (Fig. 5b–g). In addition, we analyzed mitotic cells depleted of endogenous Cnd1 and expressing wild-type or mutant Cnd1 from plasmids and observed that Cnd1-K658E showed reduced enrichment compared to wild-type Cnd1 (Supplementary Fig. 8a–g). Furthermore, we investigated condensin enrichment in mitotic cells depleted of Pmc4 and found that Cnd1 enrichment at gene regions was impaired by Pmc4 depletion (Fig. 5a–f, h). These results suggest that mediator localizes to gene regions to recruit condensin via the Cnd1-Pmc4 interaction.
a Genome-wide distributions of condensin were determined by ChIP-seq using mitotic cells expressing wild-type Cnd1 (Cnd1-WT-Myc) or Cnd1-K658E mutant (Cnd1-K658E-Myc) from the endogenous cnd1 locus (top two rows). Mitotic cells were prepared in YEA medium using the cdc25-22 G2/M block-release method and harvested 40 min after release. In addition, the genome-wide profiles of Cnd1-Pk in mitotic cells expressing wild-type Pmc4 from a plasmid (Pmc4 WT) or lacking exogenous Pmc4 expression (Pmc4 depletion) are shown (bottom two rows). Endogenous Pmc4 was depleted by the AID system. Mitotic cells were prepared in EMM media by the cdc25-22 G2/M block-release procedure and harvested 60 min after release. Condensin enrichment across a 130-kb genomic region on chromosome I (b), the Ace2-regulated eng1 gene locus (c), and the highly transcribed rps23 gene locus (d). Average condensin enrichment at Ace2 target and highly transcribed Pol II genes (e) and at tRNA and 5S rRNA Pol III genes (f) in the indicated samples. Highly transcribed Pol II genes were classified as described in Fig. 1e, and genes bound by both Cnd1 and Cut14 (defined in Fig. 1f) were designated as condensin-associated genes. Genes overlapping between these two categories were annotated as the highly transcribed, condensin-binding genes. g ChIP-qPCR analysis verifying enrichment of Cnd1-WT-Myc and Cnd1-K658E at the eng1, rps23, and tRNA-ser loci in mitotic cells. Experiments were performed in triplicate as described in (a). h ChIP-qPCR analysis verifying the enrichment of Cnd1-Pk at the eng1, rps23, and tRNA-ser loci in mitotic cells expressing wild-type Pmc4 from a plasmid (Pmc4 WT) or lacking exogenous Pmc4 expression (Pmc4 depletion). Experiments were performed in triplicate as described in (a). Data in (g, h) are represented as mean ± SD. Source data are provided as a Source Data file.
We also examined condensin distribution in cells depleted of endogenous Cnd2 and expressing wild-type or mutant Cnd2 from plasmids (Supplementary Fig. 8a). We observed that Cnd2-C703R enrichment was impaired at mitotically activated Ace2-target genes, highly transcribed Pol II genes, and Pol III genes (Supplementary Fig. 8b–f, h). This result suggests that TBP is also involved in condensin loading onto gene regions across the genome.
Pmc4 depletion impairs expression of mitotically activated genes
To explore the potential role of the condensin-mediator interaction in transcription, we examined global expression profiles in cells expressing Cnd1-K658E. For this purpose, we employed the nascent RNA labeling as described previously61, followed by deep sequencing (nascent RNA-seq) to quantify nascent transcripts across the genome. We found that Cnd1-K658E expression and Cnd1 depletion had only modest effects on transcription (Supplementary Fig. 9a–e). We also performed nascent RNA-seq experiments using Pmc4-depleted cells (Supplementary Fig. 9a–d) and found that eng1 transcription was almost completely abolished by Pmc4 depletion (Supplementary Fig. 9c). Among 87 significantly down-regulated genes, 8 were Ace2 target genes (Supplementary Fig. 9f). We confirmed that condensin genes were not significantly affected by Pmc4 depletion.
Next, we compared the nascent RNA-seq expression profiles with ChIP-seq enrichment of Cnd1-Pk and Cut14-Pk (condensin), Pmc4-Pk (mediator), and Tbp1-Pk (TBP). We observed that gene expression levels were tightly correlated with ChIP enrichment of these factors, as strong nascent RNA-seq signals coincided with high ChIP enrichment (Supplementary Fig. 9g). Intriguingly, genes with the highest enrichment of these factors were typically mitotically activated Ace2-targeted genes, which were down-regulated by Pmc4 depletion (Supplementary Fig. 9h, i). Moreover, approximately 300 genes with high enrichment of condensin, mediator, and TBP were generally highly transcribed (Supplementary Fig. 9h). Expression of 341 highly transcribed genes was slightly enhanced in Pmc4-depleted cells (Supplementary Fig. 9h, j). These observations predict that the mediator promotes the expression of mitotically activated Ace2-target genes located at domain boundaries.
Mediator localization disrupted by 1,6-hexanediol treatment
Human Med1, a component of the mediator complex, contains intrinsically disordered regions (IDRs) and is known to form droplets through liquid-liquid phase separation62,63. Such droplet formation by Med1 has been linked to transcriptional initiation64. In fission yeast, the mediator subunit Pmc4 is predicted to possess a disordered region at its C-terminal portion (Supplementary Fig. 10a). Based on this observation, we hypothesized that Pmc4 might form liquid droplets. To test this hypothesis, we first purified Pmc4 proteins expressed in Escherichia coli and observed that Pmc4 formed droplets when exposed to PEG concentrations exceeding 5%, implying that Pmc4 may undergo droplet formation (Supplementary Fig. 10b, c).
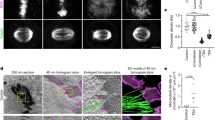
We examined Pmc4 localization in fission yeast cells and observed that treatment with 1,6-hexanediol (HD), a compound known to suppress liquid-liquid phase separation, disrupted the nuclear focal localization of Pmc4, whereas Rbp1 Pol II foci remained nuclear under the same conditions (Fig. 6a). These results imply that mediator, including Pmc4, is prone to forming droplets via phase separation. To gain further insight into the subnuclear localization of mediator and condensin, we visualized these factors using Zeiss Airyscan super-resolution microscopy and observed that the Cnd1 and Cnd2 condensin subunits formed fiber-like structures along mitotic chromosomes (Fig. 6b and Supplementary Fig. 11a, b). In contrast, the mediator appeared as spherical foci on mitotic chromosomes. Notably, mediator foci did not strongly overlap with condensin signals but were instead often localized at the termini of condensin-associated fibrous structures, potentially reflecting their respective distributions at promoters and transcribed regions. Moreover, we generated Pmc4 variants carrying amino acid substitutions predicted to disrupt the disordered region (Supplementary Fig. 10d). Spherical foci were observed for both wild-type and variant Pmc4, suggesting that the disordered region of Pmc4 may not be directly responsible for mediator droplet formation (Supplementary Fig. 10e). Our current hypothesis is that Med1 may mediate droplet formation as observed in human cells62,63.
a Effects of 1,6-hexanediol (HD) treatment on Pmc4 localization. Fission yeast cells were treated with 5 or 10% HD for 1 or 5 min and subjected to immunofluorescence analysis to visualize Pmc4-Pk and Rpb1 (RNA Pol II subunit). Two biological replicates showed similar results. b Subnuclear localization of Pmc4-Pk and Cnd2 visualized using Zeiss Airyscan super-resolution microscope. Three independent experiments showed similar results. c Genome-wide contact maps of mitotic cells, either untreated (left) or treated (right) with 5% HD. Mitotic cells were prepared in YEA medium using the cdc25-22 G2/M block-release method and harvested 40 min after release. At 35 min post-release, cells were treated with 5% HD for 5 min prior to collection. d Difference map showing changes in contact scores between 5% HD-treated and untreated mitotic cells. e Relationships between contact scores and genomic distances in mitotic cells either untreated or treated with 5% HD. f Enlarged difference map between 5% HD-treated and untreated mitotic cells for the 1.9 Mb region of chromosome I (dotted box in (d). Border strength scores in the indicated samples and their difference were estimated as described in Fig. 3i. g Quantification of distances between inter-domain (left) and intra-domain (right) probes in mitotic cells either untreated or treated with 5% HD. FISH experiments and data analysis were carried out as described in Fig. 4. Probes #1 and #2 span adjacent condensin domains, while probes #2 and #3 are positioned within a single domain. Mitotic cells were prepared in YEA medium using the cdc25-22 block-release method. The central bar represents the median, with the boxes indicating the upper and lower quartiles. The whiskers extend to data points within 1.5× the interquartile range from the box. P-values were calculated by a two-sided Mann–Whitney U test. Source data are provided as a Source Data file. h Average enrichment of the condensin subunit Cnd2-Pk (left), the mediator subunit Pmc4-Pk (middle), and RNA polymerase II Rpb1 (right) at Ace2 target genes (top) and highly transcribed, condensin-binding genes (bottom) in mitotic cells either untreated or treated with 5% HD.
Effects of HD treatment on mitotic chromosome architecture
We examined how HD treatment affects mitotic chromosome architecture. To this end, we performed in situ Hi-C experiments using mitotic cells treated with 10% HD (Supplementary Fig. 11c–f). Treatment with 10% HD increased genomic contacts over distances of 75 kb or greater (Supplementary Fig. 11d, e). We also observed that genomic contacts within condensin domains were relatively unaffected, whereas genomic contacts across domain boundaries were enhanced by HD treatment (Supplementary Fig. 11f). These results indicate that 10% HD treatment weakens domain boundaries, yet chromosome compaction proceeds. Consistently, FISH results indicated that the distances between the inter-domain probes were shorter in mitotic cells treated with 10% HD, compared to those without HD treatment (Supplementary Fig. 11g). Moreover, the distances between the intra-domain probes were comparatively less affected (Supplementary Fig. 11g).
Because 10% HD treatment can perturb mediator as well as numerous other proteins and biological processes, we applied a milder 5% HD treatment to probe its effects on mitotic chromosome architecture and on the distribution of mediator and condensin. We performed in situ Hi-C analysis using mitotic cells treated with 5% HD and observed that 5% HD treatment increased genomic contacts between 75 to 800 kb (Fig. 6c–f). Especially, genomic contacts across domain boundaries were clearly enhanced by HD treatment (Fig. 6f). Consistently, FISH results showed that the distances between the inter-domain probes were shorter in mitotic cells treated with 5% HD than in untreated control cells (Fig. 6g). ChIP-seq analysis further demonstrated that enrichment of Cnd2-Pk and Pmc4-Pk was reduced in mitotic cells treated with 5% HD (Fig. 6h and Supplementary Fig. 11h–k). Notably, Pol II occupancy declined at mitotically activated Ace2-target genes and highly transcribed genes, but enrichment near the 3’ ends of these genes was relatively preserved, suggesting that a brief 5-minute pulse treatment with 5% HD partially lowers transcription frequency and turnover (Fig. 6h and Supplementary Fig. 11h–k). Taken together, these results suggest that 5% HD treatment diminishes mediator and condensin localization at their target genes, particularly mitotically activated Ace2-target genes, and weakens domain boundaries (“Discussion”).
Potential conservation of condensin-mediator interaction in human cells
Since condensin and mediator are conserved in both fission yeast and humans, the condensin-mediator interaction might also be conserved in human cells. To test this possibility, we employed the Y2H assay to examine the potential interaction. Human cells contain two condensin complexes, condensin I and II, and we tested the interactions of MED4 (human ortholog of Pmc4) with CAP-D2 and CAP-D3, the human homologs of Cnd1 in condensin I and II, respectively (Supplementary Fig. 12a). Interestingly, we observed the MED4 interaction with CAP-D3 (condensin II subunit) but not with CAP-D2 (condensin I subunit). Moreover, the interaction was disrupted by the CAP-D3-V628E mutation, which corresponds to the fission yeast cnd1-K658E mutation, and by the CAP-D3-∆613-635 deletion that encompasses the point mutation (Supplementary Fig. 12b).
In addition, mitotic defects, such as misaligned chromosomes during metaphase and micronuclei, were detected in human U2OS cells expressing the CAP-D3-V628E, and the frequency of these mitotic defects was significantly higher in mitotic cells expressing CAP-D3-V628E than those expressing the wild-type CAP-D3 (Supplementary Fig. 12c, d). Furthermore, misaligned chromosomes and micronuclei were also detected in human 293FT cells depleted of MED4, and the frequency of these mitotic defects was significantly increased in MED4-depleted cells compared to controls (Supplementary Fig. 12e–g). Based on these results, we speculate that a mitotic mechanism analogous to chromosome organization in fission yeast, involving the condensin-mediator interaction, may also function in human cells.
Discussion
Condensin recruitment through interactions with mediator and TBP
Our ChIP-seq data indicate that condensin strongly co-localizes with the Pmc4 mediator subunit and Tbp1 (fission yeast TBP) at highly transcribed genes and mitotically activated Ace2 target genes (Supplementary Fig. 1). While condensin preferentially accumulates at the 3’ ends of these genes, Pmc4 and Tbp1 enrichment peaks are located at gene promoters. Mechanistically, the cnd1-K658E and cnd2-C703R condensin mutations disrupt the condensin interactions with Pmc4 and Tbp1, respectively, and diminish condensin localization at its target genes (Fig. 5 and Supplementary Fig. 8)49. These results suggest that mediator and TBP contribute to condensin recruitment at promoters, after which transcription guides its redistribution along gene bodies toward the 3’ end. In this regard, depletion of Pol II has been reported to diminish condensin association with Pol II-transcribed genes, supporting the hypothesis that Pol II can stall condensin at gene regions65. Moreover, a similar mechanism has been reported for cohesin, which is relocated by transcription66. Although the precise coordination between the Cnd1-Pmc4 and Cnd2-Tbp1 interactions remains unclear, it is notable that TBP within the TFIID complex and Med4 (Pmc4) within the mediator complex are positioned in proximity to where Pol II transcription initiates67,68. Therefore, we speculate that the Cnd1-Pmc4 and Cnd2-Tbp1 interactions may jointly facilitate condensin recruitment to gene promoters.
Mitotic chromatin boundaries formed by mediator-driven transcription
In the cnd1-K658E and cnd2-C703R mutants, condensin-mediated genomic contacts within mitotic chromatin domains are disrupted (Fig. 3d and Supplementary Figs. 3e; 4c), likely due to diminished condensin localization at highly transcribed genes and mitotically activated genes across the genome (Fig. 5 and Supplementary Fig. 8). In contrast, Pmc4 mediator depletion leads to increased inter-domain genomic contacts while imparing condensin-mediated genomic contacts and mitotic chromatin domains (Fig. 3i). The reduction in condensin-mediated genomic contacts and chromatin domains likely result from compromised Pmc4-dependent condensin loading; however, residual condensin at gene regions may still mediate inter-domain genomic contacts in Pmc4-depleted cells (Fig. 5). Moreover, Pmc4 depletion, but not the cnd1-K658E mutation, diminishes expression of Ace2 target genes located at chromatin boundaries and weakens boundary integrity (Supplementary Fig. 9c, f). These results support a model in which mediator-driven transcription of Ace2 target genes contributes to the formation of mitotic chromatin domain boundaries.
Potential involvement of phase separation in mitotic chromosome organization
Mediator was initially identified as a component of the RNA polymerase II holoenzyme complex and is known to be involved in transcriptional initiation and repression69,70. Mediator also plays a role in transcriptional regulation in fission yeast58. It has recently been proposed that the mediator engages in transcription via phase separation62,63, prompting us to investigate its potential involvement in mitotic chromosome organization via phase separation. In human cells, Med1 is known to form liquid droplets. In contrast, Pmc4 (fission yeast Med4) contains an IDR at its C-terminal portion. Furthermore, HD treatment disrupts mediator localization in the nucleus, implying that the fission yeast mediator is regulated through phase separation. Because our in vivo data suggest that the disordered region of Pmc4 is unlikely to be solely responsible for mediator droplet formation, we predict that Med1 may mediate droplet formation in fission yeast as observed in human cells62,63. Consistently, we find that mitotic chromosome architecture is disrupted in both Pmc4-depleted and HD-treated cells, where chromosome compaction still occurs but to a lesser degree, and domain boundaries are disrupted (Figs. 3i and 6f; Supplementary Fig. 11f). Mechanistically, HD treatment interferes with Pmc4 mediator binding to mitotically activated Ace2-target genes, and Pmc4 depletion diminishes transcription of these genes located at domain boundaries (Fig. 6h and Supplementary Fig. 9i). These results collectively suggest that mediator-dependent transcription of mitotically activated genes involves a phase separation process, which may contribute to the establishment of mitotic chromatin domain boundaries.
Cooperation of condensin and mediator to form mitotic chromosome architecture
By comparing Hi-C data from Cnd1-Pmc4 double depletion with that from Cnd1 or Pmc4 single depletion, we find that condensin primarily contributes to genomic contacts within approximately 800 kb, whereas mediator appears to exert an inhibitory effect on genomic contacts within roughly 800 kb (Supplementary Fig. 6). Notably, genomic contacts within this range can still form in the absence of condensin when mediator is depleted. In this context, it has previously been proposed that mitotic chromosomal compaction can occur through a condensin-independent mechanism, including post-translational modifications of histones, particularly histone deacetylation71,72,73. Our results are consistent with the idea that the mediator may inhibit this condensin-independent compaction process. In support of this notion, prior studies have shown that growth defects caused by condensin mutations can be partially suppressed by mutations in mediator components59,60. Our findings further support that condensin mutations may be partly suppressed because mediator mutations can enhance condensin-independent chromosome compaction. Although the detailed mechanisms remain to be fully elucidated, this study highlights that condensin-mediated contacts are demarcated by mediator-driven transcription of mitotically activated genes located at boundaries, while mediator also appears to limit condensin-independent long-range genomic contacts.
Taking all these results together, our mechanistic view is as follows: (1) Mediator and TBP recruit condensin to highly transcribed genes and mitotically activated Ace2 target genes; (2) Condensin at gene regions mediates genomic contacts; (3) Mediator activates transcription of Ace2 target genes during mitosis likely through phase separation; (4) Transcription of Ace2-target genes defines the boundaries of condensin-mediated genomic contacts; and (5) Domain-based chromosome compaction promotes faithful chromosome segregation.
Methods
Fission yeast strains and culture conditions
C-terminal tagging of Cnd1, Cnd2, and Pmc4 with Myc, Flag, or Pk was performed using a PCR-based module method74,75. The tagged proteins were expressed from their endogenous genomic loci under the control of their native promoters. DNA constructs for gene deletions, point mutations, and auxin-inducible degron (AID)-based targeted protein degradation (Cnd1-IAA17 and Pmc4-IAA17) were generated by PCR and introduced into fission yeast (Schizosaccharomyces pombe) cells by standard chemical transformation. Additional strains were obtained through conventional genetic crosses. Cells were cultured in either yeast extract with adenine (YEA) or in Edinburgh minimal medium (EMM).
Auxin-inducible degron (AID) approach for targeted protein degradation
The AID system was used as described previously76,77. The Cnd2-AID strain was constructed and used as previously described49. A Skp1-fused F-box protein (atAFB2, atTIR1, and/or osTIR1) containing a nuclear localization signal and a 9×Myc epitope tag was expressed from the ade6 locus under the control of the adh1 promoter. Cnd1 fused to the IAA17 degron was expressed from the endogenous cnd1 locus under the control of the endogenous cnd1 promoter. To induce post-translational degradation of endogenous Cnd1 proteins, cells were cultured in YEA or EMM medium supplemented with 0.5 mM auxin [1-naphthaleneacetic acid (Sigma)]. Cnd1 depletion efficiency was evaluated by western blotting using an anti-mini-AID-tag antibody (clone IE4, MBL). When wild-type or mutant Cnd1 proteins under the control of the endogenous cnd1 promoter were expressed from LEU2-based plasmids, EMM-Leu medium containing 0.5 mM auxin was used to maintain the LEU2 plasmids. For depletion of endogenous Cnd1 in the absence of plasmid-derived Cnd1, cells were cultured in YEA medium containing 0.5 mM auxin.
Pmc4 fused to the IAA17 degron was expressed from the endogenous pmc4 gene locus. For Pmc4 depletion, cells were cultured in YEA or EMM medium supplemented with 0.5 mM auxin. When wild-type Pmc4 proteins were expressed from a LEU-based plasmid, EMM-Leu medium containing 0.5 mM auxin was used. For depletion of endogenous Pmc4 in the absence of plasmid-derived Pmc4, cells were cultured in YEA medium containing 0.5 mM auxin.
Generation of cnd1 mutations using yeast two-hybrid (Y2H) approach
A schematic procedure is shown in Supplementary Fig. 2a. A genetic screen was performed using the Matchmaker Gold Y2H system (Clontech) to generate cnd1 mutations that disrupt the Cnd1-Pmc4 interaction without affecting the Cnd1-Cnd2 interaction between the condensin subunits. To this end, the cnd1+ gene was first amplified by error-prone PCR78, and the mutated cnd1 gene was cloned into pGBKT7, which carries the TRP1 marker gene, enabling Mat α budding yeast cells to express GAL4-BD fused to the mutant Cnd1 (GAL4-BD-mutant Cnd1). The pGADT7 derivatives expressing GAL4-AD fused to either Pmc4 or Cnd2 (GAL4-AD-Pmc4 or GAL4-AD-Cnd2), along with the LEU2 marker gene, were independently transformed into the Mat a Y2HGold budding yeast strain (Clontech). After mating the Mat α and Mat a strains on YEA plates, diploid cells carrying both pGBKT7 and pGADT7 derivatives were selected by replica plating onto Synthetic Defined (SD) -Trp (tryptophan) -Leu (leucine) plates. Protein interactions were assessed by monitoring growth on SD -Trp -Leu -His (histidine) -Ade (adenine) plates containing 125 ng/ml aureobasidin A, which indicates expression of the HIS3, ADE2, and AUR1-C reporter genes. More than 3000 diploid colonies that grew on SD-Trp-Leu plates were screened, and the GAL4-BD-Cnd1-m1 clone was identified. Notably, diploid cells carrying pGBKT7 GAL4-BD-Cnd1-m1 and pGADT7 GAL4-AD-Pmc4 failed to grow on SD-Trp-Leu-His-Ade plates containing aureobasidin A, whereas those carrying pGBKT7 GAL4-BD-Cnd1-m1 and pGADT7 GAL4-AD-Cnd2 were able to grow, indicating that the cnd1-m1 allele selectively disrupts the Cnd1 interaction with Pmc4. The cnd1-m1 allele, which harbors several point mutations, was identified by Sanger sequencing. To dissect the contribution of individual mutations, a series of pGBKT7 GAL4-BD-mutant Cnd1 constructs carrying individual point mutations was generated (Supplementary Fig. 2b).
Co-immunoprecipitation (co-IP)
Co-IP experiments were performed as previously described49. Logarithmically growing fission yeast cells cultured in 10 ml YEA (1 × 108 cells at OD595 = 0.5) were collected and washed with 1 ml ice-cold H2O. Cell pellets were resuspended in 500 μl of IP buffer 1 [50 mM HEPES pH 7.6, 125 mM KCl, 0.1% NP-40, 20% glycerol, 1 mM EDTA, 1 mM PMSF, and Complete protease inhibitor cocktail (Sigma)] and disrupted using Mini-Beadbeater-16 (BioSpec Products). The soluble fraction was collected by centrifugation, and MgCl2 (final 5 mM) and 5 units of RQ1 RNase-free DNase I (Promega) were added. After incubation at 37 °C for 30 min, the DNase I reaction was terminated by adding EDTA (final 10 mM). After DNase I treatment, the soluble fraction was mixed with Dynabeads Protein G (Life Technologies) coupled with mouse monoclonal anti-Pk antibody (BioRad) and incubated at 4 °C for 2 h. The beads were washed five times, each for 5 min, with 800 μl of IP buffer 2 (50 mM HEPES pH 7.6, 125 mM KCl, 0.2% NP-40, 20% glycerol, 1 mM EDTA, and 1 mM PMSF). Proteins were eluted by boiling in 10 μl of 1× Laemmli sample buffer.
GST pull-down
The GST pull-down experiment was performed as previously described79. Full-length Pmc4 and Cnd1 were amplified by PCR and cloned into pFastBacGST (Fisher Scientific) and pFastBacHTB (Fisher Scientific), respectively. GST-Pmc4 and His-Cnd1 proteins were co-expressed in baculovirus-infected Sf9 insect cells and purified using GST pull-down. The resulting sample was subjected to size exclusion chromatography using a Superose 6 column (Sigma). Fractions were separated by SDS-PAGE, and proteins were stained with Coomassie Brilliant Blue G-250 (Sigma).
Immunofluorescence (IF) microscopy
Fission yeast cells (1 × 108 cells at OD595 = 0.5) were fixed with 3% paraformaldehyde (pFA) at 26 °C for 10 min, and fixation was quenched by incubation with glycine (final 0.25 M) for 5 min. Fixed cells were washed with PEM buffer (100 mM PIPES pH 6.9, 1 mM MgCl2, and 1 mM EGTA), incubated in PEMS buffer (100 mM PIPES pH 6.9, 1 mM MgCl2, 1 mM EGTA, and 1 M sorbitol) at room temperature for 5 min, and permeabilized with 1 mg/mL Zymolyase 100 T (Seikagaku) in PEMS buffer at 37 °C for 30 min. Cells were treated with PEMS buffer containing 1% Triton X-100 at room temperature for 1 min, washed twice with PEM buffer, and incubated in PEMBAL buffer (100 mM PIPES pH 6.9, 1 mM MgCl2, 1 mM EGTA, 1 % BSA, and 0.1 M L-lysine) at room temperature for 1 h with gentle rotation.
Primary antibodies, mouse monoclonal antibody [anti-Myc (Takara Bio), anti-Pk (BioRad), anti-Pol II antibody CTD4H8 (Abcam), or anti-tubulin antibody TAT-180], and/or rabbit polyclonal antibody [anti-Myc (Abcam) or anti-Cnd2 (Cosmo Bio USA)] were added at 1/1000 dilution in PEMBAL buffer, and cells were incubated at room temperature overnight. After three washes with PEMBAL buffer, secondary antibodies [Alexa Fluor 488-conjugated anti-rabbit IgG (Life Technologies) and/or Cy3-conjugated anti-mouse IgG (Jackson ImmunoResearch)] were added at 1/1000 dilution in PEMBAL buffer, and cells were incubated at room temperature for 3 h. Cells were washed three times with PEMBAL buffer at room temperature for 15 min each with gentle rotation.
Prior to imaging, DNA was stained with 1 μg/mL DAPI (Fisher) in PBS solution, and stained cells were mounted on coverslips using antifade solution (1 mg/ml p-phenylenediamine, 90% glycerol, and 100 mM Tris pH 8.0). Fluorescent signals were detected using a DeltaVision deconvolution microscope (GE Healthcare). Immunofluorescence experiments were repeated at least three times, and more than 30 mitotic cells were analyzed per condition unless otherwise noted, with a total of more than 100 mitotic cells examined. Super-resolution images were acquired using Zeiss LSM 880 with Airyscan running ZEN v. 2.3 (Zeiss). Imaging was conducted using the 405 nm diode laser for DAPI excitation, the 488 nm argon laser for AlexaFluor 488, and the 561 nm DPSS laser for Cy3. Three-dimensional Airyscan processing was performed using ZEN v2.3 using “Auto” filter strength setting.
Fluorescence in situ hybridization (FISH) microscopy
FISH probes were prepared using PCR-amplified DNA fragments as described previously81. Probes for specific genomic loci are indicated in Fig. 4a and Supplementary Fig. 7a. To detect a specific condensin domain on chromosome 1 of fission yeast, five pairs of primers were designed to amplify five 5 kb DNA fragments spanning the target domain (Supplementary Fig. 7c). PCR products were digested with 5 restriction enzymes (AluI, RsaI, DdeI, HaeIII, and BfuCI) and labeled with Cy3-dCTP (GE HealthCare) or Cy5-dCTP (GE HealthCare) using a random primer DNA labeling kit (Takara). A total of 100 ng labeled probes were suspended in hybridization buffer (50% formamide, 2×SSC, 5×Denhardt’s solution, and 10% dextran).
FISH experiments were performed as previously described82. Exponentially growing cells (approximately 1 × 108 cells at OD595 = 0.5) were processed for immunofluorescence with anti-tubulin TAT1 antibody as described above. After washing with PEMBAL buffer, cells were rinsed three times with PEM buffer and fixed with 3% pFA at room temperature for 20 min. Cells were incubated with 0.25 M glycine, washed twice with PEM buffer, treated with 0.1 N HCl at room temperature for 5 min, and washed again with PEM buffer. Cellular RNAs were digested with 50 µg RNase A (Takara Bio) in PEMBAL buffer at 37 °C for 2 h, followed by a single wash with PEM buffer. FISH probes were denatured at 75 °C for 15 min and mixed with cells, whose genomic DNA was subsequently denatured at 75 °C for 5 min. Hybridization was carried out at 40 °C for 12 h with gentle rotation. After hybridization, cells were washed three times with 2×SSC at room temperature for 30 min with gentle rotation. DNA was stained with 1 μg/mL DAPI in PBS buffer, and cells were mounted on coverslips with antifade solution. Fluorescent signals were detected using a DeltaVision deconvolution microscope. FISH experiments were repeated at least three times, and more than 30 cells per condition were analyzed unless otherwise noted, with a total of more than 100 cells examined.
ChIP-seq
Fission yeast cells (approximately 5 × 108 cells at OD595 = 0.5) were fixed with 3% pFA at 26 °C for 30 min, washed twice with PBS buffer, and further crosslinked with 10 mM dimethyl adipimidate (Sigma Aldrich) at room temperature for 45 min. After two washes with PBS buffer, cells were lysed using glass beads and Mini-Beadbeater-16 (BioSpec Products) in ChIP lysis buffer [50 mM HEPES pH 7.5, 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate, 0.1 mM PMSF, and EDTA-free Protease Inhibitor Cocktail (Roche)]. Chromatin DNA was sheared into 100–500 bp fragments using Bioruptor UCD-200 (Diagenode), and the soluble fraction was collected by centrifugation at 13,000 rpm for 15 min. A total of 50 μl ChIP lysis buffer containing 5 µl mouse monoclonal antibody [anti-Pk (BioRad), anti-Myc (Takara Bio), anti-Flag (Sigma), or anti-RNA polymerase II C-terminal domain (Abcam)] and 15 µl Dynabeads protein G (Fisher Scientific) was added to the soluble fraction, and the bead suspension was incubated at 4 °C for 2 h with gentle rotation. Beads were washed twice with ChIP lysis buffer, once with ChIP lysis buffer containing 0.65 M NaCl, once with washing buffer (10 mM Tris pH 8.0, 250 mM LiCl, 0.5% NP-40, 0.5% sodium deoxycholate, and 1 mM EDTA), and once with TE (10 mM Tris pH 8.0, and 1 mM EDTA), each for 5 min with gentle rotation. The protein-DNA mixture was eluted by incubating the beads with TES (10 mM Tris pH 8.0, 1 mM EDTA, and 1% SDS) at 65 °C for 30 min with vigorous agitation at 1000 rpm using ThermoMixer (Eppendorf) and subjected to reverse crosslinking at 68 °C overnight in the presence of 0.5 M NaCl. DNA was purified using QIAquick PCR purification kit (Qiagen). Purified DNA was subjected to library construction using NEBNext Ultra II DNA library prep kit (NEB). Sequencing libraries were amplified by PCR using NEBNext Ultra II Q5 Master Mix (NEB) and NEBNext Multiplex Oligos for Illumina (NEB) and sequenced on NextSeq 500, NextSeq 2000, or NovaSeq 6000 (Illumina) to obtain paired-end reads.
ChIP-seq data processing
ChIP-seq data were processed as described previously47,83 with the following modifications. ChIP-seq reads were aligned to the S. pombe genome (2018 version) using Bowtie2 (version 2.4.4) in paired-end mode. Read pairs that were correctly oriented and aligned at the expected distance were extracted using Samtools (version 1.19) with the -f 0 × 2 option. BigWig files were generated using the bamCoverage function in deepTools (version 3.5.4). Normalization was performed to an average signal of 1, excluding chromosome III to minimize the influence of rDNA repeats. For this purpose, the specific options used were “--ignoreForNormalization III --normalizeUsing RPGC --effectiveGenomeSize 10118937”. ChIP-seq peaks were defined by MACS3 (version 3.0.1) with the “bdgpeakcall” subcommand. The option -c 5 or 4, -l 220, -g 300. -c was determined based on the mean value of scores of each sample and a p-value < 0.1. For the ChIP-seq data shown in Supplementary Fig. 8, Cnd2-FLAG enrichment scores were calculated by subtracting the read density of the no-tag control from that of the indicated sample.
In situ Hi-C
Fission yeast cells (approximately 5 × 108 cells at OD595 = 0.5) were fixed with 3% pFA at 26 °C for 10 min and disrupted using glass beads and Mini-Beadbeater-16 (BioSpec Products) in lysis buffer [50 mM HEPES pH 7.5, 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate, 0.1 mM PMSF, and EDTA-free Protease Inhibitor Cocktail (Roche)]. After washing with 1 ml PBS solution, cells were further crosslinked with 3 mM disuccinimidyl glutarate (Fisher Scientific) in 2.5 ml PBS solution at 30 °C for 40 min. Fixed cells were incubated at 62 °C for 7 min with 0.1% SDS and subsequently with 1% Triton-X 100. Permeabilized cells were incubated with 25 units of MboI, HinfI, and MluCI at 37 °C overnight. The resulting pellet containing restriction enzyme-digested DNA fragments was incubated with 150 μM biotin-14-dATP, dCTP, dGTP, and dTTP, and 20 units of Klenow fragment (NEB) at 37 °C for 45 min, followed by proximity ligation using 2500 units of T4 DNA ligase (NEB) at room temperature for 4 h with gentle rotation. After ligation, the pellet was treated with proteinase K at 68 °C overnight in the presence of 0.5 M NaCl. DNA was purified by phenol/chloroform extraction and ethanol precipitation, and sheared into 100–500 bp fragments using Bioruptor UCD-200 (Diagenode). Biotin-labeled DNA fragments were purified using Dynabeads MyOne Streptavidin T1 (Fisher Scientific). While bound to the beads, sequencing adapters were ligated using NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB). Sequencing libraries were amplified by PCR using the NEBNext Ultra II Q5 Master Mix (NEB) and NEBNext Multiplex Oligos for Illumina (NEB). Libraries were sequenced on NextSeq 500, NextSeq 2000, or NovaSeq 6000 (Illumina) to obtain paired-end reads.
In situ Hi-C data processing
Data processing was performed using the rfy_hic2 package (https://github.com/rafysta/rfy_hic2)38, which is available at Zenodo (https://doi.org/10.5281/zenodo.18264493). In brief, paired-end sequence reads were individually aligned to the S. pombe genome (2018 version) using an iterative alignment strategy with Bowtie2 (version 2.4.4). Redundant paired reads derived from PCR bias, reads aligned to repetitive sequences, and reads with low mapping quality (MapQ < 30) were removed. Read pairs with inward or outward orientations within 10 kb were excluded, as these are likely to represent undigested products or self-ligation artifacts. Using paired reads aligned in the same orientation, the number of correct Hi-C reads with inward and outward orientations was estimated. The fission yeast genome was divided into non-overlapping bins of 5 kb, 10 kb, and 20 kb, and raw contact matrices were generated by counting the number of read pairs mapping to each bin pair. Hi-C contact map biases were corrected using the ICE method, with normalization repeated 30 times84.
Nascent RNA labeling followed by deep sequencing (nascent RNA-seq)
The nascent RNA detection protocol used in this study was based on the previously reported precision run-on sequencing (PRO-seq) method61 with some modifications. A total of 1 × 108 cells were permeabilized in 0.5% N-Lauroylsarcosine sodium salt (sarkosyl; Sigma) on ice for 20 min, and a nuclear run-on reaction was carried out at 30 °C for 5 min in a reaction buffer [20 mM Tris pH 7.7, 200 mM KCl, 5 mM MgCl2, 1.25 mM dithiothreitol, 0.4 U/µL SUPERase-In RNase Inhibitor (Fisher), 0.5% sarkosyl, 12.5 µM biotin-11-CTP (Enzo Life Sciences), 12.5 µM biotin-11-UTP (Biotium), 62.5 µM ATP (Fisher), and 62.5 µM GTP (Fisher)] to label newly transcribed RNA with biotin. After pulse labeling, cells were disrupted using Mini-Beadbeater-16 in the presence of AES buffer [50 mM sodium acetate pH 5.5 (Fisher), 10 mM EDTA, and 1% SDS], neutral Phenol Chloroform Isoamyl alcohol (PCI), and glass beads. Nucleic acids were purified by additional neutral PCI extraction followed by ethanol precipitation. RNA was heat-denatured at 65 °C for 1 min and fragmented by treatment with 0.2 N sodium hydroxide (NaOH; Fisher) on ice for 10 min. After neutralization with 0.5 M Tris pH 6.8, fragmented RNA was purified with RNeasy Mini Kit (Qiagen), and residual DNA was removed by treatment with 0.05 U/µL RQ1 RNase-Free DNase (Promega) at 37 °C for 30 min, followed by neutral PCI extraction and ethanol precipitation. Biotinylated RNA was selectively pulled down using Dynabeads MyOne Streptavidin T1 magnetic beads (Fisher) at 25 °C for 15min, and first- and second-strand cDNA syntheses, end repair, and adaptor ligation reactions were performed sequentially using NEBNext Ultra II RNA Library Prep Kit for Illumina (NEB), according to the manufacturer's instructions. Adaptor-ligated cDNA was amplified using NEBNext Ultra II Q5 Master Mix and NEBNext Multiplex Oligos for Illumina. PCR-amplified libraries were purified with 0.9 × volume of AMPure XP before sequencing runs on NextSeq 500 or NextSeq 2000 to generate paired-end reads.
Nascent RNA-seq analysis
Sequenced reads were aligned to the Schizosaccharomyces pombe genome (2018 version) using STAR version 2.7.685. Reads assigned to exons were quantified using RSEM version 1.3.386. Normalization of read numbers and identification of differentially expressed genes, comparing cells expressing WT or mutant proteins, as well as cells expressing WT proteins versus target protein-depleted cells, were performed using edgeR version 4.0.1687. The dispersion values for Cnd1 expression (vector, WT, and K658E mutant) and Pmc4 (WT and depletion) were 0.017 and 0.037, respectively. FDR < 0.05 was used as the threshold to define significantly up- or down-regulated genes.
Pmc4 purification from E. coli
To induce expression of GST-mCherry-Pmc4, Escherichia coli BL21 cells carrying the pGEX6P-mCherry-Pmc4 plasmid were cultured in LB medium supplemented with 10 µM IPTG (Wako) at 16 °C for 20 h. Cellular proteins were extracted with extraction buffer (50 mM Tris pH 8.0, 500 mM NaCl, 10 mM β-mercaptoethanol, and 10% glycerol), and affinity purification was performed using GST-accept resin (Nakarai). Bound proteins were eluted with glutathione buffer [10 mM reduced glutathione (Wako), 50 mM Tris pH 8.0, 500 mM NaCl, 10 mM β-mercaptoethanol, and 10% glycerol]. Purified proteins were treated with precision protease at 4 °C for 16 h, followed by desalting using a HiTrap Q HP column (GE Healthcare) and removal of cleaved GST using GST-accept resin. The buffer was exchanged to LLPS buffer (50 mM Tris pH 8.0, 100 mM NaCl, and 10 mM β-mercaptoethanol) using VivaSpin 6 Centrifugal Concentrators (Sartorius). For droplet formation assays, the purified protein solution was mixed at a 1:1 ratio with PEG3350 buffer (100 µM Bis-Tris pH 6.0 containing the indicated concentrations of PEG3350). Droplet formation was observed using a Leica STELLARIS 5 confocal microscope equipped with HC PL APO CS2 63×/1.40 OIL objective lens (Leica). Images were captured and analyzed using LAS × 4.4.0.24861 software (Leica).
Human cell lines and culture conditions
Plasmids expressing eGFP-CAP-D3-WT or eGFP-CAP-D3-V628E under the control of the TRE3GS doxycycline-inducible promoter were transfected into U2OS cells along with pTORA14_AAVS using Lipofectamine LTX Reagent (Fisher Scientific). To establish stable cell lines, transfected cells were cultured in DMEM supplemented with 10% FBS and 10 μg/mL puromycin for 14 days, with the medium replaced every 3 days. For MED4 depletion, 293FT cells were transfected with TRIPZ plasmids expressing MED4 shRNAs and cultured in DMEM supplemented with 10% FBS and 10 μg/mL puromycin for 48 h.
DAPI staining with human cells
To observe mitotic chromosomes and micronuclei, U2OS cells stably expressing eGFP constructs (eGFP-CAP-D3-WT or eGFP-CAP-D3-V628E) or 293FT cells with or without MED4 knockdown were cultured on cover glasses. For eGFP-CAP-D3 expression, doxycycline (2 µg/mL) was added 24 h before fixation. Cells were fixed with 4% pFA in PBS buffer at 37 °C for 10 min, followed by permeabilization with 0.15% Triton-X in PBS buffer at room temperature for 2 min. After washing with PBS buffer, cells were mounted on glass slides using an Antifade mounting medium (Vectashield) containing 1 μg/mL DAPI. Images were acquired using a confocal microscope (Olympus, FV1000) with a 60× lens.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ChIP-seq, Hi-C and nascent RNA-seq data reported in this paper have been deposited in NCBI’s Gene Expression Omnibus with the GEO series accession codes GSE270574, GSE270576 and GSE270577. Source data are provided with this paper.
References
Koshland, D. & Strunnikov, A. Mitotic chromosome condensation. Annu. Rev. Cell Dev. Biol. 12, 305–333 (1996).
Hirano, T. Chromosome cohesion, condensation, and separation. Annu. Rev. Biochem 69, 115–144 (2000).
Nasmyth, K. Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet 35, 673–745 (2001).
Yanagida, M. Clearing the way for mitosis: is cohesin a target? Nat. Rev. Mol. Cell Biol. 10, 489–496 (2009).
Hagstrom, K. A. & Meyer, B. J. Condensin and cohesin: more than chromosome compactor and glue. Nat. Rev. Genet. 4, 520–534 (2003).
Losada, A. & Hirano, T. Dynamic molecular linkers of the genome: the first decade of SMC proteins. Genes Dev. 19, 1269–1287 (2005).
Michaelis, C., Ciosk, R. & Nasmyth, K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 91, 35–45 (1997).
Guacci, V., Koshland, D. & Strunnikov, A. A direct link between sister chromatid cohesion and chromosome condensation revealed through the analysis of MCD1 in S. cerevisiae. Cell 91, 47–57 (1997).
Tomonaga, T. et al. Characterization of fission yeast cohesin: essential anaphase proteolysis of Rad21 phosphorylated in the S phase. Genes Dev. 14, 2757–2770 (2000).
Saka, Y. et al. Fission yeast cut3 and cut14, members of a ubiquitous protein family, are required for chromosome condensation and segregation in mitosis. EMBO J. 13, 4938–4952 (1994).
Freeman, L., Aragon-Alcaide, L. & Strunnikov, A. The condensin complex governs chromosome condensation and mitotic transmission of rDNA. J. Cell Biol. 149, 811–824 (2000).
Lavoie, B. D., Tuffo, K. M., Oh, S., Koshland, D. & Holm, C. Mitotic chromosome condensation requires Brn1p, the yeast homologue of Barren. Mol. Biol. Cell 11, 1293–1304 (2000).
Ouspenski, I. I., Cabello, O. A. & Brinkley, B. R. Chromosome condensation factor Brn1p is required for chromatid separation in mitosis. Mol. Biol. Cell 11, 1305–1313 (2000).
Hirano, T. Condensins: universal organizers of chromosomes with diverse functions. Genes Dev. 26, 1659–1678 (2012).
Uhlmann, F. SMC complexes: from DNA to chromosomes. Nat. Rev. Mol. Cell Biol. 17, 399–412 (2016).
Noma, K. -i The yeast genomes in three dimensions: mechanisms and functions. Annu. Rev. Genet. 51, 23–44 (2017).
Davidson, I. F. & Peters, J. M. Genome folding through loop extrusion by SMC complexes. Nat. Rev. Mol. Cell Biol. 22, 445–464 (2021).
Ong, C. T. & Corces, V. G. CTCF: an architectural protein bridging genome topology and function. Nat. Rev. Genet. 15, 234–246 (2014).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Szabo, Q., Bantignies, F. & Cavalli, G. Principles of genome folding into topologically associating domains. Sci. Adv. 5, eaaw1668 (2019).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. USA 112, E6456–E6465 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Ryu, J. K. et al. Bridging-induced phase separation induced by cohesin SMC protein complexes. Sci. Adv. 7, eabe5905 (2021).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Pope, B. D. et al. Topologically associating domains are stable units of replication-timing regulation. Nature 515, 402–405 (2014).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Heinz, S. et al. Transcription elongation can affect genome 3d structure. Cell 174, 1522–1536 e22 (2018).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Mizuguchi, T. et al. Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe. Nature 516, 432–435 (2014).
Kim, K. D., Tanizawa, H., Iwasaki, O. & Noma, K. Transcription factors mediate condensin recruitment and global chromosomal organization in fission yeast. Nat. Genet. 48, 1242–1252 (2016).
Van Bortle, K. et al. Insulator function and topological domain border strength scale with architectural protein occupancy. Genome Biol. 15, R82 (2014).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Kakui, Y., Rabinowitz, A., Barry, D. J. & Uhlmann, F. Condensin-mediated remodeling of the mitotic chromatin landscape in fission yeast. Nat. Genet. 49, 1553–1557 (2017).
Tanizawa, H., Kim, K.-D., Iwasaki, O. & Noma, K. -i Architectural alterations of the fission yeast genome during the cell cycle. Nat. Struct. Mol. Biol. 24, 965–976 (2017).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359, eaao6135 (2018).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Kim, E., Kerssemakers, J., Shaltiel, I. A., Haering, C. H. & Dekker, C. DNA-loop extruding condensin complexes can traverse one another. Nature 579, 438–442 (2020).
Tang, M. et al. Establishment of dsDNA-dsDNA interactions by the condensin complex. Mol. Cell 83, 3787–3800 e9 (2023).
Dowen, J. M. et al. Multiple structural maintenance of chromosome complexes at transcriptional regulatory elements. Stem Cell Rep. 1, 371–378 (2013).
Li, W. et al. Condensin I and II complexes license full estrogen receptor alpha-dependent enhancer activation. Mol. Cell 59, 188–202 (2015).
Hocquet, C. et al. Condensin controls cellular RNA levels through the accurate segregation of chromosomes instead of directly regulating transcription. eLife 7, e38517 (2018).
Swygert, S. G. et al. Condensin-dependent chromatin compaction represses transcription globally during quiescence. Mol. Cell 73, 533–546.e4 (2019).
Iwasaki, O. et al. Involvement of condensin in cellular senescence through gene regulation and compartmental reorganization. Nat. Commun. 10, 5688 (2019).
Lancaster, L., Patel, H., Kelly, G. & Uhlmann, F. A role for condensin in mediating transcriptional adaptation to environmental stimuli. Life Sci. Alliance 4, e202000961 (2021).
Iwasaki, O. et al. Interaction between TBP and condensin drives the organization and faithful segregation of mitotic chromosomes. Mol. Cell 59, 755–767 (2015).
Kornberg, R. D. Mediator and the mechanism of transcriptional activation. Trends Biochem. Sci. 30, 235–239 (2005).
Malik, S. & Roeder, R. G. The metazoan mediator co-activator complex as an integrative hub for transcriptional regulation. Nat. Rev. Genet. 11, 761–772 (2010).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Rustici, G. et al. Periodic gene expression program of the fission yeast cell cycle. Nat. Genet. 36, 809–817 (2004).
Petit, C. S., Mehta, S., Roberts, R. H. & Gould, K. L. Ace2p contributes to fission yeast septin ring assembly by regulating mid2+ expression. J. Cell Sci. 118, 5731–5742 (2005).
Nakazawa, N. et al. Dissection of the essential steps for condensin accumulation at kinetochores and rDNAs during fission yeast mitosis. J. Cell Biol. 180, 1115–1131 (2008).
Onn, I., Aono, N., Hirano, M. & Hirano, T. Reconstitution and subunit geometry of human condensin complexes. EMBO J. 26, 1024–1034 (2007).
Tsai, K. L. et al. Mediator structure and rearrangements required for holoenzyme formation. Nature 544, 196–201 (2017).
Linder, T. et al. Two conserved modules of Schizosaccharomyces pombe mediator regulate distinct cellular pathways. Nucleic Acids Res. 36, 2489–2504 (2008).
Robellet, X. et al. A genetic screen for functional partners of condensin in fission yeast. G3 4, 373–381 (2014).
Sutani, T. et al. Condensin targets and reduces unwound DNA structures associated with transcription in mitotic chromosome condensation. Nat. Commun. 6, 7815 (2015).
Mahat, D. B. et al. Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq). Nat. Protoc. 11, 1455–1476 (2016).
Cho, W.-K. et al. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361, 412–415 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 e16 (2018).
Lebreton, J., Colin, L., Chatre, E. & Bernard, P. RNAP II antagonizes mitotic chromatin folding and chromosome segregation by condensin. Cell Rep. 43, 113901 (2024).
Lengronne, A. et al. Cohesin relocation from sites of chromosomal loading to places of convergent transcription. Nature 430, 573–578 (2004).
Schilbach, S. et al. Structures of transcription pre-initiation complex with TFIIH and mediator. Nature 551, 204–209 (2017).
Richter, W. F., Nayak, S., Iwasa, J. & Taatjes, D. J. The Mediator complex as a master regulator of transcription by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 23, 732–749 (2022).
Kim, Y. J., Bjorklund, S., Li, Y., Sayre, M. H. & Kornberg, R. D. A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell 77, 599–608 (1994).
Ding, N. et al. Mediator links epigenetic silencing of neuronal gene expression with X-linked mental retardation. Mol. Cell 31, 347–359 (2008).
Hudson, D. F., Vagnarelli, P., Gassmann, R. & Earnshaw, W. C. Condensin is required for nonhistone protein assembly and structural integrity of vertebrate mitotic chromosomes. Dev. Cell 5, 323–336 (2003).
Samejima, K. et al. Functional analysis after rapid degradation of condensins and 3D-EM reveals chromatin volume is uncoupled from chromosome architecture in mitosis. J. Cell Sci. 131, jcs210187 (2018).
Schneider, M. W. G. et al. A mitotic chromatin phase transition prevents perforation by microtubules. Nature 609, 183–190 (2022).
Bahler, J. et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943–951 (1998).
Gadaleta, M. C., Iwasaki, O., Noguchi, C., Noma, K. & Noguchi, E. New vectors for epitope tagging and gene disruption in Schizosaccharomyces pombe. Biotechniques 55, 257–263 (2013).
Kanke, M. et al. Auxin-inducible protein depletion system in fission yeast. BMC Cell Biol. 12, 8 (2011).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
McCullum, E. O., Williams, B. A., Zhang, J. & Chaput, J. C. Random mutagenesis by error-prone PCR. Methods Mol. Biol. 634, 103–109 (2010).
Ricketts, M. D. et al. Ubinuclein-1 confers histone H3.3-specific binding by the HIRA histone chaperone complex. Nat. Commun. 6, 7711 (2015).
Woods, A. et al. Definition of individual components within the cytoskeleton of Trypanosoma brucei by a library of monoclonal antibodies. J. Cell Sci. 93, 491–500 (1989).
Iwasaki, O., Corcoran, C. J. & Noma, K. Involvement of condensin-directed gene associations in the organization and regulation of chromosome territories during the cell cycle. Nucleic Acids Res. 44, 3618–3628 (2016).
Kim, K. D., Iwasaki, O. & Noma, K. An IF-FISH approach for covisualization of gene loci and nuclear architecture in fission yeast. Methods Enzymol. 574, 167–180 (2016).
Tanaka, A. et al. Epigenetic regulation of condensin-mediated genome organization during the cell cycle and upon DNA damage through histone H3 Lysine 56 acetylation. Mol. Cell 48, 532–546 (2012).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Acknowledgements
We would like to thank the Yeast Genetic Resource Center (Osaka City University) for the pTN-TH7 cDNA library and Drs. Keith Gull and Jack Sunter for the anti-tubulin TAT1 antibody, and the University of Oregon Genomics & Cell Characterization Core Facility for next-generation sequencing and microscopic analysis. We also thank Drs. Jeannie and Eric Selker for scientific and humane guidance, and Ms. Yuko Tsukamoto for technical assistance. This work was supported by the National Institutes of Health/National Institute of General Medical Sciences R01GM124195, National Institutes of Health/National Institute of Aging P01AG AG031862, JSPS KAKENHI grants JP20K23376 and JP25K02249, Takeda Science Foundation, Grant for Basic Science Research Projects from The Sumitomo Foundation, and The Mitsubishi Foundation (K.N.). Finally, we sincerely appreciate the generous support from the Medical Research Foundation, the University of Oregon OVPRI Bridge Funding program, and the University of Oregon Research Resilience Committee during a particularly challenging period.
Author information
Authors and Affiliations
Contributions
O.I., S.T., X.W., S.O., Y.F., J.H., G.T., and M.K. conducted genetics, genomics, and/or cell biology experiments. C.Y.C., T.H., and H.T. performed bioinformatics analyses. R.M. and N.N.N. provided project supervision. K.N. conceived and designed the study and was in charge of the overall direction. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Shiv Grewal and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Iwasaki, O., Tashiro, S., Chung, C.YL. et al. A role for condensin-mediator interaction in mitotic chromosome organization. Nat Commun 17, 2509 (2026). https://doi.org/10.1038/s41467-026-69270-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69270-x