Abstract
HORMAD1 expression is usually restricted to germ-line cells but is also aberrantly expressed in 60% of triple-negative breast cancers (TNBCs), where it is bi-modally expressed and associated with genomic instability. Here, we show that out-of-context HORMAD1 expression in mitotic cells perturbs mitotic arrest and generates aneuploidy. These phenotypes are caused by out-of-context HORMAD1 expression driving a weakening of the spindle assembly checkpoint (SAC) and/or in kinetochore-microtubule error correction. These mitotic effects of HORMAD1 are MAD2L1-independent, but instead caused by a HORMAD1/Aurora B interaction and defective Aurora B/INCENP signalling. Consistent with this mechanism, aberrant HORMAD1 expression causes sensitivity to MPS1, Aurora B or BUB1 inhibitors currently being investigated as cancer treatments. Our data suggests how out-of-context expression of a meiotic gene imparts genomic instability upon tumour cells and also identifies several associated dependencies as mechanism-based therapeutic targets for a large, biomarker-defined, subset of cancers.
Similar content being viewed by others
Introduction
Triple-negative breast cancers (TNBC) represent a sub-group of breast cancers that lack ER, PR and HER2-positivity, and which therefore do not respond to endocrine treatments or conventional anti-HER2 therapies1. Although targeted therapy is available for TNBC patients with either BRCA1/2 mutations2,3 or PD-L1 expressing lymphocytic infiltrates4, prognosis is often poor owing to the lack of recurrent targetable mutations in the disease1.
HORMAD1 is a meiotic recombination promoting gene whose expression is usually restricted to germ-line cells5, where it has effects on the regulation of unsynapsed chromatin, DNA strand breaks, homologous recombination (HR) template choice, and meiotic cell cycle checkpoints6. In addition to its meiotic functions, out-of-context expression of HORMAD1 is detected in 60% of TNBCs, where it is associated with signatures of genomic instability7. We previously found that this out-of-context HORMAD1 expression caused genomic instability in breast cancer and mouse embryonic stem (ES) cell lines, at least in part through effects on non-homologous end-joining (NHEJ) and Rad51-mediated homologous recombination (HR)7. The effects of HORMAD1 on HR appear context-dependent and others have found evidence that HORMAD1 can enhance HR8,9,10,11.
HORMAD1 is part of a family of HORMA domain containing proteins. Additional HORMA domain-containing proteins, MAD2L1 and P31COMET, are key regulators of the spindle assembly checkpoint (SAC)12, which functions in mitotic cells to delay the onset of anaphase until all microtubules are correctly attached to kinetochores13. To promote bipolar (amphitelic) spindle attachments and ensure correct chromosome segregation, improper microtubule attachments must be corrected. The process of error correction involves the phosphorylation of kinetochore proteins that bind microtubules to modify the stability or affinity of the kinetochore-microtubule interaction. Through error correction, microtubules detach from the kinetochore allowing new rounds of attachments until all are amphitelic. Detaching the microtubules from kinetochores generates a SAC signal, which delays anaphase until amphitelic attachment is achieved; this activation of the SAC requires a number of mitotic kinases including MPS1 and Aurora B. MPS1 is the master regulator of SAC activity, and is required to activate and maintain the SAC by phosphorylating a number of MELT motifs on the KMN network of proteins that are an essential component of the kinetochore14. Aurora B is the active subunit of the chromosome passenger complex (CPC)15 and is a key component of the error-correction pathway16,17,18. Aurora B has also been shown to regulate the SAC directly by promoting MPS1 activation16. A third kinase, BUB1, has also been shown to regulate MPS1 and Aurora B activity during mitosis19,20.
Defects in the SAC and error correction are thought to drive cancer progression21. Based on this premise, a number of small molecule kinase inhibitors, including those targeting MPS1, Aurora B or BUB1 are being assessed as cancer treatments in either pre-clinical or clinical studies (e.g., NCT03328494, an MPS1 inhibitor combination trial)22,23,24.
Here, we demonstrate that the aberrant expression of HORMAD1 in tumour cells perturbs the ability of cells to delay mitosis in response to improperly attached kinetochores via a mechanism that is independent of MAD2L1. HORMAD1 exerts these effects by binding to Aurora B, disrupting its normal association with its co-factor, INCENP, and impairing its ability to phosphorylate substrates. This, in turn, impairs the error correction and/or the strength of the SAC. Consistent with this mechanism, aberrant expression of HORMAD1 drives cellular sensitivity to either MPS1, BUB1 or Aurora kinase inhibitors. Taken together, these data suggest that the out-of-context expression of HORMAD1 in tumour cells drives a weakening of the processes that control mitotic fidelity, which contributes to chromosomal instability and induces dependency on a number of clinically relevant therapeutic targets. As such, tumoral HORMAD1 expression has potential as a patient selection biomarker for synthetic lethal sensitivity to Aurora B, MPS1 or BUB1 inhibitors. Indeed, we show that three patient derived breast cancer xenograft models expressing HORMAD1 are sensitive to Aurora B inhibitor AZD2811 in vivo.
Results
HORMAD1 expression drives chromosomal instability in mitotic cells
HORMAD1 is expressed in a number of solid tumour subtypes, including TNBC, where we previously demonstrated that the out-of-context expression of this meiotic gene is associated with genomic instability7. Among TNBC, HORMAD1 expression is enriched in homologous recombination defective (HRD + ) tumours, which include most BRCA1-mutated breast cancers (Supplementary Fig. 1A, B). ER+ and HER2+ breast cancers generally do not express HORMAD1, and where it is expressed this is not associated with BRCA2 mutation (Supplementary Fig. 1C). In order to understand the mechanisms by which HORMAD1 drives genomic instability, we created an isogenic RPE1 cell line expressing a doxycycline-inducible HORMAD1 cDNA fused to a 3’-terminal HA epitope tag coding sequence. RPE1 cells are non-transformed, and have low basal levels of chromosomal instability and a stable karyotype, and have been widely used as a model system for perturbations that increase chromosomal instability25,26,27. Doxycycline-induced HORMAD1 protein expression was comparable to the level of HORMAD1 expression observed in MDA-MB-436 cells, a breast cancer cell line that naturally expresses HORMAD1 (Fig. 1A). HORMAD1 expression in RPE1 cells led to a decrease in the viable cell number (Fig. 1B).
A Inducible HORMAD1 expression in RPE1 TRE2-HA-HORMAD1 exposed to doxycycline or vehicle (water, –DOX) for three days and analysed by western blotting using indicated antibodies. B RPE1 TRE2-HA-HORMAD1 cells were exposed to doxycycline or vehicle (water, –DOX). 5 days post transfection, cellular viability was estimated using Cell Titre Glo (CTG) reagent and surviving fractions were calculated relative to DMSO treated cells. n = 6 technical replicates, p < 0.0001, t-test (two sided). C Segregation errors in RPE1 TRE2-HA-HORMAD1 cells marked with H2B-GFP. Cells were exposed to doxycycline for 3 days and analysed by time-lapse microscopy. Total segregation errors of each type are shown, scored from at least 500 anaphase cells, across 3 biological repeats. Bridge p = 0.998, Lagging p = 0.0414 and Multipolar p = 0.0128; pairwise t-tests with Bonferroni correction (two sided). D RPE1 TRE2-HA-HORMAD1 cells were exposed to doxycycline for 3 days and anti-ACA antibody (Centromere, green) used to distinguish centric and acentric micronuclei. E Centric/acentric micronuclei were quantified from >1000 cells in (D). The % micronuclei was calculated and presented as the median from 4 biological replicates. Acentric p = 0.2468 and centric p = 0.0062, pairwise t-test with a Bonferroni correction (two sided). F Induction of micronuclei in SUM159 TRE2-HA-HORMAD1 cells after doxycycline exposure for 7 days. G Quantification of experiment shown in F. At least 511 cells were scored, across 3 biological repeats. p = 0.0197, pairwise t-test (two sided). H Induction of micronuclei following doxycycline-induced HORMAD1 expression in SUM149 cells after 7 days. At least 145 cells were scored, across 3 biological repeats. p = 0.0118, pairwise t-test (two sided). I Induction of micronuclei following doxycycline-induced HORMAD1 expression in MCF10A cells after 7 days. At least 394 cells were scored, across 3 biological repeats. p = 0.0001, pairwise t-test (two sided). J Increased aneuploidy in inducible HORMAD1 expressing RPE1 cells. Cells were exposed to doxycycline for 3 days, and metaphase spreads generated. K Percentage of cells with abnormal number of chromosomes was counted from at least 100 cells per condition across 4 biological replicas from the experiment in panel J. p = 0.0229, pairwise t-test (two sided). L Tumour types with a positive association between HORMAD1 expression and aneuploidy (Data from TCGA). Aneuploid genomic signatures from tumour sequencing data, described previously29, were dichotomised based on their HORMAD1 mRNA expression status. P values calculated with Mann-Whitney-U test (two sided). Box plots show the first quartile (Q1), median and third quartile (Q3), with whiskers extended to points closest to Q1 − 1.5x the IQR (interquartile range) and Q3 + 1.5x the IQR. The following abbrevations are used; Breast Cancer (BRCA), Lung Adenocarcinoma (LUAD), Ovarian Serous Adenocarcinoma (OV), Sarcoma (SARC), Stomach Adenocarcinoma (STAD), Testicular Germ Cell Tumour (TGCT), Uterine Corpus Endometrioid Carcinoma (UCEC). Source data are provided as a Source Data file.
We first investigated the impact of HORMAD1 on chromosomal segregation, using time-lapse microscopy in HORMAD1 inducible RPE1 cells expressing GFP-tagged histone H2B. In cells not expressing HORMAD1, a small percentage of cells displayed chromatin bridges, lagging chromosomes, or multipolar spindles during anaphase. The expression of HORMAD1 caused a significant increase in lagging chromosomes and multipolar spindles but did not appear to increase the levels of chromatin bridges (Fig. 1C). To better understand the nature of these segregation errors, we stained centromeres in RPE1 cells with anti-ACA antibodies and assessed whether HORMAD1 expression induced centric or acentric micronuclei. HORMAD1 expression induced a significant increase in micronuclei containing centromeres (Fig. 1D, E), indicating defects in mitotic segregation. We also observed increased levels of micronuclei upon HORMAD1 expression in a number of breast cancer cell line models, (SUM159, SUM149) and the breast epithelial cell line MCF10A (Fig. 1F–I). This led us to ask whether HORMAD1 expression in mitotic cells would lead to alterations in chromosome number (aneuploidy). To address this, we quantified chromosome numbers by metaphase analysis of RPE1 cells expressing doxycycline inducible HORMAD1. HORMAD1 expression caused increased levels of aneuploidy (Fig. 1J, K). We concluded from these experiments that aberrant HORMAD1 expression in mitotic cells can lead to segregation errors, structural and numerical chromosomal instability, and aneuploidy.
Given our observations in isogenic model systems, we assessed whether HORMAD1 expression was associated with high levels of aneuploidy28 in cancer. To assess this, we used a previously published data set that estimated levels of aneuploidy using DNA sequence data from a wide variety of human tumours (The Cancer Genome Atlas; TGCA)29. Using this and publicly available RNA sequencing data from the same tumour samples, we dichotomised samples from each cancer subtype based on their HORMAD1 expression status. We found that HORMAD1 was expressed in significant subsets of many cancer histological subtypes, and was associated with higher levels of aneuploidy in a number of tumour types (Fig. 1L and Supplementary Fig. 2). The increased aneuploidy index associated with HORMAD1 expression in cancers (Fig. 1L) was consistent with prior functional experiments that show that induced expression of HORMAD1 causes a similar form and degree of genomic instability and aneuploidy as seen in HORMAD1-positive breast cancers7,30. We note that some HORMAD1-negative cancers also show evidence of aneuploidy, which may be driven by other features such as loss of function of p53, RB or p16.
HORMAD1 impairs mitotic arrest in response to spindle poisons
In the absence of full amphitelic kinetochore attachments, mitotic arrest is mediated by the SAC; this process is critical to prevent premature sister chromatid separation and mitotic exit, and works by preventing the ubiquitin-mediated destruction of securin and Cyclin B. Exposing mitotic cells to nocodazole or paclitaxel, drugs that depolymerise or stabilise microtubules respectively, causes SAC-dependent mitotic arrest. The SAC cannot block all Cyclin B proteolysis however, and prolonged mitotic arrest eventually leads to exit from mitosis without proper chromosome segregation28, a process known as mitotic slippage31.
Like HORMAD1, MAD2L1 and P31COMET are HORMA domain proteins that regulate the SAC via HORMA domain-mediated protein interactions12. Since HORMAD1 controls cell cycle checkpoints in meiosis6, we wondered whether dysregulated HORMAD1 expression might also affect the SAC in mitosis. Cells with a defective SAC are unable to maintain nocodazole or paclitaxel-induced mitotic arrest for prolonged periods of time and undergo mitotic slippage sooner than cells with a fully functional SAC32. We used our RPE1 model to measure the impact of HORMAD1 expression on the duration of mitotic arrest (Fig. 2A). These experiments revealed that HORMAD1 expression significantly decreased the time taken to exit mitosis via mitotic slippage after exposure to either nocodazole or paclitaxel (Fig. 2B, C). A similar phenotype is observed when SAC components are perturbed28,33,34. These data indicated that HORMAD1 perturbs the ability of cells to delay in mitosis in response to improperly attached kinetochores; either by weakening the spindle assembly checkpoint or by impairing error correction and the associated SAC activation.
A Experiment outline. B RPE1 TRE2-HA-HORMAD1 were assessed for mitotic slippage as shown in (A). The mean percentage of cells arrested in mitosis at each time point is shown -/+ SEM. A total of 90 cells from 3 biological repeats were analysed. p = 0.0001 (HORMAD1) and p = 0.1563 (n.s., empty), Mantel-Cox test (two sided). C As B but cells were exposed to 100 nM paclitaxel. A total of 125 cells from 3 biological repeats was analysed. p = 0.0001, Mantel-Cox test (two sided). D Western blot illustrating HORMAD1 depletion in H1299 cells expressing two individual shRNA constructs that target HORMAD1 or nontargeting control shRNA (NT). Cells were grown for three days in the presence of doxycycline. E Curve indicating delayed mitotic slippage following nocodazole exposure in H1299 cells depleted of HORMAD1. H1299 cells with shRNA constructs as in (D) were grown for three days in the presence of doxycycline at which point cells were exposed to 300 nM nocodazole and analysed as in (B). A total of 90 cells from 3 biological repeats were analysed. p = 0.0022, 0.0038 for shRNA 1 and 2 respectively compared to NT, Mantel-Cox test with Bonferroni correction (two sided). F Experimental outline for assessing mitotic slippage in the presence of MPS1 inhibitor. G Enhanced mitotic slippage following nocodazole and reversine exposure in RPE1 TRE2-HA-HORMAD1. Cells were exposed to doxycycline or vehicle (water, –DOX) for three days, followed by nocodazole and reversine as in (F). A total of 150 cells from 3 biological repeats were analysed and plotted as in (B). 100 nM Reversine p < 0.0001 and 200 nM Reversine p < 0.0001, Mantel-Cox test with Bonferroni correction (two sided). H As (G) but cells exposed to MPS1 inhibitor BOS172722. p < 0.0001, Mantel-Cox test with Bonferroni correction (two sided). I RPE1 TRE2-HA-HORMAD1 were exposed to doxycycline or vehicle (water) and to 200 nM of the MPS1 inhibitor, BOS172722. The percentage of cells with centric or acentric micronuclei was quantified by immunofluorescence from at least 1000 cells across 3 biological replicates. p = 0.8362 (acentric) and p = 0.0296 (centric), t-test with Bonferroni correction (two sided). J H1299 cells expressing shRNA targeting HORMAD1 or a non-targeting control were analysed for mitotic slippage as in (F). Data reflects a total of 60 cells from 2 biological repeats. p = 0.0022, 0.0038 for shRNA1 and 2 respectively compared to NTC; Mantel-Cox test with Bonferroni correction (two sided). (A) and (F) Created in BioRender. Tutt, A.N.J. (https://BioRender.com/3qiwaov). Source data are provided as a Source Data file.
We investigated whether these effects were seen in HORMAD1-positive tumour cells. HORMAD1 has presumably been expressed in these cells through many cell cycles during tumour formation, and thus cells may have adapted their response to mitotic arrest. To exclude the possibility that the effects seen in RPE1 were private to experimentally-induced ectopic HORMAD1 expression, we depleted endogenous HORMAD1 in tumour cell lines and assayed mitotic arrest in response to nocodazole. HORMAD1 depletion significantly increased the duration of mitotic arrest in cervical cancer-derived HeLa cells using two different siRNAs (Supplementary Fig. 3A, B). Depleting HORMAD1 in the TNBC cell line MDA-MB-436 and the lung cancer cell line H1299 using either of two different doxycycline-inducible shRNAs (Fig. 2D, E; and Supplementary Fig. 3C, D) increased the duration of mitotic arrest in response to nocodazole (Fig. 2E).
Nocodazole exposure leads to very high levels of SAC stimulation, which can mask incomplete and more subtle underlying defects in the SAC28. Exposing cells to a low concentration of a small molecule MPS1 kinase inhibitor, such as reversine, leads to a partial weakening of the SAC16 and allows such underlying defects in the mitotic checkpoint to be observed35. We exposed nocodazole-arrested RPE1 cells expressing inducible HORMAD1 cDNA to reversine and measured the time taken for cells to exit mitosis (Fig. 2F). These experiments revealed that HORMAD1 expression exacerbated the effects of reversine on inducing mitotic slippage (Fig. 2G). We validated these results with an investigational agent MPS1 inhibitor, BOS172722 (Fig. 2H). The profound perturbation of the SAC driven by the combination of MPS1 inhibition and HORMAD1 expression made it likely that these cells would have defects in mitotic segregation, leading to the mis-segregation of whole chromosomes and formation of micronuclei. We therefore exposed RPE1 cells expressing HORMAD1 to the investigational agent MPS1 inhibitor, BOS172722, and measured the frequency of centric and acentric micronuclei after three days. Consistent with a prominent defect in the SAC and/or error correction and consequent chromosome missegration, HORMAD1 expression caused an increase in centric micronuclei that represent whole chromosomes that fail to segregate to daughter nuclei (Fig. 2I). We validated our observation that HORMAD1 cooperates with MPS1 inhibition to perturb mitotic arrest by expressing an inducible HA-tagged HORMAD1 transgene in MCF10A breast epithelial cells (Supplementary Fig. 3E, F), and an untagged HORMAD1 transgene in the TNBC cell line, SUM159 (both of which are otherwise HORMAD1-negative, Supplementary Fig. 3G, H). We then tested whether endogenous HORMAD1 affected mitotic slippage in cancer cells exposed to reversine. Depleting endogenous HORMAD1 from HeLa (cervical carcinoma), MDA-MB-436 (TNBC) and H1299 (non-small cell lung carcinoma) cell lines delayed mitotic slippage in the presence of reversine (Fig. 2J; Supplementary Fig. 3I-J, Supplementary Fig. 4A, B), indicating that long-term endogenous HORMAD1 expression in tumour cells also impairs mitotic arrest. These data indicated that HORMAD1-driven mitotic arrest defects do indeed persist in MDA-MB-436 cells, but require the more sensitive assay associated with partial MPS1 inhibition for this to be revealed. Importantly, re-expression of shRNA-resistant HORMAD1 cDNA in H1299 cells reversed the effect of HORMAD1 shRNA (Supplementary Fig. 4A, B), indicating this effect was indeed caused by HORMAD1 depletion. Taken together, these results demonstrated that HORMAD1 expression caused prominent defects in mitotic arrest when MPS1 is partially inhibited and that this effect is independent of the cell context in which HORMAD1 is expressed. From these data we, therefore concluded that HORMAD1 expression perturbs the ability of cells to arrest in mitosis because of partial abrogation of the SAC and/or incomplete error correction.
HORMAD1 does not profoundly disrupt MAD2L1 function
We sought to understand the mechanism underpinning HORMAD1-driven mitotic slippage. The HORMA domain containing protein, MAD2L1, is a key regulator of the SAC. Since HORMA domain containing proteins interact12, we hypothesised that HORMAD1 expression might disrupt the function of MAD2L1. A critical step in the activation of MAD2L1 is its recruitment to the kinetochore36, which leads to the formation of the mitotic checkpoint complex (MCC) that inhibits the APC/C (Fig. 3A). We generated an additional HORMAD1 inducible RPE1 cell line in which the endogenous MAD2L1 gene had been engineered to express MAD2L1 tagged with the Venus fluorescent protein at its N-terminus28 (Fig. 3B). We used immunocytochemistry to measure the recruitment of MAD2L1 to kinetochores (marked by CREST) in nocodazole-arrested and reversine-exposed RPE1 cells and found that HORMAD1 did not affect MAD2L1 recruitment to the kinetochore in nocodazole-treated cells (Fig. 3C, D). These experiments indicated that HORMAD1-driven mitotic slippage operates downstream of MAD2L1 recruitment or retention at the kinetochore.
A Diagram of MAD2L1-dependent SAC pathway. B RPE1 TRE2-HA-HORMAD1 expressing MAD2L1-Venus were grown for three days in the presence of doxycycline and analysed by western blot using indicated antibodies. C Immunofluorescence showing MAD2L1 localisation at kinetochores. RPE1 TRE2-HA-HORMAD1 expressing MAD2L1-YFP were exposed to doxycycline or vehicle for 3 days at which point cells were treated with 300 nM nocodazole for 3 h followed by 100 nM reversine for 1 h. Cells were fixed and stained with anti-ACA antibody. D Quantification of MAD2L1-positive kinetochores in (C). At least 50 cells per condition from 3 biological replicates, p = 0.9798, non-parametric one-way ANOVA with Kruksal-Wallis correction. E Experimental outline. F CDC20 immunoprecipitations (IP) illustrating assembly and stability of the MCC in HORMAD1 expressing RPE1 cells. RPE1 TRE2-HA-HORMAD1 cells were cultured as shown in (E). Cell lysates were generated, immunopurified using anti-CDC20 antibodies and analysed using SDS-PAGE and western blotting using anti-CDC20, anti-BUBR1 and anti-MAD2L1 antibodies. Representative western blot is shown. Data reflective of 3 replicate experiments. G Western blot illustrating doxycycline-inducible HORMAD1 expression in RPE1 TRE2-HA-HORMAD1 cells expressing Cyclin B1-Venus. Cells were grown for three days in the presence of doxycycline, and analysed by western blot using indicated antibodies. H Curve illustrating increased Cyclin B1 degradation kinetics in HORMAD1 expressing RPE1 cells. RPE1 TRE2-HA-HORMAD1 cells were grown for three days in the presence or absence of doxycycline, at which point cells were treated with 300 nM nocodazole for 6 h, followed by the addition of 100 nM reversine for 1 h to partially inhibit MPS1. Fluorescence intensity of Cyclin B1 signal was measured over time and normalised to T0. Data reflects 30 cells from 3 biological replicates (data points show mean -/+ SEM). p < 0.0001, two-way ANOVA. I Western blot demonstrating lack of HORMAD1:MAD2L1interaction. HEK293T cells were transfected with HA-tagged HORMAD1 and MAD2L1, alongside GFP-tagged HORMAD1. Asynchronous and mitotic enriched (nocodazole-exposed) lysates were collected, and HA immunoprecipitations were performed. Blot is representative of 2 biological repeats. A, E Created in BioRender. Tutt, A.N.J. (https://BioRender.com/3qiwaov). Source data are provided as a Source Data file.
MAD2L1 is recruited to kinetochores, where it undergoes a conformational change to bind to CDC20 and BUBR1 to form the MCC37,38,39. Therefore, we tested whether HORMAD1 alters the levels of the MCC complex by either reducing its assembly or destabilising the complex. To do so, we treated HORMAD1 expressing RPE1 cells with nocodazole for 8 h, collected mitotic cells, and added either DMSO or reversine for 30 min to perturb the SAC. We then immunoprecipitated CDC20 and immunoblotted for BUBR1 and MAD2L1 (Fig. 3E) to assay for the MCC. Whilst HORMAD1 did not dramatically affect the level of the MCC complex in cells arrested in mitosis with nocodazole, we did notice a small reduction in the amount of MAD2L1 remaining bound to CDC20 when we added reversine compared to adding the DMSO control (Fig. 3F). This could indicate that HORMAD1 partially reduces the stability of the MCC.
Since HORMAD1 increased the rate of mitotic exit after adding reversine, we predicted that the degradation of Cyclin B1 would be enhanced. To test this, we expressed doxycycline-inducible HORMAD1 in RPE1 cells in which the endogenous CCNB1 gene had been engineered to express cyclin B1 tagged with Venus28 (Fig. 3G). We arrested cells in mitosis for eight h and then exposed them to reversine before analysing Cyclin B1 fluorescence using time-lapse fluorescence microscopy. Consistent with HORMAD1 driving or phenocopying SAC perturbations, HORMAD1 expression caused Cyclin B1 to be degraded faster than in control cells (Fig. 3H, p < 0.0001). We concluded from these experiments that when MPS1 is partially inhibited, HORMAD1 reduces the strength of the SAC and/or error correction, as assayed by time-lapse microscopy of mitotic slippage, reduced MCC levels, and accelerated Cyclin B1 degradation.
To understand the molecular interactions responsible for these phenotypes, we tested whether HORMAD1 interacted with MAD2L1 by expressing HA-tagged HORMAD1 and MAD2L1 constructs alongside GFP-tagged HORMAD1. We immunoprecipitated HORMAD1 with anti-HA antibody and found that whilst we could detect GFP-tagged HORMAD1 in our HA-tagged HORMAD1 immunoprecipitates (indicating self-binding), we failed to observe HORMAD1 interactions with MAD2L1 (Fig. 3I). These experiments indicated that HORMAD1 affects the ability of cells to arrest in mitosis, but this was unlikely to be through binding to MAD2L1.
HORMAD1 interacts with, and causes inhibition of, Aurora B
To understand the mechanism by which HORMAD1 perturbed the integrity of the SAC, we sought to identify HORMAD1 interactors in mitotic cells. To efficiently immunoprecipitate HORMAD1 we generated a triple-FLAG-tagged HORMAD1 cDNA construct and expressed this in RPE1 cells. We used mitotic shake-off and immunoprecipitation with the FLAG tag followed by mass spectrometry to identify HORMAD1 interactors from cell in mitosis (Fig. 4A). We identified 13 HORMAD1 interacting proteins using a SAINT40 probability cut-off of >0.7, one of which was Aurora B (Supplementary Data 1, Fig. 4B). Purified recombinant HORMAD1 and Aurora B proteins bound to each other in vitro (Fig. 4C). We confirmed the Aurora B interaction with HORMAD1 in RPE1 cells by immunoprecipitation using a Myc-tagged Aurora B construct and HA-tagged HORMAD1 (Fig. 4D). We also confirmed this interaction in H1299 cells that endogenously express both HORMAD1 and Aurora B (Fig. 4E). We also observed interaction of endogenous Aurora B with dox-induced HORMAD1 in the nucleus of RPE1 cells by proximity ligation assay (PLA) (Fig. 4F, G; Supplementary Fig. 5A), as well as between the endogenously expressed HORMAD1 and Aurora B in H1299 cells (Fig. 4H). Finally, we confirmed the interaction could be observed by PLA in a patient derived organoid (PDO) from an HORMAD1-positive TNBC tumour (KCL014A) but not an HORMAD1-negative tumour (KCL008, Fig. 4I, and Supplementary Fig. 5B).
A Experimental scheme. Created in BioRender. Tutt, A.N.J. (https://BioRender.com/3qiwaov). B HORMAD1 interacting proteins from HORMAD1-FLAG immunoprecipitation with mass spectrometry from experiment shown in (A). SP = SAINT probability score, FC=Fold change. C Western blot illustrating interactions between HORMAD1 and Aurora B in vitro. Recombinant GST-tagged Aurora B was incubated with His-tagged HORMAD1 for 2 h at room temperature and analysed using western blotting with indicated antibodies. Blot is representative of 2 biological repeats. D Western blot illustrating HORMAD1 interactions with Aurora B in mitotic cells. RPE1 TRE2-HA-HORMAD1 cells expressing a Myc-tagged Aurora B construct were exposed to doxycycline or vehicle for 3 days. Subsequently, cells were treated with 300 nM nocodazole for 8 h, and mitotic cells collected using mitotic shake-off. HA immunoprecipitations were performed, followed by western blotting with indicated antibodies. Results representative of 3 biological repeats. E Western blot illustrating endogenous HORMAD1 interactions with Aurora B in H1299 cells. H1299 cells were treated with 300 nM nocodazole for 8 h, and mitotic cells collected using mitotic shake-off. Aurora B or IgG control immunoprecipitations were performed as indicated, followed by western blotting with anti-HORMAD1 and anti-Aurora B antibodies. Blot is representative of 2 biological repeats. F PLA showing interaction between Aurora B and HORMAD1 in HORMAD1 expressing RPE1 cells. RPE1 TRE2-HA-HORMAD1 cells were exposed to doxycycline or vehicle for one day at which point cells were fixed then performed PLA followed by DAPI staining. Representative images are shown; full images with controls are shown in Supplementary Fig. 5. G Quantification of PLA experiment in (H). Graph shows the number of HORMAD1-Aurora B PLA foci. At least 150 cells from each of 3 biological replicates were analysed. p = 0.0001, unpaired t-test (two sided). For panels G-K, red line indicates mean and error bars SEM. H HORMAD1-Aurora B PLA in H1299 cells. H1299 cells containing a doxycycline-inducible shHORMAD1 construct or non-targeting control (NTC) were cultured and analysed as in (F). At least 50 cells were counted from 3 biological replicas. shNTC p = 0.479 (ns) and shHORMAD1 p = 0.0001, unpaired t-test (two sided). I HORMAD1-Aurora B PLA in patient-derived organoid models KCL008 (HORMAD1-negative) and KCL014A (HORMAD1-positive). PLA was carried out using the indicated antibody (ab) combinations. Representative images are shown in Supplementary Fig. 5B. At least 304 cells were analysed per condition. KCL008 p=ns and KCL014A p < 0.0002 (two sided t-test). J Molecular model of complex between Aurora B (yellow) and HORMAD1 (cyan) generated using AlphaFold2. Interaction is predicted between the N-terminal 12 residues of Aurora B (orange) and the safety-belt region of HORMAD1 (purple). The image was produced using PyMOL. K HORMAD1-SNAP proximity ligation assay (PLA) in H1299 cells (expressing endogenous HORMAD1) transfected with the indicated mutant or wild type (WT) Aurora B-SNAP expression constructs. Control samples were stained with the indicated single antibodies. At least 58 cells analysed from 3 biological replicates. KE(4,5) AA p < 0.0001 and N12 del p < 0.0001, unpaired t-test. Source data are provided as a Source Data file.
We used AlphaFold41 to model how Aurora B and HORMAD1 might interact. This modelling suggested that the N terminus of Aurora B could bind to the safety belt region of HORMAD1 (Fig. 4J, and Supplementary Fig. 6). The safety belt region of HORMAD1 normally binds its own C terminus closure motif to form a closed conformation; the binding of HORMAD1 to other partner proteins at the safety belt region would require loss of this intramolecular interaction42 (Fig. 4J). Our modelling predicted that Aurora B binds to the safety belt of HORMAD1 in a similar position to how the closure motif of HORMAD1 binds HORMAD1’s safety belt when in its closed conformation (Supplementary Fig. 6A–C)42. This shared binding mode could be explained by conservation in the three dimensional structure between HORMAD1’s closure motif and Aurora B’s N terminus (Supplementary Fig. 6). Based on this, we predicted that mutating Aurora B residues K4 and E5 would disrupt the HORMAD1/Aurora B interaction (Supplementary Fig. 6C). Using HORMAD1/Aurora PLA, we found that mutants of Aurora B with a 12 amino acid N terminal deletion (residues 1-12, including residues K4 and E5) or a (4,5) KE > AA double point mutation had significantly lower PLA signal with endogenous HORMAD1 in H1299 cells, when, compared to cells transfected with a wild type Aurora B expression vector (Fig. 4K). Taken together, these data suggest that HORMAD1, when expressed in cancer cells, could bind to Aurora B in a manner dependent on the N-terminus of Aurora B.
We next investigated whether the HORMAD1 / Aurora B interaction might affect Aurora B function. Aurora B kinase activity depends on its co-factor, INCENP, which binds Aurora B and promotes its auto-phosphorylation and activation (Fig. 5A)43. We hypothesised that HORMAD1 binding to Aurora B might disrupt the formation of this complex in mitotic cells. To test this, we immunoprecipitated endogenous Aurora B and immunoblotted with antibodies against INCENP (note that two Aurora B isoforms can be detected in RPE1 cells at ~40 kDa and ~35 kDa, which likely correspond to isoform 1 (Q96GD4-1) and isoform 2 (Q96GD4-2) in the ProteomicsDB database44. The validation of this doublet as two Aurora B isoforms has been confirmed by shRNA45). We found that expressing HORMAD1 decreased the levels of INCENP bound to Aurora B (Fig. 5B). This was supported by a PLA experiment where we observed a significant decrease in PLA foci representing INCENP / Aurora B interactions upon induction of HORMAD1 expression (Fig. 5C, D).
A Mechanism of Aurora B activation. B Reduced INCENP:Aurora B interactions. RPE1 TRE2-HA-HORMAD1 cells were exposed to doxycycline or vehicle for 3 days, then 300 nM nocodazole for 8 h, and cells collected by mitotic shake-off. Aurora B immunoprecipitations were performed, followed by western blotting with indicated antibodies. Representative blot shown alongside quantification from 4 biological repeats. p = 0.0063, t-test (two sided). C Representative images of Aurora B-INCENP PLA in RPE1 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle for one day, then 300 nM nocodazole for 6 h. D Quantification of experiment in (C). The number of Aurora B-INCENP PLA foci was calculated from at least 150 cells from 3 biological replicas. p < 0.0001, unpaired t-test. Error bars show mean -/+ SEM. E Western blot showing reduced Aurora B phosphorylation at threonine 232 in mitotic lysates from RPE1 TRE2-HA-HORMAD1 cells cultured as in (B) and analysed using western blotting with indicated antibodies. Representative blot shown alongside normalised densitometry quantification from 5 biological repeats. p = 0.0094, t-test (two sided). F Western blot showing reduced DSN1 phosphorylation at serine 100 in lysates from RPE1 TRE2-HA-HORMAD1 cells cultured as in (B). Representative blot shown alongside normalised densitometry quantification from 5 biological repeats. p < 0.0001, t-test (two sided). G Western blot showing reduced H3 phosphorylation at serine 10 in mitotic lysates from RPE1 TRE2-HA-HORMAD1 cells cultured as in (B). Representative blot shown alongside quantification from 5 biological repeats. p = 0.0083, t-test (two sided). H Immunofluorescence showing defective CENPA phosphorylation at Serine 7 in RPE1 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle for 3 days and then arrested in mitosis with with 300 nM nocodazole, and stained with indicated antibodies. 50 cells from 3 biological replicates were analysed. Nocodazole only p < 0.0001 and +AZD2811 p < 0.0001 (one-way ANOVA). RPE1 cells exposed to the Aurora B inhibitor, AZD2811 (100 nM, 1 h), are included as a positive control. I Mitotic slippage assay in RPE1 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle (water) for three days, then 300 nM nocodazole for 6 h and then 100 nM AZD2811 or vehicle (DMSO). The percentage of cells arrested in mitosis at each time point is shown. A total of 125 cells from 3 biological repeats were analysed. p < 0.0001, Mantel-Cox test with Bonferroni correction (two sided). Error bars show mean -/+ SEM. J Model of HORMAD1 influence on Aurora B. A, J Created in BioRender. Tutt, A.N.J. (https://BioRender.com/3qiwaov). Source data are provided as a Source Data file.
INCENP binding promotes Aurora B auto-phosphorylation at threonine 23246. Immuno-blotting mitotic cells with a phosphorylation-specific Aurora B T232 antibody showed that HORMAD1 expression reduced Aurora B phosphorylation (Fig. 5E) and mitotic cell extracts derived from HORMAD1 expressing RPE1 cells showed reduced phosphorylation of two Aurora B substrates – Serine 100 of DSN1 (Fig. 5F)47, and serine 10 of histone H3 (Fig. 5G). Moreover, HORMAD1 expression significantly impaired phosphorylation of CENP-A–a well-defined Aurora B substrate—at centromeres in mitotic arrested cells (Fig. 5H). Taken together, we found evidence of decreased Aurora B signalling induced by HORMAD1 at centromeres, kinetochores and more widely across chromosomes.
Based on our data, we hypothesised that the HORMAD1 driven effect on the ability of cells to delay mitosis in response to improperly attached kinetochores was a consequence of perturbing Aurora B activity. Indeed, akin to the effect of HORMAD1 expression, inhibition of Aurora B is known to lead to only modest defects in nocodazole-induced mitotic arrest, but profound defects when low doses of MPS1 inhibitor are added16. Since HORMAD1 partially rather than fully blocked Aurora B activity, we wondered whether the presence of HORMAD1 might render cells sensitive to Aurora B inhibition. To test this, we arrested cells in nocodazole, exposed cells to the Aurora B inhibitor AZD281148,49, and then measured the time taken to exit mitosis via slippage. Consistent with other results, we found that HORMAD1 expression exacerbated the modest SAC perturbation caused by Aurora B inhibition (Fig. 5I). These data are consistent with a model whereby HORMAD1 binds to Aurora B, disrupts INCENP binding, and leads to defects in Aurora B signalling (Fig. 5J). We concluded from these experiments that HORMAD1 perturbs the control of mitotic progression at least in part by disrupting Aurora B signalling that normally regulates error correction and reinforces the SAC.
HORMAD1 drives cellular sensitivity to Aurora B, MPS1 and BUB1 inhibitors
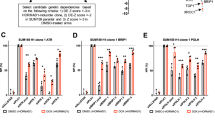
We demonstrated earlier that HORMAD1 expression enhances mitotic slippage induced by either MPS1 or Aurora B inhibition. This raised the hypothesis that somatic cells expressing HORMAD1 would be hyper-dependent upon MPS1 and residual Aurora B kinase activity for cell fitness, proliferation and clonal survival. We therefore inhibited MPS1 or Aurora B with their respective investigational inhibitors, BOS172722 and AZD2811, and measured cellular proliferation using Incucyte live cell microscopy. HORMAD1 expression resulted in a marked decrease in cellular growth when RPE1 cells were exposed to BOS172722 or AZD2811 (Fig. 6A, B). HORMAD1 also increased sensitivity to MPS1 or Aurora B inhibitor in RPE1, SUM159 and SUM149 cells (Fig. 6C–G; and Supplementary Fig. 8A–D). To ensure these were not compound specific effects we confirmed these findings with alternative MPS1 and Aurora kinase inhibitors, AZ-3146 and MK-5108, respectively (Supplementary Fig. 8E, F)50,51.
A Growth curve of RPE1 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle (water) and 200 nM MPS1 inhibitor, BOS172722. Data are from 3 biological replicates and analysed using a two-way ANOVA for vehicle p < 0.0001 and for +BOS172722 p < 0.0001. Error bars show mean -/+ SEM. B Growth curve for RPE1 TRE2-HA-HORMAD1 cells exposed to 20 nM of the Aurora B inhibitor, AZD2811, plotted as (A). Vehicle p < 0.0001 and for +AZD2811 p < 0.0001, ANOVA, error bars show mean -/+ SEM. C Clonogenic survival assay of RPE1 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle (H2O) and to indicated concentration of MPS1 inhibitor, BOS172722. Data show 3 technical replicates, normalised to DMSO, representative of 3 biological replicates. p = 0.0041, two-way ANOVA; error bars show mean -/+ SD. D Clonogenic survival assay of SUM159 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle (water) and to indicated concentration of BOS172722. Data show 3 technical replicates, normalised to DMSO, representative of 3 biological replicates. p = 0.0015, two-way ANOVA; error bars show mean -/+ SD. E–G Clonogenic survival assay of RPE1 (E), SUM159 (F) or SUM149 (G) TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle (water) and to indicated concentration of AZ2811. Data show 3 technical replicates, normalised to DMSO, representative of 3 biological replicates. P values from two-way ANOVA; error bars show mean -/+ SEM. H BUB1 knock-down in RPE1 cells transfected with 2 individual siBUB1 or control siRNA for 3 days. Cell lysates were analysed by western blotting using indicated antibodies. I Graph indicating reduced cellular survival in HORMAD1 expressing RPE1 cells depleted of BUB1 using siRNA. RPE1 TRE2-HA-HORMAD1 were exposed to doxycycline or vehicle, and were reverse transfected with indicated siRNA. 5 days post transfection, viability was measured using Cell Titre Glo reagent and surviving fractions calculated. Data representative of 3 biological replicas, each with 5 technical replicates. For BUB1.1 p < 0.0001 and for BUB1.2 p = 0.001, one-way ANOVA with Sidak’s correction. J Western blot illustrating expression of HORMAD1 in wild-type RPE1 TRE2-HA-HORMAD1 cells and cells depleted of BUB1 using CRISPR. K Graph indicating reduced cellular survival in HORMAD1 expressing RPE1 cells depleted of BUB1 using CRISPR. WT and BUB1 depleted RPE1 TRE2-HA-HORMAD1 were exposed to doxycycline or vehicle. These data are representative of 3 biological replicas, each with 3 technical replicates per replicate. Error bars show mean -/+ SD. p = 0.8666 (WT) and p = 0.0183 (sgBUB1); one-way ANOVA with Sidak’s correction. L RPE1 TRE2-HA-HORMAD1 cells were exposed to doxycycline or vehicle and indicated concentration of the BUB1 inhibitor, BAY-1816032. M Quantification of experiment in (L). Cellular viability was estimated using SRB absorbance readings. Data show 3 technical replicates, normalised to DMSO, and representative of 3 biological replicas. Error bars show mean -/+ SEM. p = 0.0086, one-way ANOVA. N Growth curve of RPE1 TRE2-HA-HORMAD1 cells exposed to doxycycline or vehicle (H2O) and 3 µM BAY-1816032 for 14 days. Data represent 3 biological replicas. Error bars show mean -/+ SEM. p < 0.0001 and for +BAY1816032 p < 0.0001; two-way ANOVA with Bonferroni correction. Source data are provided as a Source Data file.
Aurora B and MPS1 are master regulators of mitotic checkpoints and chromosome segregation, whose inhibition has significant effects on both malignant and non-malignant normal tissue cells. Conversely, BUB1, which regulates MPS1 and Aurora B activity during mitosis, is not widely essential in human cells32,52 and may be a more clinically tractable target predicted to have less toxicity to proliferative, but non-malignant, cells. To test whether HORMAD1 expression caused a dependency upon BUB1, we exposed RPE1 cells to doxycycline to express HORMAD1, then depleted BUB1 with two individual siRNAs and estimated cellular viability after five days in culture, revealing a highly selective addiction to BUB1 in HORMAD1 expressing RPE1 cells (Fig. 6H, I). We validated these findings in RPE1 cells depleted of BUB1 using CRISPR-Cas9 gene editing (Fig. 6J, K). Importantly, these cells still expressed low levels of BUB1 protein that appears to be functionally important53. These data indicated that HORMAD1 expression is synthetic lethal with BUB1 depletion in somatic cells.
More recently, small molecule BUB1 inhibitors have been developed as potential cancer therapeutics, with the intention to use them in combination with microtubule interacting chemotherapies24. Biomarker patient selection strategies that could direct the use of BUB1 inhibitors do not as yet exist. To test whether HORMAD1 expression creates highly selective sensitisation in cells to small molecule mediated BUB1 inhibition, we first performed clonogenic survival assays using RPE1 cells expressing doxycycline-inducible HORMAD1. These experiments revealed HORMAD1-driven BUB1 inhibitor sensitivity (Fig. 6L, M). We also used Incucyte-based time-lapse imaging to measure cell population growth during BUB1 inhibitor treatment. These results demonstrated that whilst HORMAD1-negative non-malignant cells show very little BUB1 inhibitor induced toxicity and can reach cellular confluence, HORMAD1-positive cells fail to grow past 20% confluence in the presence of a single agent BUB1 inhibitor (Fig. 6N).
HORMAD1 expressing tumours are sensitive to Aurora B inhibition in vivo
To assess whether inhibition of these mitotic kinases might form part of a viable targeted treatment for HORMAD1-expressing cancers, we used TNBC breast cancer patient derived xenograft models (PDX) to investigate the in vivo response to Aurora B inhibition. Both AZD2811 and its prodrug barasertib are Aurora B inhibitors that have been investigated in early stage clinical trials49, therefore we chose to investigate these agents in PDX models characterised for HORMAD1 expression (Fig. 7A, B, and Supplementary Table 1). A nanoparticle formulation of AZD2811 (NP50) caused a reduction in tumour growth in mice bearing PDX, or xenografts formed from patient derived organoids (PDOX), from HORMAD1-positive TNBC patients (KCL010, KCL003 and KCL625, Fig. 7C–K and Supplementary Tables 2, 3, Supplementary Fig. 8). Consistent with this, there was no significant effect of the same dose of inhibitor on tumour growth of a PDX from a HORMAD1-negative TNBC patient (KCL002; Fig. 7L–N). Although one HORMAD1-positive model had a significantly higher growth rate and mitotic index (Supplementary Fig. 8E, F), this did not correlate with response. We also observed significantly slower tumour growth under AZD2811 treatment in an isogenic SUM159 cell line xenograft with inducible HORMAD1 expression, compared to uninduced control (Supplementary Fig. 9). These data demonstrated that out of context expression of HORMAD1 in tumour cells is associated with in vivo Aurora B inhibitor sensitivity, consistent with the biological effects of HORMAD1 we demonstrated on mitotic slippage in the presence of these agents.
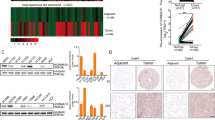
A HORMAD1 RNA expression in patient-derived models assessed by RT-qPCR. Tonsil RNA is included as a non-HORMAD1 expressing somatic tissue control. Error bars show mean -/+ SEM. B HORMAD1 mRNA expression in RNA-seq samples from patient derived xenograft models (PDX, blue). Where available, patient tumour sample from which the PDX was derived is shown in red. C–N Representative HORMAD1 immunohistochemistry images, individual tumour growth curves and survival assays based on tumours reaching size limit are shown for four PDX models: endogenous HORMAD1-positive KCL010 (C–E), KCL003 (F–H) and KCL625OX (organoid-derived xenograft; I–K), and HORMAD1-negative KCL002 (L–N). White scale bar = 50 μm P values calculated with log-rank test. KCL010 p = 0.001, KCL003 p = 0.0002, KCL625OX p < 0.0001, KCL002 p = 0.3983. HORMAD1 H-score for IHC images was calculated (based on whole slides) as described in Methods. Loss-of-function mutations detected in TP53 or breast cancer driver genes are annotated. gPALB2, germline PALB2 mutation. Source data are provided as a Source Data file.
Discussion
Here, we report a mechanism by which somatic expression of the meiotic protein HORMAD1 drives chromosomal instability in cancer. We show that expression of ectopic or endogenous HORMAD1 leads to aberrations in the activation of the SAC as measured by mitotic slippage and Cyclin B1 degradation. Despite shared possession of HORMA domains, this defect was independent of effects on MAD2L1 recruitment to kinetochores, and was rather a consequence of Aurora B binding, disruption of INCENP interactions, and the suppression of Aurora B signalling. We noted that the extent to which elevated HORMAD1 expression perturbed mitotic arrest was not as profound as, for example, MAD2L1 inactivation32; this is perhaps unsurprising, given profound SAC perturbations are generally not compatible with cell viability54, as shown by the embryonic lethality of Mad2l1 or Bubr1 knockout mice55,56. Whilst full abrogation of the SAC is usually incompatible with cell viability, partial inhibition of the SAC has been linked to the induction of cancer, as illustrated by mice with either Mad2l1 (Michel et al., 2001) or Bubr1 (Dai et al., 2004) haploinsufficiency. Notably, naturally HORMAD1-expressing breast cancer cell lines (e.g., MDA-MB-436 and SUM159) generally displayed less slippage with nocodazole alone, and HORMAD1-dependent mitotic slippage was only observed in combination with sensitisation of the SAC using reversine (Supplementary Fig. 3H, J), suggesting that tumours may adapt to long term disruption of the SAC in the presence of HORMAD1.
In addition, out-of-context expression of HORMAD1 sensitises cells to Aurora B, MPS1 and BUB1 inhibitors, findings that have clear implications for the use of drugs currently in clinical development for the treatment of many cancers, including TNBC. Taken together, our data point towards a model in which HORMAD1 binds to Aurora B and inhibits its activity on the SAC and/or error correction, leading to chromosomal instability. This binding requires the N terminus of Aurora B which we predict interacts with the safety belt region of HORMAD1 in a similar manner to intramolecular binding of the HORMAD1 C terminus closure motif (Fig. 4J, and Supplementary Fig. 6). Consistent with this, HORMAD1-expressing cells were sensitive to MPS1, BUB1 or Aurora B inhibitors (Fig. 6), which may be driven by exacerbated chromosome instability. Given that HORMAD1 is not expressed in non-germline normal tissues but is expressed in 60% of TNBCs7 and a significant number of other high unmet need malignancies (Fig. 1L), this indicates a selective dependence on several druggable regulators of mitotic segregation identifiable by a bimodally expressed biomarker in malignant cells.
Our findings are consistent with and provide mechanistic explanations for our prior observation of a selective dependency on mitotic checkpoint regulatory kinases in a number of models of TNBC in a RNA interference screen informed by integration of genomics data from primary TNBCs57. In addition to potentially informing patient stratification in breast cancer, our data reveal a mechanism by which chromosomal instability and aneuploidy may be induced by HORMAD1 in malignancy more broadly. We found that HORMAD1 expression causes the mis-segregation of chromosomes and that HORMAD1 expression positively associates with genomic signatures of aneuploidy in numerous tumour histological sub-types.
At present, it is difficult to know what other molecular features might modify the relationship between HORMAD1 and mitosis. For example, although RB1 loss is common in TNBC and present in about 40% of HORMAD1-positive basal-like cancers58, there seems to be little evidence to suggest that those tumour cell lines that express HORMAD1 and lack RB1 are those that have the most severe mitotic phenotypes. Given this is an important issue to address, our future work will focus on identifying the genes that when perturbed, increase or decrease mitotic slippage in the presence of HORMAD1.
Aurora B is known to regulate several aspects of mitosis, including error-correction and cytokinesis, in addition to contributing to the SAC. Our data indicate that HORMAD1 perturbs the CPC complex, and thus likely impairs the regulation of these functions as well as the SAC. This raises the possibility that disruption of the SAC by HORMAD1 expression may drive genomic instability, and thus tumourigenesis. Although loss of function mutation of SAC components is not often seen in cancer, smaller changes in the levels of SAC components have been previously observed to result in functional SAC defects34,59. It is possible that a partial disruption of Aurora B function in the presence of HORMAD1 results in similar changes in tumours.
One remaining question raised by our work is why this mechanism of HORMAD1 induced Aurora B inhibition might exist. In meiosis, Aurora B activity drives chromosome cohesion resolution and de-synapsis60,61. Meiotic HORMA domain proteins, on the other hand, are known to regulate synapsis and cohesion62,63,64, though the mechanisms by which this is achieved remain incompletely understood in mammalian cells. Although beyond the scope of this work, HORMAD1 could play a similar role in silencing Aurora B activity in meiosis. Indeed, it has previously been reported that Aurora B is negatively regulated by HORMA domain proteins in meiosis of lower eukaryotic species65. Meiotic regulation of Aurora B by HORMAD1 might therefore prevent premature Aurora B-mediated cohesion resolution.
The bimodality of the HORMAD1 expression in TNBC makes it a highly attractive biomarker for patient stratification, easing the definition and identification of a biomarker positive group. In addition to being expressed in a large fraction (~ 60%) of TNBCs, it is expressed in a significant fraction of other malignancies that also have a highly unmet clinical need such as non-small cell lung carcinoma. We propose that aberrant expression of the meiotic protein HORMAD1 might be used as an indicator of partial SAC impairment, or a phenocopy of the same through effects on Aurora B-mediated error correction. It is worth setting out what subsequent work is required to translate our observations that HORMAD1 weakens the SAC and causes sensitivity to mitotic checkpoint inhibitors. Whilst we see both in vitro and in vivo sensitivity to Aurora B inhibition when HORMAD1 is expressed, there are still several challenges to translating this observation. One challenge is extending the therapeutic window achieved by Aurora B inhibition and another is minimising the impact of deleterious side effects that have previously challenged the widespread application of Aurora kinase inhibitors66,67. With both of these issues in mind, our future work will focus on using pre-clinical systems to identify drug combination approaches with that widen the therapeutic window of MPS1, Aurora B and BUB1 inhibitors in HORMAD1 positive breast cancers enhancing anti-tumour efficacy at doses of such inhibitors that might be significantly below a maximum tolerated dose (MTD). Identifying such combinations could be achieved using large-scale chemical or genetic in vitro perturbation screens involving Aurora B, MPS1 or BUB1 inhibitors in HORMAD1 isogenic systems, followed by an extensive assessment of promising combinations in in vivo models. Since we find HORMAD1 to exacerbate MPS1 inhibition-induced SAC perturbations in lung cancer and cervical cancer models, our findings may also be relevant to other cancers.
It is important to state that we do not regard HORMAD1 expression in cancer to be an absolute necessity for sensitivity to inhibitors of mitotic checkpoint kinases. For example, loss of the RB1 tumour suppressor has been shown to induce Aurora B inhibitor sensitivity in models that lack HORMAD1 expression23,68. Conversely, we show in our work that Aurora B inhibitor sensitivity can be induced via HORMAD1 expression in cell lines that have no defect in RB1, such as MCF10A and RPE1 cells. As such, the most parsimonious explanation would be that Aurora B inhibitor sensitivity has multiple drivers, some of which work independently of one another.
We do note a number of limitations of our study. For example, whilst we established the link between out of context HORMAD1 expression and sensitivity to mitotic checkpoint inhibitors in multiple cell lines and show that HORMAD1-positive TNBC PDX are also sensitive to an investigational Aurora B inhibitor, the in vivo responses seen were not curative. This is common when targeted agents are used as single agents in cancer. Despite this limitation with an appropriate biomarker-based patient selection strategy they can extend life and improve symptoms. In future work it may be possible to identify drugs, such as microtubule disrupting agents used as standard of care in TNBC69, that could be combined with Aurora B, MPS1 or BUB1 inhibitors to target HORMAD1-positive cancers in a way that achieves sustained anti-tumour responses.
Methods
Ethical regulations
PDX and PDO models used in this study were derived from the BTBC (Breakthrough Breast Cancer) study. BTBC is a longitudinal blood and tissue sampling protocol conducted alongside standard of care therapy in breast cancer patients. Written informed consent was given by patients and the study conducted according to national ethics committee approved protocol (REC No: 13/LO/1248, Guy’s & St Thomas’ Hospital). Animal studies received ethical approval from King’s College London (KCL) Animal Welfare Ethical Review Body (AWERB) and the UK Home Office (PPL PF642A32A).
Cloning and production of lentiviral vectors
Wild-type HORMAD1 cDNA was purchased from GeneART (Thermo Scientific). HORMAD1 cDNA was cloned into the pINDUCER20 lentivirus expression vector, as described previously70. Lentiviral vectors were produced in HEK293T cells by transfection of pINDUCER20 plasmid, alongside pMD2.G and psPAX2 packaging vectors using Lipofectamine 2000 (Thermo Fisher) according to the manufacturer’s protocol. Virus containing media was collected, filtered, and stored at −80 °C.
Cell culture
Cell lines obtained from ATCC: RPE1 hTERT (CRL-4000), HeLa (CCL-2), MCF10A (CRL-10317), MDA-MB-436 (HTB-130) NCI-H1299 (CRL-5803). Cell lines obtained from BioIVT/Asterand: SUM159PT (HUMANSUM-0003006), SUM149PT (HUMANSUM-0003004). RPE1 and HeLa cells were grown in Dulbecco’s Modified Essential Media (DMEM) supplemented with 10% foetal bovine serum (FBS). MCF10A were grown in DMEM/F-12 (Gibco) supplemented with 5% Horse Serum, EGF (20 ng/mL), Hydrocortisone (0.5 µg/mL), Cholera Toxin (100 ng/mL), and Insulin (10 µg/mL). SUM159 and SUM149 cells were grown in Ham’s F-12 (Gibco) supplemented with 5% FBS, HEPES (10 nM), Hydrocortisone (1 µg/mL), and Insulin (5 µg/mL). MDA-MB-436 cells were grown in RPMI supplemented with 10% foetal bovine serum (FBS) and Insulin (5 µg/mL). H1299 cells were grown in RPMI supplemented with 10% foetal bovine serum (FBS). To induce HORMAD1 cDNA or shRNA expression, doxycycline was added at a concentration of 0.5 µg/mL. For dose-response survival assays, cells were exposed to drug for 7-10 days and viability measured using sulphrhodamine B (SRB) staining. Cell lines were authenticated by STR typing. All cells were tested regularly for mycoplasma contamination using MycoAlert (Lonza).
Generation of inducible HORMAD1 cell-lines
MCF10A, SUM159 and RPE1 cells were infected with P20-HORMAD1 lentivirus vectors and were grown in 0.5 mg/mL Geneticin (Gibco). All model systems were validated using western blotting for doxycycline-induced HORMAD1 expression.
Depletion of HORMAD1 or BUB1 using RNAi
RPE1 cells were reverse transfected (cell suspension pipetted onto the lipofectamine siRNA mixture) with 20 nM non-targeting, or BUB1-targeting siRNA, using Lipofectamine RNAi Max (Thermo Fisher). HeLa cells were reverse transfected with 20 nM non-targeting, or HORMAD1-targeting siRNA, using Lipofectamine 2000 (Thermo Fisher). For shRNA mediated depletion of HORMAD1, shRNA vectors were cloned into LT3GEPIR and lentiviral particles were generated to tranduce cell lines. After transduction, cells were selected using 1.5 µg/mL puromycin and induced to express shRNA constructs using 1 µg/mL doxycycline. HORMAD1/BUB1 depletion was confirmed using Western blotting. Full list of RNAi sequences is in Supplementary Table 4.
Small molecule inhibitors of MPS1, Aurora B and BUB1
All drugs were reconstituted in DMSO, aliquoted, and stored at −20 °C. Full list of drugs is in Supplementary Table 5.
Time-lapse microscopy
Bright field time-lapse microscopy was performed using a Nikon Eclipse microscope, with a 20x or 10x objective. Measurement of time arrested in mitosis was performed manually. Time lapse microscopy using H2B-GFP labelled cells was performed using a Marianas spinning-disk microscope system (Intelligent Imaging Innovations), using a 488 nm laser and equipped with a 40x objective. Assessment of segregation errors was performed manually. Time lapse microscopy using Venus labelled Cyclin B1 was imaged using a Nikon Eclipse microscope, with a 20x objective and 2 × 2 binning. Images were acquired every 5 min, and intensity values were calculated using imageJ.
Immunocytochemistry and Confocal microscopy
RPE1 cells were plated onto glass coverslips and grown in the presence of 0.5 µg/mL doxycycline for 3 days. Cells were treated with 300 nM nocodazole for 3 h, washed with PBS, and fixed in 4% paraformaldehyde for 10 min at room temperature. Subsequently, cells were washed in PBS, permeabilised with 0.5% Triton-X100 (in PBS) for 5 min at room temperature, and blocked in 3% BSA (in PBS) for 20 min at room temperature. Primary antibodies were then added to cells and incubated overnight at 4 °C. Full list of antibodies and concentrations used are in Supplementary Table 6. Cells were then washed with PBS and incubated with secondary fluorescent antibodies for 1 h at room temperature. All antibodies were diluted in 3% BSA/PBS. Finally, cells were washed with PBS and cover slips were mounted onto glass slides.
Confocal images were collected on a Marianas spinning-disk microscope system (Intelligent Imaging Innovations), with 445/488/561/650 nm lasers, an Observer Z1 inverted microscope (Zeiss), and equipped with a 100x lens. Images were analysed using Cell Profiler71: Single cells were selected, and kinetochores were marked using CREST staining28 to determine the number of kinetochores positive for MAD2L1 signal. Alternatively, the intensity of protein-of-interest (e.g., p-CENPA) was normalised to the CREST signal. For analysis of centric and acentric micronuclei, cells were stained with CREST antibodies and DAPI before high throughput confocal microscopy was also performed using the ImageXpress (Molecular Devices) platform with a 20x lens. Z-stacked images with 1 µm intervals were obtained that spanned the nucleus and were analysed manually. Analysis of MAD2L1 and p-CENPA was performed using CellProfiler and ImageJ, respectively. Analysis of centromeric micronuclei was performed manually.
HORMAD1 immunohistochemistry
FFPE sections (4 µm thick) were stained on a Ventana Discovery Ultra (Roche). Antigen unmasking was performed using the low pH CC2 citrate buffer (Roche) for 48 min at 98 °C followed by a further 24 min at 65 °C. Primary antibody incubation was performed at room temperature with rabbit anti-HORMAD1 (HPA037850, Sigma) at a dilution of 1:20 for 92 min. Binding of the primary antibody was detected using UltraMap anti-rabbit antibody (Roche) incubated for 32 min at room temperature, visualised with DAB and counterstained with haematoxylin. Images were obtained using a Hamamatsu Nanozoomer analysed in QuPath72. Colour vectors were adjusted automatically before nuclei were segmented using StarDist73. Nuclear mean DAB optical density was used to threshold cells as positive and negative with pathologist assessment. The positive cells were then subdivided as 1 + , 2+ or 3+ in 1 s.d. increments. H-scores74 for all samples were calculated in QuPath.
Immunoprecipitation
RPE1 cells were plated into 5-layer cell stacks (Corning Falcon) and grown in the presence of 0.5 µg/mL doxycycline until 70% confluent. Cells were synchronised to mitosis by adding 300 nM Nocodazole for 8 h, when cell become rounded and can be dislodged by shaking off. After the shake off the isolated mitotic cells were collected and washed with PBS and lysed in lysis buffer NP150 (20 mM Tris pH 7.6, 1 mM EDTA pH 8, 0.5% IGEPAL CA-630, and 150 mM NaCl) supplemented with protease inhibitor cocktail (Roche) and left on ice for 1 h. Cell debris was then pelleted by centrifugation (14,000x g for 10 min at 4 °C), and the supernatant containing cell lysates was collected. Antibodies against the bait protein were incubated with protein A- or G-coupled magnetic beads (Dynabeads, Thermo Fisher) for 30 min at the indicated concentration (Supplementary Table 6). After three rounds of washes with lysis buffer 500 µg of cell lysates were added to the antibody-bead conjugation for 2 h at room temperature or overnight at 4 °C. Beads were washed 5 times with cold NP150, and interacting proteins were eluted by boiling beads in the presence of Laemmli buffer (2% SDS, 10% glycerol, 0.002% bromophenol blue, 0.06 M Tris-Cl, pH 6.8, 5% 2-mercaptoethanol) (Thermo Fisher).
SDS:PAGE and Western blotting
Immunoprecipitations, whole cell extracts and chromatin extracts were all diluted in 5x Laemmli buffer to give a final concentration of 1x (2% SDS, 10% glycerol, 0.002% bromophenol blue, 0.06 M Tris-Cl, pH 6.8, 5% 2-mercaptoethanol). Extracts were then loaded into the wells of precast polyacrylamide (4–12%) gels (Thermo Fisher) and separated by SDS:PAGE. Western blot transfers were performed using wet transfer cassettes (BioRad). Membranes were blocked in 5% milk in TBS-T for 20 min and incubated with primary antibodies at the indicated concentration (Supplementary Table 6) for 2 h at room temperature or overnight at 4 °C. Membranes were washed 3 times with PBS and incubated with secondary antibodies conjugated to HRP for 1 h at room temperature. For immunoprecipitation, protein G conjugated to HRP was used in place of a secondary antibody, in order to remove the binding between the secondary antibody and the immunoglobulin chains of the bait antibody75. Membranes were then washed 3 times with TBS-T before imaging bands using electrochemiluminescence with the LiCor imaging platform. All western blot images were non-saturated. Full list of antibodies in Supplementary Table 6.
Metaphase spreads for chromosome counts
RPE1 cells were synchronised in G1 by growth at high density for 48 h and HORMAD1 induction was induced by addition of 0.5 µg/mL doxycycline. Cells were then split into low density and grown to reach confluency of 70–80%. To enrich for metaphase cells, 0.2 μg/mL of colchicine was added to the cells and left for 4 h. Metaphase cells were harvested by trypsinization, washed two times with PBS and slowly resuspended in pre-warmed 75 mM KCl and incubated 10 min at 37 °C. After centrifugation at 420 x g for 5 min, cells were fixed in 3:1 methanol: glacial acetic acid (fixation buffer). After incubating for 15 min at room temperature, cells were centrifuged and resuspended again in fixation buffer. After two additional rounds of resuspension and fixation, cell pellets were kept at −20 °C in a small volume of fixation buffer. Metaphase spreads were made by dropping the cells from ~50 cm height to slides tilted by 45°. The slides were allowed to air-dry before mounting with Vectashield containing DAPI. Chromosome spreads were then imaged by a spinning disc confocal microscope.
HORMAD1-FLAG interactome using mass spectrometry
Immunoprecipitation was performed as described above but without eluting peptides with Laemmli buffer. On-bead digestion was as described previously76. Beads were then washed 3 times with 50 mM ammonium bicarbonate. Peptides were then digested, whilst on beads, by incubating with 1 µg trypsin (Roche, sequencing grade) overnight at 37 °C. The supernatant containing peptide fragments was collected and filtered (Millipore Multiscreen HTS plate, pre-washed with 60% acetonitrile). Peptides were dried using a speed vacuum, re-solubilised in 20 mM TCEP and acidified to 0.5% formic acid. Two biological replicates were processed per condition.
The LC-MS/MS analysis was performed on the Orbitrap Fusion Lumos mass spectrometer coupled with U3000 RSLCnano UHPLC system. All instruments and columns used were from Thermo Fisher. The peptides were first loaded to a PepMap C18 trap (100 µm i.d. x 20 mm, 100 Å, 5 µm) at 10 µL/min with 0.1% FA/H2O, and then separated on a PepMap C18 column (75 µm i.d. × 500 mm, 100 Å, 2 µm) at 300 nL/in and a linear gradient of 4-32% ACN/0.1% FA in 90 min with the cycle at 120 min. Briefly, the Orbitrap full MS survey scan was m/z 375 – 1500 with the resolution 120,000 at m/z 200, with AGC (Automatic Gain Control) set at 40,000 and maximum injection time at 50 ms. Multiply charged ions (z = 2-5) with intensity above 8,000 counts were fragmented in HCD (higher collision dissociation) cell at 30% collision energy, and the isolation window at 1.6 Th. The fragment ions were detected in ion trap with AGC at 10,000 and 35 ms or 50 ms maximum injection time. The dynamic exclusion time was set at 30 s with ±10 ppm. Raw mass spectrometry data files were analysed with Proteome Discoverer 2.3 (Thermo). Database searches were carried out using Mascot (version 2.4) against the Uniprot human reference database (January 2018, 21123 sequences) appended with the cRAP database (ftp://ftp.thegpm.org/fasta/cRAP), with the following parameters: Trypsin was set as digestion mode with a maximum of two missed cleavages allowed. Precursor mass tolerance was set to 20 ppm, and fragment mass tolerance set to 0.5 Da. Acetylation at the protein N terminus, oxidation of methionine and deamidation of asparagine and glutamine were set as variable modifications. Peptide identifications were determined at 1% FDR using Mascot Percolator. SAINTexpress was used to score interaction specificity and calculate fold enrichments40. Proteins with SAINT probability score > 0.7 (FDR < 5%) were deemed specific interactors. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE77 partner repository with the dataset identifier PXD023666.
Analysis of cell viability after genetic perturbations or drug treatments
In order to assess cellular viability of RPE1 cells expressing doxycycline-inducible HORMAD1 after siRNA depletion of BUB1, cells were reverse transfected into the wells of 384 well plates (200 cells per well) with RNAi particles, and simultaneously treated with 0.5 µg/mL doxycycline. 5 days post transfection, cellular viability was estimated using Cell Titre Glo (CTG) reagent. CTG luminescence was measured using a Victor X5 plate reader (Perkin Elmer). Data was presented as the surviving fraction, relative to control siRNA particles.
In order to assess the cellular viability of parental RPE1 cells or RPE1 cells depleted of BUB1, we stably expressed doxycycline-inducible HORMAD1 in both models and plated cells into the wells of 6-well plates (500 cells per well). Cells were grown (in the presence of 0.5 µg/mL doxycycline) for 10 days and fixed using 3.3% acetic acid. Cells were stained with sulforhodamine B (SRB) and imaged. SRB was dissolved using 10 mM Tris pH 10–12 and the absorbance (510 nm) given from dissolved SRB was used to assess cell density78.
Clonogenic survival assays following MPS1, BUB1 and Aurora B inhibitors were performed by plating cells into the wells of 6 well plates (500 cells per well) and adding 0.5 µg/mL doxycycline. The following day, increasing doses of small molecule inhibitors were added and cells were grown for 10 days in the presence of drug and fixed using 3.3% acetic acid. For RPE1 experiments, -DOX conditions were fixed after 8 days and +DOX after 10 days so that cells achieved similar growth. Cells were stained with SRB and imaged using the GelCount platform. Colonies were counted automatically using Cell Profiler.
In order to monitor cellular proliferation over a 2 week period, RPE1 cells expressing doxycycline-inducible HORMAD1 were treated with 0.5 µg/mL doxycycline and plated into the wells of a 24 well-plate (500 cells per well). The following day, BUB1/Aurora B/MPS1 inhibitor was added and cells were placed into an Incucyte microscope platform. Cells were then imaged every 4 or 6 h for 2 weeks, and growth kinetics were assessed using confluence as a measure of cellular division.
Purification of recombinant HORMAD1 protein
His-tagged HORMAD1 bacterial expression constructs were described previously (Kim et al., 2014), expressed in BL21 cells treated with 0.1 mM IPTG overnight at room temperature. His-tagged HORMAD1 was then purified using the NTA Ni2+ purification kit (Qiagen) according to the manufacturer’s protocol. ~100 ng HORMAD1 was then incubated with an equal amount of recombinant GST-tagged Aurora B protein (Abcam, ab51435) diluted in 0.5 mL PBS with 0.1% Tween−20. HORMAD1:Aurora B complexes were then purified using Ni2+ spin columns.
Structural modelling
Molecular modelling of the Aurora B/HORMAD1 complex used AlphaFold2-multimer, implemented by ColabFold79. Ten models of human Aurora B (aa 1-344) and HORMAD1 (aa 1-382) were generated, all of which predicted the interaction of Aurora B N-terminus with HORMAD safety belt region (aa 179-229)42. AlphaFold2 modelling using only the N-terminus of Aurora B (aa 1-13) also predicted an interaction with the HORMAD1 safety belt region, and the best model was of moderately high confidence (interface predicted template modelling score, ipTM = 0.714).
Proximity ligation assay (PLA)
Cells (on coverslips for cell line experiments and cytospin slides for organoid material) were fixed in 4% paraformaldehyde for 10 minutes at room temperature. Subsequently, cells were permeabilized with 0.2% Triton-X100 for 10 minutes at room temperature and processed following the manufacturer’s instructions (Duolink Proximity Ligation Assay, Sigma). Briefly, samples were blocked for 1 h at 37 °C followed by primary antibody incubation at room temperature (see list of antibodies in Supplementary Table 6). Samples were washed twice with Buffer A for 5 min and afterwards incubation with PLA probes for 1 h at 37 °C. After washing samples twice with Buffer A for 5 min, ligation reaction was added and incubated for 30 min at 37 °C. After washing samples twice with Buffer A for 5 min amplification reaction was added and incubated for 90 min at 37 °C. This was followed by two final washes with Buffer B for 10 min. Samples were mounted in mounting media and imaged on confocal microscopy as described above.
In silico analysis of HORMAD1 expression and aneuploidy signatures from tumour biopsies
HORMAD1 bimodal status was established in TCGA mRNA datasets using the R package cutoff v0.1.0 (R v4.1.0). TCGA datasets were downloaded from (http://gdac.broadinstitute.org) (release 28 January 2016). Aneuploidy scores were obtained from Taylor et al.29. Briefly, the aneuploidy score (AS) is defined as the number of altered (lost or gained) chromosome arms, ranging from 0 (no arms altered) to 39 (all measured arms altered) based on copy number data obtained from Affymetrix SNP 6.0 arrays.
PDX Experiments
Human breast tumour samples were obtained from adult female patients after informed consent as part of a non-interventional clinical study (BTBC study REC no.: 13/LO/1248, IRAS ID 131133; Principal Investigator: Prof. Andrew Tutt; Study Title: “Analysis of functional immune cell stroma and malignant cell interactions in breast cancer in order to discover and develop diagnostics and therapies in breast cancer subtypes”). This study had local research ethics committee approval and was conducted adhering to the principles of the Declaration of Helsinki. Specimens were collected from surgery and transported immediately to cut up. A clinician histopathologist or pathology-trained technician identified and collected tumour material into basal culture media.
All animal experiments described below were carried out under Home Office PPL PF642A32A using NOD.Cg-PrkdcscidIl2rgtm1Wjl/SzJ (NSG) mice which were obtained commercially from Charles River UK and housed under UK Home Office regulations. Miced were housed in GM500 individually ventilated cages at a temperature range of 20 °C - 23 °C, humidity range of 45% – 55% and a light/dark cycle of 12 h light/ 12 h dark. All animal studies were conducted under PPL PF642A32A Protocol 3 (Subcutaneous or Mammary Fad Pad Models) which received ethical approval from King’s College London (KCL) Animal Welfare Ethical Review Body (AWERB) and the Home Office. The maximum tumour size permitted on this licence (as approved by KCL AWERB and the Home Office) was a maximum of 15 mm tumour diameter for unilateral tumours or 12 mm diameter for bilateral tumours. Tumour limits were not exceeded for any of the experiments perfomed. Only female mice were used for the animal studies detailed in this paper as studies utilised patient derived xenograft (PDX) models or cell line xenografts derived from female breast cancer patients.
To generate patient derived xenograft (PDX) models, ~2 mm fragments of patient tumours were orthotopically implanted under anaesthesia into the fourth, right hand side mammary fat pad of female NSG mice via trocar. Tumour presence was monitored by palpation and calliper measurement. Mice were culled once tumours reached 12 mm in diameter. Tumours were harvested and dissected into 2 mm fragments and either implanted fresh into a new cohort of NSG mice or viably frozen and subsequently implanted into new hosts from frozen.
Four PDX models (KCL010, KCL003, KCL625OX and KCL002) were used for experiments to test response to the Aurora B inhibitor AZD2811. For each PDX model 22–49 day old female NSG mice were implanted via trocar in the 4th right hand side mammary fat pad with 2 mm pieces of viably frozen PDX tumour tissue from a single mother tumour. For each model the following number of mice were implanted with tumours: KCL010 = 25 mice (22–26 days old); KCL003 = 30 mice (all 45 days old); KCL625OX = 30 mice (all 39 days old) and KCL002 = 25 mice (all 33 days old). Following implantation, tumours size was assessed twice weekly using callipers. Tumour volume was calculated using the following equation: Volume = π/6 × Length × Width2, where length represents the largest tumour diameter and width represents the perpendicular tumour diameter.
Mice implanted with tumours from each of the 4 PDX models used were distributed equally into two experimental groups, AZD2811 Nanoparticles (AZD2811 NP) or vehicle control, once tumours reached a range of 45-95 mm3 in volume. Tumour volume measurements at study entry are shown in Supplementary Table 2. Randomisation to treatment groups was performed using a random number generator. The number of mice treated for each PDX model are as follows: KCL010 vehicle = 9, AZD2811 NP = 8; KCL003 vehicle = 11, AZD2811 NP = 10; KCL625OX vehicle = 10, AZD2811 NP = 10; KCL002 vehicle = 7, AZD2811 NP = 6. Mice were treated once weekly by intravenous injection with 25 mg/kg AZD2811 NPs, diluted in saline, or with a corresponding volume of saline vehicle control. Mice were culled once tumours reached 11 mm in diameter. The antitumour activity was determined by calculating the percentage change in tumour volume at each timepoint relative to tumour volume at treatment day: Percentage change in tumour volume = (Volume day X – Volume day 0) / Volume day 0 ×100. Difference in survival was assessed using a log-rank (Mantel-Cox) test. AZD2811 NP treatment was generally well-tolerated; out of a total of 34 mice treated across all 4 PDX models, a single mouse was culled due to weight loss (Supplementary Fig. 8A–D).
Cell line xenograft experiments
One million SUM159 cells expressing a doxycycline-inducible HORMAD1 expression construct were injected into the fourth, right hand side mammary fat pad of 50 female NSG mice; mice were 47 days old at time of inoculation. Cells were suspended in a 1:1 ratio of PBS and Matrigel; 50 µl was injected per mouse. Following inoculation, tumour size was assessed twice weekly using calipers. Tumour volume was calculated as described for PDX tumours above. Once tumours reached a range of 45–115 mm3 in volume, mice were randomised into four experimental groups: no doxycycline + vehicle control (n = 9), no doxycycline + AZD2811 NP (n = 10), doxycycline + vehicle control (9) or doxycyline + AZD2811 NP (n = 9). Randomisation to treatment groups was performed using a random number generator. Doxycline was delivered via the feed ad libitum using modifed PicoLab® Rodent Diet 20 (5053) containing 0.625 g/kg doxycycline (0.0625%; LabDiet®). Mice on the no doxycycline arms received PicoLab® Rodent Diet 20 (5053; LabDiet®). Mice received doxycycline feed for 24 h (Day -1) before the first dose of AZD2811 NP or vehicle control was delivered (Day 0). Mice were treated once weekly by intravenous injection with 25 mg/kg AZD2811 NPs, diluted in saline, or with a corresponding volume of saline vehicle control. Mice were culled once tumours reached 13 mm in diameter. The antitumour activity and survival was assessed as described for PDX experiments above.
Statistical analysis
All data is reflective of at least 3 biological replicates, unless otherwise stated. Data in which individual cells were scored was analysed using non-parametric t-tests (Mann-Whitney) or non-parametric one-way ANOVA. Dose response curves were analysed using 2-way ANOVA. Summary of all statistics used can be found in Source Data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Mass Spectrometry data have been deposited in PRIDE, accession number PXD023666. The publicly available TCGA data were downloaded from gdac.broadinstitute.org (release 28 January 2016). All other data are provided in the manuscript or Supplementary Information. Source data for all Figures are provided in the Source Data file accompanying the manuscript. Source data are provided with this paper.
References
Lee, A. & Djamgoz, M. B. A. Triple negative breast cancer: emerging therapeutic modalities and novel combination therapies. Cancer Treat. Rev. 62, 110–122 (2018).
Tutt, A. N. J. et al. Adjuvant olaparib for patients with BRCA1- or BRCA2-mutated breast cancer. N. Engl. J. Med 384, 2394–2405 (2021).
Zimmer, A. S., Gillard, M., Lipkowitz, S. & Lee, J. M. Update on PARP inhibitors in breast cancer. Curr. Treat. Options Oncol. 19, 21 (2018).
Dirix, L. Y. et al. Avelumab, an anti-PD-L1 antibody, in patients with locally advanced or metastatic breast cancer: a phase 1b JAVELIN Solid Tumor study. Breast Cancer Res. Treat. 167, 671–686 (2018).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Sci. (N. Y.) 347, 1260419 (2015).
Daniel, K. et al. Meiotic homologue alignment and its quality surveillance are controlled by mouse HORMAD1. Nat. Cell Biol. 13, 599–610 (2011).
Watkins, J. et al. Genomic complexity profiling reveals that HORMAD1 overexpression contributes to homologous recombination deficiency in triple-negative breast cancers. Cancer Discov. 5, 488–505 (2015).
Liu, K. et al. Aberrantly expressed HORMAD1 disrupts nuclear localization of MCM8-MCM9 complex and compromises DNA mismatch repair in cancer cells. Cell Death Dis. 11, 519 (2020).
Wang, X. et al. Epigenetic activation of HORMAD1 in basal-like breast cancer: role in Rucaparib sensitivity. Oncotarget 9, 30115–30127 (2018).
Nichols, B. A. et al. HORMAD1 is a negative prognostic indicator in lung adenocarcinoma and specifies resistance to oxidative and genotoxic stress. Cancer Res. 78, 6196–6208 (2018).
Gao, Y. et al. The Cancer/Testes (CT) antigen HORMAD1 promotes homologous recombinational DNA Repair and Radioresistance in Lung adenocarcinoma cells. Sci. Rep. 8, 15304 (2018).
Rosenberg, S. C. & Corbett, K. D. The multifaceted roles of the HORMA domain in cellular signaling. J. Cell Biol. 211, 745–755 (2015).
Saurin, A. T. Kinase and phosphatase cross-talk at the kinetochore. Front. Cell dev. Biol. 6, 62 (2018).
Pachis, S. T. & Kops, G. Leader of the SAC: molecular mechanisms of Mps1/TTK regulation in mitosis. Open Biol. https://doi.org/10.1098/rsob.180109 (2018).
Hindriksen, S., Lens, S. M. A. & Hadders, M. A. The ins and outs of aurora B inner centromere localization. Front. Cell Dev. Biol. 5, 112 (2017).
Saurin, A. T., van der Waal, M. S., Medema, R. H., Lens, S. M. & Kops, G. J. Aurora B potentiates Mps1 activation to ensure rapid checkpoint establishment at the onset of mitosis. Nat. Commun. 2, 316 (2011).
Nezi, L. & Musacchio, A. Sister chromatid tension and the spindle assembly checkpoint. Curr. Opin. Cell Biol. 21, 785–795 (2009).
Krenn, V. & Musacchio, A. The Aurora B Kinase in chromosome Bi-orientation and spindle checkpoint signaling. Front. Oncol. 5, 225 (2015).
Baron, A. P. et al. Probing the catalytic functions of Bub1 kinase using the small molecule inhibitors BAY-320 and BAY-524. Elife https://doi.org/10.7554/eLife.12187 (2016).
Ikeda, M. & Tanaka, K. Plk1 bound to Bub1 contributes to spindle assembly checkpoint activity during mitosis. Sci. Rep. 7, 8794 (2017).
Bharadwaj, R. & Yu, H. The spindle checkpoint, aneuploidy, and cancer. Oncogene 23, 2016–2027 (2004).
Anderhub, S. J. et al. High proliferation rate and a compromised spindle assembly checkpoint confers sensitivity to the MPS1 inhibitor BOS172722 in triple-negative breast cancers. Mol. Cancer Ther. 18, 1696–1707 (2019).
Oser, M. G. et al. Cells lacking the RB1 tumor suppressor gene are hyperdependent on aurora B kinase for survival. Cancer Discov. 9, 230–247 (2019).
Siemeister, G. et al. Inhibition of BUB1 kinase by BAY 1816032 Sensitizes Tumor Cells toward Taxanes, ATR, and PARP Inhibitors In Vitro and In Vivo. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 25, 1404–1414 (2019).
Noordermeer, S. M. et al. The shieldin complex mediates 53BP1-dependent DNA repair. Nature 560, 117–121 (2018).
Olivieri, M. et al. A genetic map of the response to DNA damage in human cells. Cell 182, 481–496 e421 (2020).
Zou, X. et al. Validating the concept of mutational signatures with isogenic cell models. Nat. Commun. 9, 1744 (2018).
Collin, P., Nashchekina, O., Walker, R. & Pines, J. The spindle assembly checkpoint works like a rheostat rather than a toggle switch. Nat. cell Biol. 15, 1378–1385 (2013).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer cell 33, 676–689.e673 (2018).
Tarantino, D. et al. Functional screening reveals HORMAD1-driven gene dependencies associated with translesion synthesis and replication stress tolerance. Oncogene 41, 3969–3977 (2022).
Chen, J. G., Yang, C. P., Cammer, M. & Horwitz, S. B. Gene expression and mitotic exit induced by microtubule-stabilizing drugs. Cancer Res. 63, 7891–7899 (2003).
Raaijmakers, J. A. et al. BUB1 Is Essential for the Viability of Human Cells in which the Spindle Assembly Checkpoint Is Compromised. Cell Rep. 22, 1424–1438 (2018).
Dick, A. E. & Gerlich, D. W. Kinetic framework of spindle assembly checkpoint signalling. Nat. cell Biol. 15, 1370–1377 (2013).
Heinrich, S. et al. Determinants of robustness in spindle assembly checkpoint signalling. Nat. cell Biol. 15, 1328–1339 (2013).
Marcozzi, C. & Pines, J. Assays for the spindle assembly checkpoint in cell culture. Methods Cell Biol. 144, 1–13 (2018).
Moyle, M. W. et al. A Bub1-Mad1 interaction targets the Mad1-Mad2 complex to unattached kinetochores to initiate the spindle checkpoint. J. Cell Biol. 204, 647–657 (2014).
Chao, W. C., Kulkarni, K., Zhang, Z., Kong, E. H. & Barford, D. Structure of the mitotic checkpoint complex. Nature 484, 208–213 (2012).
Tipton, A. R. et al. BUBR1 and closed MAD2 (C-MAD2) interact directly to assemble a functional mitotic checkpoint complex. J. Biol. Chem. 286, 21173–21179 (2011).
Sudakin, V., Chan, G. K. & Yen, T. J. Checkpoint inhibition of the APC/C in HeLa cells is mediated by a complex of BUBR1, BUB3, CDC20, and MAD2. J. Cell Biol. 154, 925–936 (2001).
Teo, G. et al. SAINTexpress: improvements and additional features in Significance Analysis of INTeractome software. J. Proteom. 100, 37–43 (2014).
Varadi, M. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res 50, D439–D444 (2022).
Wang, H. et al. Structural and biochemical insights into the interaction mechanism underlying HORMAD1 and its partner proteins. Structure 31, 1578–1588 e1573 (2023).
Jeyaprakash, A. A. et al. Structure of a Survivin-Borealin-INCENP core complex reveals how chromosomal passengers travel together. Cell 131, 271–285 (2007).
Samaras, P. et al. ProteomicsDB: a multi-omics and multi-organism resource for life science research. Nucleic Acids Res 48, D1153–D1163 (2020).
Moran, E. C. et al. Mitotic R-loops direct Aurora B kinase to maintain centromeric cohesion. Preprint at bioRxiv https://doi.org/10.1101/2021.01.14.426738 (2021).
Yasui, Y. et al. Autophosphorylation of a newly identified site of Aurora-B is indispensable for cytokinesis. J. Biol. Chem. 279, 12997–13003 (2004).
Welburn, J. P. et al. Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore-microtubule interface. Mol. Cell 38, 383–392 (2010).
Wilkinson, R. W. et al. AZD1152, a selective inhibitor of Aurora B kinase, inhibits human tumor xenograft growth by inducing apoptosis. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 13, 3682–3688 (2007).
Zekri, A. et al. AZD1152-HQPA induces growth arrest and apoptosis in androgen-dependent prostate cancer cell line (LNCaP) via producing aneugenic micronuclei and polyploidy. Tumour Biol.: J. Int. Soc. Oncodev. Biol. Med. 36, 623–632 (2015).
Shimomura, T. et al. MK-5108, a highly selective Aurora-A kinase inhibitor, shows antitumor activity alone and in combination with docetaxel. Mol. Cancer Ther. 9, 157–166 (2010).
Hewitt, L. et al. Sustained Mps1 activity is required in mitosis to recruit O-Mad2 to the Mad1-C-Mad2 core complex. J. Cell Biol. 190, 25–34 (2010).
Raaijmakers, J. A. & Medema, R. H. Killing a zombie: a full deletion of the BUB1 gene in HAP1 cells. EMBO J. 38, e102423 (2019).
Zhang, G. et al. Efficient mitotic checkpoint signaling depends on integrated activities of Bub1 and the RZZ complex. EMBO J. 38, EMBJ2018100977 (2019).
Kops, G. J., Foltz, D. R. & Cleveland, D. W. Lethality to human cancer cells through massive chromosome loss by inhibition of the mitotic checkpoint. Proc. Natl. Acad. Sci. USA 101, 8699–8704 (2004).
Wang, Q. et al. BUBR1 deficiency results in abnormal megakaryopoiesis. Blood 103, 1278–1285 (2004).
Dobles, M., Liberal, V., Scott, M. L., Benezra, R. & Sorger, P. K. Chromosome missegregation and apoptosis in mice lacking the mitotic checkpoint protein Mad2. Cell 101, 635–645 (2000).
Patel, N. et al. Integrated genomics and functional validation identifies malignant cell specific dependencies in triple negative breast cancer. Nat. Commun. 9, 1044 (2018).
Brough, R. et al. Identification of highly penetrant Rb-related synthetic lethal interactions in triple negative breast cancer. Oncogene 37, 5701–5718 (2018).
Suijkerbuijk, S. J. et al. Molecular causes for BUBR1 dysfunction in the human cancer predisposition syndrome mosaic variegated aneuploidy. Cancer Res. 70, 4891–4900 (2010).
Rogers, E., Bishop, J. D., Waddle, J. A., Schumacher, J. M. & Lin, R. The aurora kinase AIR-2 functions in the release of chromosome cohesion in Caenorhabditis elegans meiosis. J. Cell Biol. 157, 219–229 (2002).
Wellard, S. R., Schindler, K. & Jordan, P. W. Aurora B and C kinases regulate chromosome desynapsis and segregation during mouse and human spermatogenesis. J. Cell Sci. 133, jcs248831 (2020).
Kim, Y. et al. The chromosome axis controls meiotic events through a hierarchical assembly of HORMA domain proteins. Dev. Cell 31, 487–502 (2014).
Shin, Y. H. et al. Hormad1 mutation disrupts synaptonemal complex formation, recombination, and chromosome segregation in mammalian meiosis. PLoS Genet. 6, e1001190 (2010).
Kogo, H. et al. HORMAD1-dependent checkpoint/surveillance mechanism eliminates asynaptic oocytes. Genes Cells.: Devoted Mol. Cell. Mech. 17, 439–454 (2012).
Ferrandiz, N. et al. Spatiotemporal regulation of Aurora B recruitment ensures release of cohesion during C. elegans oocyte meiosis. Nat. Commun. 9, 834 (2018).
Bavetsias, V. & Linardopoulos, S. Aurora kinase inhibitors: current status and outlook. Front. Oncol. 5, 278 (2015).
Kovacs, A. H., Zhao, D. & Hou, J. Aurora B inhibitors as cancer therapeutics. Molecules 28, 3385 (2023).
Soria-Bretones, I. et al. The spindle assembly checkpoint is a therapeutic vulnerability of CDK4/6 inhibitor-resistant ER(+) breast cancer with mitotic aberrations. Sci. Adv. 8, eabq4293 (2022).
Cortes, J. et al. Eribulin monotherapy versus treatment of physician’s choice in patients with metastatic breast cancer (EMBRACE): a phase 3 open-label randomised study. Lancet 377, 914–923 (2011).
Meerbrey, K. L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl. Acad. Sci. USA 108, 3665–3670 (2011).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinforma. 22, 433 (2021).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Zhao, Y. et al. Automatic segmentation of cervical cells based on star-convex polygons in pap smear images. Bioengineering (Basel) 10, 47 (2022).
Meyerholz, D. K. & Beck, A. P. Principles and approaches for reproducible scoring of tissue stains in research. Lab Invest 98, 844–855 (2018).
Lal, A., Haynes, S. R. & Gorospe, M. Clean Western blot signals from immunoprecipitated samples. Mol. Cell. probes 19, 385–388 (2005).
Hillier, C. et al. Landscape of the plasmodium interactome reveals both conserved and species-specific functionality. Cell Rep. 28, 1635–1647 (2019).
Wang, R. et al. PRIDE Inspector: a tool to visualize and validate MS proteomics data. Nat. Biotechnol. 30, 135–137 (2012).
Vichai, V. & Kirtikara, K. Sulforhodamine B colorimetric assay for cytotoxicity screening. Nat. Protoc. 1, 1112–1116 (2006).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Acknowledgements
This work was funded by Breast Cancer Now through The Breast Cancer Now Toby Robins Research Centre (CTR-Q5-Y3) to ANJT and CJL, a Programme Grant from Cancer Research UK to CJL and ANJT (DRCRPGNov21y100001) and through Breast Cancer Research Foundation (BCRF-23-199) to ANJT. We thank Prof. Iain Cheeseman (MIT, Cambridge, USA) for providing phospho-DSN1 antibodies. We thank Prof. Andrew McAinsh (Univeristy of Warwick, UK) for providing BUB1 knock-out cells. We thank Prof. Keven Corbett (UCSD, USA) for providing the bacterial HORMAD1 expression construct. We thank Prof. Stephen Taylor (University of Manchester, UK) for providing the mammalian Aurora B expression construct. The authors acknowledge the expert technical training and support received from Kai Betteridge and Fredrik Wallberg in image capture as well as the support received in image processing and analysis at the Light Microscopy Facility at the ICR, Chelsea. We thank Jelena Urosevic at AstraZeneca for the provision of Aurora B inhibitor AZD2811 NP for in vivo experiments. CMG and DM were supported by Wellcome Trust grant 090532/Z/09/Z. AG and JQ were supported by Breast Cancer Research Trust (now part of Breast Cancer Now), Cancer Research UK and Breast Cancer Now (and their legacy charity Breakthrough Breast Cancer) [CRUK/07/012, KCL-BCN-Q3].
Author information
Authors and Affiliations
Contributions
Performed experiments and analysed data: C.W., G.K., D.W., J.T., R.M.H., A.M., R.F., B.N.-L., L.H., M.P.C., N.B., H.K., A.C., K.D., S.N., N.D.'U., R.L., C.S., M.O., N.I., C.M., L.Y., D.B.K., V.A., I.R. Analysed data: I.R., J.Q., A.G., R.B., J.C., S.H., S.J.P., C.J.L. Supervised the study: J.P., S.J.P., C.J.L., A.N.J.T. Manuscript drafting: C.W., G.K., D.W., S.J.P., C.J.L., A.N.J.T. All authors read and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.J.L. makes the following disclosures: receives and/or has received research funding from: AstraZeneca, Merck KGaA, Artios, Neophore. Received consultancy, SAB membership or honoraria payments from: FoRx, Syncona, Sun Pharma, Gerson Lehrman Group, Merck KGaA, Vertex, AstraZeneca, Tango, 3rd Rock, Ono Pharma, Artios, Abingworth, Tesselate, Dark Blue Therapeutics, Pontifax, Astex, Neophore, Glaxo Smith Kline, Dawn Bioventures. Has stock in: Tango, Ovibio, Hysplex, Tesselate. C.J.L. is also a named inventor on patents describing the use of DNA repair inhibitors and stands to gain from their development and use as part of the ICR “Rewards to Discoverers” scheme and also reports benefits from this scheme associated with patents for PARP inhibitors paid into CJL’s personal account and research accounts at the Institute of Cancer Research. A.N.J.T. is/has been a consultant for AstraZeneca, Merck KGaA, Artios, Pfizer, Vertex, GE Healthcare, Inbiomotion, Prime Oncology, Medscape Education, EMPartners, VJ Oncocogy, Gilead and MD Anderson Cancer Centre; has received grant/research support from AstraZeneca, Myriad, Medivation and Merck KGaA, CRUK and Breast Cancer Now; is a stockholder in Inbiomotion; and is also a named inventor on patents describing the use of PARP repair inhibitors in DNA repair deficient cancers and stands to gain from their development and use. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Gerburg Wulf, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Walker, C., Kollarovic, G., Weekes, D. et al. Tumour specific HORMAD1 expression perturbs mitotic arrest and drives sensitivity to mitotic kinase inhibitors. Nat Commun 17, 2157 (2026). https://doi.org/10.1038/s41467-026-69561-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69561-3