Abstract
Secondary resistance to the amino-acid–depleting agent L-asparaginase (ASNase) remains poorly understood. Using ASNase-sensitive B-cell lymphoma (BCL) models, we investigate tumor relapse during treatment. Through in vitro and in vivo metabolic profiling, here we show that ASNase triggers a metabolic reprogramming characterized by increased de novo serine biosynthesis driven by phosphoglycerate dehydrogenase (PHGDH). This response mitigates treatment-induced oxidative stress and associated DNA damage, enabling malignant cells to survive. We evidence that ASNase-treated malignant cells exhibit features of replication stress and increase activity of poly(ADP-ribose) polymerase (PARP), revealing a dependence on DNA repair. Combining ASNase with the clinically approved PARP inhibitor Olaparib enhances the antineoplastic effect of each monotherapy in vitro and in vivo. Moreover, this combination shows effectiveness in homologous recombination-proficient colorectal cancer cells, suggesting broader therapeutic potential. Overall, our study identifies tumor metabolic and genomic vulnerabilities induced by ASNase and supports a rational combination strategy using clinically approved drugs.
Similar content being viewed by others
Introduction
Cancer cells exhibit enhanced metabolism to meet expanding energetic and biosynthetic demands, a vulnerability that prompted several laboratories to develop metabolic inhibitors for clinical use in oncology1,2,3. Despite successful evaluations on preclinical models of cancers, only a few of these inhibitors have achieved FDA approval, due in part to significant toxicities and development of resistance mechanisms1,4. The intrinsic mechanisms by which tumor cells adapt to metabolic targeting in vivo remain poorly understood, thereby limiting the development of additional therapeutic strategies for patients who exhibit resistance to routinely used anti-metabolic agents in the clinic.
L-asparaginase (ASNase) is the only clinically approved drug targeting cancer cell addiction for a particular amino acid, successfully integrated into multi-agent chemotherapy regimens to treat both childhood and adult T-cell and B-cell acute lymphoblastic leukemias (T-ALL, B-ALL) and NK/T-cell lymphomas (NKTCL). By catalyzing the hydrolysis of the non-essential amino acid asparagine in the bloodstream, ASNase triggers starvation and selective apoptosis of asparagine-addicted cancer cells. To date, clinical use of ASNase is restricted to ALL and NKTCL, as these hematological malignancies commonly harbor an epigenetic silencing of the gene encoding the asparagine synthetase that catalyzes the ATP-dependent conversion of L-aspartate (Asp) and L-glutamine (Gln) into L-asparagine (Asn) and L-glutamate (Glu)5,6,7. Consequently, most ALL and NKTCL are asparagine-auxotrophic, which makes them highly vulnerable to ASNase treatment. Therefore, asparagine synthetase has long been considered a primary determinant of cancer cells’ sensitivity to ASNase treatment6. However, recent studies have shown that ASNase treatment also exhibits anti-tumor efficacy in a variety of other malignancies displaying basal heterogenous asparagine synthetase expression8,9,10,11,12. Moreover, our recent work demonstrated that extracellular asparagine prevents its de novo biosynthesis in B-cell lymphomas, regardless of asparagine synthetase expression levels9. These findings challenge the conventional view that asparagine synthetase expression alone is a reliable factor predicting tumor sensitivity to ASNase treatment in asparagine synthetase-expressing malignancies9.
Several mechanisms involved in tumor resurgence during ASNase treatment have been described. Due to its bacterial nature, immunization against ASNase may occur during therapy, leading to neutralization of the enzyme activity. This issue is being addressed through the development of 2nd generation of ASNase formulations with reduced immunogenicity13,14,15. Additional mechanisms primarily involve the upregulation and/or activation of asparagine synthetase in malignant cells, driven by activation of multiple distinct signaling pathways that converge to an ATF4 (Activating Transcription Factor 4)-dependent transcriptional program promoting cell growth10,11,12. Nutrient release by stromal cells, which fuels leukemic blasts16,17 along with specific tumor metabolic adaptations, also accounts for additional reported mechanisms of resistance to ASNase treatment18,19,20. However, none have yet led to clinically viable alternative treatments for patients who fail ASNase therapy.
We recently demonstrated the antitumor efficacy of an ASNase-based anti-metabolic strategy in patients with refractory/relapsed (R/R) diffuse large B-cell lymphoma (DLBCL)8. Despite complete responses achievement, all patients eventually relapsed during or after completed therapy, suggesting that a subset of malignant cells survived and adapted to circumvent treatment.
Our study aimed to uncover additional cancer cell intrinsic metabolic alterations that contribute to tumor relapse during ASNase treatment, with the intent to reveal unknown targetable vulnerabilities. Using a preclinical mouse model of B-cell lymphomas (BCL), we modeled secondary resistance following an initial antineoplastic response to ASNase treatment both in vitro and in vivo. Through comprehensive metabolomic profiling of BCL, complemented by in-depth analysis of altered metabolic pathways, using the stable isotope tracing in both in vitro and in vivo settings, we evidenced that ASNase treatment induces a dynamic metabolic reprogramming characterized by increased de novo serine biosynthesis in malignant cells. Using pharmacological inhibition, shRNA-mediated silencing, and overexpression of PHGDH—the rate-limiting enzyme of the de novo serine biosynthesis—we demonstrated that PHGDH plays a critical role in mediating the tumor’s adaptive response to ASNase. Mechanistically, PHGDH activity mitigates ASNase-induced oxidative stress and related DNA damage, thereby enabling malignant cells to resume proliferation during treatment. Importantly, ASNase-induced DNA damage enhances reliance of malignant B cells on poly(ADP ribose) polymerase 1/2 (PARP) activity, revealing a therapeutic vulnerability to the clinically approved PARP inhibitor, Olaparib, which impairs DNA repair mechanisms.
Results
ASNase-sensitive B-cell lymphomas undergo metabolic reprogramming during ASNase therapy both in vitro and in vivo
Previously, we demonstrated that B-cell lymphomas (BCL) relying on oxidative phosphorylation (OxPhos) metabolism for energy production are sensitive to E-coli L-asparaginase (ASNase) therapy, whereas glycolytic-dependent BCL exhibit resistance8. In the present study, we engrafted wild-type C57BL/6 mice with OxPhos-dependent Eµ-Myc cells (malignant B cells) isolated from two individual transgenic Eµ-MycTg/+ mice (#506 or #688). Seven days later, mice were treated either with Vehicle or ASNase every 48 h until the lymphoma reached the ethical endpoint (Fig. 1a, b). ASNase treatment showed significant anti-tumor efficacy resulting in delayed BCL development (Figs. 1a, b and S1a, b). This represents the initial antineoplastic response to treatment. However, despite continued treatment, all mice eventually developed BCL, a result consistently observed across two independent Eµ-Myc clones (#506 or #688) showing equivalent initial sensitivity to ASNase treatment in vivo. This secondary response reflects a phase of therapeutic failure, mimicking tumor relapse following an initial favorable response, as observed in patients with R/R DLBCL treated with innovative ASNase-based anti-metabolic therapy8.
a Survival curves of mice bearing OxPhos-dependent Eμ-Myc #506 cell-derived B-cell lymphomas (BCL), treated with Vehicle or ASNase every 48 h till disease endpoint (Vehicle, n = 25 mice; ASNase, n = 26 mice, n = 3 independent experiments). b As in a with OxPhos-dependent Eµ-Myc #688 cells (n = 16 mice/group, n = 2 independent experiments). c Principal-component analysis (PCA) of 136 detected metabolites in BCL from C57BL/6 mice engrafted with Eµ-Myc #506 cells and treated with Vehicle or ASNase from day 7 until endpoint (n = 10 mice/group). d As in (c) using mice engrafted with Eµ-Myc #688 cells (based on 131 metabolites detected) (n = 12 mice/group). e Volcano plot highlighting significantly deregulated metabolites (Fold change (ASNase/Vehicle)>1.25 and raw P-value < 0.05) in Eµ-Myc #506 cell-derived BCL described in (c). (n = 10 mice/group). Nucleotide precursors in blue, proteinogenic amino acid in pink. f Venn diagram illustrating the number of statistically significant deregulated metabolites in ASNase-treated BCL highlighted in (e), S1e. g Overview of top statistically significant enriched KEGG metabolite sets from 19 commonly deregulated metabolites highlighted in (f) in ASNase-treated BCL. Relative abundance (peak area) of total intracellular serine (h) and glycine (i) in BCL presented in (c) and in (d) (from Eµ-Myc #506 cells, n = 10 mice/group; from Eµ-Myc #688 cells, n = 12 mice/group). j Percentage of dead Eμ-Myc #506 cells (DAPI+) cultivated for 24 h in Asn-free medium supplemented with (+) or without (−) Asn (0.37 mM) and ASNase (0.003 IU/ml) (n = 6 independent experiments). k Proliferation of Eµ-Myc (#506) cells cultivated as in (j). (n = 6 independent experiments). Relative abundance (peak area) of total intracellular serine (l) and glycine (m) in live Eµ-Myc (#506) cells cultivated for 24 h as in (j) (n = 4 biological and two technical replicates). Data are expressed as mean ± SD (h, i, j, l, m) or ± SEM (k). P-values are from log-rank test (a, b), non-parametric Wilcoxon rank-sum test (e), Fisher’s exact test followed by Bonferroni correction (g), t-test (h, i), one-way Anova followed by Tukey’s test (j, l, m) and indicated as **, p < 0.01; ***, p < 0.001; ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
One mechanism of tumor resurgence during ASNase treatment in clinical settings is the production of anti-ASNase antibodies, which neutralize the enzyme and are often associated with allergic episodes in patients. To rule out decreased ASNase activity as the cause of tumor relapse, mice received a final bolus of Vehicle or ASNase at endpoint. Four hours post administration, BCL-bearing mice treated with ASNase exhibited undetectable plasma asparagine concentration and elevated plasma aspartate concentration compared to Vehicle-treated controls, indicating sustained ASNase activity despite disease relapse (Fig. S1c).
Regardless of the tumor entity or energetic dependency, therapy-resistant cancer cells frequently enhance mitochondrial energetic functions to promote survival21,22,23,24. Consistently, we observed a significant increase in the contribution of OxPhos to ATP production in BCL that progressed during ASNase treatment in vivo, suggesting a metabolic reprogramming to meet elevated ATP demands (Fig. S1d).
We next conducted targeted metabolomic analysis in BCL undergoing Vehicle or ASNase treatment in vivo. Principal component analysis (PCA) and Venn diagrams showed that the metabolic profile of BCL treated with ASNase in vivo was distinct from that of Vehicle-treated BCL, irrespective of the Eµ-Myc clone used (Figs. 1c–e and S1e). Nineteen (14%) significantly deregulated metabolites were common to both datasets (Eµ-Myc #506 cell- or Eµ-Myc #688 cell-derived BCL) and exhibited similar directional regulation in response to ASNase treatment, with 14 metabolites increased and 5 metabolites decreased (Fig. 1f and Supplementary Data S1). These shared metabolites likely reflect core metabolic responses to ASNase therapy that are independent of the Eµ-Myc clone origin, suggesting conserved metabolic vulnerabilities in our in vivo model.
Metabolite Set Enrichment Analysis (MSEA) using the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway computational tool, revealed deregulated nucleotide and amino acid (AA) metabolism in ASNase-treated BCL (Fig. 1g). Among proteinogenic AA, serine, glycine, tyrosine, and asparagine were the only significantly and consistently deregulated AA in ASNase-treated BCL, meeting the fold change threshold (ASNase/Vehicle) > 1.25 and raw P-value < 0.05, across both datasets (Figs. 1e and S1e, Tables S1 and S2). Indeed, BCL treated with ASNase in vivo exhibit a two- to three-fold increase in steady-state serine and glycine levels in ASNase-treated BCL compared to Vehicle-treated BCL (Fig. 1h, i).
To confirm that ASNase treatment triggered serine metabolism remodeling in malignant cells, Eµ-Myc cells were cultured in DMEM medium lacking asparagine but containing supraphysiological concentrations of serine and glycine, with (+) or without (−) supplementation of asparagine and ASNase. A concentration of 0.003 IU/ml ASNase was used to avoid hydrolysis of glutamine into L-glutamate (Fig. S1f). ASNase treatment significantly increased the death of Eµ-Myc cells in 24 h (Figs. 1j and S1g, h). However, Eµ-Myc cells surviving this nutritional stress resumed proliferation over time (Figs. 1k et S1i), a process accompanied by a significant increase in intracellular serine and glycine levels following 24 h of ASNase treatment (Figs. 1l, m and S1j, k). To specifically evaluate metabolic adaptations to asparagine deprivation—since ASNase-mediated release of aspartate and ammonium can each influence the tumor metabolic response—we cultured Eµ-Myc cells in asparagine-free medium. This experimental setup ensured that the observed metabolic changes were directly attributable to asparagine depletion, thereby reinforcing the robustness of our conclusions. Our findings indicate that ASNase treatment induces a metabolic response in BCL, both in vitro and in vivo, characterized by elevated serine and glycine levels in malignant B cells.
Increased de novo serine biosynthesis in ASNase-treated B-cell lymphomas, in vitro and in vivo
Increased total serine levels in malignant cells may result from either enhanced serine import via active transport or increased de novo serine biosynthesis from the glycolytic intermediate 3-phosphoglycerate and glutamine-derived glutamate through a 3-step enzymatic process catalyzed by phosphoglycerate dehydrogenase (PHGDH), phosphoserine aminotransferase-1 (PSAT1), and phosphoserine phosphatase (PSPH). Eµ-Myc cells proliferation was unaffected by extracellular L-serine/glycine withdrawal, regardless of asparagine availability (Fig. S2a). In line with a previous report describing asparagine as an amino acid exchange factor facilitating serine import25, asparagine-restricted Eµ-Myc cells exhibited significantly reduced extracellular serine uptake, and an increase in total intracellular serine levels (Figs. 2a, 1l and S1j). Moreover, incubation of Eµ-Myc cells in serine/glycine-free medium still resulted in a marked and significant increase in intracellular serine and glycine abundance in ASNase-treated cells compared to control cells, suggesting activation of the de novo serine and glycine biosynthesis pathway upon treatment (Fig. 2b, c). Supporting this hypothesis, asparagine-deprived Eµ-Myc cells upregulated PHGDH, PSAT1, PSPH mRNA and protein expression, along with the expected induction of asparagine synthetase (Figs. 2d, e and S2b). The glutamine synthase (GLUL) that has been associated with resistance of sarcoma cells to ASNase18,19, was upregulated only in glutamine-deprived, but not in asparagine-deprived Eµ-Myc cells, suggesting a weak contribution of GLUL to secondary resistance to ASNase treatment in malignant B cells. The expression level of the glutamate oxaloacetate transaminase 1 (GOT1), which remained unchanged upon nutritional stress, indicates a selective regulation of specific metabolic enzymes consistent with upregulation of the transcription factor ATF4 (Figs. 2d, e and S2c), as already described26,27,28,29.
a L-[3H(G)]-serine transport rate (cpm/106 cells) in Eµ-Myc (#688) cells following 24 h incubation in serine/glycine-containing medium, with (+) or without (−) Asn (0.37 mM) and ASNase (0.003 IU/ml) (left, 15 min uptake, n = 4 independent experiments; right, 3 h uptake, n = 3 independent experiment). Relative abundance (peak area) of total intracellular serine (b) and glycine (c) levels in Eµ-Myc (#688) cells cultivated for 24 h in Asn/serine/glycine-free medium, supplemented (+) or not (−) with serine, glycine (Ser/Gly; 0.4 mM/0.4 mM), Asn (0.37 mM) and ASNase (0.003 IU/ml) (n = 3 biological replicates). d Total protein extracts prepared from live Eµ-Myc #506 (left) and #688 (right) cells cultivated for 24 h in glutamine (Gln) and Asn-free medium supplemented (+) or not (−) with Gln (2 mM), Asn (0.37 mM) or ASNase (0.003 IU/ml), were immunoblotted for the indicated proteins. The samples derive from the same experiments, but different gels for PHGDH, PSPH, ERK2, another for PSAT1, ASNS, another for ATF4, another for GLUL, and another for GOT1 were processed in parallel. e Relative quantification of immunoblots presented in (d). (#506 cells, square, n = 2 independent experiments; #688 cells, triangle, n = 2 independent experiments). Relative abundance (peak area) of 13C-labelled serine M + 3 (f) and glycine M + 2 (g) isotopologues in Eµ-Myc (#688) cells cultivated for 24 h in glucose and asparagine-free medium supplemented with 25 mM [U-13C]-glucose and with (+) or without (−) Asn (0.37 mM) and ASNase (0.003 IU/ml) (n = 4 biological and two technical replicates). h Total protein extracts prepared from BCL harvested in Eµ-Myc (#506) cells-bearing C57BL/6 mice, treated with Vehicle or ASNase every 48 h till disease endpoint, were immunoblotted for the indicated proteins (Vehicle, n = 4 mice; ASNase, n = 10 mice). The samples derive from the same experiments, but different gels for PHGDH, PSAT1, ERK2, another for PSPH, another for ASNS, and another for ATF4 were processed in parallel. i Percentage of PHGDH activity in BCL harvested in Eµ-Myc cells-bearing C57BL/6 mice treated as in (h) (n = 9 mice/group). j Schematic representation of in vivo [U-13C]-glucose consecutive bolus protocol performed in Vehicle and ASNase-treated C57BL/6 mice bearing OxPhos-dependent Eµ-Myc #688 cell-derived BCL. k Relative abundance (peak area) of 13C-labelled serine (M + 3) and glycine (M + 2) isotopologues in BCL of mice presented in (j) (n = 4 mice/group). For all graphs, data are expressed as mean ± SD. P-values are from one-way Anova followed by Tukey’s test (a, f, g), or followed by Dunnett’s test (e), 2way Anova followed by Tukey’s t-test (b, c), t-test (i., k.) and indicated as ns, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
Given that the expression level of metabolic enzyme does not necessarily reflect its activity9, we assessed the activity of PHGDH—the rate-limiting enzyme in de novo serine biosynthesis—as well as the level of de novo-synthesized serine and glycine by tracing [U-13C]-glucose in Eµ-Myc cells in vitro. We observed a significant increase in PHGDH activity (Fig. S2d), along with elevated levels of de novo 13C-labelled serine (M + 3) and 13C-labelled glycine (M + 2) derived from [U-13C]-glucose, in asparagine-deprived Eµ-Myc cells, even in medium containing supraphysiological concentrations of both amino acids (Figs. 2f, g and S2e). These cells also exhibited increased levels of 13C-labelled asparagine (M + 2 and M + 3), consistent with active asparagine synthetase under conditions of extracellular asparagine withdrawal (Fig. S2f). Similarly, using α−15N-L-glutamine, we detected increased levels of 15N-labelled serine (M + 1) and 15N-labelled asparagine (M + 1) (Figs. S2e, g, h), further supporting increased de novo serine biosynthesis in malignant B cells exposed to an asparagine-free environment.
In vivo, BCL harvested from ASNase-treated mice also exhibits upregulated PSAT1, PSPH, ASNS, and ATF4 expression, though PHGDH expression was not (Figs. 2h and S2i). Nevertheless, PHGDH activity was significantly increased in BCL of ASNase-treated mice (Fig. 2i). Following several weeks of Vehicle or ASNase treatment, BCL-bearing mice received two consecutive boluses of [U-13C]-glucose at a 20 min interval to quantify de novo-synthesized serine, glycine, and asparagine in tumors (Fig. 2j). Equivalent enrichment of fully carbon-labeled glucose (M + 6) was detected in the plasma of both Vehicle and ASNase-treated mice bearing BCL (Fig. S3a). Consistent with in vitro data, levels of 13C-labelled serine (M + 3), glycine (M + 2) and asparagine (M + 2) were significantly increased in ASNase-treated BCL, accompanied by elevated total levels of serine, glycine, and a decrease in total levels of asparagine, suggesting increased de novo serine/glycine and asparagine biosyntheses in BCL exposed to ASNase treatment in vivo (Figs. 2k and S3b-e). As previously reported in WT mice and in multiple murine cancer models30, we also confirmed elevated plasma serine concentration in BCL-bearing C57BL/6 mice treated with ASNase (Fig. S3f).
Altogether, our results demonstrate that malignant B cells exposed to an asparagine-restricted environment support de novo serine biosynthesis, despite available environmental serine and glycine.
PHGDH supports ASNase-sensitive malignant B cell proliferation during therapy
We next investigated whether ASNase-sensitive malignant B cells increase de novo serine biosynthesis to facilitate their outgrowth in an asparagine-deprived environment. First, we tested several commercially available PHGDH inhibitors—including three allosteric compounds (NCT-503, CBR-5884, and PKUMDL-WQ2101) and one competitive inhibitor (BI-4916)31,32,33,34 in Eµ-Myc cells to evaluate their effects on PHGDH enzymatic activity and cellular viability. While all compounds exhibited concentration-dependent cytotoxicity in vitro (Fig. S4a–d), only BI-4916, a cell-permeable and selective NAD⁺/NADH-competitive inhibitor of PHGDH34, effectively inhibited PHGDH activity in Eµ-Myc cells (Fig. S4e). Based on these results, BI-4916 was selected for further investigations. Notably, our findings are consistent with prior studies demonstrating the superior efficacy of BI-4916 in inhibiting de novo serine biosynthesis in the breast cancer cell line MDA-MB-468, compared to allosteric inhibitors35. Treatment with 10 µM of BI-4916 inhibited PHGDH activity by 80% (Fig. 3a), significantly reduced total intracellular serine levels (Fig. S4f), and induced an average significant 30% increase in Eµ-Myc cell death within 24 hours (Figs. 3b and S4d). When combined with ASNase, BI-4916 further enhanced cell death in a caspase-dependent manner within 24 h, compared to either treatment alone (Figs. 3b and S4g). Moreover, ASNase/BI-4916 co-treatment significantly delayed the outgrowth of Eµ-Myc cells, underscoring the contribution of PHGDH activity to malignant B cells’ adaptation during ASNase treatment (Fig. 3c). Similar effects were obtained with independent ASNase-sensitive Eµ-Myc cells (Fig. S4h, i) and with the NAD+/NADH competitive PHGDH inhibitor BI-4924, which was chemically modified to generate the esterified prodrug form, BI-491634 (Fig. S4j, k).
a Percentage of PHGDH activity in Eµ-Myc (#688) cells treated with DMSO or indicated concentrations of the PHGDH inhibitor BI-4916 for 24 h in Asn-containing medium (n = 3 independent experiments). b Percentage of dead Eµ-Myc (#688) cells (DAPI+) following 24 h of treatment with DMSO, ASNase (0.003 IU/ml) and/or BI-4916 (10 µM) in Asn-containing medium (n = 4 independent experiments). c Four-day proliferation of Eµ-Myc (#688) cells treated as in (b) (n = 5 independent experiments). d Total protein extracts prepared from Eµ-Myc (#506) cells stably expressing shRNA targeting the firefly luciferase (shSCR) or the murine Phgdh mRNA (two independent shPhgdh #1 and #2) were immunoblotted for indicated proteins. e Relative quantification of immunoblots presented in (d). (n = 4 independent experiments). f Percentage of dead Eµ-Myc (#506) cells (DAPI+) silenced (shPhgdh #1 or #2) or not (shSCR) for Phgdh mRNA following 24 h incubation in Asn-containing medium supplemented (+) or not (−) with ASNase (0.003 IU/ml) (n = 4 independent experiments). g Four-day proliferation of Eµ-Myc cells silenced or not (shSCR) for Phgdh mRNA, treated as in (f) (n = 5 independent experiments). h Survival curves of WT C57BL/6 mice intravenously injected with Eµ-Myc (#506) cells stably expressing shRNA control (shSCR) or targeting the murine Phgdh mRNA (shPhgdh #1), treated with Vehicle or ASNase every 48 h from day 7 until disease endpoint (n = 10 mice/group). i Total protein extracts prepared from Eµ-Myc cells isolated from BCL of C57BL/6 mice presented in h, were immunoblotted for the indicated proteins (shSCR-Vehicle, n = 3; shSCR-ASNase, n = 3; shPhgdh #1-Vehicle, n = 4; shPhgdh #1-ASNase, n = 4 mice). V Vehicle, A ASNase. The samples derive from the same experiments but different gels for PHGDH, PSAT1, ERK2, and another for ASNS and ERK2 were processed in parallel. j Relative quantification of PHGDH protein levels presented in (i) Data are normalized to the control condition shSCR-Vehicle. Data are expressed as mean ± SD (a, b, e, f, j) or ± SEM (c, g). P-values are from one-way Anova followed by Tukey’s test (a, e), 2-way Anova followed by Tukey’s t-test (b, f, j), t-test (c, g), log-rank test (h), and indicated as ns, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
Due to the intrinsic instability of BI-4924 and BI-4916, which precludes their use in in vivo studies34, we sought to confirm that the observed effects were attributable to on-target PHGDH inhibition by stably expressing shRNA targeting the murine Phgdh mRNA in Eµ-Myc cells. Two shRNA constructs (#1 and #2) effectively reduced PHGDH protein levels without altering PSAT1 or PSPH expression levels (Fig. 3d, e). Consistent with pharmacological inhibition, silencing of Phgdh sensitized Eµ-Myc cells to ASNase-induced cell death and prevented the expansion of surviving cells in vitro (Fig. 3f, g). In vivo, ASNase-treated mice engrafted with Eµ-Myc-shPhgdh #1 cells displayed significantly delayed lymphoma progression compared to those bearing control Eµ-Myc-shSCR cells (Fig. 3h), confirming PHGDH as a key mediator of BCL relapse during ASNase therapy. Of note, at endpoint, we confirmed downregulated levels of PHGDH protein in BCL of Eµ-Myc-shPhgdh #1 cells-bearing mice, regardless of treatments (Fig. 3i, j). On the contrary, PHGDH overexpression in Eµ-Myc cells accelerated BCL development upon ASNase treatment (Fig. S4l–n), further validating PHGDH’s role in the adaptive resistance mechanisms.
Collectively, our results highlight the critical role of PHGDH activity and de novo serine biosynthesis in the proliferation of malignant B cells exposed to an asparagine-deprived environment.
ASNase treatment induces a reversible tumor metabolic response in vivo
Prolonged nutritional stress, such as that imposed by weeks of treatments with the clinically used ASNase, raises questions about the persistence of mechanisms driving tumor relapse. To assess the reversibility of this adaptation, Eµ-Myc cells that survived 4 days of ASNase treatment in vitro (1st challenge) were released from therapy (C1 cells). Upon re-exposure to ASNase (2nd challenge), C1 cells displayed an ASNase sensitivity comparable to chemo-naïve parental (WT) Eµ-Myc cells, suggesting that the adaptive response is transient (Fig. 4a–c). This result was consistent across other independent ASNase-sensitive Eµ-Myc cells (Fig. S5a, b).
a Schematic representation of the experimental design. Following 4 days of ASNase (0.003 IU/ml; 1st challenge), wild-type (WT) Eµ-Myc (#506) cells were washed and re-seeded in Asn-containing medium (drug holiday). The resulting cell population (C1 cells) was treated with ASNase (2nd challenge). b Percentage of dead (DAPI+) WT and C1 cells (from Eµ-Myc #506 cells) incubated in Asn-containing medium with or without (CTL) ASNase (0.003 IU/ml) for 24 h (n = 4 independent experiments). c Proliferation of cells presented in (b) (n = 4 independent experiments). d Total protein extracts prepared from Eµ-Myc (#506) cells treated as in (a) were immunoblotted for the indicated proteins. The samples derive from the same experiments, but different gels for PHGDH, another for PSAT1, another for PSPH, ASNS, ERK2, and another for ATF4 were processed in parallel. e Schematic representation of the two successive therapeutic challenges with Vehicle or ASNase in C57BL/6 mice bearing Eµ-Myc #506 cell-derived BCL. The 1st therapeutic challenge resulted in VBCL and ABCL (n = 6 mice/group). Malignant B cells from VBCL or ABCL were transferred into secondary recipient WT C57BL/6 mice. Seven days later, mice were treated with Vehicle or ASNase (2nd therapeutic challenge), resulting in Vehicle-VBCL, Vehicle-ABCL, ASNase-VBCL, and ASNase-ABCL (n = 6 mice/group). Survival curve of WT C57BL/6 mice intravenously injected with VBCL (f) or ABCL (g) malignant cells and treated with Vehicle or ASNase every 48 hours until disease endpoint (n = 6 mice/group). h Principal-component analysis (PCA) of metabolites abundance (130 metabolites detected) in BCL harvested from Vehicle and ASNase-treated C57BL/6 mice bearing VBCL cells or ABCL cells (n = 6 mice/group). i Heatmap representation of asparagine and serine relative abundance in BCL presented in (e) (n = 6 mice/group). Malignant cells isolated from BCL of the Vehicle-treated mouse #738 and of the ASNase-treated mouse #750 were engrafted into secondary recipient WT C57BL/6 mice for the 2nd ASNase challenge. j Total protein extracts prepared from BCL described in (i) (VBCL, n = 1; ABCL, n = 1; Vehicle ABCL, n = 6; ASNase-VBCL, n = 6), were immunoblotted for the indicated proteins. The samples derive from the same experiments, but different gels for PHGDH, another for PSAT1, PSPH, and another for ASNS and ERK2 were processed in parallel. Data are expressed as mean ± SD (b, c). P-values are from 2way Anova followed by Tukey’s test (b), t-test (c.), log-rank test (f, g), and indicated as ns, not significant, **, p < 0.01; ***, p < 0.001 ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
The expression levels of PHGDH, PSAT1, PSPH, ASNS and of the transcription factor ATF4, which were initially upregulated within the first 24 h, remained elevated through 4 days of ASNase treatment (Figs. 4d and S5c). Following therapy withdrawal, ASNS and PSPH protein levels returned to baseline within 24 h, while PSAT1 and PHGDH levels modestly decreased at 72 h and 96 h (control condition of the 2nd challenge), respectively, consistent with a time-dependent decreased expression level of ATF4. Upon ASNase re-exposure (2nd challenge) these enzymes were re-induced within 24 h (Figs. 4d and S5c), confirming a dynamic and reversible regulatory process.
In vivo, Eµ-Myc cells isolated from BCL treated with either Vehicle or ASNase (1st in vivo challenge) were re-engrafted into WT C57BL/6 mice and subjected to a second round of ASNase therapy (2nd in vivo challenge). Mice engrafted with ASNase-treated cells still exhibited a significant survival benefit upon re-treatment (Fig. 4e–g), supporting a model of flexible, non-permanent metabolic adaptation.
To assess whether the metabolic response to ASNase was reversible in vivo, we performed metabolomic profiling of BCL harvested after the 1st or 2nd ASNase treatment (in vivo challenge). Tumors from Vehicle-treated mice (Vehicle-VBCL and Vehicle-ABCL) exhibited distinct metabolic profiles compared to those from ASNase-treated mice (ASNase-VBCL and ASNase-ABCL), regardless of tumor origin (VBCL or ABCL). Importantly, tumors harvested from Vehicle-treated mice (Vehicle-VBCL and Vehicle-ABCL) showed a highly similar metabolic signature, even though Vehicle-ABCL originated from ABCL cells, i.e., malignant B cells that had already adapted to ASNase treatment, suggesting a transient resistance phenotype (Fig. 4h). Among the most significantly deregulated metabolic pathways in ASNase-ABCL compared to Vehicle-ABCL, serine/glycine metabolism was prominent, consistent with previous findings (Fig. S5d). ASNase-treated BCLs showed elevated serine and reduced asparagine levels, whereas the opposite pattern was observed in Vehicle-treated BCLs, underscoring dynamic metabolic reprogramming (Fig. 4i). In line with in vitro results (Figs. 4d and S5c), induction of PSAT1, PSPH, and ASNS expression in BCL following 1st in vivo challenge with ASNase was transient, as their expression levels decreased in Vehicle-ABCL of Vehicle-treated mice (or were maintained in ASNase-ABCL of ASNase-treated mice) following the 2nd in vivo challenge (Figs. 4j and S5e). Collectively, our data highlight the metabolic plasticity of malignant B cells exposed to ASNase therapy.
PHGDH activity limits ASNase-induced oxidative stress in malignant B cells
Given that amino acid or glucose starvation induces oxidative stress36,37, we determined whether ASNase treatment induces oxidative stress in Eµ-Myc cells and how these ASNase-sensitive Eµ-Myc cells adapt. Asparagine-deprived Eµ-Myc cells displayed elevated ROS levels and a higher oxidized to reduced glutathione ratio (GSSG/GSH) (Figs. 5a, b and S6a, b). This oxidative stress was accompanied by NRF2 (Nuclear Factor Erythroid 2-Related Factor 2) protein stabilization that in turn activated the transcription of canonical antioxidant response genes, including Glutamate-cysteine ligase catalytic subunit (Gclc), Glutamate-cysteine ligase modifier subunit (Gclm)—the first rate-limiting enzymes of glutathione biosynthesis—as well as the NAD(P)H quinone dehydrogenase 1 (Nqo1) (Fig. 5c–e).
a Relative ROS levels in Eµ-Myc (#688) cells cultivated for 15 h in Asn-free medium with (+) or without (–) Asn, ASNase (0.003 IU/ml) and N-acetyl-L-cysteine (NAC, 10 mM, 1 h) (n = 4 independent experiments). b GSSG/GSH ratio in Eµ-Myc (#688) cells cultivated in Asn-free medium, with (+) or without (−) Asn for 24 h (n = 4 biological replicates). c NRF2 and ATF4 expression levels in Eµ-Myc cells cultivated for 24 h as in (a). d Relative quantification of (c) (#506 cells, square, n = 2 independent experiments; #688 cells, triangle, n = 2 independent experiments). e Relative mRNA expression levels in Eµ-Myc (#688) cells cultivated as in (c) for 48 h (n = 3 independent experiments). f, g. Relative ROS levels in Eµ-Myc (#688) cells treated for 20 h in Asn-free medium with (+) or without (−) Asn (0.37 mM) and/or BI-4916 (10 µM) (f) or in Asn-containing medium with (+) or without ((−), DMSO), ASNase (0.003 IU/ml) and/or BI-4916 (10 µM) (g) (n = 4 independent experiments). h Percentage of dead Eµ-Myc (#506) cells (DAPI+) treated with (+) or without (−) DMSO, ASNase (0.003 IU/ml), BI-4916 (7.5 µM) and/or NAC (5 mM) for 48 h in Asn-containing medium (n = 4 independent experiments). i Heatmap representation of mRNA expression levels for the indicated genes, determined by RNAseq from Eµ-Myc cells isolated from 12 individual Eµ-MycTg/+ mice. j As in (i) in BCL harvested from Eµ-Myc cells-bearing mice treated with Vehicle or ASNase (n = 1 /group). k slc7a11 mRNA expression levels (transcript per million) in selected tumor entities available on the TCGA database. l Total protein extracts from Eµ-Myc (#506) cells stably expressing control (CTL) or murine SLC7A11-encoding (SLC7A11 OE) vectors were immunoblotted for the indicated proteins. The samples derive from the same experiments but different gels for SC7A11, ERK2, and another for CD98 were processed in parallel. m Relative quantification of immunoblots presented in (l). Data normalized to SLC7A11 OE condition (n = 3 independent experiments). n Percentage of dead CTL and SLC7A11 OE-overexpressing Eµ-Myc (#506) cells (DAPI+) treated as in (h) (n = 5 independent experiments). Data are expressed as mean ± SD (a, d–h, m, n) or ± SEM (b). P-values are from 2-way Anova followed by Tukey’s test (a, f–h, n), t-test (b, m), one-way Anova followed by Tukey’s test (d, e), and indicated as ns, not significant, *, p < 0.05; **, p < 0.01; ***, p < 0.001 ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
Given the induction of Gclc, Gclm, and the NRF2-targeted gene slc6a9 (solute carrier family 6 member 9)38, encoding a glycine transporter (Fig. S6c), we verified whether glutathione synthesis, which requires cysteine, glutamate, and ultimately glycine, was enhanced in asparagine-deprived Eµ-Myc cells. We detected increased levels of 13C-labelled GSH (M + 2) derived from 13C2-Glycine (Fig. S6d, e), indicating active glutathione synthesis in asparagine-deprived Eµ-Myc cells. Of note, glycine appears to be specifically catabolized in glutathione biosynthesis, as evidenced by a significant reduction in the abundance of downstream products of glycine metabolism and diminished 13C-ATP (M + 2) levels (Fig. S6f).
Since de novo serine biosynthesis also contributes to glycine for antioxidant defense, and given the observed increased levels of 13C3-serine and 13C2-glycine derived from [U-13C]-glucose in ASNase-treated cells (Fig. 2f, g), we hypothesized that this pathway supports redox homeostasis. Inhibition of PHGDH with BI-4916 significantly increased ROS levels, indicating that de novo serine biosynthesis plays a critical role in maintaining redox homeostasis in Eµ-Myc cells under basal conditions (Fig. 5f, g). Furthermore, cells treated with BI-4916 in Asn-free medium or with BI4916/ASNase combination in Asn-containing medium exhibit higher ROS levels compared to those observed with each single treatment, supporting the contribution of de novo serine biosynthesis to redox homeostasis in ASNase-treated Eµ-Myc cells (Fig. 5f, g). Accordingly, scavenging ROS with N-acetyl-L-cysteine (NAC) prevented cell death induced by BI-4916, either alone or in combination with ASNase (Fig. 5h).
To promote an effective antioxidant response, cancer cells must not only meet a high demand for glycine but also cysteine, the rate-limiting precursor of glutathione biosynthesis39,40,41. Due to the oxidizing tumor microenvironment, extracellular cysteine is unstable and rapidly converts into cystine. Therefore, most cancer cells rely on cystine transport, which is imported and reduced to cysteine in the cytosol, or on de novo cysteine biosynthesis. Surprisingly, Eµ-Myc cells harvested from 12 individual transgenic Eµ-Myc Tg/+ mice lacked detectable levels of slc7a11 mRNA, the solute carrier family 7 member 11 (also called xCT), which is the main plasma membrane antiporter mediating extracellular cystine uptake in exchange for glutamate (Fig. 5i). slc7a11 mRNA remained undetectable by qPCR in Eµ-Myc cells treated with ASNase for 72 h in vitro or by RNAseq in BCL exposed to several weeks of ASNase treatment in vivo (Figs. 5j and S6g). Other cystine transporters, slc7a9 and slc3a1, were also absent, while slc3a2 (also called cd98), encoding the chaperone for xCT and LATs transporters, was ubiquitously expressed (Fig. 5i, j). Analysis of The Cancer Genome Atlas (TCGA) database confirmed that human DLBCL exhibits low slc7a11 mRNA expression (Fig. 5k), a finding reinforced by RNA-seq data from 775 human DLBCL samples42, which showed that 7.28% expressed little to no detectable levels of slc7a11 mRNA, despite heterogeneous expression of slc3a2 mRNA (Fig. S6h, i). Overexpression of the murine SLC7A11 in slc7a11-deficient Eµ-Myc cells attenuated the death induced by inhibition of PHGDH alone or in combination with ASNase (Fig. 5l–n). Collectively, our results highlight the role of PHGDH in mitigating ASNase-induced oxidative stress in malignant B cells.
ASNase-induced ROS revealed a therapeutic vulnerability to the clinically used PARP1/2 inhibitor in ASNase-sensitive malignant B cells in vitro
We next aimed to broaden our investigation into the possible consequences of perturbed redox homeostasis in ASNase-treated malignant B cells. Given that ROS accumulation can induce single-strand or double-strand DNA breaks, we sought to determine if ASNase-induced oxidative stress triggers DNA damage.
Asparagine-restricted Eµ-Myc cells exhibited increased phosphorylation of histone H2AX at Ser139 (γH2AX), along with activation of the ATR-dependent DNA damage response (DDR) pathway (Fig. 6a–d). This was evidenced by phosphorylation of ATR at Ser428 and its downstream substrates Chk1 at Ser345 and RPA32 at Ser33, in vitro and in vivo (Figs. 6c, d and S7a, b), indicating the presence of DNA damage. In contrast, the ATM-dependent DDR was not activated, as KAP1, a downstream substrate of ATM, was not phosphorylated at Ser824 in Asn-restricted Eµ-Myc cells in vitro (Fig. 6c, d).
a Total protein extracts prepared from Eµ-Myc #688 cells cultured in Asn-free medium, supplemented (+) or not (−) with Asn (0.37 mM) and ASNase (0.003 IU/ml) for 24 h, were immunoblotted for the indicated proteins. Etoposide treatment (1 µg/ml) for 3 h was used as a positive control. b Relative quantification of immunoblots presented in (a). (#506 cells, square, n = 2 independent experiments; #688 cells, triangle, n = 3 independent experiments). c As in a for the indicated proteins expressed in Eµ-Myc #506 and #688 cells. From #506 cells, the samples derive from the same experiments, but different gels for P-ATR (S428), P-Chk1 (S345), KAP1, ERK2, another for ATR, another for Chk1, P-RPA32 (S33), P-KAP1 (S824), another for RPA32, and another for γH2AX were processed in parallel. From #688 cells, the sample derived from the same experiments, but different gels for P-ATR (S428), P-Chk1 (S345), P-KAP1 (S824), ERK2, another for ATR, Chk1, KAP1, another for P-RPA32 (S33), another for RPA32, and another for γH2AX were processed in parallel. d As in (b) for immunoblots presented in (c) (#506 cells, square, n = 2 independent experiments; #688 cells, triangle, n = 3 independent experiments). e γH2AX expression levels in Eµ-Myc cells (#506 and #688) following 24 h incubation in Asn-containing medium supplemented (+) or not (−) with ASNase (0.003 IU/ml) and/or N-acetyl-L-cysteine (NAC, 10 mM). f Relative quantification of γH2AX expression in Eµ-Myc cells presented in (e) (#688 cells, triangle, n = 2 independent experiments; #506 cells, square, n = 1). g γH2AX expression levels in Eµ-Myc (#688) cells treated (+) or not (−) for 24 h with DMSO (−/−), ASNase (0.003 IU/ml) and/or BI-4916 (10 µM). h Relative quantification of γH2AX expression levels in Eµ-Myc cells presented in (g). (#688 cells, triangle, n = 2 independent experiments; #506 cells, square, n = 1). i Total protein extracts prepared from BCL harvested from C57BL/6 mice engrafted with control (CTL) or V5-tagged murine PHGDH-overexpressing (PHGDH OE) Eµ-Myc (#506) cells and treated with Vehicle or ASNase from day 7 until disease endpoint, were immunoblotted for the indicated proteins (CTL-Vehicle, n = 3; CTL-ASNase, n = 3; PHGDH OE-Vehicle, n = 4; PHGDH OE-ASNase, n = 4 mice). The samples derive from the same experiments, but different gels for γH2AX, ERK2 (upper), another for P-ChK1 (S345), ERK2 (middle), another for Chk1, another for PHGDH, ERK2 (lower), and another for V5 were processed in parallel. j Relative quantification of γH2AX and P-Chk1 (Ser345) expression levels in BCL presented in (i). V Vehicle, A ASNase. For all graphs, data are expressed as mean ± SD. P-values are from one-way Anova followed by Tukey’s test (b, d), 2-way Anova followed by Tukey’s test (f, h, j) and indicated as ns, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001 ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
Scavenging ROS with NAC reduced γH2AX levels, confirming that oxidative stress contributes to ASNase-induced DNA damage (Fig. 6e, f). Conversely, PHGDH inhibition modestly increased γH2AX in ASNase-treated cells (Fig. 6g, h). Consistently, PHGDH overexpression in Eµ-Myc cells (Fig. S4l, m) significantly attenuated γH2AX expression levels and activation of the ATR-dependent DDR during ASNase therapy in vivo (Fig. 6i, j). This supports a protective role for de novo serine biosynthesis in preserving genome stability.
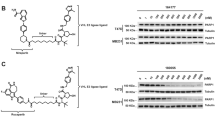
Following an ATR-dependent DNA damage response, the chromatin-associating enzymes poly (ADP-ribose) polymerase 1 (PARP1) and PARP2 catalyze the synthesis and transfer of ADP-ribose polymer (PARs) from nicotinamide adenine dinucleotide (NAD+) onto their serine, tyrosine, and glutamate residues (auto-PARylation), activating them. In turn, activated PARPs enable the PARylation of target proteins, promoting their recruitment to DNA breaks for effective DNA repair. We aimed to investigate whether PARP was activated in ASNase-treated Eµ-Myc cells, as this could represent a therapeutically exploitable tumor vulnerability using clinically approved PARP1/2 (PARP) inhibitors. We observed increased expression of proteins harboring poly(ADP-ribose) motifs, indicating elevated PARP activity in asparagine-restricted Eµ-Myc cells (Fig. 7a). PARP inhibitors, such as Olaparib, compete with NAD+ for the catalytically active site of PARP, thereby preventing the initiation of DNA repair mechanisms. To interfere with DNA repair in ASNase-treated cells, we sought to test the effects of ASNase/Olaparib co-treatment in Eµ-Myc cells, in in vitro and in vivo settings. This therapeutic combination enhanced caspase-dependent cell death in two OxPhos-dependent Eµ-Myc lines (#506 and #688), surpassing the effect of either agent alone (Fig. 7b–d). When co-treated with Olaparib (0.3 µM), ASNase-treated Eµ-Myc cells did not resume proliferation in vitro (Fig. 7e, f). In vivo, co-treatment with ASNase (every 48 h) and Olaparib (daily) significantly delayed weight loss and BCL onset in C57BL/6 mice, compared to monotherapies (Fig. 7g–i). ASNase/Olaparib co-treatment antineoplastic effect was confirmed in a human colon cancer cell line grown in 3D (Fig. S8a, b).
a Total protein extract prepared from Eµ-Myc cells (#506 or #688) cultivated for 10 h in Asn-free medium, supplemented (+) or not (−) with Asn (0.37 mM) alone or in combination with ASNase (0.003 IU/ml), was immunoblotted using an anti-poly(ADP-ribose) antibody. Treatment with H2O2 (0.5 mM) for 5 min was used as a positive control. The samples derive from the same experiments but different gels for poly(ADP)-ribose, ERK2 and another for ATF4 were processed in parallel. b, c Percentage of dead Eµ-Myc cells (DAPI+) treated for 24 h with DMSO, ASNase (0.003 IU/ml) and/or the PARP inhibitor Olaparib (at indicated concentrations) in Asn-containing medium. (b, from #506 cells, n = 3 independent experiments; c, from #688, n = 5 independent experiments). d Percentage of dead Eµ-Myc (#688) cells (DAPI+) treated with DMSO, ASNase (0.003 IU/ml) and/or Olaparib (0.3 µM) for 24 h in Asn-containing medium and in the presence (+) or absence (−) of the caspase inhibitor qVD-OPh (20 µM) (n = 3 experiments). e, f Four-day proliferation of Eµ-Myc (#506) cells (e) and of Eµ-Myc (#688) cells (f) treated with DMSO, ASNase (0.003 IU/ml) and/or Olaparib (0.3 µM) in Asn-containing medium (n = 5 independent experiments). g Schematic representation of in vivo co-treatment with ASNase/Olaparib. h Body weight of Eµ-Myc (#506) cells-bearing C57BL/6 mice from the start of treatment (day 7 post tumor cell inoculation) (n = 10 mice/group). i Survival curves of WT C57BL/6 mice intravenously injected with Eµ-Myc (#506) cells and treated 7 days later with Vehicle, Olaparib (daily), ASNase (every 48 h) or their combination, for several weeks, until lymphoma development reached endpoint (n = 10 mice/group). Data are expressed as mean ± SD (b–f) ± SEM (h). P-values are from 2-way Anova followed by Tukey’s test (b–f), log-rank test (i) and indicated as ns, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001 ****, p < 0.0001. For detailed individual P-value, please refer to the Source data table.
Overall, these results demonstrate that ASNase-induced oxidative stress creates a vulnerability to the PARP inhibitor Olaparib in malignant B cells, and that combining ASNase with Olaparib holds potential as an effective anti-cancer strategy.
Discussion
Our study reveals that the clinically used ASNase treatment induces oxidative stress and associated DNA damage in malignant B cells, therefore triggering a compensatory reliance on PHGDH-driven serine biosynthesis to restore redox stress and on PARP-mediated DNA repair to resolve the damage. Pharmacological inhibition of PARP with Olaparib significantly sensitizes malignant B cells to ASNase, confirming the functional importance of this adaptive response. This metabolic-DNA damage crosstalk not only supports tumor cell survival under treatment-induced nutritional stress but also exposes a therapeutically actionable liability. Our findings thus provide a strong rationale for combinatorial strategies targeting PARP in ASNase-treated malignant B cells using clinically approved PARP inhibitors, to improve the efficacy and durability of ASNase-based therapies.
Most studies investigating mechanisms of resistance to ASNase treatment in solid and hematological cancer cell lines have focused on primary resistance, aiming to identify therapeutic strategies to eliminate ASNase-resistant cancer cells10,11,18,19,20,43,44,45,46. Several CRISPR-Cas9 screens have identified determinants of ASNase sensitivity in long-term ASNase-exposed resistant cell lines (10–20 days in vitro)43,45,46 or in vivo in immunodeficient mice20. One such screen highlighted the overexpression of PHGDH, PSAT1 and PSPH, the three enzymes involved in de novo serine biosynthesis, in ASNase-resistant melanoma cells knocked-out for the asparagine synthetase45. However, whether de novo serine biosynthesis is functionally enhanced in resistant cells and whether its enzymes drive resistance remained not established.
Using ASNase-sensitive B-cell lymphomas, our study identified other key determinants of tumor relapse during ASNase therapy, following an initial antineoplastic response, resembling secondary resistance as observed in R/R relapsed DLBCL patients8. We observed elevated levels of de novo-synthesized serine and increased oxidative stress in ASNase-treated malignant B cells. We identified PHGDH as a key mediator of this adaptive response by mitigating ROS accumulation, thereby enabling the outgrowth of therapy-stressed malignant B cells. These findings are consistent with a recent study showing that Eµ-Myc-driven lymphomas rely on de novo serine biosynthesis for disease progression, highlighting the functional importance of PHGDH in human B-cell lymphomas47. Given that the asparagine synthetase is a key determinant in ASNase resistance, it remains to be addressed whether activation of de novo serine biosynthesis in ASNase-treated cells depends on their ability to replenish sufficient asparagine levels through upregulation and/or activation of asparagine synthetase on treatment in vitro and in vivo.
Interestingly, cycles of ASNase therapy and drug withdrawal in chemo-naïve malignant B cells, demonstrated that their progeny was still sensitive to rechallenge with therapy, a transient tumor adaptation observed in vitro and in vivo. Moreover, this tumor-intrinsic adaptative response involved a flexible metabolic remodeling, which engage de novo serine, glycine, glutathione, and asparagine biosyntheses, likely driven by a dynamic ATF4 and NRF2-dependent transcriptional program that counteracts ASNase-induced oxidative stress and facilitates cell cycle re-entry. Consistent with our results, asparagine restriction induces oxidative stress and an NRF2-dependent stress response in CD8+ T cells, enhancing their proliferation and effector functions48,49,50,51.
We also demonstrated that asparagine deprivation induces oxidative stress and DNA damage. ASNase-treated malignant B cells activated the ATR-dependent DNA damage response pathway and exhibited elevated PARP activity, suggesting replication stress. Consequently, ASNase-treated BCL are highly sensitive to the PARP inhibitor Olaparib, both in vitro and in vivo, implicating PARP as a determinant of BCL relapse during ASNase treatment. Our findings are consistent with prior studies showing that PARP plays a key role in DNA repair mechanisms following hydrogen peroxide-induced single-strand DNA breaks52 and for maintaining genomic integrity in B cells53. Notably, PARP2 loss increases replication stress and impedes BCL progression in the Eµ-Myc model53.
Overall, our study revealed a crosstalk between amino acids availability, redox metabolism, and genome integrity, with PHGDH and PARP as critical regulators. In line with our findings, the NRF2-targeted glucose-6-phosphate dehydrogenase (G6PD), was recently shown to regulate NAD metabolism and basal PARylation, linking redox control to DNA replication integrity in p53-mutant colorectal cancer cells54. Whether ASNase-induced metabolic reprogramming influences NAD⁺ biosynthesis to regulate PARP activation and DNA repair mechanisms in malignant cells warrants further investigations.
Importantly, human ALL samples resistant to ASNase treatment displayed enrichment of genes involved in double-strand break repair via break-induced replication55, reinforcing our observation that asparagine preserves genome stability in malignant cells.
A key feature of PARP inhibitors is their effectiveness in targeting solid cancers with defects in homologous recombination (HR), most often due to germline or somatic BRCA1/2 mutations. HR deficiency is prevalent in advanced breast, ovarian and metastatic pancreatic cancers, and is a key factor predicting tumor response to PARP inhibition56,57. Although BRCA1/2 mutations are rare in hematological malignancies, a recent study provided strong rationale for PARP inhibitor use in DLBCL subsets with HR deficiency. Specifically, PARP inhibition was shown to synergize with anti-CD20-based chemotherapies in a subtype of human DLBCL characterized by LIM-domain 2 (LMO2)-driven homologous recombination deficiency, a feature not observed in DLBCL lacking LMO258. LMO2 is a cysteine-rich protein preventing BRCA1 recruitment to DNA-strand breaks58. Malignant Eµ-Myc cells used in our study express variable levels of lmo2 mRNA (Fig. 5i–j), which may, in part, explain their sensitivity to Olaparib treatment, particularly in combination with ASNase. Further investigations would be needed to comprehensively characterize the HR status of Eµ-Myc lymphomas and the mechanisms regulating it. Nevertheless, the antineoplastic effect of ASNase/Olaparib co-treatment was confirmed in HR-proficient human colorectal cancer cells (HCT-116) grown in 3D, suggesting that other HR-proficient cancers may benefit from this combination strategy. Assessing toxicity in non-cancerous cells (e.g., immune cells, hepatocytes, cardiomyocytes) or normal tissues in treated WT mice would require dedicated pharmacological studies to ensure cancer cell-specific effects. That said, Olaparib is well-tolerated in normal cells with functional HR59, possibly offering a therapeutic window in which malignant cells would be preferentially targeted by ASNase/Olaparib co-treatment.
Overall, our study sheds light on the dynamic and flexible tumor metabolic response during ASNase therapy. These insights not only enhance our understanding of the metabolic mechanisms facilitating resistance to a clinically approved drug, but also unveil clinically actionable combination strategy to enhance the therapeutic index of ASNase and tackle aggressive B-cell lymphomas by combining ASNase and Olaparib. While PARP inhibitors are currently being evaluated in combination with radiation, alkylating agents, topoisomerase I inhibitors, PI3K inhibitors, and immunotherapies60, our findings establish a proof-of-concept that PARP inhibition can be effectively combined with a metabolic drug—ASNase—in B-cell lymphomas. Collectively, our findings revealed a clear path for developing an anti-tumor strategy that combines ASNase and PARP1/2 inhibitor, offering an alternative therapeutic opportunity for patients treated with ASNase or Olaparib, to prevent tumor relapse.
Methods
Mice
C57BL/6 Eµ-Myc transgenic mice (Eµ-MycTg/+) were purchased from the Jackson Laboratory (002728) and are bred and maintained at our local animal facility (C3M, INSERM U 1065, Nice, France, B 06-088-20). Wild-type (WT) C57BL/6JOlaHsd female were purchased from Envigo and housed at our local animal facility. All mice were maintained in specific pathogen-free conditions and kept in the 12 h light/night cycles at 20–24 °C and 50–55% of humidity. For all mice experiments, food (SAFE R04, Safe Lab) was given ad libitum. All animal care procedures, experiments and maintenance complied with the protocols (n° APAFIS#25228-2020032917261458 v9) approved by the Institutional Animal Care and Use Committee (IACUC of University Nice, Côte d’Azur) and the Ministère de la Recherche et de l’Innovation.
Cell lines
Primary Eµ-Myc cells were isolated from individual male or female Eμ-MycTg/+ mice as previously described and characterized8,61. In particular, the OxPhos-dependent Eμ-Myc #506 and Eμ-Myc #688 cells were derived from B-cell lymphoma of Eμ-MycTg/+ females #506 and #688, respectively. These cells were maintained in DMEM-GlutaMAX medium (31966047, Thermo Fisher Scientific) supplemented with 10% of fetal bovine serum (FBS) (F7524, Sigma), 50 µM of 2-Mercaptoethanol (31350010, Thermo Fisher Scientific), 0.37 mM of L-asparagine (A0884, Sigma), 10 mM of HEPES pH 7.4 (15630056, Thermo Fisher Scientific) and 100 U/ml of Penicillin-Streptomycin (15140122, Thermo Fisher Scientific). 293 T cells (CRL-1573, ATCC) were maintained in DMEM-GlutaMAX media (31966047, Thermo Fisher Scientific) supplemented with 10% FBS and 100 U/ml of Penicillin-Streptomycin and used to produce VSV-G retroviral vectors. The murine Lewis Lung carcinoma (LLC) cell line (CRL1642, ATCC) was maintained in DMEM-GlutaMAX medium (31966047, Thermo Fisher Scientific) supplemented with 10% FBS and 100 U/ml of Penicillin-Streptomycin. The murine colon carcinoma cell line CT26 (CRL-2638, ATCC) was maintained in RPMI-GlutaMAX medium (61870044, Thermo Fisher Scientific) supplemented with 10% FBS, 100 U/ml of Penicillin-Streptomycin and 1 mM of sodium pyruvate (11360070, Thermo Fisher Scientific). The murine colon adenocarcinoma cell line MC38 (SCC172, Sigma) was maintained in DMEM-F12-GlutamaX medium (31331028, Thermo Fisher Scientific) supplemented with 10% FBS, 1 mM of sodium pyruvate and 100 U/mL of Penicillin-Streptomycin. The human colon carcinoma cell line HCT116 (CVCL_0291) was obtained from ATCC, routinely cultured in McCoy medium (16600082, Thermo Fisher Scientific), supplemented with 10% FBS and 100 U/ml of penicillin/streptomycin. All cell lines used in this study are mycoplasma-free and routinely tested for mycoplasma.
Media for in vitro studies
Asparagine-free medium refers to DMEM high D-glucose (4.5 g/L), no L-glutamine, no pyruvate medium (11960044, Thermo Fisher Scientific), supplemented with 10% FBS (F7524, Sigma), 3–50 µM of 2-Mercaptoethanol, 10 mM of HEPES pH 7.4, 100 U/ml of Penicillin-Streptomycin, 1 mM of sodium pyruvate (11360070, Thermo Fisher Scientific), 2 mM of L-glutamine (25030081, Thermo Fisher Scientific). This medium contains supraphysiological concentration of serine and glycine.
Asparagine-containing medium refers to the asparagine-free medium described above, supplemented with L-asparagine (0.37 mM).
Asparagine, serine, glycine-free medium refers to MEM no L-glutamine medium (21090022, Thermo Fisher Scientific), supplemented with 10% FBS (F7524, Sigma) dialyzed (132724, Spectra/Por®3 Dialysis Membrane Standard RC Tubing, MWCO 3.5 kDa), 50 µM of 2-Mercaptoethanol, 10 mM of HEPES pH 7.4, 100 U/ml of Penicillin-Streptomycin, 20 mM of D-glucose (J60067, Thermo Fisher Scientific), 1 mM of sodium pyruvate, 2 mM of L-glutamine, 1% of vitamins cocktail (11120052, Gibco).
Glucose and asparagine-free medium refers to DMEM no L-glutamine, no pyruvate medium (A14430-01, Thermo Fisher Scientific), supplemented with 10% FBS (F7524, Sigma), 50 µM of 2-Mercaptoethanol, 10 mM of HEPES pH 7.4, 100 U/ml of Penicillin-Streptomycin, 1 mM of sodium pyruvate, 2 mM of L-glutamine. This medium contains supraphysiological concentration of serine and glycine.
ASNase formulations
For all in vitro experiments, the native Escherichia-coli L-asparaginase II (ASNase) (Kidrolase®, Jazz Pharmaceutical) was used. For in vivo studies presented in Figs. 1, 2, 4, S3 and S5, we used Kidrolase®, and for in vivo studies in Figs. 3, 6, 7 and S3, we used the native Escherichia-coli L-asparaginase II recombinant protein (Spectrila®).
In vivo transfer of Eµ-Myc cells and survival experiments
Mouse primary Eµ-Myc #506 or #688 cells were transferred via intravenous (I.V) injection (100.000 live cells per recipient mouse) into syngeneic, nontransgenic, 6-weeks-old C57BL/6JOlaHsd females (Envigo), which are less prone to fighting compared to males. Seven days later, mice were treated every 48 h via intraperitoneal (I.P) injection with Vehicle (NaCl 0.9%) or 2500 IU/kg of the native (Kidrolase®, Jazz Pharmaceutical) or recombinant (Spectrila®) E-coli L-asparaginase (ASNase) until lymphoma reached the ethical limit. ASNase is diluted in NaCl 0.9%. Tumor onset was assessed by palpation of the inguinal lymph node. The ethical limit (disease endpoint) was defined using a disease-associated clinical score approved by the Institutional Animal Care and Use Committee (IACUC of University Nice, Côte d’Azur) and the Ministère de la Recherche et de l’Innovation (n° APAFIS#25228-2020032917261458 v9). Once inguinal tumor onset was detected, disease-associated clinical scores were recorded daily for each mouse. When the maximal score was reached, Vehicle- and ASNase-treated mice received a final injection of Vehicle or ASNase, respectively and were sacrificed three to four hours later by cervical dislocation. Blood was then collected through cardiac punction, placed in lithium heparinate coated tubes (BD Microtainer), centrifuged 6000 × g for 2 min to isolate the plasma. The plasma samples were collected, snap-frozen and stored at −80 °C for metabolomic analysis. Mice survival defined as survival probability (%) was determined as the time between the I.V injection of Eµ-Myc cells and the time when mice had to be sacrificed for ethical reasons when lymphomas progression reached the endpoint. Tumor-free mice (%) was determined as the time between the I.V injection of Eµ-Myc cells and the time when one inguinal lymphoma was palpable.
For the second in vivo therapeutic challenge, syngeneic, nontransgenic, 6-weeks-old C57BL/6JOlaHsd females (Envigo) were intravenously (I.V) injected with 100.000 live Eµ-Myc cells per recipient mouse. Eµ-Myc cells used were harvested from BCL obtained after the 1st in vivo therapeutic challenge. Seven days later, mice were treated with Vehicle or ASNase at same concentration, frequency, and administration route as mentioned above.
For in vivo co-treatment with ASNase and Olaparib, syngeneic, nontransgenic, 6-weeks-old C57BL/6JOlaHsd females (Envigo) were intravenously injected with 100.000 live Eµ-Myc cells per recipient mouse. Seven days later, mice were treated with either Vehicle or 50 mg/kg of Olaparib (daily I.P) or 2500 IU/kg of Spectrila (I.P every 48 h) or the combination of both drugs (daily I.P Olaparib 50 mg/kg and Spectrila, 2500 IU/kg, I.P every 48 h). Olaparib (HY-10162, MCE) was dissolved in DMSO at a concentration of 100 mg/ml and then diluted at 5 mg/ml in 20% sulfobutylether-β-cyclodextrin solution (HY-17031, MCE, diluted in NaCl 0.9%). Vehicle consists in daily I.P injections of a 5% DMSO/95% sulfobutylether-β-cyclodextrin (in 20% in NaCl 0.9%) solution and/or I.P of NaCl 0.9% every 48 h.
Survival functions were estimated using the Kaplan-Meier method and compared with the log-rank test.
RNA extraction, and real-time quantitative PCR
Total RNA was extracted from Eµ-Myc cells using the RNeasy mini kit (74104, Qiagen) according to manufacturer’s instructions. Reverse transcription was performed from 2 µg of RNA using the Omniscript RT kit (205113, Qiagen). Relative mRNA levels were obtained by real-time quantitative PCR (qPCR) on the StepOne™ Real-Time PCR System (Thermo Fisher Scientific) using the TaqMan assay primer set (Thermo Fisher Scientific) for murine Phgdh (Mm01623589_g1), Psat1 (Mm01613328_g1), Psph (Mm01197775_m1), Gls (Mm01257297_m1), Gclc (Mm00802658_m1), Gclm (Mm01324400_m1), Nqo1(Mm01253561_m1), slc7a11 (Mm00442530_m1), slc6a9 (Mm00433662_m1), Asns (Mm01137310_g1), and the TaqMan Universal PCR Master Mix (4304437, Fisher Scientific) according to manufacturer’s instructions. The qPCR was performed from 8 ng of cDNA, except for slc7a11 amplification for which we used 100 ng of cDNA. Relative mRNA levels were normalized to mouse Rplp0 (Mm00725448_s1, Thermo Fisher Scientific) and to the control condition (+Asn, 0.37 mM).
RNAseq data
RNA sequencing (RNA-seq) alignment files and clinical information from the EGAS00001002606 study42 were downloaded from the European Genome-phenome Archive. Quantification of genes was performed using featureCounts v1.5.0-p3 based on Ensembl GTF release 75 annotations. RNA-seq data were then normalized using the Bioconductor package DESeq2. Eighteen (out of 775) samples showing no expression of Rplp0 were removed from the analysis.
RNA-seq data on human samples (solid cancers and hematological cancers) were retrieved and downloaded from The Cancer Genome Atlas program (TCGA, https://portal.gdc.cancer.gov/) through the Gene Expression Profiling Interactive Analysis website (http://gepia.cancer-pku.cn). The levels of slc7a11 mRNA were represented in transcript per million (TPM). HNSC, Head and Neck squamous cell carcinoma; LUSC, Lung squamous cell carcinoma; LUAD, Lung adenocarcinoma; LGG, Brain Lower Grade Glioma; GBM, Glioblastoma multiforme; STAD, Stomach adenocarcinoma; SKCM, Skin Cutaneous Melanoma; COAD, colon adenocarcinoma; DLBCL, Diffuse Large B-cell Lymphoma; LAML, Acute Myeloid Leukemias.
The RNA sequencing from Eµ-Myc cells cultivated in vitro and from Eµ-Myc cell-derived BCL treated with Vehicle and ASNase in vivo, was performed on Illumina NextSeq500 at the IPMC functional genomics platform (France). Libraries were generated using Truseq Stranded mRNA kit (Illumina), from 1 µg of total RNA extracted from different Eµ-Myc cell lines isolated from 12 individual transgenic Eµ-MycTg/+ mice and from B-cell lymphomas harvested from mice engrafted with Eµ-Myc #506 cells, and treated Vehicle or ASNase for several weeks. Libraries were then quantified with Qubit dsDNA High Sensitivity Assay Kit (Invitrogen) and pooled. 4 nM of this pool were loaded on a NextSeq 500 High output Flowcell and sequenced on a NextSeq 500 platform (Illumina) with 2 × 75 bp paired-end chemistry. Reads were mapped with STAR 2.4.0a on mouse genome (build mm10). Counting was performed with featureCounts (subread-1.5.0-p3-Linux-x86_64, “—primary -g gene_name -p -s 1 -C” options) using Ensembl GRCm38.83 release. Raw counts were transformed into fpkm (fragments per kilobase of transcript per million mapped reads) values to normalize for both sequencing depth and gene length. Expressions values are shown as log2(fpkm).
DNA constructs
To amplify mouse Asns or Phgdh cDNA, total RNA was extracted from Eµ-Myc cells and reverse transcription was performed as mentioned above. The murine Phgdh coding sequence (without stop codon) was amplified by PCR using a recombinant Taq DNA polymerase (10342020, Thermo Fisher Scientific) according to manufacturer’s instructions and with the following primers: forward 5′-GCAATGGCCTTCGCAAATCTGC-3′ and reverse 5′-GGAGCCAGCCCAGGCGT-3′. The PCR product was cloned into the pCR™ 2.1-TOPO® TA vector using the TOPO™ TA Cloning™ Kit (K455040, Thermo Fisher Scientific) according to manufacturer’s instructions. The resulting plasmid TOPO-Phgdh was amplified in One Shot TOP10 chemically competent E.Coli (C404003, Thermo Fisher Scientific) with ampicillin (A9518, Sigma) selection, purified (740414.50, Macherey-Nagel) and sequenced to confirm the cloning of the murine wild-type Phgdh coding sequence. Once confirmed, Phgdh coding sequence tagged to V5 was subcloned from the TOPO-Phgdh vector into the retroviral GFP-encoded pMIG vector (9044, Addgene), by PCR with the following primers: forward 5′-GAGAGTCGACATGGCCTTCGCAAATCTGC-3′ and reverse 5′-GGTTAACTCACGTAGAATCGAGACCGAGGAGAGGGTTAGGGATAGGCTTACCGAAGCAGAACTGGAAAGCCTCCAATAC-3′. Then, the amplicon was digested using SalI (R3136S, New England Biolabs) and HpaI (R0105S, New England Biolabs) restriction enzymes. The pMIG vector was linearized using XhoI (R0146S, New England Biolabs) and HpaI restrictions enzymes to generate compatible cohesive ends. Digested pMIG vector was dephosphorylated with Antarctic phosphatase (MO289S, Biolabs). The following double-strand DNA fragment which contains the coding sequence of the murine slc7a11 with BglII and EcoRI restriction site was synthesized (Thermo Fisher Scientific): “ GGGGAGATCTATGGTCAGAAAGCCAGTTGTGGCCACCATCTCCAAAGGAGGTTACCTGCAGGGCAATATGAGCGGGAGGCTGCCCTCCATGGGGGACCAAGAGCCACCTGGGCAGGAGAAGGTAGTTCTGAAAAAGAAGATCACTTTGCTGAGGGGGGTCTCCATCATCATCGGCACCGTCATCGGATCAGGCATCTTCATCTCCCCCAAGGGCATACTCCAGAACACGGGCAGCGTGGGCATGTCCCTGGTTTTCTGGTCTGCCTGTGGAGTACTGTCACTTTTTGGAGCCCTGTCCTATGCAGAATTAGGTACAAGCATAAAGAAATCTGGTGGTCATTACACATACATTCTGGAGGTCTTTGGTCCTTTGCTGGCTTTTGTTCGAGTCTGGGTGGAACTGCTCGTAATACGCCCTGGAGCTACTGCTGTGATATCCCTGGCATTTGGACGCTACATCCTGGAACCATTTTTTATTCAATGTGAAATTCCTGAACTTGCAATCAAGCTCGTGACAGCTGTGGGCATCACTGTGGTGATGGTCCTAAATAGCACGAGTGTCAGCTGGAGTGCCCGGATCCAGATTTTCCTAACCTTTTGCAAGCTCACAGCAATTCTGATAATTATAGTCCCTGGAGTTATACAGCTAATTAAAGGGCAAACACATCACTTTAAAGATGCATTTTCAGGAAGAGACACAAGTCTAATGGGGTTGCCCTTGGCTTTTTATTATGGGATGTATGCATATGCTGGCTGGTTTTACCTCAACTTTATTACTGAAGAAGTAGACAACCCTGAAAAAACCATCCCCCTTGCAATCTGCATCTCCATGGCTATCATCACAGTGGGCTACGTACTGACAAACGTGGCCTATTTTACCACCATCAGTGCGGAGGAGCTGCTGCAGTCCAGCGCCGTGGCGGTGACCTTCTCTGAGCGGCTGCTGGGAAAATTCTCATTAGCAGTCCCGATCTTTGTTGCCCTCTCCTGCTTCGGCTCCATGAACGGTGGTGTGTTCGCTGTCTCCAGGTTATTCTACGTCGCATCTCGAGAAGGGCACCTTCCGGAAATCCTCTCTATGATTCATGTCCACAAGCACACTCCTCTGCCAGCTGTTATTGTTTTGCATCCTCTGACGATGGTGATGCTCTTCTCCGGAGACCTCTATAGTCTTCTAAATTTCCTCAGTTTTGCCAGGTGGCTTTTTATGGGGCTGGCAGTCGCAGGACTGATTTATCTTCGATACAAACGCCCAGATATGCATCGTCCTTTCAAGGTGCCTCTCTTCATCCCGGCACTATTTTCCTTCACCTGCCTCTTCATGGTTGTCCTCTCTCTTTACTCGGACCCATTCAGCACCGGGGTCGGTTTTCTTATCACCTTGACTGGGGTCCCTGCATATTATCTCTTCATTGTATGGGACAAGAAACCCAAGTGGTTCAGACGATTATCAGACAGAATAACCAGAACATTACAGATTATACTAGAAGTTGTACCAGAAGACTCTAAAGAATTATGAGAATTCGGGG “. This sequence was then cloned in the pMIG vector using EcoR1 (RO101S, New England Biolabs) and BglII (RO143, New England Biolabs) restriction enzymes.
The shRNA targeting mouse Phgdh were obtained using the oligonucleotide sequences forward 5′- GATCCCCGTGGAGAAGCAGAACTTGATTCAAGAGATCAAGTTCTGCTTCTCCACTTTTTA-3′, reverse 5′- AGCTTAAAAAGTGGAGAAGCAGAACTTGATCTCTTGAATCAAGTTCTGCTTCTCCACGGG-3 (shPhgdh #1) and forward 5′-GATCCCCGGAGGAGCTGATAGCTGAATTCAAGAGATTCAGCTATCAGCTCCTCCTTTTTA-3′, reverse 5′-AGCTTAAAAAGGAGGAGCTGATAGCTGAATCTCTTGAATTCAGCTATCAGCTCCTCCGGG-3′′ (shPhgdh #2). As a control, shRNA targeting firefly luciferase were obtained using the oligonucleotides sequences forward 5′-GCGTTATTTATCGGAGTTG-3′ and reverse 5′-CAACTCCGATAAATAACGC-3′ (shSCR). 3 µg of corresponding forward and reverse oligonucleotides were annealed and cloned into the retroviral GFP-encoded pSUPER vector (VEC-PRT-0006, Oligoengine) linearized using HindIII (RO104S, New England Biolabs) and BglII restrictions enzymes according to the manufacture’s instruction.
Upon gel purification (Monarch DNA Gel Extraction Kit, T10205, New England Biolabs), ligation was performed using the T4 DNA ligase (MO202S, Biolabs) with a ratio vector:insert of 1:4. The generated vectors were amplified in MAX Efficiency Stbl2 competent cells (10268019, Thermo Fisher Scientific) with ampicillin selection, purified and sequenced.
Generation of stable genetically modified Eµ-Myc cells
Self-inactivating retroviruses were generated following transient co-transfection of 293 T cells (CRL-1573, ATCC) with 8.6 µg of one of the following vectors, pMIG empty vector (CTL), pMIG-Asns-v5 (ASNS OE), pMIG-Phgdh-v5 (PHGDH OE), or pMIG-slc7a11 (SLC7A11 OE), pSUPER-shSCR or pSUPER-shPhgdh (#equence #1 or #2), 3 µg of envelope plasmid phCMV-VSV-G and 8.6 µg of MLV-Gag-Pol, using the classical calcium phosphate method. After 15 h, transfected 293 T cells were incubated in Opti-MEM reduced serum medium (11058021, Thermo Fisher Scientific) for 32 h. Cell supernatant was centrifuged 3000 × g at 4 °C overnight to concentrate retroviral particles prior titration. Eµ-Myc cells were then transduced with the resulting retroviruses using a multiplicity of infection (MOI) of 10. 72 h following transduction, GFP+ Eµ-Myc cells were sorted (SH800S Cell Sorter, Sony).
Determination of glutamine and glutamate concentrations in medium
The native E-Coli L-asparaginase (ASNase, Kidrolase) was incubated in asparagine-containing medium in the absence of cells, at indicated concentrations and at 37 °C, 5% CO2. The concentrations of glutamine and glutamate (mM) in the medium were electro-enzymatically determined overtime, using the YSI 2950 Biochemistry Analyzer (Yellow Springs Instruments).
Western blot analysis
Eµ-Myc cells isolated ex-vivo from inguinal B-cell lymphomas or cultivated in vitro were washed in PBS and lysed in laemmli buffer to extract total proteins. Frozen axillary B-cell lymphomas were homogenized using a stainless-steel tissue grinder (1292, BioSpec Products) and proteins were extracted using the laemmli lysis buffer. After protein quantification (23225, Pierce BCA Protein Assay kit, Thermo Fisher Scientific), 15–30 µg of whole-cell protein lysates were separated on 8–15% SDS polyacrylamide gels and transferred onto polyvinylidene difluoride membranes (Millipore). Membranes were then blotted with antibodies against ASNS (14681-1-AP, Proteintech, RRID: AB_2060119, dilution 1/2000), PHGDH (14719-1-AP, Proteintech, RRID: AB_2283938, dilution 1/1000), PSAT1 (10501-1-AP, Proteintech, RRID: AB_2172597, dilution 1/1000), PSPH (14513-1-AP, Proteintech, RRID: AB_2171464, dilution 1/1000), NRF2 (12721, Cell Signaling, RRID: AB_2715528, dilution 1/1000), GLUL (11037-2-AP, Proteintech, RRID: AB_2110650, dilution 1/1000), GOT1 (14886-1-AP, Proteintech, RRID: AB_2113630, dilution 1/1000), γH2AX (ab81299, Abcam, RRID: AB_1640564, dilution 1/2000), ATF4 (11815, Cell Signaling, RRID: AB_2616025, dilution 1/500) for Figs. 2h, 4d and S5c, ATF4 (81798-2-RR, Proteintech; RRID: AB_3670506, dilution 1/500) for Figs. 2d, 5c, d, 7a, S2c and S7a, xCT/SLC7A11 (98051, Cell Signaling, RRID: AB_2800296, dilution 1/1000), Poly/Mono-ADP Ribose (89190, Cell Signaling, dilution 1/1000); Phospho-ATR (Ser 428) (2853, Cell Signaling, RRID: AB_2290281, dilution 1/500), ATR (13934, Cell Signaling, RRID: AB_2798347, dilution 1/1000), Phospho-Chk1 (Ser345) (2348, Cell Signaling, RRID: AB_331212, dilution 1/1000), Chk1 (2360, Cell Signaling, RRID: AB_2080320, dilution 1/1000), Phospho-RPA32 /RPA2 (Ser33) (10148, Cell Signaling, RRID: AB_3099645, dilution 1/1000), RPA32/RPA2 (52448, Cell Signaling, RRID: AB_2750889, dilution 1/1000), Phospho-KAP1 (Ser824) (A300-767A, Thermo Fisher Scientific, RRID: AB_669740, dilution 1/1000), KAP1 (4123, Cell Signaling, RRID: 2256670, dilution 1/1000); V5 (R96025, Life Technologies, RRID: AB_159313, dilution 1/5000) and ERK2 (sc-1647, Santa Cruz, RRID: AB_627547, dilution 1/2000). Immunoreactive bands were detected with anti-mouse (7076, Cell Signaling Technology, RRID: AB_330924, dilution 1/3000) or anti-rabbit (7074S, Cell Signaling Technology, RRID: AB_2099233, dilution 1/3000) IgG horseradish peroxidase (HRP)-linked antibodies.
Immunoblots were visualized (FUJIFILM LAS4000) by chemiluminescence using Pierce ECL Western Blotting substrates (32106, Pierce ECL, Thermo Fisher Scientific). ERK2 was used as a loading control in all immunoblotting experiments. When indicated, quantification of the level of indicated proteins was performed using ImageJ software. Relative quantification was obtained by normalizing data to ERK2 levels and to indicated control conditions. Whole-cell lysate obtained from Eµ-Myc cells treated with 1 µg/ml of etoposide (HY-13629, MCE) for 3 h or stimulated with H2O2 (0.5 mM) for 5 min was used as a positive control for γH2AX expression and PARylation, respectively.
PHGDH activity
For in vitro samples, live Eµ-Myc cells were lysed for 10 min on ice, in buffer A, containing 20 mM of Tris pH 7.4, 5 mM of EDTA pH 8.0, 150 mM of NaCl, 0.5 % of NP40 (UN3500, EUROMEDEX) and 0.01 % of cocktail of protease and phosphatase inhibitors (1861280, Thermo Fisher Scientific). For ex-vivo samples, frozen B-cell lymphomas were homogenized using a stainless-steel tissue grinder (1292, BioSpec Products) and proteins were extracted in buffer A. Then, 10 to 30 µg of proteins were incubated in PHGDH activity buffer containing 51.5 mM of Tris pH 8.0, 10 mM of MgCl2, 0.05% of BSA (P06-1403500, Panbiotech), 0.01 % of Tween 20 (2001-B, Euromedex), 1.25 mM of L-glutamate (49449, Sigma), 0.3 mM of NAD+ (N1511, Sigma) and 0.1 mM of 3-phosphoglycerate (HY-141412, MedChemExpress) in a black 96-well plate (655079, Greiner). PHGDH activity was measured on a fluoroscan (Synergy H1, Biotek) at 460 nm after excitation at 355 nm as the increase in fluorescence related to NADH accumulation. PHGH activity is determined as the delta absorbance, representing the speed of NADH production. Data are normalized to milligrams of protein and expressed as mean ± SD of the percentage relative to the control condition (+Asn, in vitro or Vehicle, in vivo).
L-[3H(G)]-Serine uptake
A total of 250.000 live Eµ-Myc cells/ml were seeded in asparagine, serine, glycine-free medium supplemented with 0.4 mM of L-serine (S4311, Sigma), 0.4 mM of L-glycine (G8790, Sigma), and with or without 0.37 mM of L-asparagine (A0884, Sigma) and 0.003 IU/ml of E-coli L-asparaginase (Kidrolase) for 24 h at 37 °C, 5% CO2. The number of live cells was determined by 4′,6-diamidino-2-phenylindole (DAPI, D9642, Sigma) (0.5 µg/ml) staining and flow cytometry analysis (Miltenyi Biotech). Then, 1 × 106 of live Eμ-Myc cells (DAPI negative) were washed twice with PBS and incubated in a 12-well plate in 1 mL of pre-warmed asparagine, serine, glycine-free medium supplemented as described above, with L-serine being replaced by 1 μCi/ml or 0.4 μCi/ml of L-[3H(G)]-Serine (NET248250UC, PerkinElmer), for 15 min or 3 h at 37 °C, 5% CO2, respectively. Subsequently, cells were washed twice with pre-chilled PBS and lysed with 120 μl of 0.1 N NaOH and mixed with 4 ml of Ultima Gold (6013321, PerkinElmer). Radioactivity (count per minute, cpm) was measured using a β-scintillation counter (Tri-Carb 2810 TR, Perkin Elmer). L-[3H(G)]-serine transport rate was expressed as mean ± SD of cpm/106 cells.
Cell death assay
A total of 250.000 live Eµ-Myc cells/ml were treated with either DMSO (0.075%), or E-Coli L-asparaginase (0.003 IU/ml), or indicated concentration of PHGDH inhibitor BI-4916 (HY-126253, MCE), BI-4924 (HY-126254, MCE), NCT-503 (HY-101966, MCE), CBR-5884 (HY-100012, MCE), PKUMDL-WQ2101 (HY-123269, MCE), or Olaparib (AZD2281, HY-10162, MCE), or the combination of indicated drugs, in asparagine-free medium supplemented or not with 0.37 mM of L-asparagine (A0884, Sigma) at 37 °C, 5% CO2 for the indicated time. For quenching ROS, 5 mM of N-acetyl-L-cysteine (A9165, Sigma) was added simultaneously. For inhibition of caspases, 20 µM of the pan-caspase inhibitor qVD-OPh (A1901, APExBIO) was added simultaneously. Following 24 or 48 h of treatment, cells were labeled with 0.5 µg/ml of DAPI (D9642, Sigma) and immediately analyzed by flow cytometry (Miltenyi Biotec). Data represents the percentage of dead cells (DAPI+) and are expressed as mean ± SD.
Cell proliferation assay
A total of 250.000 live Eµ-Myc cells/ml were treated with either DMSO (0.075%), or 0.003 IU/ml of E-Coli L-asparaginase, or 10 µM of BI-4916 (HY-126253, MCE), or 60 µM of BI-4924 (HY-126254, MCE), or 0.3 µM of Olaparib (AZD2281, HY-10162, MCE), or the combination of indicated drugs, in asparagine-free medium supplemented or not with 0.37 mM of L-asparagine, at 37 °C, 5% CO2 for the indicated period.
For cell proliferation in asparagine, serine, glycine-free medium, a total of 250.000 live Eµ-Myc cells/ml were seeded in the presence or absence of L-asparagine (0.37 mM), and/or L-serine (0.4 mM)/L-glycine (0.4 mM) at 37 °C, 5% CO2 for the indicated period.
Cells were diluted in the medium of the corresponding condition when they reached 1.106 cells/ml. The number of live cells was determined overtime by DAPI (0.5 µg/ml) staining and flow cytometry analysis (Miltenyi Biotec). Data are expressed as the mean number of live cells (DAPI−) ± SD or ± SEM.
Measurement of reactive oxygen species
A total of 250.000 live Eμ-Myc cells per ml were treated for 15 or 20 h (as indicated in the legends) with either DMSO (0.075%), or 0.003 IU/ml of E-Coli L-asparaginase, or 10 µM of BI-4916 (HY-126253, MCE), or the combination of both drugs, in asparagine-free medium supplemented or not with L-asparagine (0.37 mM) at 37 °C, 5% CO2. Cells were then counted by flow cytometry following DAPI staining (0.5 µg/ml) and 1 × 106 live cells (DAPI negative) were seeded in 2 ml of their respective medium in 12-well plates and treated or not with 10 mM of N-acetyl-L-cysteine (A9165, Sigma) 1-hour prior addition of 2.5 μM of CellROX green probe (C10444, Invitrogen) for additional 30 minutes incubation at 37 °C, 5% CO2. Cells were then collected and washed once with PBS, resuspended in 200µL of PBS, labelled with 0.5 µg/ml of DAPI (D9642, Sigma) and immediately analyzed by flow cytometry (Miltenyi Biotec). CellROX green is a cell-permeable and non-fluorescent reagent when in a reduced state. Upon oxidation, CellROX green exhibits a strong fluorogenic signal (excitation: 485 nm; emission: 520 nm) that is localized primarily in the nucleus and in the mitochondria, providing a reliable measure of reactive oxygen species (ROS) in live cells. Data are represented as the Median Fluorescence intensity (MFI) of the probe detected in live (DAPI-) cells, normalized to control condition (+Asn) and expressed as mean ± SD.
3D cell culture of HCT116 cells
HCT116 spheroids were prepared by seeding 3000 cells/well in 96-well ultra-low attachment plates (650970, Greiner Bio-One). After 72 h, the indicated treatments were applied. Cell viability was assessed by adding propidium iodide (final concentration 0.3 mg/L) to each well at the endpoint. After a 30-min incubation, brightfield and fluorescence images were captured using a fluorescence microscope. The images were then analyzed using the ImageJ software (National Institutes of Health) by calculating the corrected total cell fluorescence (CTCF) using the formula CTCF = integrated density – (area of fluorescent object × mean fluorescence of background readings). The images were artificially colored and cropped for publication.
In vitro stable isotope tracing
For α−15N-L-glutamine tracing experiment, 300.000 live Eµ-Myc cells/ml were incubated in asparagine-free medium supplemented with 2 mM of α−15N-L-glutamine (NLM-1016-PK, Cambridge Isotope Laboratories) instead of unlabeled L-glutamine, and with or without 0.37 mM of L-asparagine (A0884, Sigma), at 37 °C, 5% CO2, for 24 h.
For 13C2-Glycine tracing experiment, 300.000 live Eµ-Myc cells/ml were incubated in asparagine, serine, glycine-free medium supplemented with 0.4 mM of L-serine (S4311, Sigma), 0.4 mM of 13C2-Glycine (CLM-1017, Cambridge Isotope Laboratories), and with or without 0.37 mM of L-asparagine, at 37 °C, 5% CO2, for 24 h.
For [U-13C]-glucose tracing experiment, 300.000 live Eµ-Myc cells/ml were incubated in glucose and asparagine-free medium supplemented with 25 mM of [U-13C]-glucose (CML-1396, Cambridge Isotope Laboratories) and with or without L-asparagine (0.37 mM) and E-Coli L-asparaginase (0.003 IU/ml), at 37 °C, 5% CO2, for 24 h.
At endpoint, Eµ-Myc cells were analyzed by flow cytometry to determine the number of live cells (DAPI-) per mL. Then, 1·106 live Eµ-Myc cells per replicate were collected, washed twice in pre-chilled PBS (300 × g, 5 min, 15 °C centrifugation) and snap-frozen. Samples were stored at −80 °C for further LC-MS-based metabolomic analyses. Results have been corrected for the presence of naturally occurring 15N or 13C stable isotopes using Metabolite AutoPlotter62.
Ex-vivo ATP analysis
ATP was measured using the Cell Titer Glo kit (G7570, Promega). Briefly, 20,000 malignant B cells harvested from freshly isolated B-cell lymphomas were resuspended in 80 µL of DMEM medium supplemented with 10% FBS, 2 mM of L-Glutamine and distributed in a 96 well-plate. Cells were then treated in triplicates for 1 h with either control (DMSO), or 10 μg/ml of oligomycin (O4876, Sigma-Aldrich) or 100 M iodoacetate (I9148, Sigma-Aldrich), or the combination of both metabolic inhibitors to obtain the residual amount of ATP. Short time treatment (1 h) was chosen to avoid cell death upon inhibition of both metabolic pathways. 100 µl of Cell Titer Glo reaction mix was then added in each well for a final volume of 200 µl. Plates were analyzed for luminescence with a Luminoscan (Berthold Technologies). We verified that ATP measurements were in the linear range of the detection. The difference between total ATP production and the ATP produced under oligomycin treatment results in OxPhos ATP contribution. OxPhos ATP is represented by percentage of total ATP produced by the cells.
In vivo stable isotope tracing
When lymphoma development reached the ethical endpoint, mice received two consecutive intraperitoneal discrete boluses injections at 20 min interval of 1 g/kg of [U-13C6]-Glucose (CLM-1396, Cambridge Isotope Laboratories) in sterile NaCl 0.9% (100 mg/ml solution). Blood was collected from the tail-vein 40 min after the first [U-13C6]-Glucose bolus and plasma was isolated by centrifugation at 6000 × g for 2 min and snap-frozen. Mice were immediately sacrificed by cervical dislocation. All B-cell lymphomas were harvested and snap-frozen. All samples collected were stored at −80 °C and axillary BCL were later used for LC-MS-based metabolomic, immunoblot and other indicated biochemical analyses. Results have been corrected for the presence of naturally occurring 13C stable isotopes using Metabolite AutoPlotter62.
Steady state metabolomics in vitro
250.000 live Eµ-Myc cells/ml were treated with indicated drugs for 24 h, in asparagine-containing medium or in asparagine, serine, glycine-free medium supplemented with or without L-asparagine (0.37 mM), L-serine (0.4 mM) and glycine (0.4 mM) at 37 °C, 5% CO2. The number of live cells was determined overtime by DAPI (0.5 µg/ml) staining and flow cytometry analysis (Miltenyi Biotec). Then, 1·106 live cells (DAPI-) per replicate were collected, washed twice in pre-chilled PBS (300 × g, 5 min, 15 °C centrifugation) and snap frozen. Samples were stored at −80 °C for further LC-MS-based metabolomic analyses. The abundance of metabolite is represented as peak area. The GSSG/GSH ratio was calculated from total GSH and GSSG levels determined by LC-MS.
Targeted LC/MS metabolites analyses
For in vitro samples, frozen pellets of 1·106 Eµ-Myc cells were used for metabolite extraction. For lymphomas, frozen axillary B-cell lymphomas were homogenized using a stainless-steel tissue grinder (1292, BioSpec Products) and 10 mg of tissue were used for metabolite extraction. Extraction solution was composed of 50% methanol, 30% acetonitrile (ACN) and 20% water. The volume of the extraction solution was adjusted to the cell number or tissue weight (1 ml per 1 million cells or 1 ml per 50 mg of tissues, respectively). Plasma was diluted 20 folds with the same extraction solvent. After addition of extraction solution, samples were vortexed for 5 min at 4 °C and centrifuged at 16,000 × g for 15 min at 4 °C. The supernatants were collected and stored at −80 °C until analysis.
Targeted metabolomic analyses of in vitro experiments were performed using a minimum of three biological replicates per condition, with the exact numbers of biological and technical replicates provided in the corresponding figure legends. Targeted metabolomic profiling of B-cell lymphomas grown in vivo included 6 to 12 individual samples per group. Stable isotope tracing in vivo was conducted using 4 independent mice per group. Plasma samples analysis included 20 samples per group. Appropriate controls included blank extraction controls, pooled QC samples, and reference standard.
LC/MS analyses were conducted on a QExactive Plus Orbitrap mass spectrometer equipped with an Ion Max source and a HESI II probe coupled to a Dionex UltiMate 3000 UPLC system (Thermo Fisher Scientific). External mass calibration was performed using a standard calibration mixture every seven days, as recommended by the manufacturer. The 5 µl samples were injected onto a ZIC-pHILIC column (150 mm × 2.1 mm; i.d. 5 µm) with a guard column (20 mm × 2.1 mm; i.d. 5 µm) (Millipore) for LC separation. Buffer A was 20 mM ammonium carbonate, 0.1% ammonium hydroxide (pH 9.2), and buffer B was ACN. The chromatographic gradient was run at a flow rate of 0.200 µl min−1 as follows: 0–20 min, linear gradient from 80% to 20% of buffer B; 20–20.5 min, linear gradient from 20% to 80% of buffer B; 20.5–28 min, 80% buffer B. The mass spectrometer was operated in full scan, polarity switching mode with the spray voltage set to 2.5 kV and the heated capillary held at 320 °C. The sheath gas flow was set to 20 units, the auxiliary gas flow to 5 units and the sweep gas flow to 0 unit. The metabolites were detected across a mass range of 75–1000 m/z at a resolution of 35,000 (at 200 m/z) with the automatic gain control target at 106 and the maximum injection time at 250 ms. Lock masses were used to ensure mass accuracy below 5 ppm. Data were acquired with Thermo Xcalibur software (Thermo) and processed using Thermo TraceFinder 5.2 software. The peak areas of metabolites and isotopologues were determined, identified by the exact mass of each singly charged ion and by the known retention time on the HPLC column. For absolute quantification, the standard addition method was used63. For Serine quantification in plasma, d-serine was added to pooled samples, and endogenous levels were subtracted from the spiked standards. Calibration curve was prepared in concentration range of 0.05-1.5 mM in a mixed sample to overcome matrix effects. Importantly, the preparation and processing of the calibration curve samples mirrored that of the experimental samples to ensure consistency.
Software, graphs, and illustrations
ImageJ 2.0 software was used for immunoblots and immunofluorescence relative quantification. All charts and statistics presented in this article were built using GraphPad Prism 9 software (RRID: SCR_002798). Flow cytometry data were analyzed using MacsQuantify 2.13.3 (Miltenyi Biotec). Volcano plots and MSEA analyses from targeted metabolomics were produced with MetaboAnalyst 5.0 (RRID: SCR_015539), and volcano plot colors were edited using Affinity designer 1.6.1 software (RRID: SCR_016952). Data from stable isotope tracing experiments were corrected for the naturally abundance of 15N or 13C stable isotopes using Metabolite AutoPlotter (https://mpietzke.shinyapps.io/AutoPlotter/)62. Mice illustrations are publicly available on https://pixabay.com/fr/vectors/souris-le-noir-animal-rongeur-297524/.
Statistics and reproducibility
All graphical representations and statistical analyses were performed using GraphPad Prism 9 software and MetaboAnalyst 5.0 software. Data are presented as mean ± SD or ± SEM, as indicated in the figure legends. Statistical differences were assessed using the appropriated tests, including two-sided Student’s t-test, one-way or two-way ANOVA followed by Tukey’s or Dunnett’s multiple comparison tests and log-rank test. A P-value < 0.05 was considered statistically significant. No data were excluded from the analyses. For in vitro experiments, samples sizes were chosen according to the common practice of the described assays and are indicated by n or by the number of data points in the graphs. For survival experiments in mice engrafted with Eµ-Myc cells, our previous studies have shown statistical significance with a minimum of 6 mice/group9, a number confirmed using the G*power software as sufficient to detect significant biological meaningful differences between two (or more) experimental groups. All in vivo experiments randomly allocated mice into experimental groups and disease endpoints were evaluated in a blinded manner. Metabolites multivariate statistical analysis was performed with the Metaboanalyst 5.0 software (https://www.metaboanalyst.ca/)64,65. Metabolite relative concentration was normalized by median, and the data was auto-scaled (mean-centered and divided by the standard deviation of each variable) prior to analysis. After scaling the data, we performed principal component analysis. Univariate analysis was performed comparing metabolite levels between groups where metabolite differences were defined by the following fold change threshold 0.8 <Fold change (ASNase/Vehicle) > 1.25 (which corresponds to log2 transformed threshold −0.3219 <log2 (Fold change (ASNase/Vehicle)) > +0.3219) and significance as raw P-value < 0.05 from the non-parametric Wilcoxon rank-sum test, assuming unequal variance. Volcano plots representation and Metabolite Set Enrichment Analysis (MSEA) were performed based on statistically significant discriminant metabolites and built using the Metaboanalyst software and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway computational tool. P-values for MSEA were from Fisher exact test65. For clarity, some graphs display statistics only for the most relevant comparisons due to the large number of groups. For detailed individual p-values, please refer to the Source data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data used to generate the graphs and their respective individual P-values are available in the Source Data file. The RNA-sequencing data (from Eµ-Myc cells cultivated in vitro and from Eµ-Myc cell-derived BCL treated with Vehicle or ASNase in vivo) generated in this study have been deposited in the GEO repository under accession code GSE312117 https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE312117. Processed data are also available at GEO at the same accession. To facilitate data reuse, the targeted metabolomic data generated in this study are provided in the Source Data file, as excel tables containing all the raw values (peak areas) of all detected metabolites. Other remaining data are available within the Article, Supplementary Information, or Source Data file. Source data are provided with this paper.
Code availability
R codes to determine FPKM values from RNA-seq data on Eµ-Myc model or to analyze patients DLBCL RNA-seq data from Reddy et al42. can be accessed on Zenodo66 (https://doi.org/10.5281/zenodo.17737822).
References
Schmidt, D. R. et al. Metabolomics in cancer research and emerging applications in clinical oncology. CA Cancer J. Clin. 71, 333–358 (2021).
Lemberg, K. M., Gori, S. S., Tsukamoto, T., Rais, R. & Slusher, B. S. Clinical development of metabolic inhibitors for oncology. J. Clin. Investig. 132, e148550 (2022).
Stine, Z. E., Schug, Z. T., Salvino, J. M. & Dang, C. V. Targeting cancer metabolism in the era of precision oncology. Nat. Rev. Drug Discov. 21, 141–162 (2022).
Vasan, K. & Chandel, N. S. Molecular and cellular mechanisms underlying the failure of mitochondrial metabolism drugs in cancer clinical trials. J. Clin. Investig. 134, e176736 (2024).
Akagi, T. et al. Methylation analysis of asparagine synthetase gene in acute lymphoblastic leukemia cells. Leukemia 20, 1303–1306 (2006).
Touzart, A. et al. Epigenetic silencing affects l-asparaginase sensitivity and predicts outcome in T-ALL. Clin. Cancer Res. 25, 2483–2493 (2019).
Kucuk, C. et al. Global promoter methylation analysis reveals novel candidate tumor suppressor genes in natural killer cell lymphoma. Clin. Cancer Res. 21, 1699–1711 (2015).
Chiche, J. et al. GAPDH expression predicts the response to R-CHOP, the tumor metabolic status, and the response of DLBCL patients to metabolic inhibitors. Cell Metab. 29, 1243–1257 e1210 (2019).
Grima-Reyes, M. et al. Tumoral microenvironment prevents de novo asparagine biosynthesis in B cell lymphoma, regardless of ASNS expression. Sci. Adv. 8, eabn6491 (2022).
Pathria, G. et al. Translational reprogramming marks adaptation to asparagine restriction in cancer. Nat. Cell Biol. 21, 1590–1603 (2019).
Gwinn, D. M. et al. Oncogenic KRAS regulates amino acid homeostasis and asparagine biosynthesis via ATF4 and alters sensitivity to L-asparaginase. Cancer Cell 33, 91–107 e106 (2018).
Nakamura, A. et al. Inhibition of GCN2 sensitizes ASNS-low cancer cells to asparaginase by disrupting the amino acid response. Proc. Natl. Acad. Sci. USA 115, E7776–E7785 (2018).
Cecconello, D. K. et al. Asparaginase: an old drug with new questions. Hematol. Transfus. Cell Ther. 42, 275–282 (2020).
Mondelaers, V. et al. Prospective, real-time monitoring of pegylated Escherichia coli and Erwinia asparaginase therapy in childhood acute lymphoblastic leukaemia and non-Hodgkin lymphoma in Belgium. Br. J. Haematol. 190, 105–114 (2020).
Schore, R. J. et al. Plasma asparaginase activity and asparagine depletion in acute lymphoblastic leukemia patients treated with pegaspargase on Children’s Oncology Group AALL07P4. Leuk. Lymphoma 60, 1740–1748 (2019).
Iwamoto, S., Mihara, K., Downing, J. R., Pui, C. H. & Campana, D. Mesenchymal cells regulate the response of acute lymphoblastic leukemia cells to asparaginase. J. Clin. Investig. 117, 1049–1057 (2007).
Ehsanipour, E. A. et al. Adipocytes cause leukemia cell resistance to L-asparaginase via release of glutamine. Cancer Res. 73, 2998–3006 (2013).
Tardito, S. et al. The inhibition of glutamine synthetase sensitizes human sarcoma cells to L-asparaginase. Cancer Chemother. Pharm. 60, 751–758 (2007).
Rotoli, B. M. et al. Inhibition of glutamine synthetase triggers apoptosis in asparaginase-resistant cells. Cell Physiol. Biochem. 15, 281–292 (2005).
Guarecuco, R. et al. Dietary thiamine influences l-asparaginase sensitivity in a subset of leukemia cells. Sci. Adv. 6, eabc7120 (2020).
Krall, A. S. et al. Asparagine couples mitochondrial respiration to ATF4 activity and tumor growth. Cell Metab. 33, 1013–1026 e1016 (2021).
Farge, T. et al. Chemotherapy-resistant human acute myeloid leukemia cells are not enriched for leukemic stem cells but require oxidative metabolism. Cancer Discov. 7, 716–735 (2017).
Denise, C. et al. 5-fluorouracil resistant colon cancer cells are addicted to OXPHOS to survive and enhance stem-like traits. Oncotarget 6, 41706–41721 (2015).
Ippolito, L. et al. Metabolic shift toward oxidative phosphorylation in docetaxel resistant prostate cancer cells. Oncotarget 7, 61890–61904 (2016).
Krall, A. S., Xu, S., Graeber, T. G., Braas, D. & Christofk, H. R. Asparagine promotes cancer cell proliferation through use as an amino acid exchange factor. Nat. Commun. 7, 11457 (2016).
Chen, H., Pan, Y. X., Dudenhausen, E. E. & Kilberg, M. S. Amino acid deprivation induces the transcription rate of the human asparagine synthetase gene through a timed program of expression and promoter binding of nutrient-responsive basic region/leucine zipper transcription factors as well as localized histone acetylation. J. Biol. Chem. 279, 50829–50839 (2004).
Gao, S. et al. PSAT1 is regulated by ATF4 and enhances cell proliferation via the GSK3beta/beta-catenin/cyclin D1 signaling pathway in ER-negative breast cancer. J. Exp. Clin. Cancer Res. 36, 179 (2017).
Riscal, R. et al. Chromatin-bound MDM2 regulates serine metabolism and redox homeostasis independently of p53. Mol. Cell 62, 890–902 (2016).
Tameire, F. et al. ATF4 couples MYC-dependent translational activity to bioenergetic demands during tumour progression. Nat. Cell Biol. 21, 889–899 (2019).
Bollino, D. et al. Erwinia asparaginase (crisantaspase) increases plasma levels of serine and glycine. Front. Oncol. 12, 1035537 (2022).
Pacold, M. E. et al. A PHGDH inhibitor reveals coordination of serine synthesis and one-carbon unit fate. Nat. Chem. Biol. 12, 452–458 (2016).
Mullarky, E. et al. Identification of a small molecule inhibitor of 3-phosphoglycerate dehydrogenase to target serine biosynthesis in cancers. Proc. Natl. Acad. Sci. USA 113, 1778–1783 (2016).
Wang, Q. et al. Rational design of selective allosteric inhibitors of PHGDH and serine synthesis with anti-tumor activity. Cell Chem. Biol. 24, 55–65 (2017).
Weinstabl, H. et al. Intracellular trapping of the selective phosphoglycerate dehydrogenase (PHGDH) inhibitor BI-4924 disrupts serine biosynthesis. J. Med. Chem. 62, 7976–7997 (2019).
Kiweler, N. et al. Mitochondria preserve an autarkic one-carbon cycle to confer growth-independent cancer cell migration and metastasis. Nat. Commun. 13, 2699 (2022).
Graham, N. A. et al. Glucose deprivation activates a metabolic and signaling amplification loop leading to cell death. Mol. Syst. Biol. 8, 589 (2012).
Hamano, M. et al. Enhanced vulnerability to oxidative stress and induction of inflammatory gene expression in 3-phosphoglycerate dehydrogenase-deficient fibroblasts. FEBS Open Bio 8, 914–922 (2018).
Zgheib, E. et al. Investigation of Nrf2, AhR, and ATF4 activation in toxicogenomic databases. Front. Genet. 9, 429 (2018).
Hayes, J. D., Dinkova-Kostova, A. T. & Tew, K. D. Oxidative stress in cancer. Cancer Cell 38, 167–197 (2020).
Cheung, E. C. & Vousden, K. H. The role of ROS in tumour development and progression. Nat. Rev. Cancer 22, 280–297 (2022).
Bansal, A. & Simon, M. C. Glutathione metabolism in cancer progression and treatment resistance. J. Cell Biol. 217, 2291–2298 (2018).
Reddy, A. et al. Genetic and functional drivers of diffuse large B-cell lymphoma. Cell 171, 481–494 e415 (2017).
Sun, J. et al. SLC1A3 contributes to L-asparaginase resistance in solid tumors. EMBO J. 38, e102147 (2019).
Hinze, L. et al. Synthetic lethality of Wnt pathway activation and asparaginase in drug-resistant acute leukemias. Cancer Cell 35, 664–676 e667 (2019).
Apfel, V. et al. Therapeutic assessment of targeting ASNS combined with l-asparaginase treatment in solid tumors and investigation of resistance mechanisms. ACS Pharm. Transl. Sci. 4, 327–337 (2021).
Butler, M. et al. BTK inhibition sensitizes acute lymphoblastic leukemia to asparaginase by suppressing the amino acid response pathway. Blood 138, 2383–2395 (2021).
D’Avola, A. et al. PHGDH is required for germinal center formation and is a therapeutic target in MYC-driven lymphoma. J. Clin. Investig. 132, e153436 (2022).
Hope, H. C. et al. Coordination of asparagine uptake and asparagine synthetase expression modulates CD8+ T cell activation. JCI Insight 6, e137761 (2021).
Wu, J. et al. Asparagine enhances LCK signalling to potentiate CD8(+) T-cell activation and anti-tumour responses. Nat. Cell Biol. 23, 75–86 (2021).
Gnanaprakasam, J. N. R. et al. Asparagine restriction enhances CD8(+) T cell metabolic fitness and antitumoral functionality through an NRF2-dependent stress response. Nat. Metab. 5, 1423–1439 (2023).
Chang, H. C. et al. Asparagine deprivation enhances T cell antitumour response in patients via ROS-mediated metabolic and signal adaptations. Nat. Metab. 7, 918–927 (2025).
Strom, C. E. et al. Poly (ADP-ribose) polymerase (PARP) is not involved in base excision repair but PARP inhibition traps a single-strand intermediate. Nucleic Acids Res. 39, 3166–3175 (2011).
Galindo-Campos, M. A. et al. Distinct roles for PARP-1 and PARP-2 in c-Myc-driven B-cell lymphoma in mice. Blood 139, 228–239 (2022).
Elfar, G. A. et al. p53-dependent crosstalk between DNA replication integrity and redox metabolism mediated through a NRF2-PARP1 axis. Nucleic Acids Res. 52, 12351–12377 (2024).
Ferguson, D. C. et al. Amino acid stress response genes promote L-asparaginase resistance in pediatric acute lymphoblastic leukemia. Blood Adv. 6, 3386–3397 (2022).
Bryant, H. E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Parvin, S. et al. LMO2 confers synthetic lethality to PARP inhibition in DLBCL. Cancer Cell 36, 237–249 e236 (2019).
Ricci, A. D. et al. Specific toxicity of maintenance olaparib versus placebo in advanced malignancies: a systematic review and meta-analysis. Anticancer Res. 40, 597–608 (2020).
Rose, M., Burgess, J. T., O’Byrne, K., Richard, D. J. & Bolderson, E. PARP inhibitors: clinical relevance, mechanisms of action and tumor resistance. Front. Cell Dev. Biol. 8, 564601 (2020).
Chiche, J. et al. GAPDH enhances the aggressiveness and the vascularization of non-Hodgkin’s B lymphomas via NF-kappaB-dependent induction of HIF-1alpha. Leukemia 29, 1163–1176 (2015).
Pietzke, M. & Vazquez, A. Metabolite AutoPlotter - an application to process and visualise metabolite data in the web browser. Cancer Metab. 8, 15 (2020).
Luo, B., Groenke, K., Takors, R., Wandrey, C. & Oldiges, M. Simultaneous determination of multiple intracellular metabolites in glycolysis, pentose phosphate pathway and tricarboxylic acid cycle by liquid chromatography-mass spectrometry. J. Chromatogr. A 1147, 153–164 (2007).
Chong, J., Wishart, D. S. & Xia, J. Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr. Protoc. Bioinforma. 68, e86 (2019).
Xia, J. & Wishart, D. S. MSEA: a web-based tool to identify biologically meaningful patterns in quantitative metabolomic data. Nucleic Acids Res. 38, W71–W77 (2010).
Nottet, N. NicolasNottet/Article_ASNase-lymphosome_B_codes_R: Code and data for submitted article. (2025).
Acknowledgements
We gratefully acknowledge the Centre Méditerranéen de Médecine Moléculaire (C3M) animal facility. This study was supported by research fundings from la Ligue Nationale Contre le Cancer “Équipe Labellisée” (EL2021- J-E.R.), the Institut National du Cancer (INCa PLBIO—PLBIO21-062—J-E.R.; INCa PLBIO—PLBIO24-196-2024-158—V.P.; INCa PLBIO—PLBIO2022-113—C.C.), la Fondation ARC pour la recherche sur le cancer (ARCPJA22020060002303—J.C.), le Cancéropôle Provence Alpes Côte d’Azur Institut National du Cancer and Région Sud (J.C. and J-E.R.) and l’Agence Nationale de la Recherche (LABEX SIGNALIFE ANR-11- LABX-0028-01—J-E.R.). C.C. also received fundings from the Agence Nationale de la Recherche (ANR-22-CE51-0043-07) and the Ligue Contre le Cancer comités du Rhône et de la Drôme. A.A. was funded by the University of Nice Côte d’Azur and the Ministère de la Recherche et de l’Enseignement. Samples acquisition and data analysis were performed on the C3M Cytometry Core Facility financed by Conseil Général CG06 and Conseil Régional PACA.
Author information
Authors and Affiliations
Contributions
J.C. performed or participated in all experimental work, data analysis and data interpretation. A.A., A.V., and N.D. performed research experiments and were assisted by P.M., R.P., L.H., L.E., and S.M. I.N. performed LC-MS analysis. A.A., N.D., J.C. and I.N. performed the revision experiments. N.N. performed bioinformatic analysis of DLBCL dataset. C.C. performed research experiments on solid cancer cell lines. C.C., V.P., E.C. and N.M. provided valuable reagents and intellectual inputs for experiments and data interpretation. J.C. and C.C. conceived and designed the study. J.C and J.-E.R secured funding, supervised the study, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aussel, A., Nemazanyy, I., Vandenberghe, A. et al. Tumor metabolic adaptation induced by L-asparaginase reveals a vulnerability to PARP1/2 inhibitor in B-cell lymphomas. Nat Commun 17, 3305 (2026). https://doi.org/10.1038/s41467-026-70066-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70066-2