Abstract
Fibroblast growth factor receptor (FGFR) alterations are targetable in metastatic urothelial carcinoma (mUC). Identifying FGFR alterations currently requires tissue testing, which is limited by sample availability and cancer heterogeneity. Plasma circulating tumor DNA (ctDNA) offers a complementary testing strategy, but the added value of ctDNA relative to conventional FGFR alteration assessment remains unclear. Here, in a prospective, multicentre study, we show that ctDNA testing has high concordance with tissue FGFR testing and identifies additional actionable FGFR alterations. We profile plasma from 208 patients with mUC undergoing clinical FGFR tissue testing for erdafitinib eligibility. In evaluable baseline samples, FGFR alteration frequency is 26% in either tissue or ctDNA. Among 125 patients with baseline detected ctDNA and paired tissue results, FGFR status is concordant in 90%, and ctDNA has an 84% sensitivity for tissue-detected alterations while also identifying 7 additional cases. Serial plasma collections post-baseline further clarify FGFR status. In 21 patients who received erdafitinib after testing, the median progression-free survival is 7.5 months, and one patient with a ctDNA-exclusive FGFR alteration remained on erdafitinib for 33 months. Our results support clinical uptake of ctDNA FGFR testing in combination with tissue-based approaches in mUC.
Similar content being viewed by others
Introduction
Activating alterations in the fibroblast growth factor receptor (FGFR) genes are oncogenic drivers in several cancers, including urothelial cancer where FGFR3 alterations are present in 20-60% of patients depending on stage and presentation1,2,3,4. FGFR3 alterations introduce a therapeutic vulnerability, and the pan-FGFR small-molecule inhibitor erdafitinib is an approved treatment option in metastatic urothelial cancer (mUC)5. Second-generation agents with improved FGFR3 specificity and reduced toxicity are in clinical development for mUC, and there are ongoing efforts to test FGFR inhibition earlier in disease6,7. However, a barrier to effective implementation of FGFR inhibition in mUC is the need for accurate and timely identification of FGFR3 alterations.
Constitutive FGFR3 activation in mUC commonly arises through hotspot mutations in the ligand-binding or transmembrane domains, or through gene fusions involving the intracellular domain8. In the United States, the approved companion diagnostic for erdafitinib is a qualitative RT-PCR tumor tissue test that detects nine recurrent FGFR alterations (4 hotspot mutations and 5 fusions)5. Outside the US, local diagnostic labs use a variety of tests, often amplicon-based due to cost, speed, and reliability. However, no internationally standardized testing approach exists, contributing to assay-dependent variability in patient selection and access to FGFR-targeted therapies. Reliance on single-foci, archival tissue samples for testing is a further limitation given the well-documented intra-patient heterogeneity of FGFR3 alteration status, and a tissue test failure rate of 14-34% depending on the population9,10,11,12.
To address these challenges, we designed a pan-Canadian prospective study (NCT06129084) to evaluate the potential added value of plasma-derived circulating tumor DNA (ctDNA) testing to standard tissue-based approaches for FGFR gene alteration detection in mUC. Plasma ctDNA offers a minimally-invasive, real-time snapshot of the dominant tumor population, particularly at the time of clinical disease progression, and has prognostic value based on ctDNA detectability and dynamics9,13,14,15,16,17,18. However, its role as an adjunct to tissue-based FGFR testing in mUC has not been formally assessed. The primary objective of our study therefore was to assess FGFR alteration status in patient ctDNA-tissue pairs to determine the concordance and sensitivity of each analyte. Our results support the integration of ctDNA testing into contemporary clinical workflows to improve the precision and timeliness of FGFR-targeted treatment decisions in mUC.
Results
Study cohort characteristics
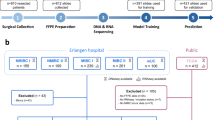
We initiated a study in patients with mUC that were under consideration for erdafitinib treatment, as per Canadian standard-of-care guidelines. Between January 2021 and April 2024, 265 patients were screened and 244 were prospectively enrolled at 12 sites across Canada; of these, 208 had both clinical-grade tumor tissue testing and at least one blood sample available for ctDNA testing using a custom targeted sequencing approach (Fig. 1a). Patients known to have no tissue available were not screened. Clinical characteristics were typical for mUC (Table 1). No patient had started FGFR-targeted therapy before or at the time of enrollment and/or initial ctDNA collection. Although erdafitinib is commonly administered in the second or third-line setting, most patients were screened at initiation of first- or second-line systemic therapy, as is common in mUC due to the requirement for rapid treatment decisions in later lines.
a Consort diagram of all patients screened. b Frequency of FGFR alterations in diagnostic tissue and baseline ctDNA samples (samples without detected ctDNA are excluded from the ctDNA denominator). Total patient number is included above the bars. c FGFR status in diagnostic tissue and ctDNA in the 125 evaluable tissue-ctDNA sample pairs. Each patient is represented as a column. ctDNA and tissue sensitivity are calculated against patient alteration status. d Confusion matrix of FGFR status in the 125 evaluable tissue-ctDNA sample pairs. e Clinical features of patients with FGFR alterations compared to wild-type (WT) cases. Two-sided Fisher’s exact tests were used to calculate p-values; no adjustments for multiple comparisons were made. Cases with missing clinical data are excluded. Box and whisker plots display median, interquartile range (IQR), and IQR ± 1.5*IQR. f Overall survival Kaplan-Meier model stratified by ctDNA FGFR status. The ctDNA detected group used for the Cox proportional hazard ratio modeling consists of both FGFR altered and wild-type. g Overall survival Kaplan-Meier model stratified by tissue FGFR status. Only patients with clinical follow-up and conclusive tissue testing were included (n = 197). Hazard ratios and p-values in (f, g) were calculated using Cox proportional hazards and two-sided log-rank tests, respectively, with no multiple comparison correction applied.
FGFR alteration status via tissue testing
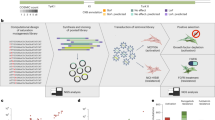
For each patient, formalin-fixed paraffin-embedded (FFPE) tissue from routine clinical procedures was submitted for clinical diagnostic standard-of-care FGFR testing, with no restrictions on tissue source (Supplementary Data 1–2). Most tissue (61%, 127/208) was centrally tested using the amplicon-based Oncomine Focus assay. An additional 27% (56/208) of tissues were screened with amplicon-based tests, at the enrolling site, or via interventional trial screening (NCT03390504, NCT04197986, NCT05614739). The remaining 12% (25/208) of tissues were tested via other clinically-approved approaches (Supplementary Data 1). Tumor tissue source was reported for 66% of patients (138/208), with 75% (103/138) obtained from transurethral resection or radical surgery and 20% (27/138) obtained via biopsy or surgical removal of distant metastases (Table 1). The median time between tissue collection and study enrollment was 296 days (IQR: 115–589 days). Tissue testing was partially successful in 12/208 cases (10 failed RNA fusion detection; 2 failed DNA mutation detection).
Activating FGFR3 gene alterations were detected in 52 patients via clinical tissue testing (26.4%, 52/197; excluding the eleven patients with partially successful testing and no alterations detected) (Fig. 1b). As expected, FGFR3 hotspot mutations were more common than gene fusions (19.4% vs 6.6%), with S249C and FGFR3-TACC3 most frequent in each class, respectively (Supplementary Fig. S1a; Supplementary Data 1). The only oncogenic alteration outside FGFR3 was a single FGFR2-CASP3 fusion. The specific test used for clinical-grade tissue testing was not associated with the rate of FGFR alteration detection (Supplementary Data 2).
To corroborate clinical sequencing results, we re-retrieved archival tissue blocks and performed an additional round of targeted sequencing on tumor tissue from 81 patients using the same custom research-based approach as applied to cell-free DNA (cfDNA; see “Methods”). 75 of these 81 patients had conclusive clinical diagnostic testing results (Supplementary Data 1). Overall, 71 of 75 (95%) patients had concordant FGFR alteration status across the research and diagnostic testing approaches (Supplementary Fig. S1b). Two FGFR3 S249C mutations and one fusion were detected only via the research approach, while one fusion was detected only via clinical testing. Interestingly, for the two mutation-discordant cases, research hybrid-capture sequencing revealed somatic in-phase intronic variants within 21 bp of the FGFR3 S249C mutation (Supplementary Fig. S1c). These variants are predicted to interfere with primer binding for the amplicon-based clinical tests, inhibiting amplification (and subsequent detection) of the mutant allele. These data suggest that amplicon-based clinical tissue testing approaches miss oncogenic FGFR3 alterations in a small subset of tumor samples.
FGFR alteration status via ctDNA testing
In total, 208 enrolled patients provided 319 blood samples in the mUC setting for cfDNA analysis. Participating sites were encouraged to collect samples while patients were therapy-naive or at times of clinical disease progression, but samples collected while patients were receiving first- or second-line systemic treatment were permitted in order to maintain trial feasibility. Serial blood samples were optional and requested at times of treatment change. cfDNA and patient-matched leukocyte DNA was subjected to targeted sequencing using a custom panel that included FGFR1-4 and intronic regions of common breakpoints in TACC3 and FGFR3 (see “Methods”). Samples which failed targeted sequencing or did not have detected ctDNA were categorized as unevaluable. Across all samples, median cfDNA sequencing depth was 1639x (Supplementary Data 3) and ctDNA was detected in 58% (185/319; ctDNA fraction range 0.5–88%). The relatively low ctDNA detection rate compared to prior studies was due to inclusion of on-treatment samples, where ctDNA is commonly suppressed13,19. Indeed, ctDNA was detected in 23.6% (17/72) of on-treatment samples compared to 72% (118/164) of samples from treatment-naive or clinically progressing mUC (p = 1.4 × 10−11 Fisher’s Exact, Supplementary Fig. S2a). Collection timing relative to treatment was unknown for 70 samples.
To standardize analyses, the first blood sample per patient was used to define baseline FGFR alteration status, even if the sample was taken at a sub-optimal collection time point for ctDNA detection. Among these, 63% (130/208) had ctDNA detected and were evaluable for FGFR alterations (median ctDNA fraction when detected: 8.4%; Supplementary Fig. S2b, Fig. 1a). FGFR alterations were detected in 26% of evaluable baseline ctDNA samples (34/130), with a similar frequency of mutations and fusions as per tumor tissue (Fig. 1a, b; Supplementary Fig. S1a; Supplementary Data 4, 5). Three FGFR3 mutations were detected at less than 1% VAF, using reduced thresholds for hotspot mutations (Methods). Six coding variants of unknown significance were identified in FGFR2/3/4. There was no difference in ctDNA fraction in FGFR altered or wild-type patients defined by ctDNA or tissue sequencing (Supplementary Fig. S2c).
Concordance between tissue and ctDNA testing
At baseline, 125 patients had tissue test results and evaluable ctDNA-detected samples (i.e., evaluable pairs) to assess concordance. FGFR status (altered or wild-type) was concordant in 90% (113/125; Fig. 1c, d, Supplementary Fig. S2d, e). ctDNA testing identified 7 patients with FGFR3 alterations that were reported as wild-type by tissue testing (5 centrally tested with the Oncomine platform, 2 via external laboratories): representing 18% of the 39 FGFR altered patients in the 125 with evaluable paired samples (and 12% (7/59) of FGFR altered patients across the entire cohort). In cases where an alteration was detected, mutation status was more frequently concordant between ctDNA and tissue (82%, 23/28) than fusion status (25%, 3/12). 5 FGFR3 fusions identified by tissue testing were undetected in evaluable ctDNA samples, encompassing 13% (5/39) of FGFR alterations in the 125 patients (Fig. 1c). There was no read support in ctDNA for 4/5 of these fusions despite using multiple breakpoint detection tools and sufficient ctDNA (median ctDNA fraction 0.22), raising the possibility that archival tumor tissue was not representative of the dominant metastatic genotype in these cases. The remaining fusion did have subthreshold support in the context of low ctDNA fraction (i.e., near the limit of detection).
Sensitivity for ctDNA testing to identify FGFR alterations reported through archival clinical tissue testing was 84% (27/32) among evaluable pairs (Fig. 1d, Supplementary Fig. S2f). Conversely, if ctDNA results were considered the reference, tissue testing had a sensitivity of 79% (27/34) due to the seven patients identified with ctDNA testing alone (Supplementary Fig. S2f). When taking the union of FGFR alterations identified by either method (n = 39/125), ctDNA sensitivity rose to 87% (34/39) and tissue sensitivity to 82% (32/39).
Among the 72 patients with tissue test results but inconclusive ctDNA results (due to undetected ctDNA), there were a further 20 (27.8%) patients with FGFR alterations (Supplementary Fig. S2d, e). While this may dampen enthusiasm for ctDNA testing as a sole replacement for tissue testing, it is important to note that sample collection timing was not always optimal for maximizing ctDNA fraction. Notably, when considering only the samples collected prior to treatment or at time of progression (i.e., optimal collections), the proportion of evaluable samples increased to 74% and sensitivity for detection of FGFR alterations identified via tissue testing was 88% (21/24, Supplementary Fig. S2b, f–g). Furthermore, our testing approach employed a conservative detection threshold of 0.5% VAF. Sequencing reads supporting tissue-detected FGFR mutations were observed in 7/16 “inconclusive” ctDNA samples, suggesting presence below our conservative limit of detection.
Patient clinical characteristics were broadly comparable by FGFR status, although more patients with FGFR alterations presented with upper tract disease, as expected20 (Fig. 1e, Table 1, Supplementary Data 6). We observed no significant difference in overall survival (OS) from metastatic diagnosis based on FGFR alteration status in ctDNA or tissue (Fig. 1f, g). Patients with undetected ctDNA had the best outcomes (Fig. 1f), as previously reported21,22,23.
Longitudinal analysis of FGFR alterations by serial ctDNA testing
A key advantage of ctDNA is minimally-invasive serial sampling to reevaluate inconclusive results, monitor temporal heterogeneity, and detect treatment-emergent resistance alterations. In this study, 52 of 67 (78%) patients who provided serial plasma samples had ctDNA detected in at least one sample (Supplementary Fig. S3a). Serial samples collected while disease was progressing on systemic therapy were more likely to have ctDNA detected than those collected during therapy response (69% vs 33%, p = 0.0003; Supplementary Fig. S3b). Among the 52 patients, FGFR alterations were identified in 14 patients (Fig. 2a). Of the 29 patients with initially inconclusive FGFR status due to undetected ctDNA but who provided serial collections, 48% (14/29) had detected ctDNA in a subsequent sample, including 8 patients whose initial sample was collected while treatment naive/progressing. Notably, 4 of these 14 (29%) were found to have FGFR alterations upon resampling, all of which were concordant with tissue. These results demonstrate that repeat ctDNA testing can uncover clinically-relevant FGFR alterations in the event of earlier samples with no or low ctDNA fraction.
a FGFR status in patients with multiple ctDNA collections and an FGFR alteration identified in at least one ctDNA sample. Samples were grouped by patient and ordered chronologically. All patients with multiple samples can be found in Fig. S4. b Clinical timeline of a patient with heterogeneous ctDNA FGFR3 status. Bar plots provide the variant allele frequency (VAF) of all somatic alterations detected across 1 tissue and 3 ctDNA collections, indicated on the timeline as a tissue block or blood drop, respectively. Yellow bars are the FGFR3 p.S249C hotspot mutation while gray bars are other protein-altering mutations. This patient had a good response to initial platinum/gemcitabine and was therefore re-treated with platinum/gemcitabine followed by maintenance avelumab before receiving single agent enfortumab vedotin at progression. Unfortunately, their clinical condition did not allow for erdafitinib after progression on enfortumab vedotin. The blood drop is Public Domain, and we thank the artist for their work.
Among patients with FGFR alterations identified in at least one ctDNA sample and multiple collections with ctDNA detected, alterations were consistently detected across timepoints in 4/5 cases. In the remaining patient, an FGFR3 S249C mutation detected in the first ctDNA sample was not detected in 2 subsequent samples (Fig. 2b). Interestingly, clinical tissue testing of an upper tract biopsy identified an FGFR3-TACC3 fusion in the same patient, which was not detected in any ctDNA sample, suggestive of FGFR spatiotemporal heterogeneity.
Outcomes and genomic resistance in erdafitinib-treated patients
Twenty-one patients had received erdafitinib at data cut-off, with additional FGFR altered patients eligible for erdafitinib yet to progress on their current line of treatment (Fig. 3a; Table 2). Median time to disease progression or toxicity was 6.0 months (95% CI 2.7–7.8 months; median PFS 7.5 months). Treatment cessation due to toxicity mostly occurred within 4 months of treatment start. Median OS from the start of erdafitinib was 10.6 months (95% CI: 6.0–16.3 months). 6 patients achieved a complete or partial response, and 8 had stable disease (disease control rate = 67%). Erdafitinib was discontinued due to toxicity in 6 (29%) patients, while 3 patients (14%) remained on treatment at the data cutoff.
a Swimmer’s plot of all patients who received erdafitinib by data cut-off (n = 21). Tissue and ctDNA FGFR status are indicated on the left. Best response, discontinuation reason, and patient status at last follow-up are shown. 19–345 and 18–470 were not considered FGFR altered via baseline tissue and ctDNA and were only identified as FGFR altered via serial sampling. Cumulative incidence function, accounting for progression-free survival (PFS) and discontinuation due to toxicity, is inset. Patients with ongoing treatment are indicated by tick marks along the x-axis. b Oncoprint displaying metastatic urothelial cancer (mUC) driver genomic alterations in plasma sample pairs for all patients with a ctDNA-detected sample collected at progression on erdafitinib (n = 11). For 2 patients, the corresponding ctDNA results provided are from a follow-up plasma sample (serial sample) because the baseline sample was inconclusive, i.e., no ctDNA detected. ctDNA FGFR alterations are shown, along with all genes covered by the targeted sequencing approach with at least one alteration detected in a progression sample. c A proposed workflow for FGFR testing incorporating upfront ctDNA testing, with reflex tissue testing when plasma samples are inconclusive due to undetected ctDNA (i.e., ctDNA-). The blood drop is Public Domain, and we thank the artist for their work. 1 L = first line; 2 L = second line. d Theoretical sensitivity of tissue-only testing (n = 198) versus the proposed workflow with upfront ctDNA reflex testing approach using data from 202 patients with conclusive ctDNA or tissue results.
Notably, 1 patient classified as FGFR altered solely by ctDNA testing experienced an exceptionally durable clinical benefit, remaining on therapy for 33 months. This case highlights the potential clinical utility of ctDNA-identified FGFR alterations that may be missed by tissue testing.
Thirteen patients provided a blood sample at the time of erdafitinib progression. ctDNA was detected in 11 of 13 (85%) samples (Fig. 3b). The original FGFR alterations were re-detected in all 11 progression ctDNA samples, consistent with prior work examining samples at erdafitinib progression9,24. In two patients, additional putative resistance alterations in FGFR3 emerged. Both cases had known resistance-associated “gatekeeper” mutations24. In one patient, 20-383, three distinct base substitutions, all resulting in V555M or V555L gatekeeper mutations, were observed at varying allele frequencies: suggesting the presence of polyclonal resistance of unequal size. While on erdafitinib therapy, this patient also acquired an FGFR3-TACC3 fusion and an FGFR3 N540K “molecular brake” mutation, which has been previously implicated in resistance to FGFR inhibition (Supplementary Fig. S4)24.
Discussion
In this prospective, multicenter study, we evaluated the utility of ctDNA testing for identifying actionable FGFR alterations (predominantly FGFR3 mutations and fusions) in mUC, using standard archival tumor tissue testing as the reference. Among patients with evaluable results from both platforms, FGFR status was concordant in 90% of cases, and ctDNA testing identified clinically actionable FGFR3 gene alterations in 7 additional patients that were not detected in tissue. Serial ctDNA testing also enabled reassessment of initially inconclusive cases and captured mechanisms of resistance at the time of progression on FGFR-targeted therapy. Our findings support ctDNA as a complementary tool for informing FGFR-directed treatment strategies in mUC.
A key limitation of ctDNA-based testing is that a subset of patients with low or undetected ctDNA levels cannot be evaluated for FGFR alterations, particularly when blood samples are collected during treatment response13,25. In our cohort, ctDNA was undetected in ~27–42% of baseline samples, depending on collection timing; this limitation is likely partly biological, reflecting the fact that cancers have variable ctDNA shedding dynamics26, but also assay-dependent. Our ctDNA assay employed a relatively conservative detection threshold for FGFR3 mutations (0.5% VAF; see “Methods”), prioritizing specificity and cost practicalities. Contemporary commercial ctDNA genotyping platforms (e.g., offered by Foundation Medicine, Tempus, Guardant) have limits of detection between 0.2% and 0.8% VAF depending on the mutation, but reducing sequencing costs and improved error suppression mean that these limits are regularly improving27. In future, ctDNA platforms with improved sensitivity at low tumor fractions may reduce the rate of non-evaluable samples and broaden the applicability of a mUC ctDNA genotyping approach (although ultra-sensitive tumor-informed ctDNA detection platforms are not appropriate for mUC FGFR genotyping). Importantly, our study design included only patients with available archival tissue, which inherently biases the non-evaluable category toward ctDNA. In real-world practice, where tissue may be unavailable or inadequate, ctDNA testing could offer greater incremental value than observed here. In a recent real-world study of over 3600 patients with tumor tissue FGFR testing, failure rates were 14% and 34% for mUC and non-muscle-invasive bladder cancer, respectively12. Notably, we did not observe a difference in FGFR alteration frequency or distribution between patients with and without detected ctDNA, indicating that our study was not biased towards a specific FGFR status. However, ultimately, it remains unclear whether excluding patients without tumor tissue introduced a biological or clinical bias, and this is a key limitation of our study.
We deliberately focused our primary analysis on patients with both successful tissue testing and detected ctDNA to provide the most controlled, interpretable comparison between testing approaches. Within this subset, ctDNA identified additional FGFR alterations that were not detected by tissue, including in one patient who experienced a 33-month durable response to erdafitinib. Our findings align with prior reports of spatial and temporal heterogeneity in FGFR alterations. For example, Clinton et al. found that 9% of FGFR alterations were detected exclusively in metastatic tissue and not in the primary tumor, while Guercio et al. reported a 26% discordance between primary and metastatic tumor FGFR status9,11. In 2 discordant cases in our cohort, mutations detected in ctDNA were not reported by amplicon-based tissue testing likely due to adjacent intronic variants predicted to disrupt primer binding, highlighting that technical aspects of assay design can also contribute to false negative results. Taken together, our data supports the biological and clinical validity of ctDNA-detected alterations and highlights potential for ctDNA to expand access to FGFR-targeted therapy. Given that archival tissue is often limited in quantity, of poor quality, or unrepresentative of the metastatic genotype, ctDNA may be particularly valuable when rapid treatment decisions are needed or repeat biopsy is infeasible.
The dynamic nature of ctDNA sampling also enabled assessment of temporal heterogeneity and treatment-emergent resistance. In our study, serial blood samples especially those collected at systemic therapy progression converted a substantial number of initially inconclusive cases to evaluable (i.e., ctDNA detected) and revealed FGFR3 alterations not detected at baseline. Moreover, in patients progressing on erdafitinib, ctDNA re-identified the original FGFR alteration and uncovered additional putative resistance mechanisms such as FGFR3 gatekeeper mutations (V555M/L) and molecular brake mutations (N540K): as previously reported and validated in several recent studies9,24. In 1 case, we observed clear evidence of polyclonal resistance, suggesting continued pathway dependence. Interestingly, we observed on-treatment emergence of an FGFR3-TACC3 fusion in the same case; this fusion typically qualifies patients for FGFR inhibition and to our knowledge, an emergent fusion on erdafitinib has not previously been described, and should be investigated as a potential resistance mechanism.
Collectively, the treatment-emergent FGFR alterations align with observations from other cancers where targeted therapies can be associated with acquired genomic alterations (e.g., EGFR T790M mutations following treatment with first generation EGFR tyrosine kinase inhibitors)28,29. These on-target resistance alterations can signal dependency and have been associated with good initial response and continued pathway reliance30,31. Continued FGFR dependency is particularly relevant in the context of next-generation FGFR inhibitors currently in development including more selective FGFR3-targeted agents (e.g., TYRA-300, LY3866288)32,33. Analogous to the use of osimertinib in EGFR-mutant non–small cell lung cancer following the development of T790M mutations, patients progressing on erdafitinib with acquired FGFR3 resistance mutations may represent an enriched population for subsequent FGFR-targeted therapies.
Our results reinforce prior retrospective analyses and provide prospective evidence supporting integration of ctDNA into FGFR biomarker testing workflows for mUC24,34. At present, it is most likely that ctDNA testing is considered as a back-up or repeat testing option for scenarios where tissue testing fails or is inappropriate. However, an option for clinical integration of ctDNA in patients with treatment-naive mUC is as an upfront testing approach, with reflex tissue testing in case of inconclusive ctDNA results, as was also proposed by the European Society for Medical Oncology recommendations for ctDNA testing (Fig. 3c, d)35. While previous studies have shown a degree of concordance between plasma and tissue FGFR testing9,13,24,34,36, our study is the largest prospective evaluation to date, implemented across multiple centers and using harmonized clinical data and centralized molecular profiling. Importantly, our study incorporated both diagnostic and research sequencing platforms, allowing for orthogonal validation of tissue results and enhanced confidence in ctDNA findings.
Our study has several other limitations. Firstly, heterogeneity in tissue testing methods across participating centers may have influenced tissue-based detection rates, although this was mitigated by central testing and orthogonal research sequencing in a large subset of cases. Given the limited genomic information obtained via clinical testing we also were unable to explore FGFR3 mutation (sub)clonality as a variable influencing spatio-temporal concordance. Additionally, while our cohort included patients eligible for erdafitinib, only a limited number received the drug to date, and thus our findings on treatment response and resistance are exploratory. The progression-free survival in our study was 7.5 months, higher than the 5.6 reported in the erdafitinib registration trial5, possibly because 43% of patients in our cohort received erdafitinib as second-line rather than third-line therapy. Finally, for our overall tested population certain clinical variables such as metastatic volume were not collected, limiting our ability to relate clinical features to FGFR or ctDNA status.
In conclusion, ctDNA FGFR testing showed high concordance with archival tumor tissue and identified actionable FGFR alterations not captured by tissue analysis in a clinically meaningful subset of patients. Our data support the use of ctDNA as a complementary tool to tissue testing in the management of mUC.
Methods
The study was conducted in accordance with the Declaration of Helsinki, and written informed consent was obtained from all participants prior to enrollment. Study approval was granted by the BC Cancer or University of British Columbia Research Ethics Board (REB #H20-02794, #H23-02533, #H18-00488), the Western University Health Sciences Research Ethics Board (#121410), the Ottawa Health Science Network Research Ethics Board (#20220431-01H), Le Comité d'éthique de la recherche du CHU de Québec-Université Laval (#2022-5986), the Health Research Ethics Board of Alberta Cancer Committee (HREBA.CC-21-0110), the University Health Network Research Ethics Board (#21-5393), and the Health Research Ethics Board of Alberta Cancer Committee (HREBA.CC-22-0160).
Study cohort and eligibility
We conducted this prospective, multicentre study at 12 sites across Canada (Supplementary Data 7).
Eligible patients had mUC and were candidates for erdafitinib treatment per standard of care practices. In Canada, FGFR-targeted therapy is predominantly administered as second- or subsequent-line systemic treatment, although diagnostic FGFR testing may be ordered earlier at the discretion of the treating physician. Patients known to have no available or accessible tissue samples prior to intention to test were excluded. Patients were required to have tissue-based testing attempted and to have provided at least one blood sample in the mUC setting for plasma cell-free DNA (cfDNA) testing. Blood samples collected or analyzed prior to study enrollment were permitted. When multiple blood samples were available, the first sample collected in the mUC setting was used as the representative sample. For patients treated with erdafitinib, an additional blood sample was collected at progression as determined by the treating physician.
Clinical data were abstracted from electronic medical records and managed using Research Electronic Data Capture (REDCap37). Metastatic volume was not collected.
Tissue FGFR clinical diagnostic testing
FFPE tissue from routine clinical procedures was submitted for diagnostic standard of care FGFR testing, with no restrictions on tissue source (Supplementary Data 1, 2). Local clinical-grade FGFR testing and tissue test results obtained prior to study enrollment were permitted. Otherwise, central clinical-grade prospective FGFR testing was offered at Lady Davis Institute for Medical Research, Jewish General Hospital, Montreal, Quebec, Canada using the Oncomine Focus Assay (Ion Torrent platform). Briefly, total nucleic acids were extracted using the Promega Maxwell® system according to the manufacturer’s protocol. DNA and RNA concentrations were quantified by TaqMan quantitative PCR. Libraries were prepared from 10 ng of input DNA or RNA using the Ion Chef™ System with DL8 kits. Next-generation sequencing (NGS) was performed on the Ion GeneStudio™ S5 Prime system, and data analysis was conducted using the Ion Reporter™ software platform.
Patients were considered FGFR altered by tissue testing if oncogenic activating mutations or fusions in FGFR2/3 were reported. Qualifying alterations included FGFR3 mutations (R248C, S249C, G370C, Y373C) and FGFR gene fusions (FGFR3-TACC3, FGFR3-BAIAP2L1, FGFR2-BICC1, FGFR2-CASP7). If tissue testing returned a partial failure (either for the DNA (mutation) or RNA (fusion) sequencing component), patients were either considered FGFR altered if an FGFR alteration was identified in the successful modality or inconclusive if no FGFR alteration was reported. Mutations in FGFR1-4 were evaluated for oncogenicity using COSMIC, ClinVar, OncoKB, and CancerHotspot databases.
Plasma circulating tumor DNA testing
Whole blood samples were collected in 2–3 × 10 ml Streck cfDNA collection tubes and shipped for central processing. Participating sites were encouraged to collect samples while patients were therapy-naive or progressing, but on-treatment samples were allowed. Serial samples were optional and requested at time of treatment change. Blood processing, DNA extraction and library preparation was performed as previously described13. In summary, whole blood was collected in 3 × 9 mL Streck Cell-Free DNA BCT® tubes. Blood collected in Streck tubes was kept at room temperature prior to and during processing where samples were centrifuged at 1600 r.c.f for 15 min, after which plasma was transferred to a new tube and spun for an additional 10 min at 3200 r.c.f. Aliquots of buffy coat (leukocytes for germline DNA; gDNA) and plasma were obtained simultaneously and stored at −80 °C prior to DNA extraction. Plasma cfDNA was extracted from up to 6 mL of input with the QIAGEN Circulating Nucleic Acids Kit, and quantified with the Quantus Fluorometer and QuantiFluor ONE dsDNA system or Qubit 2.0 Fluorometer and Qubit dsDNA HS Assay Kit. Matched gDNA was extracted from the buffy coat fraction/PBMC using the QIAGEN DNeasy Blood and Tissue Kit, or the Promega Maxwell® RSC Blood DNA Kit and Promega Maxwell® RSC Instrument. Extracted gDNA was quantified with a NanoDrop spectrophotometer. cfDNA libraries were generated using the Roche KAPA HyperPrep Kit, for gDNA libraries the Roche KAPA HyperPlus Kit was used. We aimed for 25 ng input DNA for cfDNA and 50 ng for gDNA; however, library preparation was attempted with as little as 10 ng per sample if necessary. Sample processing, sequencing, and bioinformatics were performed centrally at the Vancouver Prostate Centre, Canada.
We performed targeted hybridization-capture sequencing on plasma cfDNA and patient-matched leukocyte DNA using one of five sequential versions of a custom panel (Roche) covering the exonic regions of 50-60 bladder cancer driver genes, including FGFR1-4 and intronic regions of common breakpoints in TACC3 and FGFR3 (see coverage details in Supplementary Fig. S5 and Supplementary Data 2). Earlier versions of the panel did not cover FGFR2 or FGFR4. Later panel versions included a low-pass whole-genome backbone of regularly spaced probes capturing heterozygous germline SNPs of common frequencies across various ancestral backgrounds (IDT xGen Human CNV Backbone), to aid in tumor fraction estimation and chromosome arm copy number calling. Libraries were sequenced on Illumina instruments. Sequencing alignment, quality control, detection of variants (mutations, copy number alterations, and structural rearrangements), and tumor fraction estimation followed our published approaches13,38,39,40. Briefly, tumor fraction estimation used two orthogonal approaches, first using genome-wide copy number and heterozygous germline SNP allele frequency changes to estimate tumor fraction and ploidy, in addition to tumor fraction estimation using somatic mutation allele frequency. In samples without somatic alterations passing detection thresholds (lower limit of variant detection corresponding to ~1% ctDNA), ctDNA was considered undetected. Germline variants and those related to clonal hematopoiesis were excluded using patient-matched leukocyte DNA.
Plasma samples were first categorized as inconclusive (i.e., non-evaluable), defined by no detected ctDNA or failed sequencing. Among samples with detected ctDNA, FGFR alteration status was assigned based on the presence of oncogenic variants and compared to tissue results. Samples were considered FGFR alteration-positive if an oncogenic variant was detected. Mutations within FGFR1-4 were considered oncogenic based on ClinVar annotations of “Pathogenic” or “Likely Pathogenic”. Detection required a minimum of 8 independent supporting sequencing reads and a variant allele frequency (VAF) of above 1%, or a VAF of ≥0.5%, with at least 4 mutant reads for known FGFR3 hotspot mutations (R248C, S249C, G370C, Y373C). Thresholds were selected based on 100% specificity in 25 healthy volunteer cfDNA samples41 and were based on our published methods22,41. FGFR3-TACC3 gene fusions with at least 4 supporting split unaligned reads were inferred by structural rearrangement breakpoint analysis and were considered oncogenic if the open reading frame was predicted to be maintained. Other FGFR gene fusions were considered oncogenic if previously annotated as such, or if the fusion partner included a coiled-coil dimerization domain (analogous to TACC3) and the open reading frame was maintained. In patients with structural variants identified in at least one diagnostic tissue or baseline ctDNA sample, GRIDSS, GRIPSS, and AnnotSV were also run using default parameters on the cfDNA sequencing data42,43. Treating physicians were notified of positive ctDNA FGFR screening results and were able to prescribe erdafitinib based on ctDNA alone.
Tissue correlative research testing
For patients enrolled at sites in British Columbia, we re-retrieved archival tissue after clinical testing to perform orthogonal FGFR testing using a laboratory-developed research-use only assay (the same targeted sequencing approach as described for plasma cfDNA). When possible, the same blocks used for clinical testing were retrieved. Tumor regions were reviewed and marked by a board-certified pathologist (G.W.). Nucleic acids were extracted from pooled cores taken from the same FFPE tissue block (Maxwell® RSC RNA/DNA FFPE Kits) followed by RNAse treatment and DNA quantification. DNA was processed and libraries were constructed using the NEBNext UltraShear FFPE DNA Library Kit according to the manufacturer’s protocol; 100 ng DNA was used as input.
Outcomes and statistical analysis
The prespecified enrollment target was 210 patients with tissue and plasma cfDNA collection. Prior work suggested 15% of samples (n = 20) collected at time of clinical mUC progression would have undetected ctDNA with the thresholds of our test, and an FGFR alteration rate of 20% among ctDNA detected samples: resulting in 35 patients with ctDNA detected and FGFR alterations identified9,13.
The primary objective was to assess the diagnostic performance of ctDNA testing relative to standard-of-care tumor tissue testing for the detection of FGFR gene alterations in mUC. Diagnostic performance was evaluated through concordance analysis and sensitivity of ctDNA testing for detecting FGFR alterations identified in tissue. Concordance of FGFR alteration status (categorized as altered or wild-type) was assessed among patients with evaluable results from both tissue testing and ctDNA analysis.
Secondary objectives included determining whether ctDNA testing could identify actionable FGFR alterations not detected by tissue testing, including through analysis of serial blood samples, and evaluating ctDNA at the time of disease progression in patients treated with FGFR inhibitors to investigate potential mechanisms of acquired resistance. FGFR3 alterations that emerged at progression on erdafitinib but were absent at baseline were considered candidate resistance mechanisms.
Python (v3.12.8) and scipy (v1.15.1) were used for most statistical analysis. All tests were two-sided with p < 0.05 considered significant. Lifelines (v0.30.0) was used for Kaplan-Meier visualization and Cox proportional hazard models; scikit-survival (v0.25.0) was used for cumulative incidence functions. Graphical figures were made using matplotlib (v3.10.0). Overall survival (OS) was defined as the time from metastatic diagnosis to death from any cause. Patients without documented events were censored at the date of the last follow-up. Progression-free survival (PFS) on erdafitinib was calculated from erdafitinib initiation to disease progression or death. Cumulative incidence functions were used to show PFS, due to the high percentage of toxicity-related discontinuations. OS on erdafitinib was calculated from erdafitinib therapy initiation to death.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
De-identified targeted DNA sequencing data produced using the research assays have been deposited in the European Genome Archive (EGA) and can be accessed under accession number: EGAS50000001450. Controlled release is guided by the following data access policy: https://ega-archive.org/datasets/EGAD50000002089. We aim to complete data access approval and release within 1 month of the initial request. Unprocessed sequencing data generated during clinical-grade tissue testing is not available due to privacy laws. All other data supporting the findings of this study are available within the article and Supplementary files. Source data are provided with this paper.
Code availability
Custom computer code utilized for analysis is available on Github at https://github.com/wyatt-lab/fgfr_ctdna_tissue/44. Custom python code for downstream figure generation and data processing is not included.
References
van Rhijn, B. W. G. et al. FGFR3 and P53 characterize alternative genetic pathways in the pathogenesis of urothelial cell carcinoma. Cancer Res. 64, 1911–1914 (2004).
Pouessel, D. et al. Tumor heterogeneity of fibroblast growth factor receptor 3 (FGFR3) mutations in invasive bladder cancer: implications for perioperative anti-FGFR3 treatment. Ann. Oncol. 27, 1311–1316 (2016).
Knowles, M. A. FGFR3—a central player in bladder cancer pathogenesis? Bladder Cancer 6 403–423 (2020).
Dyrskjøt, L. et al. Bladder cancer. Nat. Rev. Dis. Prim. 9, 1–21 (2023).
Loriot, Y. et al. Erdafitinib or chemotherapy in advanced or metastatic urothelial carcinoma. N. Engl. J. Med. 389, 1961–1971 (2023).
Iyer, G. et al. A first-in-human phase 1 study of LY3866288 (LOXO-435), a potent, highly isoform-selective FGFR3 inhibitor (FGFR3i) in advanced solid tumors with FGFR3 alterations: initial results from FORAGER-1. J. Clin. Oncol. 43, 662–662 (2025).
Loriot, Y. et al. TYRA-300, an oral, FGFR3-selective inhibitor: preliminary pharmacokinetic and pharmacodynamic analysis from SURF301, the multicenter open-label phase 1/2 study of TYRA-300 in advanced urothelial carcinoma and other solid tumors with activating FGFR3 alterations. Eur. J. Cancer 211, 114599 (2024). 72 (PB060).
Krook, M. A. et al. Fibroblast growth factor receptors in cancer: genetic alterations, diagnostics, therapeutic targets and mechanisms of resistance. Br. J. Cancer 124, 880–892 (2021).
Clinton, T. N. et al. Genomic heterogeneity as a barrier to precision oncology in urothelial cancer. Cell Rep. 41, 111859 (2022).
Bacon, J. V. W. et al. Somatic features of response and relapse in non-muscle-invasive bladder cancer treated with Bacillus Calmette-Guérin immunotherapy. Eur. Urol. Oncol. 5, 677–686 (2022).
Guercio, B. J. et al. Clinical and genomic landscape of FGFR3-altered urothelial carcinoma and treatment outcomes with erdafitinib: a real-world experience. Clin. Cancer Res. 29, 4586–4595 (2023).
Matsubara, N. et al. Fibroblast growth factor receptor alteration testing for >3600 Patients with locally advanced/metastatic urothelial cancer and non-muscle-invasive bladder cancer: an analysis of the Global ANNAR Biomarker Study. Eur. Urol. Oncol. 8, 1558–1565 (2025).
Vandekerkhove, G. et al. Plasma ctDNA is a tumor tissue surrogate and enables clinical-genomic stratification of metastatic bladder cancer. Nat. Commun. 12, 184 (2021).
Birkenkamp-Demtröder, K. et al. Monitoring treatment response and metastatic relapse in advanced bladder cancer by liquid biopsy analysis. Eur. Urol. 73, 535–540 (2018).
Vandekerkhove, G. et al. Circulating tumor DNA reveals clinically actionable somatic genome of metastatic bladder cancer. Clin. Cancer Res. 23, 6487–6497 (2017).
Christensen, E. et al. Liquid biopsy analysis of FGFR3 and PIK3CA hotspot mutations for disease surveillance in bladder cancer. Eur. Urol. 71, 961–969 (2017).
Christensen, E. et al. Cell-free urine and plasma DNA mutational analysis predicts neoadjuvant chemotherapy response and outcome in patients with muscle-invasive bladder cancer. Clin. Cancer Res. 29, 1582 (2023).
Fonseca, N. M. et al. Prediction of plasma ctDNA fraction and prognostic implications of liquid biopsy in advanced prostate cancer. Nat. Commun. 15, 1828 (2024).
Tolmeijer, S. H. et al. Early on-treatment circulating tumor DNA measurements and response to immune checkpoint inhibitors in advanced urothelial cancer. Eur. Urol. Oncol. 7, 282–291 (2024).
Sfakianos, J. P. et al. Genomic characterization of upper tract urothelial carcinoma. Eur. Urol. 68, 970–977 (2015).
van Dorp, J. et al. High- or low-dose preoperative ipilimumab plus nivolumab in stage III urothelial cancer: the phase 1B NABUCCO trial. Nat. Med. 29, 588–592 (2023).
Vandekerkhove, G. et al. ERBB2/HER2 alterations in ctDNA and metachronous tissues of patients with metastatic urothelial cancer. Clin. Cancer Res 31, 3725–3741 (2025).
Shohdy, K. S. et al. Serial ctDNA analysis predicts clinical progression in patients with advanced urothelial carcinoma. Br. J. Cancer 126, 430–439 (2022).
Facchinetti, F. et al. Resistance to selective FGFR inhibitors in FGFR-driven urothelial cancer. Cancer Discov. 13, 1998–2011 (2023).
Helal, C. et al. Clinical utility of plasma ctDNA sequencing in metastatic urothelial cancer. Eur. J. Cancer 195, 113368 (2023).
Andersen, L. et al. Exploring the biology of ctDNA release in colorectal cancer. Eur. J. Cancer 207, 114186 (2024).
Vandekerkhove, G. et al. Toward informed selection and interpretation of clinical genomic tests in prostate cancer. JCO Precis Oncol. 8, e2300654 (2024).
Pao, W. et al. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med. 2, e73 (2005).
Goodall, J. et al. Circulating cell-free DNA to guide prostate cancer treatment with PARP inhibition. Cancer Discov. 7, 1006–1017 (2017).
Oxnard, G. R. et al. Association between plasma genotyping and outcomes of treatment with osimertinib (AZD9291) in advanced non–small-cell lung cancer. J. Clin. Oncol. https://doi.org/10.1200/JCO.2016.66.7162 (2016).
Loehr, A. et al. Emergence of BRCA reversion mutations in patients with metastatic castration-resistant prostate cancer after treatment with rucaparib. Eur. Urol. 83, 200–209 (2023).
Hudkins, R. L. et al. Discovery of TYRA-300: first oral selective FGFR3 inhibitor for the treatment of urothelial cancers and achondroplasia. J. Med Chem. 67, 16737–16756 (2024).
Ballard, J. A. et al. Abstract P141: preclinical characterization of LOX-24350, a highly potent and isoform-selective FGFR3 inhibitor. Mol. Cancer Ther. 20, P141–P141 (2021).
Grivas, P. et al. Spectrum of alterations in cell-free DNA of patients with advanced urothelial carcinoma. Bladder Cancer 7, 143–148 (2021).
Pascual, J. et al. ESMO recommendations on the use of circulating tumour DNA assays for patients with cancer: a report from the ESMO Precision Medicine Working Group. Ann. Oncol. 33, 750–768 (2022).
Santiago-Walker, A. E. et al. Analysis of FGFR alterations from circulating tumor DNA (ctDNA) and Tissue in a phase II trial of erdafitinib in urothelial carcinoma (UC). J. Clin. Oncol. 37, 420–420 (2019).
Harris, P. A. et al. The REDCap consortium: Building an international community of software platform partners. J. Biomed. Inform. 95, 103208 (2019).
Warner, E. W. et al. Multiregion sampling of de novo metastatic prostate cancer reveals complex polyclonality and augments clinical genotyping. Nat. Cancer 5, 114–130 (2024).
Herberts, C. et al. Deep whole-genome ctDNA chronology of treatment-resistant prostate cancer. Nature 608, 199–208 (2022).
Tolmeijer, S. H. et al. Early on-treatment changes in circulating tumor DNA fraction and response to enzalutamide or abiraterone in metastatic castration-resistant prostate cancer. Clin. Cancer Res. 29, 2835–2844 (2023).
Virtanen, T. et al. Repertoire and clinical hierarchy of AR locus alterations in castration-resistant prostate cancer. Ann. Oncol. 37, 388–402 (2026).
Cameron, D. L. et al. GRIDSS2: comprehensive characterisation of somatic structural variation using single breakend variants and structural variant phasing. Genome Biol. 22, 202 (2021).
Geoffroy, V. et al. AnnotSV: an integrated tool for structural variations annotation. Bioinformatics 34, 3572–3574 (2018).
Muller, D. C. et al. Prospective multicenter study of ctDNA versus tumor tissue guiding FGFR-targeted therapy in metastatic urothelial cancer. GitHub repository: wyatt-lab/fgfr_ctdna_tissue https://doi.org/10.5281/zenodo.18158103 (2026).
Acknowledgements
This work was supported by a Canadian Institutes of Health Research (CIHR) Project Grant (#PJT180417; to A.W.W., B.E., G.V.), a bladder cancer research grant from Bladder Cancer Canada (to B.E., A.W.W.), and an innovation grant from the Bladder Cancer Advocacy Network (BCAN; to A.W.W., B.E.). D.C.M. received salary support from the Swiss National Science Foundation, Swiss Society for Urology, and The Voluntary Academic Society Basel. A.M. was supported by a BC Cancer Rising Star Award through the BC Cancer Foundation. This study utilized biospecimens and clinical data from the BC Cancer Genitourinary Biobank and we acknowledge the staff and investigators for their support, as well as funding support for the biobank and this study from the BC Cancer Foundation. The authors gratefully thank all contributing patients and their families.
Author information
Authors and Affiliations
Contributions
D.C.M., C.W., K.N.C., G.V., A.W.W., and B.J.E. made substantial contributions to the conception or design of the work. D.C.M., A.J.M., J.V.W.B., M.S., C.W., C.V.R., K.R., L.R., E.S., J.N., N.A., N.S.B., D.F., J.J.K., J.M.L., L.N., K.N., M.O., G.O., S.P., M.S., S.S., P.T., E.W., C.Q.B., M.K., J.A., E.F., L.K., B.E., D.O., T.A.B., G.W., R.M., A.I.P., A.S., C.K., C.M.D., M.A., G.V., A.W.W., and B.J.E. made substantial contributions to the acquisition, analysis, or interpretation of data. A.J.M. and M.A. contributed to the creation of new software used in the work. D.C.M., A.J.M., G.V., A.W.W., and B.J.E. have drafted the work or substantively revised it.
Corresponding authors
Ethics declarations
Competing interests
D.C.M.: Honoraria—EMD Serono; Travel, Accommodations, Expenses—EMD Serono. A.J.M.: Honoraria—EMD Serono; Travel, Accommodations, Expenses—EMD Serono. L.N.: Honoraria—Bayer; Ipsen; Janssen; Pfizer; Tersera; Consulting or Advisory Role—Bayer; Eisai; Ipsen; Novartis; Pfizer; Research Funding - Bayer (Inst); Travel, Accommodations, Expenses—Pfizer, Ipsen. P.T. has served on advisory boards and/or received honoraria from Bayer, Astellas and has contract research agreements (via institution) with AstraZeneca and Janssen. E.W.: Honoraria—EMD Serono; Travel, Accommodations, Expenses—Eisai; Consulting or Advisory Role—Bayer, Eisai, EMD Serono, Merck, Roche; Research Funding—Lilly, Novartis, Merck. A.S. has served on advisory boards to Janssen. C.M.D.: Pfizer, Merck, Astra Zenca, Novartis, consulting/advisory: Biomica Ltd, Merck, Pfizer and Eisai, Novartis, travel expenses: Janssen, GSK, Pfizer. G.V.: research funding from Gilead Sciences; travel expenses, honoraria and/or advisory board roles from Gilead Sciences, Janssen, and EMD Serono. A.W.W. has served on advisory boards and/or received honoraria from AstraZeneca, Astellas, Bayer, EMD Serono, Janssen, Merck, and Pfizer; and has contract research agreements (via institution) with ESSA Pharma, Tyra Biosciences, and Promontory Therapeutics. The remaining authors declare no potential conflicts of interest.
Peer review
Peer review information
Nature Communications thanks Zachary McCaw, David McConkey and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Müller, D.C., Murtha, A.J., Bacon, J.V.W. et al. Prospective multicenter study of ctDNA versus tumor tissue guiding FGFR-targeted therapy in metastatic urothelial cancer. Nat Commun 17, 3263 (2026). https://doi.org/10.1038/s41467-026-69927-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69927-7