Abstract
Pulmonary vascular development is essential for alveolarization, and disruption of this process contributes to pathogenesis of bronchopulmonary dysplasia (BPD). Proper vascular development requires an orchestration of many cell types within the lung. However, the transcriptional mechanisms by which pericytes support the endothelium in the postnatal lung remain poorly understood. Herein, we identify FOXF2 as a critical transcription factor that governs pericyte maturation and function during postnatal lung development and regeneration. FOXF2 expression in pericytes increases postnatally and is selectively downregulated after neonatal hyperoxic injury. Pdgfrb-CreER mediated Foxf2 deletion in pericytes leads to pericyte hyperplasia, impaired migration, and reduced expression of angiogenic factors such as ANGPTL4. Transcriptomic and genomic studies demonstrate that FOXF2 maintains chromatin accessibility at pro-angiogenic loci and modulates paracrine signaling essential for endothelial regeneration. Loss of FOXF2 disrupts pericyte–endothelial crosstalk, leading to impaired angiogenesis and alveolarization as well as increased vascular permeability after neonatal lung injury. Altogether, FOXF2 acts as a key transcriptional regulator of the pericyte-driven vascular niche in the neonatal lung, highlighting the pathogenic role of pericyte dysfunction in BPD.
Similar content being viewed by others
Introduction
The lung’s respiratory function relies on an extensive capillary network underlying a thin epithelial layer, predominantly composed of alveolar type 1 (AT1) cells. Pulmonary alveolar development and maturation are driven by intricate interactions between parenchymal and mesenchymal cells. These interactions enable the formation of the air–blood barrier, composed of AT1 and capillary endothelial cells in close proximity1,2,3. However, in premature neonates, the need for mechanical ventilation and supplemental oxygen can injure this barrier and disrupt proper angiogenesis and alveologenesis4. These pulmonary insults, particularly in preterm infants, are linked to early-life morbidity and long-term respiratory complications5.
While the roles of mesenchymal and epithelial cells in alveolar development are well-characterized, the contribution of the vascular system to alveolarization remains poorly understood. Recent studies from our labs and others have underscored the critical roles of angiogenic factors, such as FOXF1, FOXM1, and VEGFA, in driving capillary cell proliferation and differentiation during alveologenesis6,7,8. Nevertheless, the intercellular signaling mechanisms governing the development of the alveolar microvasculature remain largely elusive. Recent studies suggest that the lung capillary network consists of two intermingled subpopulations of endothelial cells, the general capillary (gCaps or CAP1) that serve as vascular stem cells and alveolar capillary endothelial cells (aCaps or CAP2) which are terminally differentiated and located in close apposition with AT1 cells9. Intriguingly, only gCaps are in contact with pericytes, suggesting a potential role of pericytes-endothelium signaling for pulmonary angiogenesis8,9. In support of this notion, pericyte-specific inactivation of Hippo effector YAP and TAZ impaired lung capillary angiogenesis via disruption of the paracrine ANGPT1-TIE2 signaling in endothelial cells10. The roles of pericyte-derived paracrine factors have also been reported in other organs, highlighting the crucial role of pericytes as an essential orchestrator for angiogenesis and tissue homeostasis11,12.
Forkhead box F2 (FOXF2, also known as LUN and FREAC2) is a key transcription factor for mesenchymal development13,14,15. FOXF2 functions independently or partially redundantly with its paralog, FOXF1, in various developing contexts. FOXF2 has been shown to play a cell-specific role in regulating pericyte function16,17. Global deletion of Foxf2 leads to hyperplasia and defective differentiation of brain pericytes, hampering blood-brain barrier formation and homeostasis17. However, the role of FOXF2 in lung pericytes has not been investigated.
In the present study, we show that Foxf2 expression is downregulated in pericytes following postnatal hyperoxic lung injury. While Foxf2 expression in pericytes is dispensable for normal alveolar architecture, its deletion induces a premature and hyperproliferative pericyte phenotype and impairs their migratory capacity. Loss of Foxf2 in pericytes exacerbates hyperoxic lung injury, resulting in hypoxemia accompanied by pronounced alveolar simplification, increased endothelial permeability, and impaired alveolar angiogenesis. Genomic and transcriptomic profiling demonstrate that FOXF2 transcriptionally regulates pericyte genes required for angiogenesis, including Angptl4. Our findings indicate that FOXF2 is essential for pericyte maturation and pericyte–endothelial crosstalk, maintaining the homeostasis of the pulmonary capillary network required for efficient gas exchange in alveoli.
Results
Foxf2 expression correlates with pericyte maturation genes in postnatal alveologenesis
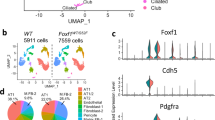
Pericytes play a critical role in postnatal alveolar formation and pulmonary angiogenesis10. To examine temporal changes in pericyte gene expression, we utilized the TDEseq package to analyze gene expression profiles across multiple developmental stages (embryonic day (E)12.5 to postnatal day (P) 42) using single-cell RNA sequencing (scRNA-seq) data18,19. We identified distinct expression patterns, categorized as “growth,” “peak,” “recession,” and “trough.” Genes exhibiting “growth” or “peak” patterns, such as Trpc6, Des, Rgs5, and Pdgfrb, are well-established pericyte identity markers. Notably, genes previously reported associated with mature pericytes, including Des, Vtn, and Higd1b, were classified under the “growth” pattern (Fig. 1a)20,21. Functional enrichment analysis revealed that “growth” pattern genes were enriched in processes related to vascular development, migration, and angiogenesis, whereas “recession” pattern genes were associated with mitotic cell cycle functions (Fig. 1b, c).
a Time course analysis of developmental gene expression pattern in pericytes using TDEseq. b Toppgene enrichment results for genes with the “growth” pattern. c Toppgene enrichment result for genes with the “recession” pattern. d Violin plot showing the expression of Foxf2 in pericytes at different time points during lung development. e UMAP plot of mesenchymal cell types of developing mouse lungs. Mes, mesenchymal; AF1, alveolar fibroblast 1; AF2, alveolar fibroblast 2; MyoFb, myofibroblast; ASMC, airway smooth muscle cell; VSMC, vascular smooth muscle cell. f Feature plots showing the expression pattern of selected mature pericyte genes that were categorized into the “growth” pattern. g Representative immunostaining images for PDGFRβ, αSMA and FOXF2 on lung sections at P7, arrows indicate pericytes. FOXF2 is expressed in PDGFRβ+ pericytes but not in vascular smooth muscle cells. Data are representative of three independent mice. Scale bar=50 μm. Arrows indicate FOXF2+ pericytes.
Interestingly, Foxf2 was among the genes displaying a “growth” pattern (Fig. 1a). Its expression remained relatively stable during embryonic development but increased markedly in postnatal lungs, reaching a higher level by P42 (Fig. 1d). UMAP plot showed higher Foxf2 expression in pericytes, which appeared to align with expression of mature pericyte marker genes, such as Higd1b, Des, and Trpc6. By comparison, Foxf2 expression was lower in alveolar fibroblasts (AF1 and AF2) and myofibroblasts (Fig. 1e, f). Immunofluorescence indicated that Foxf2 was present in PDGFRβ+ pericytes but absent in vascular smooth muscle cells (Fig. 1g). These observations suggest that Foxf2 expression is associated with the pericyte maturation gene program during postnatal lung development.
Foxf2 is required for cell quiescence of postnatal lung pericytes
Previous studies have shown that Foxf2 deficiency impairs brain pericyte function17. To determine whether Foxf2 is required for pericyte function during postnatal pulmonary alveologenesis, we generated mice with Pdgfrb-CreER-mediated Foxf2 deletion (designated Foxf2 conditional knockout (cKO)). Two doses of tamoxifen were administered at P0 and P1 to induce Cre-mediated recombination (Fig. 2a). Immunostaining confirmed selective loss of FOXF2 in PDGFRβ+ pericytes (Fig. 2b, c). Quantitative RT-PCR (qRT-PCR) revealed reduced Foxf2 mRNA levels in whole lung tissue and magnetic activated cell sorting (MACS)-purified PDGFRβ+ cells at P21 (Fig. 2d, e). Histological analysis indicated that alveolar structures in Foxf2 cKO and control mice were similar, with no differences in mean linear intercept (MLI) or blood oxygenation (Supplementary Fig. 1a-c). However, immunostaining for PDGFRβ showed an increased number of pericytes in Foxf2 cKO lungs, with PDGFRβ+ cells densely packed within alveolar walls (Fig. 2f, g). Quantification of endomucin-stained areas revealed a modest reduction in capillary density (Fig. 2f, h).
a Schematic representative of treatment of Foxf2 cKO and control (Ctrl) mice. Tamoxifen was injected at P0 and P1, tissues were harvested at P7 and P21 for analyses. b Representative immunostaining images for FOXF2 and PDGFRβ at P21. Scale bar=25 μm. c Quantification of the percentage of FOXF2-positive pericytes. n = 4 for Ctrl and 5 for Foxf2 cKO. p < 0.0001. d qPCR analysis showing mRNA level of Foxf2 from lung tissue at P21. n = 10 for Ctrl and 11 for Foxf2 cKO. p < 0.0001. e qPCR analysis showing mRNA level of Foxf2 from MACS-sorted pericytes. n = 4 for Ctrl and 4 for Foxf2 cKO. p < 0.0001. f Representative immunostaining images for Endomucin and PDGFRβ at P21. Arrows indicate pericytes. Scale bar = 100 μm. g Quantification of the percentage of PDGFRβ-positive cells (pericytes). n = 10 for Ctrl and 10 for Foxf2 cKO. p = 0.0079. h Quantification of the percentage of endomucin covered area per field of view. n = 5 for Ctrl and 7 for Foxf2 cKO. p = 0.0563. i Representative FACS panel showing CD140b+ (PDGFRβ+) cells (pericytes) at P21. j Quantification of the percentage of CD140b+ cells. n = 6 for Ctrl and 7 for Foxf2 cKO. p = 0.0130. k Representative FACS panel showing cell cycle distribution of CD140b+ cells at P7. l Quantification of percentage of CD140b+ cells in each cell cycle phase. n = 7 for Ctrl and 6 for Foxf2 cKO. p = 0.0104 for G0/G1 phase, 0.0008 for S phase, and 0.1723 for G2/M phase. m Representative immunostaining images for PDGFRβ and Ki-67 on lung sections at P7. Arrows indicate Ki-67+ pericytes. Scale bar = 50 μm. n Quantification of the percentage of Ki-67 positive pericytes. n = 9 for Ctrl and 10 for Foxf2 cKO. p < 0.0001. All p-values were determined by an unpaired 2-tailed t test. Data are presented as mean values ± SEM. Source data are provided as a Source Data file.
Fluorescence-activated cell sorting (FACS) analysis confirmed a higher proportion of CD140b+ (PDGFRβ+) cells in Foxf2 cKO lungs, with no changes in PDGFRα+ (CD140a+) fibroblasts, epithelial cells (CD326+) and endothelial cells (CD31+) at P21 (Fig. 2i, j; Supplementary Fig. 2a, b). FACS analysis of EdU incorporation revealed a nearly 2-fold increase in the number of S-phase cells and a corresponding decrease in G0/G1-phase cells among CD140b+ cells in Foxf2 cKO lungs (Fig. 2k, l). We observed a slight shift in cell cycle distribution in CD31+ cells, with a trend toward increased S-phase cells and decreased G2/M-phase cells, while G0/G1-phase cells remained stable following Foxf2 knockout (Supplementary Fig. 3a-c). This observation is consistent with the unchanged number of CD31+ cells seen at P21 (Supplementary Fig. 2a, b). Co-immunostaining for PDGFRβ and Ki-67 revealed a higher number of Ki-67-positive pericytes in Foxf2 cKO mice compared to controls (Fig. 2m, n). In cultured PDGFRβ+ cells, siRNA-mediated Foxf2 knockdown led to increased Bromodeoxyuridine (BrdU) incorporation, indicating that Foxf2 inactivation cell autonomously increases pericyte proliferation (Supplementary Fig. 1d-f). While pulmonary vasculature was slightly affected in Foxf2 cKO lungs, the alveolar epithelial differentiation program appeared normal, with AT1 marker T1α and AT2 marker proSPC normally presented, quantification of AT2 cells revealed no change after pericyte-specific deletion of Foxf2 (Supplementary Fig. 1g, h). Together, these data suggest that FOXF2 is required to maintain quiescence of pericytes during postnatal alveologenesis.
Loss of Foxf2 led to reduced expression of genes associated with normal pericyte function
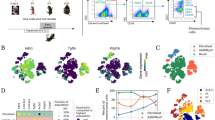
We next performed single-cell RNA sequencing (scRNA-seq) analysis of non-immune (CD45-) cells to interrogate the cellular and molecular changes, which captured all major cell types in control and Foxf2 cKO lungs at P7 (Fig. 3a, b). While no change in the proportion of endothelial cells was detected (Supplementary Fig. 2c), we found that expression of selected genes associated with the “growth” pattern and angiogenesis, including Angpt2 and Angptl4 were downregulated in Foxf2 cKO pericytes (Fig. 3c). Due to the limited ability to determine gene expression changes by scRNA-seq, we also performed deep bulk RNA-seq on MACS-sorted PDGFRβ+ pericytes (Fig. 3d). Downregulated genes in Foxf2 cKO pericytes included those associated with pericyte function (Yap1 and Wwtr1), maturation (Itga8 and Lama4), migration (Itga8 and Amotl1), and angiogenesis (Angptl4 and Angpt2). In contrast, genes linked to cell proliferation, such as Foxm1 and Top2a, were upregulated (Fig. 3e). Consistent with these findings, functional enrichment analysis showed that downregulated genes were related to Gene Ontology (GO) terms such as vascular development and cell migration, while upregulated genes were associated with ATP synthesis and oxidative phosphorylation, processes tied to cell proliferation capacity (Fig. 3f). qRT-PCR analysis confirmed reduced expression of mature pericyte genes, including Itga8, Lama4, and Trpc6, and angiogenesis-related genes like Angptl4 in sorted CD140b+ cells at P7 (Fig. 3g)22,23. In agreement with decreased genes associated with cell migration, isolated PDGFRβ+ pericytes from Foxf2 cKO mice showed reduced migration capacity in vitro (Fig. 3h, i). A previous study reported that increased PI3K activity inhibits pericyte maturation20. Consistent with this observation, we detected elevated p-AKT intensity in Foxf2-deficient pericytes in the alveolar region (Fig. 3j, k). Cultured CD140b+ cells isolated from Foxf2 cKO lungs exhibited a marked increase in AKT phosphorylation, indicating that FOXF2 suppresses PI3K/AKT activation in a cell-autonomous manner (Fig. 3l-n). These findings indicate that FOXF2 is essential for gene programs governing pericyte maturation and function.
a UMAP plots of cell clusters in Ctrl and Foxf2 cKO mice lung at P7 under room air. b Cell-type proportion is shown by different colors, as same color-coding scheme shown in UMAP plot. c Violin plot showing expression of Foxf2, Higd1b, Angptl4 and Angpt2 in pericytes of lung from Ctrl and Foxf2 cKO mice. d Experimental schematics for analyses on P7. e Genes differentially expressed in Foxf2 cKO vs. Ctrl pericytes. Y-axis denotes − log10(P.adj), and x-axis shows log2 fold change. f Gene Ontology enrichment for genes altered in pericytes of Foxf2 cKO mice. g qRT-PCR analysis showing mRNA level of Itga8, Lama4, Trpc6 and Angptl4 from MACS purified pericytes at P7. n = 11 for Ctrl and 9 for Foxf2 cKO for Itga8, p = 0.0022; n = 10 for Ctrl and 11 for Foxf2 cKO for Lama4, p = 0.0012; n = 10 for Ctrl and 9 for Foxf2 cKO for Trpc6, p = 0.0021; n = 9 for Ctrl and 9 for Foxf2 cKO for Angptl4, p = 0.0023. h Representative images for wound healing assay of MACS sorted pericytes. Scale bar=200 μm. i Quantification of the percentage of wound healing recovery distance. n = 8 for Ctrl and 7 for Foxf2 cKO. p = 0.0464. j Representative images for PDGFRβ and p-AKT immunostaining on lung sections. Scale bar = 50 μm. Arrows indicate p-AKT hyperactivation in Foxf2 cKO pericytes. k Quantification of p-AKT intensity in each PDGFRβ+ cells. n = 424 cells from 3 Ctrl and 1496 cells from 3 Foxf2 cKO at P7. p < 0.0001. l Representative western blot images for protein bands of p-AKT, AKT, FOXF2 and β-actin in cultured CD140b+ cells. m Quantification of relative ratio of p-AKT/AKT and n FOXF2/β-actin. n = 4 for Ctrl and 6 for Foxf2 cKO, p = 0.0015 for p-AKT/AKT and 0.0017 for FOXF2/β-actin. All p values were determined by an unpaired 2-tailed t test. Data are presented as mean values ± SEM. Source data are provided as a Source Data file.
Pericytes have been reported to send signals that support AT2 cell proliferation10, while AT2 cell counts remained unchanged following pericyte-specific Foxf2 deletion (Supplementary Fig. 1g, h), we observed downregulation of host-defense and lipid metabolism genes linked to canonical AT2 cell function, along with upregulation of ER stress-related genes in AT2 cells from scRNA-seq data (Supplementary Fig. 2d, e). These findings suggest that Foxf2 loss in pericytes may also influence certain aspects of AT2 cell biology.
Pericyte Foxf2 loss worsens neonatal lung injury, impairs angiogenesis and increases vascular permeability
To further investigate the role of FOXF2 in postnatal alveolar development, we analyzed scRNA-seq datasets from human BPD samples (Supplementary Fig. 4a, b)24. Consistent with murine data, FOXF2 was highly expressed across multiple mesenchymal subtypes (Supplementary Fig. 4c), but its expression was markedly reduced in pericytes from BPD lungs (Supplementary Fig. 4d). Similarly, in a mouse model of BPD induced by postnatal hyperoxia25, Foxf2 expression was selectively downregulated in pericytes compared to room air (RA) controls, with minimal changes observed in other mesenchymal populations (Supplementary Fig. 4e, f). These transcriptomic findings were further supported by immunostaining of lung tissue sections, which revealed a reduced number of FOXF2+ cells among PDGFRβ+ pericytes in hyperoxia-exposed lungs (Fig. 4a-c), as well as by western blot analysis of whole lung lysates, which showed decreased FOXF2 protein in hyperoxia-treated lungs (Fig. 4d-e).
a Schematics of hyperoxia treatment of neonatal mice. b Representative images for FOXF2 and PDGFRβ immunostaining in pericytes of mice in room air and hyperoxia. Scale bar = 50 μm. Arrows indicate pericytes. c Quantification of the percentage of FOXF2+ pericytes. n = 6 for room air and 7 for hyperoxia group. p < 0.0001. d Representative western blot images for protein bands of FOXF2 and β-actin in whole lung lysates from mice pups under room air and hyperoxia. e Quantification of relative ratio of FOXF2/β-actin. n = 4 for room air and 3 for hyperoxia group. p = 0.0023. f Table shows the 10-day mortality in Ctrl and Foxf2 cKO mice treated with room air or hyperoxia. g Measurements of arterial oxygenation showing decreased oxygen saturation in Foxf2 cKO mice compared to Ctrl, when exposed to hyperoxia at P7. n = 6 for Ctrl and 4 for Foxf2 cKO. p = 0.0117. h Representative H&E staining showing P7 lungs of Ctrl and Foxf2 cKO mice treated with room air or hyperoxia. Scale bar = 100 μm. i Quantification of MLI. n = 13 for Ctrl with room air, 9 for Foxf2 cKO with room air, 9 for Ctrl with hyperoxia, and 10 for Foxf2 cKO with hyperoxia. p = 0.9998 for Ctrl vs. Foxf2 cKO at room air, p < 0.0001 for Ctrl +hyperoxia vs. Ctrl + room air, p < 0.0001 for KO + hyperoxia vs. KO + room air, and p < 0.0001 for Ctrl vs. Foxf2 cKO at hyperoxia. j Representative images for dextran-FITC for mice at P7 in room air and hyperoxia. Scale bar=50 μm (k) Quantification of dextran-FITC area. n = 6 for Ctrl with room air, 3 for Foxf2 cKO with room air, 3 for Ctrl with hyperoxia, and 6 for Foxf2 cKO with hyperoxia. p = 0.5495 for Ctrl vs. Foxf2 cKO at room air, p = 0.3496 for Ctrl +hyperoxia vs. Ctrl + room air, p = 0.0052 for KO + hyperoxia vs. KO + room air, and p = 0.0102 for Ctrl vs. Foxf2 cKO at hyperoxia. p-values in (c), (e) and (g) were determined by an unpaired 2-tailed t test, and p-values in (i) and (k) were determined by one-way ANOVA followed by Sidak’s multiple comparison test. Data are presented as mean values ± SEM. Source data are provided as a Source Data file.
Decreased FOXF2 expression in pericytes, observed in both human BPD and a mouse hyperoxia model through scRNA-seq datasets, indicates that FOXF2 function in pericytes might be essential for endothelial regeneration in postnatal lung injury. To test this hypothesis, Ctrl and Foxf2 cKO pups administered tamoxifen were exposed to 75% O2 for 7 days (Fig. 4a). While Foxf2 deletion did not affect pup survival under room air, survival rates in Foxf2 cKO mice decreased following hyperoxia treatment (Fig. 4f). Consistent with reduced survival, lower oxygen saturation was observed in hyperoxia-treated Foxf2 cKO mice compared to controls (Fig. 4g). Histological analysis revealed exacerbated alveolar simplification in Foxf2-deficient mice under injury conditions (Fig. 4h, i). Given the high mortality rate following injury, we investigated whether barrier function was impaired in Foxf2 cKO lungs. A vascular leakage assay using FITC-conjugated dextran was conducted to evaluate blood vessel permeability in Foxf2 cKO lungs compared to controls under hyperoxia treatment. The results showed increased dextran leakage into the pulmonary parenchyma under hyperoxia, indicating that Foxf2 deficiency compromises vascular integrity in the alveoli after hyperoxic injury (Fig.4j, k).
Pericytes accumulated in the alveoli of hyperoxic Foxf2 cKO lungs, similar to what is seen under normal conditions. (Fig. 5a, b). Endomucin coverage was reduced in Foxf2-deficient lungs (Fig. 5c), indicating a potential decrease in alveolar endothelial cells. Consistent with impaired angiogenesis, Foxf2 cKO lungs exhibited lower Pecam1 expression (Supplementary Fig. 5a). Hif1a mRNA was upregulated in Foxf2 cKO lungs, and immunostaining confirmed elevated HIF1α expression, aligning with reduced oxygen saturation (Supplementary Fig. 5b–d). FACS analysis verified a decrease in endothelial cell number and an increase in pericytes in Foxf2 cKO lungs under neonatal hyperoxia treatment (Fig. 5d–f). Co-staining for Ki-67 and PDGFRβ showed increased pericyte proliferation and reduced endothelial cell proliferation in Foxf2 cKO mice during hyperoxic injury (Fig. 5g–i). In line with the observed changes in pericyte and endothelial cell populations, FACS analysis of the cell cycle showed an approximately two-fold increase in the percentage of CD140b+ cells in the S phase in hyperoxic Foxf2 cKO mice. (Supplementary Fig. 6a, b). Conversely, CD31+ cells exhibited a reduction in S-phase cells and an increase in cells in the G0/G1 phase (Supplementary Fig. 6c, d). Although capillary endothelial cell numbers were reduced during injury, the proportions of gCap and aCap cells remained unchanged in both normal and hyperoxic conditions, indicating that endothelial cell differentiation was largely unaffected upon pericyte loss of Foxf2 (Supplementary Fig. 7a-d). These findings indicate that FOXF2 expression in pericytes is essential for pulmonary angiogenesis following perinatal hyperoxic injury.
a Representative images for endomucin and PDGFRβ immunostaining on lung sections from hyperoxia-treated mice. Scale bar=100 μm. Arrows indicate pericytes. b Quantification of the percentage of PDGFRβ-positive cells (pericytes). n = 7 for Ctrl and 9 for Foxf2 cKO with hyperoxia. p = 0.0009. c Quantification of the percentage of endomucin area. n = 8 for Ctrl and 8 for Foxf2 cKO. p = 0.0288. d Representative FACS panel showing CD140b positive cells (pericytes) and CD31 positive cells (endothelial cells) from mice at P7 under hyperoxia. e Quantification of the percentage of CD31-positive cells. n = 5 for Ctrl and 6 for Foxf2 cKO with hyperoxia. p = 0.0483. f Quantification of the percentage of CD140b-positive cells. n = 5 for Ctrl and 6 for Foxf2 cKO. p = 0.0088. g Representative images for PDGFRβ, ERG, and Ki-67 staining in hyperoxia-treated lungs at P7. Scale bar=50 μm. White arrows indicate PDGFRβ+ pericytes. Yellow arrows indicate ERG+ endothelial cells. h Quantification of the percentage of Ki-67 positive pericytes. n = 6 for Ctrl and 6 for Foxf2 cKO with hyperoxia. p = 0.0003. i Quantification of the percentage of Ki-67 positive endothelial cells. n = 4 for Ctrl and 4 for Foxf2 cKO with hyperoxia. p = 0.0054. All p values were determined by an unpaired 2-tailed t test. Data are presented as mean values ± SEM. Source data are provided as a Source Data file.
FOXF2-dependent pericyte gene expression is essential for pulmonary angiogenesis
Paracrine factors from pericytes are known to be essential for pulmonary angiogenesis and homeostasis. To investigate changes in signaling interactions between pericytes and endothelial cells following Foxf2 deletion, we performed CellChat analysis on scRNA-seq data26. The analysis of overall signaling flow revealed that pro-angiogenic pathways, such as ANGPTL and EDN27, were enriched in control lungs, whereas pathways associated with stress and inflammation, including TRAIL and GDF, were elevated in Foxf2 cKO lungs (Fig. 6a)28,29. By analyzing communication probabilities mediated by ligand-receptor pairs from pericytes to capillary endothelial cells, we showed that Foxf2 cKO lungs exhibit increased TGFβ signaling and reduced paracrine signaling interactions associated with angiogenesis, including KITL-KIT, ANGPT1-TEK, and multiple ANGPTL4-related pairs, with ANGPTL4 showing the most pronounced decrease (Fig. 6b, c). To investigate whether ANGPTL4-mediated signaling was reduced in Foxf2 cKO lungs, we utilized a proximity ligation assay (PLA) to detect ligand-receptor interactions. As a positive control, we used PECAM1 and VE-Cadherin, known interactors that form a mechanosensory complex in endothelial cells30, to validate PLA signals. Consistent with previous study showing that ANGPTL4 interacts with integrins to promote cell migration31, we detected the interaction between ANGPTL4 and integrin β1 and showed that this interaction is decreased in Foxf2 cKO lungs, as evidenced by a reduced proportion of endothelial cells with PLA signals (Supplementary Fig. 8a-c).
a CellChat analysis of changes in overall signaling flow in Foxf2 cKO mice. b, c Dot plot showing up-regulated and down-regulated ligand–receptor pairs sending from pericytes to capillary endothelial cells. d Representative images for endothelial tube formation assay on HPMEC cells cultured with conditioned medium from Ctrl or Foxf2 cKO mice treated with PBS or rhANGPTL4. Scale bar= 200 μm. e Quantification of junction number per field of view. n = 5 for Ctrl + PBS, 4 for Ctrl +rhANGPTL4, 9 for Foxf2 cKO + PBS and 10 for Foxf2 cKO+ rhANGPTL4. p = 0.0349 for Ctrl +rhANGPTL4 vs. Ctrl + PBS, 0.0002 for Foxf2 cKO + PBS vs. Ctrl + PBS, and 0.0017 for Foxf2 cKO +rhANGPTL4 vs. Foxf2 cKO + PBS. f Heatmaps for CUT&Tag peaks for FOXF2 binding sites and chromatin accessibility detected by CUTAC-H3K4me2, peaks were all centered to FOXF2 CUT&Tag peaks. Foxf2 cKO mice exhibited a broad decrease in chromatin accessibility at FOXF2 binding sites. g DiffTF analysis of RNA-seq and CUTAC-H3K4me2 data showing increased and decreased transcription factor activity with classification of transcription factor into repressors or activators. h Motif analysis in FOXF2 binding genomic regions. i Tracks for FOXF2 binding and chromatin accessibility measured by CUTAC-H3K4me2 at the genomic locus of Angptl4. Red boxes indicate regions bound by FOXF2, with decreased accessibility. p values in (e) were determined by one-way ANOVA followed by Sidak’s multiple comparison test. Data are presented as mean values ± SEM. Source data are provided as a Source Data file.
Recent studies have identified ANGPTL4 as a potent pro-angiogenic factor with a critical role in regulating vascular permeability22,23,32,33. To investigate whether FOXF2 is essential for pericyte-endothelial signaling during angiogenesis, we isolated and cultured CD140b+ pericytes from control and Foxf2 cKO mice and collected their conditioned media. Enzyme-linked immunosorbent assay (ELISA) confirmed a reduction in ANGPTL4 protein in media from Foxf2 cKO pericytes (Supplementary Fig. 9a). Human pulmonary microvascular endothelial cells (HPMECs) demonstrated a reduction in tube formation when incubated with conditioned media from Foxf2-deficient pericytes. The addition of recombinant ANGPTL4 enhanced HPMEC tube formation in control pericyte conditioned media, and partially restored the deficit observed in the Foxf2-deficient conditioned media. (Fig. 6d, e). These data support a FOXF2-ANGPTL4 axis in pericytes that regulates pulmonary angiogenesis.
To elucidate how FOXF2 regulates pericyte gene transcription, we performed Cleavage Under Targets and Tagmentation (CUT&Tag) to map FOXF2 binding and H3K4me2-mediated Cleavage Under Targeted Accessible Chromatin (CUTAC) to assess chromatin accessibility near H3K4me2 in MACS-sorted pericytes34. Plots of heatmaps of FOXF2 CUT&Tag and H3K4me2 CUTAC signals centered on FOXF2 peaks revealed that FOXF2 and H3K4me2 CUTAC signals co-occupy the nearby genomic regions, indicating an association between FOXF2 and open chromatin (Fig. 6f). In Foxf2-deficient pericytes, overall CUTAC signal was reduced, with loss of FOXF2 leading to decreased chromatin accessibility in over half (9085 of 16,197) of FOXF2-binding regions (Fig. 6f; Supplementary Fig. 9b). Motif analysis revealed that less accessible sites were enriched for motifs including FOXF1/2, SP3, ELK4, CTCF, and TEAD4, suggesting that FOXF2 may collaborate with these transcription factors to regulate pericyte gene expression (Supplementary Fig. 9c). Integrated analysis of transcriptomic and CUTAC data using diffTF revealed that Foxf2-deficient pericytes exhibit reduced transcription factor activity in the ETV/ELF and KLF families, which share similar binding motifs with ELK and SP, respectively (Fig. 6g). These transcription factors were predicted to function as either repressors or activators (Fig. 6g). Conversely, increased activity of transcription factors such as FOXM1, SMAD1/3, and CTCF was observed, consistent with enhanced proliferation programs (Fig. 6g). In line with the above data, motif analysis of FOXF2-binding sites revealed enrichment for ELKs, CTCF, and TEAD4 binding motifs, in addition to FOXF motifs (Fig. 6h).
Genomic Regions Enrichment of Annotations Tool (GREAT) analysis showed that regions with reduced CUTAC signals were associated with genes involved in cell division, cell migration, and vascular development (Supplementary Fig. 9d). Decreased chromatin accessibility was observed near FOXF2-binding peaks in promoter and enhancer regions of gene loci, such as Angptl4, Angpt1, Hgf, and Wwtr1 (Fig. 6i, Supplementary Fig. 9e-g). Collectively, these findings indicate that FOXF2 acts as a co-regulator with other transcription factors, binding cooperatively across the genome to modulate chromatin organization and define pericyte properties critical for pericyte-endothelial signaling, which is essential for neonatal pulmonary angiogenesis and lung repair after neonatal lung injury.
Discussion
Pericytes are not only integral components of the vasculature but also coordinate with other cell types to regulate organ development and regeneration. Dysfunction of pericytes disrupts tissue homeostasis, contributing to various pulmonary diseases. Our study demonstrates that FOXF2 is essential for proper pericyte maturation and function, maintaining their cellular properties to support pulmonary angiogenesis and vascular integrity following postnatal lung injury. FOXF2 binds to and modulates chromatin accessibility of proangiogenic genes, such as Angptl4, promoting pericyte-endothelium crosstalk essential for angiogenesis and barrier function in postnatal lung injury.
Postnatal alveolar formation, which results in a 16-fold increase in surface area, is accompanied by a significant expansion of the pulmonary capillary network adjacent to the alveolar surface35. Impaired angiogenesis is implicated in the pathogenesis of alveolar capillary dysplasia with misalignment of pulmonary veins (ACDMPV) and bronchopulmonary dysplasia (BPD)36,37,38. Although studies in other organs demonstrate that newly formed vessels recruit pericytes to stabilize vasculature and promote angiogenesis, the molecular mechanisms governing pericyte-capillary interactions in the lung remain poorly understood39. In Foxf2-deficient pericytes, we observed increased proliferation coupled with reduced expression of genes characteristic of mature pericytes, including Angptl4, Lama4, Itga8, and Trpc6. These genes regulate critical pericyte functions, such as angiogenesis, endothelial permeability, cell adhesion, and vascular tone modulation. This altered gene expression profile suggests a less mature pericyte phenotype. Consistent with prior studies, elevated proliferation in Foxf2 cKO mice was associated with enhanced PI3K/AKT signaling20. While enhanced PI3K activity promotes hyperplasia and suppresses endothelial outgrowth in the retina20, Foxf2 cKO lungs displayed reduced endothelial cell numbers and diminished vascular coverage after hyperoxic injury, indicating organ-specific mechanisms underlying angiogenic regulation.
Since defects in pulmonary angiogenesis often manifest as chronic conditions, we propose that additional aspects of vascular biology may contribute to the reduced survival of Foxf2 cKO mice during injury. Consistent with this notion, our data indicate a loss of endothelial barrier integrity in hyperoxic Foxf2 cKO mice. In line with previous studies showing that increased vascular permeability disrupts gas exchange and decreases oxygenation40, the elevated permeability observed in Foxf2 cKO lungs likely contributes to hypoxemia, which may underlie the poor survival of these animals. Given the systemic deletion of Foxf2 in Pdgfrb+ cells, it is also possible that defects in other organs contribute to this survival phenotype. For instance, loss of Foxf2 in brain pericytes has been shown to cause hemorrhage17. Defining the expression patterns and functional roles of Foxf2 in pericytes across different organs will provide further mechanistic insight into its role in vascular homeostasis and survival.
Our findings highlight a pivotal paradigm, demonstrating that competent pericytes are crucial for sustaining a vascular niche required for homeostasis and regeneration after perinatal lung injury. We further identify ANGPTL4 as one of the key niche factors mediating endothelial angiogenesis. Although we identified decreased ANGPTL4-integrin interaction by PLA assay, the functional significance of this interaction still warrants future investigation given various receptors for ANGPTL4 have been reported. Moreover, whether the observed impairment in barrier function results directly from the loss of ANGPTL4 remains unclear, as ANGPTL4 has been shown to exert context-dependent and sometimes opposing effects on endothelial permeability23,33. Nevertheless, ANGPTL4 likely represents only one component of a broader network of regulatory signals within the pericyte–endothelial niche. Other signaling molecules may also contribute to maintaining vascular integrity and modulating pulmonary angiogenesis as demonstrated by the partial restoration of HPMEC tube formation from recombinant ANGPTL4 in Foxf2-deficient pericyte conditioned media. Understanding how these pathways collectively orchestrate cell-cell interactions may shed light on the complex pathophysiology of BPD and identify new therapeutic targets.
While our findings elucidate the role of pericyte-derived angiogenic signaling in hyperoxic lung injury, specifically ANGPTL4, ectopic accumulation of pericytes in lung parenchyma may also impair functions of endothelial or other cell types. In healthy lungs, pericytes are sparsely distributed in pulmonary capillaries, whereas aerocytes (aCaps) cover most of the gas exchange surface. An increased presence of dysfunctional pericytes may enhance extracellular matrix deposition or signaling factor secretion, thereby disrupting the intrinsic properties of neighboring cells and compromising gas exchange and alveolar architecture during hyperoxic injury. Supporting this hypothesis, CellChat analysis revealed elevated GDF and TGFβ signaling from pericytes to endothelial cells, alongside heightened stress-related pathways in global signaling flow in Foxf2-deficient mice. Similarly, increased numbers and misallocation of pericytes have been associated with endothelial dysfunction in pathological conditions such as pulmonary arterial hypertension, underscoring the broader implications of pericyte dysregulation in lung diseases41,42.
Our work integrates FOXF2 into the known transcriptional network that regulates pericyte properties. FOXF2 binding sites are enriched with motifs of ELK, SP and TEAD, etc., suggesting that FOXF2 functions in a cooperative manner with other transcription factors. In alignment with this notion, deletion of Hippo pathway component YAP/TAZ, two upstream regulators of TEAD, in pericytes causes loss of Angpt1 and Hgf expression, which are required for endothelial and alveolar type 2 (AT2) cell proliferation in pulmonary alveologenesis10. While Foxf2-deficient mice exhibited normal alveolar structure, our transcriptomic and genomic analyses revealed alterations in a comparable gene expression program. Notably, FOXF2 binding was identified at the genomic loci of Wwtr1, Angpt1, and Hgf, where chromatin accessibility was markedly reduced following Foxf2 deletion. Despite stable mRNA levels of key genes such as Angpt1, diminished chromatin accessibility may impair the activation of proangiogenic genes during regeneration, potentially increasing susceptibility to hyperoxia-induced injury. Enrichment of the FOXF1 motif within FOXF2 binding sites suggests partial redundancy in their regulation of gene expression in pericytes. Further genetic studies are needed to elucidate their overlapping and distinct roles in pericyte function under physiological and pathological conditions.
In conclusion, our study establishes FOXF2 as a pivotal regulator of pericyte function in the postnatal lung. By orchestrating paracrine signaling, FOXF2 empowers pericytes to promote endothelial repair and preserve alveolar integrity during pathological challenges. These insights identify molecular targets for therapeutic strategies aimed at mitigating vascular injury and impaired angiogenesis in neonatal lung disorders such as BPD.
Methods
Animals
All animal experiments were performed in compliance with the Institutional Animal Care and Use Committee (IACUC) protocol approved by the West China Second University Hospital, Sichuan University (2021-054) and Cincinnati Children’s Hospital (2019-0017). Mice were bred on a mixed background and housed with food and water ad libitum on a 14/10 (CCHMC) and a 12/12 (WCSUH) hour light/dark cycle with humidity levels maintained between 40 and 60%. Pdgfrb-CreER mice were obtained from The Jackson Laboratory (Strain No.029684). The Foxf2fl/fl mouse line was generated as previously described43. Foxf2fl/fl mice were crossed with Pdgfrb-CreER mice to produce Foxf2 cKO offspring. Mice were maintained on a C57BL6 genetic background. To induce pericyte-specific Foxf2 deletion, tamoxifen (10 mg/ml, 10 μl per pup; Sigma) was administered via intraperitoneal injection at postnatal day 0 (P0) and P1. Both male and female mice were included in all experiments, with littermates serving as controls.
Postnatal hyperoxia injury was induced by exposing newborn mice to 75% O2 from P0 to P7. To ensure the well-being of breeding females during hyperoxic exposure, they were returned to normoxic conditions every 48 h and substituted with alternate lactating females. Wild-type C57BL/6 mice, used for primary lung pericyte isolation, were sourced from GemPharmatech (Nanjing, China). Arterial oxygenation was measured using the MouseOx Plus system (STARR Life Sciences) according to the manufacturer’s instructions. To assess pulmonary vascular permeability, 50 μL of 50 mg/mL FITC-dextran (70 kDa, F491425, Aladdin Scientific) in PBS was injected via the facial vein 0.5 h prior to euthanizing the animals. The lungs were subsequently embedded in OCT and cryosectioned at 10 μm thickness.
Histology and Immunostaining
Lungs were harvested, perfused with phosphate-buffered saline (PBS) via the right ventricle, and inflated with 2% paraformaldehyde (PFA) at 25 cm H2O pressure. Tissues were immersion-fixed in 2% PFA at 4 °C overnight, washed in PBS, and dehydrated through a graded ethanol series (30%, 50%, 70%, 90%, 100%). Lungs were cleared in xylene, embedded in paraffin, and sectioned at 5 μm thickness. Hematoxylin and eosin (H&E) staining was performed to evaluate lung morphology. Brightfield images were acquired using a Zeiss Axioplan2 or Olympus APX-100 microscope with a 20× objective. MLI was quantified using ImageJ, as previously described44. H&E-stained sections were analyzed by overlaying a 10 × 10 grid on images, counting intersections between alveolar walls and grid lines, excluding those crossing major blood vessels or airways. MLI was calculated as: MLI = total line length/total number of intercepts.
For immunostaining, paraffin-embedded lung sections were blocked in PBS containing 5% donkey serum and incubated with primary antibodies (diluted in PBS) in a humidified chamber at 4 °C overnight (see Supplementary Table 1 for antibody details). Sections were then incubated with secondary antibodies conjugated to Alexa Fluor 488, 594, or 647 (Jackson ImmunoResearch), and mounted using ProLong Gold Antifade Mountant (Thermo Fisher, P36934). For antigens requiring tyramide signal amplification (TSA), biotin-conjugated secondary antibodies were applied, followed by incubation with biotinylated tyramide (Aladdin Scientific, cat. no. B288371) and subsequent labeling with streptavidin-fluorophore conjugates. Fluorescence images were acquired using a Nikon A1R or Olympus FV3000 confocal microscope. The intensity of phosphorylated AKT (p-AKT) in PDGFRβ+ cells was quantified automatically using CellProfiler. PDGFRβ+ cells were identified with the “RunCellpose” module (cyto2 model) in CellProfiler45,46, and the mean fluorescence intensity of p-AKT within PDGFRβ-marked regions was calculated. Endomucin-covered areas were quantified in FIJI using the default automatic thresholding method.
FACS and MACS
FACS analysis was performed as previously described47. Lung lobes were minced and enzymatically digested with 5 µg/mL Dispase (Corning, 354235), 0.025 mg/mL DNase I (Roche, 10104159001), and 480 µg/mL Collagenase Type I (Thermo Fisher, 17100017) in PBS at 37 °C for 30 min with intermittent agitation. A gentleMACS Dissociator (Miltenyi Biotec) was used to generate single-cell suspensions. Digested tissues were washed with FACS buffer (PBS containing 1 mM EDTA, 25 mM HEPES, and 0.5% BSA), filtered through a 70-µm cell strainer, and incubated with 1× RBC Lysis Buffer (BioLegend, 420301) for 5 min on ice to remove erythrocytes. Samples were blocked with anti-mouse CD16/32 (FcR) antibody (BioLegend, 101301) at 4 °C for 1 h, followed by staining with FACS antibodies indicated in the figures. Dead cells were identified by DAPI staining (1:10,000). Data were acquired using an LSRII flow cytometer (BD Biosciences). For EdU incorporation assay, EdU (Aladdin Scientific, 61135-33-9) was injected at 50 μg/g body weight 2 h before euthanization. Single cell suspension was prepared as described above and blocked with anti-mouse CD16/32 (FcR) antibody. Cells were then stained with FACS antibodies indicated in the figures. Stained cells were then fixed with 1% PFA in PBS for 15 min and permeabilized with PBS with 0.2% tween 20 for 10 min. Alexa Fluor 488- conjugated Azide was applied to cells with a click-reaction using aladdin®488 Click-iT EdU kit (Aladdin Scientific, A598377) according to specifications provided by manufacturer. Cells were counter stained with DAPI and analyzed on a CytoFlex SRT system (Beckman). All detailed information for FACS antibodies was listed in Supplementary Table 1. For PDGFRβ+ pericyte isolation, FcR-blocked single-cell suspensions were incubated with biotin-conjugated anti-CD140b (PDGFRβ) antibody at 4 °C for 1 h, followed by anti-biotin microbeads (Miltenyi Biotec, 130-090-485) for 30 minutes. Bead-bound PDGFRβ+ cells were enriched using LS columns (Miltenyi Biotec) and washed with FACS buffer. CD45− cells for scRNA-seq were isolated by magnetic-activated cell sorting (MACS). Single-cell suspensions were incubated with biotin-conjugated anti-mouse CD45 antibody, followed by anti-biotin microbeads (Miltenyi Biotec). The flow-through fraction, containing CD45−cells, was collected for downstream analysis.
Quantitative PCR
Total RNA was extracted from whole lung tissue and isolated pericytes using the RNeasy Micro Kit (Qiagen). Complementary DNA (cDNA) was synthesized with iScript Reverse Transcription Supermix (Bio-Rad, 1708840). Quantitative PCR (qPCR) was performed on a StepOnePlus (Applied Biosystems) or LightCycler 96 (Roche) system. Gene-specific primers used for amplification are listed in Supplementary Table 2. Actb served as the internal reference gene, and relative gene expression was quantified using the 2−ΔΔCt method.
Bulk RNA-seq
RNA-seq libraries were prepared using the NEBNext Ultra RNA Library Prep Kit (New England Biolabs) and sequenced on an Illumina HiSeq 4000 platform. Raw FASTQ files were aligned to the mouse mm10 genome using HISAT248. Read counts were generated with the Subread package49. Differential expression analysis was performed using DESeq2, with genes exhibiting an adjusted P-value < 0.05 and |log2 fold change| > 0.58 considered differentially expressed50. Functional enrichment analysis was conducted using ToppGene (https://toppgene.cchmc.org/enrichment.jsp)51.
scRNA-seq
scRNA-seq libraries were prepared from CD45− cells isolated from control and Foxf2 cKO lungs using the Chromium Next GEM Single Cell 5’ Reagent Kit v2 (10x Genomics). The scRNA-seq libraries were constructed and sequenced on a Novaseq 6000 platform by Annoroad Gene Technology (Beijing) Co., Ltd. Raw FASTQ files were aligned to the mouse mm10 reference genome using CellRanger (10x Genomics). Downstream analysis was performed with Seurat v552. Cells meeting the following criteria were retained: 1,000 <nFeature_RNA < 8000 and percent mitochondrial gene expression <15%. Data integration was conducted using the reciprocal principal component analysis (RPCA) method, followed by clustering at a resolution of 0.4. Cell types were annotated based on markers from LungMAP53. Ligand-receptor interactions were analyzed using the CellChat package, focusing on “secreted signaling” pathways with default parameters26. For human scRNAseq analysis, two term infants and four BPD infants were included in the analysis24. Visualizations were generated using the scCustomize package (https://samuel-marsh.github.io/scCustomize/index.html).
CUT&Tag and CUTAC
CUT&Tag and CUTAC experiments were performed following established protocols54. For FOXF2 CUT&Tag and H3K4me2 CUTAC, approximately 30,000 cells per reaction were used. Briefly, cells were incubated with concanavalin A-coated magnetic beads (Vazyme, N515) for 15 min at room temperature. Primary antibodies against FOXF2 or H3K4me2 were applied, and samples were rotated overnight at 4 °C. Secondary antibodies were added and incubated for 30 min at room temperature. Unbound antibodies were removed using Triton wash buffer, followed by incubation with pG-Tn5 adapter complex (1:25 dilution) for 1 h at room temperature55.
For CUT&Tag, samples were resuspended in 50 µL tagmentation buffer (1 ml Triton-300 wash buffer, 10 µL 1 M MgCl2) and incubated at 37 °C for 1 h. For CUTAC, samples were resuspended in 50 µL CUTAC-DMF tagmentation buffer (10 mM TAPS, 5 mM MgCl2, 20% DMF, 0.05% Triton X-100) and incubated at 37 °C for 20 min. Post-tagmentation, DNA was extracted using SDS-Proteinase K solution and amplified via PCR with TruePrep CUT&Tag Amplification Mix (Vazyme, TD612) using Illumina i5 and i7 indices. Amplified libraries were purified with VAHTS DNA Clean Beads (Vazyme, N411) and sequenced on an illumina NovaSeq 6000 platform by Annoroad Gene Technology (Beijing) Co., Ltd.
FASTQ files were trimmed using TrimGalore and aligned to the mouse mm10 genome with Bowtie2 using the parameters: --local --very-sensitive --no-mixed --no-discordant --phred33 -I 10 -X 100056. Peaks were called using MACS2. Differentially accessible peaks were identified with DiffBind57. Peak heatmaps were generated using deepTools58. Differential transcription factor activity and classification of repressors and activators were determined using diffTF, integrating RNA-seq and CUTAC data59. Motif analysis was performed with the monaLisa package using the JASPAR 2020 database60. Functional annotation of peaks was carried out using GREAT analysis61.
Cell culture
Primary PDGFRβ+ pericytes were cultured in DMEM GlutaMAX supplemented with 10% fetal calf serum (Gibco, Life Technologies). Primary human pulmonary microvascular endothelial cells (HPMEC, Meisen CTCC, CTCC-HUM-0027) were cultured in EGM-2 Endothelial Cell Growth Medium-2 BulletKit (Lonza, CC-3162). Cultures were maintained at 37 °C in a humidified incubator with 5% CO2, with media refreshed every 48 h. No cell authentication was conducted for the primary HPMEC. For the scratch wound healing assay, 50,000 pericytes, isolated from control and Foxf2 cKO lungs, were seeded in 24-well plates and cultured to confluence. A linear scratch was created using a pipette tip, and detached cells were removed by washing with medium. Images were captured at 0 and 24 h post-scratch using an Olympus APX-100 microscope. Wound closure was quantified as: Wound recovery (%) = (D0 − Dn) / D0 × 100, where D0 is the initial wound distance and Dn is the wound distance at 24 h.
To assess pericyte proliferation upon siRNA-mediated Foxf2 knockdown, 5000 pericytes were seeded per well in 96-well plates, transfected with 50 nM si-NC or si-Foxf2 (5’-GCGTGTGCCAAGATATTAA-3’) (RiboBio) using Lipofectamine RNAiMAX (Life Technologies, 13778030), and cultured for 48 h. BrdU (Beyotime, ST1056) was added to the medium at 10 μM, and cells were incubated for 2 h. Cells were then fixed, permeabilized with 0.2% Triton X-100 on ice, and blocked with PBS containing 5% donkey serum for 1 h at room temperature. Samples were incubated with primary antibodies against BrdU and PDGFRβ at 4 °C overnight, followed by secondary antibodies and DAPI for nuclear staining. BrdU+ cells were quantified using ImageJ, and the proliferation rate was calculated as the percentage of BrdU+ cells relative to total nuclei. All images were acquired with an Olympus APX-100 microscope.
Endothelial tube formation assay
To assess the effect of Foxf2 in pericytes on endothelial cell (EC) tube formation, 50,000 pericytes, isolated from control and Foxf2 cKO lungs, were cultured in 24-well plates for 72 h, and the conditioned medium from day 1 to day 3 of culture was collected. Subsequently, 10,000 HPMECs cells were seeded onto solidified matrigel (Corning, 356231) and supplemented with the conditioned medium in 96-well plates. Recombinant human ANGPTL4 (25 ng/ml, MedChemExpress, HY-P7507) or an equivalent volume of PBS (vehicle control) was added. After 6 h of culture, cells were incubated with Calcein AM (Beyotime, C2012) for 20 min, and vascular network formation was imaged using an Olympus APX-100 microscope with a 4× objective. The number of junctions was counted in ImageJ software.
Quantification of ANGPTL4 by ELISA
ANGPTL4 concentration was measured using a commercial ELISA kit (Novus Biologicals, NBP3-06850) according to the manufacturer’s instructions. Briefly, 100 μl of conditioned medium was added to each antibody-precoated well and incubated for 2 h at room temperature. After several washing steps, colorimetric detection was performed using an HRP-based substrate for 30 min at room temperature, followed by the addition of stop solution. Absorbance was measured using a BioTek Synergy plate reader, and ANGPTL4 concentrations were calculated based on a standard curve generated in each assay.
Proximity ligation assay (PLA)
Protein–protein interaction between ANGPTL4 and Integrin β1 was assessed using the Duolink PLA system. Paraffin-embedded tissue sections were dewaxed and subjected to antigen retrieval in Tris/EDTA buffer (pH 9.0) using microwave heating. Sections were then incubated overnight at 4 °C with rabbit anti-ANGPTL4 (Thermo Fisher, 40-9800, 1:100) and goat anti-Integrin β1 (R&D Systems, AF2405, 1:200) antibodies diluted in the antibody diluent provided with the kit. After primary antibody incubation, sections were incubated at 37 °C for 1 h with Duolink In Situ PLA Probe anti-Goat MINUS (Sigma, DUO92006) and Duolink In Situ PLA Probe anti-Rabbit PLUS (Sigma, DUO92002). Following washes with Duolink In Situ Wash Buffers (Sigma, DUO82049), ligation and rolling-circle amplification (RCA) were performed using reagents from the Duolink In Situ Detection Reagents Green kit (Sigma, DUO92014). After completion of the PLA procedure, sections were counterstained with additional antibodies as indicated. Known interactions between VE-Cadherin (R&D Systems, AF1002, 1:200) and PECAM1 (CST, 77699S, 1:200) were used as positive controls.
Western blot
A standard Western blot procedure was performed. Briefly, 20 μg of total protein extracted in RIPA lysis buffer was loaded into each well, separated on a 12% SDS–PAGE gel, and transferred onto a PVDF membrane. The membranes were blocked with 5% milk and incubated with primary antibodies in PBS overnight at 4 °C. After washing, the membranes were incubated with Biotin-conjugated secondary antibody followed by 30 min incubation in Strepavidin-HRP reagent (ApexBio, K1229, 1:2000) and developed using an enhanced chemiluminescence (ECL) substrate, and signals were detected with a G:Box digital imaging system (Syngene). Band intensities were quantified using ImageJ software. Antibody details for western blot was provided in Supplementary Table 1.
Statistical analysis
Data are presented as mean ± standard error mean (SEM). Statistical analyses and graph plotting were performed using GraphPad Prism 8.0. Statistical significance was determined using an unpaired two-tailed Student’s t test or one-way ANOVA followed by Sidak’s multiple comparison test, with p < 0.05 considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA sequencing, single-cell RNA sequencing, CUT&Tag and CUTAC data have been deposited in the NCBI Gene Expression Omnibus with accession numbers GSE297573, GSE297575 and GSE297570. Human BPD scRNA-seq data were from GSE275938. Source data are provided with this paper.
References
Whitsett, J. A., Kalin, T. V., Xu, Y. & Kalinichenko, V. V. Building and regenerating the lung cell by cell. Physiol. Rev. 99, 513–554 (2019).
Vila Ellis, L. & Chen, J. A cell-centric view of lung alveologenesis. Dev. Dyn. 250, 482–496 (2021).
Morrisey, E. E. & Hogan, B. L. M. Preparing for the first breath: genetic and cellular mechanisms in lung development. Dev. Cell 18, 8–23 (2010).
Jobe, A. H. Mechanisms of lung injury and bronchopulmonary dysplasia. Am. J. Perinatol. 33, 1076–1078 (2016).
Thébaud, B. et al. Bronchopulmonary dysplasia. Nat. Rev. Dis. Prim. 5, 78 (2019).
Ren, X. et al. Postnatal alveologenesis depends on FOXF1 signaling in c-KIT1 endothelial progenitor cells. Am. J. Respir. Crit. Care Med. 200, 1164–1176 (2019).
Bolte, C. et al. Nanoparticle delivery of proangiogenic transcription factors into the neonatal circulation inhibits alveolar simplification caused by hyperoxia. Am. J. Respir. Crit. Care Med. 202, 100–111 (2020).
Vila Ellis, L. et al. Epithelial Vegfa specifies a distinct endothelial population in the mouse lung. Dev. Cell 52, 617–630.e6 (2020).
Gillich, A. et al. Capillary cell-type specialization in the alveolus. Nature 586, 785–789 (2020).
Kato, K. et al. Pulmonary pericytes regulate lung morphogenesis. Nat. Commun. 9, 2448 (2018).
Bergers, G. & Song, S. The role of pericytes in blood-vessel formation and maintenance. Neuro-Oncology 7, 452–464 (2005).
Sweeney, M. D., Ayyadurai, S. & Zlokovic, B. V. Pericytes of the neurovascular unit: Key functions and signaling pathways. Nat. Neurosci. 19, 771–783 (2016).
Xu, J. et al. A Shh-Foxf-Fgf18-Shh molecular circuit regulating palate development. PLoS Genet. 12, e1005769 (2016).
Nik, A. M., Johansson, J. A., Ghiami, M., Reyahi, A. & Carlsson, P. Foxf2 is required for secondary palate development and Tgfβ signaling in palatal shelf mesenchyme. Dev. Biol. 415, 14–23 (2016).
Van Den Brink, G. R. & Rubin, D. C. Foxf2: a mesenchymal regulator of intestinal adenoma development. Gastroenterology 144, 873–876 (2013).
Graff, M. F. E., Heeg, E. E. M. & Childs, S. J. Progressive mural cell deficiencies across the lifespan in a foxf2 model of cerebral small vessel disease. Elife 14, RP106720 (2025).
Reyahi, A. et al. Foxf2 is required for brain pericyte differentiation and development and maintenance of the blood-brain barrier. Dev. Cell 34, 19–32 (2015).
Fan, Y., Li, L. & Sun, S. Powerful and accurate detection of temporal gene expression patterns from multi-sample multi-stage single-cell transcriptomics data with TDEseq. Genome Biol. 25, 96 (2024).
Zepp, J. A. et al. Genomic, epigenomic, and biophysical cues controlling the emergence of the lung alveolus. Science. 371, eabc3172 (2021).
Figueiredo, A. M. et al. Phosphoinositide 3-kinase-regulated pericyte maturation governs vascular remodeling. Circulation 142, 688–704 (2020).
Klouda, T. et al. Specialized pericyte subtypes in the pulmonary capillaries. EMBO J. https://doi.org/10.1038/s44318-024-00349-1 (2025).
Hung, C. F. et al. Pericyte-like cells undergo transcriptional reprogramming and distinct functional adaptations in acute lung injury. FASEB J. 35, e21323 (2021).
Chaube, B. et al. Suppression of angiopoietin-like 4 reprograms endothelial cell metabolism and inhibits angiogenesis. Nat. Commun. 14, 8251 (2023).
Shirazi, S. P. et al. Bronchopulmonary dysplasia with pulmonary hypertension associates with semaphorin signaling loss and functionally decreased FOXF1 expression. Nat. Commun. 16, 1–14 (2025).
Hurskainen, M. et al. Single cell transcriptomic analysis of murine lung development on hyperoxia-induced damage. Nat. Commun. 12, 1–19 (2021).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Salani, D. et al. Endothelin-1 induces an angiogenic phenotype in cultured endothelial cells and stimulates neovascularization in vivo. Am. J. Pathol. 157, 1703–1711 (2000).
Schwarz, A., Kinscherf, R. & Bonaterra, G. A. Role of the stress- and inflammation-induced cytokine GDF-15 in cardiovascular diseases: from basic research to clinical relevance. Rev. Cardiovasc. Med. 24, 81 (2023).
Sullivan, G. P. et al. TRAIL receptors serve as stress-associated molecular patterns to promote ER-stress-induced inflammation. Dev. Cell 52, 714–730.e5 (2020).
Woodfin, A., Voisin, M. B. & Nourshargh, S. PECAM-1: a multi-functional molecule in inflammation and vascular biology. Arterioscler. Thromb. Vasc. Biol. 27, 2514–2523 (2007).
Goh, Y. Y. et al. Angiopoietin-like 4 interacts with integrins β1 and β5 to modulate keratinocyte migration. Am. J. Pathol. 177, 2791 (2010).
Babapoor-Farrokhran, S. et al. Angiopoietin-like 4 is a potent angiogenic factor and a novel therapeutic target for patients with proliferative diabetic retinopathy. Proc. Natl. Acad. Sci. USA 112, E3030–E3039 (2015).
Galaup, A. et al. Angiopoietin-like 4 prevents metastasis through inhibition of vascular permeability and tumor cell motility and invasiveness. Proc. Natl. Acad. Sci. 103, 18721–18726 (2006).
Henikoff, S., Henikoff, J. G., Kaya-Okur, H. S. & Ahmad, K. Efficient chromatin accessibility mapping in situ by nucleosome-tethered tagmentation. Elife 9, e63274 (2020).
Mund, S. I., Stampanoni, M. & Schittny, J. C. Developmental alveolarization of the mouse lung. Dev. Dyn. 237, 2108–2116 (2008).
Sun, F. et al. Nanoparticle delivery of STAT3 alleviates pulmonary hypertension in a mouse model of alveolar capillary dysplasia. Circulation 144, 539–555 (2021).
Pradhan, A. et al. The S52F FOXF1 mutation inhibits STAT3 signaling and causes alveolar capillary dysplasia. Am. J. Respir. Crit. Care Med. 200, 1045–1056 (2019).
Baker, C. D. & Abman, S. H. Impaired pulmonary vascular development in bronchopulmonary dysplasia. Neonatology vol. 107, 344–351 (2015).
Ribatti, D., Nico, B. & Crivellato, E. The role of pericytes in angiogenesis. Int. J. Dev. Biol. 55, 261–268 (2011).
Xu, J. et al. Excess neuropeptides in lung signal through endothelial cells to impair gas exchange. Dev. Cell 57, 839–853.e6 (2022).
Ricard, N. et al. Increased pericyte coverage mediated by endothelial-derived fibroblast growth factor-2 and interleukin-6 is a source of smooth muscle-like cells in pulmonary hypertension. Circulation 129, 1586–1597 (2014).
Kim, H. et al. Pericytes contribute to pulmonary vascular remodeling via HIF2α signaling. EMBO Rep. 25, 616–645 (2024).
Bolte, C. et al. Forkhead box F2 regulation of platelet-derived growth factor and myocardin/serum response factor signaling is essential for intestinal development. J. Biol. Chem. 290, 7563–7575 (2015).
He, H. et al. Heparan sulfate regulates myofibroblast heterogeneity and function to mediate niche homeostasis during alveolar morphogenesis. Nat. Commun. 16, 1834 (2025).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinform. 22, 433 (2021).
Pachitariu, M. & Stringer, C. Cellpose 2.0: how to train your own model. Nat. Methods 19, 1634–1641 (2022).
Wang, G. et al. Endothelial progenitor cells stimulate neonatal lung angiogenesis through FOXF1-mediated activation of BMP9/ACVRL1 signaling. Nat. Commun. 13, 2080 (2022).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550(2014).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305-11 (2009).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Sun, X. et al. A census of the lung: cellcards from LungMAP. Dev. Cell 57, 112–145.e2 (2022).
Henikoff, S., Henikoff, J. & Ahmad, K. CUT&Tag-direct for whole cells with CUTAC v5. https://doi.org/10.17504/protocols.io.x54v9mkmzg3e/v5 (2023).
Soroczynski, J. et al. OpenTn5: open-source resource for robust and scalable Tn5 transposase purification and characterization. bioRxiv Preprint at https://doi.org/10.1101/2024.07.11.602973 (2024).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Ramírez, F. et al. deepTools2: a next-generation web server for deep-sequencing data analysis. Nucleic Acids Res 44, W160–W165 (2016).
Berest, I. et al. Quantification of differential transcription factor activity and multiomics-based classification into activators and repressors: diffTF. Cell Rep. 29, 3147–3159.e12 (2019).
Machlab, D. et al. monaLisa: an R/Bioconductor package for identifying regulatory motifs. Bioinformatics 38, 2624–2625 (2022).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Acknowledgements
This research was supported by grants from NSFC (82271748 to F.S., 82570003 and 82200098 to H.H.), the National Key R&D Program of China (2022YFA0806200) to F.S. and H.H., Science and Technology Projects of Xizang Autonomous Region, China (XZ202502ZY0008) to F.S. and H.H., and the research funds from the University of Arizona and Cincinnati Children’s Hospital (V.V.K).
Author information
Authors and Affiliations
Contributions
F.S., H.H. and V.V.K. conceived experiments and prepared the manuscript. F.S. and H.H. performed data analyses. F.S., H.H., Y.Z., P.C. and E.L. conducted the in vivo experiments, and H.H., Y.Z. and Y.L. conducted the in vitro experiments. H.H., S.P.S. and J.D. performed bioinformatic analyses. T.V.K., V.V.K., F.S., J.M.S.S. and H.H. provided critical insights. All authors discussed the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dritan Agalliu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, F., Zhao, Y., Do, J. et al. FOXF2 regulates pericyte–endothelial signaling required for vascular homeostasis after neonatal hyperoxic lung injury. Nat Commun 17, 2691 (2026). https://doi.org/10.1038/s41467-026-69525-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69525-7