Abstract
A common host response to pathogen infection involves the production of robust interferons or proinflammatory cytokines to activate the JAK-STAT pathway, thereby limiting pathogen replication. The bacterial pathogen Legionella pneumophila creates an intracellular niche and evades host immunity utilizing a cohort of effectors by diverse biochemical activities, thereby permissive for its intracellular replication. However, roles of the JAK-STAT pathway during bacterial infection remain elusive. Here, we identify for the first time that L. pneumophila acetyltransferase effector Lem17 acts as a negative regulator of the JAK-STAT signaling. Lem17 directly interacts with JAK1 through a JAK1-binding Box1-like motif, preventing its recruitment by cytokine receptors. As a YopJ-family acetyltransferase, Lem17 catalyzes Nε-lysine acetylation of JAK1 and impairs its kinase activity, thereby disrupting JAK1-mediated signaling transduction. Our findings provide insights into the mechanism by which L. pneumophila subverts host immunity through acetylation and underscore the role of the JAK-STAT pathway against bacterial infection.
Similar content being viewed by others
Introduction
Protein acetylation, one of the most vital post-translational modifications (PTMs), is catalyzed by acetyltransferases that divert acetyl groups from acetyl co-enzyme A (acetyl-CoA) molecules to specific residues of substrates. Protein acetylation occurs on histone and non-histone proteins and plays a critical regulatory role in various cellular processes, e.g., signal transduction and immune response1. An accumulating body of evidence demonstrates that acetylation is one of the potent strategies for various intracellular pathogens to resist host immunity and promote their replication in host cells. One such example is found in bacterial virulence factors such as the Yersinia outer protein J (YopJ) family, which functions as Ser/Thr-acetyltransferases2. The primary function of these virulence factors is believed to play a pivotal role in evading the host immune response and establishing a persistent infection in host cells after bacterial infection3.
The innate immune system is the first line of host defense against pathogen infection. In response to pathogen infection, host cells produce robust cytokines such as interferons (IFNs) and interleukins4. The Janus kinase (JAK) /signal transducer and activator of transcription (STAT) pathway is important in orchestrating immune responses, which mediates the functions of IFNs and other cytokines5. IFNs bind to the cognate IFN receptor (IFNAR) and induce IFNAR dimerization to recruit JAKs, thus activating the JAK/STAT signaling. The activated JAKs phosphorylate downstream STATs. The phosphorylated STATs then form dimers and subsequently translocate into the nucleus, where they bind to DNA elements and trigger the transcription of IFN-stimulated genes6. These genes play an important role in establishing the immune response against pathogens7,8. Although the JAK/STAT pathway is well documented to be mainly activated through interferons for host defenses, many pathogens have evolved mechanisms to combat host defense and establish an efficient infection. Zika viruses (ZIKV) and Influenza A viruses (IAV) target JAK1 for proteasomal degradation, blocking JAK/STAT signaling and impairing IFN-mediated antiviral response9,10. Bacterial pathogen Shigella binds calmodulin (CaM) to inhibit the CaM/CaMKII pathway and block STAT1 phosphorylation, thereby dampening host IFN response11.
Legionella pneumophila, an intracellular bacterial pathogen, causes Legionnaires’ disease. This bacterium creates an intracellular niche and evades host immunity utilizing a large cohort of effectors by diverse biochemical activities, thereby permissive for its replication12,13. One of these effectors functions by mediating post-translational modifications that perturb host signaling pathways. Effector LnaB, as a phosphoryl-AMPylase, AMPylates the Src family of kinases, thereby inhibiting host immunoreceptor signaling14,15. Effector LphD, a eukaryotic-like histone deacetylase, targets H3K14ac and works in synergy with another effector, RomA, to promote H3K14 methylation, thereby counteracting host immune responses16,17. Notably, L. pneumophila encodes several effectors homologous to enzymes in either the eukaryotic Gcn5-related N-acetyltransferase family or prokaryotic Serine/Threonine acetyltransferase family. This suggests that effector-mediated acetylation may play a vital role in L. pneumophila infection. However, only L. pneumophila effector VipF has been identified as an acetyltransferase containing dual GNAT domains and targeted the eukaryotic eIF3 complex to modulate protein translation18,19. Nevertheless, the roles of these effectors in L. pneumophila infection have remained limited.
Here, we identified L. pneumophila effector Lem17 (Lpg1949) as a negative regulator of the JAK1-STAT pathway to attenuate the host immune response. Lem17 acts as an acetyltransferase that shuts down JAK1-STAT signaling through direct interactions and acetylation of JAK1, thereby facilitating the creation of a permissive intracellular environment for L. pneumophila replication in host cells.
Results
Lem17 is a bona fide acetyltransferase containing a conserved C-H-D motif
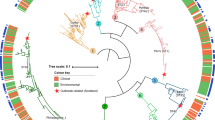
Bioinformatics analysis reveals that the YopJ family proteins are encoded by diverse bacterial pathogens of a wide range of hosts, particularly several proteins, including Lem17 (Lpg1949), Lpg2538, and Lpg2907, found in L. pneumophila species (Supplementary Fig. 1a). Notably, L. pneumophila deficient in the lpg1949 gene exhibited impaired replication within host cells, demonstrating that effector Lem17 contributes to the intracellular growth of L. pneumophila (Fig. 1a, b). Sequence analysis further revealed that Lem17 harbors a conserved C-H-D (C174-H105-D124) motif associated with the YopJ family members (Fig. 1c). To uncover whether Lem17 possesses an acetyltransferase activity, we ectopically expressed the wild-type Lem17 or its catalytic mutants (C174S, H104A and D124A) in HEK293T cells. Interestingly, we found that the global acetylation signaling increased in cells expressing Lem17 using an anti-pan acetyl-lysine antibody, instead, the indicated mutants are not (Fig. 1d). These results demonstrate that Lem17 is an acetyltransferase facilitating the intracellular growth of L. pneumophila.
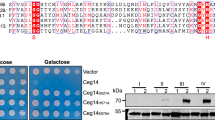
a Construction of a lem17-deficient L. pneumophila strain. The expression of effector Lem17 was probed by immunoblotting using Lem17-specific antibody, and isocitrate dehydrogenase (ICDH) was probed as a loading control. WT: wild type L. pneumophila. b Lem17 benefits to the intracellular growth of L. pneumophila. Bacteria of the indicated strains were used to infect mouse macrophages Raw264.7 at an MOI of 0.05, and the growth of the bacteria was determined at the indicated time points. WT: wild type L. pneumophila. hpi: hours post-infection. CFUs: Colony-forming units. The data represent three independent experiments; the data are mean ± s.d. Source data are provided as the Source Data file. c Multiple sequence alignment of the C-H-D/E motifs from YopJ family members and Lem17. Listed from left to right are the protein name and the amino acids. Identities are highlighted in red, and residues of the C-H-D/E motif are marked by a red star. Lp: Legionella pneumophila; Yp: Yersinia pseudotuberculosis; Rs: Ralstonia solanacearum; Ps: Pseudomonas syringae pv. Syringae; Se: Salmonella enterica subsp. enterica serovar Typhimurium. d HEK293T cells transfected to express Flag-Lem17 showed an increased global acetylation profile. HEK293T cells were transfected to express 3 × Flag-Lem17 or its indicated mutant for 24 h, and the samples were probed with the appropriate antibodies. GAPDH as loading control. Residues C174, H104 and D124 of the C-H-D motif are highlighted in red. WT: wild type Lem17. The results shown were from 3 independent experiments with similar results.
Lem17 requires host inositol hexaphosphate for activation
Given the Ser/Thr acetyltransferase family members generally require an essential cofactor inositol hexaphosphate (IP6) as an activator20, we next examined whether Lem17 binds IP6. We found that IP6 shows high affinity for Lem17, with a binding constant (Kd) of approximately 2 μM (Fig. 2a). Specifically, Lem17 only binds acetyl-CoA in the presence of IP6 (Kd ≈ 20 μM), whereas no binding was detected in its absence (Fig. 2a). These results suggest that Lem17 requires IP6 for activation.
a ITC experiments verifying that IP6 is required for Lem17 binding the ligand CoA. Raw ITC curves (top panel) and binding isotherms with fitting curves (bottom panel) of Lem17 titration by IP6. Lem17 + IP6: Lem17 is saturated with IP6 at a 1: 5 molar ratio for 60 min before titration. IP6: inositol hexaphosphate; CoA: Co-enzyme A. The data represent three independent experiments; the data are mean ± s.d. b Schematic diagram of the Lem17 region for crystallization. The catalytic core domain is shown in cyan, and the Lid domain is colored in green. c Two views of the overall structure of Lem17 in complex with IP6. The IP6 molecule is shown in gray stick. Lem17 is represented by a cylindrical cartoon, colored as panel (b). d The conserved residues C174, H105 and D124 in the structure of Lem17. Residues are represented as sticks with the annealing omit map (colored in gray) contoured at the 2.0σ level, and proteins were depicted in cyan cartoon. e The IP6-binding pocket. Residues important for IP6 binding were shown as sticks. Hydrogen bonds were marked by blue dashed lines. The refined 2mFo-Fc map (colored in pink) of IP6 is contoured at the 2.0σ level. The six phosphate groups of IP6 are designated the labels P1–P6. f The affinity of IP6 with Lem17 and its indicated mutants was determined using isothermal titration calorimetry (ITC). WT: wild type Lem17; Kd: the dissociation constant. The data represent three independent experiments; the data are mean ± s.d. g Lem17 mutants were evaluated for the ability to increase gthe lobal acetylation profile. HEK293T cells were transfected to express 3 × Flag-Lem17 or its indicated mutant for 24 h, and the samples were stained with the appropriate antibodies. GAPDH was probed as a loading control. Residue C174 of the C-H-D motif is highlighted in red. WT: wild type Lem17; Results shown were from 3 independent experiments with similar results.
To elucidate the IP6-mediated activation mechanism of Lem17, we have devoted significant efforts to determining the crystal structures of distinct forms: apo-Lem17, IP6-bound Lem17, and IP6-bound Lem17 in complex with the ligand CoA. However, our attempts have so far only succeeded in obtaining the IP6-bound Lem17 structure (Supplementary Fig. 1b). As the structural observation, Lem17 is divided into two subdomains: catalytic core domain (CD) and Lid domain (Lid). IP6 binds to a positive charge cleft in the YopJ-like domain, where is covered by the Lid domain, thus locking IP6 in place (Fig. 2b, c). The catalytically critical residues C174, H105 and D124 form a continuous platform (Fig. 2d), which may be the site for interactions between Lem17 and its host targets. Structural alignment reveals that Lem17 shares high structural similarity with the catalytic domain of the YopJ family members except for the additional Lid domain, for example, HopZ121 (PDB ID: 5KLP) with RMSD of 3.4 Å and PopP222 (PDB ID: 5W3T) with RMSD of 3.6 Å, (Supplementary Fig. 1a, b). Specifically, residues within the conserved C-H-D motif exhibit a nearly identical spatial pattern (Supplementary Fig.2c). The presence of this Lid domain may confer Lem17 with greater specificity or affinity for its host targets.
Moreover, structural alignment between the Lem17-IP6 complex and the AlphaFold3-predicted Lem17 model suggested that IP6 binding might cause a conformational change in its Lid domain (Supplementary Fig. 2d–f), potentially enabling the binding of ligand acetyl-CoA or substrates. In addition, structural data revealed that IP6 contacts Lem17 extensively via polar interactions (Fig. 2e and Supplementary Fig. 2g). Residues K248 and R222 coordinate with P1 in IP6 by hydrogen bonds. Residues K278 and R281 hydrogen bonds with P2 in IP6. Residues R45 and K46 coordinate with P3 and P4 in IP6 by hydrogen bonds. Residues R50, R53 and K187 coordinate with P5 in IP6 by hydrogen bonds. Residues K187 and K247 coordinate with P6 in IP6 by hydrogen bonds. As structural observations, alanine substitutions at the indicated residues severely impaired IP6 binding to Lem17 (Fig. 2f and Supplementary Fig. 3a–c). Consistent with these findings, these mutants deficient in IP6-binding exhibited a severe reduction of the global acetylation signaling in cells (Fig. 2g). Taken together, these data suggest that Lem17 is a bona fide acetyltransferase toward lysine residues. Of course, serine- or threonine-acetylation cannot be ruled out.
Lem17 mediates Nε-acetylation of host JAK1 that impairs its kinase activity
To elucidate the specific physiological function of Lem17 in the host, we next sought to identify host proteins that are targeted by Lem17. We ectopically expressed Flag-Lem17 in HEK293T cells and performed an immunoprecipitation (IP) after formaldehyde using an anti-Flag antibody, followed by mass spectrometric analysis, and another L. pneumophila effector Lpg2538 as a control (Fig. 3a and Supplementary Data 1). Since YopJ family members target host kinases, we focused on candidate proteins harboring kinase activity. The top 10 selected kinases are presented in Fig. 3b. We first examined their acetylated modification in cells using an anti-pan acetyl-lysine antibody. A distinct acetylation signal was observed at the position corresponding to Janus kinase 1 (JAK1) in the presence of Lem17, whereas no such signal was readily detectable for the other selected proteins examined (Supplementary Fig. 4a). Notably, PTK7, TAOK3, PACSIN3 and GRK6 from the top 10 list failed to be expressed under our experimental conditions. Consistently, we purified HA-tagged JAK1 by HA-beads after co-expression with Flag-Lem17 or its indicated mutant in HEK293T cells. Acetylated-JAK1 indeed was detectable in the IPed profile of cells expressing Flag-Lem17, while it was not in cells harboring the empty vector or Lem17 C174S mutant (Fig. 3c). In addition, the Flag-Lem17 or its C174S mutant were substantially enriched with JAK1 in the IPed fraction. Collectively, these data indicated that JAK1 is one host target of Lem17. In addition, we also examined the top 20 proteins from the proteomics data and found that Lem17 modified host protein EPB41L2 (Supplementary Fig. 4b, c). EPB41L2, which also contains a FERM domain, is required for dynein-dynactin complex and NUMA1 recruitment at the mitotic cell cortex during anaphase23. However, whether EPB41L2 engages in the JAK1-STAT pathway warrants further investigation.
a Identification of host candidates for Lem17. Flag-Lem17 expressed in HEK293T cells was isolated by immunoprecipitation, and the bound proteins were identified by mass spectrometry. Similarly, obtained samples with Flag-Lpg2538 were used as a control. HCHO: Formaldehyde. The results shown were from 3 independent experiments with similar results. b Putative host kinases of Lem17. A full list of proteins is provided in Supplementary Data 1 and the top 10 kinases are shown here. c Acetylation of JAK1 by Lem17 requires the C-H-D motif. HEK293T cells were co-transfected to express HA-JAK1 and 3 × Flag-Lem17 or its mutant C174S for 24 h, and then were IPed with the HA-specific antibody. Acetylation of JAK1 was detected using a pan-acetyl-lysine antibody. Residue C174 of the C-H-D motif is highlighted in red. WT: wild type Lem17. The results shown were from 3 independent experiments with similar results. d Schematic of Lem17-dependent JAK1 acetylation sites identified by mass spectrometry. Acetylated residues are indicated by a red line and labeled; their identifies are provided in Supplementary Data 1. FERM: four-point-one, ezrin, radixin, moesin domain; SH2: Src Homology 2 domain; PK: Pseudokinase; TK: Tyrosine kinase domain. e Validation of JAK1 acetylation sites by immunoblotting. HEK293T cells were co-transfected to express HA-Lem17 and 3 × Flag-JAK1 or its indicated mutants for 24 h, and then were IPed with the Flag-specific antibody. Acetylation of JAK1 was detected using a pan-acetyl-lysine antibody. The results shown were from 3 independent experiments with similar results. f Lem17-mediated acetylation impairs the kinase activity of JAK1. The kinase activity of JAK1 is determined by the remaining amount of ATP in the reaction using an ATP assay kit. The IRS-1 peptide (KKHTDDGYMPMSPGVA) is used as a substrate. Results are indicated as luminescence. WT: wild type Lem17; C174S: Lem17 C174S mutant. The data represent three independent experiments; the data are mean ± s.d. Two-sided p-values were generated using a Student’s t test. Source data are provided as the Source Data file. g Electrostatic surface potential surrounding the conserved C-H-D triad in Lem17 and the YopJ family. Residues are shown in sticks and proteins are depicted in surface, colored according to the electrostatic surface potential [contoured from − 5 kBT (red) to + 5 kBT (blue)].
JAK1, which belongs to the JAK family of non-receptor tyrosine protein kinases, has four domains. Starting from the carboxyl terminus, these include: a tyrosine kinase domain (TK) that phosphorylates substrates, a pseudokinase domain (PK) that regulates the activity of the TK domain, a Src-homology 2 (SH2) domain and a four-point-one, ezrin, radixin, moesin domain (FERM) domain5. We next analyzed Lem17-targeted acetylation sites on JAK1 using mass spectrometry. This analysis revealed multiple lysine residues were modified by Nε-acetylation, including K205, K213, K263, and K493 in the FERM-SH2 domain, and K911, K941 and K953 in the TK domain (Fig. 3d, Supplementary Figs. 5, 6 and Supplementary Data 2). Consistent with these findings, strong acetylation signal was detectable in JAK1 and its FERM-SH2 and TK domains in the IPed profiles (Supplementary Fig. 4d). Alanine substitutions in the FERM domain (individual: K205; triple: K205A/K213A/K267A) or the TK domain (individual: K911; triple: K911A/K941A/K953A) caused detectable decreases in JAK1 acetylation. Notably, the quadruple substitution (K205A/K267A/K911A/K941A) exhibited a substantial decrease in acetylation signal (Fig. 3e). In addition, due to the non-expression of the K213A and K267A mutations in cells, we further constructed these substitutions in the truncated FERM-SH2 and TK domains of JAK1, respectively. Reduced acetylation signals were detected for the K213A and K267A substitutions within the FERM-SH2 domain (Supplementary Fig. 4e). These results suggest that Lem17 could acetylate multiple residues on JAK1.
Notably, these acetylated residues cluster in two functional domains: the FERM-SH2 domain, which mediates cytokine-associated receptor binding, and the kinase domain (TK), which is essential for substrate phosphorylation. Specifically, residues K267 and K493, positioned adjacent to the JAK1-IFNλR1 interaction interface, potentially disrupt cytokine receptor binding, whereas K911, K941 and K953, localized within the N-lobe of the kinase domain, are likely to impair kinase activity (Supplementary Fig. 4f). Consistently, Lem17-mediated acetylation of JAK1 led to a significantly reduced kinase activity (Fig. 3f). Although Lem17 belongs to the Ser/Thr acetyltransferase family characterized by acetylating substrates’ Ser/Thr residues, MS analysis failed to detect any Ser/Thr-acetylated residues in JAK1. This discrepancy may arise from differences in the electrostatic potentials surrounding their C-H-D catalytic triads: Lem17 exhibits a negatively charged microenvironment, whereas YopJ family proteins such as PopP222 and HopZ1a21 display a distinct bias toward positively charged regions (Fig. 3g).
Lem17 blocks JAK1 recruitment to cytokine receptors
To elucidate the mechanism by which Lem17 recognizes and acetylates JAK1, we have made extensive efforts to resolve the structure of the Lem17-JAK1 complex; however, these attempts have thus far been unsuccessful. Therefore, we predicted the structural model of the Lem17-JAK1 complex (Supplementary Fig. 7a, b), and showed that the Lid domain of Lem17 binds to the FERM domain of JAK1 through polar and hydrophobic interactions (Fig. 4a and Supplementary Fig. 7c, d). To validate this model, we performed alanine substitutions at residues involved in the hydrophobic interaction between JAK1 and Lem17 and conducted the co-immunoprecipitation assay. As the predicted structural observation, alanine mutations in JAK1 significantly reduced its affinity for Lem17, particularly in the LVL (L190A/V194A/L195A) and VFF (V243A/F244A/F247A) mutants (Fig. 4b). Similarly, alanine mutation F368A/I371A/F375A (termed FIF) in Lem17 also markedly impaired its interaction with JAK1 (Fig. 4c). It notes that the predicted JAK1-Lem17complex structure is limited by a moderate ipTM score, despite our point-variant data supports this prediction. In addition, Lem17 was observed in the IPed profile of cells harboring the full-length and FERM-SH2 of JAK1 (Supplementary Fig. 7e). These results indicate that the FERM domain of JAK1 plays a dominant role in binding to Lem17. Further sequence alignment revealed that three hydrophobic residues in Lem17 (F368, I371, F375) are conserved and correspond to characteristic residues of the PxxLxF motif within the Box1 of JAK1-associated cytokine receptor cytoplasmic domains (Fig. 4d). This conserved PxxLxF motif, corresponding to each cytokine receptor, is documented to be functionally essential for JAK1 binding24.
a Close-up view of the hydrophobic interaction between Lem17 and JAK1. JAK1 (green) and Lem17 (Cyan) are shown in cartoon representation. Residues are depicted in sticks. b Validation of residues involved in hydrophobic interaction between JAK1 and Lem17 by co-immunoprecipitation. WT: wild type JAK1; FF: F247A/F251A mutant; LVL: L190A/V194A/L195A mutant; VFF: V243A/F244A/F247A mutant. The data represent three independent experiments with similar results. c Validation of residues involving interaction between Lem17 and JAK1 by co-immunoprecipitation. WT: wild type JAK1; FIF: F368A/I371A/F375A mutant. The data represent three independent experiments with similar results. d Multiple sequence alignment of Lem17 with the JAK1-binding motif Box1 from JAK1-associated upstream receptors by PROMALS3D. Listed from left to right are the protein name and the amino acids. Identities of residues important for JAK1 binding are highlighted in red, and a line marks the Box1 motif. IL10RA: Interleukin 10 receptor subunit α; IFNAR2: Interferon α/β receptor subunit 2; IFNLR1: Interferon λ receptor 1; IL22RA1: Interleukin 22 receptor subunit α1; IFNGR1: Interferon γ receptor 1; IL20RA: Interleukin 20 receptor subunit α. e Structural alignment of the predicted JAK1-Lem17 complex with the JAK1-IFNλR1 complex revealed steric hindrance between Lem17 and IFNλR1. Proteins were depicted in a cylindrical cartoon (Lem17 in cyan, JAK1 in green and IFNλR1 in orange). IFNλR1: Interferon λ receptor 1. f Close-up view of the steric hindrance between Lem17 and IFNλR1. Residues that cause steric hindrance between Lem17 and IFNλR1 were shown in sticks. Lem17 (cyan) and IFNλR1 (orange) were depicted in a cartoon, JAK1 was depicted in surface, colored according to the electrostatic surface potential [contoured from −5 kBT (red) to + 5 kBT (blue)]. g Effect of the important residues of Lem17 in JAK1 interaction on bacterial replication. Bacteria of the indicated strains were used to infect macrophage-like U937 cells at an MOI of 0.05, and the growth of the bacteria was determined at the indicated time points. WT: wild type L. pneumophila; ∆Lem17: lem17-deficient L. pneumophila. FIF: F368A/I371A/F375A mutant. The data represent three independent experiments; the data are mean ± s.d. Source data are provided as the Source Data file. h Ectopic expression of Lem17 impaired JAK1 recruitment by upstream cytokine-associated receptors. HeLa cells expressing 3 × Flag-Lem17 for 24 h were stimulated with IFN-β for another 6 h and immunoprecipitated using a JAK1-specific antibody. The IPed profiles were detected using an approximate antibody. The results shown were from 3 independent experiments with similar results.
Subsequently, structural alignment of the Lem17-JAK1 complex with the JAK1-IFNλR1 complex revealed that the binding interface between Lem17 and JAK1 sterically clashes with the Box1 motif of IFNR1, which resides within the FERM domain of JAK1 (Fig. 4e and Supplementary Fig. 7f). This clash primarily involves three hydrophobic residues in Lem17 Lid domain (F368, I371, and F375) that occupy equivalent spatial positions to three residues in IFNλR1 (P264, L267, and F269) (Fig. 4f). Consistently, the mutant FIF in Lem17 impaired the intracellular replication of L. pneumophila (Fig. 4g). These results suggested that Lem17 might sterically hinder JAK1 from recognizing its cognate cytokine receptors by preemptively occupying the cytokine receptor’s binding pocket before JAK1 engagement. To clarify this hypothesis, we immunoprecipitated endo-JAK1 in IFN-λ-primed HEK293T cells with ectopically expressing Flag-lem17. Immunoprecipitation profiles demonstrated reduced enrichment of the JAK1-upstream IFNAR2 and JAK1-downstream STAT1, while these proteins were significantly enriched in Lem17-deficient cells (Fig. 4h). Collectively, these results indicated that Lem17 impairs the ability of JAK1 to recognize its associated cytokine receptors through a dual mechanism involving direct interaction and acetylation.
Lem17 dampens the JAK1 signaling that facilitates bacterial intracellular replication
Given that JAKs function as the central signaling hub in the JAK-STAT pathway, transducing the cytokine-mediated extracellular stimuli into intracellular STAT activation response5,25, we subsequently detected the role of Lem17 in the JAK1-mediated signaling transduction. Phosphorylation levels of STAT1 and STAT2 were strongly elevated in the cells primed with IFN-β. However, this increase was remarkably suppressed by the ectopic expression of Lem17. In contrast, the C174S mutant reversed the suppression caused by Lem17 (Fig. 5a). Consistently, we performed RNA-sequencing (RNA-seq) analysis on total RNA isolated from IFN-primed cells ectopically expressing Lem17 or an empty vector. The distinct gene expression profiles (comprising approximately 500 differentially expressed genes) were observed in Lem17-expressing cells compared to those transfected with empty vector (Fig. 5b). Gene Ontology (GO) analysis revealed that these genes are enriched in multiple signaling pathways, including the interferon-mediated antiviral pathway (Fig. 5c). Genes associated with interferon-mediated antiviral pathways – including those involved in the ISG15 antiviral mechanism, antiviral mechanism by IFN-stimulated genes, and interferon signaling, demonstrate significant enrichment in negatively correlated phenotypes, as evidenced by negative normalized enrichment scores (Fig. 5d, e and Supplementary Fig. 8a). Collectively, these results demonstrate that Lem17 dampens the JAK signaling.
a Ectopic expression of Lem17 suppressed IFN-β-primed JAK1 signaling. HeLa cells were treated with IFN-β for 6 h following transfection to express 3 × Flag-Lem17 or its indicated mutant for 24 h. The samples were probed with the appropriate antibodies. GAPDH was probed as a loading control. Residue C174 of the C-H-D motif is highlighted in red. WT: wild type Lem17; IFN-β: Interferon β. The results shown were from 3 independent experiments with similar results. b A volcano plot of differentially expressed genes following ectopic Lem17 expression in IFN-β-primed cells. HeLa cells expressing Lem17 for 24 h were treated with 1μg/mL IFN-β for 6 h, followed by harvesting for RNA-seq analysis, an empty vector was used as a control. The statistical test used was two-sided. c GO enrichment analysis of differentially expressed genes. GO: Gene Ontology. Red star indicates the pathways associated with IFN-β signaling. d–e, GSEA profiling of Genes pertinent to antiviral mechanisms by IFN-stimulated genes (d) and Interferon signaling (e). GSEA: Gene Set Enrichment Analysis. NES: Normalized Enrichment Score. f JAK1 knockout enhanced the intracellular growth of L. pneumophila. Bacteria of the indicated strains were used to infect human HeLa expressing FcγΙΙ at an MOI of 0.05, and the growth of the bacteria was determined at the indicated time points. HeLaWT: wild type HeLa cells; HeLa∆jak1: jak1-deficient HeLa cell; LpWT: wild type L. pneumophila; Lp∆dotA: dotA-deficient L. pneumophila. Data represent mean values ± s.d. n = 3 biological replicates, Two-sided p-values were generated using a Student’s t test. Source data are provided as the Source Data file.
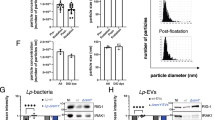
A common host response to pathogen infection is the production of IFNs or proinflammatory cytokines to limit pathogen replication. Similarly, host cells produce robust proinflammatory cytokines like interferon and interleukins, particularly IFN-β and IL-6, in response to L. pneumophila infection (Supplementary Fig. 8b–e). These cytokines exert antibacterial effects through their downstream JAK-STAT pathway26. As expected, JAK1 knockdown promoted the intracellular proliferation of L. pneumophila in both HeLa and U937 cells (Fig. 5f and Supplementary Fig. 8f–h). Therefore, these results suggest that Lem17 disrupts JAK1-mediated signal transduction, which contributes to the intracellular replication of L. pneumophila in host cells.
Discussion
In this study, we describe Legionella effector Lem17 as an IP6-dependent acetyltransferase that targets host JAK1-mediated signaling, thereby suppressing host immunity. Homologues of Lem17 have been described in other enteric and plant pathogens, including YopJ, VopA and AopP, harboring acetyltransferase activities that require host IP6 as an activator2. Similarly, the kinase activity of Legionella effector Lpg2603 requires IP6 for activation27.
The requirement of a host cell-specific co-factor allows pathogens to restrict the activity of virulence factors within target cells. OspC family members of Shigella28 and CyaA of Bordetella pertussis29 employ CaM as the co-factor to ensure that their functions exactly execute only in host cells. The requirement of CaM by L. pneumophila effector SidJ is to prohibit untimely inactivation of SidEs in the bacterial-infected cells30. The requirement of actin by L. pneumophila effector LnaB is to initiate its ATPase activity, thereby converting PR-Ub to ADPR-Ub14,15. Moreover, Ub is essential for the activation of the Legionella phospholipase effector VpdC to promote the expansion of its phagosome31 and for PtpA and PtpB of Mycobacterium tuberculosis to suppress host innate immunity32,33. Given that ligand CoA did not bind to Lem17 in the absence of IP6 (Fig. 2a), we therefore propose that IP6 binding induces conformational changes in Lem17 that enable ligand/substrate binding. This is consistent with the IP6-induced conformational changes occurring globally among YopJ family proteins21,22,34.
The hijacking of host immunity-associated kinases to subvert host immunity has been exemplified by the functional characterization of YopJ family members2. Similarly, the Lem17 catalytic function suppressed host kinase JAK1 function through acetylation, thereby subverting host immunity. Unlike most proteins in the YopJ family that use O-acetylation, Lem17 mediates N-acetylation of substrate JAK1. This difference may arise from variations in the electrostatic surface potential on the surface around the conserved catalytic triad in Lem17 and other YopJ family effectors (Fig. 3g). This analysis revealed that HopZ1a, a YopJ effector produced by the phytopathogen Pseudomonas syringae, features a distinct positively charged catalytic center conducive to catalyzing O-acetylation. In contrast, Lem17 harbors a negatively charged catalytic center that preferentially binds positively charged lysine. This characteristic aligns with PopP2, another YopJ effector from Ralstonia solanacearum, which has been documented to catalyze N-acetylation35, also possesses a negatively charged catalytic center.
Innate immunity plays a key role in host defense against pathogen infection. As is often the case with pathogen infections, the IFNs or proinflammatory factors are secreted and bind to the cognate receptor to activate the JAK/STAT signal pathway, thereby inducing the expression of ISGs that inhibit pathogen replication26. In order to survive successfully in host cells, intracellular pathogens have been in an evolutionary arms race with their host cell to block signaling of the JAK-STAT pathway, thus hindering ISGs production. The bacterial pathogens are also capable of the same functions to limit ISGs production by diverse mechanisms. Shigella employs the OspC family of type-III-secreted effectors OspC1 and OspC3 to block calcium signaling, thus inhibiting CAMKII and its downstream STAT1 phosphorylation11. Helicobacter pylori regulates the JAK-STAT signal transduction by the phosphorylation status of CagA, acting through the gp130 receptor36. In response to L. pneumophila infection, host cells produced proinflammatory factors like IFN- and IL-6 (Supplementary Fig. 8), thereby activating JAK-STAT signaling to limit bacterial replication. However, effector Lem17 suppressed JAK1-mediated signaling by targeting JAK1 (Figs. 3 and 5), and JAK1 deficiency facilitated the intracellular growth of L. pneumophila (Fig. 5f and Supplementary Fig. 8h). Therefore, one consequence of Lem17-mediated blockage of JAK1 signaling is to inhibit host innate immunity, thereby facilitating L. pneumophila intracellular replication.
In summary, we report a working model of the regulatory role of the L. pneumophila effector Lem17 in the JAK1-mediated signaling pathway (Fig. 6). Lem17 functions as an acetyltransferase and suppresses JAK1-STAT signaling by perturbing JAK1 function through a dual mechanism involving both lysine acetylation and Lem17 binding, thereby impairing host IFN response. These findings enhance understanding of the pathogen immune evasion mechanism, and highlight the role of effector-mediated acetylation in L. pneumophila pathogenesis and JAK-STAT signaling in host antibacterial immunity.
ACK: Acetyl-CoA; Ac: Acetylation; P: phosphorylation; LCV: Legionella-containing vacuole; T4SS: Type IV Secretion System of Legionella. TK: Kinase domain; PK: Pseudokinase domain. ISGs: Interferon-stimulated genes. ISRE: Interferon-Stimulated Response Element. This Figure was created in BioRender. Yang, Y. (2026) https://BioRender.com/o2d84se.
Methods
Cell lines
HEK293T, HeLa, U937 and mouse RAW264.7 cells purchased from the ATCC were cultured in Dulbecco’s modified minimal Eagle’s medium (DMEM) supplemented with 10% FBS at 37 °C. All mammalian cell lines were regularly checked for potential mycoplasma contamination by the universal mycoplasma detection kit from ATCC (Cat# 30-1012 K).
Bacterial strains
All L. pneumophila strains were derived from the Philadelphia-1 strain Lp0237. The bacteria were grown on a charcoal-yeast extract (CYE) plate or in N-(2-acetamido) 2-aminoethanesulfonic acid (ACES)–buffered yeast extract (AYE) broth with additional thymidine (200 μg/mL) for the thymidine auxotrophic mutant and its derivatives. The Lp02∆lem17 mutant was described earlier38.
Escherichia coli (E.coli) strains DH5α and BL21 Rosetta DE3 were grown in lysogeny broth (LB) shaking at 37 °C. Antibiotics were added to culture media at the following concentrations: ampicillin (100 μg/mL), kanamycin (30 μg/mL), and streptomycin (50 μg/mL), unless specified. All strains were stored at − 80 °C in LB supplemented with 10% (v/v) glycerol.
Plasmid constructs
To express Lem17 in mammalian cells, the ORF of lem17 was amplified from L. pneumophila genome and inserted into vectors pCMV-3 × Flag or pCMV-HA at EcoRI/NotI sites. To express JAK1 in mammalian cells, the coding regions of JAK1 or its fragments were amplified from a human cell complementary DNA library, digested with XhoI II/NotI, and inserted into vector pcDNA3.1 with an HA-tag or pCMV-3 × Flag.
To produce Lem17 protein in E.coli, the lem17 gene and its fragments were amplified from L. pneumophila genome, digested with BamHI/NheI, and inserted into vector pET-his harboring a 6 × His-tag.
For jak1-knockout HeLa cells, the sense and antisense chains of human jak1 gRNA were annealed, and then inserted into vector pLentiCRISPR-v2 that was pre-digested with BsmBI, to generate pLentiCRISPR-Δjak1.
All expression constructs for Lem17 or JAK1 point mutations were generated by the modified site-directed mutagenesis protocol with corresponding primers. All primers used in this study are synthesized by a local company (Beijing Tsingke Biotech Co., Ltd, Chain) and listed in Supplementary Table 1.
L. pneumophila preparation and infection
L. pneumophila used for infection were grown in AYE broth to their post-exponential phase, as judged by both optical density [optical density at 600 nm (OD600), 3.2 to 4.0] and bacterial motility.
For intracellular growth assays, cells were seeded into 12- or 24- well plates or 24 at 5 × 105 per well respectively. Cells infected with L. pneumophila at a multiplicity of infection (MOI) of 0.05 were synchronized at 2 hours by washing three times with fresh medium to remove all extracellular bacteria. Infected macrophages were continued to culture at 37 °C in the presence of 5% CO2. To measure the bacterial growth, cells were lysed with 0.02% digitonin. Colony forming unit of infected bacteria was determined by plating a series of lysate dilutions on solid CYET medium. HeLa cells were transfected with the plasmid expressing the FcγΙΙ receptor for 20-24 h before infection. U937 cells were differentiated in fresh medium by phorbolmyristate acetate (1 μM) for 12 h.
For the transcription assay, mouse RAW264.7 cells were seeded into 6-well plates at 2 × 106 and 1 × 106 per well, respectively, for 1 day before infection. Cells infected with L. pneumophila at a multiplicity of infection (MOI) of 50 were synchronized at 2 h by washing three times with fresh medium to remove all extracellular bacteria. Infected macrophages were continued to culture at 37 °C in the presence of 5% CO2.
Formaldehyde-cross linking in vivo and mass spectrometry
HeLa cells were plated in six 10-cm dishes at a density of 2 × 106 cells per dish, three of which were transfected with plasmid pCMV-3 × Flag-Lem17, while the other three were transfected with empty vector pCMV-3 × Flag vector as a control. The cells after 18 h of transfection were washed twice by PBS, and cross-linked with 1% (v/v) formaldehyde for 10 min at 37 °C, and terminated by 1.25 mM glycine for 10 min at room temperature. The cells were then harvested by centrifugation at 1500 × g for 3 min, washed twice by PBS, and lysed by sonication for 1 min in a lysis buffer, containing 50 mM Tris (pH 7.5), 150 mM NaCl, 10% glycerol, 1% NP-40, 0.2 mM EDTA, and 1 mM PMSF. The cell lysate was rocked for another 4 h at 4 °C and centrifuged at 16,200 × g for 30 min. Supernatant of the lysate was collected into a new tube and incubated with 50 μL of the Flag affinity resins at 4 °C overnight. The affinity resin was then collected by centrifugation at 400 × g for 2 min, washed in lysis buffer three times, and boiled for 10 min with 1.25 × SDS loading buffer. All IPed proteins were separated on 12% SDS–polyacrylamide gel electrophoresis (SDS-PAGE) gel and further identified by mass spectrometric analysis.
Co-Immunoprecipitation
HEK293T or HeLa cells were plated at a density of 2 × 106 cells per 100 mm dish and transfected with plasmid pCMV-3 × Flag-Lem17 or pCMV-HA-JAKs using a liposomal transfection reagent (Yeasan). The cells were collected at 24 h post-transfection and were incubated for 30 min on ice with 1.0 mL of a lysis buffer after sonication. The cell lysis supernatant was collected by centrifugation at 16,200 × g for 10 min, and was transferred to a new 15 ml tube. The supernatant was incubated with 20 μL of the Flag affinity resins in IP buffer including 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 0.02% Triton X-100, 10% glycerol, 0.2 mM EDTA at 4 °C for 6 h, followed by three washes with the IP buffer. The captured proteins were eluted with 30 μL of Flag peptide (50 μg/mL) for 30 min on ice. The elution was boiled with 1.25 × SDS loading buffer and separated on 12% SDS-PAGE, followed by immunoblotting with appropriate antibodies.
Immunoprecipitation
HeLa cells were plated in ten 10-cm dishes at a density of 2 × 106 cells per dish, and five of which were transfected with plasmid pCMV-3 × Flag-Lem17 or its mutants, while the other five were transfected with empty vector pCMV-3 × Flag vector as a control. The cells were collected at 24 h post-transfection and were incubated for 30 min on ice with 1.0 ml of a lysis buffer after sonication. The cell lysis supernatant was collected by centrifugation at 16,200 × g for 10 min, and was transferred to a new 15 ml tube. The supernatant was incubated with 20 μL of the Flag affinity resins in IP buffer containing 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 0.02% Triton X-100, 10% glycerol, 0.2 mM EDTA at 4 °C for 6 h followed by three washes with the IP buffer. The captured proteins were eluted with 30 μL of Flag peptide (50 μg/mL) for 30 min on ice. The elution was boiled with 1.25 × SDS loading buffer and separated on 12% SDS-PAGE, followed by immunoblotting with appropriate antibodies.
Liquid chromatography-tandem mass spectrometry analysis
After separation by SDS–PAGE, gel slices containing the protein detected by CBB stain were digested using 20 ng/L trypsin in 25 mM NH4HCO3 overnight at 37 °C39. The digested peptides were analyzed on a C18 reversed-phase column connected to a UPLC (Acquity, Waters) coupled to an Orbitrap mass spectrometer (Q-Exactive Plus, Thermo Fisher Scientific). The mass spectrometer was operated in positive ion and standard data-dependent acquisition mode with the Advanced Peak Detection function activated for the top 10n. The resolution of the Orbitrap mass analyzer was set to 60,000 and 15,000 for MS1 and MS2, respectively. The full scan MS1 spectra were collected in the mass range of 350-1400 m/z, with an isolation window of 1.2 m/z and a fixed first mass of 100 m/z for MS2. The spray voltage was set at 2.2 kV and the Automatic Gain Control (AGC) target was 300% for MS1 and standard for MS2, respectively. Maximum ion injection time was set as 50 ms for MS1 and 110 ms for MS2, respectively. Dynamic exclusion was set with an exclusion duration of 6 s.
For protein identification, the raw data were processed with the software MaxQuant40 (version 1.6.3.3) against the Homo sapiens database (Uniprot, UP000005640) or L. pneumophila database (Uniprot, UP000000609). For acetylation identification, the raw data were converted to peak lists using DeconMSn41 and submitted for blind post-translational modification search using MODa42 against. Post-translational modification candidates were confirmed by manual inspection, looking for consistent mass shifts in b and y fragment series, and by reprocessing the data with MaxQuant, considering the specific modifications. MaxQuant was set to search with the following parameters: peptide tolerance at 10 ppm, MS/MS tolerance at 0.02 Da, carbamidomethyl (C) as a fixed modification, oxidation (M) as a variable modification, and a maximum of two missed cleavages. The false-discovery rates (FDR) were controlled at < 1%.
Quantitative real-time PCR analysis
Total RNA was isolated from RAW264.7 cells infected with L. pneumophila using RNAiso Plus (Takara, 9109) as per the manufacturer’s instructions. Then reverse transcribed into cDNA by PrimeScript™ RT reagent Kit with gDNA Eraser (Takara, RR047A). Real-time PCR was performed with TB Green® Premix Ex Taq™ (Tli RNaseH Plus) (Takara, RR420A) in QuantStudio 6 Flex Real-Time PCR System (ABI) according to the standard procedure. The fold change expression (-∆∆Ct) was calculated after normalization with GAPDH expression.
Generation of CRISPR-Cas9 genome-edited cell lines
The lentiviral particles were packaged by co-transfecting 1.5 μg of pLentiCRISPR-Δjak1 together with 1.5 μg of lentiviral packaging plasmids (pMDL:pVSV-G:pREV = 0.5:0.3:0.2) into HEK293T cells plated at 2 × 106 cells each well by liposome transfection reagent Turbofect (Thermo Fisher Scientific). The cells were changed into fresh medium after 8 to 12 h and continued to culture for another 24–48 h. The supernatant containing lentivirus particles was then centrifuged to remove cell debris at 1500 × g for 5 min at room temperature. A new batch of HeLa cells were infected for 48 h by the lentivirus in 60-mm dishes and continued to culture for several generations with puromycin (2 μg/mL) until the knockout cell line was obtained. The efficiency of the jak1 knockout was evaluated by Western blot using anti-JAK1 antibody.
JAK1 knockdown in U937 cells by shRNA
shRNAs were designed to target human JAK1 using a design tool provided by Sigma (https://www.sigmaaldrich.cn/). The corresponding sequences were cloned into the pLKO.1-puro vector, and the insert sequences were verified by Sanger sequencing. The knockdown procedure was performed following the protocol from our previous method38. The lentiviral particles were packaged by co-transfecting 1.5 μg of pLKO-JAK1 together with 1 μg of lentiviral packaging plasmids (psPAX2:pMD2G = 3:1) into HEK293T cells plated at 2 × 106 cells each well by liposome transfection reagent. The cells were changed into fresh medium after 8 to 12 h and continued to culture for another 24–48 h. The supernatant containing lentivirus particles was then centrifuged to remove cell debris at 1500 × g for 5 min at room temperature. U937 cells were infected by the lentivirus in the presence of 10 µg/mL polybrene. 48 h post-infection, the cells were selected with 2 μg/mL puromycin to generate stable pools. After the mock-infected control cells were completely eliminated, the knockdown efficiency of each shRNA construct was evaluated by Western Blot using anti-JAK1 antibody. The cell line exhibiting the most potent JAK1 knockdown was selected for subsequent experiments. Primers for cloning shRNA-JAK1 as follows:
F: 5′-CCGGGAGACTTCCATGTTACTGATTCTCGAGAATCAGTAACATGGAAGTCTCTTTTTG-3′
R: 5′-AATTCAAAAAGAGACTTCCATGTTACTGATTCTCGAGAATCAGTAACATGGAAGTCTC-3′
Kinase activity assay of JAK1
Flag-tagged JAK1 was purified from HEK293T cells co-expressing either Lem17 or its mutant C174S. The IRS-1 peptide (KKHTDDGYMPMSPGVA) synthesized by a local company (Sangon Biotech, China) was used as the substrate in the kinase activity assay. The kinase reaction was performed in a reaction buffer containing 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 10 mM MgCl₂. Briefly, 100 μL of reaction mixture containing 1 mM ATP, 1 mM IRS-1 peptide and 0.1 μM purified JAK1 was incubated at 37 °C for 1 h. Then, 100 μL of ATP detection reagent was added, followed by incubation at room temperature for another 30 min. The remaining amount of ATP in the reaction was monitored using an ATP Assay Kit (Beyotime, S0027) according to its manual, with a fluorescence microplate reader (Thermo Fisher Varioskan LUX, USA). Results are indicated as luminescence, where greater luminescence indicates lower kinase activity of JAK1.
Protein expression and purification
E. coli strain BL21 (DE3) transformed with an expression construct was cultivated in LB medium at 37 °C till its optical density OD600 of 0.8, and induced by 400 μM isopropyl β-D-1-thiogalactopyranoside (IPTG) at 18 °C overnight. The cell pellets were harvested by centrifugation at 3470 × g for 15 min and stored at − 20 °C.
His6-tagged proteins were purified using the following protocol. The cell pellets were suspended in a lysis buffer containing 50 mM Tris pH 8, 500 mM NaCl, 10% (v/v) glycerol, 5 mM β-mercaptoethanol, 20 mM imidazole and protease inhibitors, and lysed using sonication on ice. The cell lysate was centrifuged for 30 min at 34,572 × g and its supernatant was collected and incubated with nickel sepharose beads for 1 h at 4 °C. The beads were washed by the lysis buffer and eluted by the lysis buffer added with a gradient of imidazole: 50, 100, 200, and 500 mM. The best fractions that contained our protein of interest were pooled, concentrated, and run through a Superdex 200 size-exclusion column (GE Healthcare) in buffer containing 20 mM Tris-HCl pH 8.0, 150 mM NaCl and 2 mM DTT. The best fractions of protein peak were pooled, concentrated to 10 mg/mL with an Amicon Centrifugal filter (Millipore), flash-frozen in liquid nitrogen and stored at − 80 °C for crystallization and activity assay. Protein concentration was measured at A280 and calculated using their theoretical extinction coefficients.
Protein crystallization and structural determination
The truncated Lem1715-446 was pre-incubated with 1 mM IP6 for 30 min on ice and the crystallization was performed at 16 °C using a sitting drop diffusion method by mixing 0.4 μL of protein (10 mg/mL) with an equal volume of reservoir solution. The crystals initially grew in 0.1 M MES pH 6.0, 5% (w/v) polyethylene glycol PEG 3000, and 30% (v/v) PEG 200 at 16 °C after 3 days. After optimization, the best crystallization buffer was 0.1 M MES pH 6.2, 25% PEG200 and 5% PEG 3,000. The Lem17-IP6 binary complex crystals were cryoprotected by brief soaking in the crystallization buffer with an additional 1 mM IP6 and flash-frozen in liquid nitrogen. Diffractions of crystals were collected at beamline BL02U1 of Shanghai Synchrotron Radiation Facility (SSRF) (Shanghai, China). Data sets were indexed, integrated, and scaled with xia2 software43. Lem17-IP6 binary structure was determined by the molecular replacement method with program Phaser in PHENIX44, and the AlphaFold3 model of Lem17 15-320 was employed as a template. The subsequent model was manually built using COOT45 and refined with the program Refine in PHENIX44. The data collection and structure refinement statistics are summarized in Supplementary Table 2. Structural figures were generated using PyMol (http://pymol.org/2/).
Isothermal titration calorimetry (ITC)
The ITC experiments were performed using a microcalorimeter Affinity ITC (Waters, USA) at 20 °C. The Lem17 protein or its mutants purified above were diluted into a binding buffer including 50 mM Tris-HCl (pH 8.0), 150 mM NaCl to a final concentration of 50 μM. IP6 or Acetyl-CoA stocks were also diluted in the same buffer at a final concentration of 500 μM, separately. Titrations are set for 20 injections of 2 μL each with 200-s intervals except from 0.2 μL of the first injection. Baseline subtraction and data analysis are performed using NanoAnayze. Heat spikes are integrated and fit with 1:1 binding model. The first injection of each experiment is excluded from the analysis.
To test the impact of IP6 on acetyl-CoA binding to Lem17, Lem17 was pre-incubated with IP6 at a molar ratio of 1: 5 for 60 min before the ITC experiments. The ITC experiment was performed as above.
RNA-seq analysis
Total RNA was isolated using R (Qiagen) following the manufacturer’s instructions. DNase I in column digestion was included to ensure RNA quality. RNA library preparation was performed by using NEBNext@ UItraTM Directional RNA Library Prep Kit for Illumina (E7420L), and paired-end sequencing was performed with Illumina HiSeq 4000 platform. Low-quality reads were cleaned through FastQC (version 0.11.9) and Fastp (version 0.22.0), two packages for sequence pre-disposal. Processed reads were aligned to the human reference genome hg38 using STAR (version 2.7.11b). The alignment results were processed using the featureCount (version 2.0.6) for gene quantification with the option -primary (count primary alignments only) -B (only count read pairs that have both ends aligned) -C (do not count read pairs that have their two ends mapping to different chromosomes or mapping to the same chromosomes but on different strands) -Q 20 (the minimum mapping quality score 20) -s 0 (unstranded). Differential gene analysis was performed using limma (version 3.54.1), using P-value < 0.05 and FC ≥ 1 as cutoff. Then, volcano plots were drawn using the ggplot2 (version 3.5.1) package of R software to depict these differentially expressed genes. Reactome enrichment analysis was performed using the R package clusterProfiler (version 4.6.2) and ReactomePA (version 1.42.0). GSEA analysis was performed using the R package GSVA (version 1.46.0).
Antibodies and immunoblotting
For immunoblotting, samples resolved by SDS-PAGE were transferred onto 0.2 μm Polyvinylidene Fluoride (PVDF) membranes (Millipore, 0000183267), which were blocked with 5% non-fat milk at room temperature for 1 h prior to being incubated with the appropriate primary antibodies: anti-DYKDDDDK tag (Proteintech, 66008-4-Ig), 1:5000; anti-HA (Proteintech, 51064-2-AP), 1:5000; anti-JAK1 (CST, 3344 T), 1:1000; anti-STAT1 (Abcam, ab109320), 1:2000; anti-STAT1 (phospho Y701) (Abcam, ab109457), 1:1000; anti-STAT2 (Abcam, ab32367), 1:2000; anti-STAT2 (phospho Y690) (Abcam, ab191601), 1:1000; anti-Acetyllysine (PTM, PTM-105RM), 1: 2000; anti-ICDH (Sigma-Aldrich, ABS2090), 1:10000; anti-IFNAR2 (Proteintech, 10522-1-AP), 1: 1000; anti-GAPDH (Proteintech, 60004-1-Ig), 1:5000; anti-Lem17(prepared from Biodragon), 1:1000. Membranes were then incubated with appropriate secondary antibodies conjugated to HRP (Proteintech, SA00001-2 or SA00001-1, 1:5000), and scanned by an imaging system (ImageQuant800, Cytiva).
Bioinformatic identification of members of the YopJ family
We retrieved the YopJ protein from the Pfam database (ID: PF03421) and obtained all sequences under this record. Additionally, protein sequences from Legionella pneumophila were included. Sequences were deduplicated at 90% identity using CD-HIT, followed by manual removal of excessively long and short sequences, resulting in a final set of 240 sequences. Corresponding protein structures were acquired by querying the AlphaFold Database and performing predictions using AlphaFold3. Multiple structural alignments were conducted with US-align. The resulting structure-based sequence alignment served as a reference profile, which was then used to guide multiple sequence alignment with Clustal Omega. Phylogenetic analysis was performed using FastTree 2.1.11 with 1000 bootstrap replicates. Results were visualized in iTOL v6, with branch coloring applied according to bacterial family classifications.
Data analysis
The significant difference analysis of one-way analysis of variance (ANOVA) and Student’s t test were calculated all in GraphPad Prism 8 software (San Diego, USA). P < 0.05 was considered a significant difference.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Coordinate and structure factor for the Lem17-IP6 complex included in this study have been deposited in the Protein Data Bank (PDB) under accession code 9VA7. The mass spectrometry proteomics data identifying host targets of Lem17 and Lem17-mediated acetylation sites on JAK1 have been deposited to the ProteomeXchange Consortium via the iProX partner repository (https://proteomecentral.proteomexchange.org) via the iProX partner repository)46,47 with the dataset identifiers: PXD070394 and PXD073153, respectively. RNA_seq data have been deposited in the Gene Expression Omnibus (GEO) database under accession code: GSE316818. All data and the related. Source data are provided in this paper.
References
Xue, M. et al. Protein acetylation going viral: implications in antiviral immunity and viral infection. Int. J. Mol. Sci. 23, https://doi.org/10.3390/ijms231911308 (2022).
Ma, K. W. & Ma, W. YopJ Family Effectors Promote Bacterial Infection through a Unique Acetyltransferase Activity. Microbiol. Mol. Biol. Rev. 80, 1011–1027 (2016).
Grabowski, B., Schmidt, M. A. & Ruter, C. Immunomodulatory Yersinia outer proteins (Yops)-useful tools for bacteria and humans alike. Virulence 8, 1124–1147 (2017).
Kaempfer, R. Subduing the Inflammatory Cytokine Storm. Int. J. Mol. Sci. 25, https://doi.org/10.3390/ijms252011194 (2024).
Hu, X., Li, J., Fu, M., Zhao, X. & Wang, W. The JAK/STAT signaling pathway: from bench to clinic. Signal Transduct. Target Ther. 6, 402 (2021).
Lv, Y. et al. The JAK-STAT pathway: from structural biology to cytokine engineering. Signal Transduct. Target Ther. 9, 221 (2024).
Crosse, K. M., Monson, E. A., Beard, M. R. & Helbig, K. J. Interferon-stimulated genes as enhancers of antiviral innate immune signaling. J. Innate Immun. 10, 85–93 (2018).
Schoggins, J. W. Interferon-stimulated genes: what do they all do? Annu. Rev. Virol. 6, 567–584 (2019).
Wu, Y. et al. Zika virus evades interferon-mediated antiviral response through the co-operation of multiple nonstructural proteins in vitro. Cell Discov. 3, 17006 (2017).
Yang, H. et al. The influenza virus PB2 protein evades antiviral innate immunity by inhibiting JAK1/STAT signalling. Nat. Commun. 13, 6288 (2022).
Alphonse, N. et al. A family of conserved bacterial virulence factors dampens interferon responses by blocking calcium signaling. Cell 185, 2354–2369 (2022).
Iyer, S. & Das, C. The unity of opposites: Strategic interplay between bacterial effectors to regulate cellular homeostasis. J. Biol. Chem. 297, 101340 (2021).
Qiu, J. & Luo, Z. Q. Legionella and Coxiella effectors: strength in diversity and activity. Nat. Rev. Microbiol. 15, 591–605 (2017).
Wang, T. et al. Legionella effector LnaB is a phosphoryl-AMPylase that impairs phosphosignalling. Nature 631, 393–401 (2024).
Fu, J. et al. Legionella maintains host cell ubiquitin homeostasis by effectors with unique catalytic mechanisms. Nat. Commun. 15, 5953 (2024).
Rolando, M. et al. Legionella pneumophila effector RomA uniquely modifies host chromatin to repress gene expression and promote intracellular bacterial replication. Cell Host Microbe 13, 395–405 (2013).
Schator, D. et al. Legionella para-effectors target chromatin and promote bacterial replication. Nat. Commun. 14, 2154 (2023).
Chen, T. T., Lin, Y., Zhang, S. & Han, A. Structural basis for the acetylation mechanism of the Legionella effector VipF. Acta Crystallogr. D Struct. Biol. 78, 1110–1119 (2022).
Syriste, L. et al. An acetyltransferase effector conserved across Legionella species targets the eukaryotic eIF3 complex to modulate protein translation. mBio 15, e0322123 (2024).
Pruitt, R. N. et al. Structure-function analysis of inositol hexakisphosphate-induced autoprocessing in Clostridium difficile toxin A. J. Biol. Chem. 284, 21934–21940 (2009).
Zhang, Z. M. et al. Structure of a pathogen effector reveals the enzymatic mechanism of a novel acetyltransferase family. Nat. Struct. Mol. Biol. 23, 847–852 (2016).
Zhang, Z. M. et al. Mechanism of host substrate acetylation by a YopJ family effector. Nat. Plants 3, 17115 (2017).
Kiyomitsu, T. & Cheeseman, I. M. Cortical dynein and asymmetric membrane elongation coordinately position the spindle in anaphase. Cell 154, 391–402 (2013).
Ferrao, R. et al. The structural basis for class II cytokine receptor recognition by JAK1. Structure 24, 897–905 (2016).
Villarino, A. V., Kanno, Y. & O’Shea, J. J. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat. Immunol. 18, 374–384 (2017).
Morris, R., Kershaw, N. J. & Babon, J. J. The molecular details of cytokine signaling via the JAK/STAT pathway. Protein Sci. 27, 1984–2009 (2018).
Sreelatha, A. et al. A Legionella effector kinase is activated by host inositol hexakisphosphate. J. Biol. Chem. 295, 6214–6224 (2020).
Li, Z. et al. Shigella evades pyroptosis by arginine ADP-riboxanation of caspase-11. Nature 599, 290–295 (2021).
Shen, Y. et al. Physiological calcium concentrations regulate calmodulin binding and catalysis of adenylyl cyclase exotoxins. EMBO J. 21, 6721–6732 (2002).
Gan, N. et al. Regulation of phosphoribosyl ubiquitination by a calmodulin-dependent glutamylase. Nature 572, 387–391 (2019).
Li, X., Anderson, D. E., Chang, Y. Y., Jarnik, M. & Machner, M. P. VpdC is a ubiquitin-activated phospholipase effector that regulates Legionella vacuole expansion during infection. Proc. Natl. Acad. Sci. USA 119, e2209149119 (2022).
Chai, Q. et al. A bacterial phospholipid phosphatase inhibits host pyroptosis by hijacking ubiquitin. Science 378, eabq0132 (2022).
Wang, J. et al. Mycobacterium tuberculosis suppresses innate immunity by coopting the host ubiquitin system. Nat. Immunol. 16, 237–245 (2015).
Xia, Y. et al. Secondary-structure switch regulates the substrate binding of a YopJ family acetyltransferase. Nat. Commun. 12, 5969 (2021).
Le Roux, C. et al. A receptor pair with an integrated decoy converts pathogen disabling of transcription factors to immunity. Cell 161, 1074–1088 (2015).
Lee, I. O. et al. Helicobacter pylori CagA phosphorylation status determines the gp130-activated SHP2/ERK and JAK/STAT signal transduction pathways in gastric epithelial cells. J. Biol. Chem. 285, 16042–16050 (2010).
Berger, K. H. & Isberg, R. R. Two distinct defects in intracellular growth complemented by a single genetic locus in Legionella pneumophila. Mol. Microbiol. 7, 7–19 (1993).
Chen, T. T. et al. Atypical Legionella GTPase effector hijacks host vesicular transport factor p115 to regulate host lipid droplet. Sci. Adv. 8, eadd7945 (2022).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Mayampurath, A. M. et al. DeconMSn: a software tool for accurate parent ion monoisotopic mass determination for tandem mass spectra. Bioinformatics 24, 1021–1023 (2008).
Na, S., Bandeira, N. & Paek, E. Fast multi-blind modification search through tandem mass spectrometry. Mol. Cell. Proteom. 11, M111 010199 (2012).
Winter, G., Lobley, C. M. & Prince, S. M. Decision making in xia2. Acta Crystallogr. Section D Biol. Crystallogr. 69, 1260–1273 (2013).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. Sect D Biol. Crystallogr. 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect. D Biol. Crystallogr. 60, 2126–2132 (2004).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China grants 82172287 (S.O.), U25A20131 (S.O.), 82225028 (S.O.) and 82572582 (T.-T.C.), Prevention and Control of Emerging and Major Infectious Diseases-National Science and Technology Major Project 2025ZD01903600 (S.O.), and High-level personnel introduction grant of Fujian Normal University Z0210509 (S.O.), and Fujian Province Natural Science Foundation 2025J01639 (T.-T.C.), and Fujian provincial guiding funds of central government for supporting the development of the local science and technology 2024L3008 (S.O.), and Fujian provincial funds for promoting high-quality development of marine and fishery sectors FJHYF-L-2025-24 (S.O.). The diffraction data were collected at beamlines BL-02U1 and BL-10U2 of Shanghai Synchrotron Radiation Facility (SSRF). Mass spectrometric analysis was performed in the core facility of Xiamen University.
Author information
Authors and Affiliations
Contributions
T.-T.C. performed crystal collection and structural determination, analyzed the data and wrote the manuscript; S.-R.Z. and B.Y. performed the cellular experiments; W.Z., Q.L., and S.-R.Z. performed the bacterial infection experiments; W.Z., X.L., and Z.J. performed protein purification, crystallization, and mutagenesis; F.L. performed structural refinement and bioinformatic analysis; J.S. and G.H. performed RNA-seq analysis. J.S., Y.F., and C.H. revised the manuscript; S.O. conceived the ideas for this work and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, TT., Zheng, SR., Yang, B. et al. A bacterial YopJ-family acetyltransferase suppresses host immune response by Nε-acetylation of JAK1. Nat Commun 17, 2910 (2026). https://doi.org/10.1038/s41467-026-69623-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69623-6