Abstract
Aerobic glycolysis and lactate have been shown to modulate tumor microenvironment (TME) and disease progression. Lactate-mediated histone lysine lactylation (Kla) is a newly recognized epigenetic modification whose biological function remains poorly understood. Here, through integrated bioinformatic and experimental analyses, we demonstrate that glycolysis-derived lactate induces histone H3 lysine 18 lactylation (H3K18la) and up-regulates the expression of chemokine C-X-C motif Ligand 1 (CXCL1), thereby recruiting neutrophils and inducing immunosuppression in pancreatic cancer. Moreover, our data suggest that p300/CBP-associated factor (PCAF) functions as a histone lactyltransferase that transcriptionally activates CXCL1 expression. Finally, we reveal that combinational treatment with bromosporine (a PCAF inhibitor) and anti-PD-1 antibody exhibits a synergistic antitumor effect on both subcutaneous and orthotopic tumor models of pancreatic cancer. Taken together, our study not only identifies a mechanism by which the aerobic glycolysis-induced Lactate-PCAF-H3K18la-CXCL1 pathway mediates neutrophil infiltration and immunosuppression, but also develops a potential therapeutic strategy for pancreatic cancer.
Similar content being viewed by others
Introduction
Pancreatic adenocarcinoma (PAAD), of which over 80% of cases are pancreatic ductal adenocarcinoma (PDAC), is the third-leading cause of cancer-related death due to limited therapeutic options1. Immune escape mediated by an immunosuppressive microenvironment is a key reason for the ineffective immune checkpoint blockade (ICB) therapy in PDAC2,3,4. Neutrophils, the most abundant myeloid cells in human blood and a critical component of the tumor microenvironment (TME), have been suggested to play important roles in tumor progression and therapy resistance5,6,7. Therefore, understanding the immune suppression mechanism may enhance the therapeutic effectiveness of ICB and develop novel therapeutic strategies against PDAC.
Metabolic reprogramming to aerobic glycolysis is a hallmark of tumor cells8. Accumulating evidence suggests that aerobic glycolysis may promote tumor immune escape through metabolic communication between tumor and immune cells, mediated by glycolytic products. For example, tumor-cell-derived lactate stimulates the differentiation of CD4+ T cells9,10, promotes the polarization of tumor-associated macrophage (TAM)9,11, and inhibits natural killer (NK) cells9,12 and effector T cells (Teffs)9,12. Moreover, glycolysis pathways in tumor cells may also regulate the expression of immunomodulators, such as CXCL1013, PD-L114, and TNF-α15. However, the potential roles of these glycolysis pathways in regulating neutrophil-mediated immune escape remain largely unclear.
Epigenetic remodeling has emerged as a newly recognized feature of tumors in recent years16. Histone modification is one of the most fundamental epigenetic types and plays a key role in physiological and pathological conditions17. As the end product of aerobic glycolysis, lactate was first identified as a mediator of histone L-lactylation in 2019, which promotes macrophage polarization18. Histone L-lactylation has been associated with tumor cell proliferation, metastasis, and immune escape in many tumors, such as ocular melanoma, lung cancer, head and neck squamous cell carcinoma, and colorectal cancer19,20,21,22. However, the function of histone L-lactylation in pancreatic cancer remains to be further elucidated. Another pivotal issue related to histone L-lactylation is the identification of its writer. Several histone acetyltransferases, such as P300/CBP18,23 and GCN523,24, along with alanyl-tRNA synthetase, AARS125,26, and AARS227,28 have been shown to function as lactyltransferase; whether there are new lactyltransferases in pancreatic cancer requires further investigation.
In this study, we conduct bioinformatic analysis and experimental studies to assess the effect of aerobic glycolysis on the tumor immune microenvironment, and comprehensively analyze the relationship between the levels of glycolysis and immune cell infiltrations. We reveal that aerobic glycolysis promotes tumor immune evasion by promoting neutrophil infiltration through lactate-mediated H3K18la in pancreatic cancer. In addition, our data suggest that PCAF functions as a potential histone lactyltransferase and directly activates the Cxcl1 expression at the transcriptional level. Furthermore, we demonstrate that a combinational treatment with a PCAF inhibitor and an anti-PD-1 antibody induced a synergistic antitumor efficacy in the orthotopic mouse model of pancreatic cancer. Our study has identified a novel mechanism that the Lactate-PCAF-H3K18la-CXCL1 pathway mediates the immune evasion in pancreatic cancer, which may also help develop novel therapeutic strategies against pancreatic cancer.
Results
Glycolysis mediates neutrophil recruitment and promotes the progression of pancreatic cancer
To evaluate the effect of glycolysis on the tumor immune microenvironment, we first comprehensively analyzed the relationship between the immune cell infiltration and glycolysis. The results showed that neutrophils were enriched in tumor patients with high glycolysis, whereas CD8+ T cells were enriched in patients with low glycolysis (Fig. 1A). Correlation analysis revealed that the infiltration of neutrophils and CD8+ T cells was positively and negatively correlated with glycolysis, respectively (Fig. 1B and Supplementary Fig. S1A). In addition, the levels of glycolysis and neutrophil infiltration were negatively correlated with patient survival probability (Fig. 1C). Interestingly, a worse prognosis was observed in patients with high neutrophil infiltration, along with high CD8+ T cell infiltration, but not in patients with low CD8+ T cell infiltration (Supplementary Fig. S1B). Moreover, in addition to pancreatic cancer, we also observed similar results in other tumor types (Supplementary Fig. S1C–F).
A The expression of neutrophils and CD8+ T cells in low- and high-glycolysis clusters, divided by ssGSEA score of glycolysis-related genes, in TCGA and CPTAC PAAD database. Data are presented as box plots showing the median (center line), the first and third quartiles (box bounds), and the whiskers extend to 1.5 times the interquartile range from the box. B The correlation of glycolysis level and neutrophils or CD8+ T cells expression in TCGA (n = 179 patients) and CPTAC (n = 140 patients) PAAD database. C Kaplan–Meier plots representing survival probabilities in TCGA-PAAD patients according to the relative level of glycolysis-related or neutrophil-related gene expression. D The diagram illustrating the targets of 2DG and shRNA in the glycolysis process. E CyTOF analysis of tumor-infiltrating immunocytes in subcutaneous tumor models with or without 2DG treatment: tSNE plots showing 12 meta-clusters based on the expression of 41 markers for the immunocytes. F Bar chart of the frequencies of the immune cell subsets in two experimental groups. G A schematic diagram showing the orthotopic PDAC model (n = 6 mice per group in one experiment) with or without 2DG treatment. H, I Image and weights of the orthotopic tumors in the experimental groups from (G) at the end of the experiments. (n = 6 mice per group). (J–M) Tumor-infiltrating neutrophils, CD8+ T cells, GZMB+CD8+ T cells, and PD-1+CD8+ T cells isolated from the orthotopic tumors were analyzed using flow cytometry: representative graphs of flow cytometry (left panel) and statistical analysis of the cell ratio (right panel) (n = 6 mice per group). N A schematic diagram showing the orthotopic tumor model in C57BL/6 J mice (n = 6 mice per group in one experiment) treated with or without anti-Ly6G/anti-CD8 antibody. O, P Image and weights of the orthotopic tumors in the experimental groups from (N) at the end of the experiments (n = 6 mice per group). Q–S Tumor-infiltrating neutrophils, CD8+ T cells, and GZMB+CD8+ T cells isolated from the orthotopic tumors were analyzed using flow cytometry: Representative graphs of flow cytometry (left panel) and statistical analysis of the cell ratio (right panel) (n = 6 mice per group). T, U Representative luminescence images of the mouse model in (N) and statistical analysis of the total flux (n = 6 mice per group). Data represent mean ± SEM. Statistical analysis was conducted using the two-tailed unpaired Student’s t test (A, I–M, P–S, and U), two-tailed Pearson’s correlation analysis (B), and log-rank test (C). Source data are provided as a Source Data file.
To further analyze whether glycolysis affects neutrophil infiltration in pancreatic cancer, we used a hexokinase (HK) inhibitor (2-deoxy-D-glucose, 2DG) and the shRNA targeting LDH (silenced both Ldha and Ldhb, LDH-knockdown) to inhibit glycolysis in tumor cells (Fig. 1D). We found that tumor growth in the 2DG treatment group was significantly inhibited compared to the control group in the KPC subcutaneous implantation tumor model (Supplementary Fig. S2A–G). Treatment with 2DG significantly decreased intratumoral lactate levels (Supplementary Fig. S2H). Then, we conducted mass cytometry (CyTOF) and identified 12 distinct cell clusters based on the differential expression profile of 41 markers (Supplementary Fig. S2I). The t-distributed stochastic neighbor embedding (t-SNE) analysis showed that tumor-infiltrating neutrophils decreased, whereas CD8+ T cells increased significantly in the 2DG-treatment group (Fig. 1E, F, and Supplementary Fig. S2J–L). Additional analysis by flow cytometry confirmed the reduction in tumor-infiltrating neutrophils after 2DG treatment (Supplementary Fig. S3A, C). In contrast, the proportions of CD8+ T cells were significantly enriched in tumor tissues treated with 2DG (Supplementary Fig. S3B, D). To rule out the influence of tumor size on the number of infiltrating immune cells, we examined the infiltration levels of neutrophils and CD8+ T cells at the early stage of 2DG treatment (Supplementary Fig. S2A). After 2 days of 2DG treatment, the tumor size showed little difference between the 2DG and control treatment groups (Supplementary Fig. S3E–G). However, the lactate levels had already decreased after 2 days of 2DG treatment (Supplementary Fig. S3H). Flow cytometry analysis showed that the proportion of neutrophils in the tumor microenvironment was significantly decreased following 2 days of 2DG treatment, whereas CD8+ T cells were markedly enriched in tumors (Supplementary Fig. S3I, J). Furthermore, in the orthotopic transplanted tumor model, we also found that 2DG treatment inhibited tumor growth, with reduced infiltration of neutrophils and increased infiltration of CD8+ T cells (Fig. 1G–K). Flow cytometry results revealed that the percentage of tumor-infiltrating GZMB+CD8+ T cells was significantly higher, but exhausted PD-1+CD8+ T cells were unchanged (Fig. 1L, M).
To exclude the effect of 2DG on non-tumor cells in vivo, we double-silenced Ldha and Ldhb (LDH-knockdown) in KPC cells using the shRNAs, and then subcutaneously injected control or LDH-knockdown KPC cells into the C57BL/6 J mice (Supplementary Fig. S4A). Compared with the control group, mice that received LDH-knockdown KPC cells showed a significantly smaller tumor burden (Supplementary Fig. S4B–G). We also observed similar changes (i.e, reduced neutrophils and enriched CD8+ T cells) in tumor tissues from the subcutaneous tumor model of LDH knockdown KPC cells (Supplementary Fig. S4H, I). Therefore, our findings suggest that high glycolysis may enrich neutrophils and inhibit the infiltration of CD8+ T cells, subsequently altering the tumor microenvironment and promoting tumor progression in PDAC.
Glycolysis induces immunosuppression mediated by neutrophils
To further investigate whether glycolysis-induced immunosuppression depends on neutrophils in pancreatic cancer, we depleted CD11b+Ly6G+ neutrophils in an orthotopic transplanted tumor model using anti-Ly6G antibody (Fig. 1N). Similar to the results of subcutaneously transplanted tumors, the LDH-knockdown group showed a significant reduction in tumor burden (Fig. 1O, P). However, there was less change in tumor burden between the control and LDH-knockdown groups treated with anti-Ly6G antibody compared with their isotype control (Fig. 1O, P). Flow cytometry results showed that the proportion of neutrophils in the tumor microenvironment of the LDH knockdown group was significantly decreased, whereas CD8+ T cells and GZMB+CD8+ T cells were markedly enriched in tumors (Fig. 1Q–S). Furthermore, neutrophil depletion also restored the abundance and functional status of CD8+ T cells (Fig.1Q–S). Previous studies have shown that neutrophils contribute to pancreatic cancer progression by inhibiting the recruitment and function of CD8+ T cells29,30. To examine whether neutrophil depletion can reverse glycolysis-mediated tumor-promoting effects through reactivating CD8+ T cells in vivo, we further depleted CD8+ T cells in mice (Fig. 1N). The results showed that depleting CD8+ T cells rescued the tumor-suppressing effect of neutrophil depletion (Fig. 1T, U, and Supplementary Fig. S4J, K). Taken together, these findings suggest that glycolysis may induce immunosuppression mediated by neutrophils and play an oncogenic role in the tumor progression of pancreatic cancer.
Glycolysis enhances neutrophil infiltration by up-regulation of Cxcl1 expression
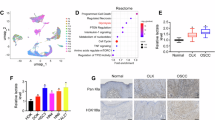
We next investigated the mechanism underlying the recruitment of neutrophils induced by glycolysis. Since chemokines are well known to regulate the migration of immune cells31, we performed RNA-seq in PANC1 and BxPC3 cells with or without treatment of 2DG and found that Cxcl1 was the most significant differentially expressed chemokine (Fig. 2A). Additional analysis of TCGA data also revealed that Cxcl1 was highly expressed in pancreatic cancer patients and was associated with poor prognosis (Supplementary Fig. S5A, B). We also examined the expression level of Cxcl1 in vitro in human PANC1 and BxPC3 cell lines, as well as in mouse KPC and PANC02 cell lines, following glycolysis inhibition with either 2DG or shRNA silencing of LDH (Supplementary Fig. S5C). The results indicated that inhibition of glycolysis significantly reduced the intracellular Cxcl1 mRNA levels (Fig. 2B, C) and extracellular CXCL1 protein levels (Fig. 2D, E). Immunohistochemical staining showed that CXCL1 expression was decreased in tumor tissues treated with 2DG or LDH-knockdown compared with the controls (Fig. 2F). We also found that secreted CXCL1 in the serum of tumor-bearing mice treated with 2DG or LDH-knockdown was significantly reduced compared with the control mice (Fig. 2G and Supplementary Fig. S5D). Moreover, the serum CXCL1 level in pancreatic cancer patients was significantly higher than in healthy controls (Fig. 2H). Taken together, these findings suggest that glycolysis may regulate Cxcl1 expression in pancreatic cancer cells.
A The flowchart illustrates that RNA sequencing was conducted using two PDAC cell lines with or without 2DG treatment. Subsequently, the intersection of detected genes within the chemokine family in both cell lines is identified, and a heatmap is generated to display the log2 fold change (2DG versus vehicle) of these genes. B, C Relative mRNA levels of Cxcl1 in PDAC cell lines following glycolysis inhibition, either by using 2DG or silencing LDH, were analyzed by qRT-PCR (n = 3 independent experiments). D, E Relative protein levels of CXCL1 in PDAC cell lines following glycolysis inhibition, either by using 2DG or silencing LDH, were determined using ELISA assay (n = 3 independent experiments). F Representative IHC images of subcutaneous tumors (n = 6 mice for 2DG treatment, n = 5 mice for shLDH/shNTC groups) stained by CXCL1 antibody (scale bars = 100 μm). G Relative serum CXCL1 levels in mice treated with or without 2DG were measured by ELISA (n = 6 mice per group). H Relative serum CXCL1 levels in healthy donors and PDAC patients were measured by ELISA (22 healthy samples and 27 PDAC samples). I Schematic diagram showing the in vitro migration assay: human/mouse neutrophils were co-incubated with the culture medium supernatant from PDAC cells treated with 2DG or LDH-knockdown. J Neutrophils were co-cultured with CD8+ T cells in different proportions, and the proliferation of CD8+ T cells was detected with the CFSE assay (n = 6 biologically independent samples). K The migratory activity of neutrophils co-incubated with the culture medium supernatant from PDAC cells treated with shLDH and recombinant CXCL1 was analyzed by counting the penetrated cell numbers (n = 3 biologically independent samples). L The neutrophils were treated with SX-682 or Navarixin for 1.5 h in advance, and the migratory activity of neutrophils co-incubated with the culture medium supernatant from PDAC cell lines treated with shLDH was analyzed by counting the penetrated cell numbers (n = 3 biologically independent samples). M Representative luminescence images of the orthotopic tumor in mice and statistical analysis of MFI (n = 5 mice per group). N Tumor-infiltrating neutrophils isolated from the orthotopic tumors were analyzed using flow cytometry (n = 5 mice per group). Data represent mean ± SEM. Statistical analysis was conducted using the two-tailed unpaired Student’s t test. Source data are provided as a Source Data file.
Furthermore, we performed the cell migration assays to evaluate the ability of glycolytically regulated CXCL1 to recruit neutrophils (Fig. 2I). We isolated neutrophils from human peripheral blood and mouse bone marrow (Supplementary Fig. S5E). Firstly, we confirmed the immunosuppressive activity of neutrophils by measuring the T cell proliferation (Fig. 2J). The recombinant human and mouse CXCL1 proteins were confirmed by western blot, ELISA, and the migration ability of neutrophils (Supplementary Fig. S5F–I), respectively. Subsequent chemotaxis assays revealed that conditional culture medium from cancer cells treated with 2DG or LDH-knockdown recruited significantly fewer neutrophils compared to the control groups which could be effectively restored by the addition of recombinant human or mouse CXCL1 protein (Fig. 2K and Supplementary Fig. S6A). Moreover, the conditional culture medium from the treated cancer cells did not affect neutrophil survival, ROS production, and degranulation (Supplementary Fig. S6B–D). CXCL1 mainly mediates neutrophil migration through the receptor CXCR1/232,33. We found that following treatment with the CXCR1/2 inhibitor SX-682 or Navarixin, the difference in neutrophil migration between the 2DG treatment or LDH-knockdown group and the control group was significantly reduced or eliminated (Fig. 2L and Supplementary Fig. S6E). Furthermore, the tumor growth inhibition observed following LDH-knockdown could be partially reversed by restoring Cxcl1 expression in vivo (Fig. 2M and Supplementary Fig. S6F). Finally, flow cytometry results indicated that the reduction in neutrophil infiltration caused by LDH-knockdown was restored after overexpression of Cxcl1 (Fig. 2N). Collectively, our data suggest that up-regulation of Cxcl1 expression and secretion by glycolysis may enrich neutrophils in tumor tissues and induce an immunosuppressive tumor microenvironment.
Glycolysis-derived lactate enhances H3K18la modification and promotes the expression of Cxcl1
We next investigated the mechanism underlying the transcriptional activation of Cxcl1 by glycolysis. Lactate has been suggested to promote the expression of Cxcl134. In addition, recent studies have identified lactate-mediated histone lactylation as a post-transcriptional modification to direct activation of gene transcription18. There are two stereoisomeric forms of lysine lactylation (Kla): L-lactyl-lysine and D-lactyl-lysine35. Recent studies have clearly demonstrated that L-lactyl-lysine is the primary responder to glycolysis and the Warburg effect35. Therefore, we speculated that histone L-lactylation may directly activate Cxcl1 expression at the transcriptional level. Indeed, western blot showed increased L-lactyl-lysine (PanKla) in pancreatic cancer tissues compared with normal tissues, and H3K18la showed the most significant changes (Fig. 3A and Supplementary Fig. S7A). Immunohistochemical (IHC) results also showed that H3K18la levels were up-regulated in pancreatic cancer tissues and cell lines (Fig. 3B and Supplementary Fig. S7B). We then examined the effects of aerobic glycolysis on histone lactylation using glycolysis inhibitor (2DG), glycolysis agonist (rotenone), and LDH-knockdown in human and mouse pancreatic cancer cell lines, respectively (Fig. 3C). The levels of PanKla and H3K18la consistently reduced in the cancer cells treated with 2DG or LDH-knockdown (Fig. 3D and Supplementary Fig. S7C). In contrast, the levels of PanKla and H3K18la consistently increased in cancer cells treated with rotenone (Fig. 3E). Consistent with the results in vitro, PanKla and H3K18la levels were also significantly reduced in mouse tumor tissues treated with 2DG or LDH-knockdown (Fig. 3F, G and Supplementary Fig. S7D, E). Furthermore, the reduction in H3K18la levels induced by 2DG or LDH-knockdown was recovered after treatment with sodium lactate (Fig. 3H, I). These findings suggest that aerobic glycolysis-derived lactate can enhance H3K18la modification.
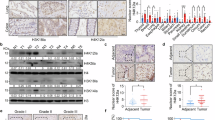
A Lactylation levels of pan-lysine and different lysine sites in histone H3 of 7 pairs of PDAC tissue and adjacent normal tissue were detected by western blot. B Representative IHC images of clinical PDAC and adjacent normal tissue stained by H3K18la antibody (n = 79 patients). C The diagram illustrating the targets of 2DG, shRNA, and rotenone in the glycolysis process. D, E Relative levels of H3K18la and PanKla in PDAC cells treated with gradient doses of 2DG (D) or rotenone (E) for 24 h were detected by western blot. F Relative levels of H3K18la and PanKla in subcutaneous tumors treated with or without 2DG were detected by western blot. (n = 4 biologically independent samples per group). G H3K18la levels of subcutaneous tumors treated with or without 2DG were detected by immunofluorescence staining. H The H3K18la levels in PDAC cells treated with 2DG or sodium lactate were detected by western blot. I The H3K18la level in PANC1 and PANC02 cells following LDH-knockdown and sodium lactate treatment by western blot. J Schematic diagram of the CUT&Tag assay for PanKla or H3K18la in BxPC3 cells with or without 2DG treatment. K TSS heatmaps presenting the binding density of PanKla and H3K18la with or without 2DG treatment. L Venn diagram indicating the overlap of differentially expressed genes in TCGA-PAAD between PDAC and adjacent normal tissue, differentially expressed genes in RNA-seq after 2DG treatment, and differentially bound genes of H3K18la and PanKla after 2DG treatment. M Genome browser track analysis shows the PanKla and H3K18la levels in the Cxcl1 promoter region with or without 2DG treatments. N The Cxcl1 promoters that are bound by H3K18la and PanKla in PDAC cells were quantified using ChIP-qPCR assays (n = 3 independent experiments). O Relative mRNA levels of Cxcl1 in PDAC cells treated with 2DG or sodium lactate were detected by qRT-PCR (n = 3 independent experiments). P Representative IHC images stained by CXCL1 or H3K18la antibody and correlation analysis of H-scores of CXCL1 and H3K18la in PDAC samples (n = 21 patients). Scale bar = 100 μm. Data represent mean ± SEM. Statistical analysis was conducted using the two-tailed unpaired Student’s t test (N–O) and two-tailed Pearson’s correlation analysis (P). Unless otherwise indicated, all western blots had three independent experimental repetitions with consistent results, where Histone H3, H3K9la, H3K14la, H3K18la, and PanKla blots are from parallel-processed separate gels (size conflict) with samples from the same experiment. Source data are provided as a Source Data file.
To understand the potential functional significance of H3K18la modification, we performed CUT&Tag assays using antibodies against PanKla and H3K18la to identify candidate genes regulated by H3K18la in pancreatic cancer (Fig. 3J). The 2DG-treated group showed a noticeable reduction of PanKla peaks and H3K18la peaks in the transcription start site (TSS) region compared with the control (Fig. 3K). KEGG analysis of the down-regulated H3K18la binding gene at the promoter regions in the 2DG-treated group revealed enrichment in the pathways associated with cancer (Supplementary Fig. S7F). Moreover, we identified 9 genes that exhibited significantly reduced mRNA levels and reduced enrichment of the H3K18la signal at the promoter region upon 2DG treatment but up-regulated mRNA levels in PDAC tissues by analyzing the overlapping genes among CUT&Tag, RNA-seq, and TCGA database (Fig. 3L). In particular, Cxcl1 was one of the most significantly changed genes. Furthermore, we investigated whether H3K18la could directly regulate Cxcl1 expression at the transcriptional level. Specifically, the called peaks showed that the levels of PanKla and H3K18la at the promoter regions of Cxcl1 were noticeably reduced in cancer cells treated with 2DG (Fig. 3M). A quantitative chromatin immunoprecipitation analysis indicated that the PanKla and H3K18la levels on Cxcl1 promoters were significantly enriched (Fig. 3N). In addition, the decreased mRNA levels of Cxcl1 by 2DG treatment could be recovered after sodium lactate treatment (Fig. 3O). We then performed IHC analyses of 21 human PDAC tissues and revealed that the levels of CXCL1 and H3K18la were positively correlated (Fig. 3P). These results indicate that H3K18la modification may directly activate the transcription of Cxcl1 in pancreatic cancer cells.
PCAF functions as a lactyltransferase for H3K18la modification
A number of histone acetyltransferases (HATs) have been reported to function as lactyltransferase23. However, the writer who promotes histone lactylation in pancreatic cancer cells remains unclear. Here, we screened the existing histone acetyltransferase inhibitors to identify lactyltransferase that may regulate H3K18la in pancreatic cancer cells (Supplementary Fig. S8A). The western blot results showed that the levels of H3K18la decreased significantly in tumor cells treated with bromosporine (Supplementary Fig. S8B), an inhibitor for PCAF, suggesting that PCAF may function as a potential lactyltransferase in pancreatic cancer. To further explore whether PCAF, a member of the GNAT (Gcn5-related N-acetyltransferase) sub-family of histone acetyltransferases, is a potential lactyltransferase, we performed in silico molecular docking analysis. We docked lactyl coenzyme A (lactyl-CoA) into the structure of PCAF (Protein DataBank (PDB):1CM0). Lactyl-CoA was well accommodated in the cofactor pocket of PCAF, similar to the structure of the acetyl coenzyme A (acetyl-CoA)–PCAF complex (Fig. 4A). The co-immunoprecipitation (Co-IP) assays showed that H3K18la interacted with PCAF (Fig. 4B). We then treated multiple pancreatic cancer cell lines with two inhibitors of PCAF, bromosporine and Embelin, and both of them exhibited a dose-dependent decrease in H3K18la (Fig. 4C, D). In addition, when Pcaf was knocked down in PANC1, BxPC3, and KPC cells, the levels of H3K18la were decreased (Fig. 4E and Supplementary Fig. S8C). Moreover, PCAF-mediated histone H3 K18 lactylation was confirmed with an in vitro lactylation assay (Fig. 4F). Previous studies have reported that the acetyltransferases P300 and GCN5 can catalyze the lactylation of histone H3 K1823. Our findings demonstrated that PCAF exhibited similar enzymatic activity with P300 and GCN5, and was able to catalyze H3 K18 lactylation in pancreatic cancer cells (Supplementary Fig. S8D, E). The mRNA and secreted protein levels of Cxcl1 in the PCAF-inhibited group were also significantly reduced compared with the control (Fig. 4G, H), consistent with previous results. We also observed similar results in cancer cells after Pcaf knockdown (Fig. 4I and Supplementary Fig. S8F). In addition, we examined the effects of bromosporine on lysine lactylation and lysine acetylation. The western blot results indicated that the levels of H3K18la and H3K18ac decreased in a dose-dependent manner with the treatment of bromosporine, but the reduction level of H3K18la was the most significant (Supplementary Fig. S8G). Interestingly, H4K8ac remains unchanged at the current concentration gradient (Supplementary Fig. S8G). The role of H3K18la, H3K18ac, and H4K8ac in the direct transcriptional regulation of Cxcl1 was further analyzed following bromosporine treatment. The results showed that the enrichment effect of H3K18la on Cxcl1 was significantly reduced after bromosporine treatment (Supplementary Fig. S8H). However, H3K18ac and H4K8ac did not show any differences (Supplementary Fig. S8H).
A Molecular docking simulated the binding affinities of PCAF for acetyl-CoA and lactyl-CoA. B Co-IP was performed to confirm the interaction between H3K18la and PCAF in PANC1 and KPC cells. PCAF and H3K18la were detected on the same gel. C, D Western blot analysis demonstrates the level of H3K18la in PDAC cells with or without the treatment of PCAF inhibitors (bromosporine or Embelin). Histone H3 and H3K18la blots are from parallel-processed separate gels (size conflict) with samples from the same experiment. E Western blot analysis demonstrates the level of H3K18la in PDAC cells following the knockdown of Pcaf. Histone H3, PCAF, β-actin and H3K18la blots are from parallel-processed separate gels with samples from the same experiment. F Western blots of in vitro histone acetylation or lactylation assay, the samples as indicated. Blots are from parallel-processed separate gels with samples from the same experiment. G, H Relative Cxcl1 RNA and protein levels with the vehicle or bromosporine treatment were analyzed by qRT-PCR and ELISA (n = 3 independent experiments), respectively. I Relative Cxcl1 RNA levels after Pcaf knockdown were analyzed by qRT-PCR (n = 3 independent experiments). J A schematic diagram showing the orthotopic tumor construction in C57BL/6 J mice (n = 5 mice per group in one experiment, 2 × 106 KPC-luc cells per mouse) with or without bromosporine treatment. K, L Image and weights of the orthotopic tumors in the experimental groups from (J) at the end of the experiments (n = 5 mice per group). M Lactylation levels of H3K18 and PanK in orthotopic tumors with or without bromosporine treatment were detected by western blot (n = 4 biologically independent samples per group). Histone H3, H3K18la, and PanKla blots are from parallel-processed separate gels (size conflict) with samples from the same experiment. N Serum CXCL1 levels in mice with or without bromosporine treatment were measured by ELISA (n = 5 mice per group). O–R Tumor-infiltrating neutrophils, CD8+ T cells, GZMB+CD8+ T cells, and PD-1+CD8+ T cells isolated from the orthotopic tumors were analyzed using flow cytometry. Representative graphs of flow cytometry (left panel) and statistical analysis of the cell ratio (right panel) (n = 5 mice per group). S The migratory abilities of neutrophils co-incubated with the culture medium supernatant from shNTC/Pcaf PDAC cells treated with recombinant CXCL1 were analyzed by counting the penetrated cell numbers (n = 3 biologically independent samples). T Representative IHC images stained by CXCL1 or H3K18la antibody (scale bar = 100 μm, left panel), correlation analysis of H-scores of CXCL1 and H3K18la in PDAC samples (n = 21 patients, right panel). Data represent mean ± SEM. Statistical analysis was conducted using the two-tailed unpaired Student’s t test (G–I, L, and N–S) and two-tailed Pearson’s correlation analysis (T). Unless otherwise indicated, all western blots had three independent experimental repetitions with consistent results. Source data are provided as a Source Data file.
We further explored the function of PCAF in vivo by an orthotopic transplantation tumor model of pancreatic cancer (Fig. 4J). The tumor growth in the bromosporine-treated group was significantly inhibited compared to that in the vehicle-treated group (Fig. 4K, L). Consistent with the results in vitro, H3K18la levels were also significantly reduced in bromosporine-treated mouse tumor tissues, while H3K18ac and H4K8ac showed no significant changes (Fig. 4M, and Supplementary Fig. S8I). We also found that the secreted CXCL1 in the serum of tumor-bearing mice treated with bromosporine decreased significantly compared with the vehicle (Fig. 4N). Flow cytometry analysis showed that the number of tumor-infiltrating neutrophils decreased significantly, whereas the number of CD8+ T cells increased in the bromosporine-treated group (Fig. 4O, P). Furthermore, flow cytometry results revealed that the percentage of tumor-infiltrating GZMB+CD8+ T cells was significantly higher, but exhausted PD-1+CD8+ T cells were unchanged (Fig. 4Q, R). Similar results were observed in the KPC subcutaneous implantation tumor model, and the mouse body weight was not affected (Supplementary Fig. S8J–U).
Cell migration assays indicated that the conditioned culture medium from cancer cells with Pcaf-knockdown recruited fewer neutrophils compared with the control, which could be rescued by recombinant human CXCL1 protein (Fig. 4S). Moreover, we performed IHC analyses of 21 human PDAC tissues and revealed that the levels of PCAF and H3K18la were positively correlated (Fig. 4T). Taken together, these data suggest that PCAF functions as a lactyltransferase for H3K18la in pancreatic cancer.
PCAF inhibitor combined with anti-PD-1 antibody may become a novel therapeutic strategy for pancreatic cancer
Clinical studies have shown that ICB therapy does not benefit patients with pancreatic cancer2,4, emphasizing the necessity of formulating rational combinational therapies. To visualize the immune contexture shaped by H3K18la, we performed multiplex immunofluorescence on human PDAC tissues and observed that regions with higher H3K18la and CXCL1 signals often coincided with increased neutrophils but fewer CD8+ T cells in the same patient tissue (Supplementary Fig. S9). Conversely, H3K18la-low regions showed the opposite immune cell landscape. We investigated whether the inhibition of PCAF could enhance the efficacy of anti-PD-1 antibodies in two pancreatic cancer mouse models. In the PANC02 subcutaneous implantation tumor model, we combined PCAF inhibitor bromosporine with anti-PD-1 antibody and revealed an additive effect on tumor growth inhibition (Fig. 5A–D), without affecting mouse body weight (Supplementary Fig. S10A–D). Consistently, IHC staining showed that combinational therapy with bromosporine and anti-PD-1 antibody synergistically suppressed Ki67 positive tumor cells (Supplementary Fig. S10E), decreased neutrophil infiltration, and enhanced CD8+ T cell infiltration in tumor tissues (Fig. 5E and Supplementary Fig. S10E). Moreover, bromosporine alone or in combination with anti-PD-1 antibody reduces the levels of H3K18la in tumor tissues (Fig. 5F) and the levels of secreted CXCL1 in serum (Fig. 5G).
A A schematic diagram showing the subcutaneous tumor model (n = 6 mice per group in one experiment, 6 × 106 PANC02 cells per mouse) treated with bromosporine and anti-PD-1 antibody. B. The tumor volume growth curves of subcutaneous tumors from (A). C, D Image and weights of the subcutaneous tumors at the end point of experiments (n = 6 mice per group). E The statistical analysis of the cell ratio of tumor-infiltrating neutrophils and CD8+ T cells isolated from subcutaneous tumors (n = 3 mice per group). F Western blot analysis demonstrates H3K18la levels in the subcutaneous tumors (n = 3 biologically independent samples per group) from (A). Histone H3 and H3K18la blots are from parallel-processed separate gels (size conflict) with samples from the same experiment. G Relative serum CXCL1 levels in mice treated with or without bromosporine/anti-PD-1 antibody were measured by ELISA (n = 6 mice per group). H A schematic diagram showing the combinational treatment schedule for the orthotopic KPC-luc tumor model (n = 21 mice per group in one experiment, 2 × 106 KPC-luc cells per mouse). I, J Image and weights of the orthotopic tumors at the end point of experiments (n = 6 mice per group). K–N Tumor-infiltrating neutrophils, CD8+ T cells, GZMB+CD8+ T cells, and PD-1+CD8+ T cells were analyzed using flow cytometry. Representative graphs of flow cytometry (left panel) and statistical analysis of the cell ratio (right panel) (n = 6 mice per group). O Western blot analysis showing the H3K18la levels in the orthotopic tumors (n = 3 biologically independent samples per group) from (H). Histone H3 and H3K18la blots are from parallel-processed separate gels (size conflict) with samples from the same experiment. P Representative luminescence images and Quantification of radiance intensity of the mouse model in (H). Q Survival probability of mice with orthotopically transplanted PDAC (n = 15 mice per group). R A working model displaying the signaling pathway through which the aerobic glycolysis-mediated Lactate-PCAF-H3K18la-CXCL1 axis modulates the tumor microenvironment in pancreatic cancer, and the scientific basis for the development of a novel therapeutic strategy for PDAC. Data represent mean ± SEM. Statistical analysis was conducted using the two-tailed unpaired Student’s t test (B, D, E, G, J–N, P) and the log-rank test (Q). Source data are provided as a Source Data file.
Then, we further assessed the therapeutic effect of bromosporine combined with anti-PD-1 antibody in a pancreatic cancer orthotopic transplantation tumor model (Fig. 5H). Simultaneous blockade of PCAF and PD-1 could markedly attenuate the tumor burden compared to the monotherapy group (Fig. 5I, J). Flow cytometric analysis showed that the combination of bromosporine and PD-1 blockade synergistically decreased the proportion of infiltrated neutrophils and increased CD8+ T cell infiltration (Fig. 5K, L). Meanwhile, flow cytometric results revealed that the percentage of tumor-infiltrating GZMB+CD8+ T cells was significantly higher and exhausted PD-1+CD8+ T cells were significantly lower in the combinational therapy compared to control (Fig. 5M, N). Moreover, western blot analysis also showed that the levels of H3K18la were decreased in bromosporine alone or in combination with anti-PD-1 antibody in tumor tissues (Fig. 5O). Furthermore, compared to other groups, the survival time was also significantly prolonged after combinational treatment with bromosporine and anti-PD-1 antibody (Fig. 5P, Q). Taken together, our findings demonstrate that PCAF inhibitors could significantly enhance the efficacy of anti-PD-1 antibodies by reducing neutrophil infiltration in pancreatic cancer.
Discussion
In this study, we found that aerobic glycolysis-generated lactate could increase the levels of H3K18la, which subsequently upregulate Cxcl1 expression, enrich neutrophil infiltration, and induce immunosuppression in pancreatic cancer. Moreover, our data suggest that PCAF functions as a histone lactyltransferase and directly activates the Cxcl1 expression at the transcriptional level. Furthermore, we demonstrated that a combinational treatment with a PCAF inhibitor and an anti-PD-1 antibody induced a synergistic antitumor efficacy in an orthotopic mouse model of pancreatic cancer (Fig. 5R).
Pancreatic cancer cells have been shown to exhibit a high level of glycolysis36, suggesting that lactylation may occur frequently in PDAC. Two other groups have reported high levels of histone lactylation in pancreatic cancer. Li et al.37 reported that H3K18la modification transcriptionally activated TTK/BUB1B to promote the oncogenic processes of PDAC. Wang et al.38 reported H4K12la-mediated immune escape in pancreatic cancer. In addition, H3K18la has also been reported to mediate immune escape in non-small cell lung cancer20. Our study is the first to report that H3K18la can mediate immune escape in pancreatic cancer. These results suggest that H3K18la can play important roles in regulating the tumor microenvironment and mediating tumor immune escape.
Here, 2DG, a classic inhibitor of glycolysis, has been used to dissect how histone lactylation modulates immune cell infiltration in tumors. Based on the Warburg effect in tumors, radiolabeled 2DG (2-deoxy-2-[18F]fluoro-D-glucose) has been commonly used as a tracer in Positron Emission Tomography (PET) scans due to its selective uptake and accumulation in malignant tissues39. CD8+ T cells undergo metabolic rewiring to meet their proliferation and effector functions40. At the same time, 2DG might have an impact on T cells as well. Interestingly, although glycolysis supports effector T cell expansion41, its potentiation paradoxically compromises the development of early memory CD8+ T cells42,43. Consistent with this, ex vivo priming in the presence of 2DG enhances the generation of long-lived memory CD8+ T cells with superior antitumor activity upon adoptive transfer42,43. In contrast, Greco et al.44 demonstrated that while CAR-T cells cultured alone readily take up the fluorescent glucose analog 2-NBDG and exhibit dose-dependent suppression of lactate production in response to 2DG, these effects are abolished in the presence of tumor cells, suggesting that tumor engagement may shield T cells from glycolytic inhibition. In addition, elevated lactate within the acidic tumor microenvironment is known to suppress CD8+ T cell cytotoxicity40. Our data show that 2DG treatment significantly reduces intratumoral lactate levels, potentially alleviating this immunosuppressive mechanism. Notably, recent studies have identified lactylation as a novel modification in immune cells including CD8+ T cells45 and neutrophils46. We observed that lactylation levels were markedly higher in KPC tumor cells compared to CD8+ T cells and neutrophils (Supplementary Fig. S7G). Furthermore, in vitro 2DG treatment selectively reduced H3K18la in KPC cells, but not in CD8+ T cells or neutrophils (Supplementary Fig. S7H), suggesting tumor-specific susceptibility to glycolysis inhibition. Moreover, we ruled out 2DG’s effect on immune cells in our study by genetically knocking down LDH in tumor cells. Collectively, these findings suggest that 2DG may modulate the immune microenvironment and enhance antitumor immunity, although the underlying mechanisms require further elucidation.
PCAF, a member of the GNAT superfamily of HAT enzymes, has been reported to mediate histone acetylation47. For example, PCAF may acetylate H4K8 to recruit MRE11 and EXO1 at the stalled replication forks48. Here, we demonstrated that PCAF may function as a lactyltransferase to catalyze histone H3K18la by conducting molecular docking, Co-IP, western blotting, and ELISA assays. Most importantly, for the first time, we directly demonstrated through in vitro histone lactylation assay that PCAF could function as a histone lactyltransferase. Recent studies have reported that P300/CBP18,23,37 and GCN523,24 could mediate H3K18la. Our results showed that when Pcaf, P300 and Gcn5 were knocked down separately in human pancreatic cancer PANC1 cells or mouse pancreatic cancer KPC cells, the levels of H3K18la were reduced to a similar level. However, additional studies are required to determine whether PCAF, P300 and GCN5 could synergistically regulate H3K18la in pancreatic cancer, or each of them may play a different role in this process. Furthermore, through subcutaneous and orthotopic tumor models of pancreatic cancer, we have demonstrated that inhibition of PCAF activity by bromosporine could suppress H3K18la, which subsequently reduced the Cxcl1 expression and promoted antitumor immunity in vivo.
As bromosporine may influence both lysine acetylation and lysine lactylation, we examined the levels of H3K18ac and H4K8ac after treatment with bromosporine in vitro and in vivo. Interestingly, at the concentration used in this study, we found that bromosporine had the most significant effect on H3K18la (Supplementary Fig. S8G). Zhu et al.24 have reported that GCN5 from the same family protein of PCAF executes distinct histone acylation in different protein complexes. We speculate that PCAF may also have different responses to bromosporine due to exerting different acylase activities in different protein complexes. Finally, we revealed that the expression level of PCAF was positively correlated with the level of H3K18la in the tissue samples from the pancreatic cancer patients, suggesting that PCAF may play an important role in the disease progression of pancreatic cancer through the regulation of H3K18la.
Numerous studies have shown that CXCL1 promotes the progression of pancreatic cancer by recruiting neutrophils29,30,49,50,51,52. However, little is known about whether tumor-intrinsic glycolysis is able to mobilize neutrophils. The expression level of Cxcl1 is regulated by several mechanisms, such as NF-κB signaling50 and H3K27me330. In this study, we delineated a new mechanism by which the glycolysis-derived lactate promoted the level of H3K18la to transcriptionally activate Cxcl1 expression. Interestingly, Dong et al. found that lactate can promote Cxcl1 expression in endothelial cells during the development of the mouse neocortex34, suggesting that histone lactylation may regulate Cxcl1 expression in a variety of physiological and pathological conditions. In addition to Cxcl1, the significance of other downstream genes of H3K18la should be investigated in the future.
Taken together, our findings revealed, for the first time, that the aerobic glycolysis-lactate-PCAF-H3K18la-CXCL1 signaling axis could modulate the disease progression of pancreatic cancer by recruiting neutrophils and inducing immunosuppression in TME. Additionally, we developed a novel therapeutic strategy for pancreatic cancer by using PCAF inhibitors combined with anti-PD-1 antibodies.
Methods
Mice
In this study, male C57BL/6 J mice (6–8 weeks old) were obtained from GemPharmatech Co., Ltd. The mice were housed in the Specific Pathogen Free (SPF) animal facility at the First Affiliated Hospital of Nanchang University. Prior to the initiation of the experiments, a one-week acclimation period was allowed for the mice to adjust to the laboratory environment. All mice were housed under controlled conditions with a 12 h light/dark cycle, a temperature range of 22-24 °C, and 40-70% humidity. Throughout the study, they had continuous free access to food (Beijing Keao Xieli Feed Co., Ltd.) and water. All the animal experiments were conducted in strict accordance with the ethical standards for the welfare of experimental animals and were approved by the Ethics Committee for Animal Welfare at the First Affiliated Hospital of Nanchang University (CDYFY-IACUC-202407QR128). The maximal tumor burden permitted was 1500 mm³ for subcutaneous tumors or a palpable diameter exceeding 1.5 cm or clear signs of abdominal distension for orthotopic tumors. According to our approved protocol, mice were monitored daily and euthanized immediately upon observation of predefined humane endpoints, including neurological deficits, persistent lethargy, impaired mobility, or seizures.
Subcutaneous pancreatic cancer mouse model
PDAC cells were mixed with Matrigel (Corning, #354234) at a ratio of 3:1, and 100 μL of the suspension was injected subcutaneously into the right flank of 6–8 weeks-old male C57BL/6 J mice (PANC02 with 6 × 106 cells per mouse, KPC-luc with 3 × 106 cells per mouse). Treatment was initiated once the subcutaneous tumor volume reached 100–200 mm3. Tumor volume was measured daily using calipers and calculated using the formula: Volume = 0.52 × length × width2. Upon completion of the treatment, mice were euthanized, and the corresponding tissues were prepared for further recording or examination. The treatment schedule is depicted in the figures in the manuscript and described as following: (1) The treatments of vehicle/2DG (800 mg/kg, i.p., once daily) was initiated on day 7 post-implantation and continued for 2 or 8 days; (2) Administration of vehicle/bromosporine (30 mg/kg, i.p., once a day) was initiated on day 9 post-implantation and continued for a total of 11 doses; (3) For combinational treatment, the administration of bromosporine (30 mg/kg, i.p., 3 times a week) or anti-PD-1 antibody (200 μg/mouse, i.p., twice a week, BioXCell, #BE0146)/IgG2a antibody (200 μg/mouse, i.p., twice a week, BioXCell, #BE0089) began on day 13 post-implantation. Mice received two 5-day cycles of therapy, separated by a 2-day rest period; (4) For the shNTC/shLDH KPC-luc subcutaneous implantation model, shNTC/shLDH KPC-luc cells were implanted subcutaneously into C57BL/6 J mice. The mice were euthanized on the 16th day after the implantation.
Orthotopic pancreatic cancer mouse model
Male C57BL/6 J mice (6–8 weeks old) were implanted orthotopically with a 50 μL cell suspension containing 2 × 106 KPC-luc cells into the pancreas. Tumors were monitored by bioluminescence imaging (D-luciferin potassium salt, Biodragon, #BDXB0116, i.p., 150 mg/kg). The IVIS Lumina Series III in vivo imaging system was employed to capture the bioluminescent signals emitted by the tumors. Then, the mice were subjected to different treatments as shown in the figures and described as following: (1) Vehicle/2DG treatment was administered (800 mg/kg, i.p., once daily) from day 7 for 9 consecutive days; (2) For neutrophil depletion, these mice received anti-IgG2a (0.25 mg/mouse, i.p.) or anti-Ly6G antibody (0.25 mg/mouse, i.p., BioXCell, #BE0075-1) treatment once a day from day 3 post-implantation. After 11 consecutive days of antibody administration, the mice were euthanized, the peripheral blood was harvested for flow cytometry analysis to assess the efficiency of neutrophil depletion; (3) For CD8+ T cells depletion, mice received an intraperitoneal injection of anti-IgG2a or anti-CD8 antibody (Selleck, #A2102) treatment (0.2 mg/mouse, i.p.) 2 days before implantation. To sustain CD8+ T cells depletion, anti-IgG2a or anti-CD8 antibody was administered on day 3, 6, 10 and 13 post-implantations. To confirm the efficacy of CD8+ T cell depletion, mice were euthanized on the 14th day after implantation, and the peripheral blood was harvested for flow cytometry analysis; (4) The administration of vehicle/bromosporine (30 mg/kg, i.p., once a day) began on day 7 post-implantation with the treatment duration lasting for 7 consecutive days; (5) For combinational treatment with bromosporine and anti-PD-1 antibody, the administration of bromosporine (30 mg/kg, i.p., 3 times a week) or anti-PD-1 antibody (0.2 mg/mouse, i.p. twice a week) began on day 6 post-implantation. The tumor was detected through bioluminescence imaging on day 12. After the administration of two 5-day cycles of therapy, separated by a 2-day rest period, six mice per group were euthanized for western blot and flow cytometry analysis. The left 15 mice in each group were used for survival analysis; (6) For the orthotopic mouse model, shNTC, shLDH, and shLDH + CXCL1 KPC-luc cells were orthotopically injected into C57BL/6 J mice. The mice were euthanized on the 16th day after the implantation.
Human samples
In this study, peripheral blood samples from one cohort (22 healthy donors and 27 patients with pancreatic cancer, aged 22–88 including 31 males and 18 females) and formalin-fixed paraffin-embedded (FFPE) PDAC tissues from a separate, non-overlapping cohort (3 male and 3 female patients, aged 54–72) were obtained from residual clinical specimens after routine diagnostic procedures at the First Affiliated Hospital of Nanchang University. The research protocol was approved by the Ethics Committee of the First Affiliated Hospital of Nanchang University ((2024) CDYFYYLK (08-099)). Human PDAC tissue microarrays consisting of 79 tumor and 79 adjacent normal tissues (aged 34–82, 40 males and 39 females) were purchased from Guilin Fanpu Biotech with the ethical approval through the Guilin Fanpu Biotech Ethics Committee (Fanpu-2018[23]). Written informed consent was obtained from all study participants.
Cell lines
PANC02 (STCC20054P) was purchased from Servicebio. BxPC3 (BNCC338477) was obtained from BNCC (Bena Culture Collection, China). PANC1 (CRL-1469) and MIA PaCa-2 (CRL-1420) were purchased from ATCC. The KPC-luc cell line was a generous gift from Prof. Baiyong Shen and Prof. Da Fu53. All the cell lines mentioned above were maintained in Dulbecco’s Modified Eagle Medium (DMEM, Servicebio, #4511) supplemented with 10% fetal bovine serum (FBS, ZETA LIFE #Z7181), 100 U/ml penicillin, and 100 mg/L streptomycin. The cells were maintained in a humidified incubator at 37 °C with an atmosphere of 5% CO2.
Reagents, Primers and shRNAs
2DG (#HY-13966), bromosporine (#HY-15815), WM-1119 (#HY-102058), MG149 (#HY-15887), WM-3835 (#HY-134901), PF-9363 (#HY-132283), CPTH2 (#HY-W013274), PU139 (#HY-124696), and SGC-CBP30 (#HY-15826) were obtained from
the MedChemExpress (MCE), China. Rotenone (#S2348) and Embelin (#S7025) were purchased from Selleck (Shanghai, China). The primers and shRNAs listed in Supplementary Table S1 were purchased from Sangon Biotech Co. Ltd., China.
Plasmids and establishment of stable cell lines
The synthetic shRNAs were annealed and cloned into the pLKO.1-TRC vector. This procedure resulted in the generation of several plasmids: pLKO.1-hPcaf, pLKO.1-mPcaf, pLKO.1-hLdha, pLKO.1-mLdha, pLKO.1-hLdhb, pLKO.1-mLdhb, pLKO.1-hGcn5, pLKO.1-mGcn5, pLKO.1-hP300, pLKO.1-mP300 and pLKO.1-NTC. Additionally, mCxcl1 was amplified by PCR using cDNA from KPC cells as a template. Subsequently, the mCxcl1 gene was inserted into the pLVX-IRES-G418 vector via 2 × In-Fusion Cloning Mix (Servicebio, #G3350). All the constructed plasmids were verified by Sanger sequencing (Sangon Biotech Co. Ltd, China). A mixture of plasmids (psPAX2: pMD2.G: pLKO.1 = 1:1:2) was transfected into HEK293T cells using polyethylenimine (PEI). The supernatant containing virus particles was collected 48 h post-transfection and filtered through a 0.22 µm membrane to infect target cells at approximately 50% confluence. Twenty-four hours post-transfection, the supernatant was replaced with fresh medium containing puromycin (Selleck, #S7417) at a final concentration of 1.5 µg/mL for BxPC3, 2 µg/mL for PANC1, 0.75 µg/mL for KPC-luc, and 1.5 µg/mL for PANC02 to screen stably transfected cell lines.
Quantitative real-time PCR (qPCR) analysis
Total RNA from cultured cells was isolated using the RNA Easy Fast Tissue/Cell Kit (TIANGEN, #DP451). The quality and concentration of the isolated RNA were determined using a NanoDrop spectrophotometer (Thermo Fisher Scientific). The cDNA was generated with 1 µg of total RNA using the HiScript Q RT SuperMix for qPCR (+ gDNA wiper) (Vazyme, #R123-01). The qPCR reactions were performed using 2 × qPCR Master Mix (Vazyme, #Q711-02) on a Bio-Rad detection system (Bio-Rad; CFX96 Touch Real-Time PCR Detection System). The sequences of the qPCR primers used are listed in Supplementary Table S1. Data analysis was performed using the comparative Ct method, and the relative expression levels of the target genes were normalized to Actb mRNA levels. Statistical significance was determined using a t test.
Co-Immunoprecipitation (Co-IP)
For Co-IP, 60 µL Protein A + G beads (Beyotime, #P2108) were washed twice with 1 × TBS and then incubated with 2 µg of anti-rabbit IgG (Abclonal, #WH220308) or anti-L-Lactyl-Histone H3K18 (PTMBIO, #PTM-1427RM) antibodies at 4 °C for 4 h. BxPC3 or KPC cells were lysed, and 100 µL lysate was set aside as input, while the remainder was divided and incubated overnight at 4 °C with the antibody-precoated beads. The next day, the beads were washed thrice with 1 × TBS to remove unbound proteins. Elution was performed by adding 50 µL of 1 × protein loading buffer and incubating at 95 °C in a heating block for 10 min. The supernatant was then collected for the western blot.
ELISA
To examine the secreted CXCL1, 5 × 105 cells per well were cultured in six-well plates for 24 h. The cell culture supernatant was centrifuged at 400 × g for 5 min and collected. The concentration of CXCL1 in cell culture supernatant and plasma of tumor-bearing mice was measured using a Mouse CXCL1 ELISA Kit (Proteintech, #KE10104). The concentration of CXCL1 from healthy individuals and PDAC patients’ plasma was measured using a Human CXCL1 ELISA Kit (Proteintech, #KE00133) according to the manufacturer’s instructions.
Protein extraction and western blot analysis
Proteins from cells and tumor tissues were extracted using RIPA lysis buffer (Beyotime, #P0013B) supplemented with Trichostatin A (TSA, Selleck, #S1045), a cocktail of protease inhibitors (Servicebio, #G2006), and phenylmethylsulfonyl fluoride (PMSF, Servicebio, #G2008). The proteins were then separated by electrophoresis on a 12% polyacrylamide gel and transferred onto a 0.45 µm PVDF membrane. The membrane was blocked with 5% skim milk in TBST to minimize non-specific binding. The membrane was incubated with the primary antibodies overnight at 4 °C, followed by multiple washes with TBST. The primary antibodies used here included anti-LDHA antibody (Proteintech, #19987-1-AP, 1:3000), anti-LDHB antibody (Proteintech, #14824-1-AP, 1:5000), anti-H3 antibody (Proteintech, #17168-1-Ap, 1:3000), anti-PCAF antibody (Cell Signaling Technology, #3378S, 1:1000), anti-P300 antibody (Proteintech, #20695-1-AP, 1:500), anti-GCN5 antibody (Proteintech, #66575-1-Ig, 1:3000), anti-beta Actin antibody (PTMBIO, #PTM-5028, 1:1000), anti-human CXCL1 Polyclonal antibody (Proteintech, #12335-1-AP), anti-mouse CXCL1 Polyclonal antibody (Proteintech, #30322-1-AP), anti-L-Lactyl Lysine antibody (PTMBIO, #PTM-1401RM, 1:1000), anti-L-Lactyl-Histone H3K18 antibody (PTMBIO, #PTM-1406RM, 1:1000), anti-L-Lactyl-Histone H3K9 antibody (PTMBIO, #PTM-1419RM, 1:1000), anti-L-Lactyl-Histone H3K14 antibody (PTMBIO, #PTM-1414RM, 1:1000), anti-H4 antibody (PTMBIO, #PTM-1009, 1:1000), anti-L-Lactyl-Histone H4K12 antibody (PTMBIO, #PTM-1411RM, 1:1000), anti-L-Lactyl-Histone H4K8 antibody (PTMBIO, #PTM-1415RM, 1:1000), anti-Acetyl-Histone H3 (Lys18) antibody (PTMBIO, #PTM-114RM, 1:1000), anti-Acetyllysine antibody (PTMBIO, #PTM-101, 1:1000), and anti-Acetyl-Histone H4 (Lys8) antibody (PTMBIO, #PTM-190, 1:1000). After incubation with the secondary antibody for 1 h, the membrane was rewashed with TBST multiple times. The secondary antibodies included Goat Anti-Rabbit IgG H&L(HRP) pAb (PTMBIO, #PTM-6261, 1:5000) and Goat Anti-Mouse IgG H&L pAb (PTMBIO, #PTM-7168, 1:5000). The immunoreactive bands were visualized using Ultra-High-Sensitive ECL Kit (Servicebio, #G2020-25ML) on an iBright CL750 imaging system (Thermo Fisher Scientific).
Measurement of lactate levels
To measure lactate concentration in tumor tissue, the tissue was homogenized in ice-cold PBS buffer. Following centrifugation at 12000 × g for 10 min at 4 °C, the supernatants were collected, and lactate levels were determined using an L-Lactate Assay Kit (BioAssay Systems, ECLC-100) according to the manufacturer’s instructions. Lactate levels of tumor tissues were normalized to the tumor weight.
In vitro histone H3 lactylation assay
To analyze PCAF-catalyzed histone H3 lactylation, the reaction was performed in 20 μL of 1 × reaction buffer (25 mM Tris-HCl (pH 7.5), 5 mM beta-glycerophosphate, 2 mM dithiothreitol (DTT), 0.1 mM Na3VO4, 10 mM MgCl2) containing 0.5 μg PCAF (Active Motif, #81142), 0.5 μg histone H3 (Active Motif, #31294) and 20 μM lactyl-CoA (MCE, #HY-141540) at 30 °C for 30 min. Then, 5 × SDS loading buffer was added to the reaction and boiled for 5 min at 95 °C. The samples were separated by SDS-PAGE and immunoblotted with the indicated antibodies.
Neutrophil isolation and in vitro migration assay
The mouse neutrophils were isolated from the bone marrow of C57BL/6 J mice using a mouse neutrophil enrichment kit (STEMCELL, #19762RF). The human neutrophils were isolated from the peripheral blood of healthy donors using the human peripheral blood neutrophil isolate kit (TBD Science, #LZS11131) and single nuclear cell isolate kit (TBD Science, #LDS1075) according to the manufacturer’s instructions. The neutrophils (1 × 105 cells per well) in RPMI 1640 (Servicebio, #G4535) supplemented with 10% FBS were seeded in the top chamber of the transwell (Corning, #3421). The conditional media from the cultured PANC1, KPC-luc, and PANC02 cell lines with treatments were collected and added to the bottom layer of the transwell.
For rescue experiments, recombinant human CXCL1/GRO-alpha (Abclonal, #RP01651, 2000 pg/mL) or mouse CXCL1/GRO-alpha (Abclonal, #RP01615, 1000 pg/mL) protein was supplemented in the medium of the bottom layer.
For treatment with inhibitor, purified neutrophils were pre-incubated for 1.5 hours in complete medium containing gradient concentrations of SX-682 (MCE, #HY-119339) and Navarixin (MCE, #HY-10198). Then, the pre-treated neutrophils were seeded in the top chamber of the transwell while the bottom layer was supplied with a corresponding dose of recombinant CXCL1. After incubation at 37°C for 4 h, cells migrated to the bottom chamber were counted.
Assessment of neutrophil viability, degranulation, and ROS
Purified mouse or human neutrophils were incubated in KPC-luc/PANC1 cell-conditional supernatant for 4 h. The cell viability and degranulation of the neutrophils were quantitatively assessed by flow cytometry after stained with the Zombie Aqua™ Fixable Viability Kit (Biolegend, #423102) and mouse or human anti-CD63 antibody (Biolegend, #353010, #143910). For ROS measurement, purified neutrophils were labeled with a fluorogenic ROS detection probe (YEASEN, #50101ES01) after incubation with the respective cell-conditional supernatant. The labeled neutrophils were then analyzed by flow cytometry.
CD8+ T cell isolation and suppression assay
Purified mouse neutrophils were incubated in KPC-luc cell-conditional supernatant or complete medium for 24 h. CD8+ T cells were isolated from C57BL/6 J mouse spleens using positive magnetic selection kit (RWD, #K1303-10). The purified CD8+ T cells (1 × 105 cells per well) were then labeled with CFSE (YEASEN, #40714ES76) to track cell division. Following incubation, the cells were stimulated by 5 µg/mL anti-CD3/CD28 antibody (Selleck, #A2104, #A2166) and 20 ng/mL IL-2 (PeproTech, #212-12) in a 96-well plate at 37 °C in 5% CO2. The aforementioned neutrophils and CD8+ T cells were mixed and co-cultured in different proportions. After 3 days of co-culture, the proliferation of CD8+ T cells was determined by measuring the dilution of CFSE using flow cytometry.
RNA sequencing and data analysis
Total RNA was extracted from BxPC3 and PANC1 cells treated with PBS or 2DG (20 mM for 24 h) using RNAeasy Mini Kit (QIAGEN, #74104), according to the manufacturer’s protocol. Libraries for RNA sequencing were constructed using the Illumina Stranded Total RNA Prep with Ribo-Zero Plus kit (#20040525). The quality and size distribution of the amplified libraries were assessed using a NanoDrop spectrophotometer (Thermo Fisher Scientific) and an Agilent 2100 Bioanalyzer, respectively. Sequencing was performed on Illumina Novaseq 6000 using a paired-end run, yielding approximately 30 million raw reads per sample. Raw sequencing data underwent quality control filtering to obtain high-quality reads. Differential gene expression analysis was performed using the DESeq2 R package. Genes with an adjusted p-value of less than 0.05 and an absolute log2 fold change greater than 1 were considered significantly differentially expressed.
CUT&Tag
The CUT&Tag experiment was performed utilizing the Hyperactive Universal CUT&Tag Assay Kit for Illumina (Vazyme, #TD903). In brief, 1 × 105 BxPC3 cells treated with or without 2DG were harvested and rinsed once with 500 μL wash buffer, followed by attachment to ConA beads at 25 °C for 10 min. Subsequently, the cells were incubated overnight at 4 °C with 1 μg of anti-L-Lactyl Lysine (PTMBIO, #PTM-1401RM) antibody or anti-L-Lactyl-Histone H3K18 (PTMBIO, #PTM-1427RM) antibody. On the following day, anti-rabbit IgG (Abclonal, #WH220308) was introduced and incubated at 25 °C for 1 h. After washing with DIG wash buffer, the cells were exposed to 0.04 μM pA/G–Tnp at 25 °C for 1 h. The washing process was repeated before the cells were resuspended in tagmentation buffer and incubated at 37 °C for 1 hour. Tagmentation was halted by adding proteinase K, buffer LB, and DNA extraction beads, followed by incubation at 55 °C for 10 min. The beads were magnetically separated, washed twice with 80% ethanol, and the DNA was subsequently eluted with double-distilled water. Library preparation was carried out using the TD903 Hyperactive Universal CUT&Tag Assay Kit for Illumina, alongside the TD202 TruePrep Index Kit V2 for Illumina from Vazyme Biotech.
CUT&Tag data processing
Raw paired-end reads underwent adapter removal and quality trimming with trim_galore (v0.6.8). The cleaned FASTQ files were then aligned to the human reference genome sequences (hg38) using BWA (v2.2.9). Duplicate reads were marked and removed with Picard. Peak calling was performed with MACS2 (v2.2.9.1). For signal normalization, Deeptools (v2.5.4) was used to generate coverage tracks with bamCoverage and to calculate signal distributions around peak centers (identified by MACS2) using computeMatrix with the ‘reference-point’ model. The processed data were visualized with plotProfile and plotHeatmap from Deeptools.
Chromatin immunoprecipitation (ChIP)-qPCR
Cells were cultured to 70-80% confluence and crosslinked with 1% formaldehyde at 37 °C for 10 min. Following the BeyoChIP™ ChIP Assay Kit (Protein A/G Bead) (Beyotime, #P2080S) protocol, cells were lysed and subjected to sonication to shear the chromatin. Then, the cell lysates were incubated with anti-L-Lactyl Lysine antibody (PTMBIO, #PTM-1401RM), anti-L-Lactyl-Histone H3K18 antibody (PTMBIO, #PTM-1406RM), anti-Acetyl-Histone H3 (Lys18) antibody (PTMBIO, #PTM-114RM), or anti-Acetyl-Histone H4 (Lys8) antibody (PTMBIO, #PTM-190) at 4 °C overnight. The DNA interacting with the target protein was captured by magnetic beads and precipitated. The immunoprecipitated DNA was reverse crosslinked at 65 °C for 4 hours in the presence of 0.1 M NaHCO3 and 1% SDS. DNA purification was performed using a phenol extraction-ethanol precipitation method, which involved mixing the sheared chromatin with an equal volume of phenol, vigorous vortexing, and subsequent centrifugation to separate phases. The aqueous phase containing DNA was transferred to a new tube, and an equal volume of chloroform mixture was added, followed by another centrifugation. The supernatant was collected, and DNA was precipitated by adding 2.5 volumes of cold 100% ethanol and incubating at − 20 °C overnight. The DNA pellet was collected after centrifugation, washed with 70% ethanol, air-dried, and resuspended in nuclease-free water. qPCR was conducted using the qPCR Master Mix (Vazyme, #Q711-02) on a Bio-Rad detection system (Bio-Rad; CFX96 Touch Real-Time PCR Detection System) with primers specific to the target genomic region. Relative enrichment was calculated using the 2-ΔΔCt method, normalized to input DNA and a negative control region. Statistical significance was assessed using a two-tailed unpaired Student’s t test.
Flow cytometry analysis
Fresh tumors were minced and incubated in RPMI 1640 medium containing mouse tissue digestive enzymes (RWD, #DHTE-5001) at 37 °C for 40 min, followed by filtering through a 40 µm cell strainer (SAINING, #5021000) to prepare a single cell suspension. The cells were incubated with fixable zombie dyes (BioLegend, #423102) at room temperature in the dark for 15 min. Then the cells were rinsed and treated with Fc receptor (BioLegend, #101320) blocking solution for 15 min. Then, the cells were incubated in fluorescence-labeled extracellular antibodies at 4 °C in the dark for 30 min followed by twice rinsing to remove unbound antibodies. For Granzyme B staining, the cells were incubated with the intracellular fixation/permeabilization buffer kit (Elabscience, #E-CK-A109). Finally, the cells were resuspended in 300 µL buffer (PBS + 1% FBS) and subjected to flow cytometry analysis performed on a DxFLEX flow cytometer (Beckman Colter). The data was then analyzed using FlowJo software (Treestar). The antibodies used here included PerCP/Cyanine5.5 anti-mouse CD45 antibody (BioLegend, #157208), Brilliant Violet 421 anti-mouse CD3 antibody (BioLegend, #100227), APC/Fire 750 anti-mouse CD8a antibody (BioLegend, #100766), PE anti-mouse CD279 (PD-1) antibody (BioLegend, #109104), APC anti-human/mouse Granzyme B Recombinant antibody (BioLegend, #372204), Brilliant Violet 421 anti-mouse/human CD11b antibody(BioLegend, #101236), APC anti-mouse Ly-6C antibody (BioLegend, #128016), PE anti-mouse Ly-6G antibody (BioLegend, #127608), APC anti-human CD45 antibody (BioLegend, #368512), FITC anti-human CD66b antibody (BioLegend, #305104), PE anti-human CD11b antibody (BioLegend, #301306), FITC anti-mouse/human CD11b antibody (BioLegend, #101206), and FITC anti-mouse CD3 antibody (BioLegend, #100204).
Mass cytometry by time of flight (CyTOF)
The subcutaneous tumor tissues were collected after vehicle/2DG treatment for 8 consecutive days. Tissues were then rinsed twice with cell culture medium. After sectioning the tissue into 1 mm³ fragments, samples were transferred to a C tube containing a digestive enzyme mixture (Miltenyi Biotec-mouse Tumor Dissociation Kit, #130-096-730) and cell culture medium, adjusted to a total volume of 5 mL. The tissue suspensions were incubated in a shaking incubator at 37 °C for 1 h, then filtered through a 70 μm cell strainer. Cells were collected by centrifugation at 300 × g for 5 min at 2–8 °C. The supernatant was removed, and cells were resuspended in CSB for counting before proceeding with staining. Cells were washed in PBS, stained with 250 nM cisplatin (Fluidigm, # 201194) for 5 min on ice to exclude dead cells, and treated with Fc receptor blocking solution before incubation with a surface antibody cocktail on ice for 30 min. Following two washes with FACS buffer (PBS + 0.5% BSA), cells were fixed overnight in 200 μL intercalation solution (Maxpar Fix and Perm Buffer with 250 nM 191/193Ir, Fluidigm, #201067). Post-fixation, cells were washed sequentially with FACS buffer and perm buffer before staining with intracellular antibody cocktails listed in Supplementary Table S2 on ice for 30 minutes. Each antibody was labeled with specific metal tags using the MaxPAR Antibody Labeling Kit (Fluidigm, #201300) and titrated to determine optimal concentrations prior to use. After final washes, cells were resuspended in deionized water, combined with 20% EQ beads (Fluidigm, #201078), and analyzed on a Helios mass cytometer (Fluidigm, Helios).
Raw data for each sample were first debarcoded using a doublet-filtering scheme54 with unique mass-tagged barcodes. To account for batch effects, .fcs files from different batches were normalized through the bead normalization method55. Data were then manually gated using FlowJo software to exclude debris, dead cells, and doublets, isolating only live, single immune cells. Following this, the X-shift clustering algorithm56 was applied to partition cells into distinct phenotypes based on marker expression levels. Each cluster was annotated by cell type according to its unique marker expression pattern, with visualizations displayed on a heatmap. To facilitate the interpretation of high-dimensional data, the dimensionality reduction algorithm t-SNE57 was employed to create two-dimensional plots showing the distribution of clusters, marker expression, and differences among groups or sample types. Finally, statistical analyses of cell population frequencies were performed using a t test to evaluate significant differences between annotated cell populations.
Immunohistochemistry (IHC) analysis
Tumor tissues were captured and fixed in 10% formalin overnight and embedded in paraffin for H&E staining, immunohistochemistry (IHC), and immunofluorescence (IF). The IHC analysis of H3K18la expression in PDAC TMA (79 paired human PDAC tumor and adjacent normal tissues) was performed by Servicebio (Wuhan, China) with anti-L-Lactyl-Histone H3K18 antibody (PTMBIO, #PTM-1406RM, 1:500). For IHC, tumor tissues were captured and fixed in 10% formalin overnight and embedded in paraffin. Next, the embedded tissues were cut into 4 mm slices, and endogenous peroxidase was eliminated with 3% hydrogen peroxide. The sections were boiled for 5 min in an electric pressure cooker for antigen retrieval in citric acid repair solution (pH 6.0) and then blocked using 3% BSA for 30 min. Then, sections were incubated with primary antibody at 4 °C overnight, followed by incubation with HRP-conjugated secondary antibody for 50 min at room temperature prior to DAB solution. The antibodies used here included anti-L-Lactyl-Histone H3K18 antibody (1:500, PTMBIO, #PTM-1406RM), anti-GRO antibody (1:200, AFFINITY, #AF5403), Ki-67 (1:400, Servicebio, #GB111499), anti-Ly6G antibody (1:500, Servicebio, #GB11229), and anti-CD8 antibody (1:200, Cell Signaling Technology, #70306), and anti-PCAF antibody(1:200, Cell Signaling Technology, #3378).
Multiplex immunofluorescence (IF) analysis
The embedded tissues were sectioned into 4 mm slices, deparaffinized, rehydrated, and antigen retrieved in citric acid repair solution (pH = 6.0) for 3 min under a pressure steam condition. The sections were blocked with 3% hydrogen peroxide and 3% bovine serum albumin (BSA) at room temperature for 30 min respectively, and incubated with anti-CXCL1 antibody (1:200, AFFINITY, #AF5403), anti-CK19 antibody (1:2000, Servicebio, #GB15198), anti-L-Lactyl-Histone H3K18 antibody (1:1000, PTM BIO, #PTM-1406RM), anti-MPO antibody (1:2000, Abcam, #ab208670) and anti-CD8 alpha antibody (1:2000, Abcam, #ab237709) overnight at 4 °C. The sections were washed with PBS and incubated with fluorescein-conjugated secondary antibody at room temperature for 1 h. The slides were imaged and visualized using a digital slide scanner (3DHISTECH, Hungary).
Bioinformatics analysis
Clinical information and RNA sequencing data for pancreatic cancer were obtained from The Cancer Genome Atlas (TCGA-PAAD, n = 179) and CPTAC Pancreatic Cancer dataset (n = 140)58, accessible via cBioPortal (https://www.cbioportal.org). Additionally, pan-cancer cohorts comprising transcriptomic data from 33 cancer types and comprehensive survival information were retrieved from TCGA. The glycolysis-related genes used in this study originated from previously published research59 and are listed in Supplementary Table S3. Based on the expression of the glycolysis-related genes, samples were classified into high and lowglycolysis-level groups according to the Single-sample Gene Set Enrichment Analysis (ssGSEA) scores, implemented via the GSEA R package. The QUANTISEQ60 algorithm was employed to estimate the infiltration proportions of various immune cells for each sample based on gene expression profiling data. Then, differences in the infiltration proportions of neutrophils and CD8+ T cells between high and low glycolysis-level groups were visualized using the ggplot2 R package. The survival and survminer R packages were utilized to compare survival probabilities. The relationships between immune cell infiltration and the expression of core glycolysis-related genes were assessed using Spearman’s correlation coefficient.
Molecular docking
The structures of acetyl-CoA and lactyl-CoA were retrieved from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Using OpenBabel 3.1.1, the structures were converted from SDF format to MOL2 format, then imported into AutoDock Tools 1.5.7 and saved as PDBQT files. The protein structure (PDB ID: 1CM0) was downloaded from the PDB database, and PyMOL was used to remove unnecessary molecules, saving the processed protein as a PDB file for molecular docking. AutoDock Tools 1.5.7 was used to prepare the protein structure by adding hydrogens and charges, and the result was saved as a PDBQT file. Molecular docking was performed using the semi-flexible docking method in AutoDock. Based on the binding affinity calculated by the scoring function, the top-ranked compound with the highest docking score was selected for visualization and analysis.
Statistics and reproducibility
Statistical analyses were conducted using a two-tailed unpaired Student’s t test to assess significant differences between groups. Survival analysis p-values were determined using the log-rank test. All statistical tests were carried out using Prism 9 software (GraphPad Software) or R version 4.4.1. The respective figure legends provide comprehensive details regarding the statistical methods, the number of observations or replicates, and definitions of center and dispersion. No data were excluded from the analyses. The data used for analysis were from three or more independent experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw sequencing data generated in this study have been deposited in the Genome Sequence Archive (GSA) at the National Genomics Data Center (NGDC, Beijing, China) under BioProject accessions: RNA-seq: PRJCA038857 and CUT&Tag-seq: PRJCA038880. All publicly released data comply with the Ministry of Science and Technology of China guidelines. The RNA sequencing data of the Cancer Genomic Atlas Pancreatic adenocarcinoma (TCGA-PAAD) cohort and pan-cancer cohort from 33 cancer types with comprehensive survival information used in this study are available at https://portal.gdc.cancer.gov/. Transcriptomic data of Clinical Proteomic Tumor Analysis Consortium (CPTAC) Pancreatic Cancer cohort are available at https://proteomics.cancer.gov/programs/cptac. Data analysis and visualization were performed using open-source software packages. Where applicable, we adapted published algorithms with proper citation. No custom code was generated for this study. Source data are provided in this paper.
References
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024).
O’Reilly, E. M. et al. Durvalumab with or without tremelimumab for patients with metastatic pancreatic ductal adenocarcinoma: a phase 2 randomized clinical trial. JAMA Oncol. 5, 1431–1438 (2019).
Bear, A. S., Vonderheide, R. H. & O’Hara, M. H. Challenges and opportunities for pancreatic cancer immunotherapy. Cancer Cell 38, 788–802 (2020).
Leinwand, J. & Miller, G. Regulation and modulation of antitumor immunity in pancreatic cancer. Nat. Immunol. 21, 1152–1159 (2020).
Jaillon, S. et al. Neutrophil diversity and plasticity in tumour progression and therapy. Nat. Rev. Cancer 20, 485–503 (2020).
Veglia, F., Sanseviero, E. & Gabrilovich, D. I. Myeloid-derived suppressor cells in the era of increasing myeloid cell diversity. Nat. Rev. Immunol. 21, 485–498 (2021).
Wang, C. et al. CD300ld on neutrophils is required for tumour-driven immune suppression. Nature 621, 830–839 (2023).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Li, F. & Simon, M. C. Cancer Cells Don’t Live Alone: Metabolic Communication within Tumor Microenvironments. Dev. Cell 54, 183–195 (2020).
Comito, G. et al. Lactate modulates CD4(+) T-cell polarization and induces an immunosuppressive environment, which sustains prostate carcinoma progression via TLR8/miR21 axis. Oncogene 38, 3681–3695 (2019).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
Brand, A. et al. LDHA-Associated lactic acid production blunts tumor tmmunosurveillance by T and NK cells. Cell Metab. 24, 657–671 (2016).
Cascone, T. et al. Increased tumor glycolysis characterizes immune resistance to adoptive T cell therapy. Cell Metab. 27, 977–987 (2018).
Guo, D. et al. Aerobic glycolysis promotes tumor immune evasion by hexokinase2-mediated phosphorylation of IkappaBalpha. Cell Metab. 34, 1312–1324 (2022).
Wu, L. et al. Tumor aerobic glycolysis confers immune evasion through modulating sensitivity to T cell-mediated bystander killing via TNF-alpha. Cell Metab. 35, 1580–1596.e1589 (2023).
Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 12, 31–46 (2022).
Yao, W., Hu, X. & Wang, X. Crossing epigenetic frontiers: the intersection of novel histone modifications and diseases. Signal Transduct. Target. Ther. 9, 232 (2024).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Yu, J. et al. Histone lactylation drives oncogenesis by facilitating m(6)A reader protein YTHDF2 expression in ocular melanoma. Genome Biol. 22, 85 (2021).
Zhang, C. et al. H3K18 Lactylation potentiates immune escape of non-small cell lung cancer. Cancer Res. 84, 3589–3601 (2024).
Wang, R. et al. H3K9 lactylation in malignant cells facilitates CD8(+) T cell dysfunction and poor immunotherapy response. Cell Rep. 43, 114957 (2024).
Xiong, J. et al. Lactylation-driven METTL3-mediated RNA m(6)A modification promotes immunosuppression of tumor-infiltrating myeloid cells. Mol. Cell 82, 1660–1677 (2022).
Li, H., Sun, L., Gao, P. & Hu, H. Lactylation in cancer: Current understanding and challenges. Cancer Cell 42, 1803–1807 (2024).
Zhu, R. et al. ACSS2 acts as a lactyl-CoA synthetase and couples KAT2A to function as a lactyltransferase for histone lactylation and tumor immune evasion. Cell Metab. 37, 361–376.e367 (2025).
Ju, J. et al. The alanyl-tRNA synthetase AARS1 moonlights as a lactyltransferase to promote YAP signaling in gastric cancer. J. Clin. Invest. 134, e174587 (2024).
Zong, Z. et al. Alanyl-tRNA synthetase, AARS1, is a lactate sensor and lactyltransferase that lactylates p53 and contributes to tumorigenesis. Cell 187, 2375–2392 (2024).
Li, H. et al. AARS1 and AARS2 sense L-lactate to regulate cGAS as global lysine lactyltransferases. Nature 634, 1229–1237 (2024).
Mao, Y. et al. Hypoxia induces mitochondrial protein lactylation to limit oxidative phosphorylation. Cell Res. 34, 13–30 (2024).
Niu, N. et al. Tumor cell-intrinsic SETD2 deficiency reprograms neutrophils to foster immune escape in pancreatic tumorigenesis. Adv. Sci. 10, e2202937 (2023).
Liu, J. et al. QDPR deficiency drives immune suppression in pancreatic cancer. Cell Metab. 36, 984–999 (2024).
Ozga, A. J., Chow, M. T. & Luster, A. D. Chemokines and the immune response to cancer. Immunity 54, 859–874 (2021).
Nywening, T. M. et al. Targeting both tumour-associated CXCR2(+) neutrophils and CCR2(+) macrophages disrupts myeloid recruitment and improves chemotherapeutic responses in pancreatic ductal adenocarcinoma. Gut 67, 1112–1123 (2018).
Kwak, J. W. & Houghton, A. M. Targeting neutrophils for cancer therapy.Nat. Rev. Drug Discov.24, 666–684 (2025).
Dong, X. et al. Metabolic lactate production coordinates vasculature development and progenitor behavior in the developing mouse neocortex. Nat. Neurosci. 25, 865–875 (2022).
Zhang, D. et al. Lysine L-lactylation is the dominant lactylation isomer induced by glycolysis. Nat. Chem. Biol. 21, 91–99 (2025).
Zhao, D. et al. Lysine-5 acetylation negatively regulates lactate dehydrogenase A and is decreased in pancreatic cancer. Cancer Cell 23, 464–476 (2013).
Li, F. et al. Positive feedback regulation between glycolysis and histone lactylation drives oncogenesis in pancreatic ductal adenocarcinoma. Mol. Cancer 23, 90 (2024).
Wang, X. et al. Histone lactylation dynamics: Unlocking the triad of metabolism, epigenetics, and immune regulation in metastatic cascade of pancreatic cancer. Cancer Lett. 598, 217117 (2024).
Gatenby, R. A. & Gillies, R. J. Why do cancers have high aerobic glycolysis? Nat. Rev. Cancer 4, 891–899 (2004).
Llibre, A., Kucuk, S., Gope, A., Certo, M. & Mauro, C. Lactate: A key regulator of the immune response. Immunity 58, 535–554 (2025).
Chang, C. H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Sukumar, M. et al. Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function. J. Clin. Invest. 123, 4479–4488 (2013).
Tang, X. et al. Glycolysis inhibition induces anti-tumor central memory CD8(+)T cell differentiation upon combination with microwave ablation therapy. Nat. Commun. 15, 4665 (2024).
Greco, B. et al. Disrupting N-glycan expression on tumor cells boosts chimeric antigen receptor T cell efficacy against solid malignancies. Sci. Transl. Med. 14, eabg3072 (2022).
Raychaudhuri, D. et al. Histone lactylation drives CD8(+) T cell metabolism and function. Nat. Immunol. 25, 2140–2151 (2024).
Ugolini, A. et al. Functional reprogramming of neutrophils within the brain tumor microenvironment by hypoxia-driven histone lactylation. Cancer Discov. 15, 1270–1296 (2025).
Nagy, Z. & Tora, L. Distinct GCN5/PCAF-containing complexes function as co-activators and are involved in transcription factor and global histone acetylation. Oncogene 26, 5341–5357 (2007).
Kim, J. J. et al. PCAF-Mediated histone acetylation promotes replication fork degradation by MRE11 and EXO1 in BRCA-deficient cells. Mol. Cell 80, 327–344 (2020).
Seifert, L. et al. The necrosome promotes pancreatic oncogenesis via CXCL1 and Mincle-induced immune suppression. Nature 532, 245–249 (2016).
Kemp, S. B. et al. Apolipoprotein E Promotes Immune Suppression in Pancreatic Cancer through NF-kappaB-Mediated Production of CXCL1. Cancer Res. 81, 4305–4318 (2021).
Yang, J. et al. KDM6A Loss recruits tumor-associated neutrophils and promotes neutrophil extracellular trap formation in pancreatic cancer. Cancer Res. 82, 4247–4260 (2022).
Bianchi, A. et al. Cell-autonomous Cxcl1 sustains tolerogenic circuitries and stromal inflammation via neutrophil-derived TNF in pancreatic cancer. Cancer Discov. 13, 1428–1453 (2023).
Chen, D. et al. Pan-cancer analysis implicates novel insights of lactate metabolism into immunotherapy response prediction and survival prognostication. J. Exp. Clin. Cancer Res. 43, 125 (2024).
Zunder, E. R. et al. Palladium-based mass tag cell barcoding with a doublet-filtering scheme and single-cell deconvolution algorithm. Nat. Protoc. 10, 316–333 (2015).
Finck, R. et al. Normalization of mass cytometry data with bead standards. Cytometry A 83, 483–494 (2013).
Samusik, N., Good, Z., Spitzer, M. H., Davis, K. L. & Nolan, G. P. Automated mapping of phenotype space with single-cell data. Nat. Methods 13, 493–496 (2016).
Laurens, V. D. M. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Cao, L. et al. Proteogenomic characterization of pancreatic ductal adenocarcinoma. Cell 184, 5031–5052.e5026 (2021).
Zappasodi, R. et al. CTLA-4 blockade drives loss of T(reg) stability in glycolysis-low tumours. Nature 591, 652–658 (2021).
Finotello, F. et al. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 11, 34 (2019).
Acknowledgements
We thank Dr. Baiyong Shen and Dr. Da Fu (Ruijin Hospital, Shanghai Jiao Tong University School of Medicine) for providing the KPC-luciferase cell line. We also thank all the members of the Center for Molecular Diagnosis and Precision Medicine for the discussions. This study was supported by the National Natural Science Foundation of China (32200639 to P.Z.), the Natural Science Foundation of Jiangxi Province, China (20224BAB216066 to P.Z.), the Jiangxi Province ‘Ganpo Talent Program’ Innovation Leadership Talent (gpyc20240198 to P.Z.), the Jiangxi Medical Academy of Nutrition and Health Management (2022-PYXM-03 to C.Z.), and the start-up fund from the First Affiliated Hospital of Nanchang University (500021010 to C.Z., 500021017 to P.Z.).
Author information
Authors and Affiliations
Contributions
C.Z. and P.Z. conceptualized the study and designed the study. C.Z. and P.Z. acquired the funding support and supervised the study. C.Z., P.Z., and C.L. wrote the manuscript. P.Z., J.R.M., and Y.W. performed most of the experiments and statistical analysis. C.L. and Z.W.Z. performed bioinformatics analysis. L.L., M.H., N. Z., Y.M., J.H., L.Z., X.L., J.G., T.Z., J.P.M., Y.D., Z.Y.Z., J.Y., X.W., and T.G. participated in human sample collection, technical assistance and manuscript revision. All authors have read and approved the final manuscript. All the authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, P., Ma, J., Wan, Y. et al. Histone lactylation increases CXCL1 expression for neutrophil infiltration and immune escape in pancreatic cancer. Nat Commun 17, 2526 (2026). https://doi.org/10.1038/s41467-026-69311-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69311-5