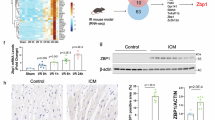
Abstract
The extensive crosstalk among pyroptosis, apoptosis, and necroptosis limits the efficacy of therapies targeting only one pathway. Here, we show that Prussian blue (PB) nanoparticles act as multi-target PANoptosis inhibitors by binding key PANoptosome components including RIPK1, ZBP1, and AIM2 through multimodal interactions, thereby concurrently suppressing pyroptosis, apoptosis, and necroptosis in myocardial ischemia-reperfusion injury (MIRI). Platelet membrane-coated PB nanoparticles (PB@PM) exhibit enhanced cardiac targeting and efficiently alleviate MIRI-induced cardiac dysfunction, adverse ventricular remodeling, and cardiomyocyte hypertrophy. Mechanistically, PB@PM disrupt PANoptosome assembly, scavenge reactive oxygen species, improve mitochondrial function, and restore immune-inflammatory homeostasis. By integrating single nucleus transcriptomics of human heart samples, molecular dynamics simulations, transcriptomics, medical imaging, and molecular validation, we systematically decipher the therapeutic mechanisms of PB-based PANoptosis inhibition. This study establishes an integrative multi-omics framework for exploring PANoptosis in cardiovascular diseases and provides a promising nanotherapeutic strategy for MIRI treatment.
Similar content being viewed by others
Introduction
Innate immunity constitutes the first-line defense against pathogenic insults and cellular stress, wherein pattern recognition receptors orchestrate inflammatory responses and programmed cell death through detection of pathogen- and damage-associated molecular patterns1. Emerging evidence from genetic, molecular, and biochemical studies reveals complex interconnectivity between cell death pathways, reshaping our understanding of inflammasomes and cell death complexes in diseases pathogenesis2,3,4. Notably, monotherapeutic targeting of individual pyroptotic, apoptotic, or necroptotic components often proves inadequate due to molecular redundancy5,6,7. The recent conceptual breakthrough of PANoptosis have unveiled an inflammation-driven programmed cell death modality governed by multilayered PANoptosome complexes. This pathophysiological process uniquely integrates molecular components from classical death pathways, with PANoptosome assembly serving as its biochemical hallmark. Extensive interactions between key molecules of pyroptosis, apoptosis, and necroptosis are the basis for PANoptosome assembly2,8. Currently, Z-DNA-binding protein 1 (ZBP1), receptor-interacting protein kinase 1 (RIPK1), absent in melanoma 2 (AIM2), and NLR family pyrin domain-containing 12 (NLRP12) PANoptosomes have been molecularly characterized3,8,9,10,11,12,13. Notably, previous studies have primarily explored the intrinsic mechanisms of PANoptosis in cancer, infectious, and inflammatory diseases.
Acute myocardial infarction persists as the global leading cause of mortality. Prompt revascularization constitutes the cornerstone for mitigating myocardial damage and optimizing long-term prognosis. However, revascularization can cause myocardial cell death and exacerbate post-ischemic myocardial injury, a phenomenon termed myocardial ischemia-reperfusion injury (MIRI), which frequently culminates in cardiac dysfunction, arrhythmia, and heart failure14,15. The pathophysiological complexity of MIRI stems from multifactorial mechanisms encompassing oxidative stress, calcium dysregulation, metabolic disturbances, and dysregulated immune-inflammatory activation16. Notably, ischemia-triggered apoptotic cascades critically dictate myocardial injury severity and propel maladaptive cardiac remodeling17,18. Given the intrinsic linkage between programmed cell death and inflammation, pyroptosis emerges as a key mediator of MIRI-related inflammatory response that potentiates detrimental ventricular remodeling19. Concurrently, robust necroptosis activity accompanies MIRI progression20,21. Despite these insights, the therapeutic potential of PANoptosis modulation in cardiovascular pathologies, particularly MIRI, remains underexplored and clinically untapped.
PANoptosis represents a pivotal therapeutic target against diseases, and its biological effects cannot be explained solely by pyroptosis, apoptosis, or necroptosis8. The intricate molecular interplay among these death pathways predisposes single-pathway inhibition strategies to compensatory pathway activation, ultimately diminishing therapeutic effectiveness. Consequently, monotherapeutic targeting of individual molecular components fails to suppress PANoptosis22,23. Experimental evidence demonstrates that pharmacological blockade of terminal effectors—including mixed lineage kinase domain-like pseudokinase (MLKL), caspase-1, gasdermin D (GSDMD), or caspase 3/7—achieves limited efficacy in curtailing overall cell death24,25. Current investigative efforts focus on multipronged strategies targeting upstream initiators, signaling hubs (RIPK1/3), protease networks, and PANoptosome complexes8,26. Nevertheless, most available inhibitors exhibit narrow target specificity, rendering them inadequate for addressing PANoptosis’ pathway interconnectivity. This underscores the urgent need for novel therapeutic entities capable of simultaneous pathway modulation to disrupt the crosstalk among pyroptosis, apoptosis, and necroptosis.
In this study, we establish Prussian blue (PB) nanoparticles as PANoptosis inhibitors capable of concurrently targeting multiple PANoptosomes, effectively disrupting the molecular crosstalk among pyroptosis, apoptosis, and necroptosis to combat PANoptosis-driven pathologies (Fig.1). Integrating single-nucleus transcriptomics of human cardiac specimens with prior experimental evidence, we validated MIRI as a representative PANoptosis-associated disease model. Molecular dynamics simulations revealed high-affinity binding between the constructed PB and core PANoptosome components RIPK1, ZBP1, and AIM2. To improve targeted delivery, we engineered platelet membrane-modified PB (PB@PM), which demonstrated efficient mitigation of MIRI-induced cardiac dysfunction and pathological ventricular remodeling, enhanced extracellular matrix regeneration and functional neovascularization, and robust suppression of cardiomyocyte hypertrophy. Mechanistic studies revealed that PB@PM modulated mitochondrial metabolism and restored immune-inflammatory balance, thereby disrupting PANoptotic circuitry. Bulk transcriptomics combined with multiplex immunoblotting confirmed PB@PM’s pan-inhibitory effects on PANoptosome activation, conclusively establishing its therapeutic efficacy against MIRI-associated PANoptosis. This study highlights a promising therapeutic strategy for MIRI and offers valuable information for developing clinical therapies for innate immunity and inflammation.
PBs act as PANoptosis inhibitors by targeting RIPK1, ZBP1, and AIM2 to suppress multiple PANoptosomes, effectively blocking crosstalk among pyroptosis, apoptosis, and necroptosis in PANoptosis-driven diseases. In addition, PB regulates mitochondrial metabolism and immune-inflammatory homeostasis, further disrupting multimodal cell death crosstalk. Some elements in the image were sourced from BioRender (Created in BioRender. xu, L. (2026) https://BioRender.com/9kka4p3).
Results
MIRI as a representative PANoptosis-related disease model
Emerging evidence demonstrates that ischemic stress induces mitochondrial genomic instability in cardiomyocytes, culminating in mitochondrial DNA (mtDNA) leakage that potently activates the STING pathway and propagates pro-inflammatory cascades27,28,29,30. Notably, mtDNA-STING axis activation serves as a master regulator of PANoptosis via AIM2/ZBP1-PANoptosome induction2,31,32. Concurrently, RIPK1 hyperactivation in MIRI pathogenesis is mechanistically linked to mitochondrial dysfunction, ZBP1 signaling, and TNF-α-mediated inflammation21,33,34,35. This compelling evidence positions MIRI as a paradigm of PANoptosis-driven cardiovascular pathology. To clinically validate PANoptosis targeting potential, we analyzed a single-nucleus transcriptomic dataset from human specimens.
We performed comparative analysis of non-transplanted donor heart (control group, Con) versus necrotic core (ischemic zone, IZ) specimens from acute myocardial infarction patients (Fig. 2A). Single-cell clustering was visualized through uniform manifold approximation and projection (UMAP), t-distributed stochastic neighborhood embedding analysis, and principal component analysis (PCA), with cell type annotation based on canonical marker gene expression. Thirteen distinct cell populations were identified: adipocytes, B cells, cardiomyocytes, doublets, endothelial cells, fibroblasts, mast cells, megakaryocytes, mononuclear phagocytes, mural cells, plasma cells, Schwann cells, and T cells (Fig. 2Band Supplementary Fig. 1A). Global cellularity was markedly reduced in IZ specimens, most pronounced in cardiomyocytes (Fig. 2Band Supplementary Fig. 1A). Subclustering analysis resolved cardiomyocytes into five subtypes, revealing substantial depletion of subclusters 1, 2, 4, and 5 in the IZ group (Fig. 2Cand Supplementary Fig. 1B). PANoptosis activation signature was mapped at single-cell resolution across all cardiac cells and subtypes. Quantitative analysis revealed cell type-specific PANoptosis reprogramming, with cardiomyocytes exhibiting the most profound pro-PANoptotic shift (Fig. 2D, E). UMAP plots and box-type statistical plots were also generated for pyroptosis, apoptosis, and necroptosis. Comparative death modality profiling demonstrated dominant apoptotic/necroptotic activation in bulk cardiac cells during acute myocardial infarction onset, and concurrent amplification of pyroptosis, apoptosis, and necroptosis specifically in IZ cardiomyocytes (Supplementary Fig. 2). Intuitively, the expression levels of multiple PANoptosis-related genes were markedly upregulated in both the overall cell population and the cardiomyocytes of the IZ group (Fig. 2Fand Supplementary Figs. 3–5).
A Schematic of ischemic zone (IZ), remote zone (RZ), and border zone (BZ) in human acute myocardial infarction. The heart element in the image was sourced from BioRender (Created in BioRender. xu, L. (2026) https://BioRender.com/9kka4p3). B UMAP visualization of cell distribution (left) and annotated cell types (right) in the control (n = 4) versus IZ (n = 11) groups. C Cardiomyocyte subpopulation quantification (left) and UMAP-based clustering (right) in the control and IZ groups. D, E UMAP projection (D) and box plots (E) showing PANoptosis activation across cardiac cell types. (The exact n in E = [left to right] 20, 84 cells; 17326, 3506 cells; 6231, 6969 cells; 9468, 12187 cells; 3386, 7142 cells; 3039, 2286 cells; 583, 254 cells; 531, 1138 cells). The minima, maxima, mean, median, bounds, whiskers, and percentile information were provided in the Source Data file. Exact P-values from left to right: 2.39E-265, 1.55E-300, 1.05E-35, 4.09E-05, 2.21E-06, 0.0002, 1.9E-11. F Violin plots comparing PANoptosis-related gene expression (AIM2, ZBP1, RIPK1, Pyrin, NLRP12, NLRP3, MLKL, GSDMD, RIPK3, caspase 3, caspase 1, and caspase 8) in total cells. (n = 40706 cells [control], 33688 cells [IZ]). The minima, maxima, mean, median, bounds, whiskers, and percentile information were provided in the Source Data file. Exact P-values from left to right (AIM2 to caspase 8): 0.2752, 0.0002, 0, 0.0854, 6.67E-09, 7.19E-82, 3.59E-37, 0.0709, 0.0269, 2.29E-280, 2.45E-30, 4.07E-09. G, H Cardiomyocyte-specific UMAP (G) and quantitative PANoptosis levels (H) of the control and IZ groups. (The exact n in H = [left to right] 6904 cells, 0; 5281 cells, 0; 1191, 3346 cells; 3192, 80 cells; 758, 80 cells). The minima, maxima, mean, median, bounds, whiskers, and percentile information were provided in the Source Data file. Exact P-values from left to right: 4.01E-05, 0.0021. I Proportional representation of cardiomyocyte subpopulations. J, K Heatmap depicting activation patterns of apoptosis/PANoptosis/pyroptosis/necroptosis (J) and PANoptosis-related genes (K) across cardiomyocyte subtypes. Significance: wilcox.test, a two-sided test; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns: no significant difference (P > 0.05). Source data are provided as a Source Data file.
Focusing on cardiomyocyte functional heterogeneity, we quantified PANoptosis activation across subtypes. Strikingly, subclusters 1/2 were nearly eradicated in IZ specimens, while subcluster 4 exhibited severe depletion (Fig. 2G–K). Residual subclusters 3-5 displayed elevated PANoptosis-related gene expression (Fig. 2H–K), and apoptosis-dominant death modality (Supplementary Fig. 6). These findings indicate that during myocardial ischemic infarction, cardiomyocytes do not exhibit uniform activity, with subclusters 1 and 2 being the most susceptible to death, followed by subcluster 4, and subcluster 3 being the most resistant to the post-ischemic microenvironment. Collectively, the single-cell atlas establishes PANoptosis as a spatially regulated therapeutic target for MIRI intervention.
Computational simulation of PB interaction with PANoptosis-related proteins
PB is an antidote approved by the US Food and Drug Administration for the clinical treatment of thallium poisoning. Previous studies reported that when its size was at the nanometer level, PBs showed various biological activities, including inhibiting cell apoptosis, pyroptosis, or necroptosis36,37,38. PBs with multitarget effects on cell death are expected to play an important role in PANoptosis. However, their role in PANoptosis remains unclear.
Molecular dynamics simulations were firstly conducted to elucidate interactions between PB and PANoptosome core components. Mechanistically, PANoptosome assembly initiates upon sensing diverse pathogenic stimuli, orchestrating PANoptosis execution. Currently, ZBP1, RIPK1, and AIM2 PANoptosomes have undergone molecular characterization8. The components of the ZBP1 PANoptosome include ZBP1, RIPK3, caspase 1/6/8, apoptosis-associated speck-like protein containing a CARD (ASC), nucleotide-binding oligomeric domain-like receptor protein 3 (NLRP3), and RIPK1. RIPK1 PANoptosome consists of RIPK1, Fas-associated death domain protein (FADD), caspase 1/8, ASC, NLRP3, and RIPK3. AIM2 regulates the innate immune sensors Pyrin and ZBP1 and forms PANoptosomes with ASC, caspase-1/8, RIPK1/3, and FADD (Fig. 1). These three hubs were selected as representative therapeutic targets due to their pivotal role in cross-regulating pyroptotic, apoptotic, and necroptotic pathways.
System validation confirmed simulation robustness via stable root mean square deviation (RMSD) equilibration in PB-RIPK1/ZBP1/AIM2 systems (Supplementary Fig. 7). For AIM2 systems, dual simulations probing PB interactions with distinct functional domains (HIN and PYD) were independently performed. The simulation results of the C-terminal HIN domain on the AIM2 structure (AIM2HIN) were shown in Fig. 3A. As known as the binding of this region with dsDNA releases AIM2 from its inhibitory state, so that the helix interaction of the PYD domain (AIM2PYD) can further bind with ASC39. Here, PB evidently interfered with the conformation of the AIM2HIN domain. Analysis of the interaction with the PB interface revealed that the contact residues were mainly concentrated on the charged Arg and Lys residues (Fig. 3B, C). The interactions between different residues and PB also indicated that Lys and Arg play important roles in PB adhesion to AIM2HIN (Fig. 3D, Eand Supplementary Movie 1). The total interaction energy between the AIM2HIN structure and PB reached -267.4 kJ/mol, which mainly refers to the electrostatic force (Fig. 3F). However, PB did not have a strong conformational effect on the PYD domain in AIM2 (Supplementary Fig. 8and Supplementary Movie 2), which may be related to the high stability of the helical structure.
A Initial and final snapshots (200 ns) of the PB-AIM2HIN simulation system. Interaction interfaces are highlighted (blue/yellow boxes). B, C The residue contact detail of two chains of AIM2HIN with PB is highlighted on the left snapshots. D, E The interaction energy analysis of these contact residues of AIM2HIN with PB (n = 10, taken from the final 10 ns interaction energy of the simulated system). F The interaction energy between PB and proteins in PB-AIM2 HIN changes with time. G–L The conformation images of the PB-ZBP1 (G) and PB-RIPK1 (J) systems at the end of the simulation (t = 200 ns) and the residue contact details. The interaction energy analysis of the contact residues of ZBP1 (H) and RIPK1 (K) with PB (n = 10, taken from the final 10 ns interaction energy of the simulated system). The interaction energy between PB and proteins in PB-ZBP1 (I) and PB-RIPK1 (L) changes with time. Data: mean ± SD. Source data are provided as a Source Data file.
Comparative analysis of PB-ZBP1 interactions revealed substantially weaker binding affinity compared to PB-AIMHIN complexes (Fig. 3Gand Supplementary Movie 3). Electrostatic forces dominated at critical interfacial residues Arg135/Lys138, serving as primary anchoring points for PB surface adhesion (Fig. 3H). Energy quantification demonstrated moderate binding strength (-157.9 kJ/mol) during the equilibrium phase (Fig. 3I), while radius of gyration (Rg) trajectory analysis indicated ZBP1 structural compaction (Supplementary Fig. 7C). This compression phenotype likely disrupts Zα domain-mediated dsDNA recognition, compromising ZBP1’s DNA-sensing capacity.
We further conducted the conformational effects of RIPK1 in an environment containing PB. The PB-RIPK1 interaction analysis showed that PB tightly binds to RIPK1, primarily through interaction with C-terminal residues of RIPK1 (Fig. 3Jand Supplementary Movie 4). A detailed residue-level analysis revealed that the binding sites between PB and RIPK1 are predominantly located within a turn region of the helical domain in the N-terminal segment, involving key residues such as Glu98, Met99, Pro102, Leu103, Thr244, Tyr246, Ser282, and Gln283 (Fig. 3K). The overall interaction energy was calculated to be − 73.2 kJ/mol, with van der Waals interactions being the dominant force, particularly the strong interaction between Tyr and PB (Fig. 3L). This intimate binding of PB to RIPK1 may markedly modulate RIPK1-mediated inflammatory signaling and cell death regulation40,41,42,43. In conclusion, the molecular dynamics simulation results elucidate the regulatory role of PB in the pathological progression of PANoptosis, providing a therapeutic strategy for the early intervention and treatment of PANoptosis-related diseases.
Construction of PB@PM for targeted treatment of MIRI in vivo
Leveraging molecular dynamics simulation data and biomedical nanotechnology requirements, we synthesized PBs through single-precursor-mediated one-step hydrothermal synthesis. Polyvinylpyrrolidone was incorporated during nanoparticle synthesis as both reducing agent and dispersion stabilizer, endowing the nanostructures with exceptional colloidal stability in physiological media - a critical parameter for clinical translation. Scanning electron microscopy and transmission electron microscopy (TEM) revealed the morphology, size, and uniformity of the nanoparticles (Fig. 4A, B). Comprehensive material characterization through elemental mapping, X-ray diffraction, X-ray photoelectron spectroscopy, ultraviolet-visible-near infrared spectroscopy, and Fourier-transform infrared spectroscopy confirmed successful construction of PB (Fig. 4Cand Supplementary Fig. 9).
A Representative SEM image of PB. Scale bar: 200 nm. B Representative TEM image of PB. Scale bar: 200 nm. C Element mapping of PB. Scale bar: 100 nm. D TEM image of PB@PM. Scale bar: 50 nm. E Western blot analysis of CD41, CD42b, CD47, and GP6 expression in PM and PB@PM. Scale bar: 50 nm. F Fluorescence colocalization of PB and PM in PB@PM, and control represents a direct mixture of FITC-labeled PB (green) and Dil-labeled PM (red). Scale bar: 10 μm. G Typical ex vivo fluorescence images of major organs at 12 and 24 h after administration of Cy5.5-labeled PB (Sham mice; Control group), Cy5.5-labeled PB (MIRI mice; PB group), and Cy5.5-labeled PB@PM (MIRI mice; PB@PM group). H, I Quantitative analysis of the fluorescence intensities from separated hearts, livers, spleens, lungs, and kidneys in each group at 12 and 24 h, respectively (n = 3 biologically independent replicates). Exact P-values from upper to lower: (12 h-heart) 8.70E-07, 9.50E-07, 0.8948; (24 h-heart) 0.0442; (12 h-liver) 0.6819; (24 h-liver) 0.9998; (12 h-spleen) 0.1353; (24 h-spleen) 0.9857; (12 h-lung) 0.0654; (24 h-lung) 0.0599; (12 h-kidney) 0.9741; (24 h-kidney) 0.8667. Data are expressed as mean ± SD. P- values were calculated by one-way ANOVA with Tukey’s correction. *P < 0.05, **P < 0.01, ***P < 0.001, and ns implies P > 0.05. Source data are provided as a Source Data file.
To endow PB with the ability to target PANoptosis-related disease sites, we encapsulated PB using platelet membranes (PM) derived from wild-type mice to construct PB@PM. The results of TEM, western blotting, fluorescence colocalization, dynamic light scattering, and zeta potential demonstrated the successful construction of PB@PM (Fig. 4D–Fand Supplementary Fig. 10). Furthermore, we systematically investigated the myocardial targeting efficacy and biodistribution profile of PB@PM in vivo. Fluorescence imaging analysis was performed on major organs harvested from MIRI mice at 12 and 24 h post-injection of Cy5.5-labeled PB or PB@PM, with sham mice receiving Cy5.5-labeled PB serving as the control group. Quantitative analysis revealed that PB@PM exhibited substantially elevated fluorescence intensities within the ischemic myocardial region compared to both PB-treated groups at both time points (Fig. 4G–I). Notably, there were no observable differences in the fluorescence intensities of the livers, spleens, lungs, and kidneys across all experimental groups. These results indicated that PB@PM exhibits superior myocardial targeting efficacy with sustained retention in infarcted cardiac tissue exceeding 24 h. The biosafety profile of PB@PM was rigorously assessed. Comprehensive hematological analysis and histopathological examination via H&E staining revealed no visible histopathological alterations in major organs or aberrations in hematological/biochemical parameters following PB@PM administration (Supplementary Fig. 11and Supplementary Table 2). This targeted delivery system addresses critical limitations in current MIRI therapeutics and establishes a robust foundation for translational applications in PANoptosis-related pathologies.
Therapeutic efficacy of PB@PM in MIRI
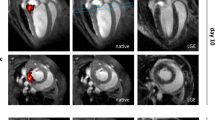
Subsequently, we evaluated the potential PANoptosis-inhibitory effects of PB@PM in MIRI. Serial echocardiographic assessments were conducted at preoperative baseline and postoperative days 1, 14, and 28 to quantify cardiac functional parameters, including the left ventricular ejection fraction (LVEF), fraction shortening (FS), LV end-diastolic volume, and LV end-systolic volume. Preoperative echocardiography confirmed the absence of preexisting cardiac conditions in all murine subjects. While no statistically significant intergroup differences in LVEF and LVFS were observed at postoperative day 1, all experimental groups exhibited marked reductions compared to baseline values, confirming MIRI-induced cardiac dysfunction (Fig. 5A–C). Notably, PB@PM treatment showed superior therapeutic efficacy versus PB treatment at postoperative days 14 and 28, as evidenced by improved echocardiographic parameters (Fig. 5A–E).
A–E Echocardiographic analysis of left ventricular ejection fraction (LVEF), LV fraction shortening (LVFS), LV end-systolic volume (LVESV), and LV end-diastolic volume (LVEDV) on preoperative day 1 and postoperative days 1, 14, and 28. Scale bar: 100 ms. Exact P-values from upper to lower: (B) day 14 (0.0012), day 28 (0.00003, 0.0028); (C) day 14 (0.0056), day 28 (0.0002, 0.039); (D) day 14 (0.0011, 0.0201), day 28 (0.00001, 0.0004); (E) day 14 (0.015, 0.0294), day 28 (0.0101). F Representative Masson and HE staining in hearts on 28 days post-surgery. Scale bar: 1 mm. G Fibrosis quantification in hearts 28 days post-surgery. Exact P-values from upper to lower: 2.28E-08, 3.28E-07, 0.0116. H Evans blue/TTC staining and quantitative analysis showing the cardiac cell survival and blood perfusion in different groups on day 3 post-surgery. AAR: area at risk. IF: infarct zones. Scale bar: 1 mm. Exact P-values from upper to lower: (AAR/LV) 0.4990 for MIRI vs. PB, 0.3119 for MIRI vs. PB@PM, 0.9269 for PB vs. PB@PM; (IF/AAR) 3.48E-08, 0.0001, 0.00002. I, J TUNEL (red) and cTnT (green) co-staining and quantification in peri-infarct zones (day 3). Scale bar: 50 μm. Exact P-values from upper to lower: 0.000005, 0.0006, 0.0154. K, L Representative immunofluorescence images and quantitative analysis of CD86 (red) in peri-infarct regions (day 3). Scale bar: 1 mm or 50 μm. Exact P-values from upper to lower: 2.17E-08, 0.00001, 0.00005. M, N Representative immunofluorescence staining and quantitative analysis of Ly-6G (red) in peri-infarct regions (day 3). Scale bar: 1 mm or 50 μm. Exact P-values from upper to lower: 8.27E-09, 5.89E-08, 9.99E-07. O–R Representative immunofluorescence staining and quantification analysis of α-SMA (red)/cTnT (green) and CD31(red)/cTnT (green) in peri-infarct region (day 28). Scale bar: 1 mm or 50 μm. Exact P-values from upper to lower: (P) 0.000002, 0.0005, 0.0026; (R) 0.000004, 0.0003, 0.016. Data: mean ± SD of 5 biologically independent replicates. Significance: one-way ANOVA with Tukey’s post-hoc test (*P < 0.05, **P < 0.01, ***P < 0.001; ns, P > 0.05). Source data are provided as a Source Data file.
Subsequent to the 28-day postoperative period, histopathological assessments were conducted on murine cardiac tissues. Infarct size quantification and collagen deposition analysis were performed utilizing Masson’s trichrome and H&E staining methodologies. The MIRI cohort exhibited pronounced myocardial atrophy, ventricular wall thinning, and excessive collagen fiber accumulation within the left ventricular wall. Notably, PB@PM administration demonstrated conspicuous attenuation of MIRI-induced cardiac dysfunction and fibrotic remodeling (Fig. 5F, G). In acute phase evaluations, dual staining protocols employing Evans blue and 2,3,5-triphenyltetrazolium chloride (TTC) were implemented for infarct characterization. Histological analysis revealed distinct colorimetric demarcation: blue-stained regions indicated viable left ventricular tissue, white-colored areas represented infarct zones (IF), while TTC-positive regions designated the area at risk (AAR)44. Quantitative assessment demonstrated a progressive reduction in the infarct-to-risk area ratio (IF/AAR) across MIRI, MIRI + PB, and MIRI + PB@PM treatment groups (Fig. 5H). These results collectively substantiate the potential therapeutic efficacy of PB@PM in mitigating post-ischemic myocardial injury progression.
Apoptotic cell death analysis through terminal deoxynucleotide transferase-mediated dUTP nick end labeling (TUNEL) staining revealed that PB@PM markedly decreased the number of TUNEL+ cardiomyocytes within the infarct margin area (Fig. 5I, J). On the third postoperative day, immunofluorescence analysis revealed that PB@PM treatment demonstrated a marked reduction in CD86+ M1 macrophages and Ly6G+ neutrophils recruitment induced by MIRI (Fig. 5K–N). Furthermore, quantitative analysis of macrophage polarization markers indicated that pro-inflammatory M1 phenotype markers (TNF-α, IL-1β, and IL-6) were distinctly downregulated, while anti-inflammatory M2 markers (IL-10 and TGF-β) exhibited marked upregulation in both LPS-stimulated RAW264.7 macrophage cultures and ischemic myocardial tissues following PB@PM administration (Supplementary Figs. 12 and 13). These results suggest that PB@PM effectively regulates the immune microenvironment after MIRI.
Post-MIRI cardiac repair unfolds through a coordinated cascade of biological processes, wherein extracellular matrix (ECM) remodeling constitutes a critical pathological determinant. Efficient repair depends on the transformation of fibroblasts into myofibroblasts, which produce large amounts of ECM proteins and facilitate wound contraction45. Collagens are the most abundant ECM proteins in the LV. Following MIRI, collagen increases in the peri-infarct region, and types I and III are the major components of the myocardium scar46,47. Quantitative fluorescence analysis revealed the highest expression levels of collagen I and III, alpha-smooth muscle actin (α-SMA)+ myofibroblasts, and CD31+ vascular endothelial cells in the peri-infarct zone of PB@PM-treated MIRI mice across the three different treated groups (Fig. 5O–R and Supplementary Figs. 14 and 15). During the reparative phase, robust angiogenesis coupled with enhanced α-SMA in infarct and peri-infarct territories facilitates the development of organized granulation tissue, characterized by increased collagen density, aligned stress fibers, and augmented mechanical strength. This structural reinforcement maintains ventricular integrity while preventing adverse remodeling manifestations including infarct expansion and ventricular rupture48,49. Morphometric analysis further demonstrated that PB@PM treatment markedly attenuated pathological myocardial cell hypertrophy compared to MIRI controls (Supplementary Fig. 16). Collectively, these experimental findings substantiate that PB@PM confers therapeutic benefits in MIRI management through synergistic mechanisms involving ECM remodeling enhancement, angiogenic activation, and suppression of maladaptive myocardial hypertrophy.
PB@PM comprehensively blocks the crosstalk between pyroptosis, apoptosis, and necroptosis
To elucidate the molecular mechanisms underlying PB-mediated suppression of PANoptosis in vivo, comprehensive transcriptomic analysis was conducted through bulk RNA sequencing of myocardial tissues harvested from the Sham, MIRI, and PB@PM treatment groups. PCA revealed distinct clustering patterns among experimental groups, with PB@PM-treated specimens demonstrating prominent transcriptional modulation compared to MIRI controls (Fig. 6A). The volcano plots showed distinct differential gene expression profiles between the MIRI and control groups (Supplementary Fig. 17A). Quantitative analysis demonstrated that PB@PM treatment induced a total of 1875 upregulated and 2752 downregulated differentially expressed genes (DEGs) in contrast to the MIRI group (Fig. 6B). Furthermore, comparative analysis between the PB@PM and Sham group identified 227 upregulated and 97 downregulated DEGs (Supplementary Fig. 17B). Notably, the 10 most prominently upregulated and downregulated genes are clearly annotated in the respective volcano plots. Heatmap analysis provided a comprehensive visualization of differential gene expression across the three experimental groups, revealing pronounced alterations in transcriptional patterns associated with immune and inflammatory responses, energy metabolism, oxidative stress response, and redox homeostasis maintenance (Supplementary Fig. 17C). These biological pathways are integral to the cardioprotective mechanisms observed during MIRI. Venn diagram analysis quantified the overlap of DEGs between the control vs. MIRI and PB@PM vs. MIRI comparisons, with 1284 upregulated and 2351 downregulated genes shared between both comparisons, representing 56.7% and 70.4% of the respective total DEG populations (Fig. 6C).
A PCA of RNA-seq data (Sham: orange; MIRI: green; PB@PM: blue; n = 5 biologically independent replicates). B Volcano plot of differentially expressed genes (DEGs; PB@PM vs. MIRI: red/blue = up/downregulated). The Wald test of DESeq2 (DESeq(dds) default test = “Wald”) was used, which is a two-sided test. Multiple comparisons correction: the FDR (False Discovery Rate) correction of the BH (Benjamini-Hochberg) method was computed to obtain P adjustment (P.adj.). C Venn diagram of shared DEGs between Sham-MIRI and PB@PM-MIRI comparisons. D GSVA of pathway enrichment across groups. E KEGG pathway enrichment of downregulated genes in PB@PM vs. MIRI. A one-sided hypergeometric test was used. Multiple comparisons correction: the FDR correction for the BH method was calculated to obtain P.adj. F, G GSEA networks (F) and protein interaction networks (G) for pyroptosis, apoptosis, and necroptosis. H GSEA plots showing marked downward trends of pyroptosis, apoptosis, and necroptosis in the PB@PM group compared to the MIRI group. A permutation test was used, based on weighted Kolmogorov-Smirnov-like statistics. It is a two-sided test (tests whether the enrichment scores are significantly deviated from zero, which can be positive or negative, corresponding to up- or down-regulated enrichment, respectively). Multiple comparisons correction: The FDR correction for the BH method was calculated to obtain P.adj. I Pearson correlation between pyroptosis, apoptosis, and necroptosis in the Sham (C), MIRI (I), and PB@PM (T) groups. Using psych R package, method = “spearman” function, t-test, two-sided test, and no correction for multiple comparisons. J Western blot of PANoptosis-related proteins (AIM2, ZBP1, Pyrin, ASC, FADD, Bax, Bcl-2, phosphorylated/total RIPK1/RIPK3/MLKL, cleaved/total caspase-1/3/8 and GSDMD) in ischemic myocardium (day 3; n = 5 biologically independent replicates). Source data are provided as a Source Data file.
The Gene Set Variation Analysis plot showed that the expression levels of pyroptosis, apoptosis, and necroptosis were dramatically elevated in the MIRI group compared to the Sham group. In stark contrast, these pathways were substantially downregulated in the PB@PM treatment group (Fig. 6D). Further Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of the downregulated genes revealed that PB@PM markedly suppressed the molecular pathways associated with pyroptosis, apoptosis, and necroptosis following MIRI (Fig. 6E). The Gene Set Enrichment Analysis (GSEA) network diagram and protein network further illustrated that pyroptosis, apoptosis, and necroptosis were tightly interconnected and mutually reinforcing pathways (Fig. 6F, G). Moreover, the GSEA plot confirmed that PB@PM treatment attenuated pyroptosis, apoptosis, and necroptosis compared to the MIRI group (Fig. 6H). Correlation analysis revealed a robust positive association among pyroptosis, apoptosis, and necroptosis across different treatment groups (Fig. 6I). Specifically, the MIRI group exhibited high panoramic expression of these pathways, whereas PB@PM treatment robustly reduced their expression levels, bringing them closer to the profile observed in the Sham group (Fig. 6I). Furthermore, we conducted a systematic investigation of PANoptosis in murine ischemic myocardial tissues through western blot analysis and histopathological evaluation. In the MIRI group, PANoptosis exhibited distinct activation, evidenced by upregulation of active forms of AIM2, ZBP1, Pyrin, ASC, FADD, apoptotic markers (caspase 3, caspase 8, Bax, and Bcl-2), necroptosis markers (RIPK1, RIPK3, and MLKL), and pyroptosis markers (caspase 1, NLRP3, and GSDMD) (Fig. 6J and Supplementary Figs. 18 and 19). Conversely, the PB@PM treatment regimen demonstrated marked downregulation of PANoptosis-associated protein expression (Fig. 6J and Supplementary Figs. 18 and 19). These findings provided compelling evidence that PB@PM exerted comprehensive inhibition of RIPK1 (RIPK1, NLRP3, RIPK3, ASC, FADD, caspase 1, and caspase 8), ZBP1 (ZBP1, NLRP3, ASC, RIPK3, RIPK1, caspase 1, and caspase 8), and AIM2 (AIM2, Pyrin, ZBP1, ASC, FADD, RIPK1, RIPK3, and caspase 8) PANoptosomes. This multi-modal suppression effectively attenuated MIRI-induced PANoptosis execution and conferred cardioprotection against myocardial injury.
Regarding the efficacy of PB in molecular dynamics simulations, we conducted supplementary validation to corroborate its inhibitory effects on the three principal effectors of PANoptosis (AIM2, ZBP1, and RIPK1). Lentiviral overexpression constructs targeting AIM2, ZBP1, and RIPK1 were systematically developed (Supplementary Fig. 20). Initial fluorescence colocalization analysis revealed that PB treatment induced evident spatial overlap with endogenous AIM2, ZBP1, and phosphorylated RIPK1 in OGD/R-stimulated HL-1 cells (Supplementary Fig. 21A). Quantitative western blot analysis demonstrated that PB exhibited comparable suppression efficacy against exogenously overexpressed AIM2 and ZBP1 in both normoxic and OGD/R conditions (Supplementary Fig. 21B), indicating that PB’s potent inhibitory activity effectively neutralized the OGD/R-induced protein expression perturbations. Notably, while RIPK1 overexpression remained unaffected by PB treatment, the OGD/R-induced phosphorylation of RIPK1 was remarkably attenuated by PB intervention (Supplementary Fig. 21B), suggesting differential regulatory mechanisms of PB on RIPK1 activation states.
We conducted additional validation of PANoptosis manifestations through in vitro cellular experiments. CCK-8 assays and Calcein AM/PI viability staining indicated that PB and PB@PM treatments produced a marked reduction in OGD/R-induced HL-1 cell death (Supplementary Fig. 22A–C). Furthermore, under equimolar low-dose conditions, the triple combination of Necrostatin-1 (necroptosis inhibitor), Disulfiram (pyroptosis inhibitor), and Z-VAD-FMK (pan-caspase inhibitor) reversed the limited therapeutic efficacy observed with monotherapy approaches (Supplementary Fig. 22D), providing evidence for crosstalk among distinct cell death modalities and indicating insufficient therapeutic outcomes from single-pathway inhibition. Notably, PB monotherapy achieved comparable therapeutic efficacy to the combination regimen (Supplementary Fig. 22D). Flow cytometric analysis using Annexin V-FITC/PI and TUNEL staining showed that PB@PM attenuated apoptosis levels in OGD/R-exposed HL-1 cells (Supplementary Fig. 22E, F). Morphological analysis further confirmed characteristic pyroptotic features - including cellular swelling, membrane rupture, and cytoplasmic vacuolation - in OGD/R-treated HL-1 cells (Supplementary Fig. 22G, H), which were substantially ameliorated by PB and PB@PM treatment, confirming their potent anti-pyroptotic activity. Consistent with in vivo findings in murine ischemic myocardium, PB@PM treatment inhibited OGD/R-induced upregulation of AIM2, ZBP1, phosphorylated RIPK1, cleaved caspase 1, cleaved caspase 8, cleaved GSDMD, and phosphorylated MLKL in HL-1 cells (Supplementary Fig. 22I, J). Besides, PB@PM or PB treatment markedly suppressed the colocalization of ASC and caspase 1/8 (Supplementary Fig. 22K, L), the assembly of which is essential for the formation of PANoptosome complex, suggesting that PB@PM can inhibit the PANoptosome assembly. Collectively, these findings provide comprehensive evidence for PANoptosis activation during MIRI and substantiate the therapeutic potential of PB@PM in targeting this multifaceted cell death pathway. In addition, we also found that PB@PM could suppress the secretion of proinflammatory cytokines in macrophages by reducing PANoptosis levels (Supplementary Fig. 23).
Inflammation and oxidative stress are associated with the crosstalk among pyroptosis, apoptosis, and necroptosis2. Gene ontology (GO) enrichment analysis showed that PB@PM predominantly modulated mitochondrial respiration function and energy metabolism pathways in MIRI mice. Conversely, downregulated genes were predominantly associated with leukocyte chemotaxis, cytokine-mediated signaling cascades, and immune-inflammation regulatory networks (Supplementary Fig. 24A). Notably, comparative GO analysis between control and MIRI groups demonstrated similar biological regulatory patterns (Supplementary Fig. 17D), indicating PB@PM’s capacity to restore aberrant MIRI-associated biological processes to homeostatic functional states. Integrated GO and KEGG pathway analyses showed that PB@PM treatment resulted in remarkable upregulation of mitochondrial metabolic processes, including oxidative phosphorylation, tricarboxylic acid cycle activity, and fatty acid β-oxidation, alongside activation of cAMP-dependent and cGMP-PKG signaling cascades (Supplementary Fig. 24C, D). The pathophysiological significance of these cyclic nucleotide systems in MIRI pathogenesis warrants emphasis: while pathological cAMP overaccumulation during myocardial ischemia exacerbates ischemia-induced cellular injury through accelerated ATP depletion and membrane depolarization50, pharmacological enhancement of cGMP/PKG signaling confers cardioprotection through vasodilatory effects on vascular smooth muscle cells. This vascular relaxation mechanism reduces coronary resistance while augmenting perfusion pressure, thereby ameliorating myocardial hypoxia and limiting apoptotic cell death51,52. Our findings demonstrate that PB@PM exerts dual regulatory effects by attenuating pathological cAMP hyperactivity while potentiating cGMP/PKG-mediated cytoprotective signaling (Supplementary Figs. 25 and 26). This pharmacological modulation demonstrates therapeutic potential for preserving cardiomyocyte viability and restoring post-ischemic myocardial functional recovery.
In addition to the three PANoptosis-associated cell death modalities, downregulated DEGs exhibited notable enrichment in various immune and inflammatory signaling pathways, including nuclear factor (NF)-κB, TNF, chemokine, and TLR signaling pathways (Fig. 6E). Furthermore, GSEA showed that PB@PM treatment in MIRI mice induced marked downregulation of NOD-like receptor signaling, TLR signaling, cytokine receptor interaction, chemotaxis, immune response activation, and chemokine signaling pathway (Supplementary Fig. 24B). The RNA-seq data provided compelling evidence for PB@PM’s dual regulatory effects in attenuating cardiac cell death while preserving mitochondrial integrity and modulating immuno-inflammatory homeostasis. Building upon the KEGG pathway analysis of downregulated genes, we further performed targeted validation of NF-κB and TLR signaling cascades. As a pivotal component of innate immunity, the TLR4 signaling pathway plays critical roles in the pathogenesis, progression, and clinical outcomes of cardiovascular disorders53,54. This pathway orchestrates downstream inflammatory responses through NF-κB activation, ultimately driving pro-inflammatory cytokine production. Our results indicated that PB@PM suppressed the aberrantly elevated protein levels of TLR4 and phosphorylated NF-κB-p65 in MIRI mice (Supplementary Fig. 13), suggesting its therapeutic potential in modulating TLR4/NF-κB-mediated inflammation. Collectively, these findings elucidate PB@PM’s capacity to restore immune-inflammatory equilibrium through multimodal regulatory mechanisms.
Cardiac tissue exhibits a high mitochondrial density, which serves as the primary source of reactive oxygen species (ROS) generation55. Pathological overproduction of ROS induces mitochondrial permeability transition pore activation, thereby initiating membrane depolarization and subsequent cardiomyocyte cell death. Mitochondrial ROS constitutes the principal pathogenic mechanism underlying oxidative stress during ischemia and reperfusion injury56. Given the central role of mitochondrial dysfunction and oxidative stress in initiating PANoptosis, we systematically evaluated the effects of PB@PM on mitochondrial health. Electron spin resonance (ESR) spectroscopy confirmed the potent ROS-scavenging capacity of PB@PM, demonstrating attenuation of hydroxyl (•OH) and hydroperoxyl (•OOH) radicals (Supplementary Fig. 27A–D). In OGD/R-injured HL-1 cells, PB@PM treatment markedly reduced intracellular ROS levels, as measured by flow cytometry using DCFH-DA probe (Supplementary Fig. 27F–H). Mitochondrial membrane potential (MMP), a classic indicator of mitochondrial energy status and intracellular homeostasis, was systematically evaluated across experimental groups. ODG/R induced MMP collapse in HL-1 cells, evidenced by increased JC-1 monomer fluorescence (green), indicating mitochondrial dysfunction (Supplementary Fig. 28A–C). Notably, PB@PM treatment efficiently restored MMP (Supplementary Fig. 28A–C). Most importantly, assessment of mitochondrial function using the Seahorse XF Analyzer revealed that PB@PM treatment restored oxidative phosphorylation, as evidenced by improving basal and maximal respiration, as well as ATP production in OGD/R-injured HL-1 cardiomyocytes (Supplementary Fig. 28D–G). Collectively, these findings delineate a dual mechanistic pathway for PB@PM: (1) Direct targeting: high-affinity binding and functional suppression of core PANoptosome components (RIPK1, ZBP1, and AIM2), disrupting the initiation of PANoptosis. (2) Indirect mitigation: potent ROS scavenging that preserves mitochondrial integrity and function, thereby eliminating a critical upstream trigger for PANoptosome activation. This synergistic action underpins the robust cardioprotection observed in our MIRI models.
Discussion
In addressing the critical unmet clinical need for effective PANoptosis inhibitors, we discovered that PBs simultaneously bind to RIPK1, ZBP1, and AIM2 PANoptosomes, regulate inflammation and oxidative stress, and comprehensively block crosstalk among pyroptosis, apoptosis, and necroptosis, thereby achieving efficient treatment of PANoptosis-related diseases. We identified PANoptosis as a potential therapeutic target for MIRI by combining previous research support and single-nuclear sequencing analysis of human heart samples, and established MIRI as a representative PANoptosis-driven disease model. Molecular dynamics simulations revealed robust interactions between PB and key PANoptosome components (RIPK1, ZBP1, and AIM2). To enhance myocardial targeting, PB was encapsulated with platelet membranes to construct PB@PM. In murine MIRI models, PB@PM exhibited selective accumulation in ischemic myocardium, efficiently improving cardiac function. Bulk transcriptomics and molecular validation confirmed that PB@PM inhibited PANoptosome activation, disrupted pyroptosis-apoptosis-necroptosis crosstalk, and restored mitochondrial metabolism and immune-inflammatory homeostasis, which collectively suppressed PANoptosis to alleviate MIRI.
While PANoptosis research has predominantly focused on cancer and infectious diseases, this study pioneers a multi-omics approach—integrating clinical sample analysis, molecular dynamics simulations, medical imaging, and transcriptomics—to elucidate PANoptosis mechanisms in cardiovascular pathology. Our findings underscore the potential of bioactive nanoparticles as a promising platform for developing PANoptosis-targeted therapies.
Nevertheless, this study has several limitations. First, whether PB modulates PANoptosis through other signaling pathways beyond those identified remains to be elucidated. Second, the effects of PB on non-cardiomyocyte populations within the heart, such as fibroblasts, endothelial cells, and immune cells, warrant further investigation. Third, to advance toward clinical translation, it is necessary to validate these findings in higher-order mammalian models and conduct comprehensive pharmacokinetic and toxicological studies of PB in vivo. Although our data indicate an improvement in mitochondrial function, the specific impact of PB@PM on mtDNA release has not been directly assessed and requires explicit experimental verification. In addition, while reduced cardiomyocyte death and attenuated inflammation may indirectly influence cardiac remodeling, potential direct effects of PB on fibroblast activation and extracellular matrix reorganization represent an important direction for future research. Together, these limitations and open questions provide a foundation for further exploration of PANoptosis in disease contexts and inform the rational design of next-generation nanoinhibitors.
Methods
Materials
Hydrochloric acid (HCl, 36 − 38%), potassium ferricyanide (K3[Fe(CN)6]), and polyvinylpyrrolidone (PVP, K30) were purchased from Adamas-beta (Shanghai, China). The chemicals were used as received and did not require further purification. Hydrogen peroxide (H2O2) in a 30 wt.% aqueous solution was purchased from Sinopharm Chemical Reagent Co., Ltd., China. All aqueous solutions were prepared using deionized (DI) water with a resistivity of 18.2 M cm. The Annexin V-FITC apoptosis detection kit (C1062M), propidium iodide (PI) and calcein acetoxymethyl ester (Calcein AM) kit (C2015M), mitochondrial membrane potential detection kit JC-1 (C2005), 2,7-dichlorofluorescein diacetate (DCFH-DA) kit (S0035M), reactive oxygen species assay kit (S0033M), DAPI (C1002), and TUNEL (C1088 and C1090) were purchased from Beyotime Biotechnology, China. Necrostatin-1 (HY-15760), Disulfiram (HY-B0240), Z-VAD-FMK (HY-16658B), ZBP1 (HY-P83326), phospho-RIPK1 (HY-P81539), phospho-MLKL (HY-P80468), cleaved caspase 1 (HY-P80622), cell counting kit-8 (CCK-8) assay kit (HY-K030), and cleaved caspase 8 (HY-P80624) were obtained from MedChemExpress, USA. LPS (L8880), PBST (P1031), BSA (SW3015), Tubulin (K200059M), Masson’s trichrome kit (G1340), fluorescein isothiocyanate (FITC; 3326-32-7), and Cy5.5 (S1061) were purchased from Solarbio, China. Claycomb medium (51800 C), Evans blue (314-13-6), and TTC (298-96-4) were purchased from Sigma, German. RIPA lysis buffer (WB3100), proteinase and phosphatase inhibitors (P002), and BCA Protein Assay Kit (WB6501) were purchased from NCM Biotech, China. CD41 (24552-1-AP), Collagen III (22734-1-AP), AIM2 (20590-1-AP), caspase 1 (81482-1-RR), and caspase 8 (13423-1-AP) antibodies were bought from Proteintech, China. 5-dimethyl-1-pyrroline-Noxide (D048) was bought from Dojindo, China. DMEM medium (11965092), penicillin/streptomycin (15140122), PBS (10010023), and FBS (A5670701) were bought from Gibco, USA. TBST (G2150), Ly6G antibody (GB11229), Hematoxylin and eosin (H&E) kit, and WGA (G1730) were purchased from Servicebio Co., China. Collagen I (ab316222), cTNT (ab209813), CD31 (ab182981), Pyrin (ab195975), a-SMA (ab124964), caspase 1 (ab179515), caspase 8 (ab108333), and GSDMD (ab219800) antibodies were purchased from Abcam, UK. TNF-α (EMC102a.96), IL-1β (EMC001b.96), IL-6 (EMC004.96), IL-10 (EMC005.96), cAMP (JL13362-96T), and cGMP (JL12644-96T) enzyme-linked immunosorbent assay (ELISA) kits were bought from NeoBioscience, China. CD86 (19589 s), Bax (5023 T), ASC (40002 T), caspase 3 (9662S), cleaved caspase 3 (9664 T), cleaved GSDMD (10137 T), TLR4 (14358S), NF-κB p65/RelA (8242 T), phospho-NF-κB p65/RelA (3033 T), phospho-RIPK3 (91702 T), and GAPDH (2118 T) antibodies were purchased from Cell Signaling Technology, USA. CD42b (A22277), CD47 (A21904), GP6 (A14999), Bcl-2 (A19693), TNF-α (A24214), IL-1β (A16288), IL-10 (A12255), ASC (A22046), FADD (A19838), MLKL (A26436), phospho-MLKL (AP1173), phospho-RIPK3 (AP1408), RIPK3 (A5431), phospho-RIPK1 (AP1230), RIPK1 (A7414), NLRP3 (A24294), cAMP (A1640), phospho-VASP (AP1510), VASP (A14217), AIM2 (A25252), and ZBP1 (A22612) antibodies were purchased from ABclonal, China. Three plasmids for the overexpression of mouse AIM2 mRNA, ZBP1 mRNA, and RIPK1 mRNA were synthesized by OBiO Technology (Shanghai) Corp., Ltd, and the corresponding plasmid maps were provided in Supplementary Fig. 20.
Synthesis of PB
PBs were synthesized by mixing PVP (8.0 g) and K3[Fe (CN)6] (696 mg) in HCl solution (1 M, 50 mL). The mixture was stirred for 1 h obtain a clear yellow solution, which was subsequently transferred to an electric oven and heated at 80 °C for 20 h to form a blue solution. The resulting PBs were washed five times with deionized water via centrifugation.
Design and construction of PB@PM
PB@PM were fabricated via an optimized ultrasonic fusion method57. Firstly, a 2 mL aliquot of PB solution with a concentration of 1 mg/mL was thoroughly mixed with 0.2 mg of isolated platelet membrane (quantified based on protein content). The resulting mixture was subjected to ice bath sonication at 75 W for 3 min using an ultrasonic processor (SB-5200DT, SCIENTZ, China) to promote effective membrane coating. To achieve uniform size distribution and nanoscale dimensions, the mixture was subsequently processed through a sequential extrusion protocol utilizing Nuclepore™ hydrophilic membranes with nominal pore sizes of 800 nm and 400 nm, yielding a final size of PB@PM with a hydrodynamic diameter of approximately 100 nm. The structural integrity and coating efficiency of the synthesized PB@PM were systematically characterized using TEM, molecular characterization techniques, and fluorescence co-localization imaging. Western blotting confirmed the retention of platelet-specific surface proteins (CD41, CD42b, CD47, and GP6) on PB@PM.
Characterization
The morphological analysis of PBs was characterized by Scanning Electron Microscope (SEM, Zeiss Merlin Compact, Zeiss, Germany) and Transmission Electron Microscopy (TEM, FEI Tecnai F20, USA). UV-Vis-NIR absorption spectra were acquired with a Shimadzu UV-3600i Plus spectrophotometer (Shimadzu, Japan). The Fourier-transform infrared (FT-IR) spectroscopy was performed on a Nicolet iS10 FT-IR spectrometer (Thermo Fisher Scientific, USA). X-ray photoelectron spectra (XPS) data were obtained using an EscaLab 250Xi electron spectrometer (Thermo Scientific K-Alpha, Thermo, USA). Powder X-ray diffraction (XRD) patterns were detected on a D8 ADVANCE X-ray diffractometer (Rigaku Ultima IV, Rigaku, Japan). The concentration of nanoparticles was measured by inductively coupled plasma atomic emission spectroscopy (ICP-AES, Thermo Fisher Scientific). Dynamic light scattering (DLS) measurements were conducted using a SZ-100 nanoparticle analyzer (HORIBA Scientific, Japan).
ROS scavenging evaluation
The hydroxyl radical (•OH) and hydroperoxyl radical (•OOH) scavenging capacities of PB and PB@PM were investigated using electron spin resonance (ESR) spectroscopy (Bruker ESR 5000 system, Bruker Inc.), employing 5-dimethyl-1-pyrroline-Noxide as the trapping agent. In standardized assays, DMPO (5 μL, 10 mM) was dissolved in PBS buffer (45 μL, 10 mM). The Fenton reaction was subsequently initiated by introducing Fe2+ (20 μL, 50 mM) to catalyze H2O2 (30 μL, 50 mM)-mediated •OH generation. Quantitative analysis of •OH levels was conducted through ESR amplitude intensity measurements, with comparative assessments performed under both NP-supplemented and NP-free conditions. For superoxide anion (•O2-) evaluation, 70 μg of potassium superoxide (KO₂) was reacted with 200 μL of 18-crown-6/DMSO solution (0.7 mM) to generate •O₂⁻ radicals, followed by DMPO-mediated trapping (50 μL). ESR spectral analysis of the spin-trapped adducts enabled comparative quantification of superoxide dismutase (SOD)-mimetic activity in PB and PB@PM systems.
MD simulation
MD simulations were performed utilizing the GROMACS 2022.2 computational suite. Four distinct simulation systems were constructed to characterize the interactions between PB and AIM2HIN, AIM2PYD, RIPK1, or ZBP1. Initial separation distances between the proteins and PB surfaces were maintained at 1.5 nm. Each system was solvated with SPC water molecules within a cubic simulation box (dimensions: 16.26 × 16.26 × 10.56 nm3), ensuring a minimum solvent buffer distance exceeding 12 Å from all protein surfaces. Physiological ionic conditions were mimicked by adding Na+/Cl− ions. Following system initialization, energy minimization was executed using an iterative steepest descent algorithm (a maximum of 5000 steps). Systems were energy-minimized using the steepest descent algorithm (5000 steps), followed by 20 ps NVT and NPT equilibrations to stabilize temperature (310 K) and pressure (1 bar). Force field parameters, including Amber99SB-ILDN for protein and published parameters58 for PB structures. Lennard-Jones59 interactions and short-range electrostatics were evaluated using a 10 Å cutoff, while long-range electrostatics employed the Particle Mesh Ewald (PME) method60. Periodic boundary conditions were applied in all directions, and hydrogen bonds were constrained using the LINCS algorithm. PB remained fixed during simulations to maintain interfacial stability. Production runs (200 ns, 2 fs time step) conducted to analyze the equilibrium properties.
Cell culture
HL-1 cells (a mouse cardiomyocyte cell line; CC-Y2145), RAW 264.7 cells (CC-Y2084) were purchased from the Shanghai EK-Bioscience Biotechnology Co. Ltd. HL-1 cells were cultured in Claycomb medium (Sigma) with reference to the previously reported protocol61, and RAW 264.7 cells were grown in DMEM medium (Gibco) with 10% FBS (Gibco) and 1% penicillin/streptomycin. Cultures were incubated at 37 °C in an atmosphere with 5% CO2, with daily medium replacement to maintain optimal growth conditions.
In vitro OGD/R model system and nanoparticle-based therapeutic interventions
The OGD/R injury model was established following a validated protocol62. Briefly, HL-1 cells underwent 8-hour hypoxic exposure in serum-free, glucose-derived DMEM medium (Gibco) within a modular hypoxic chamber (JSQA-50, JIANGETECH), maintaining standardized hypoxic conditions (5% CO2, 95% N2) at 37 °C with controlled humidity. Following OGD induction, cells were reperfused by replacing the hypoxic medium with complete growth medium supplemented with 10% fetal bovine serum and 25 mM glucose, followed by 12-hour normoxic recovery in a standard cell culture incubator (37 °C, 5% CO2). During the reperfusion phase, PB or PB@PM was administered to designated treatment groups, while the control group received equivalent volumes of untreated culture medium.
Cellular uptake, cytotoxicity, and cell viability assessment
The cellular internalization efficiency of PB@PM was systematically evaluated through a time-dependent fluorescence analysis. HL-1 cardiomyocytes were exposed to FITC-labeled PB@PM at specified time intervals (0.5, 1, 2, and 4 h). Quantitative determination of intracellular FITC fluorescence was performed using a Novocyte Advanteon flow cytometer (Agilent Technologies, USA) equipped with standardized cytometric analysis protocols. Cytotoxicological evaluation was conducted using a CCK-8 proliferation assay. Cells were seeded at a density of 1 × 104 cells/well in 96-well culture plates and treated with serial dilutions of PB@PM for 24 h. Cellular viability assessment followed a standardized protocol: after treatment, cells were washed with PBS buffer and incubated with 10 μL of CCK-8 solution supplemented to a final volume of 100 μL in culture medium for 1.5 h at 37 °C. The metabolic activity was quantified by measuring absorbance at 450 nm using a Spark multimode microplate reader (Tecan Group Ltd., Switzerland) with integrated wavelength correction and background subtraction functions.
Mitochondrial membrane potential (MMP) assessment
The MMP was quantitatively evaluated using JC-1 fluorescent dye following the manufacturer’s standardized protocol. Following 2 h pretreatment with PB or PB@PM formulations, OGD/R-exposed cells were incubated with JC-1 working solution at 37 °C for 30 min. Dual-mode analysis was performed using a BZ-X810 All-in-One Fluorescence Microscope (Keyence Corporation, Japan) for morphological assessment and a Novocyte Advanteon flow cytometer (Agilent Technologies, USA) for quantitative fluorescence intensity measurement.
Seahorse XF mitochondrial stress assay
Cellular oxygen consumption rates (OCR) were quantified in HL-1 cardiomyocytes utilizing the Seahorse XF Mito Stress Test protocol (Agilent Technologies). Cells were seeded at a density of 5000 cells per well in XF assay medium (pH 7.4) supplemented with 1 mM sodium pyruvate, 0.2 mM palmitic acid, 0.2 mM L-carnitine, and 4 mM glutamine. Baseline OCR measurements were acquired, followed by sequential pharmacological modulation: 3 μM oligomycin (ATP synthase inhibitor) to determine ATP-linked respiration, 1.5 μM carbonyl cyanide-p-trifluoromethoxy phenylhydrazone (FCCP) to induce maximal respiratory capacity, and 2 μM rotenone/antimycin A combination to achieve complete mitochondrial respiration inhibition. All experimental procedures adhered to validated methodologies63. Nuclear quantification was performed using Hoechst 33342 fluorescence staining for cell density determination and data normalization.
ELISA analysis
RAW264.7 macrophages were established in monolayer culture within six-well plates at a density of 2 × 10⁵ cells/well. Following 24 h stabilization, cells were treated with LPS (300 ng/mL) in conjunction with PB@PM at specified concentrations (20 or 40 μg/mL). Culture supernatants were harvested 24 h post-treatment and immediately processed for cytokine profiling. Quantitative determination of inflammatory mediators was conducted using commercially available ELISA kits (R&D Systems) following manufacturer protocols. The analyte panel included tumor necrosis factor-α (TNF-α), interleukin-1β (IL-1β), interleukin-6 (IL-6), interleukin-10 (IL-10), cyclic adenosine monophosphate (cAMP), and cyclic guanosine monophosphate (cGMP).
Apoptosis detection and intracellular ROS quantification
Cell apoptosis was quantitatively assessed using a FITC-conjugated annexin V/propidium iodide (PI) apoptosis detection kit following the manufacturer’s standardized protocol. Post OGD/R-induced injury, HL-1 cells were enzymatically dissociated into single-cell suspensions. The cell pellets were resuspended in binding buffer containing annexin V-FITC and PI, followed by 30 min incubation at room temperature in the dark. Flow cytometric analysis was performed using a Novocyte Advanteon flow cytometer (Agilent Technologies, Santa Clara, CA, USA), with data acquisition and analysis conducted through NovoExpress software. The TUNEL assay was employed to evaluate DNA fragmentation according to the manufacturer’s specifications, serving as an independent apoptotic index. Concurrently, intracellular ROS levels were quantified using dichlorofluorescin diacetate (DCFH-DA) fluorescent probes. Following 30-minute incubation with 10 μM DCFH-DA at 37 °C, cells were analyzed for oxidative stress parameters on a CytoFLEX XL flow cytometer (Beckman Colter, Brea, CA, USA) with CytExpert software for data interpretation.
Live/Dead cell viability assay
The HL-1 OGD/R cell model was employed. Cellular viability was assessed using calcein AM and PI fluorescence staining, following the manufacturer’s protocol. Following 30 min incubation with the fluorescent probes at room temperature in the dark, representative fluorescence micrographs of different experimental groups were captured using a BZ-X810 All-in-One Fluorescence Microscope (Keyence Corporation, Japan).
In vitro morphological analysis and SEM assessment
Following OGD/R induction, HL-1 cells treated with PB or PB@PM underwent standardized processing. Cell specimens were initially fixed in 4% paraformaldehyde (PFA) for 30 min, followed by sequential washing with PBS for 5 min per rinse (three repetitions). Tubulin cytoskeleton visualization was accomplished through immunofluorescent staining with tubulin-specific antibodies under dark conditions at ambient temperature for 1 h. Post-staining rinses were performed using PBS, with subsequent 15 min DAPI nuclear counterstaining in darkness. After final PBS washes (three times, 5 min each), morphological evaluations were conducted using an inverted fluorescence microscope (Keyence Corporation, Osaka, Japan). Concurrently, ultrastructural analysis of cellular membranes was performed utilizing a field emission SEM (Hitachi Regulus 8100, Tokyo, Japan) to assess membrane integrity and surface characteristics.
Plasmid overexpression assay
HL-1 cells were seeded into cell plates and divided into five groups: (1) negative control group (transfected with blank plasmid); (2) transfected with expression plasmid group; (3) transfected with expression plasmid+OGD/R group; (4) transfected with expression plasmid+PB intervention group; (5) and transfected with expression plasmid+OGD/R + PB intervention group. In brief, following the manufacturer’s protocols, confluent HL-1 cell cultures at approximately 70% confluency were transiently transfected with AIM2, ZBP1, or RIPK1 overexpression plasmids using opti-MEM reduced serum medium. Transfected cells were subsequently maintained under standard culture conditions for subsequent analysis. The OGD/R model was established in HL-1 cardiomyocytes following completion of transfection, with propidium bromide administered during the reperfusion phase for interventional assessment.
Experimental animals
Male C57BL/6 J mice (6–8 weeks of age, 20–25 g) were obtained from Shanghai BK/KY Biotechnology Co., Ltd. All animals were maintained in specific pathogen-free facilities under standardized conditions: 12 h light/dark cycles, libitum access to food and water, relative humidity of 30%, and ambient temperature (22–24 °C). Experimental protocols were approved by the Animal Care and Use Committee of Shanghai Sixth People’s Hospital affiliated to Shanghai Jiao Tong University School of Medicine (Approval No: DWSY2022-0085) and complied with guidelines from the American Association for Laboratory Animal Care. Euthanasia was performed by CO2 inhalation followed by cervical dislocation.
In vivo biosafety assessment of PB@PM
Healthy male wild-type mice (6- to 8-week-old) were randomized into three experimental groups. The control cohort received vehicle solution, while the treatment cohorts were intravenously administered PB@PM at a dosage of 5 mg/kg body weight via caudal vein injection. Histopathological evaluation was conducted at post-dosing intervals of 7 and 14 days through H&E staining of cardiac, hepatic, splenic, pulmonary, and renal tissue sections using standardized protocols (Sigma-Aldrich). Histotoxicological manifestations and parenchymal morphological characteristics were systematically analyzed microscopically. Concurrently, peripheral blood samples were collected for comprehensive hematological evaluation and serum biochemical profiling to assess systemic physiological status.
MIRI model
The murine MIRI model was established following previously validated methodology44. Briefly, mice were anesthetized via inhalation of 3% isoflurane in oxygen (1 L/min), and a ~ 2 cm thoracic incision was made at the apex of maximal cardiac pulsation. A 4-0 purse-string suture was placed, and the fourth intercostal space was bluntly dissected using curved forceps to access the thoracic cavity. Gentle pressure was applied to exteriorize the heart, which was temporarily secured within the sutured opening while minimizing pneumothorax. A 6-0 surgical slipknot was tied around the left anterior descending coronary artery, with the suture and externalized. The heart was repositioned into the thoracic cavity, residual air was evacuated, and the epidermis was closed with sutures. After 45 min of ischemia, the slipknot was released to initiate reperfusion. Surgical incisions were closed with sterile 5-0 sutures, and mice were monitored postoperatively under standard housing conditions.
Biodistribution analysis
Sham mice (Control group) received intravenous injection of Cy5.5-labeled PB, while MIRI mice were administered either Cy5.5-labeled PB (PB group) or Cy5.5-labeled PB@PM (PB@PM group). Injections were performed at 1 h post-ischemia induction. Mice were euthanized, and major organs (heart, liver, spleen, lungs, and kidneys) were harvested for ex vivo fluorescence imaging using VISQUE, in vivo Elite system at different time points.
Animal grouping and experimental protocol
The murine subjects were randomized into four experimental cohorts: Sham group, MIRI group, MIRI with PB treatment group, and MIRI with PB@PM treatment group. Pharmacological intervention was initiated at 1 and 24 h post-reperfusion via intravenous injection of either 0.5 mg/kg body weight PB or PB@PM solution (encapsulating 0.5 mg/kg PB) to the respective treatment groups. The Sham and MIRI control groups received an equivalent volume of 0.9% sodium chloride solution via the same administration route.
Echocardiography
Transthoracic echocardiography was conducted using a Vevo 3100 imaging system (FUJIFILM VisualSonics, Toronto, Canada) equipped with a 400 MHz transducer. Cardiac function was assessed preoperatively and on postoperative days 1, 14 and 2844. Mice were anesthetized with inhalation of 1–1.5% isoflurane in oxygen following chest hair removal via depilatory paste. Heart rates were maintained at 550–650 beats per minute (bpm) across all subjects. Long-axis views were captured to measure cardiac parameters, including LVEDV, LVESV, LVEF and LVFS, which were automatically calculated by the system’s integrated software.
Evans blue/TTC staining
Infarct size quantification was conducted via Evans blue/TTC dual staining44. Briefly, mice were anesthetized 3 days post-I/R surgery, the LAD coronary was re-ligated. Hearts were perfused with PBS to remove residual blood, followed by perfusion with 1% Evans blue (Sigma, > 75% purity). The hearts were subsequently excised and sectioned into serial 1.0 mm-thick transverse slices. Tissue sections were incubated in 2% TTC (Sigma-Aldrich, > 98% purity) solution at 37 °C for 15 min in the dark. Sections were fixed in 4% paraformaldehyde, and images were acquired. Evans blue-negative regions represented the area at risk (AAR), while TTC-negative zones indicated infarcted myocardium. Infarct area, AAR and LV area were quantified using ImageJ software (ImageJ 1.52 v).
Masson and HE staining
Hearts harvested 28 days post-surgery were perfused with PBS to clear blood, fixed in 4% formaldehyde, and embedded in paraffin. Sections were stained using Masson’s trichrome and HE kits (Sigma-Aldrich) to distinguish healthy tissue from fibrotic regions. Stained samples were imaged using a Nikon microscope (Nikon Corporation, Tokyo, Japan).
WGA staining
Tissue sections underwent wheat germ agglutinin (WGA) staining per the manufacturer’s protocol. Briefly, sections were washed to remove OCT compound, fixed in 4% paraformaldehyde, and incubated with 50 µg/mL WGA for 30 min at room temperature. Short-axis images of cardiomyocytes were captured, and cross-sectional areas were analyzed using ImageJ software.
Immunofluorescence staining
Hearts were excised after euthanasia, fixed in 4% paraformaldehyde overnight at 4 °C, and sectioned transversely. Sections were washed three times in PBS containing 0.1–0.2% Triton X-100 (PBST), blocked with 4% bovine serum albumin (BSA), and incubated with primary antibodies against Collagen I, Collagen III, Ly6G, CD86, cTNT, CD31, α-SMA, and TUNEL. Secondary antibody incubation and DAPI nuclear counterstaining were performed at room temperature. Images were acquired via Nikon widefield or confocal microscopes (CSU-W1, Nikon Corporation, Tokyo, Japan), and analyzed with NIS Elements AR and ImageJ software.
Western blot
Proteins were extracted from murine cardiac tissues with RIPA lysis buffer (Beyotime) supplemented with proteinase and phosphatase inhibitors. Protein concentrations were quantified using a BCA Protein Assay Kit. Proteins were separated on 10–12.5% SDS-PAGE gels and transferred onto Polyvinylidene fluoride (PVDF) membranes. Membranes were blocked with 5% skim milk in Tris-buffered saline/Tween20 (TBST, pH 7.4–7.6) for 1.5 h at room temperature, followed by overnight incubation with primary antibodies at 4 °C. After TBST washes, membranes were incubated with HRP-conjugated IgG antibody (AntiRabbit IgG (H + L) antibody, 1:1000, Beyotime) for 1 h at room temperature. Immunoreactive bands were visualized using an ECL substrate (Millipore, Burlington, MA) and detected with a fluorescent imaging system (ProteinSimple, Sata Clara, CA, USA). Image acquisition and densitometric analysis were performed using ImageJ software. Primary antibodies: Bax, Bcl-2, cleaved-caspase 3, Caspase 3, TNF-α, IL-1β, IL-10, phosphorylated MLKL (p-MLKL), total MLKL, phosphorylated RIPK1 (p-RIPK1), total RIPK1, phosphorylated RIPK3 (p-RIPK3), total RIPK3, NLRP3, cleaved-GSDMD, total GSDMD, TLR4, cAMP, NF-κB p65/RelA, phosphorylated NF-κB p65/RelA (p-p65), phosphorylated VASP (p-VASP), total VASP, AIM2, cleaved-caspase 8, caspase 8, cleaved-caspase 1, caspase 1, ZBP1, pyrin, ASC, FADD, and GAPDH.
RNA-seq analysis
To investigate PB@PM’s therapeutic mechanisms in myocardial ischemia-reperfusion injury, RNA sequencing was performed on cardiac tissues from three groups: Control, MIRI, and PB@PM-treated mice (n = 5 per group). Hearts were quickly collected on postoperative day 3, then washed with cold saline, frozen in liquid nitrogen, followed by kept at − 80 °C. Total RNA was extracted, and its integrity/quantity was assessed using an RNA Nano 6000 Assay Kit of the Bioanalyzer 4200 system (Agilent Technologies, CA, USA). High-quality RNA samples were used to construct the sequencing library. The library preparation for transcriptome sequencing, clustering, and sequencing was performed using Illumina NovaSeqX-Plus (Illumina) according to the manufacturer’s instructions. Differential expression analysis was conducted using the Seurat package through implementation of the Wilcoxon rank-sum test with default parameters. DEGs were identified based on three criteria: (1) expression in more than 10% of cells within each compared group, (2) average log2-fold change exceeding 1, and (3) statistical significance determined by Adjusted p-value (adjusted p < 0.05), with multiple testing correction via the Bonferroni method.
Single-nucleus RNA sequencing (snRNA-seq) analysis of human cardiac tissues
A publicly available snRNA-seq dataset from patients with acute myocardial infarction was reanalyzed (https://cellxgene.cziscience.com/collections/8191c283-0816-424b-9b61-c3e1d6258a77). Clinical sample metadata are detailed in Supplementary Table 1.
(1) Initial processing and quality control
snRNA-seq data preprocessing was performed using CeleScope v2.0.7 (Singleron Biotechnologies) with default settings. Briefly, barcodes and UMIs were extracted from R1 reads and error-corrected. Adapter sequences and poly-A tails were trimmed from R2 reads, followed by alignment to the GRCh38 transcriptome via STAR (v2.7.11a). Uniquely mapped reads were assigned to genes, and cells were defined by the co-occurrence of cell barcodes, UMIs, and gene expression. Gene expression matrices were generated for downstream analysis.
(2) Quality control, dimensionality reduction and clustering
Data processing was executed in Python 3.7 using Scanpy v1.8.1. Cells were filtered based on: (1) Minimum gene count ≥200 and exclusion of top 2% highest gene counts; (2) Exclusion of top 2% highest UMI counts; 3) Mitochondrial gene content ≤ 10%; 4) Gene detection threshold ≥5 cells. Post-filtering, 64804 cells remained (1328 genes/cell; 3963 UMIs/cell). Data were normalized to total counts per cell, log-transformed, and scaled. Top 2000 highly variable genes (HVGs) were identified using Seurat’s “seurat” flavor. Principal component analysis (PCA) was performed on scaled HVGs, with the top 20 principal components (PCs) retained for downstream analysis. Unsupervised clustering was implemented via the Leiden algorithm (resolution = 1.2), and UMAP embeddings were generated for visualization64.
(3) Batch effect correction
Cross-sample heterogeneity was mitigated using Harmony v1.0, integrating the top 20 PCs from PCA2 to generate batch-corrected embeddings65.
(4) Differential expression gene (DEG) analysis
DEGs were identified using Scanpy’s rank_genes_groups function with Wilcoxon rank-sum tests. Genes meeting criteria (detected in ≥ 10% of cells within any group; |log2(Fold Change)| > 1) were selected. Multiple testing correction was applied via Benjamini-Hochberg (FDR < 0.05).
(5) Cell-type annotation
Cell-ID annotation: Cell identity was assigned using Cell-ID, a multivariate computational approach that computes cell-type specificity via hypergeometric tests (HGT)66. Gene expression matrices were reduced to latent dimensions via multiple correspondence analysis (MCA), projecting cells and genes into a unified space. Cell-specific gene signatures were derived from SynEcoSys™ (Singleron Biotechnologies), a curated database integrating canonical markers from CellMarkerDB, PanglaoDB, and literature. HGT scores were calculated for each cluster against reference cell types, with minimal HGT P-values defining cluster identities. Reference-based annotation: Canonical marker expression was cross-referenced against SynEcoSys™, ensuring annotation consistency. Cluster-specific cell-type frequencies were computed, and predominant cell types were assigned as cluster labels.
(6) Cardiomyocyte subtyping: High-resolution subtyping of cardiomyocytes was achieved by re-clustering cells from the parent cardiomyocyte cluster using identical analytical pipelines67.
(7) Gene Set Scoring with UCell
Gene set enrichment was quantified via UCell v1.1.068, a rank-based scoring methodology advantageous for large-scale datasets. Mann-Whitney U statistics evaluated query gene expression ranks across individual cells.
Statistical analysis
Data are presented as means ± standard deviation (SD). For single-nucleus RNA-seq data, wilcox.test was applied. Intergroup differences among three or more groups were assessed by one-way ANOVA with Tukey’s post-hoc correction. Statistical significance was defined as P < 0.05. Analyses were carried out with GraphPad Prism 9.0 (GraphPad Software, San Diego, CA) and SPSS software 25.0 (SPSS Inc., USA).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All generated or analyzed data supporting the findings of this study are available within the paper, its Supplementary Information, and Source Data. The bulk RNA sequencing data generated in this study have been deposited to the Gene Expression Omnibus database under accession code GSE298751. The single nuclear data used in this study are available at: https://cellxgene.cziscience.com/collections/8191c283-0816-424b-9b61-c3e1d6258a77. Source data are provided in this paper.
References
Kanneganti, T.-D. Intracellular innate immune receptors: Life inside the cell. Immunol. Rev. 297, 5–12 (2020).
Sun, X. et al. PANoptosis: Mechanisms, biology, and role in disease. Immunol. Rev. 321, 246–262 (2024).
Sundaram, B. et al. NLRP12-PANoptosome activates PANoptosis and pathology in response to heme and PAMPs. Cell 186, 2783–2801 (2023).
Sundaram, B. et al. NLRC5 senses NAD plus plus depletion, forming a PANoptosome and driving PANoptosis and inflammation. Cell 187, 4061–4077.e17 (2024).
Wang, Y. & Kanneganti, T. D. From pyroptosis, apoptosis and necroptosis to PANoptosis: A mechanistic compendium of programmed cell death pathways. Comput. Struct. Biotechnol. J. 19, 4641–4657 (2021).
Shi, C. X. et al. PANoptosis: A cell death characterized by pyroptosis, apoptosis, and necroptosis. J. Inflamm. Res. 16, 1523–1532 (2023).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. 1, aag2045 (2016).
Pandeya, A. & Kanneganti, T. D. Therapeutic potential of PANoptosis: innate sensors, inflammasomes, and RIPKs in PANoptosomes. Trends Mol. Med. 30, 74–88 (2024).
Henkel, F. D. R. & O’Neill, L. A. J. NLRP12 drives PANoptosis in response to heme. Trends Immunol. 44, 574–576 (2023).
Lee, S. J. et al. AIM2 forms a complex with pyrin and ZBP1 to drive PANoptosis and host defence. Nature 597, 415–419 (2021).
Banoth, B. et al. ZBP1 promotes fungi-induced inflammasome activation and pyroptosis, apoptosis, and necroptosis (PANoptosis). J. Biol. Chem. 295, 18276–18283 (2020).
Karki, R. et al. ZBP1-dependent inflammatory cell death, PANoptosis, and cytokine storm disrupt IFN therapeutic efficacy during coronavirus infection. Sci. Immunol. 8, eabo6294 (2022).
Zheng, M. & Kanneganti, T. D. The regulation of the ZBP1-NLRP3 inflammasome and its implications in pyroptosis, apoptosis, and necroptosis (PANoptosis). Immunol. Rev. 297, 26–38 (2020).
Zhang, X.-J. et al. Pharmacological inhibition of arachidonate 12-lipoxygenase ameliorates myocardial ischemia-reperfusion injury in multiple species. Cell Metab 33, 2059–2075.e2010 (2021).
Thiele, H. et al. One-Year Outcomes after PCI Strategies in Cardiogenic Shock. N. Engl. J. Med. 379, 1699–1710 (2018).
Eltzschig, H. K. & Eckle, T. Ischemia and reperfusion—from mechanism to translation. Nat. Med. 17, 1391–1401 (2011).
MacLellan, W. R. & Schneider, M. D. Death by design. Programmed cell death in cardiovascular biology and disease. Cir. Res. 81, 137–144 (1997).
Zhang, T. et al. CaMKII is a RIP3 substrate mediating ischemia- and oxidative stress–induced myocardial necroptosis. Nat. Med. 22, 175–182 (2016).
Toldo, S. et al. Interleukin-1β blockade improves cardiac remodelling after myocardial infarction without interrupting the inflammasome in the mouse. Exp. Physiol. 98, 734–745 (2013).
Li, C. et al. Metformin mediates cardioprotection against aging-induced ischemic necroptosis. Aging Cell 19, e13096 (2020).
Oerlemans, M. I. F. J. et al. Inhibition of RIP1-dependent necrosis prevents adverse cardiac remodeling after myocardial ischemia–reperfusion in vivo. Basic Res. Cardiol. 107, 270 (2012).
Qi, Z. H., Zhu, L. L., Wang, K. K. & Wang, N. PANoptosis: Emerging mechanisms and disease implications. Life Sci. 333, 122158 (2023).
Pandian, N. & Kanneganti, T. D. PANoptosis: A Unique Innate Immune Inflammatory Cell Death Modality. J. Immunol. 209, 1625–1633 (2022).
Malireddi, R. K. S. et al. TAK1 restricts spontaneous NLRP3 activation and cell death to control myeloid proliferation. J. Exp. Med. 215, 1023–1034 (2018).
Kesavardhana, S. et al. ZBP1/DAI ubiquitination and sensing of influenza vRNPs activate programmed cell death. J. Exp. Med. 214, 2217–2229 (2017).
Xiang, Q. et al. PANoptosis: a novel target for cardiovascular diseases. Trends Pharmacol. Sci. 45, 739–756 (2024).
McArthur, K. et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science 359, eaao6047 (2018).
Zheng, Z. et al. Enhancing myocardial infarction treatment through bionic hydrogel-mediated spatial combination therapy via mtDNA-STING crosstalk modulation. J. Control Release 371, 570–587 (2024).
Wu, J. X. et al. Cyclic GMP-AMP Is an Endogenous Second Messenger in Innate Immune Signaling by Cytosolic DNA. Science 339, 826–830 (2013).
Xian, H. X. et al. Oxidized DNA fragments exit mitochondria via mPTP- and VDAC-dependent channels to activate NLRP3 inflammasome and interferon signaling. Immunity 55, 1370–1385.e8 (2022).
Messaoud-Nacer, Y. et al. STING agonist diABZI induces PANoptosis and DNA mediated acute respiratory distress syndrome (ARDS). Cell Death Dis. 13, 269 (2022).
Bi, Y. G. et al. FUNDC1 protects against doxorubicin-induced cardiomyocyte PANoptosis through stabilizing mtDNA via interaction with TUFM. Cell Death Dis. 13, 1020 (2022).
Zhang, T. et al. Metabolic orchestration of cell death by AMPK-mediated phosphorylation of RIPK1. Science 380, 1372–1380 (2023).
Zhang, N. et al. Palmitoylation licenses RIPK1 kinase activity and cytotoxicity in the TNF pathway. Mol. Cell 84, 4419–4435.e10 (2024).
Geng, L. L., Zhang, G., Yao, M. & Fang, Y. Rip 1-dependent endothelial necroptosis participates in ischemia-reperfusion injury of mouse flap. J. Dermatol. Sci. 97, 30–40 (2020).
Ma, X. et al. Prussian Blue Nanozyme as a Pyroptosis Inhibitor Alleviates Neurodegeneration. Adv. Mater. 34, 2106723 (2022).
Hou, R. et al. Prussian blue nanozyme promotes the survival rate of skin flaps by maintaining a normal microenvironment. ACS Nano 16, 9559–9571 (2022).
Zhang, K. et al. Hollow prussian blue nanozymes drive neuroprotection against ischemic stroke via attenuating oxidative stress, counteracting inflammation, and suppressing cell apoptosis. Nano Lett. 19, 2812–2823 (2019).
Wang, B., Bhattacharya, M., Roy, S., Tian, Y. & Yin, Q. Immunobiology and structural biology of AIM2 inflammasome. Mol. Aspects Med. 76, 100869 (2020).
Zhang, H. et al. Crucial roles of the RIP homotypic interaction motifs of RIPK3 in RIPK1-dependent cell death and lymphoproliferative disease. Cell Rep. 31, 107650 (2020).
Mompeán, M. et al. The structure of the necrosome RIPK1-RIPK3 core, a human hetero-amyloid signaling complex. Cell 173, 1244–1253.e1210 (2018).
Kelliher, M. A. et al. The death domain kinase RIP mediates the TNF-induced NF-κB signal. Immunity 8, 297–303 (1998).
Ito, Y. et al. RIPK1 mediates axonal degeneration by promoting inflammation and necroptosis in ALS. Science 353, 603–608 (2016).
Fan, Q. et al. Dectin-1 contributes to myocardial ischemia/reperfusion injury by regulating macrophage polarization and neutrophil infiltration. Circulation 139, 663–678 (2019).
Wynn, T. A. & Ramalingam, T. R. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat. Med. 18, 1028–1040 (2012).
Lindsey, M. L. et al. Effect of a cleavage-resistant collagen mutation on left ventricular remodeling. Circ. Res. 93, 238–245 (2003).
Shamhart, P. E. & Meszaros, J. G. Non-fibrillar collagens: Key mediators of post-infarction cardiac remodeling? J. Mol. Cell Cardiol. 48, 530–537 (2010).
Prabhu, S. D. & Frangogiannis, N. G. The biological basis for cardiac repair after myocardial infarction. Circ. Res. 119, 91–112 (2016).
Cao, D. J. et al. Cytosolic DNA sensing promotes macrophage transformation and governs myocardial ischemic injury. Circulation 137, 2613–2634 (2018).
Falcon, D. et al. Inhibition of adenylyl cyclase 8 prevents the upregulation of Orai1 channel, which improves cardiac function after myocardial infarction. Mol. Ther. 32, 646–662 (2024).
Hu, M. et al. Isthmin-1 alleviates cardiac ischaemia/reperfusion injury through cGMP-PKG signalling pathway. Cardiovasc. Res. 120, 1051–1064 (2024).
Chen, S. et al. PDE10A Inactivation prevents doxorubicin-induced cardiotoxicity and tumor growth. Circ. Res. 133, 138–157 (2023).
Frustaci, A. et al. Myocarditis-associated necrotizing coronary vasculitis: incidence, cause, and outcome. Eur. Heart J. 42, 1609–1617 (2021).
Balistreri, C. R. et al. Role of toll-like receptor 4 in acute myocardial infarction and longevity. J. Am. Med. Assoc. 292, 2339–2340 (2004).
Kornfeld, O. S. et al. Mitochondrial reactive oxygen species at the heart of the matter. Circ. Res. 116, 1783–1799 (2015).
Chen, Y.-R. & Zweier, J. L. Cardiac mitochondria and reactive oxygen species generation. Circ. Res. 114, 524–537 (2014).
Li, B. et al. A nanocapsule system combats aging by inhibiting age-related angiogenesis deficiency and glucolipid metabolism disorders. ACS Nano 18, 21061–21076 (2024).
Sazman, F. N. et al. First-principles approach to the structural, electronic and intercalation voltage of Prussian blue (KxFe[Fe(CN)6]) (x=1, 2) as potential cathode material for potassium ion batteries. J. Solid State Electr. 27, 1095–1106 (2023).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald - An N.Log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Papait, R. et al. Histone methyltransferase G9a is required for cardiomyocyte homeostasis and hypertrophy. Circulation 136, 1233–1246 (2017).
Ye, T. B. et al. Protective effects of Pt-N-C single-atom nanozymes against myocardial ischemia-reperfusion injury. Nat. Commun. 15, 1682 (2024).
Luongo, T. S. et al. The mitochondrial Na+/Ca2+ exchanger is essential for Ca2+ homeostasis and viability. Nature 545, 93–97 (2017).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Cortal, A., Martignetti, L., Six, E. & Rausell, A. Gene signature extraction and cell identity recognition at the single-cell level with Cell-ID. Nat. Biotechnol. 39, 1095–1102 (2021).
Zhang, Y. et al. SynEcoSys: a multifunctional platform of large-scale single-cell omics data analysis. Preprint at https://doi.org/10.1101/2023.02.14.528566 (2023).
Andreatta, M. & Carmona, S. J. UCell: Robust and scalable single-cell gene signature scoring. Comput. Struct. Biotec. 19, 3796–3798 (2021).
Acknowledgements
This work was supported by the National Natural Science Foundation of China Major Research Program (grant number T2394534, to Y.Z.), National Natural Science Foundation of China (grant number 82172074, to X.C.), Shanghai Jiao Tong University School of Medicine’s “Clinical Professional Research Team” Project (SBR2022006, to X.C.), Shanghai Jude Charitable Foundation, and the Innovative Research Team of high-level local universities in Shanghai (to Y.Z.).
Author information
Authors and Affiliations
Contributions
L.X., L.J., and R.W. contributed equally to this work. Y.Z., X.C., and B.L. conceived and designed the study; Y.Z., X.C., and B.L. interpreted the results; L.X., L.J., R.W., H.Y., and B.L. conducted experiments and performed data analysis; R.W. carried out molecular dynamics simulation analysis; X.C., B.L., and L.X. drafted the manuscript. All authors collectively discussed the results and provided critical feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Rongsheng Su and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, L., Jiang, L., Wu, R. et al. Prussian blue nanoparticles targeting multiple PANoptosome-mediated PANoptosis for myocardial ischemia-reperfusion injury therapy. Nat Commun 17, 2329 (2026). https://doi.org/10.1038/s41467-026-70012-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70012-2