Abstract
Herpesviruses encapsulate their double-stranded DNA (dsDNA) genomes within an icosahedral nucleocapsid formed in the infected cell nucleus. Four biochemically purified nucleocapsids have been characterized, but their roles in herpesvirus replication remain controversial. The status of the capsid vertex-specific component (CVSC), essential for capsid stability and dsDNA packaging and retention, is also unclear. By integrating cryogenic focused ion beam milling with electron tomography and subtomogram averaging, we derive atomic models for all protein components, including the CVSC, across different herpesvirus capsid types within infected cell nuclei. Focused classification of pentonal vertex densities reveal differences in CVSC occupancy between genome-filled capsids and capsids lacking dsDNA, highlight structural heterogeneity and providing insight into distinct capsid assembly stages in situ. These intra-nuclear findings redefine the maturation model of herpesvirus capsid assembly, advancing the understanding of herpesvirus replication, and demonstrate the effectiveness of in situ electron imaging by studying virus assembly within host cells.
Similar content being viewed by others
Introduction
The Herpesviridae encompass a large family of viruses comprised of alpha-, beta- and gamma subgenera that infect humans and most animal species with wide-ranging health and economic consequences1,2,3,4,5,6,7,8. Varicella-zoster virus (VZV), a human alphaherpesvirus, causes varicella (chicken pox) as the primary infection, and zoster (shingles) after reactivation from latency in sensory ganglia1, and can be life-threatening in immunocompromised patients9,10,11,12,13,14,15. VZV vaccines have reduced morbidity substantially16,17,18 and antiviral drugs are effective but neither prevent VZV latency in sensory neurons19,20 or post-herpetic neuralgia after zoster21,22, and antiviral resistance can emerge23,24. The other two human alphaherpesviruses, herpes simplex virus (HSV) 1 and 2 (HSV-2), also create a significant disease burden25,26. HSV vaccines are not available, and HSV persists in neurons despite antiviral treatment8. Vulnerable populations need interventions for these viruses and against the betaherpesvirus, human cytomegalovirus4, and the Epstein-Barr and Kaposi’s sarcoma gammaherpesviruses, which cause a range of cancers2,3.
Herpesviruses are classified by their distinct 4-layer morphology composed of a double-stranded DNA (dsDNA) core, capsid, tegument, and envelope27. These complex entities form through a convoluted morphogenesis process starting inside the infected cell nucleus. The dsDNA genome is packaged into newly synthesized capsids that exit the nucleus across nuclear membranes into the cytoplasm, where they traffic to sites of secondary envelopment, completing acquisition of tegument proteins and the lipid envelope containing virus glycoproteins. The nascent enveloped particles are transported in vesicular structures to the plasma membrane and released into the extracellular milieu. The glycoproteins protruding from the lipid membrane then mediate virus entry into neighboring cells, and tegument proteins support the intracellular transport of capsids to nuclear pores, where the dsDNA genome is injected; additional tegument proteins undergo nuclear translocation to initiate transcription factor functions. Since the herpesvirus capsid structure is essential for virus genome preservation during infection, high-resolution information about the progression of capsid assembly holds great potential to identify targets for next-generation pharmaceutical interventions and vaccines.
Herpesvirus capsids are ~125 nm in diameter and share a common icosahedral symmetry with a T = 16 geometry made up of 60 asymmetric units. The capsid assembles in the nucleus from 161 capsomeres, composed of 150 hexons and 11 pentons that form an icosahedron with 12 vertices, including a portal vertex for the packaging of the dsDNA genome and its release through the host cell nuclear pore in newly infected cells. Capsids undergo several transformative stages starting with immature pro-capsids, which are thought to form as capsomeres coalesce with the scaffold protein to produce segments and subsequently spherical capsids28,29. Maturation of pro-capsids occurs as the scaffold protein is proteolytically digested by its companion protease, triggering a conformational change from a more spherical shape to their final icosahedral morphology, with 20 flat triangular faces and 30 edges. Mature capsids have been conventionally classified as A-, B-, and C-capsids, as they differ in their buoyant densities and morphologies by thin-section electron microscopy (EM)30. The cavities of C-capsids contain packaged dsDNA genomes, whereas B-capsids have a persistent scaffold protein and its associated protease, and A-capsids are described as empty. While A- and B-capsids have been presumed to be dead-end assembly products29, an alternative hypothesis is that one or both might represent assembly intermediates with the potential to encapsidate dsDNA genomes31. Capsid purification can induce artefacts, such as loss of genomic dsDNA by disruption of the portal vertex cap, creating empty A-capsids from C-capsids32. B-capsids accumulated when unit-length genome formation was blocked by drug inhibition of the genome-transferase complex31, suggesting that scaffold protein removal is linked to dsDNA insertion, and capsids with scaffold protein in the cavity might be capsid intermediates rather than representing a failure of genome encapsidation.
The proteins that assemble to create a mature herpesvirus capsid have orthologues in all Herpesviridae. These include the major capsid protein (MCP), the small capsomere interacting protein (SCP), and the capsid triplex proteins 1 (TRX1) and 2 (TRX2). The multimeric subunits formed by these proteins to produce fully mature capsids have been described at near atomic resolution for all three subfamilies of herpesviruses33,34,35,36,37,38,39,40,41,42,43. Briefly, the MCP forms the capsomeres of hexons and pentons of the icosahedral capsid. Six copies of the SCP decorate the tops of the hexons. Heterotrimeric triplexes (TRX1 monomer and TRX2 dimer) occupy the canyons between the capsomere towers and cover up the holes in the MCP floor to stabilize the capsid for dsDNA genome containment33,44. At each of the 12 capsid vertices, five copies of the capsid vertex-specific component (CVSC) bind to triplexes adjacent to the pentons and the portal vertex33. In HSV, the CVSC is a heteropentamer composed of orthologues shared with VZV; the capsid vertex-specific component 1 (CVC1) monomer, two copies of CVC2, and two copies of the protein designated VP1/2. CVSC binding has been associated with reinforcing capsid stability to withstand the very high packaging pressure of the dsDNA genome45,46,47. At the portal vertex, a dodecamer (VZV ORF54; HSV-1 pUL6), replaces a penton with a portal and tentacle-like helices that ring the DNA translocation channel, and an ATP-driven terminase is recruited to cleave the herpesvirus dsDNA genome for packaging29,34,35,36,38,45,48,49,50. The CVSC is expected to form part of the portal cap that retains the dsDNA genome in the capsid29,34,35,36,38,45,48,49,50. The CVSC heteropentamer has also been called the capsid-associated tegument complex because its subunits include the large tegument protein (VP1/2), VZV ORF22 and HSV pUL3651; in HSV and PRV, these orthologues have been identified as inner tegument proteins that function in microtubule-mediated capsid transport to cell nuclei52,53. The CVSC of CMV, a betaherpesvirus, has a non-conserved tegument subunit, pp15042,54, while gammaherpesvirus CVSCs are thought to resemble those of the alphaherpesviruses51.
Critical information about the structural composition of herpesvirus capsids was gained using cryogenic-electron tomography (cryo-ET) of purified particles, but resolution was limited55,56,57,58,59,60,61,62,63. The adaptation of single particle cryo-EM enabled capsid structures of most human herpesviruses to be determined at better than 4 Å resolution33,35,36,37,38,39,40,41,47. However, knowledge about capsid assembly as it occurs in situ within the infected cell nucleus remains limited. The CVSC has been proposed to associate with the capsid inside the infected cell nucleus, but the structural organizations of their individual components have not been visualized directly in the nucleus of herpesvirus-infected cells64,65. Whether the assembly of the VP1/2 subunit of the CVSC takes place in the nucleus or cytoplasm is unclear64,66,67. Furthermore, despite previous high-resolution structures for VZV capsids, the CVSC and the portal vertex were not resolved in these studies39,41. Recent work with purified capsids partially resolved the CVSC and the portal vertex68. To better understand capsid assembly and the relationship between capsids conventionally classified as A- and B-capsid types compared to the dsDNA-containing C-capsids, we sought subnanometer resolution imaging of capsids within VZV-infected cells. Imaging inside mammalian cells is challenging because cell thickness limits the penetration of the electron beam and reduces the image contrast due to inelastic scattering and multiple electron scattering69. Even at high voltages (1MV), the electron beam can only penetrate 1 µm; the nuclei of typical mammalian cells range in size from 5 to 10 µm. To overcome this limitation, recent advances in cryogenic focused ion beam and scanning electron microscopy (cryo-FIB/SEM) have been introduced to mill away most of the thick and vitrified cell, leaving a thin lamella of 100 to 400 nm in thickness. When integrated with cryo-ET using a 300 keV electron microscope, 3D volumes can be produced at near-atomic resolution by imaging at different angles, cryo-ET reconstruction, and subtomogram averaging (STA)70. We applied this approach to visualize VZV particles within infected cells in a frozen, hydrated state, without the use of contrast agents, and determined the structures of their component proteins.
Our findings reveal that the CVSC heteropentamer assembles on capsids in the nucleus of infected cells and that the definition of the asymmetric unit of the icosahedral VZV capsid should be expanded to incorporate the CVSC interactions with neighboring MCPs and triplexes. Focused classification, including of the portal vertex, indicates that genome-filled C-capsids have full CVSC occupancy, defined as the maximum of five heteropentamers. Among capsids lacking dsDNA, empty A-capsids are not detected within the infected cell nucleus, whereas capsid assembly intermediates (CAIs) could be distinguished by differences in the number of CVSCs found at the vertices of the icosahedron, which correlate with levels of scaffold protein in the capsid cavity, suggesting a continuum along the assembly pathway. Notably, CAIs would be grouped together as B-capsids based on scaffold protein alone. These observations provide evidence that CVSC acquisition can serve as a temporal marker for the progression of intra-nuclear herpesvirus capsid assembly in situ, consistent with its biological importance for ensuring the stability of genome-containing capsids during nuclear egress and formation of complete virus particles in the cytoplasm.
Results
VZV morphogenesis visualized within infected cells in situ by cryo-FIB-SEM and cryo-ET
To visualize herpesvirus morphogenesis in 3D space, a workflow was developed to distribute VZV-infected cells evenly on EM grids suitable for cryo-FIB milling. First, because VZV does not yield high-titer cell-free virus particles1, it was critical to optimize the passage of infected cells onto uninfected cells to initiate infection prior to vitrification. Infection was monitored by the fluorescence RFP signal produced by MeWo cells infected with a recombinant VZV (pOka-TK-RFP71) expressing a thymidine kinase (TK) tagged with a monomeric red fluorescence protein (RFP; Fig. 1a [created in BioRender https://BioRender.com/c2ose1j] and Fig. 1b); the pOka-TK-RFP maintains the characteristics of the bacterial artificial chromosome-derived WT pOka71,72,73,74. Second, to overcome the propensity of VZV infected cells to adhere to the grid bars when seeded on SiO2 EM grids, islands of extracellular matrix were micropatterned within the grid squares (Fig. 1b). With this technique, infected cells were positioned centrally within the grid squares after vitrification and ideal for cryo-FIB milling to produce lamella suitable for cryo-ET (Fig. 1c, d). Tomograms of cryo-FIB milled cells revealed the complex cellular environment in VZV-infected cells (Fig. 1e; Suppl. Figs. 1 to 7; Suppl. Tables 1 and 2). Cellular features were readily visible in medium magnification (16.9 to 39.1 Å/pixel) montage maps used to facilitate cryo-ET tilt series data collection from cells thinned to a target thickness of ~175 nm; the nuclear envelope with visible nuclear pores demarcated the intranuclear and cytoplasmic compartments (Fig. 1d; Suppl. Figs. 1a, 4a to 7a). A plethora of vesicular structures, mitochondria, and, within infected cells, VZV capsids and complete virus particles, was visible, facilitating the localization of regions of interest in a tilt series collection. The higher-resolution tomographic reconstructions yielded substantial contrast enabling machine-learning-based segmentation to identify ribosomes, microtubules, microfilaments, complex vesicular structures, and assembled capsids in the nucleus, and sites of virus particle morphogenesis in the cytoplasm (Suppl. Fig. 7c).
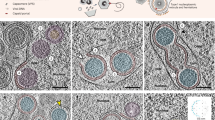
a Overview of the VZV replication cycle (created with BioRender.com). Key steps numbered 1 to 7. (1) VZV envelope glycoproteins bind cell-surface receptors. (2) Subsequent gB/gH-gL-mediated membrane fusion at the plasma membrane or within endocytic vesicles. (3) Capsid release into the cytoplasm, trafficking to the nuclear pore, ejection of dsDNA genome into the nucleus, initiating replication and capsid assembly. (4) Viral glycoproteins synthesized in the endoplasmic reticulum (ER) traffic through the Golgi and delivered to the trans-Golgi network (TGN) or nuclear envelope, including the inner (INM) and outer (ONM) nuclear membranes. (5) Envelopment and de-envelopment of newly assembled capsids by the nuclear egress complex. (6) Glycoproteins traffic to sites of secondary envelopment; capsids acquire tegument proteins producing nascent virions. (7) Vesicular structures export mature particles to the extracellular space. b Cryogenic scanning electron microscopy (cryo-SEM) of VZV-infected MeWo cells seeded on micropatterned Quantifoil grids. Insets show fluorescence microscopy prior to vitrification (red fluorescence indicates infection with recombinant VZV expressing TK-RFP) and the corresponding cryo-SEM image after vitrification. Green square marks the cell selected for lamella preparation. c Cryo-FIB–milled lamella generated from the infected cell in panel (b). d Cryogenic transmission electron microscopy (cryo-TEM) montage of the lamella (16.85 Å/pixel). Orange box outlines tilt-series acquisition area for tomogram ev. e Capsids corresponding to critical stages of VZV morphogenesis (Roman numerals i to vi; dotted ellipses in panel (a). Each image shows a single slice through a low-pass-filtered tomogram; features are identified with colored arrowheads. (i) Empty capsid docked at a nuclear pore (pink). (ii) B-capsids containing scaffold protein adjacent to a neighboring capsid along the three-fold axis (blue violet). (iii) C-capsid surrounded by the nuclear egress complex (navy blue) and primary envelope (lime green) undergoing fusion with the ONM (white). (iv) C-capsid interacting with tegument layer (bright turquoise) and glycoprotein-coated cell membranes (tangerine) during secondary envelopment. (v) Fully assembled VZV particles undergoing late morphogenesis, including formation of a light particle; tegument (bright turquoise), envelope (gorse yellow), and membrane restriction (red). (vi) Plasma membrane-bound mature particle and a partially visible light particle in the extracellular environment (EE). Scale bars, 125 nm.
Several critical steps in the herpesvirus replication cycle were captured within cryo-ET tomograms generated from cryo-FIB milled cells infected with VZV (Fig. 1e; Suppl. Figs. 1 to 8; Suppl. Movies 1 to 6). These included capsids docked at the nuclear pore (Fig. 1ei; Suppl. Fig. 1; Suppl. Movie 1), newly synthesized capsids in the nucleus (Fig. 1eii; Suppl. Figures 2 and 3; Suppl. Movie 2), nuclear egress (Fig. 1eiii; Suppl. Fig. 4; Suppl. Movie 3), secondary envelopment (Fig. 1eiv; Suppl. Fig. 5; Suppl. Movie 4), VZV particle morphogenesis (Fig. 1ev; Suppl. Figure 6 and 7; Suppl. Movie 5), and VZV particles bound to the extracellular plasma membrane surface (Fig. 1evi; Suppl. Fig. 8; Suppl. Movie 6), a typical feature of this cell associated virus1. Within the infected cell nucleus, pro-capsids and A-, B-, and C-capsids, based on conventional morphological features, were distinguishable in 3D-space at this resolution. Procapsids were spherical and contained a solid density of scaffold protein (Suppl. Figs. 2 and 3); capsids apparently devoid of scaffold protein or dsDNA (conventional A-capsids) were identified (Suppl. Fig. 3b); B-capsids were observed and defined by the presence of scaffold protein (Fig. 1eii; Suppl. Fig. 3c); and C-capsids were distinguishable by their cavities filled with dsDNA (Figs. 1eiii, 2a,b; Suppl. Figs. 4 to 8). Procapsid assembly precursors were identified within infected cell nuclei, appearing as angular segments of capsomeres associated with scaffold protein with a curvature similar to the procapsids (Suppl. Fig. 2). Importantly, high-resolution detail was evident by the visualization of individual capsid capsomeres (Fig. 1eii, Suppl. Fig. 3; Suppl. Movie 2), individual nuclear egress complexes (Fig. 1eiii, Suppl. Figure 4; Suppl. Movie 3), and individual glycoproteins on enveloped VZV particles in the cytoplasm or bound to the plasma membrane (Fig. 1eiv and vi; Suppl. Figs. 5, 8, and 9; Suppl. Movie 5 and 6). Partially assembled virus particles were observed within cytoplasmic sites of VZV secondary envelopment and morphogenesis, including sites of tegument protein accumulation, based on the electron-dense protein surrounding the capsids within the reconstructed tomograms (Fig. 1eiv and 1ev; Suppl. Figs. 5 and 6). For example, a VZV light particle, a glycoprotein-studded envelope containing tegument proteins but without a capsid, and a complete particle were observed in a vesicular structure, and in a neighboring vesicular structure, an elongated VZV particle contained a C-capsid and the accumulation of tegument protein associated with a membranous structure (Suppl. Fig. 7b). The virus envelope within these vesicular structures was detected by machine learning-based segmentation (Suppl. Fig. 7c). Collectively, these direct visualizations reveal details of the VZV replication cycle previously unobserved in its native hydrated state inside infected cells, particularly for nuclear egress, tegument acquisition during secondary envelopment, and virus glycoprotein interactions with both intra- and extracellular membranes.
Subnanometer reconstruction of VZV capsid proteins in infected cells
From 194 tilt series, 1084 capsids were identified and classified based on classical morphological features, the presence or absence of scaffold protein or the virus genome in the capsid cavity. A visual inspection of the image contrast in the cavity of capsids without the dsDNA genome revealed a distribution of scaffold protein density, filled to sparse, suggesting that completely empty A-capsids are absent or very rare within infected cell nuclei (Suppl. Fig. 10). Residual scaffold protein was limited enough in some capsids to be obscured by conventional EM techniques due to the plane of the cut in ultrathin sections in previous studies. The cellular distribution of the 1084 VZV capsids was primarily localized inside the nucleus (n = 980) with only 9.3% detected in the cytoplasm, either as naked capsids, without tegument or envelope proteins (n = 42), or as complete enveloped VZV particles (n = 59) (Suppl. Table 3). The capsid distribution was typical for peak particle synthesis of VZV-infected cells, which aligns with thymidine kinase (TK) expression, an early gene, as determined by the red fluorescence signal produced from the recombinant pOka-TK-RFP virus.
An initial 13 Å cryo-ET map of the VZV capsid was generated by subtomogram averaging (STA) of 317 capsids (Fig. 2; Suppl. Fig. 11), containing variable quantities of scaffold protein or packaged dsDNA. This icosahedral symmetry-imposed map clearly shows the T = 16 icosahedron, composed of 60 asymmetric units. Each asymmetric unit consists of a single pentonal MCP subunit (PP), one peripentonal hexon (P), one central hexon (C), and half of an edge hexon (E), along with interdigitating triplexes (Ta, Tb, Tc, Td, and Te) (Fig. 2c, d). Cryo-ET maps were produced for the vertex around the 5-fold axis (8.3 Å) and the 3-fold symmetry axis (8.4 Å) by subsequent symmetry expansion and focused refinement of the 317 capsids. (Fig. 2e, f; Suppl. Fig. 11). The 8.3 Å vertex map had sufficient resolution to perform segmentation of individual proteins for the pentonal MCP (PP), six MCPs (P1–6) for the peripentonal hexon and the associated small capsomere interacting protein (SCP), triplex proteins TRX1 (monomer) and TRX2 (dimer) that form the α2β heterotrimers of Ta and Tc, and the three proteins, CVC1 (ORF43 monomer), CVC2 (ORF34 dimer), and ORF22 dimer, that assemble to form the capsid vertex specific component (CVSC) heteropentamer (Figs. 3 and 4; Suppl. Table 4 and 5). Similar to the vertex map, individual proteins were segmented for the 8.4 Å 3-fold symmetry axis cryo-ET map for the six MCPs (C1–6) for the central hexon and associated SCP, in addition to the TRX1 and TRX2 proteins of the Te triplex (Suppl. Fig. 12; Suppl. Table 4). The densities in both 3- and 5-fold axes of symmetry were segmented using the structure derived from single particle cryo-EM of VZV capsids (7BW6)39. All of the features for the pentonal, peripentonal hexon, and central hexon MCPs were apparent, including the tower, consisting of the crown and buttress regions (Fig. 3a to c; Suppl. Fig. 12), and the HK97 phage capsid protein-like fold facing the interior chamber of the capsid (Fig. 3d), which is evolutionary conserved for herpesviruses and dsDNA bacteriophages75. The resolution of these regions in the cryo-ET map was supported by the ability to observe densities for alpha helices in each of the MCP, TRX1, and TRX2 proteins and corresponding Map-Q scores (Fig. 3c; Suppl. Fig. 12c and e; Suppl. Fig. 13; Suppl. Movie 7). The classical HK97 phage capsid protein fold of the MCP was distinguishable with the long alpha helix (helix 4) interdigitating at the pseudo 3-fold axis of symmetry between the penton and neighboring peripentonal hexon MCPs (Fig. 3d). These data provide subnanometer resolution structural details of the MCP, SCP, TRX1, and TRX2 within the nucleus of a herpesvirus-infected cell, representing an unprecedented view of capsid architecture in situ.
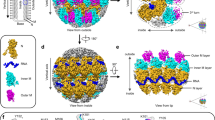
a A representative tomogram reconstruction produced from cryo-ET of a cryo-FIB/SEM milled region inside the nucleus of a vitrified, VZV-infected MeWo cell represented as a 3D volume. b A single slice from the tomogram in (a) in a region inside the nucleus of a vitrified VZV-infected MeWo cell, where a cluster of B-capsids (scaffold protein and without dsDNA labeled as B) and C-capsids (dsDNA genome labeled as C) were identified. Scale bars indicate 125 nm. c Surface representation of a cryo-ET map (13 Å) generated by subtomogram averaging with imposed icosahedral symmetry of 317 VZV capsids with a mixture of B and C capsids. The capsid is colored radially (510–700 Å from the center of the capsid). The yellow kite connects the 2-, 3-, and 5-fold axes of symmetry. d Zoom in view of a portion of (c) annotated with protein components that make up an asymmetric unit composed of the pentonal major capsid protein (PP), the three types of hexon (peripentonal (P1–P6), edge (E1–E3), and center (C1–C6)), and the five triplexes (Ta, Tb, Tc, Td, and Te). The inset (right) is a schematic of the asymmetric unit of the icosahedral capsid. e, f Cryo-ET maps generated by subtomogram averaging and focused classification of the VZV capsid vertex (e; 8.3 Å; EMD-49591) and 3-fold axis of symmetry (f; 8.4 Å; EMD-49886). The inset cryo-ET maps of the 13 Å VZV capsid cryo-ET map are projected along the 5-fold (e) and 3-fold (f) axes of symmetry. Peripentonal (P) and center (C) hexons are labeled. The color key indicates the local resolution (Å) of the cryo-ET maps.
a Segmentation of the 8.3 Å cryo-ET map based on the 7BW639 and a refined model based on the subnanometer map. The major capsid protein (MCP; ORF40) of the penton (PP) and the peripentonal hexons (P1–6), triplex proteins TRX1 (ORF20) and TRX2 (ORF41) for triplexes Ta and Tc, and small capsomere interacting protein (SCP; ORF23) are colored lavender blue (PP), very soft violet (P1,3,5), light coral (P2,4,6), deep sky blue (TRX1), Columbia blue (TRX2.1), aqua (TRX2.2), and dark gray (SCP). The colored shapes within the black outline demarcate a portion of the asymmetric unit (PP, P1–6 and associated SCP, and triplexes Ta and Tc) of the VZV capsid vertex, which is isolated in the dotted box (inset). b Extracted densities of the PP and P2 MCP with the tower, buttress and crown highlighted, and Ta triplex. c Exploded views of extracted densities for helices from the PP and P2 MCP, and Ta triplex. Each helix is numbered according to the primary structure, along with the beginning and end numbered amino acids. d Segmentation of the VZV capsid vertex without the triplexes. PP and P1–3 MCPs are identified with colored shapes. The dashed box outlines the region shown in the right-hand panel. Triangles represent the local 3-fold symmetry axes where the pentonal MCP (PP) intersects the two peripentonal hexon MCPs, P1 and P2. The black outline highlights the intersection between the HK97-like fold and the PP, P1, and P2 MCPs (inset).
The CVSC heteropentamer assembles as part of the capsid asymmetric unit in infected cell nuclei
The CVSC status as a capsid component has been difficult to assess due to its propensity to disassociate from vertices during herpesvirus capsid purification, including VZV39,41,45,76. In contrast, the CVSC density was apparent at all 12 vertices of the capsid structure in infected cell nuclei (Fig. 2c). Furthermore, the 8.3 Å cryo-ET map of the VZV capsid vertex was of sufficient resolution to extract densities for the CVSC, demonstrating that the heteropentamer is a capsid component acquired independently of the full tegument layer of the virus particle (Figs. 2e and 4a, c). An AlphaFold model of the CVSC structure oriented the head group for each CVC2 (ORF34) molecule at a downward angle compared to the extracted CVSC cryo-ET map density (Suppl. Fig. 14a, b). The CVC2 head group AlphaFold models were fitted into the cryo-EM density with the aid of molecular dynamics flexible fitting (Suppl. Fig. 14c; Suppl. Movie 8), enabling segmentation of the cryo-EM density into the five individual subunits of the CVSC heteropentamer (Figs. 4a–c). This revealed that the CVC1 subunit (ORF43 monomer), which has a limited interaction with TRX1 from triplex Te, primarily associates with all three Ta triplex subunits along with MCP P1 and P2 of the peripentonal hexon from the neighboring asymmetric unit (Fig. 4b, d, and e). A helical bundle forms between the CVC1 subunit, the two ORF22 proteins and the two CVC2 proteins, enabling the CVC2 head group to interact with the peripentonal hexon facing side of the pentonal MCP crown (Fig. 4a). Since almost all capsids analyzed were intra-nuclear, these observations definitively demonstrate that CVSC assembly at the capsid vertices occurs inside the nucleus. Furthermore, these findings enabled us to redefine the asymmetric unit of the mature C-capsid to be composed of six C hexon subunits (MCP and SCP), three E hexon subunits, and a pair of three peripentonal hexon subunits from neighboring hexons, one pentonal MCP, five interdigitating triplexes, and a single CVSC heteropentamer (Figs. 2d and 4b; Suppl. Fig. 15a). This arrangement differs from previous models of alphaherpesvirus asymmetric units, where the interaction between CVC1 with triplexes Ta and Tc and MCP P1 and P2 were not used to define the asymmetric unit, proteins were missing, or proteins from a neighboring asymmetric unit were included (Suppl. Fig. 15).
a Segmentation of the 8.3 Å cryo-ET map of the vertex around the 5-fold axis of the capsid to isolate the CVSC and fit an AlphaFold 2 model using MDFF simulation and the 7BW639 model. The MCP, SCP, TRX1, and TRX2 are colored as in Fig. 3 and the CVSC subunits are colored orange (CVC1; ORF43), red (CVC2.1; ORF34), falu red (CVC2.2; ORF34), yellow (VP1/2.1; ORF22), and gold (VP1/2.2; ORF22). The black outline highlights a portion of the VZV capsid vertex with the CVSC, which is isolated in the dashed box (inset). The lower panels show side views of the complete vertex map and the extracted region in the dotted box. b A cartoon representing the CVSC components’ locations in relation to the redefined asymmetric unit compared to Fig. 2C. Colored shapes define the pentonal major capsid protein (PP), the three types of hexon (peripentonal (P1-P6), edge (E1-E3), and center (C1-C6)), the five triplexes (Ta, Tb, Tc, Td, and Te), and the CVSC. c Exploded view of the CVSC subunits. d, e Extracted densities revealing (d) CVC1 (ORF43) interactions with triplexes Ta and Tc, and (e) peripentonal hexon MCP P1 and P2. Models of the CVC1, TRX1, TRX2, P1, and P2 within the cryo-ET map densities are shown in the left-hand panels. Right-hand panels show plane views of the molecular models (far right) that highlight likely interaction sites between CVC1, the triplexes (d), and peripentonal MCPs (e) and the boundaries of the planes (black boxes).
CVSC acquisition redefines capsid assembly intermediates formed during herpesvirus capsid assembly
Given uncertainties about the stages of capsid assembly based on studies of purified capsids31,45,76,77,78, we investigated the distribution of CVSC heteropentamers at vertices of capsids in infected cells to determine whether the CVSC occupancy could be used to differentiate capsid types and achieve a more precise characterization for the stages of capsid assembly (Fig. 5). Uniform Manifold Approximation and Project (UMAP) is a dimension reduction technique that can be used for the visualization and categorization of complex data79, including structural heterogeneity in single particle Cryo-EM80,81. Application of this unsupervised computational approach (Fig. 5a) using three-axis orthogonal projection in latent space classified the 317 capsids that were used to generate the 8.3 Å vertex cryo-ET map (Fig. 2). Each capsid image was aligned to its symmetry axes, and icosahedral symmetry was applied to individual particles before the projections were generated to compensate for the missing wedge effect intrinsic to cryo-ET data82. Three clusters were differentiated in latent space, revealing limited (i) versus extensive (ii) scaffold protein in the capsid cavity, and those with packaged dsDNA (iii) corresponding to C-capsids, as viewed along the 2-, 3-, and 5-fold axes of symmetry and the STA for each class (Fig. 5a–c). The detection of limited quantities of scaffold protein for class i capsids enabled the differentiation from conventionally designated B-capsids. The absence of any density inside the capsid cavity, previously established for classical empty A-capsids based on conventional negative-stained images30, was not detected within infected cell nuclei. Residual scaffold protein was visible in the native structures preserved by vitrification performed in this study (Suppl. Fig. 10). In addition to capsid cavity content differences, the three UMAP capsid clusters were found to differ in CVSC occupancy at the vertices (Fig. 5d–f). For each of the 317 capsids, sub-particles were extracted from the 12 vertices and aligned to the same C5 symmetry axis. These particles were initially subject to binary reference-free focused classification, resulting in two classes reflecting vertices without or with CVSC occupancy (Fig. 5d, e). The CVSC was either absent or present, with the arms and lobes of the CVSC clearly visible in the C5 symmetry STA maps (Fig. 5d). Capsid vertices were then classified into those without and with the CVSC and by their UMAP cluster, i to iii (Fig. 5e). As expected, CVSC occupancy was highest on C-capsids in infected cells, as has been reported for purified capsids33,37,40,42,47, and because they are considered complete and ready for egress from the nucleus for final tegumentation and envelopment in the cytoplasm45. Persistent scaffold protein is the conventional marker for B-capsids by EM, resulting in greater buoyant density than empty A-capsids30. Notably, capsids differed in CVSC occupancy depending on whether scaffold protein was limited or extensive within the capsid cavity, with the former having greater CVSC occupancy than the latter (Fig. 5b–f). Critically, for the 12 vertices of individual capsids, CVSC occupancy at each vertex increased with decreasing levels of scaffold protein density (Fig. 5f). Since C-capsids have the highest CVSC occupancy, these findings indicate that capsids with relatively more CVSC occupancy and less scaffold protein are capsid assembly intermediates (CAIs) further along the pathway of capsid maturation than those with less CVSC occupancy and more residual scaffold protein.
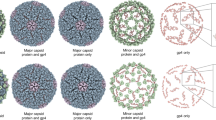
a Uniform manifold approximation and projection (UMAP) of 317 VZV capsids producing three clusters, i, ii and iii. b 2D class averages of clusters i to iii projected along 2-, 3-, and 5-fold axes of symmetry. c Subtomogram reconstruction of the three capsid classes, i, ii, and iii. Capsids are colored radially from the center of the capsid to the outer shell: beige, green, white, and red. The transverse planes show the empty cavity (i), scaffold-filled cavity (ii), and dsDNA-filled cavity (iii) of the VZV capsids. d Subtomogram reconstructions of the capsid vertex without and with the CVSC. The vertices are colored radially as for panel (b), with the CVSC colored yellow. e Normalized frequency of i, ii, and iii capsid classes without or with the CVSC. Values for each class are given above each bar of the chart. f Increased CVSC occupancy at capsid vertices is associated with decreased scaffold protein. Slice view of representative polished particles (EMAN2) extracted from tomograms and cartoons depicting the scaffold protein visible in the capsid cavity. Numbers represent the frequency of vertices occupied by the CVSC. g Subtomogram averaging after symmetry relaxation and focused classification. The frequency (n) of VZV capsid vertices without (0) or with 1 to 5 copies of the CVSC (yellow) is indicated for each vertex. h Non-consecutive distribution of the CVSC (yellow) at the VZV capsid vertices with the frequency (n) of 2 (2’) or 3 (3’) copies after subtomogram averaging and focused classification. i A violin plot representing the frequency of the CVSC (60 copies per capsid; 5 CVSC subunits per 12 vertices of the capsid) for each capsid class, i, ii and iii. The solid line represents the median, and the dotted lines represent quartiles. Asterisks above the violin plots indicate p-values (<0.0001) for significant differences from an ANOVA with Tukey’s multiple comparison test.
To further scrutinize the CVSC on capsids in infected cells, vertex particles were subjected to C5 symmetry expansion, generating five sub-particles for the asymmetric orientations. Additional focused classification using a mask covering a single CVSC site produced two classes of CVSC occupancy at each individual location: those with or without the heteropentamer. This procedure established the frequency of CVSC heteropentamers, producing vertex sub-particles with either zero copies, 1 to 5 copies, or 2 and 3 copies of non-consecutive CVSC distribution, which were used for STA without symmetry imposed (Fig. 5g, h). Each capsid has a potential maximum of 60 copies of the CVSC heteropentamer, as there are 12 capsid vertices and 5 CVSC per vertex, including the portal vertex29,34,35,36,38,45,48,49,50. For the 317 capsids in the present study, there were 3,804 vertices. Frequencies ranged from 290 vertices without a CVSC to 1,198 vertices with a full complement of five copies of the CVSC (Fig. 5g). Additionally, alternative distributions of the CVSC were observed at vertices where 2 or 3 copies of the heteropentamer were not adjacent to one another, indicating a non-cooperative attachment of the heteropentamer to these capsid vertices (Fig. 5h). In contrast to purified VZV C-capsids41, intranuclear C-capsids in infected cells had almost complete vertex occupancy (52; SEM = 0.861), demonstrating their maturity (Fig. 5i). CAIs with limited scaffold protein had greater CVSC occupancy (39; SEM = 0.859) compared to CAIs with extensive scaffold protein (32; SEM = 0.753), providing additional evidence that the former represent a stage closer to dsDNA encapsidation while the latter are at an earlier stage in nucleocapsid synthesis. Taken together, these subnanometer resolution map observations strongly support the concept that CVSC occupancy can be used as an indicator for the progression of herpesvirus capsid assembly in situ and identifies capsids with varying levels of residual scaffold protein as CAIs along a continuum, having the potential for genome packaging rather than failed assembly dead ends.
Portal vertex architecture differences between CAIs and C-capsids
In contrast to previous high-resolution single particle cryo-EM structures of biochemically purified herpesvirus capsids, the portal vertex for VZV capsids could not be reconstructed in previous reports39,41 but has been achieved by Cao et al.68; this was not performed in infected cells in situ. To address this issue, we developed a RELION pipeline to reconstruct the VZV portal vertex, which was successfully identified for CAIs without the packaged dsDNA genome, or C-capsids within a subset of cryo-FIB/ET tomograms using STA and 3D classification of capsid vertices (Fig. 6; Suppl. Figs. 16 to 18; Suppl. Table 4). An initial cryo-ET map (23.6 Å) for the portal vertex revealed a density within the capsid cavity reminiscent of the herpesvirus portal34,38,48,49,50, and an absence of density for penton proteins compared to the 8.3 Å cryo-ET map of the capsid vertex (Figs. 3 to 5; Suppl. Fig. 17a). Further STA of the portal density generated a 22.9 Å cryo-ET map of the VZV portal with C12 symmetry (Suppl. Fig. 17b). A molecular model based on an AlphaFold prediction of an individual VZV portal protein (pORF54; 769aa) and modeling of 12 subunits based on the entire HSV-1 portal was placed into the 22.9 Å cryo-ET map of the VZV portal (Suppl. Fig. 17c,d). Additional densities in the portal cryo-ET map were attributed to poor modeling by AlphaFold of amino acids 1 to 64 and 656 to 769, which were disordered and removed from the model. Architectural differences between the portal vertices of CAIs and C-capsids were visible when cryo-ET map reconstructions were performed based on capsid type (Fig. 6; Suppl. Fig. 18). The portal cap was absent for CAIs compared to C-capsids, which was apparent when the entire capsid was reconstructed (Suppl. Fig. 18). The portal structure of C-capsids was confirmed as the portal density in the capsid cavity was only present at a single vertex (Suppl. Fig. 18). Further capsid-type specific architectural differences were evident from portal vertex cryo-ET maps and fitment of the molecular models for the peripentonal hexons, Ta and Tc triplexes, CVC1, ORF22 helices, and the portal (Fig. 6). As expected, densities for the pentonal MCP and alpha helices for the CVSC were absent and a channel for dsDNA translocation was visible through the center for the CAIs capsid portal vertex cryo-ET map (Fig. 6b, c). In contrast, the C-capsid cryo-ET map had a portal cap with densities for the CVSC, which filled the portal vertex channel (Fig. 6e, f), presumably, tentacle helices and the dsDNA genome terminus as previously described for HSV-138. As expected from the VZV vertex-focused classification analysis (Fig. 5g–i), occupancy of the CVSC at the non-portal vertices of the CAI was incomplete, whereas each vertex for the C-capsid exhibited a full complement of the CVSC (Fig. 6a, d and Suppl. Fig. 16). These findings further support the CVSC as a defining element and thus a marker of herpesvirus capsid assembly and maturity inside the cell nucleus during replication.
a–f Cryo-ET maps of CAI and C-capsids and their portal vertices. a, d Cryo-ET maps of the VZV CAIs (A; EMD-49470) and C-capsid (D; EMD-49472) projected along the 3-fold axis of symmetry and colored radially (510–700 Å from the center of the capsid). The yellow triangle encompasses the 2-, 3-, and 5-fold axes of symmetry. b, e Cryo-ET maps of the portal vertices from CAIs (B; EMD-49471) and C-capsids (E; EMD-49473). The left-hand panels show the vertices along the z-axis of the cryo-ET map. The dashed black line shows the slice for views along the y-axis in the upper-right-hand panels. The lower right-hand panels for the CAIs and C-capsids show transparent views of the cryo-ET maps with the portal map (see Suppl. Fig. 15) aligned to the portal vertex map density. c, f The molecular model of the VZV capsid vertex without the pentonal MCP was fit into the cryo-ET maps of the CAIs and C-capsid portal vertices. The MCP of peripentonal hexons, triplexes Ta and Tc (TRX1, TRX2.1, and TRX2.2), small capsomere interacting protein (SCP), CVSC subunits CVC1, VP1/2.1 and VP1/2.2, and the VZV portal protein ORF54 are colored light coral or very soft violet (MCP), deep sky blue (TRX1), Columbia blue (TRX2.1), aqua (TRX2.2), dark gray (SCP), orange (CVC1), yellow (VP1/2.1), gold (VP1/2.2), and olive yellow (ORF54). Dashed lines indicate the slices used for the right-hand panels to more clearly visualize the portal. The dashed circles highlight the absence of CVSC densities in the portal vertex cryo-ET map for the B-capsid compared to the C-capsid (right-hand panels).
Discussion
The cell nucleus is a complex, crowded, dynamic environment, raising many questions about how herpesvirus capsids assemble within this large organelle. This study reveals the power of cryo-ET for the in situ investigation of herpesvirus assembly and morphogenesis at subnanometer resolution, the localization of macromolecular structures inside herpesvirus-infected cell nuclei, and the reconstruction and segmentation of individual capsid proteins, allowing the identification of capsid assembly intermediates (CAIs) in their native environment for replication. In addition, the observations of procapsid precursors and architectural differences in capsid vertices revealed by the CVSC distribution on capsids in VZV-infected cells provide a transformative insight relevant for understanding the temporal nature of herpesvirus capsid assembly inside the cell nucleus.
Herpesviruses are thought to assemble in a similar fashion to dsDNA bacteriophages, where the capsomeres coalesce with the scaffold protein to generate procapsids28,29. The high-resolution imaging inside the nucleus of VZV-infected cells in the present study revealed angular segment structures where capsomeres, formed by the MCP and SCP, were associated with electron-dense material presumed to be scaffold protein (Suppl. Figure 2). This interpretation is supported by cell-free systems that recapitulate herpesvirus capsid assembly by expression of the MCP, TRX1, TRX2, and VP22a proteins83. VP22a is the scaffold protein expressed from UL26.5 (VZV ORF33.5) cleaved by the VP24 protease during maturation84,85. VP24 is a cleavage product derived from UL26 (VZV ORF33), where the protease autocatalytically cleaves the UL26 gene product into VP24 and VP21. VP24 protease activity triggers spherical procapsids to undergo angularization, and the SCP (VP26) is thought to provide additional capsid stability33,86,87,88. For VZV, the ORF33 protease is required for replication, and an SCP (ORF23) null virus produces predominantly spherical capsids, replicates poorly in cell culture and is non-viable in human skin xenografts89,90. Furthermore, previous transient transfection studies have shown that the scaffold protein is required for intranuclear localization of the HSV-1 MCP (VP5)91, whereas nuclear translocation of the VZV MCP is impaired by deleting the SCP90. Further characterization of HSV procapsid formation in cell-free expression systems noted the fragility and production of angular segments, leading to a proposed model where capsomeres and scaffold protein aggregate to form angular segments, which in turn form spherical procapsids92. In support of this, the angular segment structures of VZV demonstrate that herpesvirus procapsids form within the nucleus through cooperative assembly of capsomere-associated scaffold protein subunits rather than directly on a spherical precursor structure that recruits individual capsomeres.
Our study demonstrates that the acquisition of high-resolution 3D structure information derived from vitrified cells is critical for differentiating between capsid types that occur during assembly. Empty capsids were not apparent in the infected cell nucleus, whereas previous observations with traditional thin-section EM of infected cell nuclei frequently identify capsid cavities devoid of protein and dsDNA30, leading to their conventional classification as A-capsids. From our 3D observations, evidence of scaffold remnants inside the capsid cavity was lost when the 2D sectional plane spanned the entire diameter of the capsid, leading to their potential mischaracterization. Furthermore, our data eliminate fixation, sectioning, and staining artefacts, which could also lead to mischaracterization of A-capsids. In addition, capsid purification methods can produce empty capsids as an artefact since the pressurized dsDNA genome can be ejected due to the instability of the CVSC at the portal vertex32. In contrast, the 3D analyses of capsid architecture coupled with the binding of CVSC heteropentamers to icosahedral vertices in the infected cell point to the need for an amended classification. In the present study, capsids with packaged dsDNA had the highest level of CVSC occupancy, consistent with previous single particle cryo-EM of purified C-capsids33,37,40,42,47, and, furthermore, assembly intermediates were identifiable by correlating CVSC occupancy with levels of residual scaffold protein. Increasing CVSC occupancy at the vertices and of heteropentamer copies (up to the maximum of five) was associated with diminished quantities of scaffold protein in the capsid cavity. Although the 3-axis orthogonal projection clusters capsids into three classes, our observations indicate that the conventionally designated A- and B-capsids (classes i and ii) represent a continuum of CAIs that can potentially progress towards dsDNA encapsidation and formation of mature capsids ready for nuclear egress.
Earlier high-resolution cryo-EM studies of VZV capsids were not able to visualize the portal vertex39,41. However, a recent study reconstructed the portal vertex from purified capsids68. In the present in situ study, reconstruction of the portal vertex for CAIs and C-capsids, along with acquisition of the VZV portal cap and packaged dsDNA for C-capsids, was performed with capsids inside the infected cell nucleus. The dodecameric portal for VZV is composed of pORF54, which is essential for the packaging and cleavage of the dsDNA genome93,94. Similar to the VZV cryo-ET maps of the portal vertex in the present study, cryo-EM studies of purified HSV, HCMV, KSHV, and EBV have described the molecular architecture of the portal and portal vertex to have a C12 to C5 symmetry mismatch of the portal and vertex constituents34,35,38,49. For HSV, the CVSC is visible at the portal vertex and the CVC2 protein is required for dsDNA packaging38. The CVC1 subunit sits atop the Ta and Tc triplexes. At the portal, for HSV-1, the Ta triplexes are rotated 120° anticlockwise about their respective centers compared to pentonal Ta triplexes38. Thus, the triplexes and the CVC1 are at an altered angle compared to the other 11 vertices. The MCP of HSV in the peripentonal hexon at position 6 has spine helices that extend into the pentonal space at non-portal vertices38. The extension of the N-terminal helix was not visible in previous VZV cryo-EM structures, and that absence can now be attributed to the loss of ‘signal’ due to the combined STA of the portal and non-portal vertices39,41. Moreover, the head groups of the CVC2 dimer, important for dsDNA genome packaging and retention, form part of the portal cap, which was not observed in previous studies of VZV, and explains the role of the CVSC in capsid maturation.
Biochemical and cryo-ET studies with HSV-1 have demonstrated either the absence or presence of multiple vertices containing portal densities using highly purified HSV-1 procapsids from a protease-deficient virus (m100) or A-capsids and pUL6-labeled B-capsids from wild-type virus, or a UL25 null virus (KUL25NS)55,56,95,96,97. Our in situ study of naturally synthesized capsids did not quantify the frequency of portals per capsid because portal structures were not observed for all dsDNA genome-containing C-capsids. Critically, a portal is required for genome packaging. The lack of at least one portal per C-capsid was attributable to a combination of incomplete angular sampling, the low signal-to-noise ratio of in situ cryo-ET, and the missing wedge artefact, leaving gaps in Fourier space, potentially obscuring the portal if the capsid is at a non-optimal orientation in the tilt series. Thus, any estimate of portal frequency per capsid would be grossly underestimated. Therefore, our data do not undermine previous portal distribution estimates. To address the observations of the previous HSV-1 studies55,56,95,96,97, future work must be performed to establish whether herpesvirus capsids form inside the nucleus without or with multiple portals.
In addition to the retention of the dsDNA genome, the CVSC has also been linked to nuclear egress, a critical mechanism enabling capsids to leave the nucleus and enter the cytoplasm to complete herpesvirus morphogenesis98,99. The process of nuclear egress is thought to be initiated by capsids budding at the nuclear envelope into the perinuclear space through interactions of CVC2 head groups with the nuclear egress complex (NEC) embedded in the INM, leading to acquisition of a primary envelope98,100. Although the NEC has been shown to induce curvature in the absence of CVC2 via a mechanism that involves a random lattice formation101, these studies were performed using a US3 mutant that causes capsids to become trapped within the perinuclear space102. It has also been proposed that the CVSC functions as a selective switch to preferentially allow the dsDNA genome-filled C-capsids to leave the nucleus45,99. Previous studies of capsid structures at subnanometer or near atomic resolution were performed with capsids purified from the nuclei of infected cells or purified herpesvirus particles, providing conflicting information about the quantities of the CVSC associated with capsid vertices33,36,37,38,39,40,41,47. For VZV, precise information about the occupancy of the CVSC was unknown, likely due to the purification of these particles by ultracentrifugation and detergent-based removal of the envelope and tegument from C-capsids39,41. Purification approaches have limitations in defining the stages of acquisition of the CVSC on capsids. However, the extensive interactions among all five subunits of the heteropentamer imply that the CVSC components co-assemble in the nucleus and act together as a functional unit in binding the capsid vertices33. The present study provides definitive proof that the CVSC can assemble as a heteropentamer and bind to capsids inside the nuclei of VZV-infected cells. Moreover, the application of STA and focused classification to the capsids further enabled the determination of the CVSC occupancy at individual sites at the vertices in relationship to different capsid types, supporting the notion that capsids with variable levels of scaffold protein are intermediates. Studies of HCMV support this concept, as the frequency of morphologically defined A-capsids within infected cell nuclei was very low compared to B- and C-capsids31,32 and C-capsids purified from infected cell nuclei ejected their genomes, leading to higher quantities of apparent A-capsids compared to those seen by TEM32. Furthermore, the A-capsids were covered with the HCMV tegument protein pp150, which is equivalent to the CVSC of alphaherpesviruses and typically associated with C-capsids32,42,103. Thus, high-resolution imaging coupled with state-of-the-art image classification approaches can now assign temporal information to the dynamic herpesvirus capsid assembly process in situ. The near-complete occupancy of the CVSC on C-capsids and the increased levels of the CVSC on capsids with limited scaffold protein further support a selective function of the CVSC for nuclear egress, as both of these capsid types are found in extracellular VZV particles.
Our study provides direct evidence that the CVSC has a critical role in herpesvirus capsid assembly within the host cell nucleus, revealing heterogeneity in CVSC occupancy associated with CAIs, with the highest frequency found on dsDNA genome-filled C-capsids. Rather than discrete A-, B-, and C-capsid categories, we propose a model of herpesvirus capsid assembly where CVSC acquisition serves as a temporal marker towards completion; CAIs progress along a continuum defined by decreasing scaffold protein and increasing CVSC occupancy prior to genome packaging. Early intermediates retain abundant scaffold and have few CVSCs, whereas later intermediates contain limited scaffold with more extensive CVSC occupancy at the vertices, culminating in C-capsids with full vertex occupancy and packaged genomes. This model integrates structural heterogeneity into a dynamic pathway, resolves the ambiguity surrounding A- and B-capsids, and highlights CVSC acquisition as a critical determinant of capsid stability, genome retention, and readiness for nuclear egress. Our observations shed light on the unresolved question about whether A- and B-capsids are assembly dead ends, instead suggesting that A-capsids represent artefacts from purification, whereas those conventionally grouped as B-capsids are assembly intermediates that can be further differentiated based on CVSC occupancy. Our findings underscore the intricate and dynamic nature of capsid assembly in situ during herpesvirus replication, providing conceptual insights for the development of therapeutic strategies targeting herpesvirus capsid maturation.
Methods
Reagents, consumables, and resources
All reagents, consumables, and resources, including supplier information where applicable, are provided in Suppl. Table 6.
Cells lines
MeWo cells (HTB-65; ATCC) were propagated in minimal essential medium (Corning Cellgro) supplemented with 10% fetal bovine serum (FBS; Invitrogen), nonessential amino acids (100 μM; Corning Cellgro), antibiotics (penicillin, 100 U/ml; streptomycin, 100 μg/ml; Invitrogen) and the antifungal agent amphotericin B (250 µg/ml; Invitrogen) at 37 °C in a humidified atmosphere with 5% CO2.
Viruses
The VZV parental Oka strain was originally cloned into a bacterial artificial chromosome (BAC) and designated pPOKA-BAC-DX104. Recombinant viruses pOka-TK-GFP (pOka-rTK) and pOka-TK-RFP were generated in a previous study71,105.
ECM micropatterning of SiO2 EM grids
To position MeWo cells centrally within EM grid squares, 300 mesh gold grids coated with a thin perforated layer of SiO2 (Quantifoil R2/2) were subjected to micropatterning following the procedure available at protocols.io (https://www.protocols.io/view/micropatterning-em-grids-for-cryo-electron-tomogra-rm7vz3wqrgx1/v1). Briefly, EM grids were exposed to atmospheric plasma, incubated in poly-L-lysine (0.01%; Sigma) and subsequently incubated with HEPES (0.1 M; Sigma) + mPEGSuccinimidyl Valerate (PEG-SVA MW 5,000; 100 mg/ml; Laysan Bio Inc.), then PLPP gel (1:6 dilution in H2O; Alveole). The coated grids were exposed to UV light according to a digitally generated micropattern (30 × 30 µm or 33 × 33 µm to match the 300 mesh spacing) that covered the EM grid square using a Primo photo-micropatterning system (Alveole) to selectively degrade the PEG coating to remove the hydrophobic antifouling layer. An extracellular matrix (ECM) surrogate, Gelatin Oregon Green 488 (1 mg/ml; Invitrogen), was applied to the grids prior to cell seeding.
Vitrification of VZV-infected MeWo cells on ECM micropatterned grids
Cell-free VZV cannot be purified to high titers, making direct infection of cells grown on EM grids with high MOIs needed for productive infection impractical. Thus, infected cells were seeded directly onto ECM micropatterned EM grids. VZV (pOKA-TK-RFP) infected MeWo cells were passaged onto freshly seeded uninfected MeWo cells 24 hours prior to the day of seeding. On the day of seeding, the VZV-infected MeWo cells were then passaged onto uninfected MeWo cells and incubated for 8 hours. The infected cells were then trypsinized, centrifuged at 400xg for 5 min, then resuspended in maintenance media to yield one cell per grid square; determined empirically on the day of seeding. Infected and uninfected cell suspensions were applied to the ECM micropatterned grids in 35 mm glass-bottomed dishes (MatTech P35G-1.5-14-C). After 2 h incubation at 37 °C in a humidified atmosphere with 5% CO2, 3 ml of maintenance media was added and incubated for an additional 16 hours. Grids were removed from the maintenance media and plunged frozen into liquid ethane using an EM GP2 (Leica) with humidity at 99% and a blot time of 9 seconds.
Cryo-FIB/SEM of vitrified VZV-infected MeWo cells
An Aquilos 2 cryo-FIB/SEM microscope (ThermoFisher) was used to generate lamella from VZV-infected cells on micropatterned grids. Prior to milling, the grids were platinum-coated. Microexpansion joints of 1 µm width and 4 µm between the milled area and the microexpansion joint were used to relieve tension on the final lamellae106. The primary milling step was performed at 1 nA, and 5 µm height and 15 µm width with two 1 µm microexpansion joints milled on either side at 4 µm from the lamella milling site. Successive milling steps of 3 µm (300pA), 1 µm (100pA), 500 nm (50pA), and a final polishing step of 175 nm (30pA) were performed to generate a lamella with a target thickness of 175 nm. Lamella were produced for 6 to 10 cells per grid.
Cryo-ET of whole or cryo-FIB/SEM milled VZV-infected MeWo cells
Tilt series were either collected at the periphery of whole vitrified VZV-infected MeWo cells on EM grids, using a 200 kV Talos Arctica (FEI) or at cryo-FIB/SEM generated lamella, using a 300 kV Titan Krios (FEI) controlled by SerialEM107 to automate the data collection procedure using the parameters outlined in Suppl. Table 1. To identify areas of interest for tilt series collection, medium magnification (4,800-11,500x) montages were generated, and imaging sites were determined empirically by eye. Due to the large size of herpesvirus capsids, a 3.44 Å/pixel size was chosen to accommodate areas where high frequencies of VZV capsids were present in the nucleus. A total of 399 tilt series were collected from cryo-FIB/SEM milled lamella across five, 3-day sessions on 300 kV Titan Krios microscopes. A total of 194 tomograms were generated for VZV capsid quantification, analysis, and cryo-ET map reconstructions.
Cryo-ET image processing
Motion correction, damage compensation, CTF estimation, and tilt series generation were performed using Warp108. Tomogram reconstruction for visualization was performed using the IMOD tomography GUI Etomo109. Tomogram processing and associated movie files were generated using 3dmod110. Segmentation of tomograms was performed using Dragonfly software (Version 2022.2 for [Windows]. Comet Technologies Canada Inc., Montreal, Canada; software available at https://www.theobjects.com/dragonfly). Movies were generated using FFmpeg (FFmpeg Developers. (2016). ffmpeg tool (Version be1d324) available from http://ffmpeg.org/).
Subnanometer resolution cryo-ET map reconstruction of the VZV capsid vertex
To reconstruct the VZV capsid vertex and 3-fold axis of symmetry, a WARP-EMAN2 data processing strategy was used (Suppl. Fig. 11). Tilt series generated by WARP were imported into EMAN2 for tomogram alignment using patch tracking, where handedness was checked and CTF measured. The convolutional neural network was employed to select particles (box size 512; n = 1084 capsids) from 194 tomograms and make sets (bin2). An initial model was generated from the sets using e2spt_sgd_new.py (--res=50.0; --niter=30; --shrink=4; --sym=icos) followed by subtomogram-subtilt refinement using e2spt_refine_new.py (--startres=30.0; --sym=icos; --iters=p3,t2,p,t,r,d; --keep=0.95; --tophat localwiener –goldstandard). Classification with icosahedral symmetry was performed on the output to remove poor-quality particles that can negatively influence the subtomogram averaging; e2spt_refinemulti_new.py (--nref 3; --maxres 40; --loadali3d; --skipali; --sym icos; --maskalign spt_01/mask.hdf; --niter 20). Further refinement of 317 capsids produced a cryo-ET map of the entire VZV capsids of 13.0 Å resolution. To generate a subnanometer resolution cryo-ET map for the VZV capsid vertex, symmetry expansion was performed with the 317 capsids using e2spt_extract.py (--postxf icos,0,0,110; --compressbits 8; --boxsz_unbin 192; --newlabel pent2; --mindist 500; --loadali2d) followed by building sets; e2spt_buildsets.py (--label=pent2; --allparticles; --spliteo). An average of the extracted particles was generated from the set using e2proc3d.py (--average; --averager mean.tomo; --avg_byxf). The average was then refined using e2spt_refine_new.py (--startres=30.0; --sym=c5; --iters=p3,t2,p,t,r,d; --keep=0.8,0.95,0.95; --tophat localwiener; --localrefine; --goldstandard) to achieve a cryo-ET map of the VZV capsid vertex at 8.3 Å resolution. A similar procedure was used for the 3-fold axis of symmetry to achieve a cryo-ET map at 8.4 Å resolution.
Segmentation of the 3-fold and 5-fold cryo-ET maps
The 3- and 5-fold cryo-ET maps of the VZV capsid were segmented based on the 7BW6 asymmetric unit model of the VZV A-capsid39 using ChimeraX111. To segment the CVSC density in the 5-fold axis cryo-ET map, AlphaFold 2112 was used to model the heteropentamer based on the full-length proteins CVC1 (ORF43; monomer) and CVC2 (ORF34; dimer), and 45 C-terminal amino acids (2724-2768) of VP1/2 (ORF22). Molecular dynamics flexible fitting (MDFF) was performed to fit the two head groups of the CVC2 dimer into the cryo-ET map using NAMD113, following the procedure outlined in the VMD114 tutorial. Conversion of the cryo-EM map to an MDFF potential, rigid body docking of the CVSC, initial structure generation, and secondary structure restraint definitions were all performed using VMD. The output files were used as input to run NAMD with the following parameters: step 1 -gscale 0.3, -numsteps 50000; step 2 -gscale 10, -numsteps 100000. A model of the asymmetric unit of the VZV capsid vertex, including the CVSC, was refined using ISOLDE115. All images and movie files were generated using ChimeraX. Movies were generated using FFmpeg.
Three-axis orthogonal projection and uniform manifold approximation projection
The 3D VZV capsid particles were aligned to the symmetrized capsid reference from the EMAN2 icosahedral refinement. The subsequent 2D sub-tilt particles at the corresponding orientation were reconstructed with icosahedral symmetry to obtain symmetrized 3D particles free from missing wedge artifacts. Each particle was projected along three orthogonal axes of thin central slabs (72 Å), using only one quarter of the volume closest to the center perpendicular to the corresponding axis. The resulting three 2D image projections per capsid were stacked and flattened into a 1D array that was used as input for uniform manifold approximation projection (UMAP). After manifold embedding, particles on the 2D UMAP space were clustered using a K-means algorithm to produce the individual classes.
Reconstruction of the VZV Portal Vertex
To reconstruct the VZV portal vertex, a WARP-IMOD-EMAN2-Relion data processing strategy was used (Suppl. Fig. 16). Tilt series were aligned using IMOD, and for the final aligned stack, unbinned CTF estimation was performed using CTFplotter with the ‘Find astigmatism option’ checked. Tomograms for Relion were generated using relion_tomo_reconstruct_tomogram and used for particle picking of capsids in EMAN2 (n = 471). Tomograms and particle coordinates were imported into Relion using Tomo Import. Pseudo-subtomos were used to make subtomograms (bin 4; 13.76 Å/pixel) using a box size (pixels) of 360 and a cropped box size (pixels) of 180. An initial model was generated using a 3D initial model, and a mask was generated using Dynamo116 and Mask Create. A 3D refinement was performed with 3D auto-refine with an initial low-pass filter of 50 Å and I3 symmetry. Particle reconstruction was performed (bin2; 6.88 Å/pixel) with Tomo reconstruct particle with a box size of 440 and a cropped box size of 220 with I3 symmetry, which was used to generate a new mask, followed by reconstruction of subtomograms using Make pseudo-subtomos. A 3D refinement was performed with 3D auto-refine with an initial low-pass filter of 30 Å and I3 symmetry. The output was used for Tomo frame alignment, particle reconstruction (bin 1; 3.44 Å/pixel; box size 440 pixels; I3 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 14 Å and I3 symmetry, generating a 12.1 Å cryo-ET map of the VZV capsid.
Symmetry expansion was performed using relion_particle_symmetry_expand with --sym I3. This process produces 5 rotations per vertex; redundant _rlnAngleRot values were removed to leave 12 unique vertices per capsid (n = 5,652). Vertices beyond the boundaries of the tomograms were removed (n = 5,318). The Z height was adjusted to center the vertices in the center of the box using relion_star_handler and --center_Z command. The output was used for particle reconstruction (bin 4; 13.76 Å/pixel; initial box size 120 pixels; cropped box size 60 pixels; C5 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 50 Å and C5 symmetry. The output was used for particle reconstruction (bin 2; 6.88 Å/pixel; box size 240 pixels; cropped box size 120 pixels; C5 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 30 Å and C5 symmetry. The output was used for particle reconstruction (bin 1; 3.44 Å/pixel; box size 240 pixels; C5 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 15 Å and C5 symmetry, generating a 10.3 Å cryo-ET map of the VZV capsid vertex.
The output was used to generate a cryo-ET capsid vertex map at bin 4 using the above processing scheme, then used to perform 3D classification (12 classes; 29 iterations). An initial 60 particles were selected based on the presence of a portal density and the absence of the penton major capsid protein, then used to generate a cryo-ET portal vertex map at bin 4 using the above processing scheme. A focused cylindrical mask was generated and used for further 3D classification of the VZV capsid vertices, yielding a total of 103 portal vertices. A final round of template-based 3D classification yielded a total of 109 portal vertices. The output was used for particle reconstruction (bin 2; 6.88 Å/pixel; box size 240 pixels; cropped box size 120 pixels; C5 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 30 Å and C5 symmetry, generating a 23.6 Å cryo-ET map of the VZV capsid portal vertex. To reconstruct the portal, particle reconstruction (bin 2; 6.88 Å/pixel; box size 90 pixels; cropped box size 30 pixels; C12 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 30 Å, C12 symmetry, and C1 symmetry relaxation were performed, generating a 22.9 Å cryo-ET map of the VZV portal.
To identify portal vertex-specific features, the 109 portal vertices were divided into CAI (n = 62) and C-capsids (n = 47) based on morphological features; either scaffold protein (CAI) or genomic dsDNA (C-capsids) inside the capsid cavity. Cryo-ET maps for the portal vertices from CAI (23.8 Å) and C-capsids (24.3 Å) were generated as described above; particle reconstruction (bin 2; 6.88 Å/pixel; box size 240 pixels; cropped box size 120 pixels; C5 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 30 Å and C5 symmetry. Cryo-ET maps for the entire capsid from CAI (32.6 Å) and C-capsids (33.6 Å) were generated by first adjusting the Z-height of the particles, then performing particle reconstruction (bin 2; 6.88 Å/pixel; box size 512 pixels; cropped box size 256 pixels; I3 symmetry), mask creation, reconstruction of subtomograms, and 3D refinement with an initial low-pass filter of 30 Å, C5 symmetry, and C1 symmetry relaxation.
The resolution of all Relion-generated cryo-ET maps was confirmed using post-processing with soft masks and assessed by Fourier Shell Correlation (FSC) curves with a cut-off at 0.143 from two independent half-sets of the pseudosubtomograms.
Statistics and reproducibility
Data were generated from 399 tilt series that were collected from cryo-FIB/SEM milled lamella across five, 3-day, cryo-ET data acquisition sessions. Figure 1b–d are representative images of 35 grids (1b), 265 lamella (1c), and 134 montages (1 d). Figure 2a, b are representative images of 73 tomograms containing capsids in the nucleus. Quantitative data (Fig. 5i) were analyzed by two-way ANOVA with Tukey’s multiple comparison test to determine statistical significance using Prism (GraphPad Software).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data generated and/or analyzed during the current study are available in the paper, appended as supplementary data, or available in public repositories. The original cryo-ET movie files, tilt series, and tomograms have been deposited in the Electron Microscopy Public Image Archive (EMPIAR) with accession code EMPIAR-12464. The atomic model used for the initial segmentation of the VZV capsid vertex (C5) and C3 cryo-ET maps was PDB 7BW6. The cryo-ET maps generated in this study have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-49591 for the vertex cryo-ET map (model PDB 9NO1), EMD-49886 for the 3-fold axis cryo-ET map. The EMDB accession codes for the cryo-ET maps of the VZV CAI and C-capsid portal processing are: whole capsid for CAI and C-capsids combined EMD-49465; capsid vertex for CAI and C-capsids combined EMD-49466; capsids with portal for CAI and C-capsids combined EMD-49467; portal vertex for CAI and C-capsids combined EMD-49468; portal for CAI and C-capsids EMD-49469; CAI-capsid whole capsid EMD-49470; CAI-capsid portal vertex EMD-49471; C-capsid whole capsid EMD-49472; C-capsid portal vertex EMD-49473. The source data underlying Figs. 5A, 5E, and 5I are provided in the associated Source Data file. Source data are provided with this paper.
References
Arvin, A. M. & Abendroth, A. Varicella–Zoster Virus. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen J. I.). 7 edn. 445–488 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Damania, B. & Cesarman, E. Kaposi's Sarcoma Herpesvirus. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J.I.). 7 edn. 513–572 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Gewurz, B. E., Longnecker, R. M. & Cohen, J. I. Epstein-Barr Virus. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J. I.). 7 edn. 324–388 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Goodrum, F., Britt, W. & Mocarski, E. S. Cytomegalovirus. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J. I.). 7 edn. 389–444 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Knipe, D. M., Heldwein, E.E., Mohr, I. J. & Odroski, C. N. Herpes Simplex Viruses: Mechanisms of Lytic and Latent Infection. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J. I.). 7 edn. 235–296 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Krug, L. T. & Pellett, P. E. The Family Herpesviridae: A Brief Introduction. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J. I.). 7 edn. 212–234 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Mori, Y., Zerr, D. M., Flamand, L. & Pellett, P.E. Human Herpesviruses 6A, 6B, and 7. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J. I.). 7 edn. 489--512 (Lippincott Williams & Wilkins, a Wolters Kluwer Business, 2022).
Whitley, R. J. & Johnston, C. Herpes Simplex Viruses: Pathogenesis and Clinical Insights. In: Fields Virology Volume 2: DNA Viruses (eds. Howley, P. M., Knipe, D. M., Damania, B. A., Cohen, J. I.). 7 edn. 297–323 (Lippincott Williams & Wilkins, a Wolters Kluwer Busines, 2022).
Gilden, D.H., Mahalingham, R., Deitch, S. & Randall C. J. Varicella Zoster Virus Neuropathogenesis and Latency. In: Alpha Herpesviruses: Molecular and Cellular Biology (ed, Sandri-Goldin, R. M.). 402 (Caister Academic, 2006).
Banovic, T. et al. Disseminated varicella infection caused by varicella vaccine strain in a child with low invariant natural killer T cells and diminished CD1d expression. J. Infect. Dis. 204, 1893–1901 (2011).
Gilden, D., Cohrs, R. J., Mahalingam, R. & Nagel, M. A. Varicella zoster virus vasculopathies: diverse clinical manifestations, laboratory features, pathogenesis, and treatment. Lancet Neurol. 8, 731–740 (2009).
Jean-Philippe, P. et al. Severe varicella caused by varicella-vaccine strain in a child with significant T-cell dysfunction. Pediatrics 120, e1345–e1349 (2007).
Johnson, R. W. Zoster-associated pain: what is known, who is at risk and how can it be managed? Herpes 14, 30–34 (2007).
Pahud, B. A., Glaser, C. A., Dekker, C. L., Arvin, A. M. & Schmid, D. S. Varicella zoster disease of the central nervous system: epidemiological, clinical, and laboratory features 10 years after the introduction of the varicella vaccine. J. Infect. Dis. 203, 316–323 (2011).
Schmader, K. Herpes zoster and postherpetic neuralgia in older adults. Clin. Geriatr. Med 23, 615–632 (2007).
Brunell, P. A., Geiser, C. F., Novelli, V., Lipton, S. & Narkewicz, S. Varicella-like illness caused by live varicella vaccine in children with acute lymphocytic leukemia. Pediatrics 79, 922–927 (1987).
Marin, M., Meissner, H. C. & Seward, J. F. Varicella prevention in the United States: a review of successes and challenges. Pediatrics 122, e744–e751 (2008).
Pickering, L. K. et al. Immunization programs for infants, children, adolescents, and adults: clinical practice guidelines by the Infectious Diseases Society of America. Clin. Infect. Dis. 49, 817–840 (2009).
Han, J. Y., Hanson, D. C. & Way, S. S. Herpes zoster and meningitis due to reactivation of varicella vaccine virus in an immunocompetent child. Pediatr. Infect. Dis. J. 30, 266–268 (2011).
Krause, P. R. & Klinman, D. M. Varicella vaccination: evidence for frequent reactivation of the vaccine strain in healthy children. Nat. Med. 6, 451–454 (2000).
Whitley, R. J. Therapy of herpes virus infections in children. Adv. Exp. Med Biol. 609, 216–232 (2008).
Whitley, R. J., Volpi, A., McKendrick, M., Wijck, A. & Oaklander, A. L. Management of herpes zoster and post-herpetic neuralgia now and in the future. J. Clin. Virol. 48, S20–S28 (2010).
Shiraki, K., Takemoto, M. & Daikoku, T. Emergence of varicella-zoster virus resistance to acyclovir: epidemiology, prevention, and treatment. Expert Rev. Anti Infect. Ther. 19, 1415–1425 (2021).
Topalis, D., Gillemot, S., Snoeck, R. & Andrei, G. Thymidine kinase and protein kinase in drug-resistant herpesviruses: heads of a Lernaean Hydra. Drug Resist Updat 37, 1–16 (2018).
Harfouche, M. et al. Estimated global and regional incidence and prevalence of herpes simplex virus infections and genital ulcer disease in 2020: mathematical modelling analyses. Sex Transm. Infect. 101, 214–223 (2024).
Chaiyakunapruk, N. et al. Estimated global and regional economic burden of genital herpes simplex virus infection among 15-49 year-olds in 2016. BMC Glob. Public Health 2, 42 (2024).
Davison, A. J. Overview of classification. In: Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis (eds. Arvin, A. et al.) 3-9 (Cambridge University Press 2007).
Baines, J. D. Herpes simplex virus capsid assembly and DNA packaging: a present and future antiviral drug target. Trends Microbiol. 19, 606–613 (2011).
Heming, J. D., Conway, J. F. & Homa, F. L. Herpesvirus capsid assembly and DNA packaging. Adv. Anat. Embryol. Cell Biol. 223, 119–142 (2017).
Gibson, W. & Roizman, B. Proteins specified by herpes simplex virus. 8. Characterization and composition of multiple capsid forms of subtypes 1 and 2. J. Virol. 10, 1044–1052 (1972).
Tandon, R., Mocarski, E. S. & Conway, J. F. The A, B, Cs of herpesvirus capsids. Viruses 7, 899–914 (2015).
Stevens, A., Cruz-Cosme, R., Armstrong, N., Tang, Q. & Zhou, Z. H. Structure-guided mutagenesis targeting interactions between pp150 tegument protein and small capsid protein identify five lethal and two live-attenuated HCMV mutants. Virology 596, 110115 (2024).
Dai, X. & Zhou, Z. H. Structure of the herpes simplex virus 1 capsid with associated tegument protein complexes. Science 360, eaao7283 (2018).
Gong, D. et al. DNA-packing portal and capsid-associated tegument complexes in the tumor herpesvirus KSHV. Cell 178, 1329–1343 e1312 (2019).
Li, Z., Pang, J., Dong, L. & Yu, X. Structural basis for genome packaging, retention, and ejection in human cytomegalovirus. Nat. Commun. 12, 4538 (2021).
Li, Z. et al. CryoEM structure of the tegumented capsid of Epstein-Barr virus. Cell Res. 30, 873–884 (2020).
Liu, W. et al. Structures of capsid and capsid-associated tegument complex inside the Epstein-Barr virus. Nat. Microbiol. 5, 1285–1298 (2020).
Liu, Y. T., Jih, J., Dai, X., Bi, G. Q. & Zhou, Z. H. Cryo-EM structures of herpes simplex virus type 1 portal vertex and packaged genome. Nature 570, 257–261 (2019).
Sun, J. et al. Cryo-EM structure of the varicella-zoster virus A-capsid. Nat. Commun. 11, 4795 (2020).
Wang, G. et al. Structures of pseudorabies virus capsids. Nat. Commun. 13, 1533 (2022).
Wang, W. et al. Near-atomic cryo-electron microscopy structures of varicella-zoster virus capsids. Nat. Microbiol. 5, 1542–1552 (2020).
Yu, X., Jih, J., Jiang, J. & Zhou, Z. H. Atomic structure of the human cytomegalovirus capsid with its securing tegument layer of pp150. Science 356, eaam6892 (2017).
Zhang, Y. et al. Atomic structure of the human herpesvirus 6B capsid and capsid-associated tegument complexes. Nat. Commun. 10, 5346 (2019).
Dai, X. et al. Structure and mutagenesis reveal essential capsid protein interactions for KSHV replication. Nature 553, 521–525 (2018).
Huet, A., Huffman, J. B., Conway, J. F. & Homa, F. L. Role of the herpes simplex virus CVSC proteins at the capsid portal vertex. J. Virol. 94, e01534-20 (2020).
Sae-Ueng, U. et al. Major capsid reinforcement by a minor protein in herpesviruses and phage. Nucleic Acids Res. 42, 9096–9107 (2014).
Wang, J. et al. Structure of the herpes simplex virus type 2 C-capsid with capsid-vertex-specific component. Nat. Commun. 9, 3668 (2018).
Li, Z. et al. Cryo-electron microscopy structures of capsids and in situ portals of DNA-devoid capsids of human cytomegalovirus. Nat. Commun. 14, 2025 (2023).
Machon, C. et al. Atomic structure of the Epstein-Barr virus portal. Nat. Commun. 10, 3891 (2019).
Schmid, M. F. et al. A tail-like assembly at the portal vertex in intact herpes simplex type-1 virions. PLoS Pathog. 8, e1002961 (2012).
Draganova, E. B., Valentin, J. & Heldwein E. E. The ins and outs of herpesviral capsids: divergent structures and assembly mechanisms across the three subfamilies. Viruses 13, 1913 (2021).
Diwaker, D. & Wilson, D. W. Microtubule-dependent trafficking of alphaherpesviruses in the nervous system: the ins and outs. Viruses 11, 1165 (2019).
Zaichick, S. V. et al. The herpesvirus VP1/2 protein is an effector of dynein-mediated capsid transport and neuroinvasion. Cell Host Microbe 13, 193–203 (2013).
Jih, J., Liu, Y. T., Liu, W. & Zhou, Z. H. The incredible bulk: Human cytomegalovirus tegument architectures uncovered by AI-empowered cryo-EM. Sci. Adv. 10, eadj1640 (2024).
Buch, M. H. C., Newcomb, W. W., Winkler, D. C., Steven, A. C. & Heymann J. B. Cryo-electron tomography of the herpesvirus procapsid reveals interactions of the portal with the scaffold and a shift on maturation. mBio 12, e03575-20 (2021).
Cardone, G. et al. Visualization of the herpes simplex virus portal in situ by cryo-electron tomography. Virology 361, 426–434 (2007).
Dai, W. et al. Unique structures in a tumor herpesvirus revealed by cryo-electron tomography and microscopy. J. Struct. Biol. 161, 428–438 (2008).
Deng, B., O’Connor, C. M., Kedes, D. H. & Zhou, Z. H. Direct visualization of the putative portal in the Kaposi’s sarcoma-associated herpesvirus capsid by cryoelectron tomography. J. Virol. 81, 3640–3644 (2007).
Deng, B., O’Connor, C. M., Kedes, D. H. & Zhou, Z. H. Cryo-electron tomography of Kaposi’s sarcoma-associated herpesvirus capsids reveals dynamic scaffolding structures essential to capsid assembly and maturation. J. Struct. Biol. 161, 419–427 (2008).
Grunewald, K. et al. Three-dimensional structure of herpes simplex virus from cryo-electron tomography. Science 302, 1396–1398 (2003).
Newcomb, W. W. et al. The primary enveloped virion of herpes simplex virus 1: its role in nuclear egress. mBio 8, e00825-17 (2017).
Si, Z. et al. Different functional states of fusion protein gB revealed on human cytomegalovirus by cryo electron tomography with Volta phase plate. PLoS Pathog. 14, e1007452 (2018).
Chang, J. T., Schmid, M. F., Rixon, F. J. & Chiu, W. Electron cryotomography reveals the portal in the herpesvirus capsid. J. Virol. 81, 2065–2068 (2007).
Vijayakrishnan, S., McElwee, M., Loney, C., Rixon, F. & Bhella, D. In situ structure of virus capsids within cell nuclei by correlative light and cryo-electron tomography. Sci. Rep. 10, 17596 (2020).
Bucks, M. A., O’Regan, K. J., Murphy, M. A., Wills, J. W. & Courtney, R. J. Herpes simplex virus type 1 tegument proteins VP1/2 and UL37 are associated with intranuclear capsids. Virology 361, 316–324 (2007).
Fan, W. H. et al. The large tegument protein pUL36 is essential for formation of the capsid vertex-specific component at the capsid-tegument interface of herpes simplex virus 1. J. Virol. 89, 1502–1511 (2015).
Schipke, J. et al. The C terminus of the large tegument protein pUL36 contains multiple capsid binding sites that function differently during assembly and cell entry of herpes simplex virus. J. Virol. 86, 3682–3700 (2012).
Cao, L. et al. Insights into varicella-zoster virus assembly from the B- and C-capsid at near-atomic resolution structures. hLife 2, 64–74 (2024).
Reimer, L. & Kohl, H. Transmission Electron Microscopy: Physics of Image Formation, 5th edn. (Springer, 2008).
Schaffer, M. et al. A cryo-FIB lift-out technique enables molecular-resolution cryo-ET within native Caenorhabditis elegans tissue. Nat. Methods 16, 757–762 (2019).
Oliver, S. L., Yang, E. & Arvin, A. M. Dysregulated glycoprotein B-mediated cell-cell fusion disrupts varicella-zoster virus and host gene transcription during infection. J. Virol. 91, e01613-16 (2017).
Oliver, S. L. et al. A glycoprotein B-neutralizing antibody structure at 2.8 A uncovers a critical domain for herpesvirus fusion initiation. Nat. Commun. 11, 4141 (2020).
Oliver, S. L. et al. The N-terminus of varicella-zoster virus glycoprotein B has a functional role in fusion. PLoS Pathog. 17, e1008961 (2021).
Zhou, M., Kamarshi, V., Arvin, A. M. & Oliver, S. L. Calcineurin phosphatase activity regulates Varicella-Zoster Virus induced cell-cell fusion. PLoS Pathog. 16, e1009022 (2020).
Baker, M. L., Jiang, W., Rixon, F. J. & Chiu, W. Common ancestry of herpesviruses and tailed DNA bacteriophages. J. Virol. 79, 14967–14970 (2005).
Homa, F. L. et al. Structure of the pseudorabies virus capsid: comparison with herpes simplex virus type 1 and differential binding of essential minor proteins. J. Mol. Biol. 425, 3415–3428 (2013).
DeRussy, B. M., Boland, M. T. & Tandon, R. Human cytomegalovirus pUL93 links nucleocapsid maturation and nuclear egress. J. Virol. 90, 7109–7117 (2016).
Evilevitch, A. & Sae-Ueng, U. Mechanical capsid maturation facilitates the resolution of conflicting requirements for herpesvirus assembly. J. Virol. 96, e0183121 (2022).
McInnes, L. & Healy, J. UMAP: uniform manifold approximation and projection for dimension reduction. https://arxiv.org/abs/1802.03426 (2018).
Chen, M. & Ludtke, S. J. Deep learning-based mixed-dimensional Gaussian mixture model for characterizing variability in cryo-EM. Nat. Methods 18, 930–936 (2021).
Zhong, E. D., Bepler, T., Berger, B. & Davis, J. H. CryoDRGN: reconstruction of heterogeneous cryo-EM structures using neural networks. Nat. Methods 18, 176–185 (2021).
Radermacher, M. Three-dimensional reconstruction of single particles from random and nonrandom tilt series. J. Electron. Microsc. Tech. 9, 359–394 (1988).
Newcomb, W. W., Homa, F. L., Thomsen, D. R., Ye, Z. & Brown, J. C. Cell-free assembly of the herpes simplex virus capsid. J. Virol. 68, 6059–6063 (1994).
Davison, M. D., Rixon, F. J. & Davison, A. J. Identification of genes encoding two capsid proteins (VP24 and VP26) of herpes simplex virus type 1. J. Gen. Virol. 73, 2709–2713 (1992).
Preston, V. G. et al. Efficient herpes simplex virus type 1 (HSV-1) capsid formation directed by the varicella-zoster virus scaffolding protein requires the carboxy-terminal sequences from the HSV-1 homologue. J. Gen. Virol. 78, 1633–1646 (1997).
Yuan, S. et al. Cryo-EM structure of a herpesvirus capsid at 3.1 A. Science 360, eaao7283 (2018).
Zhou, Z. H. et al. Assembly of VP26 in herpes simplex virus-1 inferred from structures of wild-type and recombinant capsids. Nat. Struct. Biol. 2, 1026–1030 (1995).
Cardone, G., Heymann, J. B., Cheng, N., Trus, B. L. & Steven, A. C. Procapsid assembly, maturation, nuclear exit: dynamic steps in the production of infectious herpesvirions. Adv. Exp. Med. Biol. 726, 423–439 (2012).
Zhang, Z. et al. Genome-wide mutagenesis reveals that ORF7 is a novel VZV skin-tropic factor. PLoS Pathog. 6, e1000971 (2010).
Chaudhuri, V., Sommer, M., Rajamani, J., Zerboni, L. & Arvin, A. M. Functions of Varicella-zoster virus ORF23 capsid protein in viral replication and the pathogenesis of skin infection. J. Virol. 82, 10231–10246 (2008).
Nicholson, P. et al. Localization of the herpes simplex virus type 1 major capsid protein VP5 to the cell nucleus requires the abundant scaffolding protein VP22a. J. Gen. Virol. 75, 1091–1099 (1994).
Newcomb, W. W. et al. Assembly of the herpes simplex virus capsid: characterization of intermediates observed during cell-free capsid formation. J. Mol. Biol. 263, 432–446 (1996).
Visalli, M. A., House, B. L., Selariu, A., Zhu, H. & Visalli, R. J. The varicella-zoster virus portal protein is essential for cleavage and packaging of viral DNA. J. Virol. 88, 7973–7986 (2014).
Visalli, R. J. & Howard, A. J. Non-axial view of the varicella-zoster virus portal protein reveals conserved crown, wing and clip architecture. Intervirology 57, 121–125 (2014).
Gao, M. et al. The protease of herpes simplex virus type 1 is essential for functional capsid formation and viral growth. J. Virol. 68, 3702–3712 (1994).
Newcomb, W. W. et al. The UL6 gene product forms the portal for entry of DNA into the herpes simplex virus capsid. J. Virol. 75, 10923–10932 (2001).
McNab, A. R. et al. The product of the herpes simplex virus type 1 UL25 gene is required for encapsidation but not for cleavage of replicated viral DNA. J. Virol. 72, 1060–1070 (1998).
Draganova, E. B., Zhang, J., Zhou, Z. H. & Heldwein, E. E. Structural basis for capsid recruitment and coat formation during HSV-1 nuclear egress. Elife 9, e56627 (2020).
Yang, K. & Baines, J. D. Selection of HSV capsids for envelopment involves interaction between capsid surface components pUL31, pUL17, and pUL25. Proc. Natl. Acad. Sci. USA 108, 14276–14281 (2011).
Mettenleiter, T. C., Muller, F., Granzow, H. & Klupp, B. G. The way out: what we know and do not know about herpesvirus nuclear egress. Cell Microbiol 15, 170–178 (2013).
Prazak, V. et al. Molecular plasticity of herpesvirus nuclear egress analysed in situ. Nat. Microbiol 9, 1842–1855 (2024).
Klupp, B. G., Granzow, H. & Mettenleiter, T. C. Nuclear envelope breakdown can substitute for primary envelopment-mediated nuclear egress of herpesviruses. J. Virol. 85, 8285–8292 (2011).
Dai, X. et al. The smallest capsid protein mediates binding of the essential tegument protein pp150 to stabilize DNA-containing capsids in human cytomegalovirus. PLoS Pathog. 9, e1003525 (2013).
Tischer, B. K. et al. A self-excisable infectious bacterial artificial chromosome clone of varicella-zoster virus allows analysis of the essential tegument protein encoded by ORF9. J. Virol. 81, 13200–13208 (2007).
Yang, E., Arvin, A. M. & Oliver, S. L. The cytoplasmic domain of varicella-zoster virus glycoprotein H regulates syncytia formation and skin pathogenesis. PLoS Pathog. 10, e1004173 (2014).
Wolff, G. et al. Mind the gap: Micro-expansion joints drastically decrease the bending of FIB-milled cryo-lamellae. J. Struct. Biol. 208, 107389 (2019).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Mastronarde, D. N. Dual-axis tomography: an approach with alignment methods that preserve resolution. J. Struct. Biol. 120, 343–352 (1997).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Phillips, J. C. et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 153, 044130 (2020).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38, 27–38 (1996).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr D. Struct. Biol. 74, 519–530 (2018).
Castano-Diez, D., Kudryashev, M., Arheit, M. & Stahlberg, H. Dynamo: a flexible, user-friendly development tool for subtomogram averaging of cryo-EM data in high-performance computing environments. J. Struct. Biol. 178, 139–151 (2012).
Acknowledgements
Cryo-FIB-SEM and cryo-ET were performed at the Stanford-SLAC Cryo-EM Center and the Stanford-SLAC Cryo-ET Specimen Preparation Center under the support of the National Institutes of Health Common Fund’s Transformative High Resolution Cryo-electron Microscopy Program (U24GM139166 and U24GM129541); and the NIH Shared Instrumentation Grant (S10OD021600). Research support was provided by NIH grants R01AI020459 (S.L.O.), R21AI159375 (S.L.O. and W.C.), R01GM150905 (M.C.), AI15937502 (W.C.), and R21MH125285 (M.C.). Molecular graphics and analyses performed with UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Contributions
S.L.O. conceptualization, methodology, software application, validation, formal analysis, investigation, resources, data curation, writing (original drafts and editing), visualization, supervision, project administration, funding acquisition. M.C. software application, validation, formal analysis, writing (review and editing). L.E. methodology, resources, writing (review and editing). C.W.H. and X.Z. methodology, writing (review and editing). M.F.S. validation, formal analysis, writing (review and editing). A.M.A. resources, writing (review and editing), supervision, funding acquisition. W.C. resources, writing (review and editing), supervision, funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Maud Dumoux, Xiangxi Wang, and Bernard Heymann for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Oliver, S.L., Chen, M., Engel, L. et al. Cryogenic electron tomography reveals herpesvirus capsid assembly intermediates inside the cell nucleus. Nat Commun 17, 3197 (2026). https://doi.org/10.1038/s41467-026-69811-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69811-4