Abstract
Despite advances in immunotherapy, the prognosis for patients with glioblastoma (GBM) remains poor. The efficacy of GBM-targeted immunotherapies is limited by the paucity of functional T cells in the tumor microenvironment, a consequence of the local and systemic immunosuppression prevalent in patients with GBM. To overcome these challenges, here we develop a treatment strategy we term “expand and pull,” which uses systemic administration of rhIL-7-hyFc, a long-acting recombinant human interleukin-7, to increase peripheral T cell abundance (“expand”), followed by intratumoral oncolytic virus treatment to recruit these cells to the tumor microenvironment (“pull”). We show that rhIL-7-hyFc improves the efficacy of multiple oncolytic viral therapies in syngeneic immuno-resistant mouse models of glioma. Combining rhIL-7-hyFc and Zika virus (ZIKV) increases systemic and intratumoral T cell abundance, improves cytotoxic T cell function, and delays expression of inhibitory checkpoint receptors, resulting in long-term tumor-free survival. We observe similar survival efficacy in experiments using a safer, genetically modified Δ10 3′-UTR ZIKV, as well as the clinically tested oncolytic adenovirus, Delta24-RGD. Collectively, our findings demonstrate that augmentation of both the systemic and local immune responses improves the utility of GBM-targeted immunotherapies.
Similar content being viewed by others
Introduction
Glioblastoma (GBM) is the most common and aggressive primary brain tumor; median survival is approximately one and a half years1. Current standard of care includes surgical resection followed by radiotherapy and temozolomide1. Multiple phase 3 clinical trials investigating immune checkpoint blockade in GBM have failed2,3, likely because the tumor microenvironment (TME) has a paucity of functional tumor-infiltrating lymphocytes (TILs)4. Additionally, as a side effect, chemotherapy and radiation can induce prolonged systemic lymphopenia and immunosuppression. Even in the absence of treatment, brain tumors are associated with immune dysfunction and lymphopenia5. Importantly, systemic lymphopenia is associated with shorter survival in patients with GBM6,7,8,9. Addressing systemic lymphopenia and enhancing immune function within the TME may therefore be crucial for improving treatment outcomes and extending survival in patients with GBM.
Lymphopenia in patients with GBM is associated with decreased levels of interleukin-7 (IL-7)10, a homeostatic cytokine that is critical for T cell development, survival, and proliferation via activation of the JAK-STAT pathway11. Using rhIL-7-hyFc, (NT-I7, efineptakin alfa), a long-acting homodimeric recombinant human IL-7, we previously observed an increase in systemic CD4+ and CD8+ T-cell abundance in mice bearing brain tumors12. When combined with radiation, rhIL-7-hyFc treatment resulted in a modest increase in survival using two glioma models, GL261 and CT2A. However, the addition of rhIL-7-hyFc to temozolomide and radiation did not provide an additional survival benefit12. Therefore, we hypothesized that an additional intratumoral immunogenic stimulus may be needed to attract these circulating T cells into the TME.
Oncolytic viral therapies have attracted increasing interest in multiple cancers. Specifically, for the treatment of GBM, Zika virus (ZIKV)13,14, adenovirus15, herpesvirus16, measles virus17, and poliovirus18 have all been studied. Recently, a phase 1/2 clinical trial combining oncolytic adenovirus Delta24-RGD (D24-RGD, DNX-2401, tasadenoturev) with an antibody against PD-1 was found to be safe; however, it did not meet the primary efficacy endpoint of the study15, suggesting that improvements in the function of immune cells in the TME are needed. Previously, we and others demonstrated that ZIKV, a flavivirus that causes congenital brain anomalies by targeting fetal neuronal progenitor cells, has specific oncolytic activity against glioma stem cells13,19, which are a treatment-resistant subpopulation of cells within GBMs20,21,22. Beyond its direct effect on tumor cells, intratumoral ZIKV treatment of GBM prolonged survival by promoting a CD8+ T cell-mediated anti-tumor response14,19. Combining oncolytic ZIKV therapy with anti-PD-1 checkpoint inhibition provided an additive benefit, further supporting the critical role of cytotoxic T cells in the treatment efficacy conferred by ZIKV. In preparation for clinical application, we engineered an interferon (IFN) sensitized ZIKV recombinant strain, Δ10 3’-UTR ZIKV, which cannot be transmitted by mosquitoes23. This strain also proved effective, although less potent than wild-type ZIKV in mouse models of glioma14. Δ10 3’-UTR ZIKV is safe when injected into the brains of rhesus macaques 24.
Given these findings, our current study investigates whether leveraging rhIL-7-hyFc-driven T cell expansion, in combination with oncolytic ZIKV therapy, enhances an immune-mediated anti-tumor response in immunosuppressive and treatment-resistant GBM models. The underlying rationale of this therapeutic approach, which we term “expand and pull,” involves first treating the systemic immune compartment with rhIL-7-hyFc (“expand”), followed by an intratumoral oncolytic stimulus to draw these T cells into the TME (“pull”) to engage and clear cancer cells that remain after viral oncolysis. Our results demonstrate that rhIL-7-hyFc boosts the effectiveness of multiple oncolytic viral therapies. The combination of rhIL-7-hyFc and ZIKV increases T cell abundance, both systemically and locally in the TME, enhances cytotoxic T cell function, and delays the expression of inhibitory checkpoint markers, thereby prolonging tumor-free survival. Similar improvements in survival are noted with Δ10 3′-UTR ZIKV and the oncolytic adenovirus D24-RGD. Overall, our findings suggest that the “expand and pull” treatment approach could substantially improve the efficacy of immunotherapies for GBM.
Results
Expansion of peripheral T cells with rhIL-7-hyFc prior to intratumoral oncolytic ZIKV treatment improves survival
Given that T cell-mediated tumor clearance is critical for oncolytic ZIKV therapy, we hypothesized that augmenting T cell abundance with rhIL-7-hyFc prior to intratumoral ZIKV administration could further enhance efficacy. We chose the CT2A and SB28 murine glioma models for our study due to their resistance to immunotherapy and their resemblance to the human glioma microenvironment25,26,27. After confirming tumor engraftment with bioluminescence imaging, we treated mice with a subcutaneous injection of rhIL-7-hyFc on post-implantation day 7 and 10, followed by intratumoral ZIKV on day 14 (Fig. 1A). To evaluate the systemic effects of rhIL-7-hyFc in our model, we collected blood from SB28 tumor-bearing mice and quantified CD45+, CD3+, CD4+, and CD8+ lymphocytes using flow cytometry. We collected blood samples on day 0 prior to tumor implantation (baseline), and on days 7, 14, 21, and 28. Mice treated with rhIL-7-hyFc had significant increases in CD45+ leukocytes (19.1-fold, p < 0.0001; Fig. 1B) as well as CD3+ (39.8-fold, p < 0.0001; Fig. 1C), CD4+ (7.8-fold, p < 0.0001; Fig. 1D), and CD8+ (97.2-fold, p < 0.0001; Fig. 1E) T cells on day 14 compared to diluent vehicle-treated control animals. In mice implanted with CT2A tumors, treatment with either rhIL-7-hyFc or ZIKV monotherapy prolonged survival compared to control mice. Furthermore, the combination of rhIL-7-hyFc with ZIKV further improved survival compared to rhIL-7-hyFc monotherapy alone, with approximately 60% of mice surviving long-term. (Fig. 1F). Consistent with its known immunotherapy-refractory phenotype27, the SB28 glioma model did not respond to either rhIL-7-hyFc or ZIKV monotherapy. However, the combination of rhIL-7-hyFc and ZIKV treatment significantly prolonged median survival compared to all other groups tested, with 40% long-term survivors (p < 0.05; Fig. 1G). To understand the temporal dynamics of tumor growth and regression, we performed serial bioluminescence imaging and found that most tumors regressed during the first week after ZIKV treatment (Supplementary Fig. 1A). These data suggest that increasing T cell abundance with rhIL-7-hyFc prior to ZIKV administration enhances the therapeutic efficacy in both immunotherapy-resistant CT2A and SB28 tumors. To verify that our findings were not sex-specific, we repeated the survival experiment using SB28 tumors in male mice. These results confirmed that the survival benefit observed with the combination of rhIL-7-hyFc and ZIKV was also seen in male mice (Supplementary Fig. 1B). Interestingly, when the order of treatments was reversed (intratumoral ZIKV on day 7 and rhIL-7-hyFc on day 8, Supplementary Fig. 1C) there was a significant increase in mouse mortality approximately one week after completing treatment (Supplementary Fig. 1D, E). Plasma samples collected from these mice at this time point showed markedly elevated levels of IL-1β, IL-4, IL-5, IL-6, IL-10, TNF-α, and IFN-γ, compared to plasma from control mice treated with diluent on day 8, as well as from mice receiving two doses of rhIL-7-hyFc followed by ZIKV on day 14 (Supplementary Fig. 1F). These findings suggest that the mice were developing symptoms consistent with cytokine release syndrome, underscoring the importance of treatment timing in our “expand and pull” approach.
A Experimental schematic. CT2A or SB28 cells were implanted in the right striatum at day 0. Tumor engraftment was confirmed with bioluminescence imaging on day 6. Mice then received subcutaneous injections of rhIL-7-hyFc on day 7 and 10, followed by intratumoral ZIKV injection on day 14. Blood was collected weekly and B CD45+, C CD3+, D CD4+, and E CD8+ cells were quantified via flow cytometry (center line represents mean, error bars represent standard deviation). F CT2A survival curve (representative of n = 3 independent experiments). G SB28 survival curve (representative of n = 2 independent experiments). Statistical differences were determined by (B–E) two-way ANOVA test with Tukey’s posttest (*P < 0.05; **P < 0.01; ***P < 0.001), and (F and G) log-rank test (*P < 0.05; ***P < 0.001; ****P < 0.0001). Multiplicity-adjusted log-rank tests are reported in Supplementary Table 1. Source data and exact p values are provided as a Source Data file. BLI bioluminescence imaging, ZIKV = Zika virus, IL-7 = rhIL-7-hyFc.
Long-term survivors following ZIKV and rhIL-7-hyFc treatment are protected against tumor rechallenge through a CD8+ T cell dependent mechanism
To model the tumor recurrence that inevitably occurs in patients, we performed rechallenge experiments in mice that were long-term survivors of CT2A and SB28 gliomas after rhIL-7-hyFc and ZIKV treatment. For rechallenge, we implanted the homologous CT2A or SB28 tumor cells into the contralateral side of the brain 120 days after primary tumor implantation, long after ZIKV has been cleared (Fig. 2A, Supplementary Fig. 2A)13,14. All age-matched control mice died within 40 days as expected, whereas 100% of SB28 rechallenged and 65% of CT2A rechallenged long-term survivors were protected (Fig. 2B, Supplementary Fig. 2B). Bioluminescence imaging demonstrated that tumors were cleared by post-rechallenge day 21 (at day 141, Fig. 2C). To assess the differences in circulating T cells between long-term survivors and age-matched control mice during the tumor rechallenge, we obtained blood samples 3 days prior to rechallenge (baseline), as well as at 3 and 7 days post-rechallenge. At 7 days post-rechallenge, although there was no difference in total circulating CD45+ lymphocytes (Fig. 2D), rechallenged mice had increased numbers of circulating CD3+ (Fig. 2E) and CD44+ CD8+ T cells (Fig. 2F). Additionally, a higher proportion of CD8⁺ T cells in the rechallenge group expressed PD-1 (Fig. 2G), suggesting an expansion of circulating antigen-experienced CD8⁺ T cells as PD-1 is commonly upregulated upon T cell activation and is used as a marker of antigen-experienced T cells28. Fernandez-Poma et al. demonstrated that PD-1⁺ TILs exhibit enhanced anti-tumor activity and improve the efficacy of adoptive T cell therapy29, further supporting the functional relevance of PD-1 expression in our model. Given prior findings demonstrating the necessity of CD8+ T cells for long-term protection following ZIKV therapy14, we tested whether the sustained protection after rechallenge conferred by rhIL-7-hyFc and ZIKV treatment was CD8+ T cell dependent. Following tumor clearance after the first rechallenge, the surviving mice then underwent a second SB28 tumor rechallenge in the right hemisphere at day 180 in the setting of CD8+ T cell depletion. When we depleted CD8+ T cells, the protection was lost (Fig. 2B, C). To further validate this finding, we also treated a separate cohort of long-term survivors with IgG control antibody and demonstrated that these control mice retained long-term protection against rechallenge compared to their CD8+ T cell-depleted counterparts (Supplementary Fig. 2C). Together, these data demonstrate that rhIL-7-hyFc and ZIKV treatment protects against tumor rechallenge through a CD8+ T cell-mediated mechanism. Of note, the CD8 cell surface marker is also expressed on other subsets of immune cells, including NK cells, plasmacytoid dendritic cells, and a subpopulation of conventional dendritic cells. Through immune profiling via flow cytometry, we determined that anti-CD8 antibody treatment predominantly depleted CD8+ T cells (Supplementary Fig. 2D). However, there was also a reduced percentage of cDC1s in the spleen, although this reduction was not noted in the draining cervical lymph nodes (Supplementary Fig. 2E–G).
A Experimental schematic. Surviving mice from SB28 studies with ZIKV and rhIL-7-hyFc were rechallenged 120 days later with implantation of 1000 SB28 cells in the contralateral hemisphere. Age-matched tumor-naive mice served as controls. After all rechallenged mice had cleared the tumors, a second rechallenge was performed after 60 days in combination with CD8+ T cell depletion. B Survival curve of SB28 rechallenge (representative of n = 2 independent experiments). C Bioluminescence imaging of rechallenged mice compared to age-matched controls (top) and after second rechallenge in setting of CD8+ T cell depletion (bottom). Absolute number of circulating D CD45+, E CD3+, F CD44+ CD8+ and G CD8+ PD-1+ at 3 days prior, or 3 and 7 days after tumor rechallenge (each data point represents one individual mouse; control n = 9, rechallenge n = 4). Representative of n = 2 independent experiments. Bar columns represent the mean, and error bars represent the standard deviation. Statistical differences were determined by (B) log-rank test (**P < 0.01), and (D–G) two-sided student’s t-test (*P < 0.05). Source data and exact p-values are provided as a Source Data file. ZIKV = Zika virus, IL-7 = rhIL-7-hyFc.
In vivo treatment with rhIL-7-hyFc and ZIKV enhances both the quantity and quality of tumor infiltrating T cells
To determine whether pre-treatment with rhIL-7-hyFc increases T cell infiltration into the TME, we performed immunohistochemistry staining for CD8 antigen on brain sections collected on post-implantation day 21 (Fig. 3A). Tumors treated with rhIL-7-hyFc and ZIKV combination therapy had noticeably increased intratumoral CD8 antigen staining compared to rhIL-7-hyFc or ZIKV monotherapy as well as control (Fig. 3B, Supplementary Fig. 2H, I). To determine the dynamics of the intratumoral T cell population following treatment, we performed immune profiling via flow cytometry on day 17 and 21 post tumor implantation (3 and 7 days after ZIKV injection, respectively) (Fig. 3A, Supplementary Fig. 3A). Mice treated with rhIL-7-hyFc had a significantly increased number of CD3+ T cells at day 17 (4.5-fold, p < 0.0001), regardless of intratumoral ZIKV injection or PBS control. However, at day 21, the combination of rhIL-7-hyFc and ZIKV had the greatest increase in intratumoral CD3+ T cells compared to all other groups (5.4 to 42.6-fold, p < 0.05) (Fig. 3C). The numbers of tumor-infiltrating CD4+ and CD8+ T cells with different treatments followed a similar trend (10.3 to 29.6-fold increase at day 21, p < 0.01 and 4.4 to 71.4-fold increase, p < 0.05, respectively; Fig. 3D, E). This pattern of increased T cell infiltration was also observed at day 21 in the CT2A model (Fig. 3F). At day 21, rhIL-7-hyFc and ZIKV treatment also significantly increased the abundance of activated CD44+ CD8+ T cells (2.6 to 39.3-fold, p < 0.05; Fig. 3G, Supplementary Fig. 3B). Because perforin and granzyme B are critical for CD8+ T cell-mediated cell killing30,31, we performed a peptide re-stimulation assay on isolated TILs prior to quantifying expression of these effector molecules by flow cytometry. Intratumoral ZIKV treatment significantly increased the proportion of perforin- and granzyme B-expressing CD8+ T cells (1.7-fold and 3.7-fold, respectively, p < 0.001; Fig. 3G, H). Coupled with the increased number of intratumoral CD8+ T cells, these data demonstrate that the combination of rhIL-7-hyFc and ZIKV resulted in the greatest number of perforin+ effector CD8+ T cells in the TME (Supplementary Fig. 3C). The absolute numbers of granzyme B, IFN-γ, and TNF-ɑ positive CD8+ T cells were also increased following combination therapy in both SB28 and CT2A models (Supplementary Fig. 3D–I).
A Experimental schematic. Brain, draining cervical lymph node, and peripheral blood were collected on days 17 and 21. B Representative brain sections with immunohistochemistry staining for CD8. Immunophenotyping via flow cytometry of C CD3+, D CD4+, and E CD8+ T cell tumor infiltration with ZIKV and rhIL-7-hyFc treatment in SB28 tumor-bearing mice (PBS+diluent n = 9, PBS + IL7 n = 10, ZIKV+diluent n = 9, ZIKV + IL7 n = 10) and F CT2A tumor-bearing mice (PBS+diluent n = 5, PBS + IL7 n = 5, ZIKV+diluent n = 7, ZIKV + IL7 n = 8). G Absolute number of CD44+ CD8+ T cells in the tumor microenvironment. Percentage of intratumoral CD8+ T cells expressing H perforin and I granzyme B (each data point represents one individual mouse; PBS+diluent n = 6, PBS + IL7 n = 9, ZIKV+diluent n = 9, ZIKV + IL7 n = 10). J Gating of flow cytometry data on tumor-specific CD8+ T cells as defined as CD45+ CD3+ CD8+ tetramer+ in a representative brain sample among SB28-bearing mice treated with ZIKV + rhIL-7-hyFc (right) and PBS + rhIL-7-hyFc (middle). Quantification of percentage and number of tumor-specific CD8+ T cells in the K, L brain, M–P draining cervical lymph node, and Q blood following treatment with ZIKV and rhIL-7-hyFc (each data point represents one individual mouse; PBS+diluent n = 5, PBS + IL7 n = 5, ZIKV+diluent n = 5, ZIKV + IL7 n = 5). Bar columns represent the mean, and error bars represent the standard deviation. Statistical differences were determined by (C–I, K–Q) one-way ANOVA test with Tukey’s posttest (*P < 0.05; **P < 0.01; ***P < 0.001, ****P < 0.0001). Source data and exact p values are provided as a Source Data file. BLI bioluminescence imaging, ZIKV = Zika virus, IL-7 = rhIL-7-hyFc.
We also quantified various other immune populations within the TME (Supplementary Fig. 4A); this analysis showed decreases in microglia (CD45int CD11bint) and neutrophils (CD45high CD11bhigh CD88+ Ly6G+) after ZIKV treatment compared to the PBS control in SB28 tumors (Supplementary Fig. 4B). However, these myeloid cell trends were not observed in the CT2A model, suggesting that these effects may be model-specific rather than a direct result of the treatment (Supplementary Fig. 4C). Notably, there was an increase in MHC-I expression on CD31+ endothelial cells in both models (Supplementary Fig. 4D–G), suggesting a potential mechanism by which ZIKV may mediate CD8+ T cell migration into the TME, as previous studies have demonstrated that MHC-I expression on endothelial cells can serve as an antigen-specific pathway for CD8+ T cells to cross the blood-brain barrier 32.
Given the differences in IFN-γ and cytotoxic perforin and granzyme B production, we hypothesized that treatment with ZIKV increases activation and proliferation of the rhIL-7-hyFc-expanded tumor-specific CD8+ T cells. To investigate this, we compared the percentage of tumor-specific CD8+ T cells using an MHC-I tetramer flow cytometry assay (Fig. 3J, Supplementary Fig. 5A). Indeed, at day 21, rhIL-7-hyFc and ZIKV treatment resulted in the highest number of intratumoral tetramer-positive CD8+ T cells compared to all other groups (Fig. 3K, L). Furthermore, at day 17, both the total number of tetramer-positive CD8+ T cells and their relative proportion among all CD3+ T cells in the draining cervical lymph nodes were significantly increased (Fig. 3M–P), indicating early expansion of this tumor-specific CD8+ T cell population in response to treatment. By day 21, mice treated with rhIL-7-hyFc and ZIKV combination therapy had a higher percentage of tetramer-positive CD8+ T cells in the peripheral blood, suggesting increased mobilization of tumor-specific CD8+ T cells (Fig. 3Q). Moreover, there was significant expansion of the effector memory subset in both the blood and draining cervical lymph nodes following combination treatment (Supplementary Fig. 5B, C). In summary, the combination of rhIL-7-hyFc and ZIKV not only increased the numbers of tumor-specific, functionally active intratumoral CD8+ T cells but also likely contributed to the improved anti-tumor immune response.
rhIL-7-hyFc and ZIKV increases tumor-infiltrating effector memory CD8+ T cells and expands TCR clonal populations
To further investigate the effects of the combination treatment of rhIL-7-hyFc and ZIKV on TIL phenotype, we performed scRNA-seq on CD45high cells isolated from the treated tumors of mice at post-implantation day 21 (Fig. 4A). While all the major immune cell types were present in all samples, there was enrichment in T cells following rhIL-7-hyFc and ZIKV treatment, with a concomitant decrease in the proportion of macrophage and monocytes (Fig. 4B). We divided the T cells into subsets and refined our analysis to include only those cells with a complete alpha-beta TCR sequence (including VDJ genes and CDR3 region). We performed T cell annotation by aligning with the ProjecTILS ref. 33, and the annotated CD8+ and CD4+ T cells were then analyzed independently.
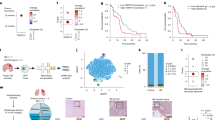
A Experimental schematic. SB28-bearing mice received rhIL-7-hyFc on post-implantation day 7 and 10, followed by intratumoral ZIKV on day 14. Brains were collected on day 21 and sorted by flow cytometry on CD45high cells for single-cell RNA sequencing (PBS+diluent n = 5, PBS + IL7 n = 5, ZIKV+diluent n = 4, ZIKV + IL7 n = 4). Created in BioRender. Chheda, M. (2026) https://BioRender.com/k7uv2i2. B UMAP representation of total cells. C UMAP representations of the CD8+ subset for each of the treatment groups. D Gene expression analysis of relevant T cell genes and pathways for each cluster. The size of the circle indicates the percentage of cells with the expression of each gene, and the color indicates the average expression across the cells in that unique cluster. E UMAP, and box plot of exhaustion score across treatment groups. F TCR clonal expansion categories superimposed onto UMAPs for each treatment and corresponding bar graphs. G, UMAP representation of CD4+ subset and bar graph. H Box plot of Tnfrsf9 expression in Treg subset. Percentage of intratumoral Tregs by flow cytometry in I SB28 (PBS+diluent n = 4, PBS + IL7 n = 5, ZIKV+diluent n = 5, ZIKV + IL7 n = 5) and J CT2A (PBS+diluent n = 4, PBS + IL7 n = 5, ZIKV+diluent n = 8, ZIKV + IL7 n = 7) models (each data point represents one individual mouse). Bar columns represent the mean, error bars represent the standard deviation. For box plots (E, H), center line = median, box limits = upper and lower quartiles, whiskers = 1.5x interquartile range, points = outliers. Statistical differences were determined by (I, J) one-way ANOVA test with Tukey’s posttest (*P < 0.05; **P < 0.01; ***P < 0.001, ****P < 0.0001). Source data and exact p-values are provided as a Source Data file. BLI bioluminescence imaging, ZIKV = Zika virus, IL-7 = rhIL-7-hyFc.
CD8+ T cells were classified as naive-like, early-activated, effector memory, progenitor-exhausted (Tpex), or terminally-exhausted (Tex) based on transcriptional patterns (Fig. 4C) and expression of cell-type markers (Fig. 4D). The naive-like population expressed the naive T cell markers SELL, IL-7R, Ccr7, and Tcf7, whereas the effector-memory population expressed the effector molecules GzmB and GzmK. The early-activated population represents an intermediate population and expresses features of both naive and effector states. The Tex population expressed canonical markers of exhaustion, including Tigit, Tox, and Havcr2, whereas the Tpex population expressed both exhaustion markers and some features of the naive and effector populations, such as Ccr7 and Ifng. Comparing treatment groups, the predominant CD8+ T cell population in tumors following rhIL-7-hyFc and ZIKV treatment was the effector memory population with a relative decrease in the naive and exhausted populations (Fig. 4C). This correlates with an overall decrease in expression of exhaustion-related genes as represented by the Exhaustion Score (factoring expression of Havcr2, Tox, Tigit, and Lag-3) (Fig. 4E). ZIKV treatment alone demonstrated a similar increase in the effector population with a decrease in exhaustion marker expression. In contrast, rhIL-7-hyFc treatment alone expanded the naive and Tpex populations. To better understand the reactivity of these CD8+ T cells, we analyzed the clonality of their TCR sequences. Following treatment with rhIL-7-hyFc and ZIKV or ZIKV alone, there was a decrease in the proportion of unique clonotypes and an increase in the proportion of medium and large clonal populations, particularly in the combination treatment group (Fig. 4F). This suggests clonal expansion of CD8+ T cell populations following rhIL-7-hyFc and ZIKV treatment.
Next, given the impact of CD4+ T cells on intratumoral CD8+ T cell function and exhaustion25, we analyzed CD4+ T cell subsets, including naive-like, helper-1 (Th1), follicular helper (Tfh), or regulatory (Treg). Of note, we observed minimal expression of markers associated with Th2 and Th17 subsets. Comparing treatment groups, the combination of rhIL-7-hyFc and ZIKV had the highest percentage of Th1 CD4+ T cells and the lowest percentage of Tregs (Fig. 4G). rhIL-7-hyFc treatment alone had a higher percentage of naive-like cells, as was observed for CD8+ T cells. We then analyzed the Treg population to compare gene expression associated with Treg activity. Tnfrsf9, the gene encoding costimulatory receptor 4-1BB, is a marker of activated (immunosuppressive) Tregs34,35. We found the lowest Tnfrsf9 expression in the combination treatment group compared to either monotherapy (Fig. 4H). We further validated these findings using flow cytometry, which confirmed that mice treated with ZIKV had a reduced proportion of Tregs in both the SB28 and CT2A models (Fig. 4I, J). Taken together, our findings indicate that treatment with rhIL-7-hyFc and ZIKV leads to a marked increase in intratumoral effector CD4+ and CD8+ T cells and a decrease in CD8+ exhausted T cells and Treg cells.
rhIL-7-hyFc-treated CD8+ T cells have increased IFN-γ secretion and delayed expression of inhibitory checkpoint receptors PD-1, TIM-3, and LAG-3
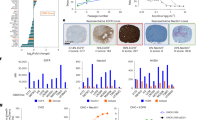
To further evaluate the effect of rhIL-7-hyFc treatment on T cell function in the TME, we co-cultured CD3+ TILs collected on post-implantation day 21 with SB28 or CT2A tumor cells ex vivo. CD3+ TILs from tumors treated with rhIL-7-hyFc and ZIKV had increased IFN-γ, IL-2, and TNF-ɑ secretion compared to TILs isolated from brains of mice treated with only ZIKV (Fig. 5A). In addition, both SB28 and CT2A tumor cells, which are constitutively MHC-I negative27, exhibited an increased upregulation of MHC-I when co-cultured with CD3+ TILs from tumors treated with rhIL-7-hyFc and ZIKV, compared to TILs from tumors treated with ZIKV alone (Fig. 5B). The addition of an IFN-γ blocking antibody abolished this effect, suggesting that the combination of rhIL-7-hyFc and ZIKV may sensitize immunologically ‘cold’ tumors to CD8+ effector T cells through an IFN-γ-dependent mechanism (Fig. 5C, D). We did not observe changes in MHC-II expression in either model (Supplementary Fig. 5D, E). Given the difference in IFN-γ and overall cytokine production by rhIL-7-hyFc and ZIKV-treated CD3+ TILs, we assessed CD8+ T cells for expression of inhibitory immune receptors including PD-1, TIM-3, and LAG-3 (Supplementary Fig. 6A). At post-implantation day 17, mice treated with rhIL-7-hyFc had a decreased proportion of PD-1+ CD8+ T cells compared to those treated with diluent control (Fig. 5E, Supplementary Fig. 6B). However, at day 21, this effect was lost, and all groups had a similar percentage of PD-1+ CD8+ T cells. Changes in the expression of TIM-3 and LAG-3 followed a similar trend (Fig. 5F, G, Supplementary Fig. 6C, D). These data suggest that pretreatment with rhIL-7-hyFc improves tumor-infiltrating T cell cytokine production and delays expression of inhibitory checkpoint receptors. Additionally, at day 21, we detected a significantly increased number of Tpex (PD-1+ TCF-1+ TIM-3-) in the brain in the combination group compared to all other groups (Supplementary Fig. 6E, F). We did not detect significant differences in T cell populations in the draining cervical lymph nodes at this time point (Supplementary Fig. 6G). In systemic circulation at day 21, mice treated with rhIL-7-hyFc and ZIKV had greater numbers of CD3+, CD4+, and CD8+ T cells (Fig. 5H). However, the percentage of PD-1+, TIM-3+, and LAG-3+ CD8+ T cells was also highest in this group (Fig. 5I). Overall, these findings suggest that the combination of rhIL-7-hyFc and ZIKV enhances T cell functionality and promotes a pro-inflammatory response in TME, while also delaying the upregulation of inhibitory checkpoint receptors.
A Multiplex cytokine array of tumor-infiltrating CD3+ T cells collected from rhIL-7-hyFc and ZIKV-treated brains on day 21 compared to those from ZIKV treatment alone (ZIKV+diluent n = 2, ZIKV + IL7 n = 2). B Representative flow cytometry histograms of MHC I and MHC II expression on SB28 cells. C Quantification of C SB28 and D CT2A tumor cell MHC I expression when co-cultured with CD3+ TILs from rhIL-7-hyFc and ZIKV-treated mice, as well as in the setting of IFN-γ blocking antibody (each data point represents one experimental replicate, n = 3). Percentage of E PD-1+, F TIM-3+, and G LAG-3+ CD8+ T cells in the brain at day 17 and 21 (each data point represents one individual mouse; PBS+diluent n = 9, PBS + IL7 n = 10, ZIKV+diluent n = 9, ZIKV + IL7 n = 10). H Absolute number of circulating CD3+, CD4+, and CD8+ T cells following rhIL-7-hyFc and ZIKV treatment (each data point represents one individual mouse). I Percentage of PD-1+, TIM-3+, and LAG-3+ CD8+ T cells in circulation at day 21 (each data point represents one individual mouse; PBS+diluent n = 8, PBS + IL7 n = 9, ZIKV+diluent n = 8, ZIKV + IL7 n = 10). Bar columns represent the mean, and error bars represent the standard deviation. Statistical differences were determined by (C–I) one-way ANOVA test with Tukey’s posttest (*P < 0.05; **P < 0.01; ***P < 0.001, ****P < 0.0001). Source data and exact p-values are provided as a Source Data file. ZIKV = Zika virus, IL-7 = rhIL-7-hyFc.
PD-1 checkpoint blockade further improves the efficacy of rhIL-7-hyFc and ZIKV combination therapy
Given the increased expression of inhibitory markers on CD8+ T cells at post-implantation day 21 compared to day 17, we hypothesized that the addition of an immune checkpoint inhibitor during this window might further increase treatment efficacy. Using the SB28 glioma model, which is resistant to immune checkpoint blockade alone27, we treated tumor-bearing mice with rhIL-7-hyFc and ZIKV, with or without anti-PD-1 and its respective isotype control (Fig. 6A). Triple therapy with rhIL-7-hyFc + ZIKV + anti-PD-1 was superior to rhIL-7-hyFc + ZIKV, rhIL-7-hyFc + anti-PD-1, and ZIKV + anti-PD-1 treatments, leading to an 80% long-term survivor rate (Fig. 6B). Thus, the efficacy of the rhIL-7-hyFc and ZIKV treatment regimen can be augmented further with immune checkpoint inhibitors.
A Experimental schematic. In addition to the previously described treatment paradigm, anti-PD-1 antibody injections were started on day 15. B Survival curve of ZIKV, rhIL-7-hyFc, and anti-PD-1 treatment (representative of n = 2 independent experiments). C Survival curve of Δ10 3′-UTR ZIKV, rhIL-7-hyFc, and anti-PD-1 treatment (representative of n = 2 independent experiments). D Survival curve of D24-RGD adenovirus, rhIL-7-hyFc, and anti-PD-1 treatment (representative of n = 2 independent experiments). Statistical differences were determined by (B–D) log-rank test (*P < 0.05; **P < 0.01; ***P < 0.001, ****P < 0.0001). Log-rank test and False Discovery Rate corrected p values are reported in Supplementary Tables 2–4. Source data are provided as a Source Data file. Abbreviations: BLI = bioluminescence imaging, ZIKV = Zika virus, IL-7 = rhIL-7-hyFc.
“Expand and pull” treatment paradigm works with immune-sensitized ZIKV and D24-RGD adenovirus
To determine whether the “expand and pull” treatment approach is generalizable, we also tested two other oncolytic viruses (Supplementary Fig. 7A–C). As safety is a priority in any future clinical application of ZIKV, we previously engineered a deletion of 10 nucleotides in the 3′ untranslated region (Δ10 3′-UTR ZIKV) of the ZIKV-Dakar cDNA clone14, resulting in an immune-sensitized virus that cannot be transmitted by mosquitoes and has a reduced ability to antagonize cell-intrinsic innate immune responses23. Treatment of SB28 tumor-bearing mice with rhIL-7-hyFc and Δ10 3’-UTR ZIKV was superior to either treatment alone (Fig. 6C). Adding an immune checkpoint inhibitor, triple therapy with rhIL-7-hyFc + Δ10 3’-UTR ZIKV + anti-PD-1, resulted in 67% long-term survivors. Next, we applied the paradigm to D24-RGD, a replication-competent, adenovirus that kills cancer cells and is safe for treatment in humans15,36,37,38. rhIL-7-hyFc and D24-RGD treatment improved survival outcome compared to D24-RGD treatment alone, and the addition of an antibody against PD-1 further improved the long-term survivor rate to 73% (Fig. 6D). In summary, using a highly aggressive, immunosuppressive tumor model that has a paucity of TILs in the TME similar to the human GBM context, we found that treating with rhIL-7-hyFc followed by an oncolytic virus leads to enhanced tumor clearance and improved mouse survival.
Discussion
Currently, several oncolytic viruses are being evaluated in clinical trials for GBM15,16,17,18. Many oncolytic viruses function through selective replication within cancer cells, triggering destruction of infected cells and the release of progeny virus that spreads to adjacent tumor cells. Viral infection of tumor cells also stimulates an immune response within the highly immunosuppressive ‘cold’ TME, shifting it towards a more immunologically ‘hot’ phenotype39,40. Despite these promising attributes, the clinical efficacy of oncolytic viruses in GBM has been limited by tumor heterogeneity and tumor-induced immunosuppression. Combining oncolytic virotherapy with other treatment modalities may be essential to overcome these limitations and enhance therapeutic outcomes in GBM. Our study highlights an “expand and pull” treatment approach, which augments the oncolytic and immunostimulatory effect of viral therapies, leading to improved survival in multiple immunotherapy-resistant models of GBM.
For GBM, oncolytic therapies face the challenges of an immunosuppressive TME and a dysfunctional host immune system. GBMs generally elicit few tumor-specific CD8+ T cells, and many of these exhibit an exhausted phenotype41. Overcoming these obstacles has proven difficult, as evidenced by the failure of D24-RGD plus anti-PD-1 to meet its primary efficacy endpoint15. Tying the TME with the systemic immunosuppression, intracranial tumors may prevent peripheral T cells from infiltrating tumors through bone marrow sequestration5. To address these issues, we developed a treatment strategy that first increases the abundance of circulating T cells and subsequently attracts these cells into the tumor to engage cancer cells. Recently, Campian et al. demonstrated that systemic administration of rhIL-7-hyFc increases the number of cytotoxic CD8+ T lymphocytes both systemically and within tumors12. Building upon these findings, our study incorporates an intratumoral oncolytic viral stimulus that both pulls in T cells and promotes a pro-inflammatory shift in the TME. Interestingly, we observed an increase in intratumoral T cells in rhIL-7-hyFc-treated mice following intracranial injection of PBS alone, indicating that perturbations of the blood-brain barrier (BBB) at the peak of circulating T cell abundance can facilitate T cell influx into the brain. However, without a conducive microenvironment, these T cells either exit the brain or undergo apoptosis by day 7 post-PBS injection. In contrast, the combination of rhIL-7-hyFc with ZIKV results in substantial proliferation of infiltrating T cells and a reduction of immunosuppressive T regulatory cells. These findings underscore the important role of both components of “expand and pull” for achieving therapeutic success.
We believe at least one reason for efficacy from the combination of rhIL-7-hyFc plus oncolytic virus is the increased level of T cell infiltration into the TME. In our model, ZIKV and rhIL-7-hyFc resulted in a 42.6-fold increase in the number of intratumoral T cells compared to control mice treated with intracranial PBS injection, including a 29.6-fold increase in CD4+ T cells and a 71.4-fold increase in CD8+ T cells. Compared to mice treated with ZIKV alone, those treated with rhIL-7-hyFc and ZIKV showed substantial infiltration and expansion with a 5.4-fold increase in intratumoral CD3+ T cells, a 10.3-fold increase in CD4+ T cells, and a 4.4-fold increase in CD8+ T cells. In comparison to other preclinical models, our approach appears to recruit more T cells, particularly CD8+ T cells, into the TME. For example, immunoprofiling analyses after treatment with oncolytic D24-RGD adenovirus42 and MV-s-NAP-uPA measles virus43 monotherapy revealed CD8+ TIL counts ranging from 1000 to 4000 and 10,000 to 30,000 cells per brain, respectively. In comparison, treatment with rhIL-7-hyFc and ZIKV resulted in a higher range of 60,000 to 170,000 T cells per brain.
The improved efficacy observed likely involves not only an increase in the number of intratumoral T cells but also a boost in the anti-tumor effector subset. In our study, pretreatment with rhIL-7-hyFc generated a substantial pool of circulating effector CD8+ T cells, which were subsequently recruited into the TME by oncolytic ZIKV. Of note, this approach bypasses concerns related to the BBB permeability of the dimerized cytokine, as T cells are exposed to rhIL-7-hyFc prior to entering the tumor site. These rhIL-7-hyFc-stimulated CD8+ T cells exhibited increased tumor-specificity, TCR clonality, expression of perforin and granzyme B, and production of IFN-γ, as well as delayed expression of inhibitory receptors such as PD-1, TIM-3, and LAG-3. Beyond the effects on cytotoxic T cells, treatment with rhIL-7-hyFc and ZIKV also induced a pro-inflammatory shift in the TME, characterized by increased helper CD4+ T cells, reduced Tregs, and decreased Treg activation. Collectively, these findings suggest that this therapeutic strategy has the potential to counteract the immunosuppressive effects of GBM, both at systemic levels and within the local TME, thus improving the anti-tumor response.
Although our study primarily examines the effects on T cells, particularly the CD8+ subset, it is important to note that the IL-7 receptor is also expressed on B cells, NK cells, and, to a lesser extent, dendritic cells, monocytes, and macrophages44. Further research is needed to investigate the impact of rhIL-7-hyFc on these additional cell types in our treatment paradigm. Another limitation of this study is that we focused exclusively on the IL-7 signaling axis, as we had previously established that rhIL-7-hyFc could independently support systemic T cell expansion in similar glioma models12. Moreover, clinical use of rhIL-7-hyFc has been effective at restoring lymphocyte counts in patients without serious adverse effects45,46,47. Other cytokines such as IL-2, IL-12, and IL-15 have also been shown to be important for T cell proliferation and function. However, systemic IL-2 and IL-12 treatment in clinical settings has been associated with dose-limiting toxicities48,49, and the practical use of IL-15 has been difficult as it requires a continuous intravenous infusion50. Recent advances in bioengineering offer potential to improve cytokine stability and safety51,52,53, encouraging further exploration of these synthetic cytokines within our treatment framework.
In summary, our findings demonstrate that rhIL-7-hyFc markedly increases tumor T cell infiltration and enhances anti-tumor activity in vivo, leading to substantial improvements in survival. Moreover, this “expand and pull” strategy has potential to generate an environment where endogenous T cells more effectively target tumor cells, though additional research on other immune subsets within the TME is needed. As more oncolytic viruses undergo clinical trials for GBM, our study highlights rhIL-7-hyFc as a clinic-ready adjunct to augment the anti-tumor immune response.
Methods
All animal procedures were performed in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the NIH (National Academies Press, 2011). Tumor growth/burden was monitored regularly, and animals were euthanized upon reaching predefined humane endpoints, including severe or persistent neurological deficits, significant body weight loss (>10–15%), impaired mobility or inability to access food and water, or other signs of pain or distress. The protocols were approved by the Institutional Animal Care and Use Committee at the Washington University School of Medicine (protocol 24-0229).
Cell lines
The murine CT2A glioma cell line was purchased from Millipore Sigma. The murine SB28 glioma cell line was generously provided by Dr. Hideho Okada (University of California, San Francisco). CT2A cells were cultured in DMEM (Invitrogen, Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (FBS) (Gibco) and 1% penicillin G-streptomycin (Corning) and SB28 cells were cultured in RPMI media supplemented with 10% FBS, 1% pen/strep, 1% HEPES (Corning), 1% non-essential amino acid (Corning), and 1% sodium pyruvate (Corning). Cells were maintained at 37 °C in an incubator with 5% humidified CO2. All cell lines were routinely tested for mycoplasma contamination by PCR and authenticated by Short Tandem Repeat (STR) profiling (Genome Technology Access Center, Washington University).
ZIKV, Δ10 3′-UTR-ZIKV, and D24-RGD virus propagation
Mouse-adapted ZIKV (Dakar strain) and Δ10 3′-UTR-ZIKV viral stocks were propagated in Vero cells after inoculating at an MOI of 0.01 and incubating for 72 h, as previously described14,54. ZIKV and Δ10 3′-UTR-ZIKV viral titers were determined by plaque assay as previously described55. D24-RGD was propagated in A549 cells after inoculating at an MOI of 0.01, as previously described37. D24-RGD viral titer was assessed in HEK293 cells using the Adeno-X Rapid Titer Kit (Takara, 632250).
rhIL-7-hyFc
rhIL-7-hyFc (NT-I7, efineptakin alfa) was generously provided by NeoImmuneTech, Inc. (Rockville, MD).
In vivo mouse experiments
Single-cell suspensions of CT2A (4×104 cells in 3 μL) or SB28 cells (1×103 cells in 3 μL) were implanted into the right cerebral hemisphere of 8- to 9-week-old C57BL6/J female and male mice (Jackson Laboratory) after mice were anesthetized with ketamine (10 mg/kg), xylazine (100 mg/kg), and buprenorphine SR (1 mg/kg). The intracranial tumor implantations were performed as previously described14. Mice were stereotactically injected with CT2A or SB28 cells in the right hemisphere with the following sets of coordinates: 2.5 mm deep and 2.5 mm lateral to bregma; or 2.5 mm deep and 2.5 mm lateral and 1.5 mm anterior to lambda. Tumor engraftment was confirmed on day 6 with bioluminescent imaging, and mice were randomly assigned to treatment groups by total photon flux. Bioluminescence images were captured on a Spectrum In Vivo Imaging System (PerkinElmer), and total bioluminescence was analyzed using Living Image software. When using cell lines without a luciferase tag, mice were cage randomized prior to starting treatment. rhIL-7-hyFc (10 mg/kg) was injected subcutaneously on days 7 and 10. On day 14 after tumor implantation, using the same burr hole, mice were treated with one intracranial injection of 1 × 105 PFU ZIKV, 1 × 106 PFU Δ10 3′-UTR-ZIKV, or 3 × 108 PFU Delta24-RGD, each in 10 µL. Checkpoint blockade antibodies against PD-1 (clone RMP1-14, Leinco Technologies), or corresponding IgG2a control (clone 1-1, Leinco Technologies), were injected intraperitoneally on post-implantation days 15, 17, 19, and 21 at a dose of 10 mg/kg. For rechallenge experiments, long-term surviving mice were implanted with homologous CT2A or SB28 tumor cells into the contralateral side of the brain 120 days after primary tumor implantation. For CD8+ T cell depletion experiments, we injected depleting antibodies (clone 2.43, Bio X Cell) intraperitoneally starting the day before tumor rechallenge with an initial dose of 25 mg/kg and followed with booster doses of 12.5 mg/kg every 5 days until post-rechallenge day 9. Representative mice were bled to confirm depletion. Mice were monitored daily for signs of neurological impairment and were euthanized when moribund. Animal caretakers were blinded to treatments.
Immunohistochemistry
Brains were collected on post-implantation day 21, fixed in 4% paraformaldehyde (Electron Microscopy Sciences), embedded in paraffin after graded ethanol dehydration, and cut into 5 μm thick sections. Tissue sections were loaded into BOND (Leica Biosystems) for staining with CD8 (Abcam, AB217344, dilution 1:1000). Slides were counter-stained with hematoxylin and then scanned by Axio Scan Z1 (Zeiss). Images were analyzed using HALO software (Zeiss).
Flow cytometry
Mice were anesthetized with ketamine (10 mg/kg) and xylazine (100 mg/kg). Blood was collected, and then mice were perfused with DPBS (Gibco) prior to the collection of the brain and draining cervical lymph nodes. For peripheral blood evaluation, 50 μl of blood was added to 1 ml of ACK lysis buffer and incubated for 10 min at room temperature, followed by a single wash and addition of Fc block (BioLegend, 156604), and then antibody staining. Brain and lymph nodes were macerated over a 70 μm cell strainer using the piston from a 3 ml syringe to create single cell suspensions. Myelin was removed from brain samples using a Percoll Plus (Cytiva, 17544501) gradient. Cell suspensions were then washed once prior to Fc block and antibody staining. For the peptide re-stimulation assay, SB28 TILs were cultured with 25 μM luciferase-peptide (ABI Scientific, LMYRFEEEL) and 25 μM GFP-peptide (ABI Scientific, DTLVNRIEL); CT2A TILs were cultured with 25 μM mEpb4 (ELEQFESTIGFKLPNLRAAKRLWK), 25 μM mPomgnt1 (ECIIPDVSLSYHFGIVGLNMNGYF), and 25 μM mPlin2 (QQLQTTCQTVLVNAQRLPQNIQDQA)56. Brefeldin A (Biolegend, 420601) was added, and TILs were cultured for 4 h at 37 °C prior to antibody staining. For MHC-I tetramer staining, class 1 monomers (H-2Db or H-2Kb) were loaded with the aforementioned peptides (Immunomonitoring Lab, Washington University) and incubated with PE-streptavidin (Biolegend, 405203) and D-biotin (Invitrogen, B20656) to generate PE-conjugated tetramers. Cells were incubated with PE-conjugated tetramers (dilution 1:50) for 15 min at 37 °C prior to other antibody staining.
Detailed phenotyping of murine lymphocytes was performed using the following antibodies (all against mouse antigens): Fixable Viability Dye-eFluor506 (Invitrogen, 65086614), CD45-APC-eFluor780 (Thermo Fisher Scientific, 47-0451-82), CD3-BV711(Biolegend, 100241), CD4-PE-Cy7 (Biolegend, 116016), CD8a-BUV395 (BD Biosciences, 563786), CD11b-BV421 (Biolegend, 101236), CD11c-BV605 (Biolegend, 117334), CD19-APC (Biolegend, 115512), CD26-PE-Cy7 (Biolegend, 137810), CD31-PE-Cy5 (Thermo Fisher Scientific, 15-0311-82), CD44-PE-Cy5 (Biolegend, 103010), CD44-BV786 (BD Biosciences, 563736), CD62L-AF700 (Biolegend, 104426), CD88-PerCP-eFluor710 (Thermo Fisher Scientific, 46-0882-82), CD127-PE/Dazzle594 (Biolegend, 135032), FoxP3-PE-Cy5 (Thermo Fisher Scientific, 15-5773-82), Perforin-PE (Biolegend, 154306), Granzyme B-BV421 (Biolegend, 396414), TNF-α-BV605 (Biolegend, 506329), interferon-γ-APC (Biolegend, 505810), PD-1-AF647 (Biolegend, 135230), CTLA-4-PE/Dazzle594 (Biolegend, 106318), TIM-3-BV605 (Biolegend, 119721), LAG-3-BV421 (Biolegend, 125221), TCF-1-PE (Cell Signaling, 14456S), TIGIT-BV786 (BD Biosciences, 744215), Ly-6C-PE/Dazzle594 (Biolegend, 128044), Ly-6G-BV785 (Biolegend, 127645), NK1.1-AF700 (Biolegend, 108730), H-2Db-PE (Biolegend, 111508), I-A/I-E-BUV395 (BD Biosciences, 743876), Ki67-PerCP-eFluor710 (Thermo Fisher Scientific, 46-5698-82), LIVE/DEAD-Fixable Blue (1:400, Thermo, L34961), CD45-BUV395 (Thermo, 363-0451-80), CD11b-BV570 (Biolegend, 101233), CD11c-PE-Cy5 (1:400, Biolegend, 117316), I-A/I-E-BUV496 (1:400, Thermo, 364-5321-80), Ly6C-AF700 (1:800, Biolegend, 128023), Ly6G-APC-Fire750 (Biolegend, 127651), CD26-PE-Cy7 (Biolegend, 137809), CD88-BUV805 (BD Biosciences, 748611), XCR1-BV421 (Biolegend, 148216), SIRPα-BUV737 (BD Biosciences, 741819), PDCA1-Spark-UV387 (Biolegend, 127044), B220-PE-Fire810 (1:400, Biolegend, 103287), CD19-PE-Ccy5.5 (1:400, Thermo, 35-0193-82), CD3-Spark-NIR685 (1:400, Biolegend, 100261), CD4-BV510 (Biolegend, 100553), CD8-BUV615 (Thermo, 366-0081-82), NK1.1-BUV661 (Thermo, 376-5941-80), TCRγδ (BD Biosciences, 748993). Cells were stained with fluorochrome-conjugated anti-mouse antibodies at a dilution of 1:200 unless otherwise noted. Corresponding isotype controls were used for setting population gates. For all staining procedures, cells were fixed (Biolegend, 420801), and permeabilization (Biolegend, 421002) was performed if intracellular staining was necessary. Absolute cell counts were determined using TruCount beads (BD Biosciences, 340334) or cytometer event counts. All samples were run on either an LSRFortessa X-20 (BD Biosciences) or an Aurora (Cytek Biosciences). Data analysis was performed using FlowJo v10.10.0 (BD Biosciences).
Single-cell RNA sequencing
For scRNA-seq experiments, mice were euthanized on post-implantation day 21. Tumors were harvested, dissociated, and filtered through a 70-micron cell strainer. A Percoll Plus (Cytiva, 17544501) density gradient was used for myelin removal. Live CD45-high cells were sorted on a BD FACS ARIA-II flow sorter. Single-cell RNA libraries were generated using the Chromium Next GEM Single Cell 5’ library and Gel Bead Kit v1.1; T Cell Receptor (TCR) enrichment was performed with the Chromium Single Cell V(D)J Enrichment Kit for Mouse T cells as per manufacturer’s protocol (10X Genomics). Library quality and concentration were determined using a Bioanalyzer Tapestation (Agilent) for library sequencing on the NovaSeq 6000 instrument at a sequencing depth of ~50 K read pairs per cell. CellRanger v8.0.0 (10X Genomics) was used to process, align, and generate feature-barcode matrices against the mouse reference genome. Transcriptomic data were analyzed with the Seurat v5 R package57. Quality controls eliminated cells with the top 1% or bottom 7% of features/counts or with >5% mitochondrial content. Presumed doublets were identified with DoubletFinder58 and removed. Normalization/variance stabilization was performed with SCTransform59, and the datasets were integrated with Harmony60. Annotation of total immune cells was performed manually by the identification of canonical genes. T cell annotation was performed by alignment with the ProjecTILs reference33. The Exhaustion Score was calculated as a module score within Seurat and included the features Havcr2, Tox, Tigit, and Lag3. V(D)J libraries were processed with CellRanger V(D)J v8.0.0(10X Genomics) and mapped onto a mouse V(D)J reference genome. Clonotype analysis was performed with scRepertoire61, and TCR were defined by “strict” criteria, including VDJC genes and CDR3 sequence. Results were visualized with the dittoSeq package 62.
The following publicly available pipelines were utilized for analysis: standard preprocessing, quality control, SCTransform, Harmony Integration, and cluster analysis of scRNAseq data was based on Seurat v5 vignettes (https://satijalab.org/seurat/), doublet identification was based on the DoubletFinder tutorial (https://rpubs.com/kenneditodd/doublet_finder_example), VDJ analysis was based on scRepertoire vignettes (https://www.borch.dev/uploads/screpertoire/), T cell subset analysis was based on the ProjecTILs tutorial (https://carmonalab.github.io/ProjecTILs.demo/tutorial.html), and visualization was based on the dittoSeq tutorial (https://bioconductor.org/packages/release/bioc/vignettes/dittoSeq/inst/doc/dittoSeq.html).
Ex vivo SB28-TIL co-culture
Tumor-infiltrating lymphocytes were collected from mice on post-implantation day 21 and processed as described above. Live CD3+ cells were sorted on a BD FACS ARIA-II flow sorter. 10,000 CD3+ T cells were co-cultured with 5000 SB28 cells for 72 h. Mouse CD3/CD28 Dynabeads (Gibco, 11452D) were added to the co-culture per the manufacturer's protocol for T cell stimulation. Anti-mouse IFN-γ (Leinco, I-1119) or IgG1 control (Leinco, I-1195) was added at 10 μg/ml. After 72 h, supernatant was collected for multiplex cytokine ELISA (V-PLEX Proinflammatory Panel 1 Mouse Kit, Meso Scale Diagnostics, K15048D-1), which was performed per manufacturer protocol on a MESO QuickPlex SQ 120 (Meso Scale Diagnostics). T cells were removed via multiple washes and SB28 cells were trypsinized and collected for flow cytometric analysis.
In vitro oncolytic viral kill assay
Wild-type ZIKV, Δ10 3′-UTR-ZIKV, and D24-RGD were incubated with 250 SB28 cells, plated 24 hours in advance, using a range of multiplicity of infection (MOI). After 72 hours, cell viability was assessed using CellTiter-Glo Luminescent Assay (Promega, G7572) per manufacturer protocol.
Statistical analysis
Data plots were generated using Prism 10.3.1 (GraphPad). All data are representative of at least two independent experiments, unless specifically noted. The determination of sample size and data analysis for this study followed the general guidelines for animal studies. Mouse survival studies were analyzed using log-rank tests. To account for multiple comparisons, we also conducted multiplicity-adjusted log-rank tests employing the Benjamini-Hochberg procedure for false discovery rate control. All other in vivo data were summarized using means and standard deviations. The differences were compared using a two-sample Student’s t test, one-way analysis of variance (ANOVA), or two-way ANOVA as appropriate, followed by post hoc Tukey’s test for multiple comparisons for between-group differences of interest. For all analyses, significance was set at a p-value of 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. The data generated in this study have been deposited in the Figshare database under accession code https://doi.org/10.6084/m9.figshare.30521282. The scRNA-seq data generated in this study have been deposited into the NGDC genome sequence archive (GSA) with the accession number CRA035950. The mouse reference genome (mm10-2020-A) required for CellRanger can be downloaded from 10× Genomics (https://www.10xgenomics.com/support/software/cell-ranger/latest/release-notes/cr-reference-release-notes). The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Stupp, R. et al. Effect of tumor-treating fields plus maintenance temozolomide vs maintenance temozolomide alone on survival in patients with glioblastoma: a randomized clinical trial. JAMA 318, 2306–2316 (2017).
Reardon, D. A. et al. Effect of nivolumab vs bevacizumab in patients with recurrent glioblastoma: the checkmate 143 phase 3 randomized clinical trial. JAMA Oncol. 6, 1003–1010 (2020).
Omuro, A. et al. Radiotherapy combined with nivolumab or temozolomide for newly diagnosed glioblastoma with unmethylated MGMT promoter: An international randomized phase III trial. Neuro Oncol. 25, 123–134 (2023).
Klemm, F. et al. Interrogation of the microenvironmental landscape in brain tumors reveals disease-specific alterations of immune cells. Cell 181, 1643–1660.e1617 (2020).
Chongsathidkiet, P. et al. Sequestration of T cells in bone marrow in the setting of glioblastoma and other intracranial tumors. Nat. Med. 24, 1459–1468 (2018).
Mendez, J. S. et al. Association between treatment-related lymphopenia and overall survival in elderly patients with newly diagnosed glioblastoma. J. Neurooncol 127, 329–335 (2016).
Balmanoukian, A., Ye, X., Herman, J., Laheru, D. & Grossman, S. A. The association between treatment-related lymphopenia and survival in newly diagnosed patients with resected adenocarcinoma of the pancreas. Cancer Investig. 30, 571–576 (2012).
Campian, J. L., Ye, X., Brock, M. & Grossman, S. A. Treatment-related lymphopenia in patients with stage III non-small-cell lung cancer. Cancer Investig. 31, 183–188 (2013).
Grossman, S. A. et al. Immunosuppression in patients with high-grade gliomas treated with radiation and temozolomide. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 17, 5473–5480 (2011).
Ellsworth, S. et al. Sustained CD4+ T cell-driven lymphopenia without a compensatory IL-7/IL-15 response among high-grade glioma patients treated with radiation and temozolomide. OncoImmunology 3, e27357 (2014).
Singh, K. et al. IL-7-mediated expansion of autologous lymphocytes increases CD8+ VLA-4 expression and accumulation in glioblastoma models. J. Clin. Invest. 135, https://doi.org/10.1172/jci181471 (2025).
Campian, J. L. et al. Long-Acting Recombinant Human Interleukin-7, NT-I7, Increases Cytotoxic CD8 T Cells and Enhances Survival in Mouse Glioma Models. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 28, 1229–1239 (2022).
Zhu, Z. et al. Zika virus has oncolytic activity against glioblastoma stem cells. J. Exp. Med. 214, 2843–2857 (2017).
Nair, S. et al. Zika virus oncolytic activity requires CD8+ T cells and is boosted by immune checkpoint blockade. JCI insight 6, https://doi.org/10.1172/jci.insight.144619 (2021).
Nassiri, F. et al. Oncolytic DNX-2401 virotherapy plus pembrolizumab in recurrent glioblastoma: a phase 1/2 trial. Nat. Med. 29, 1370–1378 (2023).
Todo, T. et al. Intratumoral oncolytic herpes virus G47Δ for residual or recurrent glioblastoma: a phase 2 trial. Nat. Med. 28, 1630–1639 (2022).
Galanis, E. et al. Carcinoembryonic antigen-expressing oncolytic measles virus derivative in recurrent glioblastoma: a phase 1 trial. Nat. Commun. 15, 493 (2024).
Desjardins, A. et al. Recurrent glioblastoma treated with recombinant poliovirus. N. Engl. J. Med. 379, 150–161 (2018).
Chen, L. et al. Oncolytic Zika virus promotes intratumoral T cell infiltration and improves immunotherapy efficacy in glioblastoma. Mol. Ther. oncolytics 24, 522–534 (2022).
Bao, S. et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 444, 756–760 (2006).
Lathia, J. D., Mack, S. C., Mulkearns-Hubert, E. E., Valentim, C. L. & Rich, J. N. Cancer stem cells in glioblastoma. Genes Dev. 29, 1203–1217 (2015).
Eramo, A. et al. Chemotherapy resistance of glioblastoma stem cells. Cell death Differ. 13, 1238–1241 (2006).
Shan, C. et al. A live-attenuated Zika virus vaccine candidate induces sterilizing immunity in mouse models. Nat. Med. 23, 763–767 (2017).
Hirsch, A. J. et al. Intracranial injection of genetically modified, mosquito non-transmissible Zika virus: Safety in primates and ramifications for brain tumor therapy. Cell Rep. Med. 6, 102509 (2025).
Khan, S. M. et al. Impact of CD4 T cells on intratumoral CD8 T-cell exhaustion and responsiveness to PD-1 blockade therapy in mouse brain tumors. J. Immunother. Cancer 10, https://doi.org/10.1136/jitc-2022-005293 (2022).
Kosaka, A., Ohkuri, T. & Okada, H. Combination of an agonistic anti-CD40 monoclonal antibody and the COX-2 inhibitor celecoxib induces anti-glioma effects by promotion of type-1 immunity in myeloid cells and T-cells. Cancer Immunol., immunotherapy: CII 63, 847–857 (2014).
Genoud, V. et al. Responsiveness to anti-PD-1 and anti-CTLA-4 immune checkpoint blockade in SB28 and GL261 mouse glioma models. Oncoimmunology 7, e1501137 (2018).
Ahmadzadeh, M. et al. Tumor antigen–specific CD8 T cells infiltrating the tumor express high levels of PD-1 and are functionally impaired. Blood 114, 1537–1544 (2009).
Fernandez-Poma, S. M. et al. Expansion of tumor-infiltrating cd8+ t cells expressing pd-1 improves the efficacy of adoptive t-cell therapy. Cancer Res. 77, 3672–3684 (2017).
Thiery, J. et al. Perforin pores in the endosomal membrane trigger the release of endocytosed granzyme B into the cytosol of target cells. Nat. Immunol. 12, 770–777 (2011).
Ravi, V. M. et al. T-cell dysfunction in the glioblastoma microenvironment is mediated by myeloid cells releasing interleukin-10. Nat. Commun. 13, 925 (2022).
Galea, I. et al. An antigen-specific pathway for CD8 T cells across the blood-brain barrier. J. Exp. Med. 204, 2023–2030 (2007).
Andreatta, M. et al. Interpretation of T cell states from single-cell transcriptomics data using reference atlases. Nat. Commun. 12, 2965 (2021).
Elpek, K. G., Lacelle, C., Singh, N. P., Yolcu, E. S. & Shirwan, H. CD4+CD25+ T regulatory cells dominate multiple immune evasion mechanisms in early but not late phases of tumor development in a B cell lymphoma model. J. Immunol. (Baltim., Md.: 1950) 178, 6840–6848 (2007).
Magnuson, A. M. et al. Identification and validation of a tumor-infiltrating Treg transcriptional signature conserved across species and tumor types. Proc. Natl. Acad. Sci. USA. 115, E10672–e10681 (2018).
Lang, F. F. et al. Phase I study of dnx-2401 (delta-24-rgd) oncolytic adenovirus: replication and immunotherapeutic effects in recurrent malignant glioma. J. Clin. Oncol. 36, 1419–1427 (2018).
Suzuki, K. et al. A conditionally replicative adenovirus with enhanced infectivity shows improved oncolytic potency. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 7, 120–126 (2001).
Fueyo, J. et al. Preclinical characterization of the antiglioma activity of a tropism-enhanced adenovirus targeted to the retinoblastoma pathway. J. Natl. Cancer Inst. 95, 652–660 (2003).
Martin, N. T. & Bell, J. C. Oncolytic virus combination therapy: killing one bird with two stones. Mol. Ther.: J. Am. Soc. Gene Ther. 26, 1414–1422 (2018).
Ilkow, C. S. et al. Reciprocal cellular cross-talk within the tumor microenvironment promotes oncolytic virus activity. Nat. Med. 21, 530–536 (2015).
Watowich, M. B., Gilbert, M. R. & Larion, M. T cell exhaustion in malignant gliomas. Trends cancer 9, 270–292 (2023).
Belcaid, Z. et al. Low-dose oncolytic adenovirus therapy overcomes tumor-induced immune suppression and sensitizes intracranial gliomas to anti-PD-1 therapy. Neuro-Oncol. Adv. 2, vdaa011 (2020).
Panagioti, E. et al. Immunostimulatory bacterial antigen–armed oncolytic measles virotherapy significantly increases the potency of anti-PD1 checkpoint therapy. The Journal of Clinical Investigation 131, https://doi.org/10.1172/JCI141614 (2021).
Eum, H. H. et al. Single-cell RNA sequencing reveals myeloid and T cell co-stimulation mediated by IL-7 anti-cancer immunotherapy. Br. J. Cancer 130, 1388–1401 (2024).
Phase I study of efineptakin alfa (NT-I7) for the treatment of Kaposi sarcoma. Ramya Ramaswami et al. PMID: 39915263 PMCID: PMC11804200 https://doi.org/10.1136/jitc-2024-010291
Ahn, S. et al. Compassionate use of recombinant human IL-7-hyFc as a salvage treatment for restoring lymphopenia in patients with recurrent glioblastoma. Cancer Med. 12, 6778–6787 (2023).
Butt, O. et al. 624 A phase I/II study evaluating the safety and efficacy of a novel long-acting interleukin-7, NT-I7, for patients with newly diagnosed high-grade gliomas after chemoradiotherapy. J. Immunother. Cancer 10, A656–A656 (2022).
Dutcher, J. P. et al. High dose interleukin-2 (Aldesleukin) - expert consensus on best management practices-2014. J. Immunother. Cancer 2, 26 (2014).
Atkins, M. B. et al. Phase I evaluation of intravenous recombinant human interleukin 12 in patients with advanced malignancies. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 3, 409–417 (1997).
Conlon, K. C. et al. IL15 by continuous intravenous infusion to adult patients with solid tumors in a phase i trial induced dramatic nk-cell subset expansion. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 25, 4945–4954 (2019).
Quijano-Rubio, A. et al. A split, conditionally active mimetic of IL-2 reduces the toxicity of systemic cytokine therapy. Nat. Biotechnol. 41, 532–540 (2023).
Romee, R. et al. First-in-human phase 1 clinical study of the IL-15 superagonist complex ALT-803 to treat relapse after transplantation. Blood 131, 2515–2527 (2018).
Wolfarth, A. A. et al. Advancements of common gamma-chain family cytokines in cancer immunotherapy. Immune Netw. 22, e5 (2022).
Gorman, M. J. et al. An Immunocompetent Mouse Model of Zika Virus Infection. Cell host microbe 23, 672–685.e676 (2018).
Lazear, H. M. et al. A mouse model of zika virus pathogenesis. Cell host microbe 19, 720–730 (2016).
Liu, C. J. et al. Treatment of an aggressive orthotopic murine glioblastoma model with combination checkpoint blockade and a multivalent neoantigen vaccine. Neuro Oncol. 22, 1276–1288 (2020).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. Doubletfinder: doublet detection in single-cell rna sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e324 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Borcherding, N., Bormann, N. & Kraus, G. scRepertoire: An R-based toolkit for single-cell immune receptor analysis [version 2; peer review: 2 approved]. F1000Res. 9, https://doi.org/10.12688/f1000research.22139.2 (2020).
Bunis, D. G., Andrews, J., Fragiadakis, G. K., Burt, T. D. & Sirota, M. dittoSeq: universal user-friendly single-cell and bulk RNA sequencing visualization toolkit. Bioinformatics 36, 5535–5536 (2020).
Acknowledgements
The authors thank the Flow Cytometry & Fluorescence Activated Cell Sorting Core and the McDonnell Genome Institute at Washington University for providing technical assistance and equipment, and the Division of Comparative Medicine at Washington University for their excellent animal care. Y.D.L. and D.A.G. were supported by the NCI R38 Stimulating Access to Research in Residency (StARR) Program at Washington University. Research reported in this publication was supported by the Alvin J. Siteman Cancer Center through philanthropic funds, Dr. Ralph and Marian Falk Medical Research Trust, Robert J. Kleberg, Jr. and Helen C. Kleberg Foundation and NINDS/NIH under award numbers R01NS107833 and R01NS117149 (M.G.C.) and Cancer Biology Pathway/Molecular Oncology T32 Program (A.A.C.). M.G.C. was supported by the Washington University Institute of Clinical and Translational Sciences, which is partly funded by the NIH/National Center for Advancing Translational Sciences (NCATS) CTSA grant (UL1TR002345), the Hope Center for Neurological Disorders, and the Alvin J. Siteman Cancer Center at Barnes-Jewish Hospital. The Siteman Cancer Center is also supported in part by an NCI Cancer Center Support Grant (P30CA091842). We are grateful to Mrs. Mary Pickle for her support in honor of Brett Pickle, and to J. Michael, Heather Locke, and family for their generous support. We also thank the Alvin J. Siteman Cancer Center at Washington University School of Medicine and Barnes-Jewish Hospital in St. Louis, MO., for the use of the Biostatistics and Qualitative Research Shared Resource (BQSR). The Siteman Cancer Center is supported in part by an NCI Cancer Center Support Grant P30 CA091842.
Author information
Authors and Affiliations
Contributions
Y.D.L. and M.G.C. conceived and designed the study. Y.D.L., D.A.G., Y.H., A.K., T.H., L.P., and A.A.C. performed the experiments and collected the data. Y.D.L., D.A.G., A.W., and M.G.C. developed the methodology and provided technical support. Y.D.L., D.A.G., Y.H., A.K., L.P., and A.A.C. analyzed and interpreted the data. J.L., D.C., M.T.D, A.H.K., J.F.D., D.T.C., M.S.D., and M.G.C. contributed materials and analysis tools. Y.D.L. and M.G.C. drafted the manuscript. All authors contributed to revising the manuscript critically for important intellectual content and approved the final version.
Corresponding author
Ethics declarations
Competing interests
A.W. and D.C. are currently employed by NeoImmuneTech, Inc. J.F.D. is a consultant or on a Scientific Advisory Board for Rivervest, Bioline, Bluebird, Vertex, Incyte, NeoImmuneTech, and Macrogenics. J.F.D. has ownership investment in Magenta and WUGEN. M.S.D. is a consultant or on a Scientific Advisory Board for Inbios, IntegerBio, Akagera Medcines, GlaxoSmithKline, Merck, and Moderna. The Diamond laboratory has received unrelated funding support in sponsored research agreements from Vir Biotechnology, Emergent BioSolutions, Bavarian Nordic, and Moderna. M.G.C. receives royalties from UpToDate, and has received research funding from Merck, Orbus Therapeutics, Incyte, and NeoimmuneTech. M.G.C. and M.S.D hold a patents on the use of oncolytic Zika virus to treat brain tumors (US12274725B2, EP3638304, EP3638304A4, JP7525853B2, AU2018283957A1). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks E.Antonio Chiocca and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y.D., Giles, D.A., Huang, Y. et al. Long-acting interleukin-7 improves the efficacy of oncolytic viral therapy in glioblastoma. Nat Commun 17, 2893 (2026). https://doi.org/10.1038/s41467-026-69606-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69606-7