Abstract
Cancer cells reprogramme translation and metabolism to fuel tumorigenesis. Here, we show that hepatocellular carcinoma (HCC) paradoxically maintains low tyrosine levels despite increased uptake and reduced metabolism, redirecting tyrosine to translation via MYC-driven upregulation of tyrosyl-tRNA synthetase 1 (YARS1) and tRNA-TyrGUA. Restricting tyrosine translation availability (RTTA) via dietary limitation, YARS1/tRNA-TyrGUA ablation, tyrosine degradation (TAL), or YARS1 inhibition (tyrosinol) disturbs this adaptation, leading to the mitigation of tumorigenesis and extension of survival. Mechanistically, RTTA reduces tyrosine codon-dependent translation of mitochondrial complex I subunit NDUFB8 and lipid regulator SCD1, causing complex I misassembly, oxidative phosphorylation failure, and lipid peroxidation-induced ferroptosis. Genome-wide CRISPR screening identifies that loss of GPX4 and BCL2 by genetic manipulation or pharmacological treatment enhances the ability of RTTA to inhibit hepatocellular carcinogenesis. Our findings establish RTTA as a therapeutic strategy targeting tyrosine dependency and highlight combinatorial targeting of translation-metabolism crosstalk and ferroptosis pathways in liver cancer.
Similar content being viewed by others
Introduction
Reprogramming of amino acid utilization pathways to optimise catabolic and anabolic processes, thus promoting cancer cell proliferation and preventing cell death, constitutes a common hallmark of tumorigenic metabolic plasticity1,2,3,4. Specifically, oncogene-driven rewiring of amino acid flux involves fine-tuning between catabolism and protein synthesis due to the increased mRNA translation requirements and the reprogramming of mRNA translation, conferring a demand for specific amino acids5,6. However, a deep understanding of how these two features coordinate in cancer pathogenesis to maintain amino acid utilisation balance is still lacking.
Tyrosine, a large neutral amino acid, is a non-essential amino acid that can be converted from phenylalanine via phenylalanine hydroxylase in vivo7,8. In mammals, some L-type amino acid transport systems, including SLC7A5 and SLC43A1, are responsible for transporting diverse neutral amino acids from the extracellular matrix into the intracellular space, including tyrosine, phenylalanine, leucine, histidine, tryptophan, valine, isoleucine, and alanine9,10. The lysosomal amino acid transporter protein SLC38A9 is required to efficiently transport neutral amino acids back into the intracellular amino acid pool after protein degradation in lysosomes11. In hepatocytes, tyrosine can be degraded, mediated by a five-enzyme cascade (TAT-HPD-HGD-GSTZ1-FAH), to generate fumaric acid and acetoacetate in addition to its role as a constituent unit of proteins12. Consequently, when there is an inborn functional deficiency of hepatic tyrosine catabolic enzymes, tyrosine and its downstream metabolites accumulate in the body, which significantly affects newborn survival and childhood development, i.e., tyrosinemia13,14. Clinically, long-term tyrosine-restricted diets are established for disease management in these patients, demonstrating the feasibility of tyrosine-restricted diets in HCC patients.
In this study, we found that HCC cells hijack tyrosine uptake and degradation to shunt tyrosine towards protein synthesis, representing a critical adaptation in response to hepatocellular carcinogenesis, thereby remodelling the balance of tyrosine utilisation. Restriction of tyrosine translation availability through dietary intervention, genetic manipulation, or pharmacological inhibition disturbs this delicate balance, which then impairs oxidative phosphorylation and triggers ferroptosis, ultimately alleviating tumorigenesis and potentiating cancer therapy combined with sorafenib or venetoclax, which represents a potential therapeutic window for patients with liver cancer.
Results
HCC hijacks tyrosine uptake and degradation to preserve oncogenic tyrosine levels
To determine whether amino-acid levels were changed in HCC, targeted metabolomics on tumour and normal liver tissues obtained from human and mouse sources was performed, of which the mouse liver tumours were isolated from H11LNL-Myc; Alb-cre mice (hereafter referred to as Myc/Alb-cre). In this mouse model, hepatocyte-specific overexpression of MYC sequentially drove the development of hepatomegaly and multiple high-grade tumours in the mouse liver within 12 weeks of age15. Irrespective of whether the tissues were from humans or mice, tyrosine was the most significantly decreased amino acid in the tumour compared to the normal liver (Fig. 1a, b). Our observations were consistent with the previous metabolomics results in human HCC patients (n = 130) and mTOR-hyperactivation-driven HCC mouse model1,16. Meanwhile, tumour tyrosine levels were negatively correlated with tumour diameter (Fig. 1c). We also isolated interstitial fluid from liver tumours of different sizes to measure tyrosine concentration. The results showed no significant correlation between tumour size and tyrosine levels in the tumour interstitial fluid, indicating that tumour size did not affect local tyrosine availability (Supplementary Fig. 1a). Therefore, we speculated whether high levels of tyrosine favour liver tumorigenesis.
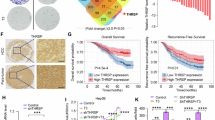
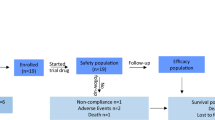
a, b Relative amino acid profile of tumour tissues relative to normal liver tissues from humans (a) (n = 6 patients) or mice (b) (n = 6 mice). c Pearson correlation between tumour diameter and amino acid level in Myc/Alb-cre mice. (n = 11 mice). d A diagram depicting the in vivo assessment of hepatocellular carcinogenesis in mice. e Photographs of the liver from the tumour-bearing mice fed either a standard diet or a tyrosine-restricted diet. f Body weight excluding liver weight, liver to body weight ratio, tumour number and maximum tumour diameter of the tumour-bearing mice were shown, including Myc/Alb-cre mice (top) (n = 6 mice), C57BL/6 J mice (middle) (n = 8 mice) and BALB/c nude mice (bottom) (n = 5 mice). g Kaplan–Meier survival graph of tumour-bearing mice, including Myc/Alb-cre mice (left) (n = 11 mice), C57BL/6 J mice (middle) (n = 12 mice), or BALB/c nude mice (right) (n = 8 mice). h Relative 3H-tyrosine uptake into normal liver and Myc/Alb-cre tumour. (n = 8 mice). i Schematic representation of changes in RNA and protein expression of tyrosine transporters and metabolic enzymes in HCC patients based on the data from CPTAC. j, k Immunoblots of tyrosine transporters and tyrosine-metabolising enzymes in tumour tissues and normal liver tissues from human (j) (n = 4 patients) and Myc/Alb-cre mice (k) (n = 4 mice). β-Actin serves as a loading control. l Kaplan–Meier survival analysis correlating tyrosine transporters expression and tyrosine-metabolising enzymes expression with overall survival in TCGA-LIHC patients (n = 165 low group and n = 165 high group). m Summary of increased tyrosine uptake and reduced tyrosine metabolism in HCC. Diethylnitrosamine (DEN). d was generated using Adobe Illustrator 2025. j, k the samples are derived from the same experiment, and all gels/blots are processed in parallel under identical conditions. Source data are provided as a Source Data file. Data are presented as means ± SD, two-tailed Student’s t test (a, b, f, h, i, j), Pearson’s coefficient with two-tailed test (c), two-sided log rank test (g, l).
To identify the role of tyrosine in hepatocellular carcinogenesis in vivo, we established three mouse HCC models, including a MYC overexpression-driven HCC model, a DEN treatment-induced HCC model, and an intrahepatic injection model of human HCC cells to validate the effect of tyrosine levels on HCC development. Mice were fed diets containing 0% or 500% of the normal tyrosine levels in the standard diet (tyrosine 100%). Surprisingly, we found that restriction of dietary tyrosine intake (Tyr 0% diet, hereafter referred to as TR diet) led to a significant decrease in characteristic hepatomegaly, liver-to-body weight ratio, tumour burdens, and diameters relative to the standard diet (Tyr 100% diet) (Fig. 1d–f). This phenomenon was opposite to the supplementation of dietary tyrosine intake (Tyr 500% diet) (Supplementary Fig. 1b–d). Remarkably, the TR diet prolonged the survival time in tumour-bearing mice, while the tyrosine-supplemented diet shortened the survival time of these mice (Fig. 1g). Moreover, at 9 weeks of age after tumours had already developed in Myc/Alb-cre mice, tyrosine-restricted diets were applied to evaluate their impact on established tumorigenesis. Results also demonstrated that restriction of dietary tyrosine intake (Tyr 0% diet) significantly inhibited liver tumour development compared to the standard diet (Tyr 100% diet) (Supplementary Fig. 1e–g). These findings indicate that tyrosine-restricted diets exert significant therapeutic efficacy against HCC development during tumour initiation and establishment. Tyrosine restriction mainly reduced the tyrosine levels in the liver, with minimal effects on other amino acids (Supplementary Fig. 1h). To consolidate the above findings, we also determined whether low tyrosine abundance inhibits cancer cell growth in vitro. Consistent with in vivo effects, tyrosine restriction (Tyr 0%) substantially reduced cell viability and induced cell cycle arrest, along with increasing the ratio of early and late apoptotic cells compared with the standard medium (Tyr 100%) (Supplementary Fig. 2a–g), whereas tyrosine supplementation promoted tumour proliferation (Supplementary Fig. 2h–j). Together, these results indicated that tyrosine abundance is critical for tumorigenicity.
Moreover, to evaluate the systemic consequences of tyrosine-deficient diets in mice, wild-type mice were fed with a 10-week tyrosine-deficient diet to assess tyrosine restriction effects on body and organ weights and food intake. The results demonstrated that tyrosine restriction increased total body weight and fat mass, along with an increase in liver weight (Supplementary Fig. 3a, b). No significant differences were observed in the weight of other organs or in daily food intake (Supplementary Fig. 3b, c). Concurrently, the levels of alanine aminotransferase (ALT), aspartate aminotransferase (AST), triglyceride (TG), total cholesterol (TC), creatinine (CREA), creatine kinase (CK), and lactate dehydrogenase (LDH), established biomarkers for liver, kidney, and myocardial injury, were measured. Tyrosine-deficient diets induced liver injury in mice, whereas no significant effects on kidney and heart functions (Supplementary Fig. 3d). Furthermore, tyrosine-deficient mice showed the lipid droplet accumulation in liver, whereas no pathological changes or tissue injury were observed in other organs (Supplementary Fig. 3e). Moreover, given tyrosine’s role as a dopamine precursor, we evaluated depression-like behaviours using the tail suspension test (TST), a method for assessing depressive states in mice. No significant behavioural differences were observed between the two groups after 10 weeks of diet intervention (Supplementary Fig. 3f). These findings indicate that tyrosine deprivation induces lipid droplet accumulation in wild-type mice, consistent with previous studies that combined dietary deprivation of tyrosine and methionine with an HFD diet disrupts lipid metabolism17.
Next, we speculated whether the low levels of tyrosine in HCC tissues were due to reduced tyrosine uptake or increased tyrosine metabolism. To investigate this hypothesis, tyrosine transport was measured by an ex vivo assay using 3H-tyrosine and an in vitro assay using 13C-tyrosine. These assays confirmed that tyrosine uptake was upregulated in HCC (Fig. 1h and Supplementary Fig. 4a). However, 13C-tyrosine metabolic tracing assay in primary hepatocytes isolated from normal or HCC mouse livers revealed a decreased conversion of tyrosine to its downstream metabolites, including fumarate and citrate, in primary cancer cells compared with primary normal hepatocytes (Supplementary Fig. 4b, c). Consistently, the downstream metabolites of tyrosine were also decreased in HCC tissues (Supplementary Fig. 4d). These results suggested that the rapid influx of tyrosine to its downstream metabolites did not occur, despite increased uptake in HCC. Consistent with our observations, transcriptomic and proteomic analyses of HCC datasets (The Cancer Genome Atlas (TCGA) and The Clinical Proteomic Tumour Analysis Consortium (CPTAC)) also showed that the levels of tyrosine transporters such as SLC7A5, SLC43A1, and SLC38A9 were increased in tumour tissues compared with adjacent normal tissues, whereas the levels of tyrosine metabolism proteins such as TAT, HPD, HGD, FAH, and GSTZ1 were decreased in HCC samples (Fig. 1i, Supplementary Fig. 4e, f, and Supplementary Fig. 5a–g). Gene set enrichment analysis (GSEA) also indicated that the tyrosine metabolism pathway was downregulated in HCC tissues and cells (Supplementary Fig. 5b, e, and g). Finally, we validated the upregulation of tyrosine transporters and loss of tyrosine metabolism proteins in tumours by Western blotting and immunohistochemistry (Fig. 1j, k, Supplementary Fig. 4g, h, and Supplementary Fig. 5h, i). Furthermore, we found that SLC7A5 transporter inhibitor (JPH203) treatment resulted in a significantly reduction tyrosine uptake in liver cancer cells to further corroborate the upregulation of tyrosine uptake in HCC (Supplementary Fig. 4i). Moreover, based on the TCGA dataset, upregulation of tyrosine transporters and loss of tyrosine metabolism genes were associated with a worse prognosis for HCC patients, with the most significant effects reflecting the protective role of tyrosine in tumorigenesis (Fig. 1l). Thus, our findings indicated that liver tumours increase tyrosine uptake and reduce tyrosine metabolism to compensate for another intracellular pathway that consumes tyrosine (Fig. 1m).
MYC fuels tyrosine translation availability via YARS1 in HCC
In addition to undergoing metabolism, another primary consumption of intracellular tyrosine is its role as a building block in protein synthesis. We presumed that the diversion of tyrosine utilisation, i.e., reduced tyrosine metabolism in HCC tissues, was attributed to accommodating the high requirements of the tumour proteome (Fig. 2a). To validate this hypothesis, an ex vivo isotopic tracer method using 3H-tyrosine confirmed that both the total amount and the relative proportion of tyrosine incorporated into protein synthesis were indeed increased in liver cancer compared to normal liver (Fig. 2b, c). Consistent with this observation, the tRNA-TyrGUA, a tyrosine cognate isoacceptor tRNA, exhibited the most marked elevation among all tRNAs in HCC relative to normal tissues by tRNA RT-qPCR (after tRNA de-modification) and Northern blotting (Fig. 2d and Supplementary Fig. 6a). In contrast, the expression of another tRNA-TyrAUA isoacceptor did not change, which was associated with the low distribution of its cognate TAT codon in the human transcriptome (Fig. 2d and Supplementary Fig. 6a, b). Moreover, tyrosyl-tRNA synthetase 1 (YARS1) expression was upregulated in HCC and correlated with poor survival among HCC patients (Fig. 2e, f and Supplementary Fig. 6c–f). Subsequently, in-depth analysis of amino acid and codon enrichment in proteins with translation upregulated and downregulated in HCC patients further confirmed that the content of tyrosine and its cognate TAC codon was significantly increased in translation-upregulated proteins (unchanged at mRNA levels but increased at protein levels) as compared with translation-downregulated proteins (unchanged at mRNA levels but decreased at protein levels) (Fig. 2g, h). Taken together, our findings indicated that unmetabolized tyrosine is utilised in protein synthesis, promoting tumorigenicity and progression.
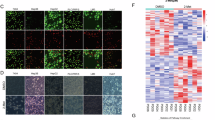
a Schematic showing the canonical pathways for intracellular utilisation of tyrosine. b Relative 3H-tyrosine levels of ribosome (left) and protein (right) in tumour tissues compared to normal tissues from Myc/Alb-cre mice. (n = 8 mice). c Percentage of 3H-tyrosine in ribosome and protein to 3H-tyrosine uptake in normal liver and tumour tissues. (n = 8 mice). d Volcano plot showing differentially expressed tRNA between HCC and normal liver tissues. (n = 18 patients). e Immunoblots of YARS1 in tumour tissues and normal liver tissues from human (top) (n = 4 patients) and Myc/Alb-cre mice (bottom) (n = 4 mice). f Kaplan–Meier survival curve for TCGA-LIHC patients with high (n = 165 patients) or low (n = 165 patients) expression of YARS1. g A graph representing gene levels detected by RNA sequencing and proteomics of HCC tissues and normal liver tissues. h A heat map representing amino acids and codon enrichment analysis in genes translating upregulated compared with the genes translating downregulated based on the data from g. i ChIP qPCR analysis in SUN398 cells detecting indicated binding sites on YARS1 promoter. (n = 3 independent experiments). j Immunoblot analysis of the protein of YARS1 in SNU398 cells. (n = 3 independent experiments). k–m Representative images (k) of the liver (n = 6 mice), tumour statistics (l) (n = 6 mice), and a survival curve (m) (n = 8 mice) in mice are shown. n–p Representative liver images (n), tumour statistics (o) (n = 6 mice), and a survival curve (p) (n = 8 mice) in Myc/Alb-cre transgenic mice infected with AAV8-shNC or AAV8-shYARS1 are shown. e, j The samples are derived from the same experiment, and all gels/blots are processed in parallel under identical conditions. Source data are provided as a Source Data file. Data are representative of three independent experiments (i, j). Data are presented as means ± SD, two-tailed Student’s t test (b–d, h, i), two-sided Wald test with Benjamini–Hochberg adjustments (g), one-way ANOVA followed by multiple comparisons (l, o), two-sided log-rank test (f, m, p).
Moreover, to further identify that the increased translational utilisation of tyrosine is a specific feature of liver tumour progression, the uptake of tyrosine, lysine, and alanine between tumour tissues and normal tissues was measured. We found that while the uptake rates of both tyrosine and lysine were significantly elevated in tumour tissues compared to normal tissues, only tyrosine exhibited a significantly higher incorporation ratio into protein synthesis within HCC (Supplementary Fig. 6g). We also found that lysine-tRNA ligase (KARS1) and alanine -tRNA synthetase (AARS1), responsible for lysine and alanine translational utilisation, respectively, were significantly downregulated in tumour tissues (Supplementary Fig. 6h). Correspondingly, the levels of tRNA-LysCUU showed a significant decrease and the levels of tRNA-Ala remained unchanged (Supplementary Fig. 6i), implying that an increase in amino acid uptake, alongside elevated levels of aminoacyl-tRNA synthetases and cognate isoacceptor tRNA, is essential for promoting translational utilisation of amino acids.
Gene amplification of MYC is observed across multiple human cancer types, explicitly accounting for 11.56% of HCC patients18. MYC hyperactivation or overexpression, alone or in combination with other oncogenic mutations, drives active translation, thus increasing total protein production in many malignancies, contributing to its ability to regulate tRNA and ribosome biogenesis19,20,21,22. Here, we found MYC potently bound to the promoter of YARS1 in multiple human and mouse cells by analysing the chromatin immunoprecipitation (ChIP)-sequencing database (Supplementary Fig. 7a, b). Indeed, ChIP and luciferase reporter assays also confirmed that a conserved element within the YARS1 promoter was responsible for its interaction with MYC (Fig. 2i and Supplementary Fig. 7c). Notably, the expression of YARS1 exhibited the most marked reduction among all aminoacyl-tRNA synthetases after knocking down MYC in SNU398 cells, indicating that tyrosine translational utilisation, for which YARS1 is responsible, plays a critical role in increased mRNA translation requirements in hepatocellular carcinoma (Supplementary Fig. 7d). Furthermore, MYC knockdown also decreased the protein levels of YARS1 (Fig. 2j). Consistent with these findings, a positive correlation was observed between the mRNA levels of MYC and YARS1 (Supplementary Fig. 7e, f). We also found that only the expression levels of TAT and FAH were negatively correlated with MYC expression, indicating that the reduced levels of tyrosine metabolic genes are not fully dependent on MYC expression (Supplementary Fig. 7g). Collectively, these results showed a direct link for oncogenic MYC signalling in increasing the expression of YARS1, which identifies a role that oncogenic transcriptional programmes fulfil requirements in protein synthesis in cancer cells by upregulating aminoacyl-tRNA synthetase biogenesis in HCC. To assess the functional contribution of tyrosine translation bioavailability in developing HCC, we first explored the results from the published genome-wide HCC screen dataset based on the Cancer Dependency Map (DepMap) project. Among all aminoacyl-tRNA synthetases, YARS1 ranked as the second cell-survival-dependent gene in multiple HCC cell screens. In contrast, the tyrosine metabolism gene set, especially TAT, exhibited a cell-survival-independent pattern, i.e., knocking out the tyrosine metabolism gene set promoted HCC survival, which is also consistent with previous reports23,24,25 (Supplementary Fig. 7h, i). This result suggests that targeting tyrosine translation bioavailability represents a vulnerability in liver cancer, wherein improving the translation utilisation of tyrosine enhances cell fitness, whereas its disruption impairs cell fitness.
Moreover, to further illustrate that tyrosine dependency is a common feature of liver tumorigenesis, the expression levels of YARS1 were first evaluated. We found that YARS1 expression was upregulated in the DEN-induced mouse liver tumour model (Supplementary Fig. 8a). Next, wild-type mice were injected with the plasmids of myr-AKT and NRASG12V via hydrodynamic tail-vein injection to induce HCC tumorigenesis, together with feeding a tyrosine-modified diet. The results demonstrated that tyrosine restriction also significantly attenuated the progression of AKT/NRASG12V-driven HCC (Supplementary Fig. 8b–d). Furthermore, we found that hepatocellular carcinoma’s dependence on tyrosine was determined by its protein synthesis activity. Specifically, tumour cells exhibiting higher protein synthesis activity showed greater inhibition of cell growth upon tyrosine restriction. We measured protein synthesis activity, the expression levels of MYC, YARS1, and the tyrosine metabolism gene set in seven HCC cell lines and one normal hepatic cell line (THLE-2). The results revealed that HCC lines demonstrated higher protein synthesis activity, elevated YARS1 expression, and reduced tyrosine metabolism gene set compared to normal hepatocytes (Supplementary Fig. 8e). Tyrosine restriction was less effective at inhibiting cell proliferation of the cell lines with low-protein synthesis activity (THLE-2 or HEPG2), compared to those with high protein synthesis activity (SNU398 or PLCPRF5) (Supplementary Fig. 8f). Moreover, we found that overexpressing MYC in SNU449 cells with low-protein synthesis activity to enhance protein synthesis activity, increased the inhibitory effect of tyrosine restriction, whereas MYC knockdown in SNU398 cells with high protein synthesis activity to reduce protein synthesis activity, decreased this inhibitory effect of tyrosine restriction (Supplementary Fig. 8g). Collectively, these results demonstrate that tyrosine dependency in HCC cells is primarily governed by global protein synthesis activity rather than being exclusively restricted to MYC-driven carcinogenesis.
Moreover, we found that knocking down YARS1 or tRNA-TyrGUA reduced the fitness of HCC cells (Supplementary Fig. 9a–p). Next, control or YARS1-depleted HCCLM3 cells were implanted in the BALB/c nude mouse liver. Consistent with in vitro effects, YARS1 depletion reduced tumour burdens and tumour diameters, which in turn prolonged the survival time of tumour-bearing mice (Fig. 2k–m). Furthermore, we injected adeno-associated virus 8 (AAV8) into Myc/Alb-cre mice at one month of age to specifically knock down YARS1 in hepatocytes. Similar to our previous results, Myc/Alb-cre mice injected with AAV8-shYARS1 developed significantly fewer tumours with smaller diameters and an extended survival time of tumour-bearing mice (Fig. 2n–p and Supplementary Fig. 9q). Taken together, the presented experiments demonstrated that liver cancer is sensitive to the deletion of YARS1 and tRNA-TyrGUA-mediated limitation of tyrosine translation availability.
RTTA decreases the translation of NDUFB8 and SCD1 in liver cancer
To demonstrate whether the limitation of tyrosine is sufficient to reduce tyrosine translation bioavailability, we validated it through two approaches. First, after the restriction of tyrosine or YARS1 depletion either in vitro or in vivo, the charged tRNA-TyrGUA levels were reduced (Supplementary Fig. 10a, b). Second, upon sensing decreased levels of tRNA-TyrGUA charging, YARS1 underwent nuclear translocation (Supplementary Fig. 10c, d)26,27. Thus, we integrated the transcriptomics, translatomics, proteomics, and metabolomics of tyrosine-limited culture (Tyr 0%) and normal culture (Tyr 100%) to systematically elucidate the mechanism that underlies the effects of restriction of tyrosine translation availability in HCC (Supplementary Fig. 10e). First, the quality of our ribosome-profiling was assured, characterised by the nucleotide length and trinucleotide periodicity distribution of coding sequences in ribosome-protected fragments (Supplementary Fig. 10f, g). Our results also revealed that tyrosine deprivation and YARS1-knockdown did not change global translation efficiency (Supplementary Fig. 10h–j). Of note, by integrative analysis of matched transcriptomic and translatomics data, we found the content of tyrosine and its codon was higher in translation-downregulated protein (unchanged at mRNA levels but reduced at translation levels) compared to translation-upregulated protein upon tyrosine deprivation treatment (Fig. 3a, b), which also suggested that translation defects were observed at tyrosine-enriched transcripts. Next, by combinatorial analysis of matched transcriptomic and proteomics data or translatomics and proteomics data, we defined the tyrosine translational sensitive signature upon tyrosine restriction as candidates with the following characteristics: (1) unchanged at the RNA levels, (2) downregulated at translation efficiencies, (3) reduced at protein levels, (4) enriched in tyrosine and its codon (Fig. 3c–e). In this tyrosine signature, we mainly focused on two proteins as the most prominent translational targets of tyrosine restriction: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 8, mitochondrial (NDUFB8) and acyl-CoA desaturase 1(SCD1), due to proteomics and metabolomics results showing that tyrosine restriction affected oxidative phosphorylation and unsaturated fatty acid metabolism (Fig. 3f). Indeed, mRNAs encoding SCD1 and NDUFB8 showed significantly decreased translation in tyrosine-restricted conditions, with minimal changes in the mRNA transcription, whereas the translation of NDUFB1 with low tyrosine codon content remained unchanged (Fig. 3g). We also found that tyrosine restriction showed a higher frequency of ribosomal footprints at the TAC codon in the SCD1 and NDUFB8 transcript, with predominant A-site stalling rather than P-site stalling (Supplementary Fig. 10k). Footprint accumulation density analysis further revealed that tyrosine restriction showed a higher P-site accumulation at the codon immediately preceding the 5’ end of the TAC codon in the SCD1 transcript, confirming that tyrosine deprivation leads to A-site stalling of ribosomes at the TAC codon in the SCD1 transcript (Supplementary Fig. 10l). However, no significant footprint accumulation density difference was observed in the NDUFB8 transcript, potentially due to insufficient number of RPFs aligned to this transcript (Supplementary Fig. 10l).
a–d The graphs representing gene levels detected by transcriptomics and translatomics (a), transcriptomics and proteomics (c) or translatomics and proteomics (d) of SNU398 tyrosine-restricted cells compared to control cells. A heat map (b) representing amino acids and codon enrichment analysis in genes translating downregulated compared to the genes translating upregulated. e The frequency of tyrosine and tyrosine cognate TAC codon in coding sequence is sorted from high to low. f The graph representing significantly altered signatures in proteomics and metabolomics data upon tyrosine restriction. g Ribosome-protected fragment and total RNA tracks at SCD1, NDUFB8, and NDUFB1 gene loci are shown. h A graphic illustration of the indicated reporter with a different tyrosine codon frequency. Quantification of luciferase activities normalised to Renilla luciferase activities in SNU398 cells. (n = 3 independent experiments). i, j Native-PAGE analysis (left) and SDS-PAGE analysis (right) of SNU398 cells (i) (n = 3 independent experiments) or the livers from Myc/Alb-cre mice (j) (n = 6 mice). k, l The activity of mitochondrial complex I-V of SNU398 cells (k) (n = 3 independent experiments) or the livers of Myc/Alb-cre mice (l) (n = 6 mice). m Seahorse assays showing the oxygen consumption rate (OCR) (left) in SNU398 cells. Quantification of maximal respiration (right). (n = 4 independent experiments). n Representative TEM images of SNU398 cells (top) (n = 3 independent experiments) or the livers from Myc/Alb-cre mice (bottom) (n = 6 mice). Morphometric analysis of maximal cristae width of cells or livers. o Representative Ki-67, TUNEL, and 4-HNE staining images and quantification statistics of liver sections. (n = 6 mice). i, j The samples are derived from the same experiment, and all gels/blots are processed in parallel under identical conditions. Source data are provided as a Source Data file. Data are representative of three independent experiments (h, i, k, m, n). Data are presented as means ± SD, two-tailed Student’s t test (b, h, k–o), two-sided Wald test with Benjamini-Hochberg adjustments (a, c, d, f).
As further support for the above observation, we found proteins reduced upon tyrosine deprivation were enriched in pathways associated with the electron transport chain (ETC), including mitochondrial respiratory chain complex I assembly, proton motive force-driven mitochondrial ATP synthesis, oxidative phosphorylation and NADH dehydrogenase (ubiquinone) activity (Supplementary Fig. 11a–c). Thus, we found that the expression of mitochondrial proteins (especially NDUFB8) enriched in tyrosine (Fig. 3e), along with several of those that were translationally reduced in tyrosine-limited conditions, was generally reduced upon tyrosine restriction, thereby impairing the mitochondrial proteostasis. Moreover, proteins upregulated upon tyrosine deprivation were clustered in the hallmarks of metabolic processes, including ferroptosis, aminoacyl-tRNA biosynthesis, amino acid metabolism, autophagy, etc (Supplementary Fig. 11c). Interestingly, ferroptosis suppressors such as GPX4 and SLC7A11 were significantly elevated in tyrosine-limited culture (Tyr 0%) (Supplementary Fig. 11d), a phenomenon explained by integrated stress response (ISR) and metabolomics results. First, we observed significant activation of the GCN2-eIF2α-ATF4 axis upon tyrosine restriction, as shown by increased phosphorylation levels of GCN2 and eIF2α, and elevated protein levels of ATF4, indicating that ISR is activated to respond to tyrosine translation limitation and reestablish physiological homoeostasis (Supplementary Fig. 11e). Next, we found that the intracellular levels of tyrosine were reduced after tyrosine limitation (Supplementary Fig. 11f). However, while tyrosine restriction enhanced the cellular capacity to convert phenylalanine to tyrosine, this increased conversion was insufficient to compensate for tyrosine consumption, ultimately resulting in a significant decrease in intracellular tyrosine levels (Supplementary Fig. 11g, h). Importantly, we noticed that metabolites increased upon tyrosine deprivation were enriched in pathways associated with the unsaturated fatty acids, including biosynthesis of unsaturated fatty acids, arachidonic acid metabolism, linoleic acid metabolism, and alpha-linolenic acid metabolism (Supplementary Fig. 11i, j). In agreement with this finding, monounsaturated fatty acids (MUFAs) represented by palmitoleic acid (C16:1) and oleic acid (C18:1) were decreased upon tyrosine deprivation, whereas polyunsaturated fatty acids (PUFAs), including linoleic acid (C18:2), arachidonic acid (C20:4) and adrenic acid (C22:4) showed an increasing trend (Supplementary Fig. 11i). These changes may be attributed to downregulation of SCD1, as well as elevated extracellular fatty acid uptake following SCD1 reduction, and ISR activation-induced PUFA accumulation28,29,30,31,32,33. Given that PUFAs are more prone to peroxidation, which can trigger ferroptosis34, our results also validated that tyrosine deprivation could effectively induce the formation of oxidised lipid species, with long-chain oxidised phosphatidylethanolamines being the most prominent (Supplementary Fig. 11k). We consistently observed greater lipid peroxidation upon RTTA treatment, using the probe C11 BODIPY 493/ 503 (Supplementary Fig. 12a–f). On the other hand, increased PUFAs were further validated with lipid droplet accumulation after RTTA treatment (Supplementary Fig. 12g–j) to sequester and store excess PUFAs into lipid droplets, where they are protected against lipid peroxidation35. Thus, tyrosine restriction reduces the tyrosine codon-dependent translation of SCD1, resulting in a reduced generation of MUFAs, thereby leading to lipid peroxidation and ferroptosis, despite the cells’ attempt to counteract ferroptosis by increasing the expression of GPX4 and SLC7A11.
To further support our sequencing results that translation of tyrosine-rich NDUFB8 and SCD1 is more sensitive to tyrosine limitation, we first developed reporters expressing luciferase in fusion with SCD1 (8.73%, tyrosine), NDUFB8 (9.17%, tyrosine) or NDUFB1 (1.72%, tyrosine), which are proteins that have different tyrosine codon frequencies. We found that the translation of the SCD1-Luc reporter and either the NDUFB8-Luc reporter was more sensitive to RTTA treatment than that of the NDUFB1-Luc reporter (Fig. 3h and Supplementary Fig. 13a, b). Moreover, compared to transfection of WT reporter constructs for NDUFB8 or SCD1, the relative luciferase activities in SNU398 cells transfected with MUT reporter constructs (mutating TAC to TAT) were increased under tyrosine-free or YARS1-knockdown conditions (Supplementary Fig. 13c–e). We also found that compared to overexpression of WT plasmids for NDUFB8 or SCD1, overexpression of MUT plasmids (mutating TAC to TAT) for NDUFB8 or SCD1 effectively improved the impaired mitochondrial complex I function induced by tyrosine restriction (Supplementary Fig. 13f) and alleviated tyrosine restriction-induced elevation in lipid peroxidation (Supplementary Fig. 13g, h). These findings indicate that restricting tyrosine translation availability has a greater translation inhibition effect on TAC-rich transcripts than on TAT-rich transcripts. Next, the reduced translation of SCD1 and NDUFB8 mediated by RTTA treatment was confirmed by ribosome immunoprecipitation assay using RPL22-Flag expressing SNU398 cells (Supplementary Fig. 13i–k). Indeed, YARS1 expression was significantly and positively correlated with NDUFB8 or SCD1 in HCC patients (Supplementary Fig. 13l). Thus, these findings further supported that tyrosine codon-dependent translation is regulated by tyrosine and YARS1 along with tyrosine cognate tRNA-TyrGUA.
Furthermore, to further authenticate the functional consequences of translational reduction of NDUFB8 and SCD1 in impairing ETC activity and inducing ferroptosis, blue native-PAGE and SDS-PAGE assays were conducted to assess the assembly profiles of the ETC complexes and the expression levels of SCD1. We found that RTTA treatment could specifically reduce the complex I assembly and the expression of SCD1 in vitro and in vivo (Fig. 3i, j, and Supplementary Fig. 14a, b). We also found that tyrosine supplementation could increase the protein expression of SCD1 and NDUFB8 in vitro and in vivo (Supplementary Fig. 14c, d), but tyrosine restriction had minimal impact on NDUFB8 and SCD1 protein levels in adjacent normal liver tissues of tumour-bearing mice (Supplementary Fig. 14e). Additionally, we selected tryptophan and serine to investigate their deprivation impacting on NDUFB8 and SCD1 expression. While the frequency of tryptophan in NDUFB8 and SCD1 ranked similarly to tyrosine, tyrosine deprivation induced a significantly greater reduction in protein levels than tryptophan deprivation, while serine deprivation showed no significant impact on these two protein levels (Supplementary Fig. 14f, g), indicating that tyrosine availability exerts more profound translational control than the availability of tryptophan and serine in liver cancer. Similarly, we found that RTTA treatment led to defective complex I activity with little effect on other ETC complexes in vitro and in vivo, as evaluated by enzyme catalytic activity (Fig. 3k, l and Supplementary Fig. 15a, b). Consistently, RTTA treatment reduced OCR with a significant decrease in the maximal respiration capacity due to a defective mitochondrial function caused by complex I misassembly (Fig. 3m and Supplementary Fig. 15c, d). Impaired oxidative phosphorylation was also associated with the generation of cytotoxic mitochondrial ROS (Mito-ROS)36, as measured by an increased Mito-ROS following tyrosine restriction and knocking down of YARS1 and tRNA-TyrGUA (Supplementary Fig. 15e–g).
In addition, given that functional defects in the respiratory chain complexes and ferroptosis can both impair the morphology and structure of mitochondria, limiting tyrosine diet and knocking down YARS1 and tRNA-TyrGUA in vitro and in vivo consistently showed severe ferroptosis and respiration defect characteristics, including mitochondrial atrophy, an increase in mitochondrial membrane density and cristae width, cristae disorganisation and extensive lipid droplet formation, suggesting morphology and structural abnormalities that reflect mitochondrial defects (Fig. 3n and Supplementary Fig. 16a–j). Of note, related pathologic changes, including reduced cell proliferation (characterised by Ki-67), increased cell apoptosis (characterised by TUNEL), and increased lipid peroxidation (characterised by 4-HNE), again indicated that restricting tyrosine translation availability resulted in defective function of mitochondria and lipid peroxidation (Fig. 3o and Supplementary Fig. 16k–n). Overall, these findings provide evidence that targeting tyrosine translation bioavailability either through tyrosine limitation or knocking down YARS1 and tRNA-TyrGUA leads to a reduction in the tyrosine codon-dependent translation of NDUFB8 and SCD1, resulting in defective assembly of mitochondrial complex I and lipid peroxidation.
Genome-wide CRISPR screen identifies factors that enhance the efficacy of tyrosine-restricted therapy
According to our proteomics results, we found that multiple proteins and pathways were upregulated following tyrosine restriction, such as GPX4 and SLC7A11 in the ferroptosis pathway and YARS1 in the aminoacyl-tRNA synthetase pathway, indicating that such genetic interactions might impinge on the anti-tumour efficacy of tyrosine restriction. Therefore, a whole-genome CRISPR–Cas9 loss-of-function screen was conducted to systematically elucidate the underlying mechanism of tyrosine dependence in HCC (Fig. 4a). By analysing sgRNAs depleted in the tyrosine-deprived condition compared with the normal condition, we identified YARS1 as the strongest negatively hit gene in the tyrosine-deprived group throughout the time course (Fig. 4b, c). This result indicated that loss of YARS1 was synthetically lethal when combined with tyrosine restriction on the one hand, and again validated our previous hypothesis that intracellular tyrosine in HCC was preferentially shunted in the direction of protein synthesis rather than metabolised as a ketogenic and glycogenic amino acid on the other hand. Specifically, we noted that sgRNAs targeting SLC7A5 were decreased after tyrosine restriction. In contrast, sgRNAs targeting tyrosine metabolism genes such as TAT and HGD were enriched, confirming the necessity of maintaining intracellular tyrosine levels in HCC cells (Fig. 4b, c).
a Schematic plot summarising the workflow of genome-wide CRISPR knockout screening in SNU398 cells cultured in different levels of tyrosine. b Gene rank of selection results for SNU398 cells cultured in either a standard tyrosine medium or a tyrosine-deficient medium for 48 h (left) or 96 h (left) in CRISPR screens. Positive Δβ-score indicates resistor genes, negative Δβ-score indicates sensitiser genes. A portion of negative score genes (blue) and positive score genes (tyrosine metabolism genes and mitochondrial respiratory chain complex genes) (red) are highlighted. c Visualisation of sgRNA targeting the indicated genes (n = 3 independent experiments). d Venn diagram showing the overlap of upregulated genes and negatively selected genes in SNU398 cells cultured in tyrosine-deficient medium. e–h Cell viability curve of GPX4 inhibitor RSL3 (e), BCL2 inhibitor venetoclax (f), SLC7A11 inhibitor Erastin (g) and sorafenib (h) treatment of HCC cell lines in either tyrosine standard or tyrosine-restricted medium (n = 3 independent experiments). i–m Myc/Alb-cre mice fed a tyrosine-modified diet are treated with or without sorafenib or venetoclax for 4 weeks, starting at the age of 8 weeks. Representative image of livers treated by sorafenib (i) or venetoclax (k), the plot of the body weight excluding liver weight, liver to body weight ratio, tumour number and maximum tumour diameter of the tumour-bearing mice treated by sorafenib (j) or venetoclax (l) (n = 6 mice), and survival curve of Myc/Alb-cre mice are shown in (m) (n = 10 mice). Data are representative of three independent experiments (e–h). Next-Generation Sequencing (NGS). Source data are provided as a Source Data file. Data are presented as means ± SD, two-tailed Student’s t test (e–h), one-way ANOVA followed by multiple comparisons (j, l), two-sided log-rank test (m).
Moreover, our finding of a decrease in sgRNAs targeting ferroptosis suppressors such as GPX4 and SLC7A11 following tyrosine restriction corresponded to their upregulated protein expression, again suggesting that tumour cells increase the expression of ferroptosis suppressors to scavenge lipid peroxidation caused by tyrosine restriction (Fig. 4b, c). Of note, BCL2, which inhibits mitochondrial damage-induced apoptosis37, was also identified as the top negatively selected gene in our screen, further supporting our previous results that impaired function and structural abnormalities of mitochondria induced cell apoptosis upon tyrosine deprivation, thus depletion of BCL2 improved the therapeutic effects with tyrosine restriction (Fig. 4b, c). Surprisingly, depletion of multiple genes encoding mitochondrial respiratory chain complex proteins was beneficial for HCC cells to adapt to complex I deficiency under tyrosine-restricted conditions (Fig. 4b), which is consistent with previous observations that mitochondrial complex I dysfunction can be alleviated by deletion of pathway-intrinsic proteins5,38. Notably, a large-scale overlap was found between negatively selected genes and genes with elevated protein levels upon tyrosine limitation, further revealing a homoeostatic feedback loop in which these candidates, whose depletion impaired cell fitness in the tyrosine-deprived condition, were elevated to resist tyrosine restriction (Fig. 4d). Collectively, our results suggested that these positive and negative genetic interactions are crucial for understanding the mechanistic underpinnings of tyrosine dependency in HCC and again indicate that HCC cells require more tyrosine to maintain increased protein synthesis.
Furthermore, several of the most negatively selected genes, such as GPX4, SLC7A11, and BCL2, were selected to corroborate our loss-of-function screen. We found that knockdown of either GPX4, SLC7A11 or BCL2 could remarkably improve the efficacy of dietary tyrosine modulation and YARS1 depletion in decreasing HCC cell fitness (Supplementary Fig. 17a–f). Consistently, pharmacological inhibition of GPX4 and BCL2 by their inhibitors RSL3 and venetoclax could also effectively enhance the inhibition effect of tyrosine restriction and YARS1 depletion (Fig. 4e, f, and Supplementary Fig. 17g, h). We also tested Erastin, a small molecule inhibitor of SLC7A11, and found that it could improve the therapeutic effect of tyrosine deprivation and YARS1 depletion (Fig. 4g and Supplementary Fig. 17i). Importantly, Sorafenib, a multi-kinase inhibitor for the treatment of advanced HCC patients, also demonstrated an improved therapeutic effect with tyrosine restriction and YARS1 depletion, which was attributed to its treatment-induced ferroptosis39,40,41 (Fig. 4h and Supplementary Fig. 17j). Moreover, knockdown of GPX4 or BCL2 in YARS1-knockdown tumour cells showed an enhanced therapeutic effect, characterising by a reduced tumour volume (Supplementary Fig. 18a, b), thereby validating that restricting tyrosine translation availability combined with ferroptosis-induced or apoptosis-induced models hold excellent therapeutic efficacy in reducing hepatocellular carcinogenesis. Furthermore, to confirm that restricting tyrosine translation availability suppresses tumorigenesis through inducing ferroptosis, rescue experiments were conducted both in vitro and in vivo. Treatment with ferroptosis inhibitor (Fer-1) significantly reversed the reduction of cell proliferation caused by tyrosine deprivation or YARS1-knockdown (Supplementary Fig. 18c, d), and mitigated YARS1-knockdown-induced tumour regression (Supplementary Fig. 18e–g). These findings collectively confirm that restricting tyrosine translation availability triggers tumour cell ferroptosis. Meanwhile, we found that depletion of NDUFB8 or SCD1 reduced the proliferation of HCC cells and inhibited tumour development (Supplementary Fig. 18h–k), thereby indicating that restricting tyrosine translation availability reduced HCC tumorigenesis and progression by decreasing the expression of NDUFB8 or SCD1. As expected, our in vivo HCC mouse model studies showed that both tyrosine restriction and YARS1 depletion combined with sorafenib or venetoclax showed an enhanced therapeutic effect, characterised by a reduced tumour burden and tumour diameter and extended survival time in Myc/Alb-cre mice (Fig. 4i–m, and Supplementary Fig. 18l–n). Thus, our results suggested that dietary tyrosine modulation and YARS1 depletion combined with targeted therapies (e.g., sorafenib and venetoclax) hold excellent therapeutic efficacy in reducing hepatocellular carcinogenesis.
Moreover, to establish the direct causality between MYC-YARS1-NDUFB8 or SCD1, we first examined ETC complex function and lipid peroxidation upon MYC knockdown. We found that MYC depletion reduced the expression of YARS1, NDUFB8, and SCD1 (Supplementary Fig. 19a), thereby inhibiting mitochondrial complex I enzyme activity (Supplementary Fig. 19b) and promoting lipid peroxidation (Supplementary Fig. 19c, d). Subsequently, we investigated whether YARS1 overexpression could rescue the defective effects in MYC-knockdown cells. Results demonstrated that YARS1 overexpression effectively improved the impaired mitochondrial complex I function induced by MYC deficiency (Supplementary Fig. 19e), while alleviating MYC knockdown-induced elevation in lipid peroxidation (Supplementary Fig. 19 f, g). Meanwhile, the lipid peroxidation levels were partially restored by re-expressing SCD1 in the YARS1-knockdown cells (Supplementary Fig. 19h, i) and the downregulation of complex I activity was reversed by expressing NDUFB8 back into YARS1-knockdown cells (Supplementary Fig. 19j). Similarly, we demonstrated that supplementation with oleic acid and palmitoleic acid (hereafter referred to as MUFA) effectively alleviated ferroptosis triggered by tyrosine restriction, YARS1-knockdown, or MYC knockdown, as evidenced by decreased lipid peroxidation levels (Supplementary Fig. 19k–m). Furthermore, SCD1 knockdown also resulted in increased lipid peroxidation (Supplementary Fig. 19n). Collectively, these findings indicate that MYC facilitates the translation of NDUFB8 and SCD1 by enhancing YARS1 transcription, thereby promoting mitochondrial complex assembly and ferroptosis resistance.
Tyrosine ammonia-lyase (TAL) exhibits potent therapeutic efficacy by degrading tyrosine in HCC
A long-term tyrosine-restricted diet in patients with tyrosinemia represents the feasibility of this dietary intervention in HCC patients42. Nevertheless, this restrictive diet poses challenges in terms of patient adherence. Thus, an approach superior to dietary intervention is to catalyse amino acids into non-toxic, readily metabolisable products through the action of an enzyme43. Clinically, L-asparaginase is widely used in the treatment of patients with acute lymphoblastic leukaemia due to its aminohydrolase function that catalyses the degradation of asparagine to ammonia and aspartic acid, thereby reducing the protein synthesis availability of asparagine44. Thus, phenylalanine/tyrosine ammonia-lyases (PAL/TALs) that convert the non-oxidative deamination of phenylalanine into cinnamic acid or tyrosine into coumaric acid (Fig. 5a), respectively, represent a potential therapeutic strategy for HCC45. Considering these enzymes with dual activity of catalysing phenylalanine/tyrosine, TAL derived from Flavobacterium johnsoniae, with the lowest catalytic activity for phenylalanine, was chosen to reduce the interference and consequence of phenylalanine degradation46. The dissociation constants (Kd) for FjTAL-tyrosine (5.88 μM) and FjTAL-phenylalanine (no binding), along with enzyme catalytic activity assay, further confirmed that FjTAL efficiently degraded tyrosine, with a minimal effect on phenylalanine (Fig. 5b and Supplementary Fig. 20a, b).
a A diagram depicting the reaction mechanism of tyrosine and phenylalanine degradation by TAL/PAL. b Purified FjTAL protein directly binds tyrosine but does not bind to phenylalanine, in ITC binding affinity analysis. c IC50 curve of FjTAL in HCC cell lines. (n = 3 independent experiments). d The activities of mitochondrial complex I-V of SNU398 cells upon FjTAL treatment. (n = 3 independent experiments). e OCR and maximal respiration of SNU398 cells with FjTAL treatment. (n = 4 independent experiments). f, g BODIPY493/503 staining (f) in HCC cell lines treated with FjTAL or vehicle and quantification of lipid peroxidation (g). (n = 3 independent experiments). h–k Graphic illustration (h), representative images of livers (i), the plots of the body weight excluding liver weight, liver to body weight ratio, tumour number and maximum tumour diameter (j) (n = 4 mice), and survival curves (k) (n = 6 mice) of Myc/Alb-cre mice with the indicated treatment are shown. l Native-PAGE analysis (left) and SDS-PAGE analysis (right) of livers from Myc/Alb-cre mice treated with FjTAL or vehicle. (n = 4 mice). m Representative TEM images of livers from Myc/Alb-cre mice treated with FjTAL or vehicle (left). Morphometric analysis of maximal cristae width of cells in 15 randomly selected mitochondria (right). n, o Representative Ki-67, TUNEL and 4-HNE staining of liver sections from Myc/Alb-cre mice treated with FjTAL or vehicle (n). Quantification of staining intensities and positive cells is shown (o). (n = 4 mice). l The samples are derived from the same experiment, and all gels/blots are processed in parallel under identical conditions. Data are representative of three independent experiments (c–g). Source data are provided as a Source Data file. Data are presented as means ± SD, two-tailed Student’s t test (d, e, g, m, o), one-way ANOVA followed by multiple comparisons (j), two-sided log-rank test (k).
Subsequently, we found that FjTAL treatment significantly inhibited the cell growth/survival in multiple HCC cells, as evaluated by half-maximal inhibitory concentration (IC50) values around 0.4–0.7 μM (Fig. 5c). Similar to tyrosine restriction and YARS1 depletion, FjTAL treatment could effectively reduce cell proliferation, induce cell apoptosis, and synergistically increase the anti-tumour efficacy of sorafenib or venetoclax in HCC cells (Supplementary Fig. 20c–n). As expected, FjTAL treatment also reduced tyrosine translation bioavailability, thereby decreasing the translation of SCD1 and NDUFB8 (Supplementary Fig. 21a–d). Defective mitochondrial function was further validated upon FjTAL treatment, characterised by reduced complex I formation and enzyme catalytic activity, downregulated OCR, and increased production of Mito-ROS (Fig. 5d, e, and Supplementary Fig. 21e, f). Similarly, FjTAL treatment also induced the formation of lipid droplets and increased lipid peroxidation in HCC cells (Fig. 5f, g, and Supplementary Fig. 21g–j), and Fer-1 treatment reduced the levels of lipid oxidation induced by TAL treatment (Supplementary Fig. 21k, l). In line with this, electron microscopy-based morphometric analysis also showed mitochondria morphology and structural abnormalities following FjTAL treatment (Supplementary Fig. 21m, n).
To further consolidate the anti-tumour efficacy of FjTAL in vivo, we administered a single intraperitoneal injection of different doses of FjTAL to tumour-bearing Myc/Alb-cre mice. The amino acid profile of FjTAL-treated liver tissue at a dosage of 50 mg/kg revealed the most pronounced and the longest-lasting reduction in tyrosine (Supplementary Fig. 21o). In contrast, the other amino acids showed little change after FjTAL treatment (Supplementary Fig. 21o). Consistently, after we treated tumour-bearing Myc/Alb-cre mice three times a week for 1 week with low-dose FjTAL (10 mg/kg), medium-dose FjTAL (25 mg/kg), or high-dose FjTAL (50 mg/kg), only the high-dose treatment group revealed reduced levels of tRNA-TyrGUA charging and an increase in the nuclear translocation of YARS1, indicating a significant inhibition of tyrosine translation availability (Supplementary Fig. 21p, q). Of note, our in vivo mouse HCC model studies also showed that while FjTAL treatment alone could remarkably reduce the weight ratio of liver/body, tumour burden, and tumour diameter, and prolong survival time in Myc/Alb-cre mice, its combination with sorafenib or venetoclax displayed an improved therapeutic efficacy (Fig. 5h–k). Consistent with the in vitro observation, FjTAL-treated tumours showed a defective mitochondrial function and severe ferroptosis phenotype, characterised by reduced complex I formation and enzyme activity, decreased expression of SCD1, and impaired morphology and structure of mitochondria (Fig. 5l, m, and Supplementary Fig. 21r). Finally, histopathological examination revealed reduced cell proliferation, increased cell apoptosis and lipid peroxidation, preferentially in FjTAL-treated tumours, indicating that therapeutic degradation of tyrosine by FjTAL treatment can antagonise hepatocellular carcinogenesis through impairing oxidative phosphorylation and inducing ferroptosis (Fig. 5n, o).
YARS1-mediated tyrosine aminoacylation in liver tumorigenesis can be inhibited by tyrosinol
In addition to limiting tyrosine uptake or pharmacologically degrading tyrosine, we propose an additional strategy to block the translational utilisation of tyrosine, i.e., the use of the structural analogue tyrosinol to competitively occupy the pocket of tyrosine-binding YARS126,47, thereby reducing YARS1-catalysed tyrosine binding to tRNA (Fig. 6a). The Kd for tyrosine-YARS1 and tyrosinol-YARS1 reached 22.6 μM and 40.2 μM, respectively, indicating a similar affinity to YARS1 (Fig. 6b). We also found that tyrosinol could reduce the binding capacity of YARS1 to tyrosine (Supplementary Fig. 22a). Moreover, the amino acid profiling in liver tissues from Myc/Alb-cre mouse treated with tyrosinol demonstrated that tyrosinol treatment had minimal effects on amino acid levels compared to the control group (Supplementary Fig. 22b). Thus, chemical inhibition of YARS1 by tyrosinol reduced the fitness of HCC cells and synergistically improved the therapy efficacy of sorafenib or venetoclax in HCC cells (Supplementary Fig. 22c–n). Consistently, tyrosinol treatment prevented tyrosine aminoacylation, supported by a reduced tRNA-TyrGUA charging and increased YARS1 nuclear translocation (Supplementary Fig. 22o, p), thereby inhibiting the translation of NDUFB8 and SCD1 (Fig. 6c, d). As a functional consequence of the reduced translation of NDUFB8 and SCD1, we also observed defective mitochondrial function and lipid peroxidation, leading to mitochondrial structural and morphological abnormalities (Fig. 6e–g and Supplementary Fig. 23a–f), and Fer-1 treatment reduced the levels of lipid oxidation induced by tyrosinol treatment (Supplementary Fig. 23g, h).
a The chemical structure of tyrosine and tyrosinol. b ITC assay of YARS1 binding to tyrosine and tyrosinol. c Relative luciferase activities in SNU398 cells with the indicated treatments. (n = 3 independent experiments). d RT-qPCR after ribosomal immunoprecipitation in SNU398 cells transfected with Flag-RPL22 and treated with tyrosinol. (n = 3 independent experiments). e The activities of mitochondrial complex I-V of SNU398 cells treated with tyrosinol or vehicle. (n = 3 independent experiments). f OCR (left) and maximal respiration (right) of SNU398 cells with tyrosinol treatment. (n = 3 independent experiments). g BODIPY493/503 staining in HCC cell lines treated with tyrosinol or vehicle (left). Quantification of lipid peroxidation (right). (n = 3 independent experiments). h–j Representative images of livers (h), the plots of the body weight excluding liver weight, liver to body weight ratio, tumour number and maximum tumour diameter (i) (n = 6 mice), and survival curves (j) (n = 10 mice) of Myc/Alb-cre mice with the indicated treatment are shown. k The activities of mitochondrial complex I-V of livers from Myc/Alb-cre mice treated with tyrosinol or vehicle. (n = 6 mice). l Representative electron micrographs of livers from Myc/Alb-cre mice treated with tyrosinol or vehicle (left). Morphometric analysis of maximal cristae width of cells in 15 randomly selected mitochondria (right). m, n Representative Ki-67, TUNEL and 4-HNE staining of liver sections from Myc/Alb-cre mice treated with tyrosinol or vehicle (m). Quantification of staining intensities and positive cells is shown (n). (n = 6 mice). Source data are provided as a Source Data file. Data are representative of three independent experiments (c–g). Data are presented as means ± SD, two-tailed Student’s t-test (c–g, k, l, n), one-way ANOVA followed by multiple comparisons (i), two-sided log-rank test (j).
Subsequently, we attempted to define whether the results obtained from human cancer cells could be translated to mouse tumours. Pharmacokinetic studies in mice revealed that when given orally at a dosage of 500 mg/kg, the duration of tyrosinol in the liver was approximately 24 hours (Supplementary Fig. 24a). We chose the tyrosinol dose (500 mg/kg/d) based on a pilot study of different doses of tyrosinol (250 or 500 mg/kg/d), which showed a reduced tyrosine aminoacylation and increased YARS1 nuclear translocation at the dosage of 500 mg/kg per day (Supplementary Fig. 24b–d). Moreover, to evaluate the safety profile of tyrosinol in mice, we found that tyrosinol treatment (repeated dosing of 500 mg/kg/d for 4 weeks, oral gavage) did not result in any significant tissue weight changes (Supplementary Fig. 24e). The levels of ALT, AST, CREA, CK, and LDH were also unchanged upon tyrosinol treatment (Supplementary Fig. 24f). Furthermore, no apparent pathological differences or tissue injury were observed after tyrosinol treatment (Supplementary Fig. 24g). Additionally, we examined the impact on the aminoacylation of the other two aromatic amino acids (phenylalanine and tryptophan) after tyrosinol treatment. Results demonstrated that the charged tRNA-PheGAA and tRNA-TrpCCA levels remained unchanged upon tyrosinol treatment (Supplementary Fig. 24h), indicating that tyrosinol treatment does not have potential off-target effects in vivo. As observed in vitro, tyrosinol treatment alone could significantly reduce liver-to-body-weight ratio, tumour burdens and tumour diameters and prolong the survival time in the tumour-bearing mice, while its combination with sorafenib or venetoclax showed an improved anti-tumour efficacy (Fig. 6h–j). Similarly, defective mitochondrial function and severe ferroptosis phenotype preferentially occurred in tyrosinol-treated tumours (Fig. 6k–n and Supplementary Fig. 24i, j). Thus, our findings showed that suppressing YARS1 with tyrosinol, both alone and in combination with sorafenib or venetoclax, uncovers attractive therapeutic options for treating HCC patients.
Discussion
Rewiring of amino acid metabolism has been identified as a metabolic adaptation during tumorigenesis and growth in response to a shift in amino acid utilisation pathways or to compensate for increased amino acid requirements in protein synthesis. Particularly, increased valine tRNA biosynthesis driven by oncogenic NOTCH1 signalling in leukaemia confers an enlarged requirement for valine in protein synthesis5. Moreover, given the increased demand for protein synthesis in glioblastoma (GBM), cancer stem cells in GBM reprogramme threonine metabolism to facilitate threonine-mediated t6A biosynthesis, thereby improving codon-dependent translational reprogramming of cell cycle-related proteins6. In line with these findings, we find that tyrosine levels are decreased in HCC, despite upregulation of tyrosine import and reduction of tyrosine metabolism, due to increased demand for tyrosine in protein synthesis, represented by MYC-driven upregulation of YARS1 and tyrosine tRNA biogenesis. Increased availability of tyrosine for translation can promote mitochondrial bioenergetics and the conversion of long-chain saturated fatty acids to MUFA by improving the expression of ETC complex I and SCD1, respectively, thereby enhancing OXPHOS and escaping ferroptosis (Supplementary Fig. 25a).
Based on our findings, targeting the vulnerability of tyrosine utilisation in HCC by restricting the availability of tyrosine for translation holds great promise for application in HCC patient treatment. First, amino acid-restricted diets are currently receiving widespread attention in oncology treatment due to the restriction of one or at most two amino acids, and a recent clinical trial has found that restricting serine and glycine intake in patients with colorectal cancer effectively inhibits tumour growth and enhances anti-tumour immunity by rewiring T lymphocytes48. Thus, building on the results of our study and the long-term use of tyrosine-restricted diets in tyrosinemia patients, we are recruiting patients to test the implementation of tyrosine-restricted diets in the clinical treatment of HCC patients. Secondly, the successful clinical application of L-asparaginase in acute lymphoblastic leukemia and of phenylalanine/tyrosine ammonia lyase (PAL/TAL) derived from Anabaena variabilis for the treatment of the metabolic disorder phenylketonuria (PKU) provides support for the future translation of TAL therapy in HCC patients49. In addition, an engineered bacterium, EcN-HT, can significantly metabolise tyrosine into levodopa (L-DOPA), thereby protecting mice from lethal liver injury caused by tyrosinemia type 1, which may represent an exploitable strategy for engineered probiotics to treat HCC by degrading tyrosine50. Moreover, the current findings highlight tyrosinol as an inhibitor of tyrosine aminoacylation, indicating a means to reduce the availability of tyrosine for translation, thus reducing hepatocellular carcinogenesis in mice without causing weight loss. However, given the high dosage of tyrosinol in targeting suppression of YARS1, further exploration of pathways to inhibit YARS1 function by optimising its structure or finding more degraders (e.g., PROTACs) is needed in future clinical applications.
However, the feasibility of applying tyrosine-restricted diets in human HCC patients still requires broader investigation. First, tyrosine-restricted diets in healthy WT mice result in the accumulation of lipid droplets (Supplementary Fig. 3), consistent with previous findings that a high-fat diet combined with methionine and tyrosine deficiency induces non-alcoholic fatty liver disease17. Second, in our short-term tyrosine-deficient dietary intervention in healthy WT mice, no depression-like behaviours or hair colour changes were observed, despite tyrosine serving as a precursor to both dopamine and melanin. However, it is noted that an 8-month tyrosine-deficient diet in our DEN-induced HCC model results in some hair appearing greyish white. Notably, we observe that tyrosine-restricted diets exhibit stronger anti-proliferative effects on tumour cells than on normal cells, attributable to the higher protein synthesis activity in malignant cells (particularly in tumours) (Supplementary Fig. 8). Meanwhile, another study finds that tyrosine restriction extends lifespan in Drosophila by activating FOXO pathway, demonstrating that the multifaceted impacts of dietary tyrosine restriction on physiological functions are in a context-dependent manner51. Therefore, clinical translation of tyrosine-deficient diets for HCC patients requires consideration of the impact on hepatic function and optimisation of the intervention time to leverage potential anti-tumour effects while minimising adverse outcomes.
Results from a genome-wide loss-of-function CRISPR screen revealed that loss of multiple targets, including GPX4, BCL2, and SLC7A11, can enhance the therapeutic effect of tyrosine restriction, indicating that tyrosine restriction in combination with targeted therapy can achieve a high therapeutic efficacy. However, the synergistic therapeutic effects between the ablation of these targets and RTTA treatment require future validation. Notably, consistent with previous findings that in MYC-driven mouse breast cancer, a low-protein diet induces cell competition between tumour-associated macrophages (TAMs) and tumour cells, thereby activating the tumour suppressor function of TAMs52, we also observed that after tyrosine restriction, TAMs surround the dying tumour cells to perform their scavenger function, manifesting an immune-activated microenvironment (Supplementary Fig. 25b). Thus, combining tyrosine restriction with immunotherapy (especially targeting TAMs and perhaps also targeting lymphocytes) might also represent a promising therapeutic strategy for patients with hepatocellular carcinoma to achieve higher efficacy and lower treatment-related toxicity.
In sum, exploiting this competitive utilisation pathway between translation and metabolism of tyrosine, both alone and in combination with other therapies, offers therapeutic options for treating patients with HCC.
Methods
Patient tissues and ethics
Informed consent was obtained from all patients in this study. No compensation was provided. Due to the limited samples available, sex and gender were not considered in the study design. This study was conducted following both the Declarations of Helsinki and Istanbul and received approval from the Ethics Committee of the Fifth Medical Centre of the Chinese PLA General Hospital. HCC tissues and adjacent liver tissues were sourced from the Fifth Medical Centre of the Chinese PLA General Hospital. The patient data were supplied in Supplementary data 6. HCC data from TCGA, CPTAC, and CNCB (the China National Centre for Bioinformation) were used for analysis.
Animals and ethics statement
All animal experiments were approved by the Animal Ethics Committees of Inner Mongolia Medical University and Beijing University of Chemical Technology and performed in accordance with the guidelines of the National Institutes of Health (NIH). The maximal tumour size permitted by the ethics committee was 2500 mm3, and the maximal tumour size was not exceeded in this study. Male C57BL6/J and 4-week-old male BALB/c nude mice were provided by Beijing Vital River Laboratory Animal Technology Co., Ltd (China). Alb-cre and H11LNL-Myc (H11-CAG-loxp-STOP-loxp-MYC) strains, based on C57BL/6 J background, were purchased from Shanghai Model Organisms Centre, Inc. Myc/Alb-cre mice were generated by crossing H11LNL-Myc and Alb-cre mice. Animals were housed in individually ventilated cages (IVC) within a specific pathogen-free (SPF) grade system at 22 °C with a 12-hour light/dark cycle and free access to water and food. The animals used in the experiments were randomly assigned to groups. In the subcutaneous tumour model, tumour growth was monitored at regular intervals, and mice were euthanized when tumour volume approached 2500 mm3, mice lost 20% of body weight, or tumours exhibited ulceration. In vivo experiments, including spontaneous and orthotopic liver tumour models, were terminated at predefined endpoints when (a) tumours exceeded 2 cm in diameter, (b) mice lost 20% of body weight, or (c) mice showed signs of severe distress such as lethargy or impaired mobility. Survival time was recorded in all experiments.
Animal experiments
Male mice only were used in the different models to avoid the influence of sex and oestrogen. For the HCC mouse model, Myc/Alb-cre transgenic mice spontaneously develop HCC within 12 weeks of age. The orthotopic HCC model induced by diethylnitrosamine (DEN) (Sigma, Cat# N0676) treatment was established following the protocol described in previous reports21. Briefly, 14-day-old male mice received an intraperitoneal injection of DEN at a dosage of 25 mg/kg. Regarding the intrahepatic injection model of human HCC cells, 4-week-old male nude mice were anaesthetised, and their livers were placed ex vivo through a subcostal incision. Then, 50 μL of PBS containing 1 × 106 cells was injected into both the left and right lobes of the liver. After bleeding from the left and right lobe wounds was stopped with sterile cotton swabs, the livers were placed back into the abdominal cavity, and the incisions were sutured. Animals were sacrificed after the indicated time points and treatments. Then, livers were photographed and weighed. Tumour numbers were counted by visual assessment, and tumour diameters were measured using calliper. Liver tissues were stored at −80 °C after being frozen in liquid nitrogen for further analysis.
For the tyrosine-modified diet experiments, mice were fed different tyrosine-modified diets for the duration shown in the articles. Special diets with varying levels of tyrosine were purchased from Xie Tong Bio-engineering (Yangzhou) Co., Ltd (China). The diet of tyrosine 100% corresponds to the concentration of tyrosine (0.4%, wt/wt) contained in the standard diet (Cat. No. A100021B). The tyrosine-restricted diet (tyrosine 0% diet) contained 0% tyrosine, and the tyrosine-supplemented diet (tyrosine 500% diet) contained 2% tyrosine. Differences in protein/nitrogen content were modulated by adjusting the proportional concentration of glutamate.
For AAV administration, AAV particles including AAV8-shNC, AAV8-shYARS1-1, and AAV8-shYARS1-2 were purchased from GenePharma (China). AAV particles were injected into four-week-old Myc/Alb-cre mice by tail-vein injection at 1 × 1012 genome copies (GC) in PBS per mouse. shRNA sequences were listed in Supplementary data 1.
For FjTAL treatment, eight-week-old Myc/Alb-cre mice were administered a single intraperitoneal injection of FjTAL at different dosages to evaluate the efficacy of the FjTAL treatment. At the indicated time points, liver tissues were collected for HPLC-MS analysis to examine the degradation efficacy of amino acids. In other FjTAL-treated experiments, mice received FjTAL by intraperitoneal injection every 2 days until sacrificed.
For tyrosinol (Sigma, Cat# 469998) treatment, tyrosinol was dissolved in PBS, and mice received tyrosinol by oral gavage at a concentration of 500 mg/kg per day.
For Sorafenib or Venetoclax treatment, drugs were dissolved in vehicle (5% (v/v) DMSO, 40% (v/v) PEG300, 5% (v/v) Tween-80 in 0.9% (w/v) NaCl). Each mouse received oral gavage at 30 mg/kg per day.
Cell lines and cell culture
Human HCC cell lines that underwent short-tandem repeat testing and were confirmed free of mycoplasma contamination, including SNU398, SNU449, PLCPRF5, and HCCLM3, were obtained from Bena Culture Collection (China). A stable YARS1-knockdown HCC cell line was generated using a lentiviral vector. All cell lines were cultured in high-glucose containing DMEM medium supplemented with 10% FBS (Vazyme, Cat# F101-01) and 1% streptomycin–penicillin (Viva Cell, Cat# C3420-0100). For experiments involving the restriction of tyrosine, cells were cultured in a high-glucose DMEM medium that only lacked tyrosine (tyrosine 0% compared to standard medium). Primary mouse hepatocytes from 6-to-8-week-old mice were isolated and cultured, as previously reported53. EBSS and type Ⅳ collagenase (Sigma, Cat# C5138) were briefly perfused through the portal vein to digest liver tissue. Hepatocytes were isolated using Percoll buffer (Sigma, Cat# P1644) and cultured in DMEM medium. Cells were cultured at 37 °C with 5% CO2.
Plasmids
The promoter sequence of YARS1 was cloned into the pGL4.10 plasmid to generate the luciferase reporter vector YARS1-WT-Luc. A mutant construct of the luciferase reporter YARS1-mutant-Luc was generated as previously reported21. The pLKO.1-shYARS1 vector was homemade, and shRNA sequences were listed in Supplementary Data 1.
Transfection, lentiviral transduction, and generation of stable cell lines
For siRNA or shRNA-mediated knockdown, cells were transfected with 50 nM siRNAs or 1.5 μg shRNA-targeting plasmids using Lipofectamine 2000 (Invitrogen, Cat# 11668500), both from GenePharma (China), in Opti-MEM, according to the manufacturer’s instructions. For lentivirus packaging, HEK293T cells were co-transfected with shRNA-targeting vectors and the helper vectors psPAX2 and pCMV-VSVG using Lipofectamine 3000 (Invitrogen, Cat# L3000150). The cell supernatants containing lentiviruses were collected and concentrated 48 h after transfection. Low-density (20%) HCCLM3 cells were infected with the concentrated supernatant in the presence of 5 μg/mL polybrene (Beyotime, Cat# C0351) for 24 hours. Infected cells were placed under puromycin selection at a 1 mg/mL concentration for 1 week to obtain stable cell lines.
Drug treatment
For drug treatment, (1S,3 R)-RSL3 (MCE, Cat# HY-100218A), Erastin (MCE, Cat# HY-15763), Sorafenib (MCE, Cat# HY-10201), and Venetoclax (MCE, Cat# HY-15531) were used to improve the anti-tumour efficacy with restriction of tyrosine translation availability. Drugs were dissolved in DMSO. Cells were treated with drugs as indicated in the figures for 48 h in combination with either tyrosine-restricted medium, shRNA-mediated knockdown, FjTAL-treated medium, or medium containing tyrosinol.
RNA extraction, cDNA synthesis, and RT-qPCR
Total RNA extraction was performed using TRIZOL (Invitrogen, Cat# 15596018CN) as previously reported21. cDNA was synthesised via reverse transcription using 3 μg of RNA as a template with the HiScript III Reverse Transcription Kit (Vazyme, Cat# R323). Real-time PCR was performed with ChamQ SYBR qPCR Master Mix (Vazyme, Cat# Q412) using an ABI 7500 Sequence Detection System. Primer sequences for quantitative PCR analysis are listed in Supplementary Data 1.
tRNA qPCR
Total RNA was extracted using TRIZOL, with integrity and purity confirmed via the A260/A280 ratio. Total tRNA quantification was performed using a tRNA quantification kit (Arraystar, AS-FS-004). Briefly, purified RNA was treated with demethylase AlkB to remove methylation. Following RNA purification via chloroform extraction, purified RNA served as a template for reverse transcription using the MLV reverse transcriptase kit (Invitrogen, Cat# 28025013) according to the manufacturer’s protocols. tRNA quantification was then performed by RT-qPCR according to the manufacturer’s specifications. Primer sequences for tRNA quantification were provided in Supplementary Data 1.
Northern blot for detecting tRNA
The tRNA Northern blot was performed using a Northern blot kit (Beyotime, Cat# R0220), according to the manufacturer’s protocol. A total of 500 ng of trizol-extracted RNA was mixed with 2× RNA native loading buffer (Beyotime, Cat# R0221), heated at 65 °C for 15 minutes, and then loaded into a 15% urea polyacrylamide gel (Beyotime, Cat# R0232S). The gel was electrophoresed in TBE buffer at 200 V for 40 minutes. The RNAs were transferred to a nylon membrane at 60 V for 1 hour and then crosslinked with the membrane under UV radiation at 120 mJ/cm2 for 30–45 s. The membrane was pre-hybridised with hybridising solution for 2 hours, then a 10 pmol/mL RNA probe labelled with biotin was added for an additional 2 hours of hybridisation. Following hybridisation, the membrane was washed twice with low-salt buffer and twice with high-salt buffer, then blocked with blocking buffer at 37 °C for 15 minutes. The membrane was incubated with a blocking buffer containing Streptavidin-HRP conjugate at a 1:2000 dilution for 15 minutes, followed by two washes with a balanced buffer. Detection was achieved using the ECL detection solution according to the manufacturer’s guidance. All probe sequences were listed in Supplementary Data 1.
Northern tRNA charging assay
The tRNA aminoacylation assay was performed as previously reported54. Cells were resuspended in 0.3 M sodium acetate buffer (pH 4.7) containing 10 mM EDTA, mixed with phenol-chloroform-isoamyl alcohol (pH 4.7), vortexed, and centrifuged at 22,000 × g for 25 min at 4 °C. RNA was precipitated with 100% ethanol, resuspended in 0.3 M acetate buffer (pH 4.5), precipitated with 100% ethanol again and dried. RNA was redissolved in acetate buffer (pH 4.5), mixed with 2×RNA loading buffer (pH 4.5), loaded onto a 15% urea polyacrylamide gel (pH 4.5), and then subjected to electrophoresis at 40 V for 6 h in 0.3 M acetate buffer (pH 4.5). Membrane transfer proceeded in the buffer containing 10 mM Tris-acetate (pH 7.8), 5 mM sodium acetate and 0.5 mM EDTA at 40 V for 2 h. Subsequent steps followed the Northern blot procedure.
Immunoblot analysis
Tissues were homogenised in lysis buffer (50 mM Tris, pH 7.8, 0.5 M NaCl, 2 mM EDTA, 2 mM EGTA, 1% Triton X-100, 0.25% Deoxycholate, 1 mM NaF and 1 mM Na3VO4) supplemented with protease inhibitors (Roche, Cat# 46931320001) using a homogeniser (IKA) and subsequently lysed for 30 min at 4 °C. Cells were lysed with 1 mL lysis buffer (50 mM Tris, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, 1 mM NaF, 1 mM Na3VO4) supplemented with protease inhibitors for 30 min at 4°C. After centrifugation at 13,000 × g at 4 °C for 10 min, protein concentration was quantified using the Bradford or BCA assay, depending on the lysis buffer used. For immunodetection, equal amounts of total protein lysate were boiled and resolved thereafter by SDS-PAGE and transferred to nitrocellulose membrane, blocked with milk, and incubated with indicated diluted primary antibodies. The signal was detected using Pierce™ ECL Western Blotting Substrate (Thermo Scientific, Cat# 32209). The antibodies used in this article were shown as follows: anti-SLC7A5 (Proteintech, Cat# 28670-1-AP), anti-SLC38A9 (Proteintech, Cat# 21871-1-AP), anti-SLC43A1 (Proteintech, Cat# 10444-1-AP), anti-TAT (Thermo fisher, Cat# PA5-26541), anti-HPD (Proteintech, Cat# 67552-1-Ig), anti-HGD (Proteintech, Cat# 16465-1-AP), anti-GSTZ1 (Proteintech, Cat# 14889-1-AP), anti-FAH (Proteintech, Cat# 14928-1-AP), anti-YARS1 (Thermo fisher, Cat# PA5-53883), OXPHOS antibody cocktail (anti-NDUFB8 (Proteintech, Cat# 14794-1-AP), anti-SDHB (Proteintech, Cat# 10620-1-AP), anti-UQCRC1 (Proteintech, Cat# 10620-1-AP), anti-MTCO2 (Proteintech, Cat# 55070-1-AP), anti-ATP5A1 (Proteintech, Cat# 14676-1-AP)), anti-SCD1 (Proteintech, Cat# 28678-1-AP), anti-4-HNE (JaICA, Cat# MHN-20P), anti- Histone H3 (CST, Cat# 4499) and anti-β-Actin (TransGen Biotech, Cat# HC201-01). Horseradish peroxidase-conjugated anti-rabbit IgG (LABLEAD, Cat# RN0100), anti-mouse IgG (LABLEAD, Cat# MN0100) was employed as secondary antibodies.
Blue native-PAGE
The BN-PAGE was performed to detect the assembly of mitochondrial respiratory complexes as previously reported55. Human cancer cells were collected, washed with cold PBS and resuspended in PBS containing digitonin (6 mg/mL) (Beyotime, Cat# ST1272) and protease inhibitors. After lysing the cells on ice for 15 min, 1 mL cold PBS was added to the samples, followed by centrifugation at 10,000 × g for 5 min at 4 °C to collect mitochondrial pellets. For liver tissues, the tissues were homogenised in MOPS buffer (440 mM sucrose, 20 mM MOPS, 1 mM EDTA, 0.2 mM PMSF and protease inhibitors) using a homogeniser and then lysed for 10 min on ice. After centrifugation at 10,000 × g for 10 min, enriched mitochondrial fractions were collected. Pellets were resuspended in 100 μL buffer (1 M aminocaproic acid, 50 mM Bis–Tris/ HCl, pH 7.0) supplemented with 20 μL dodecylmaltoside (10% w/v) and protease inhibitors, vortexed, and then incubated on ice for 5 minutes before the lysates were centrifuged at 20,000 × g for 20 min to collect solubilized mitochondrial complexes in the supernatant. Protein concentration was measured using the BCA assay. Then, equal amounts of lysates were mixed with 2× blue native-PAGE Sample Buffer (Beyotime, Cat# P0761) and loaded onto a 3–12% blue native precast gel (Beyotime, Cat# P0546) at 4 °C. Samples were electrophoresed in cathode buffer Ⅰ at 40 V for 4 h and subsequently in cathode buffer II at 40 V for 2 h until excess blue dye migrated out of the gels. Stripping buffer (100 mM 2-mercaptoethanol, 2% SDS, 62.5 mM Tris-HCl, pH 6.7) was used to wash the gel at 50 °C for 30 minutes to remove G250 and expose antigen epitopes, followed by transfer of proteins to nylon membrane. Subsequent steps followed Western Blot procedures.
Flow cytometric detection of cell apoptosis, cell cycle, lipid ROS, and mitochondrial ROS
Cell death was measured using the Annexin-V FITC kit (Beyotime, Cat# C0162). Cells were washed in PBS, resuspended in annexin-V binding buffer and incubated with staining buffer containing annexin-V-FITC and propidium iodide (PI) for 30 min at 4 °C. For cell cycle analysis, cells were washed in PBS, fixed with 70% ethanol, and incubated with staining buffer containing PI and RNase A for 30 min at room temperature. Mitochondrial reactive oxygen species (ROS) were assessed by staining cells with MitoSOX Red (MCE, Cat# HY-D1055). Lipid peroxidation was evaluated using BODIPY 581/591 C11 (Thermo Scientific, Cat# D3861) for cell staining. Flow cytometry was performed using a CytoFLEX (Beckman) instrument, and data were analysed with CyteExpert software (Beckman). Excitation/absorption wavelengths were as follows: annexin-V FITC (488 nm/525 nm, FITC channel, 488 nm laser, 525/40 BP), PI (488 nm/585 nm, PE channel, 488 nm laser, 585/42 BP), MitoSOX Red (510 nm/580 nm, PE channel, 488 nm laser, 585/42 BP), and BODIPY 581/591 C11 (500 nm/510 nm, FITC channel, 488 nm laser, 525/40 BP). Gating strategies for flow cytometry analysis were provided in Supplementary Fig. 26.
3H tyrosine uptake and tracing
For ex vivo uptake, normal liver freshly isolated from H11LNL-Myc mouse or tumour tissues freshly isolated from Myc/Alb-cre mouse were incubated in tyrosine-deficient DMEM medium containing 2 μCi L- [ring 3,5-3H]-Tyrosine (American Radiolabelled Chemicals, Inc. Cat# ART 0195) at 37 °C for 30 min. Tissues were washed thrice with cold PBS and subsequently lysed with SOLVABLE (PerkinElmer, Cat# 6NE9100). To assess tyrosine translation utilisation, normal liver or tumour tissues were incubated in tyrosine-deficient DMEM medium containing 10 μCi 3H-Tyrosine at 37 °C for 1 h. Tissues were washed thrice with cold PBS and subsequently homogenised in lysis buffer using a homogeniser. For ribosome extraction, ribosome pellets were isolated using a ribosome extraction kit (BestBio, Cat# BB-36061) according to the manufacturer’s protocol. For protein extraction, the lysate was centrifuged at 4 °C for 30 min, the supernatant was retained, and trichloroacetic acid was added to the supernatant to precipitate protein. Intracellular 3H tyrosine was quantified by using a scintillation counter (PerkinElmer).
13C tyrosine uptake and metabolic flux analysis
Primary mouse hepatocytes isolated from H11LNL-Mycmouse or Myc/Alb-cre mouse were cultured in tyrosine-deficient DMEM medium containing 250 μM L-Tyrosine (ring-13C6) (Cambridge Isotope Laboratories, Cat# CLM-1542-PK) at 37 °C for 12 h or 48 h to analyse in vitro uptake or metabolic flux of 13C tyrosine, respectively. Then, cells were collected and mixed with 80% methanol, sonicated for 1 min in an ice-water bath, followed by placing on dry ice for 1 min. Samples were stored at −80 °C for 30 min, thawed at 4 °C for 10 min, vortexed for 1 min and centrifuged at 16,000 × g at 4 °C for 10 min. The supernatants were gently dried with nitrogen. Then, samples were reconstituted with 50% ACN containing internal standards for testing by UHPLC-HRMS platform.
Ultimate 3000 Liquid Chromatography System equipped with a Waters Amide column (100 mm × 2.1 mm,1.7 μm). The injection volume was 5 μL. The flow rate was 0.3 mL per minute. The mobile phase composition consisted of (A) 10 mM ammonium acetate aqueous solution and (B) 10 mM ammonium acetate in 90% acetonitrile aqueous solution. A linear gradient elution mode was employed for the separation of metabolites, with the following gradient profile: Starting at 95% B and maintained for 0.5 minutes, linearly decreasing to 65% B over 7 minutes and held for 1 minute, further linearly decreasing to 50% B at 8 minutes and held for 1 minute, then linearly increasing to 95% B at 9.1 minutes and held for 2.9 minutes, returning to the initial gradient at 9.1 minutes and equilibrating until 12 minutes. The metabolites were ionised, and mass spectrometry data were collected using an Orbitrap Fusion Tribrid mass spectrometer. Negative ion mode ionisation (HESI-) of the metabolites was performed using a heated electrospray ionisation source. The main parameters of the ion source were set as follows: spray voltage at 2500 V, capillary temperature at 325 °C, and needle heater temperature at 300 °C. The sheath gas flow rate was 30 (arbitrary units, the same below), and the auxiliary gas flow rate was 10 (arbitrary units). The S-Lens RF was set to 60 (arbitrary units). The first-order mass spectrometry scan range was from mass-to-charge ratio (m/z) 70–800. Raw mass spectral data were preprocessed using Xcalibur software (version 4.0.27.19). Data were displayed by the ratio of labelled metabolites to all metabolites.
Amino acid profiling of liver tissues from HCC patients and mice
Amino acid profiling was carried out by targeted metabolomics. Snap-frozen liver samples with no significant difference in weight between groups were added into 80% methanol containing 0.1% formic acid and the internal standard succinic acid-2,2,3,3-d4, homogenised with two stainless steel beads using a grinder and sonicated for 10 min in an ice-water bath, followed by standing at −20 °C for 20 min. After centrifugation at 16,000 × g at 4 °C for 10 min, the supernatants were added into mixed internal standard and subsequently concentrated. Pyridine solution containing methoxamine hydrochloride was added to the samples, and the oximation reaction was carried out for 60 min at 37 °C in a shaking incubator. Then, BSTFA derivatization reagent, hexane and internal standards were added into the samples, and the reaction was carried out at 70 °C for 60 min before GC-MS analysis. GC-MS analysis was performed using a TSQ9000 triple-quadrupole mass spectrometer with MRM scan type.
Mitochondrial respiration
Oxygen consumption rates (OCRs) were measured to assess mitochondrial respiration using Seahorse XFe96 Analyzer (Agilent). Cells were plated in XFe96 Cell Culture Microplates (Agilent, Cat# 102416-100) and cultured in DMEM until confluence, after which the medium was switched to Seahorse XF Base Medium (Agilent, Cat# 102353-100) supplemented with glucose, L-glutamine, and sodium pyruvate for OCR measurement. OCR was measured beginning with basal (baseline), and then was treated with oligomycin (Abcam, Cat# ab141829), 4-(trifluoromethoxy) phenylhydrazone (FCCP) (Sigma, Cat# C2920), and rotenone (Sigma, Cat# R8875). The results were expressed in pmol/min and normalised to the cell number.
Expression and purification of FjTAL and YARS1
The coding sequence of FjTAL or YARS1 was cloned into the pET22b-(+) vector, and the recombinant vector was transformed into BL21 (DE3). The transformed strains were revived and streaked onto TB agar plates containing ampicillin to select for transformants. Selected strains were expanded in TB medium with ampicillin at 37 °C overnight. Isopropyl-β-D-thiogalactopyranoside (IPTG) with a concentration of 0.4 mM was added to induce protein expression of FjTAL for 22 h at 25 °C. Correspondingly, IPTG at 0.4 mM concentration induced the protein expression of YARS1 for 16 h at 18 °C. The cells were disrupted by ultrasonication, and the supernatant was collected and purified via a Ni - affinity column, followed by concentration to yield a protein concentrate. The concentrate was stored at −80 °C after identification through molecular sieving.
Pyrophosphate production assay
All enzymatic reactions were conducted in 50 mM HEPES (pH 7.5), 25 mM KCl, 2 mM MgCl₂, 4 mM ATP, 300 nM YARS1, protein substrates, tyrosine or tyrosinol at 37 °C for 1 h. Pyrophosphate (PPi) production was quantified using the fluorogenic PPi sensor from the PhosphoWorks™ Fluorimetric Pyrophosphate Assay Kit (Enhanced Selectivity) (AAT Bioquest, Cat#21614), according to the manufacturer’s instructions.
Whole-genome CRISPR screen and analysis
The Brunello library, comprising 76,441 sgRNAs targeting 19,114 genes, was used to perform genome-wide CRISPR screening56. Lentiviruses carrying Cas9 expression vectors were used to transduce the SNU398 cell line, establishing a stable SNU398 cell line expressing Cas9 protein (SNU398-Cas9). This SNU398-Cas9 cell line was then transduced with the Brunello library at a low MOI (0.3) for 24 hours, followed by culture in fresh DMEM supplemented with puromycin for 72 hours. After puromycin selection for 7 days, cells were harvested and divided into two groups, cultured in DMEM with or without tyrosine for either 2 days or 3 days. Genomic DNA was extracted and used as a template to amplify the sgRNA insertion region via PCR, guided by sgRNA barcode sequences, to generate the library. Following NGS deep sequencing on the Illumina platform, sgRNA abundance was quantified using Bowtie2, and essentiality β score per gene was calculated by using the MLE algorithm of MAGECK software. Candidate genes were identified and listed.
Cell viability and clone formation
Cell viability was tested using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay following the manufacturer’s instructions. MTT was added to the medium at a final concentration of 5 μg/mL for an incubation of 4 h, and the medium was replaced with DMSO. The absorbance of the solution at 570 nm was measured using a microplate reader. For the clonogenic growth assay, after being treated as indicated and cultured for a total of 12 days, cells were fixed with 4% formaldehyde, stained with crystal violet solution, photographed and quantified using Image J.
ChIP assay and Ch-IP datasets
Ch-IP assays were carried out using an EZ-CHIPTM Assay Kit (Sigma, Cat# 17-371) according to the manufacturer’s instructions. Briefly, SNU398 cells were crosslinked with 1% formaldehyde for 20 min at 37 °C, after which crosslinking reactions were quenched with 0.125 M glycine for 5 min at room temperature. The fixed cells were resuspended and lysed in SDS buffer (50 mM Tris-HCl, pH 8.0, 5 mM EDTA, 1% SDS, 100 mM NaCl) supplemented with protease inhibitors and broken by sonication to obtain DNA fragments of ~200–500 bp. The supernatant was then incubated with 4 μg anti-MYC antibody or IgG overnight at 4 °C followed by adding Protein G magnetic beads into the immune complexes with a 2-h co-incubation at room temperature. After washing, elution and de-crosslinking, immunoprecipitated DNA and input DNA (fraction reserved from the sonication step) were purified and used for PCR analysis. Data were presented as a percentage of input DNA for each condition. By mining the data from the Cistrome Data Browser datasets, the MYC-binding region of the YARS1 promoter was found, as indicated in the figures.
Luciferase reporter assay
SNU398 cells were seeded into 24-well plates. After 24 h, cells were transfected with 50 nM siRNAs or scramble siRNA control and co-transfected with the luciferase reporter and the luciferase internal control reporter phRL-TK. Cells were harvested for measurement of reporter activity using the Dual-Luciferase Reporter Assay System (Promega, Cat# E1910) 48 h post-transfection. The internal control Renilla luciferase activity was used to normalise the luciferase activity.
Reporter assay
Reporters expressing luciferase in fusion with either NDUFB8, SCD1 or NDUFB1, as indicated in the figures, were generated by gene synthesis from GeneWiz (China). SNU398 cells transfected with these reporter constructs were subjected to tyrosine restriction, FjTAL treatment, tyrosinol treatment, YRAS1 knockdown or tRNA-TyrGUA knockdown for 48 h. Then, cells were collected and lysed to measure the activity of luciferase and Renilla luciferase using the Dual-Luciferase Reporter Assay System according to standard protocols. The geometric means of luciferase activities were normalised to the internal control of Renilla luciferase activities.
Ribosome immunoprecipitation
Ribosome immunoprecipitation was performed based on previous reports and a Magna RIP ™ kit (Sigma-Aldrich, Cat# 17-371)57. Briefly, cells transfected with Flag-RPL22 plasmid were subjected to tyrosine translation utilisation restriction treatment. Then, cells were treated with CHX (100 μg/mL) for 30 min at 37 °C, followed by lysis in lysis buffer (150 mM KCl, 10 mM MgCl2, 1% NP-40, 0.5 mM DTT, 100 μg/mL CHX, 20 mM HEPES KOH (pH 7.3), protease inhibitors, and 10 μL/mL RNasin and Superasin) for 15 min on ice. After centrifugation at 20,000 × g for 10 min at 4 °C, the supernatants were incubated with Flag affinity gels (Sigma-Aldrich, Cat# A220) overnight at 4 °C and subsequently the immune complexes were washed extensively. Then, immunoprecipitated RNA and input RNA (fraction reserved from the incubation step) were eluted in elution buffer containing 10% SDS and proteinase K and purified using the RNA Clean and Concentrator columns Kit (Zymo Research Corp, Cat# R1017). Immunoprecipitated RNA was reverse-transcribed, followed by RT-qPCR to quantify gene enrichment. Data are presented as a percentage of the input RNA.
Immunohistochemistry and immunostaining assessment
For immunohistochemistry staining, the formalin-fixed paraffin-embedded liver samples from HCC patients and mice were cut into sections of 3 μm, deparaffinized, rehydrated in graded alcohols, heated at 95 °C for 30 minutes in 1 mM EDTA buffer (pH 8.0) to retrieve antigen and blocked endogenous peroxidases and non-specific binding sites using 4 % hydrogen peroxide and 5% milk, respectively. Primary antibody was incubated with the sections overnight at 4 °C, followed by washing the sections with PBS and adding secondary antibodies with a 30 min co-incubation at room temperature. The positive cells were visualised using DAB chromogen. One-step TUNEL apoptosis assay kit (Beyotime, Cat# C1088) was used to detect apoptotic cells in the liver sections from the mouse according to the manufacturer’s protocol. Briefly, after being deparaffinized, rehydrated in graded alcohols and washed in PBS, the sections were incubated with 20 μg/mL DNase-free proteinase K for 20 min at 37 °C. After an extensive washing step (four times PBS), the 50 μL detection solution for each sample, containing 5 μL TdT enzyme and 45 μL fluorescent labelling solution, was used in the reaction for 1 h at 37 °C in the dark. The nuclei were visualised with DAPI (Beyotime, Cat# P0131). Representative images were recorded using a Nikon fluorescence microscope system. For the immunoreactivity evaluation of specimens from the patient, an H-score method was used to precisely quantify the immunolabelled cells by experienced pathologists as previously described58. Briefly, staining intensity of cells was divided into four categories: no positive staining (scored 0), low staining intensity (scored 1), moderate staining intensity (scored 2) and strong staining intensity (scored 3). Immunoreactivity scores of patients were calculated according to the following formula: H-score = 100ⅹ (% cells with score 1 × 1 + %cells with score 2 × 2 + % cells with score 3 × 3), ranging from 0 to 300. For the immunoreactivity assessment of Ki-67 staining, the Ki-67-positive cells were quantified by computerised counts (Image-Pro Plus 6). The TUNEL or 4-HNE positive cells were visually counted by experienced pathologists.
Transmission electron microscopy
Cultured cells and freshly isolated liver tissues were fixed in 0.15 M sodium cacodylate buffer (pH 7.4) supplemented with 2.5% glutaraldehyde, and 2 mM CaCl2 for 2 h at 4 °C and post-fixed in 1% osmium tetroxide overnight at 4 °C and 1% potassium ferrocyanide for 12 h at room temperature. Then, samples were stained in 1% aqueous uranyl acetate for 1 hour at 4 °C in the dark, dehydrated in graded acetones and finally embedded in EMbed 812 (Electron Microscopy Sciences). Ultrathin sections (70 nm) were cut using a diamond knife (Diatome) in an ultramicrotome (Leica EM UC7), placed on copper grids and stained with uranyl acetate and lead citrate. Sections were photographed on a JEM1400 transmission electron microscope operated at 120 kV. Morphometric analysis of maximal cristae width was performed by using Image-pro plus 6 on at least 15 randomly selected mitochondria per group.
Pharmacokinetic studies
A single oral dose of tyrosinol was administered to 8-week-old male Myc/Alb-cre mice. Liver tissues were collected at the indicated time points, extracted with 50% methanol and shaken for 1 h before centrifuging at 16,000 × g for 10 min at 4 °C. LC-MS analysis was performed on the UPLC I-Class chromatography coupled to a XEVO TQ-S Micro triple-quadrupole mass spectrometer with SRM scan type. Target Lynx software was used for the quantitative analysis. Phoenix WinNonlin software (version 8.1) was used to calculate the pharmacokinetic parameters through non-compartmental pharmacokinetic analysis with the linear/log trapezoidal rule.
Oil red O staining assay
HCC cells were seeded in six-well plates, then treated as indicated in the figures. Staining was performed according to the instructions of the Oil Red O Stain Kit (Solarbio, Cat# G1262). Briefly, cells were washed with PBS three times and subsequently fixed in 4% paraformaldehyde for 20 min at room temperature. After being rinsed with 60% isopropanol for 10–20 s until the stroma was clear, cells were soaked in a newly prepared Oil Red O solution for 20 min before staining with Mayer hematoxylin solution for 1 min. Representative images were recorded using a Nikon fluorescence microscope system, and lipid droplet intensity was assessed with Image-pro plus 6 software.
Cellular fractionation
Cellular fractionation was performed as previously described21. In brief, cells were harvested, and 1/4 of the cells were removed as whole cell lysate (WCL). The remaining cells were lysed in lysis buffer containing 20 mM HEPES, pH 7.9, 10 mM KCl, 1.5 mM MgCl2, 0.5 mM dithiothreitol and protease inhibitors for 30 min on ice, and then NP-40 (0.3% final concentration) was added to the lysates. The lysates were swirled for 25 s and centrifuged at 200 × g for 10 min at 4 °C to separate the cytoplasmic fraction (Cytoplasm) and the nuclear pellet (Nucleus). The precipitates were resuspended with solution 1 (0.25 M sucrose, 10 mM MgCl2), then added onto solution 2 (0.88 M sucrose, 0.05 mM MgCl2) and centrifuged at 3500 × g for 10 min to obtain the purified nuclear fraction.
Isothermal titration calorimetry (ITC) binding assay
The binding affinity of FjTAL with tyrosine or phenylalanine and of YARS1 with tyrosine or tyrosinol was measured using a MicroCal PEAQ-ITC calorimeter. Briefly,
all proteins and compounds were dissolved in the working buffer (20 mM HEPES, pH 8.0 and 200 mM NaCl). The titration was performed with 19 successive injections of tyrosine or phenylalanine at 300 μM concentration, with a 120 s interval between each injection, into the cell solution containing FjTAL at 10 μM concentration at 50 × g at 25 °C. Correspondingly, tyrosine or tyrosinol with a concentration of 500 μM was titrated into YARS1 at 25 μM concentration. The Origin software was used for baseline correction, integration and isotherm curve fitting to a single-site binding model to calculate the equilibrium dissociation constant (Kd) values of binding reactions.
Mitochondrial complex activity
The activities of mitochondrial complex I-V from cells or liver tissues were tested using mitochondrial complex activity assay kit (Solarbio, Cat# BC0515, Cat# BC3235, Cat# BC3245, Cat# BC0945 and Cat# BC1440), respectively, according to the manufacturer’s instructions.
Single-cell RNA-seq and analysis
The single-cell RNA-seq (scRNA-seq) cohort was obtained from the Cancer Single-cell Expression Map (CancerSCEM) at the China National Centre for Bioinformation (CNCB). Raw data were sourced from the following datasets in the GEO database: GSE107747, GSE202642, GSE242889, GSE233405, and GSE155481. Analysis was conducted using the Seurat v5.2.1 R package. Hepatocyte and hepatocellular carcinoma cell clusters were extracted from CancerSCEM-LIHC (n = 10). Cell clusters were defined using the SingleR R package, with hepatocellular carcinoma cells and hepatocyte clusters identified based on MYC expression levels. The Dimplot and Featureplot were generated using SCpubr and Nebulosa R packages.
RNA sequencing and data analysis
mRNAs were enriched from total RNA using Oligo dT magnetic beads, followed by fragmentation and synthesis of first and second-strand cDNA. Terminal repair, poly-A addition, adapter ligation, fragment selection, amplification, and purification prepared the library for sequencing on SURFSeq 5000 (GeneMind). Raw reads underwent quality control and filtration before being mapped to the GRCh38/hg38 genome using HISAT2. The feature Counts tools from Subread software filtered non-uniquely mapped and unqualified reads, generating an RNA-seq count matrix. Differential expression gene (DEG) analysis was executed by the DEseq2 (V1.42.1) R packages based on the count matrix. The visualisation of the results of DEGs analysis (Volcano plot) was generated using ggplot2 R package, and the DEGs list was established by setting the log2Foldchange threshold at 1 and p value < 0.05. Data are shown in Supplementary Data 2.
Tandem mass tag-labelled (TMT) quantitative proteomics and analysis
Total proteins were extracted, quantified using BCA assay, digested into peptides and labelled with TMT. These TMT-labelled peptides were fractionated via reversed-phase chromatography on an Agilent 1100 HPLC system and analysed by LC-MS. All analyses were performed by a Q-Exactive mass spectrometer (Thermo, USA) equipped with a Nanospray Flex source (Thermo, USA). Proteome Discoverer (v.2.4) was used to search all the Q-Exactive raw data thoroughly against the sample protein database. Database search was performed with Trypsin digestion specificity. Alkylation of cysteine was considered a fixed modification during database searching. For the protein quantification method, the labelling method was selected. A global false discovery rate (FDR) was set to 0.01, and protein groups considered for quantification required at least two peptides. After peptide quality control and median normalisation, a protein expression level matrix was generated. Inter-group differential protein analysis was conducted using the rstatix (version 0.7.2) R package with a mean-based differential analysis approach. A volcano plot of differential proteins was created using the ggplot2 R package. Data are shown in Supplementary Data 3.
Ribosome profiling (Ribo-seq) and data analysis
Cells were collected and washed with ice-cold PBS containing cycloheximide, then snap-frozen in liquid nitrogen. The frozen cell pellets were lysed using ice-cold lysis buffer containing 150 mM Tris-HCl, 10 mM NaCl, 1% Triton X-100, 0.5 mM DTT, 1% NP–40, 100 μg/mL CHX and DNase. The supernatant was treated with RNase, and urea gels were used to isolate ribosome-protected fragments (RPFs). The library was successfully prepared and sequenced using the SURFSeq 5000 platform (GeneMind). The clean reads were obtained by filtering the raw reads, mapped to the GRCh38/hg38 genome, human rRNA sequences, human tRNA sequences, and human CDS or cDNA sequences. Sequencing quality was assessed by RPFs distribution analysis performed using the genomic alignments R package and riboWaltz. Ribo-seq read count matrix was generated using feature counts software by counting the number of reads mapping to the ORF of protein - coding genes. Normalisation and differential translation analysis were performed using the DEseq2 R package. Visualisation of differential translation genes (volcano plot) was generated using the ggplot2 R package. The genome visualisation was performed using the integrative genomics viewer (IGV, version 2.18.4) based on BAM files of each sample mapped to GRCh38/hg38 genome. Data are shown in Supplementary Data 4.
Metabolomics analysis using GC-MS and LC-MS, and data processing
SNU398 cells were seeded in 15-cm plates in tyrosine-restricted medium or normal medium. After 2 days, a total of 1 × 10⁸ cells were harvested from each group to generate four technical replicates. Then, pre-cooled methanol-water (V:V = 4:1) and chloroform were added to the cells, followed by sonication for 20 min in an ice-water bath and standing at −40 °C overnight. After being centrifuged at 16,000 × g for 10 min at 4 °C, 2/3 of the supernatant was removed for LC-MS detection, and the remaining supernatant was used for GC-MS. To perform LC-MS, the dried supernatants were reconstituted with methanol-water (V: V = 1:4), and mixed with the internal standard (0.1 mg/mL), vortexed for 30 s, sonicated for 3 min in an ice-water bath, and then stood for 2 hours at −40 °C. After being centrifuged again at 16,000 × g for 10 min at 4 °C, the supernatant was aspirated with a syringe, filtered using a 0.22 μm organic-phase pinhole filter, and stored at −80 °C for LC-MS analysis. The procedure of GC-MS analysis was described above in amino acid profiling. LC-MS analysis was done by Waters ACQUITY UPLC I-Class plus and Thermo QE system. GC-MS analysis was performed using an Agilent 8890-5977B quadrupole instrument time-of-flight mass spectrometer.
Progenesis QI v3.0 software (Nonlinear Dynamics, Newcastle, UK) was used to pre-process the raw data for baseline filtering, peak identification, integration, retention time correction, peak alignment and normalisation. MS-DIAL software was used to perform qualitative and quantitative analysis of small molecules from GC-MS data collected in Full Scan mode. The identification of compounds was based on multiple dimensions such as RT (retention time), exact mass number, secondary fragments, and isotope distribution. Identification analysis was performed using the Human Metabolome Database (HMDB), Lipidmaps (v2.3), METLIN database, and LuMet Animal 3.0 local database. Data are shown in Supplementary Data 5.
Data mining of genome-wide CRISPR screens
Screen results from the DepMap project (DepMap Public 24Q2+ Score Chronos), visualised through ggplot2 R package were used to identify the dependencies of YARS or tyrosine metabolism-related gene sets in HCC cell lines.
Multi-omics analysis and visualisation
The GSEA analysis based on the KEGG and GO database was performed by clusterProfiler (V4.10.1) R package, and GSEA plot was generated by ggplot2 R package and enrichplot R package. Combination analysis of TCGA with CPTAC, RNA-seq with proteomics, RNA-seq with Ribo-seq, and proteomics with Ribo-seq was performed based on the results of differential analysis of each omics (log2FoldChange) and visualised by ggplot2 R package.
Codon and amino acid frequency analysis
The complete CDS sequences of the human genome were retrieved from the NCBI CCDS database. After filtering all sequences lacking an ATG initiation codon, codon counts for all 64 codon types across CDS sequences were quantified using Biostrings R package. Then, the percentage distribution of each codon within individual CDS sequences was calculated to generate a codon frequency matrix. Subsequently, the amino acid counts and frequency matrix for all 20 amino acids were generated according to the codon table and codon count matrix. Based on the codon and amino acid frequency matrix, we ranked all CDS from high to low based on the frequency of each codon and amino acid, and generated codon TAC and tyrosine frequency distribution curve of all human genome CDS using ggplot2. Next, the differential codons and amino acids of target gene sets screened from multi-omics were calculated based on differential value (log2FoldChange) and codons and amino acids frequency matrix, and the differential codons and amino acids heatmap was generated by ggplot2.
Statistical and reproducibility
All results were shown as the mean ± standard deviation, and the data were from at least three independent experiments unless specifically stated. The statistical analyses were performed using GraphPad Prism 10 or the corresponding R packages. GraphPad Prism 10 software was used for mouse survival studies to generate a survival curve and perform statistical significance analysis. Kaplan–Meier survival curves were generated from RNA sequencing gene expression data and the outcome of 330 hepatocellular carcinomas and analysed for statistical significance using the R software package. An unpaired two-tailed Student’s t test was used to analyse the data for two-group comparisons. ANOVA analysis was used to compare multiple groups as described in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The proteomic and metabolome data generated in this study have been deposited in the CNCB database under accession codes OMIX010875 and OMIX010876. The proteomic and metabolomic data are also available under accession code IPX0014937000 in the Integrated Proteome Resources. The RNA-seq and Ribo-seq data generated in this study have been deposited in the CNCB database under accession code HRA012185. The HCC single-cell transcriptome data used in this study are available in the Gene Expression Omnibus (GEO) database under accession codes GSE202642, GSE242889, and GSE155481. The raw data of RNA-seq, Ribo-seq, proteomic, and metabolomic data are available under open access. The sequences of primers, shRNA, and probes used in this article are supplied in Supplementary Data 1. The data from transcriptomics, translatomics, proteomics, and metabolomics are supplied in Supplementary Data 2–5, respectively. The raw numbers for charts and graphs are available in the Source Data file. Source data are provided with this paper.
References
Mossmann, D. et al. Arginine reprograms metabolism in liver cancer via RBM39. Cell 186, 5068 (2023).
Yuan, H. R. et al. Lysine catabolism reprograms tumour immunity through histone crotonylation. Nature 617, 818 (2023).
Lee, M. S. et al. Ornithine aminotransferase supports polyamine synthesis in pancreatic cancer. Nature 616, 339 (2023).
Badgley, M. A. et al. Cysteine depletion induces pancreatic tumor ferroptosis in mice. Science 368, 85 (2020).
Thandapani, P. et al. Valine tRNA levels and availability regulate complex I assembly in leukaemia. Nature 601, 428 (2022).
Wu, X. J. et al. Threonine fuels glioblastoma through YRDC-mediated codon-biased translational reprogramming. Nat. Cancer 5, 1024–1044 (2024).
Calvo, A. C., Pey, A. L., Ying, M., Loer, C. M. & Martinez, A. Anabolic function of phenylalanine hydroxylase in Caenorhabditis elegans. Faseb J. 22, 3046–3058 (2008).
Flydal, M. I. & Martinez, A. Phenylalanine hydroxylase: function, structure, and regulation. IUBMB Life 65, 341–349 (2013).
Lee, Y. et al. Structural basis of anticancer drug recognition and amino acid transport by LAT1. Nat. Commun. 16, 1635 (2025).
Vumma, R., Wiesel, F. A., Flyckt, L., Bjerkenstedt, L. & Venizelos, N. Functional characterization of tyrosine transport in fibroblast cells from healthy controls. Neurosci. Lett. 434, 56–60 (2008).
Wyant, G. A. et al. mTORC1 activator SLC38A9 is required to efflux essential amino acids from lysosomes and use protein as a nutrient. Cell 171, 642–654.e612 (2017).
Wang, J. et al. Decreased SLC27A5 suppresses lipid synthesis and tyrosine metabolism to activate the cell cycle in hepatocellular carcinoma. Biomedicines 10, 2334 (2022).
Kehar, M., Sen Sarma, M., Seetharaman, J., Jimenez Rivera, C. & Chakraborty, P. Decoding hepatorenal tyrosinemia type 1: Unraveling the impact of early detection, NTBC, and the role of liver transplantation. Can. Liver J. 7, 54–63 (2024).
Beyzaei, Z., Nabavizadeh, S., Karimzadeh, S. & Geramizadeh, B. The mutation spectrum and ethnic distribution of non-hepatorenal tyrosinemia (types II, III). Orphanet J. Rare Dis. 17, 424 (2022).
Mei, Y. et al. A method to establish a c-Myc transgenic mouse model of hepatocellular carcinoma. MethodsX 7, 100921 (2020).
Fang, C. N. et al. Metabolic reprogramming and risk stratification of hepatocellular carcinoma studied by using gas chromatography-mass spectrometry-based metabolomics. Cancers 14, 231 (2022).
Sano, A. et al. Steatotic hepatocytes release mature VLDL through methionine and tyrosine metabolism in a Keap1-Nrf2-dependent manner. Hepatology 74, 1271–1286 (2021).
Dhanasekaran, R. et al. The MYC oncogene - the grand orchestrator of cancer growth and immune evasion. Nat. Rev. Clin. Oncol. 19, 23–36 (2022).
Liu, F., Liao, Z. & Zhang, Z. MYC in liver cancer: mechanisms and targeted therapy opportunities. Oncogene 42, 3303–3318 (2023).
Lourenco, C. et al. MYC protein interactors in gene transcription and cancer. Nat. Rev. Cancer 21, 579–591 (2021).
Zhang, H. L. et al. A MYC-STAMBPL1-TOE1 positive feedback loop mediates EGFR stability in hepatocellular carcinoma. Cell Rep. 43, 114812 (2024).
Hsu, T. Y. T. et al. The spliceosome is a therapeutic vulnerability in MYC-driven cancer. Nature 525, 384 (2015).
Fu, L. et al. Down-regulation of tyrosine aminotransferase at a frequently deleted region 16q22 contributes to the pathogenesis of hepatocellular carcinoma. Hepatology 51, 1624–1634 (2010).
Tong, M. et al. Loss of tyrosine catabolic enzyme HPD promotes glutamine anaplerosis through mTOR signaling in liver cancer. Cell Rep. 36, 109617 (2021).
Yang, F. et al. GSTZ1-1 deficiency activates NRF2/IGF1R axis in HCC via accumulation of oncometabolite succinylacetone. Embo J. 38, e101964 (2019).
Zhao, R. et al. Nuclear ATR lysine-tyrosylation protects against heart failure by activating DNA damage response. Cell Rep. 42, 112400 (2023).
Sajish, M. & Schimmel, P. A human tRNA synthetase is a potent PARP1-activating effector target for resveratrol. Nature 519, 370–373 (2015).
Grajchen, E. et al. Fatty acid desaturation by stearoyl-CoA desaturase-1 controls regulatory T cell differentiation and autoimmunity. Cell Mol. Immunol. 20, 666–679 (2023).
Wortel, I. M. N., van der Meer, L. T., Kilberg, M. S. & van Leeuwen, F. N. Surviving stress: modulation of ATF4-mediated stress responses in normal and malignant cells. Trends Endocrinol. Metab. 28, 794–806 (2017).
Tesfay, L. et al. Stearoyl-CoA desaturase 1 protects ovarian cancer cells from ferroptotic cell death. Cancer Res. 79, 5355–5366 (2019).
Sen, U., Coleman, C. & Sen, T. Stearoyl coenzyme A desaturase-1: multitasker in cancer, metabolism, and ferroptosis. Trends Cancer 9, 480–489 (2023).
Kamphorst, J. J. et al. Hypoxic and Ras-transformed cells support growth by scavenging unsaturated fatty acids from lysophospholipids. Proc. Natl. Acad. Sci. USA 110, 8882–8887 (2013).
Labbe, K. et al. Specific activation of the integrated stress response uncovers regulation of central carbon metabolism and lipid droplet biogenesis. Nat. Commun. 15, 8301 (2024).
Liang, D. G. et al. Ferroptosis surveillance independent of GPX4 and differentially regulated by sex hormones. Cell 186, 2748 (2023).
Dierge, E. et al. Peroxidation of n-3 and n-6 polyunsaturated fatty acids in the acidic tumor environment leads to ferroptosis-mediated anticancer effects. Cell Metab. 33, 1701 (2021).
Di Bona, M. et al. Micronuclear collapse from oxidative damage. Science 385, eadj8691 (2024).
Vogler, M. et al. The BCL2 family: from apoptosis mechanisms to new advances in targeted therapy. Signal Transduct. Target Ther. 10, 91 (2025).
To, T. L. et al. A compendium of genetic modifiers of mitochondrial dysfunction reveals intra-organelle buffering. Cell 179, 1222 (2019).
You, A. et al. Metformin sensitizes sorafenib to inhibit postoperative recurrence and metastasis of hepatocellular carcinoma in orthotopic mouse models. J. Hematol. Oncol. 9, 20 (2016).
Liu, Y., Stockwell, B. R., Jiang, X. & Gu, W. p53-regulated non-apoptotic cell death pathways and their relevance in cancer and other diseases. Nat. Rev. Mol. Cell Biol. 26, 600–614 (2025).
Chang, K. et al. DPP9 stabilizes NRF2 to suppress ferroptosis and induce sorafenib resistance in clear cell renal cell carcinoma. Cancer Res. 83, 3940–3955 (2023).
Das, A. M. et al. Diagnosis, treatment, management and monitoring of patients with tyrosinaemia type 1: consensus group recommendations from the German-speaking countries. J. Inherit. Metab. Dis. 48, e12824 (2025).
Cramer, S. L. et al. Systemic depletion of L-cyst(e)ine with cyst(e)inase increases reactive oxygen species and suppresses tumor growth. Nat. Med. 23, 120–127 (2017).
Batool, T., Makky, E. A., Jalal, M. & Yusoff, M. M. A comprehensive review on L-Asparaginase and its applications. Appl. Biochem. Biotechnol. 178, 900–923 (2016).
Hendrikse, N. M. et al. Exploring the therapeutic potential of modern and ancestral phenylalanine/tyrosine ammonia-lyases as supplementary treatment of hereditary tyrosinemia. Sci. Rep. 10, 1315 (2020).
Jendresen, C. B. et al. Highly active and specific tyrosine ammonia-lyases from diverse origins enable enhanced production of aromatic compounds in bacteria and Saccharomyces cerevisiae. Appl. Environ. Microbiol. 81, 4458–4476 (2015).
Abergel, C., Rudinger-Thirion, J., Giege, R. & Claverie, J. M. Virus-encoded aminoacyl-tRNA synthetases: structural and functional characterization of mimivirus TyrRS and MetRS. J. Virol. 81, 12406–12417 (2007).
Tong, H. et al. Dual impacts of serine/glycine-free diet in enhancing antitumor immunity and promoting evasion via PD-L1 lactylation. Cell Metab. 36, 2493–2510.e2499 (2024).
Shrestha, S. et al. Pegvaliase therapy for phenylketonuria: real-world safety, efficacy, and medication access outcomes in a pharmacist-led pegvaliase program. Genet. Med. 27, 101405 (2025).
Gu, P. et al. An engineered Escherichia coli Nissle strain prevents lethal liver injury in a mouse model of tyrosinemia type 1. J. Hepatol. 80, 454–466 (2024).
Kosakamoto, H., Sakuma, C., Okada, R., Miura, M. & Obata, F. Context-dependent impact of the dietary non-essential amino acid tyrosine on Drosophila physiology and longevity. Sci. Adv. 10, eadn7167 (2024).
Zhang, X. et al. Reprogramming tumour-associated macrophages to outcompete cancer cells. Nature 619, 616–623 (2023).
Wang, Y. D. et al. Farnesoid X receptor antagonizes nuclear factor κB in hepatic inflammatory response. Hepatology 48, 1632–1643 (2008).
Pavlova, N. N. et al. Translation in amino-acid-poor environments is limited by tRNA(Gln) charging. Elife 9, e62307 (2020).
Nijtmans, L. G., Henderson, N. S. & Holt, I. J. Blue Native electrophoresis to study mitochondrial and other protein complexes. Methods 26, 327–334 (2002).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Heiman, M., Kulicke, R., Fenster, R. J., Greengard, P. & Heintz, N. Cell type-specific mRNA purification by translating ribosome affinity purification (TRAP). Nat. Protoc. 9, 1282–1291 (2014).
Jin, H. et al. EGFR activation limits the response of liver cancer to lenvatinib. Nature 595, 730–734 (2021).
Acknowledgements
We thanked the Centre of Pharmaceutical Technology at Tsinghua University, especially Jinfeng He and Jingjing Rong, for their assistance with the ITC assay. We thanked the Institute of Clinical Medical Sciences at China-Japan Friendship Hospital, especially Jing Li and Yuting Zhang, for their help in capturing electron microscope images. We thanked Henan Cao for algorithmic support. This work was supported by the National Natural Science Foundation of China (Grant No. 82560575) to W.-D.C., Beijing Natural Science Foundation (Grant No. 7252088), the National Key R&D Programme of China (Grant No. 2021YFC2103900), the National Natural Science Foundation of China (Grant No. 81970551) to Y.-D.W., and the National Natural Science Foundation of China (Grant No. 81970726) to W.-D.C.
Author information
Authors and Affiliations
Contributions
H.L.Z. designed, performed the experiments, analysed and discussed the data, and wrote the manuscript draft; Z.X.W. analysed and discussed the data, and wrote the manuscript draft; Y.Q.Z., X.Y.D., J.Z., F.S., H.J.Z., Y.L., H.B.W., Y.T.Y., H.Y.L., B.Z., C.L., F.G., R.L.Z., Q.P.Y., Y.F., S.Z.L. performed the experiments and acquired the data; W.-D.C. and Y.-D.W. designed the experiments, analysed and discussed the data, wrote the manuscript, and secured funding for the research. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Mariia Yuneva, who co-reviewed with Ersa Gjelaj, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, H., Wang, Z., Zhao, Y. et al. Targeting tRNA-dependent tyrosine usage unveils a metabolic vulnerability in hepatocellular carcinoma. Nat Commun 17, 2244 (2026). https://doi.org/10.1038/s41467-026-70112-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70112-z