Abstract
The Late Bronze Age (ca. 1300–800 BCE) of Central Europe is often characterised as a period of increasing mobility, socioeconomic transformation, environmental fluctuations, and expanding cultural networks. However, reconstructing the demographic aspects of these changes has been hindered by cremation being the dominant mortuary practice, limiting biomolecular approaches. Here, we integrate ancient DNA, oxygen and strontium isotope analyses, and osteoarchaeology to examine rare inhumation burials from Kuckenburg and Esperstedt in Central Germany (n = 36) and compare them to contemporaneous inhumations from the neighbouring regions of South Germany, Bohemia (Czechia) and Southwest/Central Poland (n = 33). Genome-wide data show genetic continuity with preceding Early Bronze Age populations, alongside gradual increases in Early European Farmer-related ancestry, albeit with regionally different timing and extent, reflecting a nuanced pattern of mobility and admixture. Oxygen and strontium isotope data from Central Germany indicate that most individuals match the local isotope signal, including those who were cremated or had a different diet, and with only a few isotopic outliers, suggesting that mobility was present but not extensive. Overall, our findings suggest that the diverse inhumation practices at Kuckenburg and Esperstedt were culturally motivated, reflecting local traditions and ongoing regional interconnectedness rather than the influx of new genetic groups or non-local individuals.
Similar content being viewed by others
Introduction
The Late Bronze Age (LBA, ca. 1300–800 BCE) in Central Europe was marked by profound cultural and social transformations, including long-distance trade and increasing interconnectedness, climatic and ideological shifts, and mobility1,2,3,4. A change in mortuary practices was also observed, with a transition from inhumation to cremation becoming the predominant practice. This transformation was so pronounced and widespread that it defined the period through its naming as the so-called Urnfield culture5 when cremated remains were placed in urns and buried in flat cemeteries. Based on recent archaeological findings, this practice is proposed to have originated in the middle Danube region, including areas of modern-day Lower Austria, Moravia, southwest Slovakia, and parts of Hungary, in the period between ca. 2000 and 1600 BCE, before spreading more widely throughout Europe and becoming consolidated in the classic Urnfield horizon after 1300 BCE6. This mortuary tradition significantly limits our ability to reconstruct the lifeways of people, such as their mobility patterns, diet, genetic ancestry and biological relatedness during this period, mainly due to the limitations of applying biomolecular methods to cremated remains7,8,9,10.
Ancient DNA (aDNA) has emerged as a critical tool for reconstructing human population histories and their relationship with cultural, economic, and linguistic changes in the past11,12,13. From an archaeogenetic perspective, the genomic profile of Bronze Age (BA) Central Europeans was largely established in the preceding 3rd millennium BCE, as the result of genetic mixing of Middle/Late Neolithic farming groups (carrying so-called Early European Farmer (EEF)-, and Western Hunter-Gatherer (WHG)-related ancestries) and pastoralist groups from Eastern Europe carrying steppe-related ancestry14,15,16,17,18,19, resulting in largely similar but regionally slightly different ancestry profiles. Specifically, for regions in today’s southern Germany, Switzerland, Austria and Hungary, a higher amount of EEF-related ancestry (and conversely a lower amount of steppe-related ancestry) has been documented relative to more northern regions such as Central Germany, Bohemia and Poland, already since the Bell Beaker and Early Bronze Age (EBA) periods16,17,18,20,21,22. In Bohemia, the shift from the EBA/Únětice to the Middle BA/Tumulus and LBA/Knovíz culture is also associated with an increase in EEF ancestry18, whereas such a shift was not observed in data from Poland associated with the Trzciniec culture21, whose roots lie largely in the forest zone of Europe.
Meanwhile, oxygen and strontium isotope analyses have been used to investigate mobility patterns23,24,25. For example, strontium evidence from Central Germany indicates that mobility was present in the EBA, with Únětice-period sites showing a mix of local and a smaller number of non-local individuals and indications of female exogamy26,27. While in southern Germany, high numbers of non-local individuals are reported, especially among female individuals, consistent with pronounced mobility and female exogamy20,28. The lack of such datasets for the LBA of much of Central Europe remains a temporal gap. Yet, despite the extent of cremation during the LBA, this practice was not universally adopted. For instance, the Unstrut group (ca. 1325-750 BCE), located between the Thuringian Forest and the Saale River in Central Germany and the focus of this study, continued to bury their dead for nearly 500 years, even as neighbouring groups, such as the Saalemündungs Group, transitioned to cremation29,30,31. Specifically, the mortuary practices of the Unstrut group are characterised by inhumation burials, where bodies are placed in a stretched supine position, oriented NW–SE, sometimes accompanied by stone packing, but also include a variety of other ways of burial, such as settlement or skull-only depositions31. Such practices align with broader patterns of LBA ‘special’ or ‘atypical’ burials (German: Sonderbestattungen) found across Central Europe, including Central Germany32, the Upper Tisza region33, Transylvania34, Moldova34, settlement burials of the Knovíz group in Czechia35, child burials within settlements at Şoimuş–Teleghi in southwestern Transylvania36, and settlement burials in Moravia, Austria, and South Germany37 among other examples38.
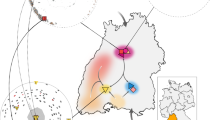
This study’s main focus lies on two archaeological sites in Saxony-Anhalt, Central Germany, associated with the Unstrut group: Esperstedt and Kuckenburg (Supplementary Note 1, Supplementary Fig. 1 and 12). Together, these sites form an extensive complex comprising a fortified settlement on a spur overlooking the Weida river (Kuckenburg)39,40,41,42,43, a second possibly fortified settlement (Esperstedt) at the opposite side of the Weida valley44, and a large bi-ritual graveyard45 situated just outside the ditch bordering the settlement to the west (Fig. 1a). For this study, we use a local, project-specific division of the LBA in two main phases: 1 (early) ca. 1300–1050 BCE and 2 (late) ca. 1050–800 BCE. In relative-chronological terms, phase 1 (early LBA) corresponds to the Central European phases Bz D and Ha A, whereas phase 2 (late LBA) corresponds to the Central European phase Ha B46. This division is based mainly on the settlement and dietary history of the Esperstedt/Kuckenburg micro-region in the LBA47, which show, at the current state of research, marked changes at the turn from the 2nd to the 1st millennium BCE.
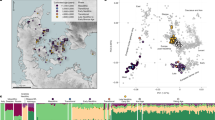
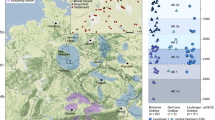
a Map of the archaeological sites of Kuckenburg and Esperstedt, Central Germany. b Map of Late Bronze Age Central European archaeological sites analysed in this study (1 Esperstedt, 2 Kuckenburg, 3 Leubingen, 4 Neckarsulm, 5 Trmice, 6 Holubice, 7 Libčice nad Vltavou, 8 Kněževes, 9 Praha-Ruzyně, 10 Slaný, 11 Łabędy Przyszówka, 12 Nowa Cerekwia, 13 Kornice; 1-4 Germany, 5-10 Czechia, 11-13 Poland). Maps (a) and (b) were generated using QGIS software (v3.12.2-București https://qgis.org/). The original base maps were extracted from (a) © GeoBasis-DE / LVerm-Geo ST, 2017, 167 (https://www.govdata.de/dl-de/by-2-0) and (b) the SRTM Data website (https://srtm.csi.cgiar.org/srtmdata/).
Mortuary practices at these two sites reflect the characteristic diversity of the northern Unstrut group, including burials in settlement pits and ditches, and the graveyard (Supplementary Figs. 1–28). While inhumation was the predominant practice, a few cremation burials have also been identified44. Additionally, the settlement of Kuckenburg includes skull-only depositions and multi-stage funerary rites39,40(Supplementary Figs. 13–28). Both the way that individuals were laid to rest and the bioarchaeological and biomolecular study of their remains can inform us about important aspects of their lived and embodied experience, such as their diet, origins, mobility patterns, social organisation, socioeconomic status, kinship practices (biological or social), degree of social integration within the community, and even societal perceptions of identity, ideas of personhood, and construction and negotiation of social memory48,49,50,51. Hence, by analysing the individuals buried at Kuckenburg and Esperstedt, we aim to address gaps in our understanding of Central European LBA communities on site-specific and regional scales, especially as forms of settlement burials are widespread across Central Germany, Bohemia, and Moravia and therefore provide a broader comparative framework. Given the general scarcity of inhumations during the Urnfield period, we generated, co-analysed and report new data from eleven additional LBA sites in today’s Bohemia/Czechia (n = 6), Southwest/Central Poland (n = 3), Thuringia/Germany (n = 1) and South Germany (n = 1) (Fig. 1b, Supplementary Notes 1.3 to 1.13, Supplementary Figs. 29-31). Leubingen in Thuringia is part of the main distribution area of the Unstrut group, whereas the sites in Bohemia and Southwest/Central Poland are attributed to the respective regional variants of the Urnfield cultural complex, such as the Knoviz and Štitary cultures in Bohemia or the Lusatian culture in Poland. Within the Urnfield complex, these represent a mosaic of regional groups differentiated by variations in burial practices, ceramic traditions, and material culture, but unified by parallel developmental trajectories and cultural traditions, including the predominance of cremation. The majority of the less common inhumations were found in settlement pits with very few or no grave goods. An exception is the LBA cemetery of Neckarsulm in South Germany, which has been assigned to the untermainisch-schwäbische Gruppe (German term), a local form of the Urnfield culture. This site is unique in that it represents a large necropolis of 32 inhumation graves, and in that it was interpreted archaeologically and anthropologically as the burial place of an all-male group. Previous interpretations have discussed whether these individuals represent retinues with a flat hierarchy or so-called warrior bands within a broadly segmentary social structure, while explicitly rejecting the concept of a hereditary warrior elite (Supplementary Note 1.4)1,52,53,54.
Taken together, adopting a primarily site-specific perspective (i.e., focusing on the sites of Esperstedt and Kuckenburg), we aim to characterise the genetic ancestry and mobility patterns of LBA individuals in Central Germany and assess their degree of continuity or change since the EBA in comparison with new and published data from contemporaneous groups from Central and South Germany, Czechia, and Poland (Fig. 1b). Specifically, we examine how factors such as age at death, genetic sex, socioeconomic status (using grave goods as a proxy), ancestry, mobility, cultural influences, and ideological shifts influenced mortuary choices in this region. Finally, we assess whether inhumation and cremation represented parallel burial traditions within the same community. We employ an interdisciplinary approach to address these questions by integrating aDNA, isotopic, and osteoarchaeological analyses. Through this approach, we aim to reach a more nuanced understanding of the lifeways of LBA communities and the motivation behind their choice of diverse mortuary practices.
Results
Genetic ancestry in the Late Bronze Age
We successfully retrieved aDNA from 75 LBA inhumations from Central Europe. All samples passed quality control, meeting criteria such as endogenous DNA content > 0.1% and characteristic aDNA damage patterns at the 5’ and 3’ ends of fragments55 in initial shotgun sequencing. After enrichment for 1.24 million single-nucleotide polymorphisms (SNPs)56, individuals (n = 69) with nuclear DNA contamination < 5%, mitochondrial DNA contamination < 10% and > 30,000 SNPs covered on the 1240k panel, were retained for downstream analysis. Genetic sex determination identified 32 female and 37 male individuals (see Methods section for details). For individuals with both genetic and osteoarchaeological sex assessments (n = 39), the results agreed in 85% of cases (Supplementary Data 1). We also called mitochondrial (mt) haplotypes (> 3000 mt reads) and Y-chromosome haplogroups for the individuals with sufficient coverage. We observed 51 different mt haplotypes in 64 individuals with enough mt reads for reliable mt haplogroup calling, broadly resembling the mt haplogroup distribution in Bronze Age Central Europe (ref. 17,57). The majority of male individuals carried either Y haplogroup I2a1 (n = 11) or R1b1 (n = 21), with two individuals carrying R1a (LBP001, NCE005), one individual (KUC023, Supplementary Note 2.4) carrying I1a3b2 and another (LEU020) I2a2a2 (Supplementary Data 1).
We conducted a Principal Component Analysis (PCA) on modern West Eurasian populations, on which we projected the newly analysed and published ancient individuals of interest (Fig. 2a), as well as ADMIXTURE analysis (Supplementary Note 2.1, Supplementary Fig. 32) and a series of f4-statistics (Supplementary Data 3), to genetically characterise the new LBA individuals and to assess their affinity to preceding EBA/Middle Bronze Age (MBA) populations (see Supplementary Data 2a and 2b for a list of populations used in this study), to nearby regional populations, and potential genetic variation linked to different mortuary practices (excluding cremated individuals). Specifically, the f4-statistics of the form f4(outgroup, X; Y, Z) use OldAfrica as an outgroup, while X represents Early European Farmer (EEF - Balkan_N, Turkey_N), Western European hunter-gatherers (WHG - WHGA and WHGB), or steppe-related ancestry (OldSteppe) (as defined in Patterson et al.18 and shown in Supplementary Data 2b, column Group ID_published), and Y and Z are the test populations of interest to assess changes in allele sharing in regards to the ancestry source X over time (see Methods for details). Positive f4-values indicate that population Z shares more alleles with X than population Y, while negative values indicate that population Y shares more alleles with X than population Z. Values near zero suggest no significant difference in allele sharing with X between Y and Z.
a Principal Component Analysis (PCA) of newly analysed individuals and relevant published ancient individuals, projected onto the genetic variation of 1252 present-day West Eurasian individuals from 77 populations (Supplementary Data 2a). Individuals identified as genetic outliers based on f4-statistics are specifically labelled and discussed in the text. Abbreviations: WHG Western Hunter–Gatherers; SHG Scandinavian Hunter–Gatherers; EHG Eastern Hunter–Gatherers; CHG Caucasus Hunter–Gatherers; EN Early Neolithic; LBK Linearbandkeramik; MN Middle Neolithic; LN Late Neolithic; PWC Pitted Ware Culture; EBA Early Bronze Age; MBA Middle Bronze Age; LBA Late Bronze Age. b Distal qpAdm model for all newly reported individuals, separated by temporal groups (early and late Late Bronze Age (LBA) and regions (Central Germany, South Germany, Bohemia (Czechia), and Southwest/Central Poland with Early European Farmer (EEF), Steppe, and Western Hunter-Gatherer (WHG) as proxies for source populations (Supplementary Data 4a). Error bars represent uncertainty (± one standard error) in ancestry estimates. Standard errors were computed with the default block jackknife approach. We use the same visual scheme as in panel (a) to indicate dietary information and geographic origin. A dashed outline indicates missing isotopic data. First-degree genetic relationships are connected with solid black lines, while more distant genetic relationships are shown with solid grey lines.
In PCA space, early LBA individuals from Central Germany plot within the genetic variation of published EBA individuals associated with the Únětice culture, suggesting regional genetic continuity27,56. This observation is supported by f4-statistics values close to zero (f4(OldAfrica, EEF; Germany_EBA_Unetice, Central_Germany_early_LBA) = 0.000345, Z = 2.659, n = 1,072,545 SNPs), which indicates no significant difference in allele sharing between the EBA individuals and the early LBA individuals from Central Germany (Supplementary Data 3, Test 1). These individuals also overlap with the only published LBA genome from Central Germany, from the archaeological site of Halberstadt56. However, most late LBA individuals from Central Germany are shifted slightly to a lower position on PC2, suggesting an increase in EEF-related ancestry and a reduction in steppe-related ancestry, which is confirmed by positive f4-statistics of the form f4(OldAfrica, EEF; Germany_EBA_Unetice, Central_Germany_late_LBA) = 0.000677, Z = 7.174, n = 1,106,172 SNPs (Supplementary Data 3, Test 2). A few of these individuals (e.g., KUC022) also overlap with individuals from the Tollense battlefield, in northern Germany58, as well as published and newly generated individuals from the LBA Knovíz culture in Czechia18 (Fig. 2a). KUC022 is a genetic outlier according to f4(OldAfrica, EEF; KUC022, Central_Germany_late_LBA) = -0.000829, Z = -3.078, n = 624,616 SNPs (Supplementary Data 3, Test 3). Due to the absence of aDNA data from the MBA in Central Germany, direct comparisons with the immediately preceding centuries are not possible.
For comparison, we also analysed 12 individuals from the unique LBA cemetery of Neckarsulm (NES) in South Germany (Supplementary Note 1.4)54,59. Genetic sex determination confirmed osteoarchaeological assignments, supporting the initial assessment that all 12 individuals were male59. A comparison of the South and Central Germany individuals in PCA space showed distinct genetic profiles: NES individuals occupy a lower position on PC2 (Fig. 2a), suggesting higher EEF-related ancestry and a difference in ancestry between the two regions, confirmed by f4(OldAfrica, EEF; Central_Germany_LBA, South_Germany_LBA) = 0.000887, Z = 6.638, n = 1,040,306 SNPs (Supplementary Data 3, Test 4). NES individuals also appear to carry more EEF-related ancestry than the local EBA populations in South Germany16,20, which is confirmed by positive f4-statistics of the form f4(OldAfrica, EEF; Germany_Lech_EBA, South_Germany_LBA) = 0.000544, Z = 3.502, n = 1,012,264 SNPs (Supplementary Data 3, Test 5). However, they overlap with MBA populations from the Lech Valley20, suggesting genetic continuity in South Germany since at least the MBA (f4(OldAfrica, EEF; Germany_Lech_MBA, South_Germany_LBA) = 0.000378, Z = 1.015, n = 179,223 SNPs) (Supplementary Data 3, Test 6). NES021, also a strontium outlier59, plots even lower on PC2, which indicates this individual may be a possible genetic outlier, which is confirmed by a negative f4-statistic (f4(OldAfrica, EEF; NES021, NES_group) = -0.001289, Z = -4.348, n = 738,215 SNPs) (Fig. 2a, and Supplementary Data 3, Test 7). The remaining strontium outliers identified by Wahl and Price59 (NES004, NES007, NES017, and NES020) do not differ in ancestry from the other NES individuals.
To broaden our regional and temporal comparison, we incorporated genomic data from 15 newly generated, as well as published, LBA Knovíz individuals from Bohemia, Czechia18 along with data from five LBA/Iron Age (IA) individuals from SW/Central Poland. On the PCA plot, the newly reported Knovíz individuals fall on a lower position on PC2 than EBA individuals from Czechia17, overlapping with published MBA and LBA genomes from the region18, and in between the LBA individuals from Central and South Germany, suggesting higher EEF-related ancestry than the EBA, which is confirmed by positive f4-statistics of the form f4(OldAfrica, EEF; Czech_EBA_Unetice, Knoviz_LBA_new) = 0.001726, Z = 16.247, n = 1,086,285 SNPs (Supplementary Data 3, Test 8). However, individuals SNY001 and LNV001 fall higher on PC2, overlapping with the earlier Czechia EBA individuals and late LBA individuals from Central Germany, making them potential genetic outliers, which is confirmed by positive f4-statistics (f4(OldAfrica, EEF; SNY001, Knoviz_LBA_new) = 0.001418, Z = 5.147, n = 889,075 SNPs) and (f4(OldAfrica, EEF; LNV001, Knoviz_LBA_new) = 0.000933, Z = 3.202, n = 883,852 SNPs) (Fig. 2a, and Supplementary Data 3, Test 9 and 10). The new LBA and IA individuals from SW/Central Poland demonstrate higher EEF- related ancestry compared to the EBA/MBA21 as showed by positive f4-statistics of the form f4(OldAfrica, EEF; Poland_MBA, SW/Central_Poland_LBA) = 0.001398, Z = 8.307, n = 948,221 SNPs (Supplementary Data 3, Test 11) and f4(OldAfrica, EEF; Poland_MBA, SW/Central_Poland_IA) = 0.0011, Z = 3.585, n = 398,011 SNPs (Supplementary Data 3, Test 12). The SW/Central_Poland_LBA individuals also seem to overlap with late LBA individuals from Central Germany. However the former carry higher WHG-related ancestry as shown by positive f4-statistics of the form f4(OldAfrica, WHGA; central_Germany_late_LBA, SW/Central_Poland_LBA) = 0.000886, Z = 3.952, n = 939,086 SNPs (Supplementary Data 3, Test 13) and f4 (OldAfrica, WHGB; central_Germany_late_LBA, SW/Central_Poland_LBA) = 0.000888, Z = 4.21, n = 953,919 SNPs (Supplementary Data 3, Test 13). Individual KOR014 falls lower on PC1, suggesting a higher proportion of WHG ancestry. This however is not confirmed by f4-statistics of the form f4(OldAfrica, WHG; KOR014, SW/Central_Poland_LBA) = -0.000774, Z = -0.811, n = 37,624 SNPs (Fig. 2a, and Supplementary Data 3, Test 14). Together, the observed patterns attest to similar shifts of genetic ancestry in all regions, characterised by increasing proportions of EEF-related ancestry between the Early and the Late Bronze Age, although the timing, extent, and mechanisms of this shift differ between regions (also see Fig. 3).
a Time series plot showing the EEF-related ancestry changes in Central Europe from the Early Bronze Age to the Iron Age calculated using qpAdm (Supplementary Data 4a). Different regions are indicated by colour and shape, individual points represent single individuals, and outliers are marked in black. Error bars represent uncertainty (± one standard error) in ancestry estimates. Standard errors were computed with the default block jackknife approach. The x-axis shows the median BCE date for each individual (Supplementary Data 4a). The number of individuals (n) per region and time period is displayed in panel b. Abbreviations: EEF, Early European Farmer–related ancestry; EBA, Early Bronze Age; MBA, Middle Bronze Age; LBA, Late Bronze Age; IA, Iron Age. b Box plots showing EEF-related ancestry changes over time in South Germany, Bohemia/Czechia, Central Germany, and Southwest/Central Poland. Each data point represents one individual. Sample sizes (n) for each period are indicated below the x-axis. The height of the boxes indicates the interquartile range (75th- 25th percentiles). The centre line indicates the median. Whiskers extend to the most extreme value within 1.5× the interquartile range. Individual data points are overlaid for periods with n < 10. Points beyond the whiskers indicate outliers and are coloured red. Statistical differences between periods were assessed using Kruskal–Wallis tests, restricted to periods with n ≥ 3 individuals, followed by two-sided pairwise Wilcoxon rank-sum tests with Bonferroni correction where applicable. Significant differences between periods are indicated by asterisks (* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001; ns = non-significant). Exact adjusted P values and detailed statistical test results are provided in Supplementary Data 12. The ancestry proportions used in both panels are the qpAdm distal estimates reported in Supplementary Data 4a. The millet icon was created by Roman from Noun Project (CC BY 3.0), https://thenounproject.com/browse/icons/term/millet/ and illustrates the attested millet consumption during the early LBA in Central Germany.
We then used qpAdm60,61,62 to model ancestry proportions by testing a set of distal source populations with reference to defined outgroup populations, ensuring comparability with previous studies18,63. All individuals were successfully modelled using qpAdm with their ancestry deriving primarily from EEF (Balkan Neolithic), Steppe populations (OldSteppe), and WHG18(Fig. 2b). We observe that late LBA individuals from Central Germany carry a slightly higher proportion of EEF-related ancestry (36.6 ± 2.9 %) than those from the early LBA in Central Germany (33.2 ± 2.7 %), followed by LBA individuals from Bohemia (43.7 ± 4.5 %) and LBA individuals from South Germany carrying the highest proportion of EEF-related ancestry (53.5 ± 5.3 %) (Fig. 2b, Supplementary Data 4a). Of all individuals analysed, genetic outlier NES021 carries the highest proportion of EEF-related ancestry with 67.2 ± 1.7 %. The four LBA individuals from SW/Central Poland also show a lower proportion of EEF-related ancestry (37.4 ± 2.9 %) similar to those from Central Germany. However, it remains open whether these individuals are fully representative of Polish LBA groups given the small sample size.
Beyond these regional differences, we also track the broader temporal trends in regions of today’s Central and South Germany, Bohemia/Czechia and SW/Central Poland, and observe a general increase in EEF-related ancestry proportions in all regions over time (Fig. 3a). We then tested whether the changes in EEF-related ancestry between the temporal cohorts of each region were statistically significant using a non-parametric Kruskal-Wallis test (Fig.3b, and Supplementary Data 12, Methods). Here, statistically significant increases occurred from the EBA to the early LBA in South Germany as well as from the EBA to the MBA/early LBA in Bohemia/Czechia. In SW/Central Poland, a potential increase, from the MBA to the early LBA is suggested, however the limited number of individuals does not allow statistical testing. Also in Central Germany, where no data from the MBA period is reported yet, EEF-related ancestry increased potentially later, from the early to late LBA (Fig. 3b).
To better understand the sources contributing to the observed increase in EEF-related ancestry during the LBA, we then used qpAdm modelling with geographically and chronologically more proximal reference populations (Fig.4) and kept the set of outgroups used for the distal models18,63. Early LBA individuals from Central Germany can be modelled with Germany EBA Únětice as the sole source, suggesting regional continuity since the EBA, as a parsimonious explanation (Supplementary Data 4b). By contrast, the ancestry of most of the late LBA individuals from Central Germany cannot be modelled as the previous EBA populations from the region alone, and are best modelled with two ancestry sources, that is with Germany EBA Únětice as the local source, and Germany_Lech_MBA, Czech_MBA_Tumulus, or Germany_SouthernGermany_Singen_EBA as the second geographically and chronologically closest non-local proxies (Supplementary Data 4c, and Figs. 4a and 4b). This pattern suggests a mixture of local continuity and gene flow from southern and/or eastern regions, consistent with increasing connectivity within the geographic distribution/extent of the Urnfield culture4,6,18,63,64, but without being able to be more specific.
Each bar represents an individual modelled as a two-way admixture using Germany Early Bronze Age (EBA) Únětice as the local ancestry source and a geographically and chronologically proximate non-local source. Ancestry proportions are shown with one standard error, calculated using the default block jackknife approach. Lighter-coloured bars indicate rejected models (p ≤ 0.05), while supported models are defined as those with p > 0.05. Detailed qpAdm modelling results are provided in Supplementary Data 4c. a Models using Germany Lech Middle Bronze Age (MBA) as the secondary source. b Models using Czech MBA Tumulus as the secondary source.
Neckarsulm individuals are best modelled with Germany_Lech_MBA as a single source proxy, suggesting regional genetic continuity with southern parts of Germany following the MBA, as a parsimonious explanation (Supplementary Data 4d). Individual NES021 is best modelled as a mixture of Germany_Lech_EBA/MBA or Germany_SouthernGermany_Singen_EBA as the local source and a population with a high level of EEF-related ancestry, such as Switzerland_EBA or Italy_EBA/EMBA as a non-local source (Supplementary Data 4d). We used the rotation approach to compare the competing models for the non-local source, by reciprocally adding a source from a model to outgroups. Both models still uphold the model fit (Supplementary Data 4d). In Bohemia, most LBA individuals are best modelled as deriving their ancestry from the preceding MBA Tumulus population, indicating genetic continuity since the MBA in the area, consistent with the overall trend observed above. However, two individuals, LNV001 and SNY001 show less EEF-related ancestry than the rest and indeed are best modelled as a mixture of EBA Únětice from Czechia and Germany and Germany_Lech_MBA (Supplementary Data 4e), confirming that they are genetic outliers, as also indicated by the f4-statistics above. Lastly, the LBA individuals from SW/Central Poland are best modelled as deriving ancestry from individuals associated with the preceding Poland MBA Trzciniec culture, together with Germany_Lech_MBA (Supplementary Data 4f).
To further investigate the geographic affinities of these genetic and isotopic outliers, we applied mobest65, a method that estimates where genetically most similar ancestry is found by comparing each individual’s genome to published ancient genomes from across Eurasia. NES021 shows highest genetic similarity to individuals from Switzerland and northern Italy, while KUC022 shows highest genetic similarity to individuals from southern Germany, possibly extending toward the Carpathian Basin (Fig. 5a). NES007 shows highest genetic similarity to southeastern European populations, though not identified as a genetic outlier from f4-statistics of the form f4(OldAfrica, X; NES007, NES_group) (Supplementary Data 3, Test 15). For NES007, NES017, and NES020, the burial location coincides with the region of highest genetic similarity (Fig. 5b). LNV001 and SNY001 from Bohemia show the highest genetic similarity to individuals from Central Germany.
a Mobest similarity maps for genetic outlier individuals from Kuckenburg (Central Germany), Neckarsulm (South Germany), Slaný (Bohemia), and Libčice nad Vltavou (Bohemia) dated to the Late Bronze Age (LBA). Each panel shows the spatial distribution of genetic similarity to ancient genomes from the same period. The orange triangle marks the point of highest similarity, while the black circle indicates the burial location. b mobest similarity maps for strontium outlier individuals from Neckarsulm (South Germany). Data visualisation in mobest is implemented in R using the ggplot2 package167. The maps use the European Terrestrial Reference System 1989 (ETRS89) Lambert Azimuthal Equal-Area (LAEA) projection (EPSG:3035). The spatial vector data was taken from the Natural Earth project (https://www.naturalearthdata.com).
Biological relatedness and social organisation in the Late Bronze Age
We also investigated possible biological relationships among individuals buried at the sites to see if these played a role in the choice of burial practices and if they could inform us about the community’s social organisation by applying methods such as READv266, BREADR67, KIN68, and ancIBD (identity-by-descent) analysis69. We detected no close biological relationships (up to the 3rd degree) among the individuals at the newly reported sites, even among single, double, or multiple burials, except for a mother (KUC004) and her two daughters (KUC005, KUC007) from a triple burial at Kuckenburg (Fig.2b, and Supplementary Data 5a, 5b, 5c, Supplementary Fig. 33). The analysis of IBD identified four more distant connections (> 4th degree, with at least two IBD fragments > 12 cM and at least one > 16 cM) among individuals from Kuckenburg, and also between individuals from Kuckenburg and Esperstedt, confirming a proposed connection41 between the closely located sites (< 2 km) (Supplementary Data 6).
Furthermore, we inferred runs of homozygosity (ROH) using hapROH70 to investigate patterns of parental background relatedness and effective population size. Most individuals show low levels of ROH, consistent with outbreeding and a larger effective population size (Supplementary Data 7, Supplementary Fig. 34). However, the female infant KUC019 demonstrates exceptionally high ROH (sum_roh > 4 = 367.08 cM), suggesting close parental relatedness, likely from a consanguineous union (e.g., first cousins). This has so far only been observed in very few aDNA studies63,71,72, with one example being the two individuals (MBG004 and APG003) in Iron Age southern Germany63. KUC027, an adult female individual, and KUC010, a young male individual, also display elevated ROH levels, though to a more moderate extent (sum_roh > 4 = 83.72 cM and 72.68 cM, respectively). In addition, KNE028, a young female from the Bohemian dataset, shows a similar moderate ROH level (sum_roh > 4 = 83.89 cM) (Supplementary Data 7, Supplementary Fig. 34).
Isotopic insights into mobility in the Late Bronze Age
To assess life history mobility patterns across the early and late LBA, and to compare these patterns with those of the preceding EBA, we measured 87Sr/86Sr ratios for 29 inhumations from the early and late LBA at Kuckenburg and Esperstedt, of which 28 are also genetically typed, as well as 21 cremated LBA individuals from Kuckenburg (n = 1) and Esperstedt (n = 20) for which no genomic data is available (Supplementary Data 8). In addition, we measured δ18O values for 24 of these individuals and six faunal specimens. To establish a local reference, we generated an 87Sr/86Sr baseline from LBA faunal remains from both sites (n = 18), defining a local 2σ range of 0.7090 to 0.7115 (Fig. 6a, and Supplementary Data 8, Supplementary Fig. 35, Methods). We additionally provide a modelled bioavailable 87Sr/86Sr isoscape for Central Europe (Supplementary Text 3.2, and Supplementary Fig. 36a, b), adapted from Bataille et al.73, to place these values within a broader regional context.
a Strontium isotope ratios (87Sr/86Sr) of human and faunal samples from Central Germany. Human individuals are grouped by period. The first three panels represent inhumations, while the fourth represents cremations. The local baseline was calculated from Late Bronze Age (LBA) faunal remains, including low-mobility taxa (e.g., snails, rodents). The solid black line marks the mean baseline, the dotted line the median, and the dashed lines the 95% confidence interval. The shaded area represents the estimated local range. Error bars represent internal analytical uncertainty (±2 standard deviations) of the 87Sr/86Sr ratios. The number of 87Sr/86Sr values (n) per period used in the analyses is displayed in panel b. The abbreviations M1, M2, M3 refer to first, second and third permanent molars, respectively. b Box plots of 87Sr/86Sr across periods in Central Germany. Each data point represents one 87Sr/86Sr value. Sample sizes (n) for each period are indicated below the x-axis. The height of the boxes indicates the interquartile range (75th–25th percentiles). The centre line indicates the median. Whiskers extend to the most extreme values within 1.5× the interquartile range. Points beyond the whiskers indicate outliers. The Kruskal-Wallis test (p = 0.074) suggests no significant differences between groups. c Scatterplot of 87Sr/86Sr vs. oxygen (δ18O) isotope values for early and late LBA individuals from Central Germany. Coloured region indicates local 87Sr/86Sr isotope ranges. 87Sr/86Sr outliers (ESP040 and KUC028) are labelled, and individuals whose sampled tooth was a first permanent molar (M1) are marked accordingly. d Scatterplot of strontium isotope ratios (87Sr/86Sr) versus carbon isotope ratios (δ13C, collagen)47 in Central Germany, showing dietary patterns47. Values of δ13C > −18 ‰ indicate consumption of C4 plants, in this case, broomcorn millet.
Most early LBA individuals fall within the local 87Sr/86Sr range (0.7107 to 0.7110, mean = 0.7110), consistent with values reported for EBA Unětice populations for the area26. We identified two isotopic outliers: ESP045 (87Sr/86Sr = 0.7095) and ESP040 (87Sr/86Sr = 0.7116) (Fig. 6a, and Supplementary Data 8). Most late LBA individuals also fall within the local range (0.7093 to 0.7110, mean = 0.7100), but five individuals demonstrate non-local 87Sr/86Sr isotopic signatures. Specifically, KUC010 (87Sr/86Sr = 0.7089) and KUC028 (87Sr/86Sr = 0.7088) likely originate from areas with calcareous bedrock or alluvial plains26. KUC017 and ESP036 (87Sr/86Sr = 0.7095) also suggest a non-local origin. Notably, KUC023, who carries the distinct Y haplogroup I1a3b2 (Supplementary Note 2.4), has an elevated Sr ratio (87Sr/86Sr = 0.7115) compared to the rest of the individuals and falls on the upper edge of the local baseline (Fig. 6a, and Supplementary Data 8).
We also tested for differences in strontium values between EBA, early LBA, and late LBA individuals. While differences were not statistically significant (Kruskal–Wallis test, χ²(3) = 6.95, p = 0.074), we observe a trend toward lower strontium values from the early to late LBA (Fig. 6b). The δ18O values for LBA individuals range from -7.3‰ to -4.9‰, with no significant differences between early and late LBA (Wilcoxon Rank-Sum Exact Test: W = 63.5, p = 0.524) (Supplementary Data 12). We applied a non-parametric 1.5×IQR approach to the human δ18O dataset to identify potential non-local individuals. In this test, we excluded first molars (M1), as their values may be affected by breastfeeding25. These individuals are nonetheless included in Fig. 6c. The lower and upper cut-off thresholds are –8.2‰ and –3.4‰ respectively (Q1 = –6.4‰, Q3 = –5.2‰, IQR = 1.2‰). We do not detect any δ18O outliers, as none of the values fall outside these limits (Supplementary Data 12, Supplementary Note 3.3). Additionally, none of the 87Sr/86Sr outliers are also δ18O outliers, suggesting that these individuals did not originate from distant climatic zones but rather from nearby geologically distinct areas (Fig. 6c). When comparing 87Sr/86Sr values by dietary patterns (δ13C from collagen47), we observe a significant difference between individuals who consumed millet (δ13C > –18‰) and those who did not (t-test, p = 0.0043). Following previous isotope studies in the region (ref. 74,75), values higher than –18‰ indicate a mixed C3/C4 diet and moderate millet intake, while values higher than –12‰ correspond to predominantly C4-based diets75,76. Individuals with millet consumption demonstrated higher 87Sr/86Sr values (Fig. 6d, and Supplementary Data 12).
Among the 21 cremated individuals, most fall within the local 87Sr/86Sr range (0.7097–0.7110) (Supplementary Data 8). However, we identified one outlier: EOR017 (87Sr/86Sr = 0.7090), radiocarbon dated to the late LBA (Supplementary Data 10). We should note here that calcined bone reflects a different period of life than teeth77, so these signals may capture mobility later in life rather than childhood origins78. Finally, we detected no significant difference in 87Sr/86Sr ratios between inhumations and cremations (Supplementary Fig. 37).
According to our results, most LBA individuals show a predominantly local strontium signal, with only a few exceptions. Interestingly, we observed a significant difference in 87Sr/86Sr values between individuals who consumed millet and those who did not. Millet consumers display higher 87Sr/86Sr values, which may suggest that millet was grown in areas with different underlying geology, whereas C3 plants would have been ubiquitous across the local landscape. Since most of the available individuals from the early LBA were millet consumers, it was not possible to test for differences in genetic profiles between millet and non-millet consumers during this period.
Mortuary practices and paleopathology in LBA Central Germany
Most individuals in this study were inhumed, in contrast to the increasing trend toward cremation during the Central European LBA. Specifically, the 36 LBA individuals from Central Germany were placed in both cemeteries and settlement pits, representing both ‘typical’ and ‘atypical’ burials, while the 12 individuals from South Germany (Neckarsulm) were exclusively male and buried in a cemetery54,59.
At Kuckenburg, we analysed 27 individuals (26 inhumations, 1 cremation), with 26 dating to the late LBA and one to the early LBA. Most individuals were buried in settlement pits, including double and triple burials, skull-only depositions, and individuals with missing vertebrae or added isolated bones40 (Supplementary Data 9). A complex mortuary feature, feature 39/2018, contained seven skull-only depositions (six from female individuals and one from a juvenile male individual), one human long bone fragment, one bovine mandible, and seven ceramic vessels, two of which contained cremated remains (Fig. 7). Radiocarbon dating places the inhumations between 978–766 cal BCE, while the cremation (KUC029) seems to be older (1049–904 cal BCE). 87Sr/86Sr analysis indicates that most individuals were local, except for KUC023 (juvenile male, 87Sr/86Sr = 0.7114) and KUC028 (young adult female, 87Sr/86Sr = 0.7088). At least two individuals showed signs of perimortem trauma. KUC025 (female individual, age at death: 40–50 years) suffered severe cranial trauma and enamel chipping, and KUC023 (male individual, age at death: juvenile) demonstrated cranial trauma and signs of possible physiological stress (cribra orbitalia, Grade 2). No close biological relationships were identified among individuals in this feature, suggesting that cultural, ritual, or opportunistic criteria, rather than biological relatedness, determined their co-burial. The integration of inhumations and cremations within the same feature suggests a hybrid, multi-stage funerary tradition.
This feature contains eight individuals, animal bones, and two cremations in vessels.
At Esperstedt, we analysed 10 inhumed individuals—eight from the early LBA graveyard phase and two (ESP036, ESP041) from late LBA settlement contexts. In contrast to Kuckenburg, skull-only burials are absent, and circular ditch graves are common. Among these, five are adults and five are non-adults, all with few grave goods. Additionally, we analysed 20 cremated individuals, most of which are adults, though age at death and anthropological sex estimations were limited due to fragmentation. We radiocarbon dated five cremations with four of them ranging from the early to late LBA and one dating to the IA (Supplementary Data 10). At Neckarsulm, all 12 individuals are male and inhumed in single, double, or triple burials. Preservation limited genetic analysis to two double burials and a few isolated individuals. Radiocarbon dating of two individuals confirms their contextual placement in the early LBA (Supplementary Data 1).
We formally tested associations between burial type (skull-only vs. whole-body), LBA phase, age at death, and genetic sex (with at least n = 3 individuals per category) (Supplementary Data 11 and 12). No graveyard burials have been identified so far from the late LBA. Within the late LBA, skull-only burials did not differ from whole-body burials in age at death (Fisher’s exact test, adjusted p = 0.2412), EEF-related ancestry (t-test, adjusted p = 0.2832), 87Sr/86Sr values (t-test, adjusted p = 0.5986), genetic sex (Fisher’s exact test, adjusted p = 0.2832), or δ13C dietary values (t-test, adjusted p = 0.822 for collagen; Wilcoxon test, adjusted p = 0.8222 for enamel). Based on these similarities, skull-only and other late LBA burials were grouped for subsequent analyses. To assess patterns in grave goods distribution, we tested whether their presence was associated with age at death category (adults vs. non-adults), genetic sex, and LBA phase (early vs. late). Only six individuals were buried with grave goods. Due to small sample sizes, we applied Fisher’s exact test, which revealed no significant differences across categories (age: adjusted p = 1; sex: adjusted p = 1; LBA phase: adjusted p = 1). This suggests that grave goods were not preferentially included based on age at death, genetic sex, or period. However, due to the small sample size, this result should be treated with caution. Non-adults were significantly more common in settlement pit burials compared to graveyard burials (Fisher’s exact test, adjusted p = 0.0234), indicating potential differences in age-related burial practices. However, it is important to note that all settlement burials excavated so far date to the late LBA, whereas all the dated graveyard burials are from the early LBA, which may confound this association. Finally, we found no significant differences in genetic sex distribution between settlement and graveyard burials (Fisher’s exact test, adjusted p = 0.434), suggesting that burial type was not strongly associated with genetic sex (see Supplementary Data 12 for raw and FDR adjusted p values).
These inhumation burials also provided us with a rare opportunity to assess the palaeopathological lesions/conditions of the people in LBA Central Germany. Among adults, degenerative diseases were most common in older individuals. Trauma was rare, with occasional long bone or rib fractures and isolated cranial impaction fractures, all of which occurred antemortem. Some individuals showed osteolytic and osteoplastic bone lesions, possibly related to nonspecific conditions or tumours. Among non-adults, non-specific markers of physiological stress such as cribra orbitalia, porotic hyperostosis, and enamel hypoplasia were most frequently observed. Non-adults also have a higher incidence of spinal tuberculosis (e.g., individual KUC012, Supplementary Data 13) and non-specific inflammatory reactions compared to adults. In addition, for the burials where dental status could be assessed, the individuals generally show good dental health. Calculus and mild caries are commonly observed, while isolated cases of severe caries also occur, ranging from pulpa aperta to pathological changes in the jawbone (e.g., individual KUC004, Supplementary Data 13). However, such severe cases are rare within the examined population. In most cases, the cause of death could not be determined. While systemic diseases or infections may have been fatal depending on severity, no definitive cases of lethal trauma (e.g., stab wounds, decapitation) have been identified in the Kuckenburg or settlement burials. However, two individuals (KUC023 and KUC025) showed perimortem cranial trauma (Supplementary Data 13).
Finally, we screened all individuals in this study (except cremated ones) for human pathogen DNA to detect diseases not identifiable through osteological analysis. We detect traces of DNA from species associated with the human oral microbiome, such as Streptococcus mutans and Parvimonas micra, which were identified as potentially authentic and ancient by the screening pipeline HOPS79 (Supplementary Data 14). These pathobionts are commonly involved in the development of dental caries80 and periodontitis81, respectively, often favoured by a carbohydrate rich diet. Their detection is in line with the palaeopathological observations. In individual ESP043, we further detect DNA of Yersinia enterocolitica, a zoonotic pathogen causing mostly acute gastrointestinal disease in humans (Supplementary Fig. 38)82. However, further analysis such as in-solution capture and phylogenetic placement is necessary for a confident confirmation of this result, as the current number of assigned reads to this species is too low (32 reads).
Overall, evidence of childhood stress, degenerative joint conditions, and occasional trauma points to physically demanding lives. Nevertheless, most individuals appear to have been in generally good condition. The rare occurrence of trauma, which can be indicative of interpersonal violence or warfare, suggests that such conflict was not a dominant feature in the lives of these communities, at least for the inhumed individuals receiving formal burial.
Discussion
The genetic analyses in this study provide insights into the population history of Late Bronze Age (LBA) communities in Central Europe, contributing to the limited genomic record for this dynamic period. The individuals from Kuckenburg and Esperstedt, in today’s Central Germany, show relative regional genetic continuity with preceding EBA Únětice populations. However, they also demonstrate a gradual increase in EEF-related ancestry over the course of the Bronze Age, likely reflecting contact with people carrying higher EEF-like ancestry, as observed in preceding and contemporaneous populations from southern and eastern Europe. Even as the Unstrut group developed and maintained connections to neighbouring, predominantly southern LBA groups, as evidenced also by the continuous increase in EEF-like ancestry, they continued to practice inhumation burial, a rite with a deep history in this part of Central Germany. Specifically, there are indications for an increased connectedness at the turn of the 2nd to the 1st millennium BCE. This increase refers only to the Middle-Elbe-Saale (MES) communities in the southern part of Saxony-Anhalt and the Thuringian Basin, for which especially the increase of cremations points to cultural changes (e.g., a southward shift of the border region between the Saalemündungs group and the northern Unstrut-Group / Helmsdorfer group)31,83,84.
A distinct north-south ancestry dichotomy is also evident: LBA individuals from Central Germany consistently show lower EEF-related ancestry compared to contemporaneous individuals from South Germany. This regional genetic differentiation is deeply rooted in the Neolithic-Bronze Age transition of the mid-3rd millennium BCE16,20,61,63 and persisted through the entire Bronze Age, indicating a lasting trend of several centuries, possibly upheld by geographic distance and/or cultural differences and/or particular cultural practices20 that, for instance, also distinguished the EBA Únětice sphere from those of the southern Straubing85 and Unterwölblingen groups16,22,86. However, a few outliers among the Central German individuals (e.g., KUC022) with elevated EEF-related ancestry hint at connections with southern groups, possibly facilitated by expanding trade networks.
The timing of the increase in EEF-related ancestry varied regionally. In South Germany and Bohemia, this shift occurred earlier, during the advanced EBA/MBA, and then stabilised. By contrast, Central Germany and SW/Central Poland, both regions north of the ranges of the Ore Mountains, appear to have experienced a delayed increase around 1000 BCE. The earlier shift in South Germany and Bohemia may be linked to stronger cultural and demographic ties with southeastern Europe. Archaeologically, we can observe an increase in the so-called “donauländische (Danubian)” influences in southern Germany and Bohemia from the advanced EBA onwards, reflected in specific axe forms (e.g., Nackenscheiben-, Nackenkamm-, Schaftröhrenäxte), pins (e.g., Kugelkopfnadeln), and hoard types (e.g., Koszider), which are much more frequent here than in the MES region, where they are rare or entirely absent. The same holds true for a wide array of ceramic vessel forms and ornaments, which clearly indicate the integration of southern Germany and Bohemia into a southeast-oriented communication network (e.g., W. Dehn’s A2/B1 ceramics). These cultural orientations continued into the MBA.
While these southeastern connections are visible earlier in the south, in Central Germany they become more apparent around 1000 BCE—both genetically and in terms of lifeways. At Kuckenburg and Esperstedt we observe a shift in subsistence strategies, where early LBA individuals consumed millet (likely in response to increasing aridity) while late LBA individuals returned to diets dominated by C3 plants, such as wheat and barley47. Early LBA individuals from Central Germany who consumed millet show a genetic profile consistent with EBA Únětice individuals from the region, supporting the interpretation that the adoption of millet was not necessarily introduced by incoming groups but likely reflected local responses to environmental or economic pressures, as discussed in our earlier work47. Intriguingly, it is the return to C3-plant-based diets that coincides with a stepwise increase in EEF-related ancestry and lower strontium isotope values and not the preceding shift to millet consumption during the early LBA (Fig. 3a), suggesting that the later changes were at least partly influenced by contact with groups practicing different subsistence strategies. However, as non-local individuals showed varied dietary profiles and local individuals also consumed millet, dietary shifts were likely not solely driven by contact but also by local cultural choices. Therefore, the association between higher EEF ancestry and the late LBA return to C3 plants does not necessarily imply causation. This pattern may reflect changes in agricultural practices, environment, culinary preferences or cultural influence rather than population movement. Proximal ancestry modelling confirms that most late LBA individuals from Central Germany cannot be modelled only with a local source, but as a mixture of local EBA groups and populations with higher EEF-related ancestry, such as those associated with Southern Germany/Switzerland but also the Knovíz culture in Bohemia, and groups from the Carpathian Basin and southern Europe. This genetic evidence reflects increasing demographic entanglement within Urnfield networks, which is also mirrored in the burial practices of the communities. At Kuckenburg and Esperstedt, we observe an increase in burials in settlement contexts around 1000 BCE. A similar practice is also observed at the Knovíz settlements in Bohemia35. Whether this corresponds to a decline in formal graveyard use remains unclear, as the Esperstedt graveyard has not yet been fully excavated, and neither is an increase in settlement contexts necessarily caused by a change in the ancestry of individuals. Importantly, settlement burials were not unusual in most prehistoric cultures from the Neolithic onwards, and do not appear to have been marginal or atypical in LBA Central German and Bohemian contexts32,38,87.
Most individuals across both LBA phases fall within the local strontium baseline, with only five individuals falling outside this range. We also found no significant difference in mobility patterns between male (n = 15) and female (n = 13) individuals, which contrasts with findings from EBA contexts, where female individuals often appear to have originated from different regions20,27,28. Additionally, strontium isotopic results indicated no significant differences between cremated and inhumed individuals, nor between skull-only and whole-body depositions, suggesting that geographical origin was not a key factor in mortuary treatment. However, this interpretation must be treated with caution due to differences in the formation periods of bone and enamel and the limited sample size. Nonetheless, one informative case is EOR021, an individual from Kuckenburg represented by a petrous portion of the temporal bone, whose formation period (0–2 years) is comparable to the enamel from the first molar. This individual falls within the local strontium range observed in the inhumed individuals. This supports the idea that geographical origin did not influence the choice between cremation and inhumation. Preliminary radiocarbon dating of five cremated individuals further suggests that cremation and inhumation were contemporaneous, indicating that these practices coexisted rather than replaced each other in chronological succession (Supplementary Data 10). Still, we cannot exclude the possibility that cremation and inhumation reflected different social or population strata, as thus far no DNA could be recovered from the cremated individuals.
The mortuary record at Kuckenburg and Esperstedt demonstrates a remarkably diverse repertoire. At these sites, we see that inhumation persisted alongside cremation, skull-only depositions, multi-staged funerary rites, and composite burials involving remains from multiple individuals. Our analysis suggests that these burial practices were not dictated solely by demographic factors. Specifically, we found no association between burial type and genetic sex, biological relatedness, strontium isotope variability, or inferred socioeconomic status (using grave goods and diet as proxies). However, we did observe that individuals buried in settlement contexts had higher EEF-related ancestry compared to those interred in the early LBA graveyard. These differences may partly reflect a chronological bias, rather than a differentiation based on ancestry, as the cemetery belongs to the early LBA while the settlement burials date mainly to the late LBA, further suggesting that cultural and social factors, rather than biological identity, played a central role in shaping burial customs. Additional sampling will be needed to assess this further. The absence of close genetic relatedness in most multiple burials further supports this view, indicating that burial group membership could have been socially constructed rather than biologically inherited51,88. A similar scenario has been proposed for Neckarsulm, where burials were interpreted as those of a warrior band54. Indeed, we did not detect any close or distant genetic relatedness among the double or multiple burials at the cemetery.
These results do not come entirely as a surprise, since limited close biological relatedness within cemeteries and mortuary contexts is increasingly recognised and described in prehistoric Europe. Although these examples come from different periods and regions, many studies now report low levels of close biological relatedness within mortuary groups. A recent study from Neolithic Ireland89, for example, shows that even long-used, well-sampled mortuary contexts often contain individuals who are only distantly related or unrelated, reflecting socially defined mortuary practices rather than biological kin groups. Similar patterns have been identified in EBA Central Europe20,28, where cemeteries often contain individuals of diverse origins with few close biological relatives. Another example comes from Knipper et al.90, which examines a 5th-century AD cemetery in Pannonia. Isotopic analyses there revealed that the community brought together people from varied geographical and cultural backgrounds, creating a socially constituted group rather than one based on biological descent. The limited biological relatedness observed at the sites studied here therefore fits well within this broader pattern and possibly demonstrates selective mortuary traditions, community belonging, and socially defined identities rather than direct biological descent. The inclusion and analysis of more individuals would refine this picture further.
Finally, feature 39/2018 at Kuckenburg, containing skull-only depositions, cremated remains, and animal bones, demonstrates another aspect of the complex mortuary practices of this period. Their burial sequence suggests multi-stage funerary treatment, supporting the idea that personhood (i.e., what it means to be a person)91,92 could have been relationally constructed and extended across time, involving both the living and the dead, as seen in other Bronze Age, but also Neolithic, contexts89,91,93,94. Additionally, missing skeletal elements (e.g., cervical vertebrae) in 18 out of 26 burials at Kuckenburg may also reflect the ongoing relationship between the living and the dead, represented by, for example, keeping these elements as relics88,95,96. Recent research has emphasised that the widespread adoption of cremation in the Late Bronze Age was not a uniform or linear shift, but rather the outcome of context-specific negotiations of identity, belief, and bodily treatment97. Our findings agree with this perspective, demonstrating that central German communities actively shaped their mortuary practices in response to growing regional connectivity and cultural entanglement. These practices did not represent rigid traditions nor passive adaptations to external influences. Rather, they represented creative negotiations of changing identities, relationships, cultural landscapes and beliefs. These negotiations are manifested in the coexistence of cremation and inhumation, the presence of multi-stage rites, and the use of settlement spaces for burial. Finally, they demonstrate that these communities were culturally flexible and adaptive, creating hybrid traditions by combining older traditions with new influences in locally meaningful ways.
Methods
Archaeological information
All studied specimens were obtained with permission to sample from the respective curating institutions, represented by collaboration partners in the author list. Excavations were carried out by the respective state heritage and equivalent organisations. No specific export/import permits are required for our study region. Detailed information on sample provenance, curating institutions, archaeological IDs, feature numbers, age-at-death, anthropological sex, and contextual and radiocarbon (14C) dates is provided in Supplementary Data 1 and Supplementary Note 1. The mortuary contexts were analysed by reviewing the anthropological reports from all sites, including detailed descriptions, drawings, and photographs, and through consultation with the responsible archaeologists (Supplementary Data 1 and Supplementary Note 1).
Osteological and palaeopathological analyses
The osteological and palaeopathological analyses of the inhumed individuals from Kuckenburg, the settlement of Esperstedt, and the burial ground of Esperstedt focused on recording individual biological profiles, such as age-at-death group and anthropological sex98,99,100,101, and assessing palaeopathological lesions/conditions102. Additional data for the Esperstedt individuals were incorporated from previous anthropological reports written by Sylvia Thüne44,45. For assessing pathological changes and their significance in relation to living conditions, only individuals with a minimum level of skeletal representation (at least 25% of the skeleton) were included103,104. The skeletal material was divided into anatomical zones (e.g., proximal, medial, and distal portions of the femur) to allow systematic morphological examination of all available skeletal elements. All morphological observations from individual skeletons were then linked, allowing for the individual analysis and comparison of collective data. This approach enabled the identification of palaeopathological lesions/conditions across the entire dataset as well as within specific subgroups defined by age-at-death, anthropological sex, or burial context103,104. For the cremated individuals from Esperstedt and Kuckenburg, anthropological reports were also reviewed. However, due to the high degree of fragmentation, age-at-death and anthropological sex estimation was not possible.
14C dating
Inhumations
Approximately 1 gram of bone from human and animal samples was sent to the Curt-Engelhorn-Centre Archaeometry gGmbH in Mannheim, Germany, for direct radiocarbon dating. This bone sample was the same one used for both aDNA and isotope analysis, ensuring that all analyses refer to the same individual. For the collagen extraction from the bone samples the lab used the modified Longin method105, and the collagen was purified by ultrafiltration (fraction > 30kD) and freeze-dried. The dates were already calibrated when received with OxCal 4.4106 using the IntCal 20 curve107. The C/N atomic ratios stay within the 2.9–3.6 range indicating good quality collagen108,109. All calibrated dates are reported in the text as 95.4% probability ranges. This study presents eleven newly generated radiocarbon dates (Supplementary Data 1), in addition to those published in Orfanou et al.47.
Cremations
We obtained six radiocarbon dates from fully calcined fragments of cremated bone (five diaphyseal fragments and one petrous portion of the temporal bone) (Supplementary Data 10). Black and grey coloured bones were avoided because of the high risk of carbon contamination110,111,112,113. These samples were also used for strontium isotope analysis. The bone samples were sent to the Royal Institute for Cultural Heritage (KIK-IRPA) in Brussels, Belgium. Their pre-treatment included CO2 extraction and graphitisation, following specific protocols114,115. The 14C/12C ratio in the graphite was measured with the AMS type MICADAS, mini carbon dating system, at the KIK-IRPA (Lab-code: RICH)116 and was converted into a radiocarbon age (expressed in years BP), after correction for isotope fractionation, using the δ13C AMS measurement. The dates were calibrated with OxCal 4.4106 using the IntCal 20 curve107.
Ancient DNA
Sample collection
We processed and analysed 75 samples from 13 Late Bronze Age archaeological sites in Central Europe. A detailed description of each site, including contextual data and available radiocarbon dates, is provided in the Supplementary Materials (Supplementary Note 1 and Supplementary Data 1). DNA sampling, extraction, and library preparation for next-generation sequencing were carried out in dedicated clean-room ancient DNA facilities at the Max Planck Institute of Geoanthropology (MPI-GEA) in Jena, Germany (formerly the Max Planck Institute for the Science of Human History), and the Max Planck Institute for Evolutionary Anthropology (MPI-EVA) in Leipzig, Germany. All samples were fully documented and photographed before processing. Additionally, all skulls from feature 39/2018 were CT-scanned before further processing.
DNA extraction and library preparation
We preferentially sampled the petrous portion of the temporal bone and teeth, and in a few cases, other elements such as scapulae (Supplementary Data 1). Samples were irradiated with ultraviolet (UV) light for 30 min on all sides. Teeth were cleaned with 3% bleach and sectioned along the cementum–enamel junction; powder was obtained from the pulp chamber. Petrous bones were drilled externally following a minimally invasive protocol117,118, after mechanical removal of the surface layer. DNA was extracted using the High Pure Viral Nucleic Acid Kit (Roche)119 with a modified silica-based protocol120, based on Dabney et al.121, as described in Ref. 122. We prepared double-indexed123, double- and single-stranded libraries using 25 μl and 20 μl of DNA extract, respectively, following established protocols for ancient DNA124,125. Details regarding library type (single-stranded or double-stranded) and uracil-DNA-glycosylase (UDG) treatment (i.e., none (minus) or partial (half-UDG), following ref. 126) are provided in Supplementary Data 1.
Sequencing
All indexed libraries were initially screened through shallow shotgun sequencing, generating approximately 5 million reads per sample using either an Illumina HiSeq 4000 or NextSeq500 platform with single-end 75 bp reads (1 × 75 bp). Quality assessment and initial screening of human DNA content and damage patterns were conducted using nf-core/eager127 (version: 2.5.0) pipeline. Libraries that met the minimum threshold of > 0.1% endogenous human DNA and had the characteristic aDNA damage patterns at the 5’ and 3’ ends of fragments55 were subsequently enriched via targeted in-solution hybridisation capture for approximately 1.24 million single nucleotide polymorphisms (SNPs) (“1240k capture”)56. These libraries were then sequenced to a target depth of 20 million reads on Illumina HiSeq 4000 or NextSeq500 platforms using single-read 75 bp (SR 75) chemistry. For libraries with fewer than 3000 mitochondrial reads, additional mitochondrial capture (“MT capture”)61,128 was performed. Additionally, for all genetically male individuals, we applied an in-house capture assay targeting ~10.4 kB within the non-recombining region of the Y chromosome (“YMCA capture”)129.
Bioinformatic processing
All raw sequence data were processed using the departmental pipelines of the Max Planck Institute for Evolutionary Anthropology (MPI-EVA) in Leipzig, Germany. These include:
Sequences were assigned to libraries based on index sequences, allowing for up to one mismatch per index. Adapter trimming and merging of paired-end reads were performed using leeHom v1.1.5130 with parameters optimised for ancient DNA (--ancientdna). For paired-end sequenced single-stranded libraries, we additionally used the following settings for ancient adapter identification:
-f
‘AGATCGGAAGAGCACACGTCTGAACTCCAGTCACIIIIIIIATCTCGTATGCCGTCTTCTGCTTG’
-s
‘GGAAGAGCGTCGTGTAGGGAAAGAGTGTAGATCTCGGTGGTCGCCGTATCATT’
-c
‘ACACTCTTTCCCTACACGTCTGAACTCCAG, ACACTCTTTCCCACACGTCTGAACTCCAGT, ACACTCTTTCCCTACACACGTCTGAACTCC, CTCTTTCCCTACACGTCTGAACTCCAGTCA, GAAGAGCACACGTCTGAACTCCAGTCACII, GAGCACACGTCTGAACTCCAGTCACIIIII, GATCGGAAGAGCACACGTCTGAACTCCAGT, AGATCGGAAGAGCACACGTCTGAACTCCAG, AGAGCACACGTCTGAACTCCAGTCACIIII, ACACGTCTGAACTCCAGTCACIIIIIIIAT, GTGCACACGTCTGAACTCCAGTCACIIIII, AGCACACGTCTGAACTCCAGTCACIIIIII, CGTATGCCGTCTTCTGCTTGAAAAAAAAAA’
The resulting reads were aligned to the human reference genome (hg19/GRCh37) using BWA v0.7.17131 with the parameters -n 0.01 -o 2 -l 16500. The mapped reads were then processed using nf-core/eager127 (version: 2.5.0), using the command: nextflow run nf-core/eager -r 2.5.0 -profile eva,archgen,medium_data,autorun,local_paths,TF -c (https://raw.githubusercontent.com/MPI-EVA-Archaeogenetics/Autorun_eager/1.6.0/conf/Autorun.config). Briefly, the reads were filtered to exclude reads below a length of 30 bp, prior to using Picard MarkDuplicates (version: 2.26.0) to remove PCR duplicates from each library. Where applicable, the unique reads from the captured double-stranded libraries were then trimmed to remove characteristic ancient DNA damage using bamUtils132 (version: 1.0.15), removing the first and last 2 bp of all UDG-half treated reads, from double-stranded libraries while all single-stranded libraries were left untrimmed. The resulting mapped reads from each sample across libraries were then merged (where applicable) and used for genotyping with pileupCaller, using the --randomHaploid calling method, applying mapping and base quality filters, and deactivating per-Base-Alignment-Quality (-Q 30 -q 25 -B --randomHaploid). Single-stranded libraries were genotyped with the additional option --singleStrandMode.
Genotype likelihoods
Deduplicated BAM files (without damage trimming) were used to estimate genotype likelihoods for every sample individually with ATLAS (v0.9; https://bitbucket.org/wegmannlab/atlas)133,134. Every library’s reads were divided between reads shorter than the number of sequencing cycles and reads equal or longer than that number with ATLAS’s splitMerge tool (75 bp). The library’s details were specified with the “readGroupSettings” setting and the option “allowForLarger” was also employed. PMD scores for each library were estimated with ATLAS’s “PMD” tool: for reads shorter than 75 bp in single-stranded single-end libraries or merged reads from paired-end double-stranded libraries, PMD patterns were estimated as a function of distance from both ends of the read; for reads longer than the expected number of sequencing cycles in single-stranded single end libraries or unmerged reads from double paired-end double stranded libraries, as a function of distance from just the beginning of the read (5’). Minimum base quality was set at 30 and maximum length of estimation from the ends at 70. The human reference genome (hg19) was specified with the “fasta = “ option. ATLAS’s “recal” tool was employed for reducing possible sequencing and reference biases. All the read groups from the samples’ sequencing plate were pooled with the option “poolReadGroups” and the invariant position for the model were specified through the “regions” options, specifying a list of 10 million highly conserved genomic positions across 88 mammal species downloaded from ensembl (https://grch37.ensembl.org/). The empiric PMD score calculation produced by the previous step was included in the recal calculation through the option “pmdFile” and the analysis was limited to chromosome 1 for computational reasons. Genotypes on the autosomal chromosomes were called with ATLAS’s “call” tool in the 1000 Genomes Phase 3 release data (1KG) 30 M panel135 employing the “MLE” algorithm. PMD Empiric pattern and recal patterns were included in the calculations with the “pmdFile” and “recal” flags respectively. The reference fasta file of the human reference genome (hg19) was specified with the “fasta” option, and the “printAll” flag was employed. Genotype calling was performed on bases with score equal or higher than 30 with the option “minQual=30”.
Imputation
Diploid autosomal genotypes produced with ATLAS were imputed to the entire 1000 Genomes Phase 3 release data (1KG) 30 M panel with GLIMPSE v1.1.1 (https://odelaneau.github.io/GLIMPSEglimpse1/)136. GLIMPSE was run with the default parameters using sex averaged genetic maps from HapMap as suggested in the original publication137. Imputation and phasing on genomic chunks of 2,000,000 base pairs (bp) with a buffer of 200,000 bp was performed with the “GLIMPSE_phase” function. The GLIMPSE_ligate and GLIMPSE_sample (with the flag --solve) functions together with bcftools v1.387,91138 were employed to obtain the final phase/imputed vcf files with the genotypes posterior probabilities (GP) at every 1KGP panel position. These regions were then subset to the overlap with the 1240k panel SNP positions for IBD inference.
Authentication and quality control
Genetic sex determination was carried out using Sex.DetERRmine139 (version: 1.1.2) on reads filtered for mapping and base quality (samtools version: 1.12; parameters -q30 -Q30) by inferring the relative sequence coverage on X, Y, and autosomal chromosomes, normalised by chromosome length139. In the absence of contamination, an X ratio of 1 and Y ratio of 0 is expected for female individuals, while male individuals are characterised by approximate ratios of 0.5 for both X and Y chromosomes140. Individuals positioned between the expected male and female ratios are indicative of contamination from the opposite sex. Then, exogenous human contamination was estimated at mitochondrial and nuclear level. Specifically, mitochondrial contamination was assessed using ContamMix141(version: 1.0.12), and individuals with mitochondrial contamination estimates >10% were excluded. For genetically male individuals, nuclear contamination was estimated with ANGSD142 (version: 0.935) and hapCon143 (version: 0.70), and samples with nuclear contamination >5% were excluded.
Reference data
We merged our newly generated aDNA data with published ancient and modern genomes from the Allen Ancient DNA Resource (AADR), version v54.1.p1144 (Supplementary Data 2b). This consists of two datasets, the Human Origins array (HO) with 593,573 SNPs, was used exclusively for PCA and ADMIXTURE analyses, as it contains a broader representation of present-day populations, both in terms of the number of groups and the number of individuals per group, allowing for more robust comparisons between ancient and modern groups. Meanwhile the 1240k dataset (TF) with 1,233,013 SNPs, was used for comparative analysis among ancient groups using f-statistics and ancestry modelling. All the population genetic analyses besides IBD analyses were conducted with pseudo-haploid genotypes produced with pileupCaller.
Principal component analysis
PCA was conducted on the Human Origins dataset (~593,000 autosomal SNPs) using smartpca v16000 from the EIGENSOFT package v6.0.1145. Principal components were calculated based on 77 present-day populations from Eurasia, North Africa, and the Caucasus to capture modern population structure (Supplementary Data 2b). Ancient individuals were then projected onto these axes using the options lsqproject: YES, and shrinkmode: YES. Individuals with <30,000 SNPs were excluded from projection.
ADMIXTURE analysis
Unsupervised model-based genetic clustering was performed using ADMIXTURE146 (v1.3.0) software. A dataset of modern populations from the Human Origins array and published ancient individuals was used for this analysis (Supplementary Data 2b). The dataset was filtered to a subset of ~143,000 ancestry-informative SNPs147. Additionally, individuals with low coverage were excluded from analysis. For each value of K (ancestral component) ranging from 1-20, the default fivefold cross-validation (--cv=5) was used to determine the coefficient of variance for each K.
F 4-statistics
We computed f4-statistics using the qpDstat programme from ADMIXTOOLS v.152060, with the parameter f4mode: YES, and printsd: YES. All tests were of the form f4(outgroup, X; Y, Z), where the outgroup was a composite “OldAfrica” population consisting of a pool of diverse ancient African individuals with no evidence of recent admixture from West Eurasian sources, following the approach outlined in ref. 18. The X population represented one of the three main West Eurasian ancestry components: Early European Farmer (EEF - Balkan_N, Turkey_N), Western Hunter-Gatherer (WHG - WHGA and WHGB), or Steppe-related ancestry (OldSteppe) (Supplementary Data 2b, column “Group ID_published”). Y and Z were test populations of interest. Z-scores were used to assess significance, with |Z|≥ 3 considered statistically significant. For comparisons involving the Neckarsulm (NES) group, the “NES_group” was defined as all NES individuals that cluster together on PCA (i.e., excluding the outlier NES021).
Ancestry modelling and calculation of ancestry proportions
We used qpWave and qpAdm (ADMIXTOOLS v1520)61,62 to model ancestry proportions and test admixture models. Analyses were performed using the 1240k panel (~1.24 million SNPs). The allsnps: NO setting was used for consistent SNP overlap; Outgroups included OldAfrica, Turkey_N, WHGB, and Afanasievo, following Patterson et al.18. Due to their close genetic similarity, individuals classified as Italy_Sardinia_EBA and Italy_Sardinia_MBA were combined into a single group labelled as Italy_Sardinia_EMBA (Supplementary Data 2b). Similarly, Spain_EBA and Spain_MBA were grouped as Spain_EMBA, while Serbia_BA_Maros and Serbia_Mokrin_EBA_Maros were combined under the label Serbia_BA (Supplementary Data 2b). Feasible models had p-values > 0.05; among these, we prioritised more parsimonious models with temporally and geographically proximal source populations.
Spatiotemporal interpolation (mobest)
We used mobest v1.0.0 to conduct spatiotemporal interpolation and probabilistic similarity search of genetic ancestry components65. We analysed outlier individuals from Kuckenburg, Neckarsulm, Slaný, and Libčice nad Vltavou. PCA coordinates (PC1 and PC2) from the above analysis were used as input, following methods outlined at (https://github.com/nevrome/mobest). The analysis used a western Eurasian base map (EPSG: 3035) and included ancient individuals from the AADR version v54.1.p1144. Sample dates were set using the calibrated radiocarbon age-at-death.
Uniparentally inherited markers
We combined shotgun, 1240k, and mitochondrial capture data to call mitochondrial haplogroups. Reads were aligned to the revised Cambridge Reference Sequence (rCRS, NC_012920.1), and a consensus sequence for each individual was generated using Geneious148 v2019.2.3. Mitochondrial haplogroups were then assigned using HaploGrep2149 and HaploGrep3150. For male individuals, Y-chromosome haplogroups were determined by manual assignment, following the approach described in ref. 129.
Biological relatedness
Biological relatedness was assessed by using the software BREADR version 1.0.367 (Supplementary Data 5a) and READv266 (Supplementary Data 5b) using default parameters, which estimate genetic relatedness (up to the second- and third-degree, respectively) by calculating pairwise mismatch rates between pairs of individuals. In addition, KIN68 (Supplementary Data 5c) was used to infer genetic relatedness through detection of identical-by-descent segments shared between individuals. We restricted our analyses to on-target sites from the 1240k capture array56 and assessed relatedness between pairs of individuals within the same genetic grouping (Central_Germany_early_LBA, Central_Germany_late_LBA, South_Germany_early_LBA, Bohemia_LBA and SW_Central_Poland). Individuals assigned to Bohemia_early_LBA (n = 13) and Bohemia_late_LBA (n = 2) were combined for relatedness estimation due to the low number of individuals in the latter group. Similarly, individuals from SW_Central_Poland were analyzed jointly.
Runs of homozygosity
We used the hapROH70 software (v1.0) to identify runs of homozygosity (ROH) and assess parental relatedness. We only applied this method to individuals that had >400,000 SNPs.
Identity-by-descent
We analysed identity-by-descent (IBD) segments among individuals by using the ancIBD69 software (v0.4). Diploid genotypes imputed with GLIMPSE and with >500,000 SNPs and genotype probabilities >0.7 were included in the analysis. The HapBLOCK function was used to estimate IBD sharing. Post-imputation, the vcf files were converted to the hdf5 format using the vcf_to_1240K_hdf command. IBD analysis was performed on a per-chromosome basis using the default parameters of the hapBLOCK_chroms command. Summary statistics were generated for each pair, retaining only those with at least one IBD segment spanning 20 cM for visualisation.
Pathogen screening
All sequencing data were further screened for the presence of pathogen DNA using the screening pipeline HOPS79. For this, adapter-trimmed reads were initially mapped with bwa aln (v0.7.12) with -n 0.01 and -l 16 to a multifasta reference containing the references of different representative human pathogenic species. Mapped reads were extracted from the bam files and transformed in fastq format with samtools (v1.3) the subcommands view and fastq. The resulting fastq files were used as input for an in-house automatic pathogen screening pipeline151 (available via Zenodo at https://doi.org/10.5281/zenodo.18403831, and GitHub at https://github.com/MPI-EVA-Archaeogenetics/Autorun_deepscreening), that utilises HOPS within nf-core/eager (v.2.4.7). Input data were mapped against the PhiX reference (NC_001422.1) and unmapped reads were then aligned and taxonomically classified with the MEGAN ALignment Tool (MALT)152,153. Assignments were then filtered and authenticated within the pipeline for potential pathogen candidates. Only candidates marked as stp3 were considered as authentic, when they fulfilled the criteria of a decreasing edit distance compared to the reference, a damage profile consistent ancient samples (C > T substitutions at the 5’ prime end) as well as a decreasing edit distance to the reference in reads with potential aDNA damage. Rma6 files of potential candidates were further visually inspected in MEGAN154.
Isotope analyses
Sample collection
We sampled 29 inhumed and 21 cremated individuals from Esperstedt and Kuckenburg. For the inhumations, our initial aim was to sample enamel from both first and third molars to assess potential movement across different life stages (childhood vs. adolescence). However, due to the limited preservation and availability of these teeth, we predominantly sampled second molars (M2), which provide information about early to mid-childhood.
Moreover, it is important to note that δ18O values from M1 should be interpreted with caution as they can be influenced by breastfeeding25. For the cremations, we sampled any available bones that were fully calcined for the cremated individuals due to the limited preservation of skeletal remains; enamel rarely survives the cremation process. Additionally, we sampled one cremated petrous part of the temporal bone available from Kuckenburg, which forms between 0 and 2 years of age and does not undergo further remodelling. It is important to acknowledge the different formation times between bones and teeth when interpreting our results. The sex and age at death of the inhumed individuals are reported in Supplementary Data 1. Sex determination for the inhumations was initially assessed anthropologically and then confirmed genetically. Due to fragmentation issues, the sex and age at death of the cremations were not assessed. The inhumations and cremations from Esperstedt were curated at the Halle Museum, and all necessary permissions for sampling were obtained.
Construction of a bioavailable Strontium baseline
The bioavailable strontium baseline for our analysis was constructed using 18 ancient faunal remains from the two sites, which could be contextually dated or radiocarbon dated to the LBA. We also integrated our baseline with the published baseline for Esperstedt during the EBA (based on archaeological fauna, three pigs), as reported by Knipper et al.26. Among the faunal remains, we also included animals with limited mobility such as hamsters, a rabbit and terrestrial snails, although the latter have their own limitations155. While modern plants are considered the most direct proxy for bioavailable strontium155, they were not available for this study. We acknowledge that some domesticated fauna (e.g., pigs, sheep or goats) may have been herded, exchanged or traded; however, we believe that the inclusion of low-mobility taxa such as rodents helps mitigate this concern and provides a more secure local signal. Additionally, the resulting baseline range agrees with previous research and published geological maps for the region26, supporting its reliability for distinguishing local from non-local individuals.
Strontium isotope analysis on tooth enamel
Tooth surfaces were mechanically cleaned using a diamond-tipped drill to remove surface contaminants. We collected ~15–20 mg of enamel powder by drilling along the full buccal surface to obtain a bulk sample. The samples were prepared and analysed at the University of Cape Town following standard protocols (e.g., Ingman et al., 2021)156.
Strontium isotope analysis on calcined bone
Cremated bone fragments were mechanically cleaned using a Dremel with a diamond burr before being rinsed three times with milliQ water. For each rinse, the samples were placed for 10 minutes in an ultrasonication bath. Cremated bone and tooth fragments were treated with 1 M acetic acid for 3 to 10 minutes in the ultrasonication bath and then rinsed again with milliQ water and 10 minutes of ultrasonication77. The petrous part from Kuckenburg was sampled and pretreated following Veselka et al.157. Strontium was extracted through column chemistry using ion exchange resin (Sr-Spec, Triskem). In short, the columns and resin were rinsed 2 x 1 mL 2 M HNO3, then 2 x 1 mL 7 M HNO3, samples loaded in 7 M HNO3, charged 4 x 1 mL 7 M HNO3, rinsed using 5 x 1 mL 7 M HNO3 and Sr was collected with 6 x 1 mL 0.05 M HNO3 and evaporated again to dryness. 87Sr/86Sr measurements were carried out on a Nu Plasma 3 MC-ICP-MS (PD017 from Nu Instruments, Wrexham, UK) at VUB. Particular attention was paid to the purity of the Argon gas used inside the spectrometer in order to avoid any interference (from Kr, for instance) on Sr isotope masses. The Sr isotopes were measured by static multi-collection. Each analysis consisted of 60 ratio measurements (2 blocks of 30 cycles). All the Sr isotopes (84, 86, 87, 88) were measured, while the masses 85 (Rb) and 83 (Kr) were simultaneously monitored, allowing for interference corrections on masses 84, 86 (Kr) and 87 (Rb). The Sr isotopic ratios were automatically normalised to 86Sr/88Sr = 0.1194 using an exponential law. During this study, repeated measurements of the NIST-SRM987 standard yielded 87Sr/86Sr = 0.710244 ± 0.000014 (2 SD for; n = 6), which is consistent with the mean value of 0.710252 ± 0.000013 (2 SD for 88 analyses) obtained by TIMS (Thermal Ionisation Mass Spectrometry) instrumentation158. All the sample measurements were normalised using a standard bracketing method with the recommended value of 87Sr/86Sr = 0.710248158. Procedural blanks were considered negligible (total Sr (V) of max 0.02 versus 10 V for analyses, i.e., ≈ 0.2%). The 87Sr/86Sr value is reported for each sample with two standard errors (absolute error value of the individual sample analysis – internal error). During the course of these analyses the matrix matched standards NIST SRM1400 (bone ash) was used and returned mean 87Sr/86Sr values of 0.713118 ± 0.000032 (2 SD; n = 4) which is close to the long term 87Sr/86Sr values of 0.713117 ± 0.000031 (2 SD; n = 345) obtained by Gerritzen et al.159.
Oxygen isotope analysis
We collected 10 mg of tooth enamel powder using a diamond-tipped drill for stable carbon and oxygen isotope analysis. We obtained bulk samples for our analysis by sampling along the entire length of the buccal surface of the tooth. The tooth enamel samples were pretreated with 1 mL of 1% NaClO for 1 h. Then, they were rinsed 3 times with ultra-pure water. 1 mL of 0.1 M acetic acid was added for 10 min, and then the samples were rinsed again 3 times. Next, the samples were lyophilised in a freeze dryer for 4 h before being weighed out (2 mg each) into borosilicate glass vials for analysis using a Thermo Gasbench II system connected to a Thermo Delta V Advantage Mass spectrometer at the Max Planck Institute of Geoanthropology in Jena, Germany. A pure CO2 reference gas was used to calculate provisional delta values of the sample peaks. Final stable carbon (δ13C, reported relative to VPDB) and oxygen (δ18O, reported relative to VSMOW) isotope values of the samples are obtained by calibrating the provisional sample data with international standards IAEA NBS 18 (δ13C -5.014 ± 0.03 ‰, δ18O -23.2 ± 0.1 ‰), IAEA 603 (δ13C + 2.46 ± 0.01 ‰, δ18O -2.37 ± 0.04 ‰), IAEA CO8 (δ13C -5.76 ± 0.03 ‰, δ18O -22.7 ± 0.2 ‰), and USGS44 (δ13C = ~ —42.1 ‰) after correcting for linearity and instrumental drift. Replicate analyses of standards suggest that machine measurement error is c. ± 0.1‰ for δ13C, and ± 0.2‰ for δ18O. Overall measurement precision was studied through the measurement of repeat extracts from a bovid tooth enamel standard (n = 20, ± 0.2‰ for δ13C, and ± 0.4‰ for δ18O).
Statistical tests
To assess if the changes in EEF proportion over time were significant, we applied a non-parametric Kruskal–Wallis160 test, when assumptions of normality and equality of variance (Shapiro–Wilk161 and Levene’s test respectively) were not met or when sample sizes were insufficient. The time series plot was created in R. For visualisation, we used a smooth.spline() function that fits a cubic smoothing spline to the data.
We also tested for associations between burial type (skull-only vs. whole body), age-at-death, genetic sex, EEF-related ancestry, 87Sr/86Sr values, and δ¹³C dietary signatures (from collagen and enamel). For continuous variables, we first assessed the distribution of each group using the Shapiro-Wilk test. If both groups met the assumptions of normality (p > 0.05), we applied Welch’s two-sample t-test162 otherwise, we used the non-parametric Wilcoxon rank-sum test. For categorical variables, we used Fisher’s exact test163 due to small sample sizes in several categories. Outliers in continuous variables were identified using the 1.5×IQR rule. All statistical tests were two-sided tests using default parameters, unless stated otherwise. All reported p-values were adjusted for multiple testing using either the Bonferroni or the Benjamini-Hochberg False Discovery Rate (FDR) correction164 within each family of related tests. The p.adjust() function in R (method = “fdr”) was used for the FDR correction. The Bonferroni correction was applied for analyses involving a small number of planned comparisons, while FDR correction was used for analyses involving a larger number of comparisons, as Bonferroni correction can be overly conservative in such contexts. We considered results significant at an adjusted p-value threshold of 0.05. All analyses were performed in R 4.2.2165. All statistical tests and adjusted p-values are provided in Supplementary Data 12.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Genomic sequence data generated in this study are available at the European Nucleotide Archive (ENA) under accession number PRJEB98508. Newly generated isotope data generated in this study are provided in Supplementary Data 8. New radiocarbon dates are given in Supplementary Data 1 and 10. Information on all individuals analysed in this study is provided in Supplementary Data 1, including archaeological identifiers (column B), site and context information (columns L-P), curating institutions (column BV), and the responsible curators (column BU), as well as the site descriptions in Supplementary Note 1. Published genomic data used in this study were obtained from public repositories, such as the Allen Ancient DNA Resource144 (AADR, v54.1.p1), accessed via the Harvard Dataverse166 (https://doi.org/10.7910/DVN/FFIDCW), and Poseidon (https://www.poseidon-adna.org/#/). The Genome Reference Consortium Human Build 38 (GrCh38) and Build 37 (GRCh37) are available via the National Center for Biotechnology Information and can be found here: (https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001405.26/), and here: (https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001405.13/). The 1000 human genomes reference panel is available here: (http://hgdownload.cse.ucsc.edu/gbdb/hg19/1000Genomes/phase3/), and the human mitochondrial revised Cambridge Reference Sequence (NC 012920.1) can be found here: (https://www.ncbi.nlm.nih.gov/nuccore/251831106).
Code availability
All software used in this work is publicly available and versions are listed in the Nature Portfolio Reporting Summary. Corresponding publications are cited in the Article and Supplementary information. Code for the pathogen screening pipeline is available here: https://zenodo.org/records/18403831.
References
Harding, A. F. European Societies in the Bronze Age. (Cambridge University Press, 2000).
Primas, M. Bronzezeit zwischen Elbe und Po. Strukturwandel in Zentraleuropa, 2200-800 v. Chr. Universitätsforschungen zur prähistorischen Archäologie 150, 223–262 (2008).
Vandkilde, H. Forward. in Rooted in Movement: Aspects in Mobility in Bronze Age Europe (eds. Reiter, S., Norgaard, H. W., Kölcze, Z. & Rassmann, C.) 9–10 (Jutland Archaeological Society Publications, Denmark, 2014).
Vandkilde, H. Bronzization: The Bronze Age as Pre-Modern Globalization. Praehist. Zeitschrift 91, 103–123 (2016).
von Schnurbein (Hrsg.), Siegmar, Bernhard Hänsel, Carola Metzner-Nebelsick, Johannes Müller, Rosemarie Müller, Susanne Sievers und Thomas Terberger. Atlas der Vorgeschichte. Europa von den Ersten Menschen bis Christi Geburt. (Verlag Konrad Theiss, Stuttgart, 2010).
Cavazzuti, C. et al. The first ‘urnfields’ in the plains of the Danube and the Po. J. World Prehistory 35, 45–86 (2022).
Fredericks, J. D., Ringrose, T. J., Dicken, A., Williams, A. & Bennett, P. A potential new diagnostic tool to aid DNA analysis from heat compromised bone using colorimetry: A preliminary study. Sci. Justice 55, 124–130 (2015).
Hansen, H. B. et al. Comparing ancient DNA preservation in petrous bone and tooth cementum. PLoS One 12, e0170940 (2017).
Gaudio, D. et al. Genome-wide DNA from degraded petrous bones and the assessment of sex and probable geographic origins of forensic cases. Sci. Rep. 9, 8226 (2019).
Snoeck, C. Isotope analysis from cremated remains. In Burnt Human Remains (ed. Ellingham, S., Adserias-Garriga, J., Zapico, S. C. & Ubelaker, D. H.) 273–289 (Wiley, 2023).
Kristiansen, K. Towards a new paradigm? the third science revolution and its possible consequences in archaeology. Curr. Swed. Archaeol. 22, 11–34 (2014).
Haber, M., Mezzavilla, M., Xue, Y. & Tyler-Smith, C. Ancient DNA and the rewriting of human history: be sparing with Occam’s razor. Genome Biol. 17, 1 (2016).
Orlando, L. et al. Ancient DNA analysis. Nat. Rev. Methods Primers 1, 14 (2021).
Mathieson, I. et al. The genomic history of southeastern Europe. Nature 555, 197–203 (2018).
Olalde, I. et al. The genomic history of the Iberian Peninsula over the past 8000 years. Science 363, 1230–1234 (2019).
Furtwängler, A. et al. Ancient genomes reveal social and genetic structure of Late Neolithic Switzerland. Nat. Commun. 11, 1915 (2020).
Papac, L. et al. Dynamic changes in genomic and social structures in third millennium BCE central Europe. Sci. Adv. 7, eabi6941 (2021).
Patterson, N. et al. Large-scale migration into Britain during the middle to Late Bronze Age. Nature 601, 588–594 (2022).
Lazaridis, I. et al. The genetic origin of the Indo-Europeans. Nature 639, 132–142 (2025).
Mittnik, A. et al. Kinship-based social inequality in Bronze Age Europe. Science 366, 731–734 (2019).
Chyleński, M. et al. Patrilocality and hunter-gatherer-related ancestry of populations in East-Central Europe during the Middle Bronze Age. Nat. Commun. 14, 4395 (2023).
Furtwängler, A. et al. Tracing social mechanisms and interregional connections in Early Bronze Age Societies in Lower Austria. Nat. Commun. 17, 131 (2025).
Bentley, R. A. Strontium Isotopes from the Earth to the archaeological skeleton: a review. J. Archaeological Method Theory 13, 135–187 (2006).
Montgomery, J. Passports from the past: Investigating human dispersals using strontium isotope analysis of tooth enamel. Ann. Hum. Biol. 37, 325–346 (2010).
Pederzani, S. & Britton, K. Oxygen isotopes in bioarchaeology: principles and applications, challenges and opportunities. Earth-Sci. Rev. 188, 77–107 (2019).
Knipper, C. et al. A distinct section of the early Bronze Age society? stable isotope investigations of burials in settlement pits and multiple inhumations of the Únětice culture in central Germany. Am. J. Phys. Anthropol. 159, 496–516 (2016).
Penske, S. et al. Kinship practices at the early Bronze Age site of Leubingen in Central Germany. Sci. Rep. 14, 3871 (2024).
Knipper, C. et al. Female exogamy and gene pool diversification at the transition from the Final Neolithic to the Early Bronze Age in central Europe. Proc. Natl. Acad. Sci. USA. 114, 10083–10088 (2017).
Schmidt, B. & Nitzschke, W. Bestattungssitten der spätbronzezeitlichen Helsmdorfer- und Saalemündungsgruppe. Ausgrab. Funde 19, 6–17 (1974).
Schmidt, B. Die jungbronzezeitlichen Stämme im Elbe-Saale-Gebiet. In Mitteleuropäische Bronzezeit. Beiträge zur Archäologie und Geschichte (eds. Coblenz, W. & Horst, F.) 136–212 (Berlin, 1978).
Maraszek, R., Muhl, A., Schwarz, R., & Zich, B. Glutgeboren: Mittelbronzezeit Bis Eisenzeit. Begleithefte Zur Dauerausstellung Im Landesmuseum Für Vorgeschichte Halle Band 5, 33–44 (Landesmuseum für Vorgeschichte Halle, 2015).
Balfanz, K. & Jarecki, H. Jung-und spätbronzezeitliche Sonderbestattungen in Mitteldeutschland. Quellen und Fragestellungen. Jahresschr. für Mitteldtsch. Vorgesch. 88, 339–378 (2004).
Urák, M. & Marta, L. Human remains of the Late Bronze Age settlements in the Upper Tisza Area. New research and new evidence. in Bronze Age rites and rituals in the Carpathian Basin 2010 (ed. Tincu, V.S. &A.) 155–162 (Editura Mega, Târgu Mureș, 2011).
Ailincăi, S.-C. Trăind cu morţii: înmormântări în aşezările de la începutul epocii fierului între Balcani, Tisa şi Nistru. Living with the dead. Burials in Early Iron Age settlements between the Balkans, Tisza and Dnestr. in Minerva, II. Fontes archaeologici 2. (Academia Română, Cluj-Napoca, 2015).
Smejtek, L. Osídlení z Doby Bronzové v Kněževsi u Prahy–Bronze Age Settlement at Kněževes near Prague. (Praha, 2011).
Rişcuţa, N. C. & Marc, A. T. Children Burials in the Bronze Age Settlement from Şoimuş–Teleghi (Hunedoara County). in Settlements of Life and Death. Studies from Prehistory to Middle Ages. Proceedings of an International Colloquium, Tulcea, 25th–28th of May 2016 (ed. Gogâltan &S.-C. Ailincăi, F.) 201–234 (Editura Mega, Cluj-Napoca, 2016).
Stapel, A. Bronzezeitliche Deponierungen im Siedlungsbereich: Altdorf-Römerfeld und Altheim, Landkreis Landshut. (Waxmann, Münster, Germany, 1999).
Berecki, S., Németh, R. E., & Rezi, B. (eds). Bronze Age Rites and Rituals in the Carpathian Basin. Proceedings of the International Colloquium from Târgu Mureş, 8–10 October 2010. (Editura Mega, Târgu Mureș, 2011).
Ettel, P. Vorbericht zu den Grabungen 2005 bis 2011 der Friedrich-Schiller-Universität Jena auf der Kuckenburg bei Esperstedt. Saalekreis. Archäologie Sachsen - Anhalt 8, 19–38 (2016).
Ettel, P., Paust, E., & Schneider, F. Kuckenburg–eine Höhensiedlung in Mitteldeutschland: Ein Beitrag zu Höhensiedlungen, Körpergräbern und Skelettfunden im Siedlungskontext der Spätbronzezeit. in WegBegleiter: Interdisziplinäre Beiträge zur Altwege- und Burgenforschung. Festschrift für Bernd W. Bahn zu seinem 80. Geburtstag (ed. M. Freudenreich, P. Fütterer, & A. Swieder) 243–268 (Beier & Beran, Langenweißbach, 2019).
Ettel, P. & Paust, E. Spätbronzezeitliche Schädeldeponierungen auf der Höhensiedlung Kuckenberg bei Esperstadt, Saalekreis. In Rituelle Gewalt - Rituale der Gewalt: 12. Mitteldeutscher Archäologentag vom 10. bis 12. Oktober 2019 in Halle (Saale) = Ritual violence - rituals of violence: 12th Archaeological Conference of Central Germany, October 12-10 [i.e. 10-12], 2019 in Halle (Saale),Tagungen des Landesmuseums für Vorgeschichte Halle 22 (ed. Meller, H., Risch, R., Alt, K. W., Bertemes, F. & Micó P. R.) 701–710 (Landesamt für Denkmalpflege und Archäologie Sachsen-Anhalt, Landesmuseum für Vorgeschichte, Halle (Saale), 2020).
Ettel, P. Die spätbronze- und eisenzeitliche Höhensiedlungen Alter Gleisberg bei Jena und Kuckenburg bei Esperstedt in Mitteldeutschland. in Die Frühgeschichte von Krieg und Konflikt: Beiträge der Vierten Internationalen LOEWE-Konferenz vom 7. bis 9. Oktober 2019 in Frankfurt/M. = The early history of war and conflict (ed. Hansen Svend, Krause Rüdiger) 461–496 (Dr. Rudolf Habelt, Bonn, 2022).
Ettel, P. et al. Die Kuckenburg bei Esperstedt und ihre Toten: bioarchäologische Untersuchungen an Sonderbestattungen aus einer spätbronzezeitlichen Höhensiedlung. in Soziale Hierarchien zwischen Tradition und Innovation in der Bronzezeit Europas: Beiträge der Sitzungen der A. G. Bronzezeit im Rahmen der 83. Tagung des NWVA vom 12.-15. September 2018 in Heide (Holstein) und der 85. Jahrestagung des WSVA und des MOVA vom 1.-5. April 2019 in Würzburg (ed. Massy Ken, Bunnefeld Jan-Heinrich, Horn Christian) 363–376 (Beier & Beran, Langenweißbach, 2023).
Grothe, A. & Bogen, C. Das spätbronzezeitliche Gräberfeld von Esperstedt. in Archäologie auf der Überholspur. Ausgrabungen an der A38. Archäologie in Sachsen-Anhalt – Sonderband 5 (ed. Meller (Hrsg.), H.) 160–194 (Halle, 2006).
Müller, U. Zufall – Absicht – Abfall? Siedlungsbestattungen im bronzezeitlichen Esperstedt. in Archäologie auf der Überholspur. Ausgrabungen an der A38. Archäologie in Sachsen-Anhalt – Sonderband 5 (ed. Meller (Hrsg.), H.) 148–159 (Halle, 2006).
Roberts, B. W., M. Uckelmann, and D. Brandherm. Old father time: The bronze age chronology of western Europe. in The Oxford Handbook of the European Bronze Age (eds. Fokkens, H. & Harding, A.) (Oxford University Press, 2013).
Orfanou, E. et al. Biomolecular evidence for changing millet reliance in Late Bronze Age central Germany. Sci. Rep. 14, 4382 (2024).
Parker Pearson, M. Archaeology of Death and Burial. (Sutton Publishing, 1999).
Schrader, S. Activity, diet and social practice: Addressing everyday life in human skeletal remains. Springer International Publishing: Cham, 2019.
Schrader, S. A., Torres-Rouff, C. Embodying bioarchaeology. Theoretical Approaches in Bioarchaeology. 15–27 (Routledge2020Routledge/Taylor & Francis Group: London; New York, (2020).
Brück, J. Ancient DNA, kinship and relational identities in Bronze Age Britain. Antiquity 95, 228–237 (2021).
Steuer, H. Frühgeschichtliche Sozialstrukturen in Mitteleuropa: Eine Analyse der Auswertungsmethoden des Archäologischen Quellenmaterials. Abhandlungen der Akademie der Wissenschaften in Göttingen. Philologisch-Historische Klasse 3. Folge, Nr. 128. (Vandenhoeck & Ruprecht, Göttingen, 1982).
Kristiansen, K. Europe before History. (Cambridge University Press, England, 1998).
Knöpke, S. Der Urnenfelderzeitliche Männerfriedhof von Neckarsulm. Mit einem Anhang von Joachim Wahl. Die Menschlichen Skelettreste aus dem Urnenfelderzeitlichen Männerfriedhof von Neckarsulm.Trendpark Süd. Forschungen und Berichte zur Vor-und Frühgeschichte in Baden-Württemberg. Konrad Theiss: Stuttgart, (2009).
Dabney, J., Meyer, M. & Pääbo, S. Ancient DNA damage. Cold Spring Harb. Perspect. Biol. 5, a012567–a012567 (2013).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015).
Brandt, G. et al. Ancient DNA reveals key stages in the formation of central European mitochondrial genetic diversity. Science 342, 257–261 (2013).
Burger, J. et al. Low prevalence of lactase persistence in Bronze Age Europe indicates ongoing strong selection over the last 3,000 years. Curr. Biol. 30, 4307–4315 (2020).
Wahl, J. & Price, T. D. Local and foreign males in a late Bronze Age cemetery at Neckarsulm, south-western Germany: strontium isotope investigations. Anthropol. Anz. 70, 289–307 (2013).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Harney, É, Patterson, N., Reich, D. & Wakeley, J. Assessing the performance of qpAdm: a statistical tool for studying population admixture. Genetics 217, iyaa045 (2021).
Gretzinger, J. et al. Evidence for dynastic succession among early Celtic elites in Central Europe. Nat. Hum. Behav. 8, 1467–1480 (2024).
McColl, H. et al. Tracing the spread of celtic languages using ancient genomics. Preprint at bioRxiv https://doi.org/10.1101/2025.02.28.640770 (2025).
Schmid, C. & Schiffels, S. Estimating human mobility in Holocene Western Eurasia with large-scale ancient genomic data. Proc. Natl. Acad. Sci. USA. 120, e2218375120 (2023).
Alaçamlı, E. et al. READv2: advanced and user-friendly detection of biological relatedness in archaeogenomics. Genome Biol. 25, 216 (2024).
Rohrlach, A. B., Tuke, J., Popli, D. R. & Haak, W. BREADR: An R package for the Bayesian estimation of genetic relatedness from low-coverage genotype data. J. Open Source Softw. 10, 7916 (2025).
Popli, D., Peyrégne, S. & Peter, B. M. KIN: a method to infer relatedness from low-coverage ancient DNA. Genome Biol. 24, 10 (2023).
Ringbauer, H. et al. Accurate detection of identity-by-descent segments in human ancient DNA. Nat. Genet. 56, 143–151 (2024).
Ringbauer, H., Novembre, J. & Steinrücken, M. Parental relatedness through time revealed by runs of homozygosity in ancient DNA. Nat. Commun. 12, 5425 (2021).
Skourtanioti, E. et al. Ancient DNA reveals admixture history and endogamy in the prehistoric Aegean. Nat. Ecol. Evol. 7, 290–303 (2023).
Ghalichi, A. et al. The rise and transformation of Bronze Age pastoralists in the Caucasus. Nature 635, 917–925 (2024).
Bataille, C. P., Crowley, B. E., Wooller, M. J. & Bowen, G. J. Advances in global bioavailable strontium isoscapes. Palaeogeogr. Palaeoclimatol. Palaeoecol. 555, 109849 (2020).
Le Huray, J. D. & Schutkowski, H. Diet and social status during the La Tène period in Bohemia: Carbon and nitrogen stable isotope analysis of bone collagen from Kutná Hora-Karlov and Radovesice. J. Anthropological Archaeol. 24, 135–147 (2005).
Filipović, D. et al. New AMS 14C dates track the arrival and spread of broomcorn millet cultivation and agricultural change in prehistoric Europe. Sci. Rep. 10, 13698 (2020).
Pospieszny, Ł et al. Isotopic evidence of millet consumption in the Middle Bronze Age of East-Central Europe. J. Archaeol. Sci. 126, 105292 (2021).
Snoeck, C. et al. Calcined bone provides a reliable substrate for strontium isotope ratios as shown by an enrichment experiment. Rapid Commun. Mass Spectrom. 29, 107–114 (2015).
Frère, A. et al. Reconstructing life histories: New insights into cremation practices, mobility, and food consumption patterns through isotope and infrared analyses of petrous parts and teeth. Am. J. Biol. Anthropol. 187, e70058 (2025).
Hübler, R. et al. HOPS: automated detection and authentication of pathogen DNA in archaeological remains. Genome Biol. 20, 280 (2019).
Lemos, J. A. et al. The biology of Streptococcus mutans. Microbiol. Spectr. 7, 1 (2019).
Colombo, A. P. V. et al. Comparisons of subgingival microbial profiles of refractory periodontitis, severe periodontitis, and periodontal health using the human oral microbe identification microarray. J. Periodontol. 80, 1421–1432 (2009).
Bancerz-Kisiel, A. & Szweda, W. Yersiniosis - a zoonotic foodborne disease of relevance to public health. Ann. Agric. Environ. Med. 22, 397–402 (2015).
Schunke, T. Der Hortfund von Hohenweiden-Rockendorf, Saalkreis, und der Bronzekreis Mittels-aale. Ein Beitrag zur jungbronzezeitlichen Kulturgruppengliederung in Mitteldeutschland. Jahresschrift für mitteldeutsche Vorgeschichte 88/2004 219–338 (2004).
Schunke, T. An der westlichen Peripherie der Lausitzer Kultur: Die mittelbronze- bis früheisenzeitlichen Gräberfelder bei Coswig, Lkr. Wittenberg. In Über den Wallrand geschaut. Der Kemberger Burgwall und sein bronze- und eisenzeitliches Umfeld. Kleine Hefte zur Archäologie in Sachsen-Anhalt 15 (eds. Meller, H. & Nebelsick, L.) 271–282 (Halle/Saale, 2018).
Lißner, B. Zu den frühbronzezeitlichen Gruppenin Süddeutschland. In Leipziger online-Beiträge zur Ur- und Frühgeschichtlichen. Archäologie 13, 1–20 (2004).
Neugebauer, J. W. Bronzezeit in Ostösterreich. (St. Pölten-Wien, 1994).
Gogâltan, F. & Ailincăi, S.-C. (eds). Settlements of Life and Death. Studies from Prehistory to Middle Ages. Proceedings of an International Colloquium, Tulcea, 25th–28th of May 2016. (Editura Mega, Cluj-Napoca, 2016).
Ion, A. Why keep the old dead around?: Bringing together theory and method in the study of human remains from Balkan (E)Neolithic settlements. Doc. Praehist. 47, 348–372 (2020).
Carlin, N. et al. Social and genetic relations in Neolithic Ireland: Re-evaluating kinship. Camb. Archaeol. J. 35, 435–455 (2025).
Knipper, C. et al. Coalescing traditions-Coalescing people: community formation in Pannonia after the decline of the Roman Empire. PLoS One 15, e0231760 (2020).
Fowler, C. The Archaeology of Personhood: An Anthropological Approach. (Routledge, London, England, 2004).
Fowler, C. From Identity and Material Culture to Personhood and Materiality. (Oxford University Press, 2012).
Thomas, J. Death, identity and the body in Neolithic Britain. J. R. Anthropological Inst. 6, 653–668 (2000).
Brück, J. Death, exchange and reproduction in the British Bronze Age. Eur. J. Archaeol. 9, 73–101 (2006).
Brück, J. & Booth, T. J. The power of relics: the curation of human bone in British Bronze Age burials. Eur. J. Archaeol. 25, 440–462 (2022).
Snoeck, C. et al. Unravelling the mysteries hidden within the cremated human remains from Belgium–The interdisciplinary CRUMBEL project. in Beyond Urnfields: New Perspectives on Late Bronze Age–Early Iron Age Funerary Practices in Northwest Europe. Schriften des Museums für Archäologie Schloss Gottorf–Ergänzungsreihe, Band 16 (eds. H. A. Rose, L. C. & Louwen, A. (2023).
Sørensen, M. L. S. & Rebay-Salisbury, K. Death and the Body in Bronze Age Europe: From Inhumation to Cremation. (Cambridge University Press, Cambridge, England, 2023).
Phenice, T. W. A newly developed visual method of sexing the os pubis. Am. J. Phys. Anthropol. 30, 297–301 (1969).
Ferembach, D., Schwidetzky, I. & Stloukal, M. Recommendations for age and sex diagnoses of skeletons. J. Hum. Evol. 9, 517–549 (1980).
Buikstra, J. E. & Ubelaker, D. Standards for Data Collection from Human Skeletal Remains. Arkansas Archeological Survey (1994).
Scheuer, L., Black, S. & Schaefer, M. C. Juvenile Osteology: A Laboratory and Field Manual. (Academic Press, San Diego, CA, 2008).
Aufderheide, A. C. & Rodriguez-Martin, C. The Cambridge Encyclopedia of Human Paleopathology. Cambridge University Press: Cambridge, England, 2011.
Brickley, M. & McKinley, J. Guidelines to the Standards for Recording Human remains, IFA Payer No. 7, British Association for biological Anthropology and Osteoarchaeology. (2004).
Trautmann, B. & Harbeck, M. Anleitung zur standardisierten Skelettdokumentation in der Staatssammlung für Anthropologie und Paläoanatomie München, Staatliche naturwissenschaftliche Sammlung BayernsTeil III: Körpergräber von Kindern und Jugendlichen. (2019).
Brown, T. A., Nelson, D. E., Vogel, J. S. & Southon, J. R. Improved collagen extraction by modified Longin method. Radiocarbon 30, 171–177 (1988).
Ramsey, C. B. Bayesian Analysis of Radiocarbon Dates. Radiocarbon 51, 337–360 (2009).
Reimer, P. J. et al. The IntCal20 Northern Hemisphere Radiocarbon Age Calibration Curve (0–55 cal kBP). Radiocarbon 62, 725–757 (2020).
Deniro, M. J. Postmortem preservation and alteration of in vivo bone collagen isotope ratios in relation to palaeodietary reconstruction. Nature 317, 806–809 (1985).
van Klinken, G. J. Bone collagen quality indicators for palaeodietary and radiocarbon measurements. J. Archaeol. Sci. 26, 687–695 (1999).
Zazzo, A., Saliège, J.-F., Person, A. & Boucher, H. Radiocarbon dating of calcined bones: where does the carbon come from? Radiocarbon 51, 601–611 (2009).
Van Strydonck, M., Boudin, M. & Mulder, G. D. The carbon origin of structural carbonate in bone apatite of cremated bones. Radiocarbon 52, 578–586 (2010).
Zazzo, A., Saliège, J.-F., Lebon, M., Lepetz, S. & Moreau, C. Radiocarbon dating of calcined bones: Insights from combustion experiments under natural conditions. Radiocarbon 54, 855–866 (2012).
Snoeck, C., Brock, F. & Schulting, R. J. Carbon exchanges between bone apatite and fuels during cremation: Impact on radiocarbon dates. Radiocarbon 56, 591–602 (2014).
Wojcieszak, M., Van den Brande, T., Ligovich, G. & Boudin, M. Pretreatment protocols performed at the royal institute for Cultural Heritage (rich) prior to AMS 14c measurements. Radiocarbon 62, e14–e24 (2020).
Capuzzo, G. et al. Final Neolithic and Bronze Age funerary practices and population dynamics in Belgium, the impact of radiocarbon dating cremated bones. Radiocarbon 65, 1–30 (2023).
Boudin, M., Van Strydonck, M., van den Brande, T., Synal, H.-A. & Wacker, L. RICH – A new AMS facility at the Royal Institute for Cultural Heritage, Brussels, Belgium. Nucl. Instrum. Methods Phys. Res. B 361, 120–123 (2015).
Pinhasi, R. et al. Optimal ancient DNA yields from the inner ear part of the human petrous bone. PLoS One 10, e0129102 (2015).
Orfanou, E., Himmel, M., Aron, F. & Haak, W. Minimally-invasive sampling of pars petrosa (os temporale) for ancient DNA extraction. protocols.io https://www.protocols.io/view/minimally-invasive-sampling-of-pars-petrosa-os-tem-bqd8ms9w (2020).
Korlević, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. Biotechniques 59, 87–93 (2015).
Rohland, N., Glocke, I., Aximu-Petri, A. & Meyer, M. Extraction of highly degraded DNA from ancient bones, teeth and sediments for high-throughput sequencing. Nat. Protoc. 13, 2447–2461 (2018).
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl. Acad. Sci. Usa. 110, 15758–15763 (2013).
Velsko, I., Skourtanioti, E. & Brandt, G. Ancient DNA Extraction from Skeletal Material. protocols.io https://www.protocols.io/view/ancient-dna-extraction-from-skeletal-material-baksicwe (2020).
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res 40, e3 (2012).
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010, db.prot5448 (2010).
Gansauge, M.-T. & Meyer, M. Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat. Protoc. 8, 737–748 (2013).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil-DNA-glycosylase treatment for screening of ancient DNA. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, 20130624 (2015).
Fellows Yates, J. A. et al. Reproducible, portable, and efficient ancient genome reconstruction with nf-core/eager. PeerJ 9, e10947 (2021).
Maricic, T., Whitten, M. & Pääbo, S. Multiplexed DNA sequence capture of mitochondrial genomes using PCR products. PLoS One 5, e14004 (2010).
Rohrlach, A. B. et al. Using Y-chromosome capture enrichment to resolve haplogroup H2 shows new evidence for a two-path Neolithic expansion to Western Europe. Sci. Rep. 11, 15005 (2021).
Renaud, G., Stenzel, U. & Kelso, J. LeeHom: adaptor trimming and merging for Illumina sequencing reads. Nucleic Acids Res 42, e141 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Jun, G., Wing, M. K., Abecasis, G. R. & Kang, H. M. An efficient and scalable analysis framework for variant extraction and refinement from population-scale DNA sequence data. Genome Res 25, 918–925 (2015).
Link, V. et al. ATLAS: Analysis tools for low-depth and ancient samples. bioRxiv https://doi.org/10.1101/105346 (2017).
Kousathanas, A. et al. Inferring heterozygosity from ancient and low coverage genomes. Genetics 205, 317–332 (2017).
1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Rubinacci, S., Ribeiro, D. M., Hofmeister, R. J. & Delaneau, O. Efficient phasing and imputation of low-coverage sequencing data using large reference panels. Nat. Genet. 53, 120–126 (2021).
Sousa da Mota, B. et al. Imputation of ancient human genomes. Nat. Commun. 14, 3660 (2023).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Lamnidis, T. C. et al. Ancient Fennoscandian genomes reveal origin and spread of Siberian ancestry in Europe. Nat. Commun. 9, 5018 (2018).
Mittnik, A., Wang, C.-C., Svoboda, J. & Krause, J. A molecular approach to the sexing of the triple burial at the upper paleolithic site of dolní věstonice. PLoS One 11, e0163019 (2016).
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013).
Korneliussen, T. S., Albrechtsen, A. & Nielsen, R. ANGSD: analysis of next generation sequencing data. BMC Bioinforma. 15, 356 (2014).
Huang, Y. & Ringbauer, H. hapCon: estimating contamination of ancient genomes by copying from reference haplotypes. Bioinformatics 38, 3768–3777 (2022).
Mallick, S. et al. The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes. Sci. Data 11, 182 (2024).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet 2, e190 (2006).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19, 1655–1664 (2009).
François, O. & Jay, F. Factor analysis of ancient population genomic samples. Nat. Commun. 11, 4661 (2020).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Weissensteiner, H. et al. HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res 44, W58–W63 (2016).
Schönherr, S., Weissensteiner, H., Kronenberg, F. & Forer, L. Haplogrep 3 - an interactive haplogroup classification and analysis platform. Nucleic Acids Res 51, W263–W268 (2023).
Andrades Valtueña, A., Michel, M. & Herbig, A. MPI-EVA-Archaeogenetics/Autorun_deepscreening: 0.0.1. Zenodo https://doi.org/10.5281/ZENODO.18403831 (2026).
Herbig, A. et al. MALT: Fast alignment and analysis of metagenomic DNA sequence data applied to the Tyrolean Iceman. bioRxiv https://doi.org/10.1101/050559 (2016).
Vågene, ÅJ. et al. Salmonella enterica genomes from victims of a major sixteenth-century epidemic in Mexico. Nat. Ecol. Evol. 2, 520–528 (2018).
Huson, D. H., Auch, A. F., Qi, J. & Schuster, S. C. MEGAN analysis of metagenomic data. Genome Res 17, 377–386 (2007).
Britton, K. et al. Sampling plants and malacofauna in 87Sr/86Sr bioavailability studies: Implications for isoscape mapping and reconstructing of past mobility patterns. Front. Ecol. Evol. 8, 579473 (2020).
Leggett, S., Rose, A., Praet, E. & Le Roux, P. Multi-tissue and multi-isotope (δ13 C, δ15 N, δ18 O and 87/86 Sr) data for early medieval human and animal palaeoecology. Ecology 102, e03349 (2021).
Veselka, B. et al. Strontium isotope ratios related to childhood mobility: Revisiting sampling strategies of the calcined human pars petrosa ossis temporalis. Rapid Commun. Mass Spectrom. 35, e9038 (2021).
Weis, D. et al. High-precision isotopic characterization of USGS reference materials by TIMS and MC-ICP-MS. Geochem. Geophys. Geosyst. 7, 1-30 (2006).
Gerritzen, C. T., Goderis, S., James, H. F. & Snoeck, C. Optimizing Zr-doped MC-ICP-MS sample-standard bracketing to simultaneously determine 87Sr/86Sr and δ88Sr for high sample-throughput. Spectrochim. Acta Part B. Spectrosc. 217, 106955 (2024).
Kruskal, W. H. & Wallis, W. A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 47, 583–621 (1952).
Shapiro, S. S. & Wilk, M. B. An analysis of variance test for normality (complete samples). Biometrika 52, 591–611 (1965).
Welch, B. L. The generalization of ‘students’ problem when several different population variances are involved. Biometrika 34, 28 (1947).
Fisher, R. A. Statistical methods for research workers. In Breakthroughs in Statistics 66–70 (Springer New York, New York, NY, 1992).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57, 289–300 (1995).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. The R Project for Statistical Computing https://www.R-project.org/ (2021).
Mallick, S. & Reich, D. The Allen Ancient DNA Resource (AADR): a curated compendium of ancient human genomes, https://doi.org/10.7910/dvn/ffidcw, Harvard Dataverse (2023).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Acknowledgements
Data was generated by the Ancient DNA Core Unit of the Max Planck Institute for Evolutionary Anthropology, as well as the Ancient DNA laboratory team in Jena, both funded by the Max Planck Society. We thank E. Krukal, A. Wissgott, G. Brandt, R. Radzeviciute, S. Nagel, E. Essel, and F. Aron for their support in the ancient DNA analyses, and S. Clayton and K. Prüfer for processing the raw sequence data. We thank the teams at the former Department of Archaeogenetics at the Max Planck Institute for the Science of Human History (MPI-SHH), now part of the Max Planck Institute for Evolutionary Anthropology (MPI-EVA); the former Department of Archaeology at MPI-SHH, now Max Planck Institute of Geoanthropology (MPI-GEA); and the Population Genetics and PALEoRIDER groups, for their continued support and valuable discussions. Finally, we thank the Landesmuseum and the Landesamt für Denkmalpflege und Archäologie Sachsen-Anhalt in Halle for providing samples and granting access to field documentation. The 2004 excavations in Esperstedt were conducted by the Landesamt für Denkmalpflege und Archäologie Sachsen-Anhalt (State Heritage Department Saxony-Anhalt), which also funded the University Jena excavations in Kuckenburg. The archaeological field works in Kuckenburg were also supported by the Agrarunternehmen Barnstädt and are possible thanks to the landowner H. Holter (Esperstedt). This research was funded by the Max Planck Society and the International Max Planck Research School (IMPRS) for the Science of Human History, and the Max Planck Institute of Geoanthropology under the Independent Research Group of Domestication and Anthropogenic Evolution. This research was also funded by the European Research Council under the European Union’s Horizon 2020 research and innovation programme under grant agreement numbers 771234-PALEoRIDER (W.H.) and 851102-FEDD (R.S.), and 948913-LUMIERE (C.S.), as well as the Deutsche Forschungsgemeinschaft under grant number HA 5407/8-1 (P.E., E.P., F.S., W.H., P.R.). M.E. was supported by the Praemium Academiae Award of the Czech Academy of Sciences.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
W.H., P.R., E.P., P.E., F.N.S., R.S., and E.O. conceived and designed the study. W.H., P.R., C.S., and J.K. provided materials and resources. E.O., J.I., M.H., M.L., L.S. (Lena Semerau), and C.S. performed the laboratory experiments. E.O., A.G. and A.B.R. conducted the data analysis with guidance and input from W.H., P.R., C.S., and S.P. The departmental pipelines were maintained and implemented by L.T. and T.L. G.U.N., and A.A.V. carried out the pathogen screening. E.P. conducted the initial osteoarchaeological assessment of Kuckenburg. M.S. conducted the osteoarchaeological assessments of individuals from Esperstedt and re-analysed those from Kuckenburg, including determinations of age-at-death, anthropological sex, and palaeopathological evaluation. D.A.M., M.E., P.E., E.P., M.F., A.H., T.H., M.K., H.M., L.P., F.N.S., L.S. (Lubor Smejtek), S.A.S., T.S., P.W.S., J.W., and B.Z. provided the archaeological material, contextual site information, and expertise on the archaeological background. E.O., F.N.S., M.S., and W.H. wrote and revised the manuscript, with additional input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Andrew Millard, Catherine Frieman and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Orfanou, E., Ghalichi, A., Rohrlach, A.B. et al. Reconstruction of the lifeways of Central European Late Bronze Age communities using ancient DNA, isotope and osteoarchaeological analyses. Nat Commun 17, 1992 (2026). https://doi.org/10.1038/s41467-026-69895-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69895-y