Abstract
Metabolic reprogramming in immune cells can determine the outcome of pathogen infection. For Pseudomonas aeruginosa, a clinically challenging pathogen, it remains unclear whether the host can exploit this strategy to combat bacterial invasion. Here, we identify phosphoglycerate dehydrogenase as a key mediator of macrophage inflammation during Pseudomonas aeruginosa infection. Pharmacological and genetic inhibition of phosphoglycerate dehydrogenase suppress macrophage hyperactivation and the production of pro-inflammatory cytokines. In a murine model of Pseudomonas aeruginosa pneumonia, myeloid-specific deletion of phosphoglycerate dehydrogenase improves survival, alleviates lung injury, and reduces bacterial load. Similarly, dietary restriction of L-serine improves prognosis in infected mice. Mechanistically, phosphoglycerate dehydrogenase fuels L-serine synthesis to augment one-carbon metabolism, which strengthens the direct interaction between histone H3 lysine 27 trimethylation and dual-specificity phosphatase 4. This cascade ultimately promotes extracellular signal-regulated kinase 1/2 phosphorylation. Our study uncovers a metabolism-epigenetics crosstalk that amplifies macrophage inflammation, proposing metabolic modulation as a therapeutic strategy for bacterial pneumonia.
Similar content being viewed by others
Introduction
Pseudomonas aeruginosa (P. aeruginosa) is a well-known Gram-negative bacterium that can lead to troublesome infections in the lower respiratory tract. The pathogen-associated molecular patterns (PAMPs) of P. aeruginosa, such as lipopolysaccharides (LPS), outer membrane proteins, and proteases, are recognized by the pattern recognition receptors (PRRs) on host innate immune cells, triggering the activation of inflammatory signaling pathways1. While appropriate inflammatory responses are crucial for host defense against P. aeruginosa2,3,4, excessive inflammation can lead to pathological tissue damage, disease progression and even death5,6,7. Lung macrophages, particularly alveolar macrophages (AMs), are integral to the innate immunity response against P. aeruginosa8,9. In response to high pathogenicity and heavy load of P. aeruginosa infection, AMs recruit large numbers of peripheral monocyte-derived macrophages to lung tissue to facilitate phagocytosis and clearance of P. aeruginosa, triggering an overly intense inflammatory response. This intense inflammatory response can lead to transient exhaustion of AMs and dysfunction of recruited macrophages, impairing their ability to facilitate tissue repair10. Therefore, controlling excessive inflammation is an effective strategy to improve the prognosis of P. aeruginosa lung infections.
Clinical studies highlight the critical role of metabolism-immune crosstalk in driving excessive inflammation during respiratory viral infections. Glucose metabolism is associated with the onset of cytokine storms in patients infected with the influenza A virus11. Circulating serum metabolites, such as arginine, tryptophan, kynurenic acid, purines, and fatty acids, may exacerbate the production of inflammatory cytokines in COVID-19 patients12,13. Whether metabolic intervention will become a strategy for pneumonia, including Gram-negative bacterial pneumonia, is anticipated. To date, numerous studies have reported that the metabolic microenvironment is closely associated with the inflammatory phenotypes of macrophages, which serve as the first defense line of pneumonia. In glycolipid metabolism, it is well established that pro-inflammatory macrophages predominantly exhibit aerobic glycolysis and fatty acid synthesis, while anti-inflammatory macrophages utilize oxidative phosphorylation (OXPHOS) and fatty acid oxidation14,15. Regarding amino acid metabolism, the conversion of nitric oxide (NO) by arginine, the generation of α-ketoglutarate by glutamine, and the transformation of citrulline via argininosuccinate synthetase often occur in inflammatory macrophages16,17,18,19. In addition to the above metabolic adaptations, there exist other potential changes of enzymes and metabolites that have not been fully explored and reported in macrophages.
Phosphoglycerate dehydrogenase (Phgdh) is the first key rate-limiting enzyme in the glycolysis-serine biosynthesis pathway. Alterations in its expression level, enzymatic activity and cellular localization can significantly influence cellular glucose and serine metabolism, as well as modulate the reduction reaction of α-KG and the oxidation reaction of malate20,21,22. Recent studies have demonstrated Phgdh modulation can alter the inflammatory response and metabolic phenotype of macrophages during viral and Salmonella infections23,24. Phgdh inhibitors can also regulate caspase3/7-mediated apoptosis in epithelial cells during COVID-19 infection25. These preliminarily revealed that Phgdh may play an important role in infectious diseases. Furthermore, it has been reported that serine can provide nutrients for one-carbon metabolism through the synthesis of GSH- or SAM-mediated histone methylation in macrophages, thereby promoting LPS-mediated interleukin (IL)−1β production26,27. These findings highlight the potential involvement of de novo serine synthesis in macrophages during Gram-negative bacterial infections. Targeting Phgdh or serine uptake and utilization may emerge as important strategies for the prognosis of P. aeruginosa pneumonia.
Here, we demonstrated that Phgdh expression and enzymatic activity are significantly upregulated during the acute phase of P. aeruginosa-induced lung infection. Our findings revealed predominant Phgdh localization in pulmonary macrophages, where it potentiates pro-inflammatory responses through enhanced serine biosynthesis. Both pharmacological inhibition of Phgdh activity and macrophage-specific genetic deletion of Phgdh improved the prognosis of P. aeruginosa infection, whereas high-L-serine diets exacerbated macrophage-mediated inflammatory responses. Mechanistically, our study reveals that Dusp4-mediated dephosphorylation of ERK1/2 acts as a critical downstream event of Phgdh-dependent serine synthesis; this cascade is directly transcriptionally repressed by the one-carbon metabolism-H3K27me3 axis. Thus, we delineate a direct pathway from metabolic reprogramming to pro-inflammatory macrophage activation during P. aeruginosa infection. This discovery not only advances the understanding of the intricate interplay between metabolism and immune response but also opens the avenues for therapeutic interventions in bacterial infections.
Results
Phgdh is upregulated after P. aeruginosa lung infection and predominantly localized in pulmonary macrophages
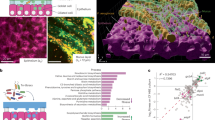
To date, little literature has reported on the changes in expression and enzymatic activity of Phgdh in lung tissue during pathogen infections. To investigate the dynamic expression of Phgdh in response to P. aeruginosa infection, we intranasally (i.n.) infected mice with varying loads of P. aeruginosa standard strain PAO1 (106CFU、2 × 106CFU、5 × 106CFU) and analyzed lung tissues at different time points (0–72 h). We found that the upregulation of Phgdh in lung tissue was dependent on the bacterial load during the acute phase of infection (Fig. 1a). Furthermore, immunohistochemical analysis of lung tissues infected with 2 × 106CFU PAO1 for 24 h, compared with normal lung tissues, revealed significant inflammatory cell infiltration and a higher proportion of Phgdh-positive cells in the infected lung tissue (Fig. 1b). To further explore the cellular location of Phgdh in mouse lung tissue, we analyzed online publicly available single-cell RNA sequencing data from normal mouse lung tissue28 (https://tabula-muris.ds.czbiohub.org/) and found that AMs exhibited the highest percentage of Phgdh-positive cells, with total Phgdh expression levels second only to stromal cells and lymphocytes (Fig. 1c). In a human lung scRNA-seq dataset, Phgdh expression was found to be the third most abundant in alveolar macrophages, after type II alveolar epithelial cells and basal cells, showing a conserved expression pattern between mice and humans29,30 (https://app.lungmap.net/app/shinycell-human-lung-cellref) (Fig. S1a). To validate the co-localization of AMs with Phgdh, we performed immunofluorescence staining with F4/80+ SiglecF+ to identify AMs, and F4/80+ to label the broader pulmonary macrophage population. In normal lung tissue, Phgdh clearly co‑localizes with AMs. Upon P. aeruginosa infection, despite a severe reduction in AM numbers, the remaining AMs exhibited Phgdh signal that maintained co-localization. Concurrently, infected lungs were infiltrated by abundant F4/80⁺SiglecF⁻ macrophages, primarily of monocytic origin, which also showed distinct Phgdh co-localization (Fig. 1d). Furthermore, we deleted AMs in vivo using intranasal administration of chlodronate liposomes (CLLs). This resulted in a significant reduction in the proportion of AMs (CD11b- SiglecF+) (Fig. S1b) and a corresponding decrease in Phgdh expression in lung tissue (Fig. 1e). To further investigate the effects of P. aeruginosa infection on Phgdh expression in AMs, we isolated AMs from bronchoalveolar lavage fluid (BALF) and infected them with PAO1 at different multiplicities of infection (MOI) for 6 h. The results demonstrated a significantly upregulation of Phgdh expression (Fig. S1c). Additionally, we isolated bone marrow cells and induced their differentiation into AMs (iAMs), followed by cell phenotyping. Similarly to AMs isolated from bronchoalveolar lavage fluid (BALF-AMs), iAMs exhibited high expression of CD11c and SiglecF (Fig. S1d). Upon infection with PAO1 (MOI = 1) for 6 h, iAMs showed a significant increase in the expression of Phgdh (Fig. 1f, g). Given that a significant number of bone marrow-derived monocytes are recruited to lung tissue and differentiate into macrophages during the acute phase of infection31, we isolated macrophages from bone-marrow-derived cells (BMDMs) for further confirmation. Consistent with previous findings, PAO1 infection led to an upregulation of Phgdh expression in BMDMs (Fig. S1e). Finally, we assessed the enzymatic activity of Phgdh in lung tissue and pulmonary macrophages during P. aeruginosa infection, revealing a dose- and time-dependent increase following PAO1 challenge (Fig. S1f, g). Collectively, these findings indicate that P. aeruginosa infection induces increased expression and activity of Phgdh in lung tissue, primarily in macrophages.
a Expression of Phgdh in mouse lung tissue after PAO1 infection; b IHC analyses of Phgdh in normal lung tissue and 2 × 106 PAO1 infected lung tissue for 24 h, Scale bar = 100 μm (n = 4, means ± SEMs). An unpaired, two-tailed Student’s t-test was used for statistical comparisons; c Single-cell sequencing analysis of the cellular location of Phgdh in mouse lung tissue; d IF analyses of co-localization of Phgdh and AMs (F4/80+SiglecF+) in normal lung tissues and 2 × 106 PAO1 infected lung tissues for 24 h, Scale bar = 20 μm (n = 4); e Expression of Phgdh in mouse lung tissue treated with Control Liposomes and Chlodronate Liposomes (CLLs) (n = 4); f Expression of Phgdh in iAMs infected with different multiplicity of PAO1 (MOI = 0, 0.1, 1, 5, 10, 20); g IF analyses of co-localization of Phgdh and iAMs after PAO1 (MOI = 1) infection, Scale bar = 20 μm (n = 3). All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
Phgdh blockage improves the prognosis of P. aeruginosa lung infection
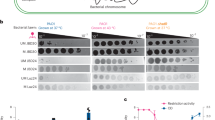
CBR-5884, a selective enzymatic activity inhibitor that suppresses the oligomerization of Phgdh, has been widely used in previous studies32. To investigate the role of Phgdh in the prognosis of P. aeruginosa lung infection, we infected mice with a lethal dose or 2 × 106CFU PAO1 and then treated them with CBR-5884 to monitor survival and disease outcome (Fig. 2a). Following a lethal intranasal challenge with 10⁷ CFU of PAO1, a significant improvement in survival was observed in mice treated with CBR-5884 (Fig. 2b). Following a sublethal infection model, histopathological analysis using hematoxylin and eosin (H&E) staining demonstrated reduced immune cells infiltration and significant reduction of lung injury in the CBR-5884-treated group (Fig. 2c). Mice treated with CBR-5884 exhibited lower lung weight and reduced protein concentration in lung tissue (Fig. 2d). Bacterial colony counts revealed significantly reduced bacterial loads in both BALF and lung tissue homogenates in the CBR-5884-treated group compared to the control group (Fig. 2e). We further assessed the mRNA expression of several pro- and anti-inflammatory cytokines, including Il6, Il1b, Tnfα, Nos2, Il10 and Arg1. Notably, CBR-5884 significantly reduced both the mRNA and protein expression of IL-6 and IL-1β after P. aeruginosa infection (Fig. 2f, g). To access the infiltration of inflammation cells in lung tissue, we stained myeloid innate immune cells in lung tissue (Fig. S2a). We observed that CBR-5884 ameliorated the reduction of AMs and decreased the proportion of monocyte derived macrophage after infection (Fig. 2h). We also compared the mean fluorescence intensity (MFI) of CD86 and CD206 in lung tissue under the CD64+ CD11c+F4/80+ gate (Fig. S2b). CBR-5884 significantly reduced the proportion of pro-inflammatory macrophages, while its effect on anti-inflammatory macrophages was not significant after P. aeruginosa infection (Fig. 2i). Together, these results indicate that inhibiting Phgdh activity significantly improves the prognosis of P. aeruginosa lung infection by alleviating the pro-inflammatory response of pulmonary macrophages.
a Schematic of Phgdh enzymatic inhibition in DMSO and CBR-5884 treated mice. Created in BioRender. Chen, R. (2026) https://BioRender.com/y1fflg1; b Survival rates of DMSO and CBR-5884 treated mice after 107 CFU PAO1 infection for 7 days (n = 10 for DMSO treatment, n = 9 for CBR-5884 treatment, log rank test [Mantel-Cox]); c Representative images of H&E-stained lung sections and lung damage scores, Scale bar = 200μm (n = 4, means ± SEMs); d Lung weight and lung protein concentration (n = 4, means ± SEMs); e CFUs of PAO1 in BALF and whole-lung homogenates (n = 4, means ± SEMs); f mRNA expression of pro-inflammation and anti-inflammation cytokines in lung tissue (n = 4, means ± SEMs); g Protein concentrations of IL-6 and IL-1β in lung tissue (n = 4, means ± SEMs); h Percentage of the cellular infiltration in lung tissue (n = 4, means ± SEMs); i Flow cytometry analysis of the MFI of CD86+ and CD206+ macrophages in lung tissue of DMSO and CBR-5884 treated mice after 2 × 106 CFU PAO1 (n = 4, means ± SEMs). An unpaired, two-tailed Student’s t-test was used for all statistical comparisons. All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
Phgdh enhances the pro-inflammatory response of macrophages
Given that Phgdh inhibitors reduced the pro-inflammatory pulmonary macrophages, mitigated the sharp decline in AMs, and suppressed the recruitment of monocyte-derived macrophages in vivo, we sought to further explore the role of Phgdh in regulating the inflammatory phenotype of macrophages in vitro. We first pre-treated the AMs cell line MH-S with CBR-5884 and stimulated it with PAO1 (MOI = 1). Cytokine levels of IL-6 and IL-1β were measured at both mRNA and protein levels after infection. In the uninfected state, CBR-5884 had no significant effect on the expression of IL-6 and IL-1β. However, it significantly reduced the production of IL-6 and IL-1β by MH-S after PAO1 infection (Fig. S3a). Next, we pre-treated the iAMs and BMDMs with CBR-5884 and stimulated them with PAO1 (MOI = 1) for 2 h. A marked reduction in IL-6 and IL-1β levels was observed in the CBR-5884 groups (Figs. 3a, S3b). To further validate these findings, we designed several siRNA sequences targeting Phgdh and screened for the silencing efficiency (Figs. 3b, S3c). Depletion of Phgdh significantly decreased Il6 and Il1b levels in MH-S after PAO1 infection (Fig. S3d). Similarly, silencing Phgdh in iAM and BMDMs using siPhgdh confirmed the decrease in mRNA expression of Il6 and Il1b (Figs. 3c, S3d). We further examined the effect of Phgdh inhibition on the pro-inflammatory macrophage phenotype. Knockdown of Phgdh in iAMs significantly reduced the proportion of CD86+ pro-inflammatory macrophages within the SiglecF+CD11c+ population, while it had no effect on the proportion of SiglecF+CD11c+ macrophages or CD206+ anti-inflammatory macrophages (Fig. 3d). Comparable results were obtained in BMDMs, where Phgdh knockdown significantly decreased the proportion of CD86+ pro-inflammatory macrophages within the CD11b+ F4/80+ population, without effecting CD206+ anti-inflammatory macrophages (Fig. S3e). Additionally, we overexpressed Phgdh in iBMDMs and found that Phgdh overexpression resulted in increased mRNA levels of Il6 and Il1b (Fig. 3e). These data indicate that Phgdh enhances the pro-inflammatory response of macrophages, including AMs and bone marrow-derived macrophages.
a Expression of IL-6 and IL-1β in iAMs pre-treated with DMSO and CBR-5884 after PAO1 infection; b Knockdown efficiency of siPhgdh in iAMs. FAM is a fluorescent dye (fluorophore) used to track and visualize the uptake, localization, and transfection efficiency of siRNA in cells; c mRNA expression of Phgdh, Il6 and Il1b in iAMs transfected with siNC and siPhgdh after PAO1 infection; d Histogram overlay of the MFI of CD86+ and CD206+ iAMs (gated on the CD11c+ SiglecF+ population) in siNC and siPhgdh transfection group after PAO1 infection; e Protein and mRNA expression of Phgdh, and mRNA expression of Il6 and Il1b in iBMDMs infected with the lentivirus expressing empty vector (EV) or Phgdh (OE) after PAO1 infection. Data are presented as mean ± SEM from 3 biologically independent experiments. An unpaired, two-tailed Student’s t-test was used for statistical comparisons. All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
L-serine in macrophages is related with hyperinflammation and poor prognosis in P. aeruginosa infection
To elucidate the underlying mechanism by which Phgdh influences the prognosis of P. aeruginosa pulmonary infection, we initially investigated the classic downstream metabolite of Phgdh. We utilized U-[13 C]-glucose tracers and found that within 6 h of tracing, macrophages under PAO1 infection had significantly higher levels of m + 3 serine and m + 6 to 9 SAM (both derived from glucose26,33) as well as the elevated total metabolite pools, indicating the activation of the de novo serine synthesis (SSP) and enhanced one-carbon unit flux for SAM production (Fig. 4a). Consistent with this, both lung tissue and iAMs exhibited marked upregulation of L-serine following PAO1 challenge (Fig. S4a). To distinguish host-derived serine from potential bacterial contributions, we used heat-killed PAO1 as a control. Although total serine and SAM levels were higher with heat-killed bacteria, the ¹³C-enrichment from glucose in these metabolites was comparable to that in live infection conditions. This demonstrates that Phgdh-mediated SSP is independent of bacterial viability (Fig. 4a). The elevated total metabolites in the heat-killed group may suggest that live PAO1 suppresses other serine sources. Furthermore, PAO1 growth in vitro was unaffected by serine supplementation (Fig. S4b), ruling out significant bacterial serine uptake or auxotrophy as a confounding factor. Together, these findings establish that the enhanced SSP and one-carbon metabolism during infection primarily result from host metabolic reprogramming.
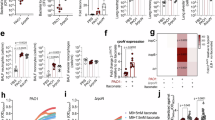
a Total abundance and 13C-enrichment of de novo serine and SAM in iAMs after PAO1 or heat-killed PAO1 infection (6 h; U-13C-glucose tracer) (n = 5 for total sample, n = 4 for U-13C-glucose tracer, n = 1 for 12C-glucose negative control, means ± SEMs); b qPCR analysis of Il6 and Il1b mRNA in iAMs treated with different concentrations of L-serine (0, 400, 800,1000 μmol/L, μM) after PAO1 infection (n = 3, means ± SEMs); c Flow cytometry detection of the MFI of CD86+ iAMs treated with different concentrations of L-serine (n = 3, means ± SEMs); d Schematic of control and high serine diet-fed mice. Created in BioRender. Chen, R. (2026) https://BioRender.com/ss002xx; e Survival rates of control (10 mg/kg serine) and high-serine diet (50 mg/kg serine)-fed mice after 107 CFU PAO1 infection for 4 days (n = 10, log rank test [Mantel-Cox]); f CFUs of PAO1 in lung tissue (n = 5, means ± SEMs); g Representative images of H&E-stained lung sections and lung damage scores, Scale bar = 100 μm (n = 5, means ± SEMs); h qPCR analysis of Il6 and Il1b mRNA in mouse lung tissue (n = 5, means ± SEMs); i Percentage of the AMs in lung tissue (n = 5, means ± SEMs); j Flow cytometry analysis of the MFI of CD86+ macrophages in mouse lung tissue between control and high-serine diet-fed mice after 2 × 106 PAO1 infection (n = 5, means ± SEMs). An unpaired two-sided Student’s t-test was used in (a, f–j); One-way ANOVA followed by Dunnett’s multiple comparison test in (b, c). All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
To assess the impact of L-serine on the inflammatory response, iAMs were starved for 12 h and then treated with 400 μM, 800 μM and 1000 μM of L-serine for 4 h, followed by 2 h of PAO1 stimulation. The results revealed that 800μM L-serine significantly enhanced the secretion of Il6 and Il1b secreted by iAMs (Fig. 4b). Similar regulatory effects of L-serine on Il6 and Il1b were also observed in MH-S and BMDMs (Fig. S4c). Consistent with these findings, 800μM L-serine markedly increased the proportion of CD86+ pro-inflammatory iAMs after PAO1 stimulation, while it had no significant effect on CD206+ anti-inflammatory iAMs (Figs. 4c, S4d). To further explore the effect of L-serine on the prognosis of P. aeruginosa lung infection, we divided mice into high-serine diet-fed and control-fed groups for three-weeks (Fig. 4d). Survival studies showed that the control diet significantly decreased the mortality compared to the high-serine diet following challenge with a lethal dose of PAO1 (Fig. 4e). There were no differences in baseline body weights between high-serine diet and control diet, whereas weight loss was significantly ameliorated in the control diet group after PAO1 infection (Fig. S4e, S4f). Compared with the control-fed mice, high-serine diet-fed mice had higher bacterial loads in lung tissue 24 h post-infection (Fig. 4f). H&E staining of the lungs after PAO1 infection showed less immune cell infiltration and tissue injury in control-fed mice compared to high-serine diet-fed mice (Fig. 4g). Consistent with this, high-serine diet-fed mice exhibited heavier lung weight and higher lung homogenate protein concentration (Figs. S4g). The expression of Il6 and Il1b was also upregulated in high-serine diet-fed mice after PAO1 infection (Fig. 4h). We observed a more pronounced depletion of AMs in the lung tissue of the high-serine-diet fed mice after PAO1 infection, while there were no significant differences in neutrophils, eosinophils, total macrophages, and trans-macrophages compared to the control-fed group (Fig. 4i, S4h). Moreover, the proportion of CD86+ pro-inflammatory macrophages in lung tissue after PAO1 infection was significantly higher in high-serine diet-fed mice than in control-fed mice, whereas the difference in the proportion of CD206+ was not significant (Fig. 4j, Fig. S4i). Together, these findings indicate that L-serine is the key metabolite that mediates the pro-inflammatory response of macrophages.
Phgdh-driven serine synthesis promotes macrophage inflammation via ERK1/2 phosphorylation
To elucidate the mechanism by which Phgdh promotes the pro-inflammatory response in macrophages, we performed RNA sequencing (RNA-seq) analysis to identify the differentially expressed genes between the siNC and siPhgdh-treated iAMs after PAO1 infection (PRJNA1357662) (Fig. S5a). Transcriptional profiling validated the role of Phgdh in macrophage pro-inflammatory activation, revealing significant downregulation of key pro-inflammatory genes (e.g., Il6, Il1b, Nos2, Cd86, Cd80, Cxcl9, Nfkb1, and Stat1); in contrast, the overall expression of anti-inflammatory genes showed no consistent trend (Fig. 5a). Among the signaling pathways associated with the differentially expressed genes, the MAPK pathway was the most significantly enriched (Fig. 5b). Gene Set Enrichment Analysis (GSEA) further confirmed that the MAPK pathway was also significantly enriched between the two groups (Fig. S5a). We then investigated whether the MAPK signaling pathways were altered upon inhibition of Phgdh. We found that CBR-5884 significantly reduced the phosphorylation level of ERK1/2 in lung tissue after PAO1 infection but had no significant effect on the activation of p38 and JNK (Fig. 5c). In vitro experiments also demonstrated that inhibiting the enzymatic activity or knocking down the expression of Phgdh led to decreased activation of ERK1/2 in iAMs (Figs. S5b, Fig. 5d). Conversely, overexpression of Phgdh significantly enhanced the phosphorylation level of ERK1/2 in macrophages (Fig. 5e).
a Transcriptional profiling of pro-inflammatory genes and anti-inflammatory genes between siNC and siPhgdh-treated iAMs after PAO1 (MOI = 1) infection for 2 h (n = 3); b KEGG pathway enrichment analysis of DEGs between siNC and siPhgdh-treated iAMs after PAO1 infection (n = 3); c Western blot analysis of MAPK signaling pathway (p-ERK/ERK1/2、p-P38/P38、p-JNK/JNK) in lung tissue of DMSO and CBR-5884 treated mice; d Western blot analysis of MAPK signaling pathway in iAMs transfected with siNC or siPhgdh; e Western blot analysis of MAPK signaling pathway in iBMDMs infected with lentivirus expressing empty vector (EV) or Phgdh (OE); f Concentration of serine in lung tissue and iAMs pre-treated with CBR-5884 or siPhgdh after PAO1 infection (n = 3, means ± SEMs); g Activation of p-ERK1/2 in lung tissue of control- (10 mg/kg serine) and high-serine (50 mg/kg serine) diet-fed mice; h Western blot analysis of p-ERK/ERK1/2 in iAMs transfected with siNC and siPhgdh and treated with or without L-serine (400 μM); i Il6 and Il1b mRNA in iAMs transfected with siNC and siPhgdh and treated with or without L-serine (400 μM) after PAO1 infection (n = 3, means ± SEMs); j qPCR analysis of Il6 and Il1b mRNA in iAMs 1 h pre-treated with ERK1/2 inhibitor (U0126-Etoh) after stimulation with 0 and 400 μM L-serine (n = 3, means ± SEMs). An unpaired two-sided Student’s t-test was used in (f, j); One-way ANOVA followed by Dunnett’s multiple comparison test in (i). All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
To elucidate the mechanism by which Phgdh-driven serine synthesis promotes macrophage inflammation via ERK1/2 phosphorylation, we measured serine levels after Phgdh inhibition both in vitro and in vivo. A consistent and significant decrease in L-serine was observed in lung and iAMs (Fig. 5f). We further also observed that L-serine treatment for more than 2 h significantly increased the phosphorylation levels of ERK1/2 in iAMs under normal physiological conditions (Fig. S5c). Compared to the L-serine-deficient group, 400μmol L-serine significantly enhanced ERK1/2 activation in iAMs after PAO1 infection (Fig. S5c). Lung tissue from high-serine diet-fed mice also exhibited higher levels of p-ERK1/2 after PAO1 infection (Fig. 5g). To determine whether L-serine serves as the key metabolite of Phgdh that exacerbates ERK1/2 phosphorylation, we pretreated Phgdh-knockdown iAMs with L-serine and found that L-serine could effectively compensate for the inhibitory effect of Phgdh knockdown on p-ERK1/2 expression (Fig. 5h). Supplementation of L-serine to Phgdh-knockdown iAMs significantly rescued the inhibitory effect on Il6 and Il1b secretion and reduced the proportion of CD86+ macrophages (Figs. 5i, S5d). Furthermore, we found that the ERK1/2-specific inhibitor U0126-Etoh blocked the L-serine-induced activation of p-ERK1/2 (Fig. S5e) and the upregulation of Il6 and Il1b expression (Fig. 5j). These results indicate that L-serine is a crucial downstream metabolite of Phgdh in driving the pro-inflammatory response of AMs. Phgdh-mediated L-serine synthesis promotes pro-inflammatory macrophage responses through the activation of the ERK1/2 signaling pathway, highlighting the critical role of ERK1/2 phosphorylation in regulating macrophage-mediated inflammatory outcomes during P. aeruginosa infection.
Serine is generally classified as a nonessential amino acid, but its functional importance, particularly in redox homeostasis, has been increasingly recognized. Previous work demonstrates that serine supports glutathione (GSH/GSSG) synthesis, thereby providing feedback inhibition of ROS production upon LPS stimulation27. Mitochondrial serine catabolism also contributes to NAD(P)H generation, aiding cellular redox balance34. In our P. aeruginosa infection model, transcriptomic analysis of Phgdh-knockdown cells revealed significant enrichment of oxidative stress and redox pathways (Fig. S5f). Phgdh-deficient macrophages exhibited elevated ROS levels as early as 0.5 h post-infection (Fig. S5f). Similarly, serine deprivation led to increased ROS production in PAO1 infected macrophages (Fig. S5f). These findings suggest that Phgdh may regulate redox homeostasis during P. aeruginosa infection by modulating serine metabolism, in addition to its role in ERK1/2 pathway activation, a possibility that merits further investigation.
Myeloid Phgdh deficiency modulates pro-inflammatory macrophages to improve the prognosis of P. aeruginosa lung infection
To further validate the impact of Phgdh in AMs on the prognosis of P. aeruginosa lung infection, we specifically deleted Phgdh in macrophages by crossing Phgdhfl/fl mice with Lyz2Cre mice (Fig. 6a). The knockout efficiency of Phgdh was confirmed in both mice and AMs (Fig. S6a, 6c). Notably, iAMs isolated from Phgdhfl/flLyz2Cre mice exhibited reduced expression of pro-inflammatory cytokines (e.g., Il6, Il1b, Nos2, Cd80, Nfkb1 and Stat1) after PAO1 infection, whereas the levels of anti-inflammatory mediators, including Arg1, Il10 and Stat6, remained unchanged between groups (Fig. 6b). For functional validation, we assessed the protein levels of iNOS and Arg-1 at 2 h post-infection. We found that Phgdh knockout significantly reduced iNOS expression, whereas Arg-1 levels remained largely unaffected. Furthermore, we observed a significant reduction in cleaved IL-1β (Fig. 6c). The phosphorylation of ERK1/2 in iAMs from Phgdhfl/flLyz2Cre mice was also lower compared to Phgdhfl/fl mice (Fig. 6c). Additionally, the proportion of CD86+ pro-inflammatory macrophages in iAMs from Phgdhfl/fl Lyz2Cre mice was significantly lower than that from Phgdhfl/fl mice after PAO1 infection, while no significant difference was observed in the proportion of CD206+ macrophages (Fig. 6d).
a Breeding scheme for the generation of Phgdhfl/flLyz2Cre mice and Phgdhfl/fl mice. Created in BioRender. Chen, R. (2026) https://BioRender.com/ml9vxvw; b qPCR analysis of Phgdh, pro-inflammatory cytokines (e.g., Il6, Il1b, Nos2, Cd80, Nfkb1 and Stat1) and anti-inflammatory cytokines (e.g., Arg1, Il10, and Stat6) mRNA in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice (n = 3, means ± SEMs); c Western blot analysis of MAPK pathway-related proteins, Arg-1, iNOS and cleaved IL-1β in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice; d Flow cytometry analysis of the MFI of CD86+ and CD206+ cells in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice (n = 3, means ± SEMs); e Survival rates of Phgdhfl/flLyz2Cre and Phgdhfl/fl mice after 107 CFU PAO1 infection for 1 week (n = 10, log rank test [Mantel-Cox]); f Representative images of H&E-stained lung sections, Scale bar = 100 μm (n = 6); g CFUs of PAO1 in lung tissue (n = 6, means ± SEMs); h qPCR analysis of Il6 and Il1b mRNA (n = 6, means ± SEMs); i Percentage of the cellular infiltration (n = 7, means ± SEMs); j MFI of CD86+ and CD206+ macrophages in lung tissue of Phgdhfl/flLyz2Cre and Phgdhfl/fl mice after 2 × 106 CFU PAO1 infection (n = 7, means ± SEMs); k Schematic of adoptive transfers of AMs (2 × 105) from Phgdhfl/flLyz2Cre or Phgdhfl/fl to WT mice. Created in BioRender. Chen, R. (2026) https://BioRender.com/a322fzj. All groups of mice were inoculated with PAO1 (1 × 106 CFU) 4 h later and were sacrificed after 24 h infection for further analysis. An equal volume of PBS was intranasally instilled as the control; l Representative images of H&E-stained lung sections, Scale bar = 200 μm (n = 4 for PBS control group, n = 7 for Phgdhfl/flrescue group, n = 6 for Phgdhfl/flLyz2Cre rescue group); m Expression of Il6 and Il1b in lung tissue from AM-rescue groups of Phgdhfl/flLyz2Cre and Phgdhfl/fl mice (n = 4, means ± SEMs). An unpaired two-sided Student’s t-test was used in (b, d, g–j); One-way ANOVA followed by Dunnett’s multiple comparison test in m. All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
In vivo Phgdh inhibition or dietary serine restriction significantly decreased P. aeruginosa burden in the lungs. To investigate if this effect is macrophage-dependent, we assessed bacterial handling in alveolar macrophages from Phgdhfl/fl mice with Lyz2Cre mice. Phgdh-deficient macrophages exhibited enhanced phagocytosis of PAO1 by 0.5 h (Fig. S6b) and more efficient intracellular bacterial clearance by 1 hour (Fig. S6c), suggesting that Phgdh loss amplifies both phagocytic and bactericidal capacities.
Next, to determine the intrinsic function of Phgdh in macrophages, we infected both Phgdhfl/fl and Phgdhfl/fl Lyz2Cre mice with PAO1. Similar to the results from in vivo experiments with the Phgdh inhibitor, Phgdhfl/fl Lyz2Cre mice exhibited lower mortality and improved prognosis after PAO1 infection (Fig. 6e). The lung injury (including lung weight and protein concentration), bacterial loads, and the mRNA expression of Il6 and Il1b in the lungs were lower in Phgdhfl/fl Lyz2Cre mice after infection with 2 × 106 CFU PAO1 (Fig. 6f, g, h). Furthermore, we observed an amelioration in the reduction of AMs and a decrease in monocyte derived macrophages in Phgdhfl/fl Lyz2Cre mice (Fig. 6i). The proportion of pro-inflammatory macrophages was significantly reduced, while the effect on anti-inflammatory macrophages was not significant after PAO1 infection (Fig. 6j). To verify the role of Phgdh in AMs during PAO1 infection, we isolated adoptive AMs from the BALF of Phgdhfl/flLyz2Cre and Phgdhfl/fl mice and transferred these cells, along with PBS, into wild-type (WT) mice via intranasal administration. We then challenged all groups of mice with a dose of 2×106 CFU PAO1 (Fig. 6k). Mice receiving PBS had the worst prognosis, while mice receiving AMs from Phgdhfl/flLyz2Cre mice showed significantly reduced severity of infection and lung injury, as evidenced by markedly reductions in bacterial loads, body loss, lung weight and expression of Il6 and Il1b in lung tissue (Fig. 6l-m, S6d-f). Similarly, tail vein transfer of BMDMs from Phgdhfl/flLyz2Cre into WT mice also resulted in improved prognosis (Fig. S6d-f). Furthermore, when comparing the prognosis between Phgdhfl/flLyz2Cre and Phgdhfl/fl mice under different concentration L-serine diets, we found that a high L-serine diet significantly abolished the anti-inflammatory effects of Phgdh deletion in macrophages during PAO1 infection (Fig. S6g).
De novo serine synthesis transcriptionally regulates Dusp4 through the one-carbon metabolism-H3K27me3 axis
To systematically identify upstream regulators connecting L-serine to ERK1/2 signaling, we performed differential gene expression analysis on transcriptomic data from Phgdh-knockdown (PRJNA1357662) and serine-deprived iAMs (PRJNA1358829). Intersection of these two gene sets identified candidate genes that commonly influence ERK1/2 phosphorylation, including direct kinases and positive/negative regulators (Fig. 7a). Subsequent transcriptional and protein-level validation of these candidates pinpointed dual-specificity phosphatase 4 (Dusp4) as a critical upstream mediator (Fig. 7b, c).
a Candidate genes that commonly modulate ERK1/2 phosphorylation under both Phgdh knockdown and serine deprivation conditions (n = 3); b qPCR analysis of the candidate genes (e.g., Dusp1, Dusp2, Dusp3, Dusp4, Dusp5, Dusp9, Dusp10, Spry2, Fgfr1) in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice (n = 3, means ± SEMs); c Western blot analysis of Dusp2 and Dusp4 in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice, and iAMs treated with or without L-serine after PAO1 infection; d qPCR analysis of Dusp4, Il6 and Il1b mRNA in iAMs transfected with siNC and siDusp4 and treated with or without L-serine after PAO1 infection (n = 3 for serine deprived group, n = 4 for serine treated group, means ± SEMs); e Schematic diagram of 13C flux in the SSP and one-carbon metabolism, and heatmap of targeted metabolites in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice after PAO1 infection (n = 5 for Phgdhfl/fl group, n = 6 for Phgdhfl/flLyz2Cre group); f Distribution of targeted metabolites in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice after PAO1 infection (n = 5 for Phgdhfl/fl group, n = 6 for Phgdhfl/flLyz2Cre group, means ± SEMs); g Distribution of 13C labels metabolites in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice after PAO1 infection (n = 4 for the 13C labeled Phgdhfl/fl group, n = 5 for the 13C labeled Phgdhfl/flLyz2Cre group, means ± SEMs); h Western blot analysis of H3K27me3 in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice, and iAMs treated with or without L-serine after PAO1 infection; i ChIP-qPCR analysis of the binding of H3K27me3 to the Dusp4 promoter regions in iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice with primers targeting distinct sites (n = 4, means ± SEMs); j ChIP-qPCR analysis of H3K27me3 enrichment at the Dusp4 promoters in uninfected and PAO1 infected iAMs derived from Phgdhfl/flLyz2Cre and Phgdhfl/fl mice (n = 3, means ± SEMs); k ChIP-qPCR analysis of H3K27me3 enrichment at the Dusp4 promoters in iAMs cultured with vehicle, serine (400 μM) or SAM (200 μM) (n = 3, means ± SEMs). An unpaired, two-tailed Student’s t-test was used in (b, f, g); One-way ANOVA followed by Dunnett’s multiple comparison test in (d, k). All experiments are repeated three times independently and similar results are obtained. Source data are provided as a Source Data file.
Dusp4 (also known as MAPK phosphatase 2, MKP2) is a member of the dual-specificity phosphatase family that negatively regulates MAPK signaling and plays important roles in cell proliferation and differentiation35. In models of LPS-induced acute lung injury, suppression of Dusp4 was shown to enhance the expression of inflammatory cytokines, including Tnfα, Il1, and Il636. Previous studies have established that Dusp4 directly interacts with ERK1/2, and emerging evidence underscores its importance in anti-parasitic and anti-influenza immunity37,38. These findings collectively suggest that Dusp4 may serve as a key regulator in innate immune responses. To investigate its role in P. aeruginosa infection model, we performed siRNA-mediated knockdown of Dusp4 in macrophages (Fig. S7a). This intervention resulted in a marked increase in ERK1/2 phosphorylation after PAO1 infection, accompanied by an elevated frequency of CD86⁺ macrophages and enhanced production of pro-inflammatory cytokines (Figs. S7b, 7d, S7c). Importantly, under Dusp4-depleted conditions, the modulatory effects of serine deprivation on ERK1/2 activation and inflammatory phenotypes were largely abolished (Figs. 7d, S7b, S7c), indicating that Dusp4 is an essential node through which serine metabolism regulates ERK1/2 signaling and macrophage inflammatory responses.
Based on established links between S-adenosylmethionine (SAM)—a core metabolite in the serine-driven one-carbon metabolism—and epigenetic regulation of Dusp4 via DNA methylation26,39, we hypothesized that Phgdh-dependent serine synthesis regulates Dusp4 expression via one-carbon metabolism and SAM production. Metabolic flux analysis in iAMs from Phgdh-knockout mice demonstrated significantly reduced overall levels of serine, glycine, methionine, and ATP (Fig. 7e, f). We also observed decreased enrichment of glucose-derived m + 3 serine, m + 2 glycine, and m + 6-9 ATP26,33, along with a reduced SAM/SAH ratio (an index of methylation potential40), collectively indicating impaired SSP and one-carbon metabolism (Fig. 7g). Consistent with these findings, competitive inhibition assays confirmed that SAM levels were significantly elevated following PAO1 infection but substantially decreased upon Phgdh deletion (Fig. S7d).
We next investigated whether Phgdh-mediated serine metabolism regulates Dusp4 expression through DNA methylation. However, bisulfite sequencing analysis under Phgdh knockout or serine deprivation conditions revealed no significant changes in methylation of CpG sites in the Dusp4 promoter (Supplementary Table 2). Given the established role of one-carbon metabolism and SAM in modulating histone methylation, along with previous evidence demonstrating that Phgdh loss or serine deprivation reduces H3K27me3 levels23,26, we explored this alternative epigenetic mechanism. We confirmed that both Phgdh knockdown and serine deprivation significantly reduced global H3K27me3 levels (Fig. 7h). Pharmacological inhibition of H3K27me3 concentration-dependently increased Dusp4 expression, whereas enhancing H3K27me3 concentration-dependently suppressed it, supporting a regulatory relationship between H3K27me3 and Dusp4 (Fig. S7e). To directly examine H3K27me3 occupancy at the Dusp4 promoter, we performed chromatin immunoprecipitation followed by quantitative PCR (ChIP-qPCR). The results showed significant H3K27me3 enrichment at the Dusp4 promoter compared to IgG control (Fig. S7f), which was markedly reduced upon Phgdh knockout (Fig. 7i, j) and restored by serine or SAM supplementation (Fig. 7k). Collectively, these findings demonstrate that Phgdh-dependent de novo serine synthesis regulates Dusp4 expression through the one-carbon metabolism–SAM–H3K27me3 axis, revealing an epigenetic mechanism that links serine metabolism to ERK1/2 signaling.
Discussion
Previous investigations into the influence of cellular metabolism on macrophage phenotypes have predominantly employed LPS stimulation models. Although LPS represents a crucial virulence factor of Gram-negative pathogens, it fails to recapitulate the intricate composition and dynamic biological activities characteristic of live pathogens41. To date, the metabolic reprogramming of immune cells during acute P. aeruginosa infection remains inadequately explored. Regarding amino acid metabolism, while essential amino acids such as L-arginine and glutamine and their derivatives have established roles in regulating classical and alternative macrophage activation16,42,43,44,45,46, our findings reveal that endogenous non-essential amino acids, particularly L-serine and its de novo synthesis enzyme Phgdh, significantly contribute to macrophage activation following P. aeruginosa infection.
Current therapeutic research for P. aeruginosa lung infection primarily emphasizes combating antibiotic resistance and developing novel antimicrobial agents47, often overlooking critical host immunomodulatory pathways48. Macrophage-mediated inflammatory responses represent a double-edged sword: moderate inflammation facilitates bacterial clearance, whereas excessive inflammation frequently culminates in severe acute lung injury49. This is particularly evident in severe infections where dysregulated cytokine storms contribute to high mortality rates. Our study demonstrates that Phgdh inhibition significantly improves survival in lethal P. aeruginosa infection models. This intervention effectively attenuated excessive pro-inflammatory responses, ameliorated lung injury, and reduced bacterial burden. Furthermore, long-term serine restriction decreased pro-inflammatory macrophage populations, suggesting potential therapeutic value for severe P. aeruginosa pneumonia.
Although serine’s importance in adaptive immunity through T cell proliferation and tumor growth is well-documented49,50,51, its role in innate immunity, particularly macrophage biology, has recently gained recognition23,26,27. Macrophages acquire L-serine through specific transporters (ASCT1/2) or via de novo synthesis through the serine synthesis pathway (SSP), which concurrently generates NADH and α-ketoglutarate52. Under nutrient-sufficient conditions, extracellular uptake predominates, while SSP becomes essential during serine deprivation or increased demand53. The regulation of serine metabolism exhibits context-dependent characteristics. In LPS-stimulated BMDMs, SSP components, including ATF4 and Phgdh demonstrate transient upregulation26,54,55, whereas RNA virus infection suppresses SSP enzymes in peritoneal macrophages23. Extended LPS or LPS + IFN-γ exposure subsequently downregulates Phgdh expression56,57, indicating that Phgdh and serine levels dynamically respond to pathogen type, duration, and infection conditions. The precise changes in Phgdh activity and expression during infection remain incompletely characterized, with limited in vivo validation available. In our P. aeruginosa lung infection model, we observed consistent upregulation of Phgdh activity and expression in pulmonary macrophages in vitro, accompanied by significant Phgdh and L-serine accumulation in infected lung tissue in vivo. These findings suggest that Phgdh expression/activity and serine levels may serve as potential diagnostic biomarkers for P. aeruginosa infection.
The signaling mechanisms underlying Phgdh-mediated macrophage activation require further elucidation. Existing evidence indicates that L-serine deprivation suppresses mTORC1 signaling, impairing inflammasome activation and IL-1β production while altering the M1 macrophage transcriptome and metabolome58. Following LPS stimulation, L-serine conversion to glycine enhances glutathione synthesis and ROS generation27. Serine deprivation also disrupts OXPHOS, reduces NADH levels and the NADH/NAD+ ratio, potentially inhibiting NF-κB activation54,59. Moreover, serine metabolism functions as an epigenetic regulator by modulating methionine, acetyl-CoA, and lactate availability, thereby influencing histone modifications and DNA regulation in macrophages23,26,54.
Beyond these established mechanisms, our study identifies an additional pathway through which Phgdh regulates macrophage activation via ERK1/2 phosphorylation within the MAPK cascade. MAPKs represent highly conserved serine/threonine kinases that transduce extracellular stimuli into diverse cellular responses, governing gene expression, metabolism, and survival60. The ERK1/2 pathway specifically modulates pro-inflammatory cytokine expression (e.g., Il1b, Il6, Tnfα) and chemokine production through AP-1 and NF-κB activation61,62, while coordinating immune cell recruitment and apoptosis resistance during infection63. In P. aeruginosa infection, we propose that ERK1/2 senses intracellular Phgdh and serine fluctuations to amplify macrophage activation. Supporting this, KEGG analysis of infection-induced transcriptional changes revealed significant MAPK pathway enrichment. Both genetic Phgdh ablation and serine deprivation downregulated ERK1/2 phosphorylation in vivo and in vitro, while pharmacological ERK1/2 inhibition reversed serine-induced pro-inflammatory cytokine enhancement.
To elucidate the specific mechanism through which serine synthesis fuels ERK1/2 phosphorylation, we integrated transcriptomic and metabolic flux analyses. Our investigation revealed that SSP-driven one-carbon metabolism significantly contributes to methylation processes, particularly through H3K27me3, a canonical repressive histone mark. Phgdh-mediated SSP activation enhanced H3K27me3 enrichment at the Dusp4 promoter region, subsequently repressing Dusp4 transcription. This epigenetic silencing attenuated Dusp4-mediated dephosphorylation of ERK1/2, thereby augmenting ERK1/2 signaling transduction. These findings provide direct mechanistic insights from a metabolo-epigenetic perspective.
The prognosis of lung infections depends not only on infection severity but also on host nutritional status, which critically regulates immune function. While our study highlights the importance of de novo serine synthesis in P. aeruginosa infection, several limitations should be acknowledged. Technical constraints prevented complete blockade of all serine synthesis and uptake pathways, though we minimized basal serine influences using serine-free media in vitro and low-serine diets in vivo. Several questions remain unresolved: whether P. aeruginosa, as a metabolically active pathogen, affects non-glucose-derived serine sources; how Phgdh-mediated serine synthesis influences redox homeostasis and its hierarchical relationship with pro-inflammatory responses in macrophages. Future studies should investigate metabolic differences between alveolar and myeloid-derived macrophages, examine pathogen-specificity of these mechanisms, and identify specific bacterial components influencing Phgdh accumulation.
Methods
Ethics statement
All animal experiments were performed according to the protocol approved by the Animal Ethics Committee of Ruijin Hospital, Shanghai Jiao Tong University School of Medicine (protocol No. RJ20250307).
P. aeruginosa strain and culture conditions
The wild-type strain of P. aeruginosa, PAO1 (Cat# ATCC-BAA-47; strain HER-1018) and GFP-PAO1 (kindly provided by Prof. Song Yuan-Lin, Zhongshan Hospital, Fudan University, Shanghai, China) were used for in vitro and in vivo infection experiments. A 10 μL aliquot of frozen bacterial stock was added to 4 mL of sterile Luria-Bertani (LB) medium and cultured for 14–18 h at 37 °C with agitation at 220 rpm. Subsequently, 100 μL of the bacterial stock was added to 4 mL of sterile LB medium and cultured for an additional 2 h at 37 °C with agitation at 220 rpm. The bacteria were washed twice with phosphate-buffered saline (PBS) and then resuspended in PBS to measure the optical density (OD). Based on previous studies in our laboratory, an OD 600 of 0.5 corresponds to ~2 × 108 CFU of bacteria. The bacterial suspension was further diluted with PBS to achieve the desired inoculum concentration for experiments. To examine the effect of L-serine on PAO1 growth, LB medium was supplemented with varying concentrations of L-serine.
Mice
7-week-old C57BL/6 J wild-type mice were obtained from Vital River Laboratory (Beijing, China). Phgdhfl/fl and Lyz2Cre (C57BL/6 J) mice were purchased from Cyagen Biosciences (Suzhou, China) using CRISPR‒Cas9 gene editing. To generate myeloid cell-specific Phgdh knockout mice, Phgdhfl/fl mice were crossed with Lyz2Cre mice, yielding Phgdhfl/flLyz2Cre mice and their littermate controls, Phgdhfl/fl. Genotyping was performed by PCR analysis of tail-snip DNA using Phgdh-flox and the Lyz2-Cre-specific primers. Experiments were conducted using sex and age-matched 7-week-old mice. All mice were housed in specified-pathogen-free (SPF) facilities at Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, China. Housing condition was as following: dark/light cycle 12/12, ambient temperature around 21–22 °C and humidity between 40% and 70%.
Mouse models
For the P. aeruginosa infection model, 7-week-old mice were anesthetized with isoflurane and intranasally (i.n.) infected with 50 μL different multiplicity of PAO1 i.n for different time points. In Phgdh inhibition experiments, mice were administered with 200 μL CBR-5884 (50 mg/kg) and an equivalent volume of DMSO (vehicle control) via gastric administration 4 h prior to PAO1 infection. To deplete AMs, mice were treated with clodronate liposomes administered intranasally (i.n.). For the L-serine dietary experiments, mice were fed either a control diet (10 mg/kg L-serine) or high-serine diets (50 mg/kg L-serine) for 3 weeks prior to the experiments. At the end of the experimental period, mice were euthanized, and lung tissues were collected for further analysis.
Primary macrophages culture and cell lines
Primary AMs were isolated from BALF of 7-week-old male mice. The cells were cultured in L-serine-free complete RPMI-1640 supplemented with 10% fetal bovine serum (FBS). Non-adherent cells were removed after 12 h, and the remaining cells were induced in RPMI-1640 medium containing 20 ng/mL granulocyte-macrophage colony-stimulating factor (GM-CSF, Proteintech), 2 ng/mL transforming growth factor-β (TGF-β, Proteintech) and 0.1 μM PPARγ agonist (Rosiglitazone, MCE). Alternatively, iAMs were generated from bone marrow cells, which were cultured in L-serine-free Dulbecco’s modified Eagle’s medium (DMEM) with GM-CSF and TGF-β for 7 days. Rosiglitazone was then added to the culture medium for an additional two days. Murine bone marrow cells were flushed from femurs using cold PBS. Red blood cells were lysed with RBC lysis buffer, and the bone marrow cells were cultured in complete DMEM medium supplemented with 20 ng/mL macrophage colony-stimulating factor (M-CSF, Proteintech) for 7 days to generate BMDMs. The MH-S cell line and iBMDMs were cultured for at least three generations in L-serine-free DMEM containing 10% FBS and 1% penicillin and streptomycin for subsequent experiments. All experimental cells were cultured at 37 °C in a humidified atmosphere with 5% CO2.
siRNA and plasmid transfection
Small interfering RNAs (siRNAs) targeting Phgdh, Dusp4 and a negative control (NC) were purchased from Genepharma (Shanghai, China). Transfection of MH-S, iAMs and BMDMs with siRNA was performed using lipofectamine RNAiMAX (Thermo Fisher Scientific, USA) according to the manufacturer’s instructions. Cells were harvested 24h-48h after transfection to assess the knockdown efficiency by qPCR (24h) and western blot analysis (48h). For plasmid transfection, Phgdh was subcloned into the pCDH-CMV-MCS-EF1-copGFP-T2A-Puro vector using standard molecular biology techniques. The empty vector and Phgdh-overexpressing plasmid were introduced into 293FT cells for lentivirus production. Immortalized Bone Marrow-Derived Macrophages (iBMDMs) were infected with the lentivirus to achieve stable Phgdh overexpression.
Cell stimulation
Before infecting macrophages with PAO1 (MOI = 1), antibiotics were removed from the culture medium and the concentration of FBS was reduced by half. If the duration of PAO1 stimulation exceeded 4 h, gentamicin (50 μg/ml) was added 1 hour after infection to eliminate the extracellular bacteria. For the Phgdh inhibition experiments, cells were pretreated with CBR-5884 (15 μM) or an equal volume of solvent control (DMSO) 12 h prior to PAO1 infection. For the L-serine deprivation and supplementation experiments, cells were cultured in L-serine-free medium, and different concentrations of L-serine were added to the culture medium 4 h before PAO1 infection.
Measurement of Phgdh activity, serine concentration, SAM concentration and cytokines
Phgdh activity was measured using a Phgdh Activity Assay Kit (Ab273328). Lung tissues and cell lysates were collected and homogenized, followed by centrifugation at 10,000 × g for 5 min. The production of NADH from Phgdh substrate in 96-well plates was quantified by measuring the absorbance at 450 nm using a BioTek Synergy Neo Multi-Mode Plate Reader, which indirectly reflects the enzymatic activity of Phgdh. The serine content in lung tissues and cell lysates was measured using a Serine Assay Kit (ab241027). The assay is based on the oxidation of intermediate metabolites in D-serine metabolism, which generates stable fluorescence upon reaction with a specific probe. By adding serine racemase, L-serine can be converted to D-serine, enabling the separate detection of total serine content (in the presence of serine racemase) and D-serine content (in the absence of serine racemase). The L-serine content was then calculated by subtracting the D-serine content from the total serine content. For SAM measurement, cell lysates were collected and centrifugation at 1000 × g for 5 min, followed by a competitive inhibition reaction (UCEG414Ge). For cytokine measurement, supernatants from homogenized tissues and cell culture media were collected and centrifuged at 1000 g for 10 min. The concentrations of cytokines (IL-6 and IL-1β) were measured using ELISA kits according to the manufacturer’s protocols.
13 C-glucose tracing experiments
U-[13 C]-glucose tracing was performed to analyze metabolic flux in iAMs. Briefly, 1 × 10⁷ cells were seeded in 10-cm dishes and infected with either live PAO1 or heat-killed PAO1 (inactivated at 65 °C for 1 h) for 1 h. The culture medium was then replaced with glucose- and serine-free DMEM containing 25 mM U-[13 C]-glucose (Cambridge Isotope Laboratories) and 50 μg/ml gentamicin. After 6 h of tracer incubation, cells were washed three times with ice-cold saline. The resulting cell pellets were dried and stored at −80 °C until analysis. All samples were subjected to liquid chromatography coupled with triple-quadrupole mass spectrometry (UPLC-TQ-MS) at Metabo-Profile Biotechnology Co., Ltd (China).
Quantitative PCR and RNA-Seq analysis
Total RNA was extracted using RNAiso Plus (Takara) and reverse transcribed into cDNA using the Evo M-MLV Reverse Transcription System (Accurate Biology) for quantitative RT-PCR analysis. RT-PCR was performed on an Applied Biosystems instrument using SYBR Green PCR Mix (Accurate Biology) with gene-specific primers. The relative expression of target genes was calculated using the 2 − ΔΔCtmethod and normalized to the expression of the housekeeping gene β-actin in each sample. For RNA-Seq analysis, total RNA was extracted from iAMs treated with siNC and siPhgdh, serine deprivation and serine stimulation. RNA sequencing was performed on an Illumina Novaseq 6000 platform. Transcription data were analyzed according to standard protocols (Oebiotech, Shanghai). Differentially expressed genes (DEGs) with a log2 (fold change) > 0.5 and a false discovery rate (FDR) < 0.005 were considered statistically significant. KEGG pathway enrichment analysis and GSEA analysis were conducted to identify the pathways associated with the DEGs.
Bisulfite sequencing PCR (BSP) and chromatin immunoprecipitation followed by quantitative PCR (ChIP-qPCR)
DNA methylation status of the Dusp4 promoter was analyzed by BSP. Genomic DNA was treated with the EpiArt® DNA Methylation Bisulfite Conversion Kit (EM112-01) to convert unmethylated cytosines to uracils, followed by purification according to the manufacturer’s instructions. PCR amplification was performed using bisulfite-converted DNA as template, EpiArt™ HS Taq DNA Polymerase (EM202-01), and specific BSP primers. The PCR products were purified using the corresponding kit (DC112-01), ligated into a T-vector (C603-01), and transformed into competent cells (C505-02) to construct a clone library. At least five positive clones were selected for sequencing, and methylation status of individual CpG sites was analyzed by sequence alignment. The enrichment of H3K27me3 at the Dusp4 promoter in iAMs was quantified using a Hyperactive pA/G-MNase CUT&RUN Assay Kit for PCR/qPCR (HD103-01). Mouse IgG was used as a negative control for immunoprecipitation. Specific promoter regions of the target gene were amplified by qPCR, while the Spike gene was amplified with corresponding primers as an internal negative control. The primers are listed in Supplementary Table 1.
Western blot analysis
Cells (106) were harvested, washed with PBS, and lysed using RIPA buffer. Protein concentrations were detected using a bicinchoninic acid (BCA) protein assay kit (Epizyme) according to the manufacturer’s instructions. Proteins were separated by SDS-PAGE electrophoresis and transferred to a PVDF membrane (pore size 0.2μm; Merck Millipore). The membranes were blocked with 5% non-fat milk in Tris-Tween-buffered saline (TBST) and subsequently incubated with primary and secondary antibodies. Primary antibodies used in this study include Phgdh (Proteintech, Cat. No. 14719-1-AP, 1:1000), Phospho-p44/42 MAPK (ERK1/2) (Thr202/Tyr204) (CST, Cat. No. 4370 T, 1:1000), p44/42 MAPK (ERK1/2) Antibody (CST, Cat. No. 9102S, 1:1000), Phospho-p38 MAPK (Thr180/Tyr182) Antibody (CST, Cat. No. 9211S, 1:1000), p38 MAPK Antibody (CST, Cat. No. 9212S, 1:1000), p-JNK (Santa, Cat. No. sc-6254, 1:500), JNK (Santa, Cat. No. sc-7345, 1:500), MKP-2/DUSP4 (Proteintech, Cat. No. 14719-1-AP, 1:1000), DUSP2 (Proteintech, Cat. No. 27327-1-AP, 1:1000), Histone H3 (tri methyl K27) (Abcam, Cat. No. ab6002, 1:1000), Histone H3 (Abcam, Cat. No. ab201456, 1:1000), iNOS (Proteintech, Cat. No. 18985-1-AP, 1:2500), Arginase-1 (Proteintech, Cat. No. 16001-1-AP, 1:1000), Recombinant Anti-GAPDH antibody (HRP Conjugated) (Servicebio, Cat. No. ZB15004-HRP-100, 1:2500), β-Actin Antibody (CST, Cat. No. 4967S, 1:1000), and HSP90 alpha/beta (Santa, Cat. No. sc-13119, 1:1000). Primary antibody binding was detected using anti-mouse IgG, HRP-linked Antibody (CST, Cat. No. 7076S, 1:5000) or anti-rabbit IgG, HRP-linked Antibody (CST, Cat. No. 7074S, 1:5000) and visualized by chemiluminescence. The expression levels of target proteins were normalized to β-actin, GAPDH or HSP90α/β as internal controls.
Tissue histological analysis
Lung samples were fixed with 4% paraformaldehyde for 24 h, which dehydrated in graded ethanol and embedded in paraffin afterwards. Samples were stained with hematoxylin and eosin (H&E) before histopathological examinations. According to the guidelines for lung injury pathology scoring proposed by the American Thoracic Society, 20 random fields of various lung tissues were selected under 10× and 20× magnification. Scoring was performed for each field according to the scoring criteria. Finally, the lung injury score was calculated using the following formula: Score = [(20×A) + (14×B) + (7×C) + (7×D) + (2×E)] / (20×100).
Immunohistochemistry and Immunofluorescence
For immunohistochemistry, paraffin-embedded lung tissue sections from mice were stained with Phgdh antibodies (Proteintech, 14719-1-AP, 1:800). The tissue sections were quantitatively analyzed using ImageJ software based on the percentage of positive cells and staining intensity. For immunofluorescence, cell samples were fixed with 4% paraformaldehyde for 20 min at room temperature. Fixed cells and tissue sections were blocked with QuickBlock™ blocking buffer (Beyotime) at room temperature for 30 minutes. The samples were then incubated with primary antibodies, including Phgdh (Proteintech, Cat. No. 14719-1-AP, 1:800), F4/80 (CST, Cat. No. 70076, 1:500) and SiglecF (BD, Cat. No. 746668, 1:100), overnight at 4 °C. Afterward, the samples were incubated with appropriate secondary antibodies for 1 h at room temperature, followed by staining with Hoechst or DAPI for 5 min at room temperature. Stained cells and tissues were visualized using a fluorescence microscope and a confocal fluorescence microscope (Zeiss).
Assessment of total protein in lung tissue
Total protein concentrations in lung tissue were measured using a BCA Protein Assay Kit according to the manufacturer’s instructions (Epizyme). Absorbance was quantified using a Synergy LX multi-mode reader (BioTek).
Flow cytometry
Multiparameter assessment and cell sorting were conducted using a BD Fortessa flow cytometer (BD Biosciences), with data analysis performed using FlowJo software (TreeStar). Single-cell suspensions were first incubated with TruStain FcX™ PLUS (anti-mouse CD16/32) to block FcγIII/II receptors. Subsequently, cells were stained with fluorochrome-conjugated antibodies in fluorescence-activated cell sorting (FACS) buffer (PBS containing 1% FBS and 2 mM EDTA) and washed twice prior to detection. The following monoclonal antibodies specific to mouse markers were used: APC/Cyanine7 anti-mouse CD45.2 (BioLegend, Cat. No. 109824, 1:500), PE-Cy™7 Rat Anti-Mouse Ly-6G (BD Pharmingen, Cat. No. 560601, 1:300), FITC anti-mouse F4/80 Antibody (BioLegend, Cat. No. 123107, 1:100), Brilliant Violet 421™ anti-mouse CD64 (FcγRI) Antibody (BioLegend, Cat. No. 139309, 1:100), Brilliant Violet 605™ anti-mouse/human CD11b (BioLegend, Cat. No.101257, 1:200), PE anti-mouse CD11c Antibody (BioLegend, Cat. No. 117307, 1:100), FITC anti-mouse CD170 (Siglec-F) Antibody (BioLegend, Cat. No.155503, 1:100), BB515 Rat Anti-CD11b (M1/70) (BD Pharmingen, Cat. No.564454, 1:200), BB700 Rat Anti-Mouse F4/80 (T45-2342) (BD Pharmingen, Cat. No.746070, 1:200), PE anti-mouse CD86 (BioLegend, Cat. No. 105008, 1:100), Brilliant Violet 421™ anti-mouse CD206 (MMR) (BioLegend, Cat. No. 141717, 1:100), Brilliant Violet 650™ anti-mouse CD11c (BioLegend, Cat. No.117339, 1:200), APC anti-mouse CD170 (Siglec-F) Antibody (BioLegend, Cat. No. 155508, 1:100), APC anti-mouse CD206 (MMR) Antibody (BioLegend, Cat. No. 321109, 1:100). Dead cells were excluded using the live/dead marker Fixable Viability Stain 510 (FVS510) (BD Biosciences, Cat. No. 564406, 1:500). For intracellular staining of CD206, cells were fixed and permeabilized using flow cytometry fixation/permeabilization concentrate, diluent, and buffer (eBioscience).
Adoptive transfer
AMs were isolated from the BALF of Phgdhfl/flLyz2Cre and Phgdhfl/fl mice and cultured in L-serine free RPMI-1640 medium supplemented with 10% fetal calf serum and 1% penicillin-streptomycin. After 2 h of adherence, non-adherent cells were removed by washing with warm media. Following 20 days of culture, 2 × 105 AMs were resuspended in 50 μL of sterile PBS and intranasally (i.n.) transferred into wild-type (WT) mice under isoflurane anesthesia. BMDMs were obtained from the bone marrow of Phgdhfl/flLyz2Cre and Phgdhfl/fl mice and cultured in L-serine-free DMEM medium supplemented with 10% FBS and 1% penicillin-streptomycin. After 7 days of culture, 5 × 105 BMDMs were resuspended in 200 μL of sterile PBS and intravenously (i.v.) transferred into WT mice.
Statistical analyses and reproducibility
All experiments were performed at least three biological and technical replicates. Sample size of biological replicates was selected as at least three to examine experimental variations. No data were excluded for analyses. Western blot assays, immunohistochemical staining, immunofluorescence staining and DNA agarose gel blot representative images are shown. Statistical analyses were performed using GraphPad Prism 9.0 software. Data were analyzed using two-tailed Student’s t-test or one-way ANOVA with Dunnett’s multiple comparison test. Survival curves were compared using the log-rank test. Results are presented as means ± standard error of the mean (SEM). A p-value < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw RNA-seq data generated in this study have been deposited in the NCBI Sequence Read Archive database under the accession code PRJNA1358829 and PRJNA1357662. All data needed to evaluate the conclusions are present in the article, the supplementary materials and source data file provided with this paper. Source data are provided with this paper.
References
Qin, S. et al. Pseudomonas aeruginosa: pathogenesis, virulence factors, antibiotic resistance, interaction with host, technology advances and emerging therapeutics. Signal Transduct. Target Ther. 7, 199 (2022).
McIsaac, S. M., Stadnyk, A. W. & Lin, T. J. Toll-like receptors in the host defense against Pseudomonas aeruginosa respiratory infection and cystic fibrosis. J. Leukoc. Biol. 92, 977–985 (2012).
Kupz, A. et al. NLRC4 inflammasomes in dendritic cells regulate noncognate effector function by memory CD8⁺ T cells. Nat. Immunol. 13, 162–169 (2012).
Sutterwala, F. S. et al. Immune recognition of Pseudomonas aeruginosa mediated by the IPAF/NLRC4 inflammasome. J. Exp. Med. 204, 3235–3245 (2007).
Cohen, T. S. & Prince, A. S. Activation of inflammasome signaling mediates pathology of acute P. aeruginosa pneumonia. J. Clin. Investig. 123, 1630–1637 (2013).
Lehman, H. K. & Segal, B. H. The role of neutrophils in host defense and disease. J. Allergy Clin. Immunol. 145, 1535–1544 (2020).
Faure, E. et al. Pseudomonas aeruginosa type-3 secretion system dampens host defense by exploiting the NLRC4-coupled inflammasome. Am. J. Respir. Crit. Care Med. 189, 799–811 (2014).
Hoyer, F. F. et al. Tissue-specific macrophage responses to remote injury impact the outcome of subsequent local immune challenge. Immunity 51, 899–914.e897 (2019).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Martin, F.P., Jacqueline, C., Poschmann, J. & Roquilly, A. Alveolar macrophages: adaptation to their anatomic niche during and after inflammation. Cells 10, 2720 (2021).
Wang, Q. et al. O-GlcNAc transferase promotes influenza A virus-induced cytokine storm by targeting interferon regulatory factor-5. Sci. Adv. 6, eaaz7086 (2020).
Shen, B. et al. Proteomic and metabolomic characterization of COVID-19 patient sera. Cell 182, 59–72.e15 (2020).
Xiao, N. et al. Integrated cytokine and metabolite analysis reveals immunometabolic reprogramming in COVID-19 patients with therapeutic implications. Nat. Commun. 12, 1618 (2021).
Van den Bossche, J., O’Neill, L. A. & Menon, D. Macrophage immunometabolism: Where are we (Going)? Trends Immunol. 38, 395–406 (2017).
Rosenberg, G., Riquelme, S., Prince, A. & Avraham, R. Immunometabolic crosstalk during bacterial infection. Nat. Microbiol. 7, 497–507 (2022).
Rath, M., Müller, I., Kropf, P., Closs, E. I. & Munder, M. Metabolism via arginase or nitric oxide synthase: two competing arginine pathways in macrophages. Front. Immunol. 5, 532 (2014).
Ren, W. et al. Glutamine metabolism in macrophages: a novel target for obesity/type 2 diabetes. Adv. Nutr. 10, 321–330 (2019).
Liu, P. S. et al. α-ketoglutarate orchestrates macrophage activation through metabolic and epigenetic reprogramming. Nat. Immunol. 18, 985–994 (2017).
Mao, Y., Shi, D., Li, G. & Jiang, P. Citrulline depletion by ASS1 is required for proinflammatory macrophage activation and immune responses. Mol. Cell 82, 527–541.e527 (2022).
Pacold, M. E. et al. A PHGDH inhibitor reveals coordination of serine synthesis and one-carbon unit fate. Nat. Chem. Biol. 12, 452–458 (2016).
Fan, J. et al. Human phosphoglycerate dehydrogenase produces the oncometabolite D-2-hydroxyglutarate. ACS Chem. Biol. 10, 510–516 (2015).
Ma, C. et al. The alternative activity of nuclear PHGDH contributes to tumour growth under nutrient stress. Nat. Metab. 3, 1357–1371 (2021).
Shen, L. et al. Serine metabolism antagonizes antiviral innate immunity by preventing ATP6V0d2-mediated YAP lysosomal degradation. Cell Metab. 33, 971–987.e976 (2021).
Jiang, L. et al. Salmonella Typhimurium reprograms macrophage metabolism via T3SS effector SopE2 to promote intracellular replication and virulence. Nat. Commun. 12, 879 (2021).
Bojkova, D. et al. Identification of novel antiviral drug candidates using an optimized SARS-CoV-2 phenotypic screening platform. iScience 26, 105944 (2023).
Yu, W. et al. One-Carbon Metabolism Supports S-Adenosylmethionine and Histone Methylation to Drive Inflammatory Macrophages. Mol. Cell 75, 1147–1160.e1145 (2019).
Rodriguez, A. E. et al. Serine metabolism supports macrophage IL-1β production. Cell Metab. 29, 1003–1011.e1004 (2019).
Schaum, N. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Gaddis, N. et al. LungMAP portal ecosystem: systems-level exploration of the lung. Am. J. Respir. Cell Mol. Biol. 70, 129–139 (2024).
Guo, M. et al. Guided construction of single cell reference for human and mouse lung. Nat. Commun. 14, 4566 (2023).
Li, F. et al. Monocyte-derived alveolar macrophages autonomously determine severe outcome of respiratory viral infection. Sci. Immunol. 7, eabj5761 (2022).
Mullarky, E. et al. Identification of a small molecule inhibitor of 3-phosphoglycerate dehydrogenase to target serine biosynthesis in cancers. Proc. Natl. Acad. Sci. USA. 113, 1778–1783 (2016).
Zhang, X. et al. PHGDH/SYK: a hub integrating anti-fungal immunity and serine metabolism. Cell Death Differ. 31, 1664–1678 (2024).
Du, C. et al. Mitochondrial serine catabolism safeguards maintenance of the hematopoietic stem cell pool in homeostasis and injury. Cell Stem Cell 31, 1484–1500.e1489 (2024).
Low, H. B. & Zhang, Y. Regulatory roles of MAPK phosphatases in cancer. Immune Netw. 16, 85–98 (2016).
Lu, Z. et al. MiR-122-5p protects against acute lung injury via regulation of DUSP4/ERK signaling in pulmonary microvascular endothelial cells. Life Sci. 256, 117851 (2020).
Jiao, H. et al. DUSP4 modulates RIG-I- and STING-mediated IRF3-type I IFN response. Cell Death Differ. 31, 280–291 (2024).
Al-Mutairi, M. S. et al. MAP kinase phosphatase-2 plays a critical role in response to infection by Leishmania mexicana. PLoS Pathog. 6, e1001192 (2010).
Ampomah, P. B. et al. Macrophages use apoptotic cell-derived methionine and DNMT3A during efferocytosis to promote tissue resolution. Nat. Metab. 4, 444–457 (2022).
Williams, K. T. & Schalinske, K. L. Homocysteine metabolism and its relation to health and disease. Biofactors 36, 19–24 (2010).
Parker, D., Ahn, D., Cohen, T. & Prince, A. Innate immune signaling activated by MDR bacteria in the airway. Physiol. Rev. 96, 19–53 (2016).
Kelly, B. & Pearce, E. L. Amino assets: how amino acids support immunity. Cell Metab. 32, 154–175 (2020).
Yeramian, A. et al. Arginine transport via cationic amino acid transporter 2 plays a critical regulatory role in classical or alternative activation of macrophages. J. Immunol. 176, 5918–5924 (2006).
Yurdagul, A. Jr. et al. Macrophage metabolism of apoptotic cell-derived arginine promotes continual efferocytosis and resolution of injury. Cell Metab. 31, 518–533.e510 (2020).
Jha, A. K. et al. Network integration of parallel metabolic and transcriptional data reveals metabolic modules that regulate macrophage polarization. Immunity 42, 419–430 (2015).
Palmieri, E. M. et al. Nitric oxide orchestrates metabolic rewiring in M1 macrophages by targeting aconitase 2 and pyruvate dehydrogenase. Nat. Commun. 11, 698 (2020).
Pang, Z., Raudonis, R., Glick, B. R., Lin, T. J. & Cheng, Z. Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 37, 177–192 (2019).
Iannitti, R. G. et al. IL-1 receptor antagonist ameliorates inflammasome-dependent inflammation in murine and human cystic fibrosis. Nat. Commun. 7, 10791 (2016).
Lear, T.B. et al. KIAA0317 regulates pulmonary inflammation through SOCS2 degradation. JCI Insight 4, e129110 (2019).
Geeraerts, S. L., Heylen, E., De Keersmaecker, K. & Kampen, K. R. The ins and outs of serine and glycine metabolism in cancer. Nat. Metab. 3, 131–141 (2021).
Ma, E. H. et al. Serine is an essential metabolite for effector T cell expansion. Cell Metab. 25, 345–357 (2017).
Murphy, J. P. et al. The NAD(+) salvage pathway supports PHGDH-driven serine biosynthesis. Cell Rep. 24, 2381–2391.e2385 (2018).
Gao, X. et al. Serine availability influences mitochondrial dynamics and function through lipid metabolism. Cell Rep. 22, 3507–3520 (2018).
Kurita, K. et al. Macrophages rely on extracellular serine to suppress aberrant cytokine production. Sci. Rep. 11, 11137 (2021).
de Goede, K. E. et al. d-2-Hydroxyglutarate is an anti-inflammatory immunometabolite that accumulates in macrophages after TLR4 activation. Biochim. Biophys. Acta Mol. Basis Dis. 1868, 166427 (2022).
Wilson, J. L. et al. Inverse data-driven modeling and multiomics analysis reveals Phgdh as a metabolic checkpoint of macrophage polarization and proliferation. Cell Rep. 30, 1542–1552.e1547 (2020).
Raines, L. N. et al. PERK is a critical metabolic hub for immunosuppressive function in macrophages. Nat. Immunol. 23, 431–445 (2022).
Chen, S. et al. Serine supports IL-1β production in macrophages through mTOR signaling. Front. Immunol. 11, 1866 (2020).
Han, W. et al. Enhanced expression of catalase in mitochondria modulates NF-κB-dependent lung inflammation through alteration of metabolic activity in macrophages. J. Immunol. 205, 1125–1134 (2020).
Cargnello, M. & Roux, P. P. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol. Mol. Biol. Rev. 75, 50–83 (2011).
Gantke, T., Sriskantharajah, S., Sadowski, M. & Ley, S. C. IκB kinase regulation of the TPL-2/ERK MAPK pathway. Immunol. Rev. 246, 168–182 (2012).
Viatour, P., Merville, M. P., Bours, V. & Chariot, A. Phosphorylation of NF-kappaB and IkappaB proteins: implications in cancer and inflammation. Trends Biochem. Sci. 30, 43–52 (2005).
Lu, Z. & Xu, S. ERK1/2 MAP kinases in cell survival and apoptosis. IUBMB Life 58, 621–631 (2006).
Acknowledgements
We express our gratitude to all members of the Tabula Muris Consortium for generously sharing their single-cell transcriptomics data of 100,000 cells from 20 organs and tissues. We also extend our appreciation to biorender.com for their figure drawing materials. This work was supported by National Key Research and Development Program of China (2023ZD0506200 to J.Q.), National Natural Science Foundation of China (No.82470002 to Y. L and No.82200006 to M.S), Shanghai Municipal Key Clinical Specialty (shslczdzk02202 to J.Q.), Shanghai Top-Priority Clinical Key Disciplines Construction Project (2017ZZ02014 to J.Q.), and Shanghai Key Laboratory of Emergency Prevention, Diagnosis and Treatment of Respiratory Infectious Diseases (20dz2261100 to J.Q.).
Author information
Authors and Affiliations
Contributions
The design, methodology, experiments, data analysis and initial draft were conducted by R.C., R.Z., Y.L. and J.Q. Review, funding acquisition and supervision were performed by Y.L. and J.Q. M.S., G.Z., F.W., Q.Y., Y.J., and Y.Z. performed in vivo experiments and statistical analyses. All authors participated in the preparation of the article and approved the final version for submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sebastian Riquelme and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, R., Zeng, R., Shi, M. et al. Phosphoglycerate dehydrogenase-mediated serine reprogramming aggravates macrophage hyperinflammation in murine Pseudomonas aeruginosa pneumonia. Nat Commun 17, 1944 (2026). https://doi.org/10.1038/s41467-026-69539-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69539-1