Abstract
Understanding how chromosome 21 gene dosage contributes to neurodevelopmental phenotypes in trisomy 21 (T21) remains a fundamental challenge. Here, we perform transcriptome-wide RNA-sequencing of fetal cortical and hippocampal tissues from T21 cases and euploid controls collected during mid-gestation, a critical window for human brain development. We identify widespread gene expression dysregulation with significant enrichment for chromosome 21 genes and perturbation of neurodevelopmental, synaptic, and immune-related pathways. Among the most strongly dysregulated genes is ADARB1, a chromosome 21-encoded RNA editing enzyme, whose overexpression associates with increased adenosine-to-inosine RNA editing, with consistent over-editing at functionally important recoding sites in glutamate and GABA receptor-related genes, including GRIK2, GRIA2, GRIA3, and GABRA3, across cortex and hippocampus. Meta-analyses across independent transcriptomic datasets validate robust chromosome 21 dosage effects, including ADARB1 overexpression and over-editing at 3′UTRs and GRIA3. These findings implicate dysregulated RNA editing as a post-transcriptional mechanism contributing to fetal neuropathology in T21.
Similar content being viewed by others
Introduction
Trisomy 21 (T21), the genetic hallmark of Down syndrome (DS), results in the triplication and overexpression of genes on chromosome 21, disrupting key neurodevelopmental processes1. Although the genetic cause of DS is well defined, the molecular mechanisms by which T21 disrupts fetal brain development remain incompletely understood and mechanistically unresolved. Transcriptomic analyses across developmental stages and brain regions have revealed widespread dysregulation of gene expression, implicating both chromosome 21 genes and broader genome-wide effects2,3,4. In fetal brain from individuals with DS, studies have identified spatially and temporally specific transcriptional changes, including impaired oligodendrocyte differentiation and disrupted myelination3, suggesting a mechanistic link to abnormalities in white matter integrity and neural circuit formation. In postnatal and adult DS brains, transcriptional dysregulation persists, extending to pathways involved in oxidative stress responses, cytoskeletal organization, and immune signaling5,6,7.
Emerging evidence suggests that post-transcriptional regulation, particularly RNA editing, may play a critical role in the neurodevelopmental phenotypes of DS. ADARB1, a chromosome 21-encoded enzyme that catalyzes adenosine-to-inosine (A-to-I) editing, modulates RNA sequence diversity, neuronal excitability, and synaptic function during brain development8,9. A dosage-sensitive gene, ADARB1 is consistently overexpressed in DS cellular models and in vivo tissues4,10,11,12,13. However, the extent to which this alters RNA editing activity in the developing human brain remains unclear. By contrast, precise editing disruptions at recoding sites have been implicated in related neurodevelopmental and neuropsychiatric disorders, including Alzheimer’s disease14, autism spectrum disorder15, and schizophrenia16, where they impair synaptic signaling and circuit maturation. Whether similar editing-dependent mechanisms contribute to neurodevelopmental abnormalities in DS remains an open question.
In parallel with transcriptional and post-transcriptional disruptions, immune dysregulation has emerged as a central and persistent feature of DS. The triplication of interferon receptor genes (IFNAR1, IFNAR2, and IFNGR2) on chromosome 21 amplifies interferon signaling pathways, leading to chronic immune activation and sustained upregulation of interferon-stimulated genes17,18,19,20. While ADAR, particularly the interferon-inducible p150 isoform, is canonically upregulated by inflammatory signals21,22, our prior transcriptomic analyses of non-DS human brain tissues revealed that interferon activation is also associated with increased ADARB1 expression23, suggesting that ADARB1 may also be responsive to immune tone in vivo. This is particularly relevant in DS, where gene dosage imbalance uniquely co-occurs with heightened interferon activity, creating a biologically distinct environment that may alter RNA editing programs. The intersection of interferon signaling, gene triplication, and post-transcriptional control represents a largely unexplored axis of pathology with the potential to disrupt neurodevelopmental trajectories and impair synaptic function.
Despite these insights, key gaps remain in our understanding of DS molecular pathology during early brain development. To date, only two transcriptomic studies have examined DS fetal brain tissue (using microarrays): one analyzing 15 DS and 15 control brains3, and another profiling 9 DS and 8 control brains4. While foundational, these studies, along with independent hiPSC-derived neuron models10,11,12,13, have been limited in both scale and resolution and were not designed to assess post-transcriptional regulation such as RNA editing. Additional efforts have focused primarily on postnatal or adult tissues using targeted PCR or, more recently, single-nucleus RNA sequencing (snRNA-seq)3,4,7,24. Although snRNA-seq provides cell-type-specific resolution, it is not well-suited for genome-wide, quantitative analysis of editing levels in the fetal brain. Furthermore, no study has systematically examined how chromosome 21 gene dosage, interferon signaling, and RNA editing intersect during early brain development. Addressing these gaps requires high-resolution transcriptomic profiling across key developmental windows and brain regions.
Here, we address these critical gaps by performing transcriptome-wide RNA sequencing of fetal prefrontal cortical and hippocampal tissues from 20 individuals with T21 and 27 euploid controls, spanning 13–22 weeks post-conception, a key window for human brain maturation (Fig. 1). By meta-analyzing these data with 9 independent transcriptomic datasets, we define robust and reproducible chromosome 21 dosage effects and reveal convergent transcriptional and post-transcriptional disruptions across diverse developmental stages and cellular contexts. Focusing on RNA editing, we demonstrate that ADARB1 is a dosage-sensitive regulator of A-to-I editing in the developing brain, driving premature and excessive recoding at key genes during neurodevelopment. Together, these findings establish RNA editing dysregulation as a conserved mechanism of T21 neuropathology and provide a framework for dissecting post-transcriptional contributions to altered brain development in DS.
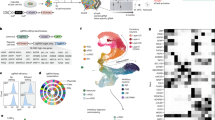
We performed deep bulk RNA sequencing of paired prefrontal cortex (PFC) and hippocampal tissue from 20 mid-gestation fetuses with trisomy 21 (T21) and 27 age- and sex-matched euploid controls (13–22 post-conception weeks [PCW]). A multi-layered analytical pipeline was applied to: (1) quantify differential gene expression and assess selective gene dosage effects on chromosome 21; (2) profile global and site-specific adenosine-to-inosine (A-to-I) editing, including recoding events that alter protein sequence; and (3) perform a harmonized meta-analysis of ten RNA-seq datasets to identify reproducibly dysregulated genes and editing sites in T21. Brain diagrams were constructed in the cerebroViz R package.
Results
We generated deep bulk RNA-sequencing profiles (34.5 M ± 5.4 M reads/sample) from fetal prefrontal cortex (PFC) and hippocampal tissues of individuals with Trisomy 21 (T21; n = 20) and age-, sex- and ancestry-matched euploid controls (n = 27) (Supplemental Data 1). To assess global transcriptomic variation, we initially applied a unified normalization strategy across all samples, quantifying expression for 16,737 genes. Rigorous quality control analyses were then conducted to identify key sources of variability in the dataset (Figs. S1–S3). First, cell type deconvolution revealed similar estimated neuronal proportions in PFC (~62.2%) and hippocampus (~62.1%), with no significant differences between T21 and control groups (Fig. S1). Next, principal component analysis (PCA) highlighted neuronal content as the dominant axis of variation (PC1; 29.2% variance explained), while PC2 (17.8% variance) clearly separated PFC and hippocampal tissues (Fig. S2). To formally assess regional differences, we performed differential expression analysis between brain regions and identified 9163 differentially expressed genes at FDR < 5%, accounting for ~54% of the transcriptome (Fig. S3). Based on these marked region-specific signatures, we conducted all downstream analyses independently for PFC and hippocampus using tailored, region-specific modeling strategies (see “Methods”).
Transcriptomic effects of T21 during early fetal brain development
To identify critical covariates for inclusion in our comparative analysis of T21 individuals and controls within each region, we used a linear mixed model to quantify the fraction of expression variance attributable to known clinical and technical factors, including neuronal cell type proportion, percentage of mRNA reads (defined as reads mapping to exons and untranslated regions), gestational age, RNA integrity number (RIN), T21 status, and sex, for each gene, separately within PFC and hippocampus (Fig. S4). Collectively, these variables accounted for ~57.1% of transcriptome-wide variation in the PFC and ~34.9% in the hippocampus. In the PFC, estimated neuronal cell type proportions exerted the largest effect, explaining a median 25.1% of the observed variation, followed by the percentage of mRNA reads (6.2%), gestational age (2.4%), and RIN (0.4%), with remaining factors contributing modestly. Expression variation attributable to T21 itself was detectable, particularly in a subset of genes located on chromosome 21, confirming the robustness of our approach. Similar patterns of variance decomposition were observed independently in the hippocampus (Fig. S4E–H).
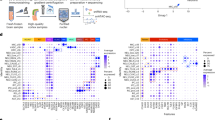
We next performed differential expression analysis comparing individuals with T21 and controls, adjusting for the possible influence of estimated neuronal cell type proportions, RIN, gestational age, sex, and percentage of mRNA reads, within each region. This analysis identified 572 differentially expressed genes (DEGs) in the PFC and 519 in the hippocampus at transcriptome-wide significance (FDR < 5%) (Fig. 2A, B, Fig. S5, Supplemental Data 2). While gene expression changes in T21 were positively correlated across regions (Spearman’s correlation, ρ = 0.40), the DEG profiles were largely region-specific. Only 155 genes were consistently dysregulated in both the PFC and hippocampus, most of which were located on chromosome 21 (Fig. 2C, D). Among the DEGs, there was significant enrichment of upregulated chromosome 21 genes in individuals with T21 in both the PFC (ngenes = 122, p = 6.7 × 10−142, one-sided FET) and hippocampus (ngenes = 118, p = 8.6 × 10−137, one-sided FET) (Fig. 2C, E), reflecting the canonical gene dosage effects associated with T21.
Differential expression between T21 (n = 20) and controls (n = 27) was tested separately in (A) PFC and (B) hippocampus using a covariate-adjusted linear model (gestational age, sex, RIN, % mRNA reads, and estimated neuronal proportion) with empirical Bayes moderation (two-sided). p Values were adjusted using the Benjamini–Hochberg false discovery rate (FDR); effect sizes are log₂ fold-changes. Each gene was plotted by log₂ fold-change (x-axis) and −log₁₀ adjusted p-value (y-axis); points are colored to indicate upregulated, downregulated, and chromosome 21-encoded genes. Brain diagrams were constructed in the cerebroViz R package. C Venn diagrams display the overlap of all differentially expressed genes (DEGs) (left) and DEGs located on chromosome 21 (right) between PFC and hippocampus. D Genome-wide concordance of expression changes between regions is shown by plotting log₂ fold-changes in PFC (x-axis) against hippocampus (y-axis); Spearman correlation coefficient (ρ) quantifies regional concordance. E Barplots show the number of DEGs (FDR < 5%) per chromosome in PFC (top) and hippocampus (bottom), separated by direction of change. F Dot plots display expression changes for all genes on chromosome 21, arranged by genomic position (x-axis) for PFC (top) and hippocampus (bottom); colors reflect FDR significance thresholds (FDR < 5%, pink; 5% <FDR < 10%, light blue; FDR > 10%, dark blue) applied to the covariate-adjusted, empirical Bayes–moderated linear model (two-sided). G For each gene, a BrainSpan-derived t-statistic summarizing prenatal versus postnatal expression bias was computed using data from matching anatomical regions (PFC and hippocampus; “Methods”). Distributions of these developmental bias scores for T21-upregulated versus T21-downregulated genes were compared using a two-sided Mann–Whitney U test. In the PFC (top) and hippocampus (bottom), a prenatal bias was observed among under-expressed genes (p = 0.02 and p = 2.3 × 10−⁵) and a postnatal bias among over-expressed genes (p = 0.01 and p = 7.6 × 10−⁷). H Normalized expression trajectories of T21-upregulated and downregulated genes were plotted across ten developmental stages in the BrainSpan bulk RNA-seq using matching PFC (top) and hippocampus (bottom) tissues, illustrating typical temporal shifts in gene expression trajectories.
Despite the strong enrichment for upregulated chromosome 21 genes, we did not observe a universal dosage effect across the entire chromosome. In the PFC, 40 genes on chromosome 21 showed no significant expression changes between T21 cases and controls, with a similar pattern observed for 50 genes in the hippocampus (Fig. 2F). These genes were distributed relatively evenly across the chromosome. Notably, chromosome 21 genes that were significantly upregulated in T21 cases were more highly expressed in T21 and control samples compared to non-significant genes (PFC: fold-change = 2.29, p = 1.1 × 10−¹⁰; hippocampus: fold-change = 2.31, p = 1.3 × 10−¹¹; Wilcoxon rank-sum test) (Fig. S6). This suggests that gene dosage sensitivity in the T21 brain preferentially affects genes that are already more actively transcribed, while lower-expressed genes may evade dysregulation, potentially due to tighter transcriptional control. Among the dysregulated chromosome 21 genes, we highlight increased expression of key interferon signaling components, including interferon gamma receptor 2 (IFNGR2) and interferon alpha/beta receptor subunits 1 and 2 (IFNAR1, IFNAR2), in both the PFC (FDR p = 6.7 × 10−⁴) and hippocampus (FDR p = 3.7 × 10−²). These genes, which canonically mediate interferon signaling in immune contexts, have also been implicated in brain development and glial function, raising the possibility that their overexpression contributes to immune-related transcriptional changes in the T21 brain. Additionally, we observed upregulation of several collagen genes critical for brain extracellular matrix integrity (COL6A1, COL6A2, COL18A1) in both regions (PFC FDR p = 1.4 × 10−²; hippocampus FDR p = 2.5 × 10−²), as well as increased expression of genes involved in ligase activity (ATP5PF, ATP5PO, GART, HLCS, DIP2A) (PFC FDR p = 2.0 × 10−²; hippocampus FDR p = 3.7 × 10−²).
Developmental imbalance of fetal gene expression patterns in individuals with T21
A substantial proportion of dysregulated T21 genes reside outside chromosome 21 and likely represent secondary, downstream effects. To assess whether non-chromosome 21 DEGs reflect broader disruptions in developmental timing, we leveraged transcriptomic data from the BrainSpan atlas to define normative gene expression trajectories in the PFC and hippocampus across prenatal and postnatal stages (Supplemental Data 3). For each gene, we computed a t-statistic capturing its expression bias toward prenatal or postnatal timepoints in typical development (see “Methods”). We then tested whether T21-overexpressed and T21-underexpressed genes were disproportionately drawn from genes with strong developmental timing biases, by comparing their BrainSpan-derived t-statistic distributions within each brain region. This analysis revealed a striking asymmetry: genes downregulated in T21 were significantly enriched for prenatally expressed genes (PFC: p = 0.02; hippocampus: p = 2.3 × 10−⁵; Mann–Whitney U), whereas upregulated genes were enriched for postnatally expressed genes (PFC: p = 0.01; hippocampus: p = 7.6 × 10−⁷; Mann–Whitney U) (Fig. 2G, H). These findings suggest that T21 disrupts the temporal coordination of gene expression, prematurely activating postnatal programs while attenuating prenatal ones during early brain development.
Systems-level dysregulation of gene expression in T21
To further characterize the biological processes underlying this temporal dysregulation, we performed functional enrichment analysis of differentially expressed genes located outside chromosome 21 (Supplemental Data 4). Using a competitive gene set ranking framework, we identified significant enrichment of downregulated genes involved in mitochondrial translation and mRNA binding in both the PFC and hippocampus (Fig. 3A, B), pointing to impaired metabolic function and post-transcriptional regulation during early neurodevelopment. This was accompanied by reduced expression of genes critical for translational initiation and elongation, ribosomal RNA processing, and nonsense-mediated mRNA decay, suggesting broader suppression of protein synthesis and RNA quality control pathways in the T21 fetal brain. In contrast, over-expressed genes were enriched for sodium channel activity across both regions and extracellular matrix (ECM) organization specifically in the hippocampus, including DCN (Decorin) and LTBP4, which regulate TGF-β signaling in the ECM microenvironment. Together, these results suggest that T21-associated gene dysregulation involves both suppression of early translational and post-transcriptional machinery and premature activation of pathways associated with neuronal excitability and structural maturation.
A Heatmap summarizing Correlation Adjusted MEan RAnk (CAMERA) gene set enrichment analysis (two-sided), a competitive method that accounts for inter-gene correlation when testing whether predefined gene sets are enriched among differentially expressed genes. P values were adjusted using the Benjamini–Hochberg false discovery rate (FDR). The y-axis lists the top five most significantly enriched gene sets for each category of DEGs within the PFC and hippocampus. B Enrichment barcode plots for selected gene sets illustrate enrichment patterns across ranked genes (x-axis) sorted by log₂ fold-change in T21. Vertical bars indicate the positions of genes from the target gene set, demonstrating enrichment for sodium channel activity, interferon receptor signaling, and mRNA binding, highlighted examples corresponding to gene sets in panel (A). C–E Gene set preservation analysis assessed whether co-expression patterns of predefined biological gene sets were maintained between T21 (n = 20) and control samples (n = 27). Preservation was evaluated in the (C, D) PFC and (E) hippocampus using a combination of Zsummary metrics with empirical permutation testing (two-sided). Multiple testing was controlled by Benjamini–Hochberg adjustment. Gene sets were classified as non-preserved if they met multiple stringent criteria: low Zsummary1 (<2), lack of statistical significance after FDR correction (p.adj > 0.1), deviation from predicted preservation by ≥4 Z-score units, and significant preservation under random permutation (p.random.adj <0.05) and are colored-coded accordingly. Inset boxplots summarize the distribution of Zsummary scores across flagged non-preserved gene sets. Boxplots show the median (center line), interquartile range (box), and whiskers extending to 1.5× IQR. D As a representative example, correlation matrices of genes within the astrocyte activation gene ontology term are visualized across 27 control samples (top, unsupervised clustering) and 20 T21 samples (bottom, force-ordered to match control structure), illustrating disrupted co-regulatory structure in T21.
While differential expression and functional enrichment analyses identify genes and pathways with altered mean expression levels, they do not capture whether the underlying coordination of gene activity within biological programs is preserved. To assess whether T21 disrupts the structural organization of transcriptional networks, we first applied weighted gene co-expression network analysis (WGCNA) to identify unsupervised gene modules in PFC and hippocampus. This analysis did not reveal major differences in overall module architecture between T21 and control samples (Fig. S7).
We next used a gene set preservation framework to test whether predefined biological gene sets retained coordinated expression across groups, an approach that can detect systems-level dysregulation even in the absence of changes in gene abundance (see “Methods”) (Supplemental Data 5). In the PFC, eight gene sets exhibited markedly reduced preservation in T21, including those involved in astrocyte activation, sodium ion transport, and macrophage differentiation, suggesting altered regulatory interactions among glial cell types (Fig. 3C, D). In contrast, the hippocampus showed more widespread co-expression disruption, with 259 gene sets demonstrating poor preservation between individuals with T21 and controls (Fig. 3E, F). These gene sets clustered into four major domains: (1) neurodevelopment and synaptic plasticity (e.g., synapse maturation, long-term synaptic depression, positive regulation of excitatory postsynaptic potential); (2) chromatin and epigenetic regulation (e.g., ATP-dependent chromatin remodeler activity, chromatin protein adapter activity); (3) mitochondrial metabolism (e.g., NADH dehydrogenase ubiquinone activity, oxidoreductase activity, NADH metabolic process); and (4) immune signaling (e.g., interleukin-4 production, regulation of neuroinflammatory response, monocyte differentiation). These findings suggest that T21 disrupts the integrity of gene co-regulation across multiple biological systems, particularly those governing neuronal signaling, energy metabolism, and glial immune responses.
Meta-analysis across developmental datasets validates chromosome 21 dosage effects in T21
To validate key findings and assess the generalizability of chromosome 21 dosage effects, we conducted a meta-analysis combining our fetal RNA-seq data with nine publicly available RNA-sequencing datasets (Table 1). Altogether, we analyzed 108 T21 and 117 euploid control samples from ten datasets, spanning early gestational stages and diverse neural lineages (e.g., iPSCs, NPCs, neurons, fibroblasts), thereby capturing diverse but developmentally relevant cellular states. To minimize inter-study bias, all datasets were reprocessed through a unified RNA-seq analysis pipeline, standardizing quantification and differential expression estimation across studies. For each dataset, a linear model with empirical Bayes moderation was applied to estimate log₂ fold-changes, adjusting for available biological and technical covariates (e.g., age, sex, cell type proportion, and sequencing depth). These covariate-adjusted estimates were then synthesized across datasets using a random-effects meta-regression framework to account for between-study heterogeneity. One previously published microarray study of T21 fetal brain3 was excluded from our meta-analysis due to technical and analytic limitations. Despite profiling overlapping regions and developmental stages, it showed only modest transcriptome-wide concordance with our RNA-seq data (PFC: r = 0.065; hippocampus vs. cerebellum: r = 0.11) and weak correlation for chromosome 21 genes (r = 0.27 and 0.10, respectively; Fig. S8, Supplemental Data 6). These discrepancies, along with the platform’s inability to assess RNA editing, precluded its inclusion in our synthesis. While cellular contexts varied across datasets, our goal was to identify genes consistently dysregulated across developmental lineages, rather than those specific to any single cell type. To ensure analytical consistency across platforms and study designs, each dataset was reprocessed from raw FASTQ files using a standardized RNA-seq pipeline for alignment, quantification, and normalization (see “Methods”).
Despite variability in sample sizes and experimental conditions, a consistent pattern emerged: genome-wide fold-change distributions revealed a distinct and reproducible elevation of gene expression on chromosome 21 in all comparisons (Fig. S9). When plotting log₂ fold changes across the transcriptome, chromosome 21 genes consistently showed an upward shift relative to the rest of the genome, reflecting their dosage-driven overexpression in T21. This global effect, observed across iPSCs, NPCs, neurons, and fibroblasts, reinforces the robustness of chromosome 21 gene dysregulation as a defining feature of T21, regardless of tissue or developmental context. Notably, this effect was not uniformly distributed across the chromosome: only a subset of chromosome 21 genes showed consistent overexpression (Fig. S10), reinforcing a model of targeted rather than global dosage sensitivity, consistent with emerging concepts of dosage compensation and chromatin architecture.
To formally synthesize findings across datasets, we performed a transcriptome-wide meta-analysis, identifying 543 upregulated and 437 downregulated genes in T21 at strict genome-wide significance (FDR < 1%) (Fig. 4A, Supplemental Data 7), reflecting a more conservative threshold appropriate for integrating heterogeneous datasets and reducing false positives. Among the upregulated genes, 149 (27.4%) were located on chromosome 21. The average log₂ fold-change for chromosome 21 genes approximated 0.58 (1.5-fold), consistent with expectations for gene dosage, while variation in effect sizes pointed to selective transcriptional sensitivity across loci (Fig. 4B). This analysis produced a robust catalog of consistently dysregulated chromosome 21 genes across diverse cellular contexts, including ANKRD18A, LNCTSI, MX1, IL10RB, and APP (Fig. 4C). Forest plots confirmed reproducible overexpression for multiple high-confidence genes, including DYRK1A, IFNAR1, IFNAR2, and ADARB1 (Fig. 4D–G). Notably, transcriptome-wide differential expression patterns in our fetal PFC and hippocampus closely mirrored those from independent datasets, underscoring the robustness and cross-context validity of our meta-analytic framework (Fig. S11).
A Volcano plot displaying transcriptome-wide differential expression results from a meta-analysis of ten independent T21 versus control RNA-seq studies. For each dataset, differential expression was estimated using a covariate-adjusted linear model with empirical Bayes moderation (two-sided). Dataset-specific log₂ fold-changes (x-axis) were synthesized using a random-effects meta-regression model to account for between-study heterogeneity; P values were adjusted using Benjamini–Hochberg FDR (y-axis). Genes are colored according to direction of dysregulation and chromosomal location, with chromosome 21 genes highlighted. The top ten T21-related over-expressed genes are labeled. B Average log₂ fold-change (y-axis) per chromosome, showing that chromosome 21 genes approximate the expected ~1.5-fold increase due to gene triplication. C Ranked plot of log₂ fold-changes for all significant chromosome 21 genes (FDR < 1%), arranged by genomic position (x-axis), illustrating selective rather than uniform dosage sensitivity. Eight well-known T21-related genes and ADARB1 are highlighted. D–G Forest plots display standardized mean differences in expression across ten datasets for four high-confidence dosage-sensitive genes: ADARB1, DYRK1A, IFNAR1, and IFNAR2. Forest plots show dataset-specific effect sizes and 95% confidence intervals, with summary estimates from a random-effects model (meta-regression framework). Summary diamonds (pink) denote the combined effect estimates derived from a random-effects meta-analysis, with axes scaled to the minimum and maximum confidence intervals for each gene to facilitate comparison.
Among these, ADARB1 was consistently overexpressed across most independent studies, making it a particularly compelling candidate for further investigation due to its known role in RNA editing and neurodevelopment. This gene encodes an adenosine deaminase involved in A-to-I editing, a post-transcriptional mechanism that modifies RNA sequences to regulate protein function and neuronal excitability during development9. Despite its consistent overexpression in both our fetal brain dataset and across external studies, ADARB1 has received relatively limited attention in the context of T21. Its dosage sensitivity, role in RNA editing, and potential interaction with interferon signaling pathways position it as a strong candidate for investigating the downstream consequences of gene dosage imbalance. We therefore next examined whether ADARB1 overexpression in T21 is associated with altered A-to-I editing in the developing brain.
ADARB1 is associated with increases in A-to-I editing and excessive RNA recoding
A-to-I editing is catalyzed by the ADAR family of enzymes, including ADAR1 and ADAR2 (ADARB1), which act on double-stranded RNA structures to diversify the transcriptome25. ADAR1, encoded on chromosome 1, exists in two isoforms: the constitutively expressed nuclear isoform p110 and the cytoplasmic, interferon-inducible isoform p150, which plays a key role in immune signaling26. ADAR2 is highly expressed in the central nervous system and mediates site-specific editing of protein-coding transcripts essential for neuronal development27. In contrast, ADAR3 (ADARB2), also brain-specific, lacks catalytic activity and is thought to act as a dominant-negative regulator of A-to-I editing through competition for RNA substrates28.
In our fetal brain RNA-seq data, we observed significant dosage-sensitive upregulation of ADARB1 in individuals with T21, in both the PFC (FDR p = 4.0 × 10−⁷) and hippocampus (FDR p = 0.04), with no significant changes in expression for ADAR or ADARB2 (Fig. 5A, Supplemental Data 8). To assess the impact of this dosage effect on global A-to-I editing activity, we computed the Alu editing index (AEI), defined as the ratio of A-to-G edited reads to total adenosine coverage in Alu elements across the transcriptome (see “Methods”). AEI values were significantly elevated in T21 compared to controls in the PFC (p = 0.01, linear regression), and showed a marginal trend in the hippocampus (p = 0.05) (Fig. 5B). To evaluate the contribution of each ADAR enzyme to global editing levels, we performed a bootstrapped linear regression of AEI values against the expression of ADAR, ADARB1, and ADARB2. Among the three, ADARB1 exhibited the strongest positive association with AEI (bootstrapped slope = 0.27), while ADAR (slope = 0.03) and ADARB2 (slope = 0.11) showed weaker and more variable effects (Fig. 5C). Confidence intervals for ADAR and ADARB2 were broad, reflecting greater inter-sample variability. These results position ADARB1 as the primary contributor to elevated A-to-I editing in T21 fetal brain, with the steepest and most consistent slope indicating the greatest per-unit effect of gene expression on editing activity, capturing regulatory influence more directly than correlation-based approaches.
A Normalized expression levels of ADAR, ADARB1, and ADARB2 in the PFC and hippocampus of T21 (n = 20) and control samples (n = 27). Differential expression of ADAR family genes was assessed using an empirical Bayes–moderated linear model (two-sided) within each region; p values were adjusted transcriptome-wide using Benjamini–Hochberg FDR. B Alu editing index (AEI), a global measure of A-to-I editing activity, calculated from bulk RNA-seq in PFC and hippocampus in T21 (n = 20) and controls (n = 27). Significance was assessed by two-sided linear regression, with exact p-values reported. C Bootstrapped two-sided linear regressions showing the relationship between AEI and normalized expression (log₂CPM) of ADAR, ADARB1, and ADARB2. D Total number of detected A-to-I editing sites per brain region (top) and the subset identified as differentially edited between T21 and control samples (bottom). Venn diagrams indicate regional overlap in total site detection (top) and differential editing (bottom). Volcano plots depicting changes in editing levels in T21 relative to controls, comparing the strength of significance (−log10 Adj. p-value; y-axis) to delta editing level (%; axis) for the E PFC and F hippocampus. Differential RNA editing was tested using a covariate-adjusted linear model (gestational age, sex, RIN, % mRNA reads, and estimated neuronal proportion) with empirical Bayes moderation (two-sided). p Values were adjusted using Benjamini–Hochberg FDR; effect sizes are Δ editing (%). G Editing level differences between T21 (n = 20) and controls (n = 27) for six functionally characterized RNA recoding sites across the PFC and hippocampus. Editing differences at recoding sites were derived from transcriptome-wide comparisons in panels (E) and (F). H Two-sided Pearson correlation coefficients (and standard error bars) between editing levels at the six recoding sites and the expression of each ADAR family member across 47 samples (20 T21 + 27 controls). I Developmental trajectories of editing levels at the six recoding sites across typical brain maturation, plotted using BrainVar data from prenatal to postnatal stages (n = 176). J Overlay of editing levels for T21 and control samples within BrainVar-defined developmental periods 5 and 6 (n = 67). K Z-score normalization of T21 and control editing levels relative to BrainVar (Periods 5 and 6) (n = 67) means and standard deviations for each site during matched developmental windows, shown for the PFC (left) and hippocampus (right). All boxplots in this figure represent the interquartile range (IQR), center lines indicate the median, and whiskers extend to 1.5× IQR.
We next profiled high-confidence A-to-I sites across the transcriptome using a curated database and a supervised detection pipeline, followed by stringent quality control filtering (see “Methods”). This approach yielded 19,346 sites in the PFC and 18,585 in the hippocampus, with ~80% of sites shared between regions (Fig. 5D). As expected, most editing sites were located within 3′UTRs and introns. Next, differential editing analyses were conducted between T21 and control samples, adjusting for estimated neuronal cell type proportions, RIN, gestational age, sex, and percentage of mRNA reads, within each region. We identified 39 dysregulated sites in the PFC and 151 in the hippocampus (FDR < 5%), the vast majority of which were over-edited in T21 (Fig. 5E, F, Supplemental Data 9), with predominant localization to 3′UTRs. In fact, 3′UTR sites showed enrichment among differentially edited sites in the PFC (1.20-fold enrichment; p = 0.049, one-sided FET) and hippocampus (1.56-fold enrichment; p = 1.6 × 10−⁵, one-sided FET). Notably, exonic sites showed significant enrichment for differentially edited sites in the PFC (~7.1-fold enrichment, p = 0.00064, one-sided FET) with a suggestive enrichment in the hippocampus (1.78-fold enrichment; p = 0.19, one-sided FET). Across genes, RNA editing in the 3′ UTR showed an inverse correlation with expression (Fig. S12), consistent with the idea that increased A-to-I editing destabilizes transcripts by altering structure, miRNA binding, or translational efficiency. Similar negative coupling across other genic regions suggests a pervasive link between editing activity and reduced mRNA abundance. Notably, intersecting differentially expressed genes with transcripts harboring over-edited sites revealed minimal overlap, with only one gene in the PFC (MRPL30) and one in the hippocampus (BASP1-AS1), indicating limited convergence between transcriptional and editing dysregulation.
Among the differentially edited sites, we identified seven well-characterized RNA recoding events where A-to-I editing results in nonsynonymous amino acid changes that were consistently over-edited in T21 and are known to play critical roles in early brain development (Fig. 5G, Fig. S13). Several of these sites reached statistical significance in both brain regions, while others were significant in one region and showed consistent trends in the other. For example, two recoding sites within GRIK2 (p.Q621R and p.Y571C) were significantly over-edited in both the PFC (FDR p = 0.001, FDR p = 0.01, respectively) and hippocampus (FDR p = 0.05, FDR p = 0.02, respectively). Over-editing was also observed at GRIA2 (p.R764G) and GRIA3 (p.R775G) in the PFC (FDR p = 0.0009, FDR p = 0.15, respectively) and hippocampus (FDR p = 0.07, FDR p = 0.01, respectively). Additional recoding sites showing over-editing included GABRA3 (p.I342M) and CYFIP2 (p.K294E) in the PFC (FDR p = 0.01 and 0.02) and hippocampus (FDR p = 0.02 and 0.15), and COG3 (p.I635V) in the hippocampus (FDR p = 0.03) (Fig. S13). Editing levels at these loci were strongly correlated with ADARB1 expression (Fig. 5H), consistent with its known regulatory role in mediating editing at recoding sites.
To further define the molecular identity of T21-associated editing changes, we integrated reference A-to-I editing maps from purified human brain cell types23. This analysis revealed that most over-edited sites in T21, particularly those within protein-coding regions, are strongly enriched in neurons, with minimal editing in glial populations (Fig. S14). This neuron-specific pattern argues against a secondary inflammatory or immune origin. Moreover, cell type deconvolution of bulk RNA-seq showed no significant differences in neuronal composition between T21 and control samples (Fig. S1), further excluding cell proportion as a confounding factor. Many of these neuron-enriched sites, especially recoding events, were also found to be highly edited in fresh brain tissue but markedly reduced in postmortem samples (Fig. S14), underscoring their dynamic regulation and supporting a functional role in neuronal plasticity rather than passive degradation artifacts.
These recoding sites undergo tightly regulated, site-specific changes during typical brain maturation, with editing levels gradually increasing and peaking postnatally, as shown using BrainVar typical cortical developmental data29 (Fig. 5I). To contextualize editing levels in T21, we overlaid our fetal brain data onto these normative trajectories, focusing on BrainVar samples from fetal Periods 5 and 6, which correspond to our gestational window. Across all six selected recoding sites, T21 samples consistently exhibited elevated editing relative to both age-matched controls and the BrainVar developmental distribution (Fig. 5J). To quantify these shifts relative to typical brain development, we normalized editing levels using z-scores calculated against the BrainVar reference dataset. Specifically, for each site and sample, we computed z-scores using the mean and standard deviation of editing levels from neurotypical BrainVar samples during developmental Periods 5 and 6 (see “Methods”). This normalization anchored our T21 and control measurements to a well-characterized mid-gestational baseline. Across all six sites, T21 samples exhibited consistently positive z-scores, indicating premature or excessive editing relative to normative expectations (Fig. 5K). In contrast, control samples were centered near the BrainVar developmental mean (Fig. 5K). These results suggest that T21 is associated with editing at key neurodevelopmental loci, potentially reflecting premature post-transcriptional maturation and altered proteomic remodeling during fetal brain development.
Meta-analysis of A-to-I editing across diverse tissues confirms over-editing in T21
Although A-to-I editing is known to vary across cell types, tissues, and developmental stages, we sought to validate our primary findings by performing a cross-study meta-analysis encompassing diverse cellular contexts. Using a harmonized editing quantification pipeline and consistent site detection criteria across datasets, we identified 68 high-confidence A-to-I editing sites under strict FDR < 5% and an additional 69 A-to-I sites under FDR < 10% that were over-edited in T21 compared to controls (Fig. 6A). These sites were selected from a transcriptome-wide screen and detected consistently across multiple studies. Totally, ~51% of these sites (71 sites) localized to 3′ untranslated regions (3′UTRs) (Fig. 6B), highlighting a potentially conserved role for post-transcriptional dysregulation in T21. Using a harmonized editing quantification pipeline and consistent site detection criteria across datasets, we identified 68 high-confidence A-to-I editing sites that were significantly over-edited in T21 compared to controls at a stringent false discovery rate (FDR < 5%), along with an additional 69 sites reaching FDR < 10% (Fig. 6A). To complement the high-confidence sites, we expanded the threshold to FDR < 10% to cast a broader net and detect consistent, lower-magnitude editing changes that may still be biologically relevant for T21. While more exploratory, these sites were retained only if consistently detected across datasets and offer candidates for future validation (see “Methods”). Notably, ~51% (71 sites) localized to 3′ untranslated regions (3′UTRs) (Fig. 6B), underscoring a potential conserved mechanism of post-transcriptional dysregulation in T21.
A Differential RNA editing meta-analysis comparing T21 and control samples across ten independent datasets. Each site is plotted by pooled delta editing level (x-axis) and significance −log₁₀ Benjamini–Hochberg FDR-adjusted p-value (two-sided; y-axis). Sites are colored by statistical confidence: FDR < 1% (red), 5–10% (orange), and >5% (gray). B Distribution of differentially edited sites by genic region, highlighting strong enrichment in 3′UTRs. Inset: Boxplot comparing miRNA binding affinity (minimum free energy, MFE) between unedited and edited 3′UTRs (n = 71) using high-confidence miRNA seed matches. Lower MFE values reflect stronger predicted binding; significance was assessed using a two-sided Mann–Whitney U test (exact p-value shown). C Average editing levels of all over-edited sites in T21 plotted across prenatal to postnatal stages using the BrainVar developmental dataset (n = 176). Inset: Boxplot showing mean editing levels for these sites in prenatal (n = 116) versus postnatal (n = 60) periods of typical development (p = 9.0 × 10−²⁴, Cohen’s D = 3.67). Boxplots in panel B and panel C represent the interquartile range (IQR), center lines indicate the median, and whiskers extend to 1.5× IQR. D Forest plot of GRIA3 (p.R775G) editing levels comparing T21 and controls across all datasets, along with corresponding sample sizes. The x-axis shows delta editing as a percentage with corresponding effect sizes (dots) and 95% confidence intervals (horizontal lines), and the pink diamond represents the combined random-effects model estimate.
We further modeled the impact of editing on miRNA–mRNA interactions using thermodynamic predictions (see “Methods”). Edited 3′UTRs exhibited significantly lower miRNA binding affinity than their unedited counterparts (p = 2.3 × 10−³², Mann–Whitney U test) (Fig. 6B), suggesting that over-editing in T21 may disrupt canonical miRNA regulation during critical developmental windows. While these findings are based on computational predictions, they support a mechanistic link between A-to-I editing and post-transcriptional gene silencing in the context of T21.
To further evaluate the developmental relevance of T21 over-edited sites, we examined their editing trajectories across typical cortical development. All sites exhibited increasing editing from the prenatal to postnatal period, with mean levels rising from ~35% to ~50% (Cohen’s d = 3.67, Fig. 6C), suggesting tight temporal regulation during maturation. Among the recoding sites elevated in T21, GRIA3 (p.R775G) emerged as the most consistently over-edited locus across both fetal brain and multiple in vitro models (Fig. 6D). This site, which regulates AMPA receptor desensitization kinetics, showed a statistically robust ~8% average increase in editing in T21 (RE model: p = 0.0011), with most datasets demonstrating concordant over-editing. Our findings suggest that in T21, premature attainment of these editing thresholds may disrupt synaptic refinement and shift excitatory–inhibitory (E/I) balance during critical periods of circuit formation. Moreover, the reproducibility of this editing phenotype across diverse cell types, from iPSCs and NPCs to fibroblasts and postmortem brain, underscores its biological relevance and supports a model in which ADARB1-driven editing at functionally constrained sites contributes to altered excitatory signaling in T21.
Peripheral blood RNA-seq reveals immune-linked ADAR activation in T21
To contextualize CNS-specific RNA editing dysregulation in T21, we analyzed matched bulk RNA-seq data from peripheral whole blood samples (n = 304 T21, n = 96 controls)20. ADARB1 expression was significantly elevated in T21 blood (FDR p = 3.14 × 10−⁸), consistent with gene dosage effects, whereas ADAR and ADARB2 expression were unaltered by karyotype (Fig. S15A). However, in contrast to the brain, global editing activity (AEI) was not elevated across all T21 samples (Fig. S15B). To evaluate immune-related editing effects, we stratified samples by transcriptional subtype (MS1–MS3), revealing that only MS3 individuals (characterized by high innate immune activation) exhibited increased AEI and elevated ADAR expression (Fig. S15C, D). Notably, ADARB1 expression tracked with karyotype but not immune tone and did not correlate with AEI (r = −0.0514, p = 0.24), whereas ADAR strongly correlated with AEI (r = 0.599, p = 1.07 × 10−⁵³) (Fig. S15E). These findings highlight that while ADARB1 is dosage-sensitive in both tissues, its role in RNA editing is CNS-specific, whereas ADAR mediates immune-linked editing in peripheral blood. This contrast underscores tissue-specific regulatory mechanisms and supports a dual model in which gene dosage and immune signaling independently shape A-to-I editing dynamics across biological systems.
Discussion
Despite the well-established genetic basis of DS as trisomy of chromosome 21, the mechanisms by which this chromosomal imbalance disrupts early human brain development remain poorly understood. Prior transcriptomic studies in fetal and postnatal tissues have reported widespread gene expression changes, implicating both direct dosage effects of chromosome 21 genes and secondary perturbations in neurodevelopmental, glial, and immune signaling pathways2,3,4,5,6,7. However, these studies have often been constrained by limited sample sizes, incomplete transcriptome surveys, postnatal sampling windows, or insufficient cell type resolution. To address these gaps, we integrated bulk RNA-seq data from the PFC and hippocampus of mid-gestation T21 fetal brains with a harmonized meta-analysis of nine additional independent RNA-seq datasets spanning early diverse T21 cellular contexts (Fig. 1, Table 1). Our findings highlight A-to-I editing, particularly through ADARB1 overexpression, as a conserved, dosage-sensitive mechanism of transcriptomic dysregulation in T21. By linking increased editing to both coding and regulatory RNA elements, this study identifies RNA editing as a previously underappreciated axis of gene dysregulation during fetal brain development. Notably, these effects converge across cell types, brain regions, and independent datasets, highlighting RNA editing as a cross-cutting mechanism that integrates gene dosage imbalance with altered post-transcriptional regulation. Given the timing and sensitivity of ADARB1-driven editing, these findings raise the possibility that RNA editing profiles may serve as early biomarkers of circuit maturation and suggest therapeutic entry points for restoring post-transcriptional balance in DS. Importantly, this work reframes T21 neuropathology through the lens of dynamic RNA regulation, complementing existing transcriptomic and epigenomic models with a mechanism tightly linked to developmental timing and neuronal function. To facilitate further research, all data from this study are available via an open-access, interactive online resource (see “Data availability”).
Our transcriptomic analysis revealed gene dysregulation across fetal PFC and hippocampus in T21, including a pronounced enrichment of chromosome 21 genes (Fig. 2). While previous studies reported variability in chromosome 21 dosage effects, ranging from global overexpression to partial compensation7,18,20,30,31, our data reinforce a model of selective dosage sensitivity. Specifically, chromosome 21 genes that are lowly expressed in the fetal brain are more likely to escape significant upregulation in T21, suggesting that transcriptional buffering or tighter regulatory control may limit their dosage response. This implies that gene expression changes in T21 are not uniformly driven by copy number, but instead shaped by context-dependent regulatory mechanisms, reinforcing a non-global model of gene dosage effects. Genes such as DYRK1A, IFNAR1, and IFNAR2 were consistently overexpressed, whereas others appeared transcriptionally buffered. Beyond chromosome 21, downregulated genes were enriched for mitochondrial translation and ribosomal biogenesis, indicative of impaired metabolic capacity during neurogenesis32,33. In contrast, upregulated non-chromosome 21 transcripts, including DCN and LTBP4, suggested atypical extracellular matrix remodeling and early electrophysiological maturation3,31. Permutation-based gene set preservation analysis revealed marked disruption of gene regulatory architecture in the hippocampus, particularly in modules governing synaptic plasticity, chromatin remodeling, and glial immune signaling (Fig. 3), suggesting region-specific vulnerability to dosage imbalance and its downstream effects.
To assess the generalizability of these patterns and to synthesize extant data, we performed a harmonized meta-analysis, integrating our fetal RNA-seq data with 9 independent RNA-seq datasets from T21 cases and controls (Fig. 4). This analysis yielded a robust signature of 543 upregulated and 437 downregulated genes (FDR < 1%), of which 30.7% of the upregulated genes were chromosome 21 encoded, closely mirroring the expected 1.5× gene dosage effects. However, consistent with our fetal brain findings and previous analyses, only a subset of chromosome 21 genes exhibited reproducible overexpression18,20,30,31. Among these, ADARB1 emerged as one of the most consistently upregulated genes across brain and non-brain tissues. In T21 fetal brains, ADARB1 was significantly elevated in both the PFC and hippocampus, while expression of other ADAR family members (ADAR1, ADARB2) remained unchanged. This association is robust and consistent across datasets and supported by independent A-to-I editing profiles in the human brain16,28,29,34. Our findings also support observations from previous in vitro studies using T21 iPSCs13, which noted ADARB1 dosage increases and potential impacts on editing fidelity and transcript diversity. While Kawahara et al. reported no changes in ADARB1 expression or GRIA2 Q/R site editing in one neonate DS brain tissue12, our brain-anchored data suggest that sample size, developmental timing and/or regional specificity may reveal editing dysregulation not captured in whole-tissue analyses.
Transcriptome-wide analyses revealed consistent over-editing at a subset of A-to-I sites, including seven protein-coding recoding events critical for neuronal signaling8,9 (Fig. 5, Fig. S13). These included the Q/R site in GRIA2, as well as homologous sites in GRIA3 and GRIK2, all of which undergo tightly regulated, developmentally timed editing that governs calcium permeability and receptor desensitization kinetics27,35. Among the consistently over-edited sites in T21 fetal brain, the GRIA3 R/G site (p.R775G) showed a ~8% increase in editing across brain regions and in vitro models (Fig. 6). Although editing at this site is conserved and has been implicated in regulating AMPA receptor desensitization kinetics36,37, its in vivo functional significance remains incompletely characterized. In contrast to the well-studied GRIA2 Q/R site, whose loss of editing accounts for the severe phenotypes in Adar2 knockout mice38,39,40, there is currently no direct evidence linking GRIA3 R/G editing to developmental phenotypes in vivo. However, increased GRIA3 editing has been observed in the context of epilepsy41,42, suggesting that it may respond to or modulate excitatory signaling. Furthermore, ADAR2 overexpression in rodents has been associated with behavioral phenotypes such as hyperphagia43, although these phenotypes have not been causally linked to GRIA3 editing or to T21. While we observe a robust and premature shift in editing at GRIA3 R/G and other recoding sites, we interpret these findings as correlative with ADARB1 upregulation rather than directly causal. Future site-directed editing studies will be needed to determine whether specific recoding changes contribute to altered neurodevelopment in T21.
Additional over-edited transcripts included GABRA3 (I342M), which modulates GABA_A receptor gating and trafficking, and CYFIP2 (K294E), a synaptic scaffolding protein involved in dendritic spine remodeling and local translation44. These alterations likely compound the effects of glutamate receptor dysregulation and disrupt synaptogenesis. Supporting this view, DS-derived astrocytes have been shown to impair synapse formation and induce mTOR hyperactivation in co-cultured neurons, a phenotype linked to E/I imbalance and reduced connectivity45. This is consistent with recent findings demonstrating that T21 iPSC-derived astrocytes secrete factors that impair neuronal mTOR signaling and reduce synaptic density in vitro46. These non-cell-autonomous effects may act synergistically with post-transcriptional disruptions, such as premature RNA editing, to alter receptor composition and shift the developmental timing of synaptic integration. Moreover, these recoding sites are also among the most highly edited in living (antemortem) human brain tissue but are markedly less edited in postmortem samples (Fig. S14)23, highlighting their reliance on intact cellular physiology. Their premature over-editing in T21 may therefore reflect a shift in the temporal regulation of editing normally restricted to viable, active neural circuits, further supporting their functional relevance during early synaptogenesis
To contextualize the CNS-specificity of our findings, we extended our analysis to peripheral blood RNA-seq data from individuals with T21 and controls20. ADARB1 expression was significantly elevated in T21 blood samples, consistent with gene dosage effects, yet global editing activity (AEI) was not increased (Fig. S15). However, when stratified by transcriptional subtype, only individuals with high interferon signaling (MS3 group) exhibited elevated AEI alongside increased ADAR expression. These observations underscore canonical immune-linked editing dynamics in blood, in contrast to the fetal brain, where AEI was driven by ADARB1 expression, independent of immune activation or ADAR levels. This tissue-specific divergence may reflect ADAR1 isoform usage, as the p150 isoform that most strongly correlates with global editing predominates in peripheral tissues, whereas the p110 isoform is dominant in the CNS and is less tightly coupled to AEI47. This divergence highlights a tissue-specific regulatory axis in T21: ADARB1 predominantly governs CNS editing, consistent with our prior reports16,28,29,34, while ADAR responds to inflammatory signaling in immune tissue21,22. Notably, we also observed a positive correlation between ADARB2 and AEI in brain tissue. Despite its lack of catalytic activity, ADARB2 may act as a dominant-negative modulator or reflect co-regulated expression patterns with ADARB1 during development. Together, these findings support a dual regulatory model of RNA editing in T21, shaped by both gene dosage and immune signaling, and emphasize the importance of tissue context in interpreting A-to-I editing dynamics.
Several limitations of this study should be acknowledged. First, while bulk RNA-seq offers robust insight into global transcriptomic changes, it lacks the resolution to resolve cell–type–specific editing patterns. Future studies leveraging single-nucleus or spatial transcriptomics will be essential to map editing dynamics across distinct neuronal and glial populations. Second, although our meta-analysis employed a harmonized processing pipeline, heterogeneity in source datasets, including differences in cell type composition, developmental stage, and technical protocols, may have reduced sensitivity for detecting low-abundance or cell type-specific editing events. To mitigate this, we prioritized the identification of T21-associated editing changes that were consistently observed across diverse cellular contexts. Finally, while the observed association between ADARB1 and aberrant editing is compelling, causal relationships remain to be established. For example, we did not perform functional assays to test the impact of individual recoding sites such as GRIA3 R/G. Experimental perturbation of individual recoding events, such as those affecting GRIA3 or 3′UTR-miRNA interactions, is required to determine their precise contribution to T21-associated phenotypes.
In sum, this study identifies RNA editing as a dosage-sensitive layer of transcriptomic dysregulation in T21, driven in part by the overexpression of ADARB1. This finding extends prior work on the dynamic regulation of A-to-I editing across brain development, where precise timing and spatial control are essential for synaptic homeostasis and neuronal plasticity. Here, we show that ADARB1-driven editing dysregulation emerges earlier than previously recognized, impacting a distinct set of neurodevelopmental targets. By integrating fetal brain sequencing, gene co-expression analysis, and cross-system meta-analysis, we define a reproducible molecular signature linking chromosome 21 dosage to post-transcriptional disruption. These findings position RNA editing as a regulatory amplifier of T21’s molecular effects, potentially exacerbating synaptic dysfunction. As such, RNA editing signatures may serve as biomarkers of circuit maturation and offer entry points for correcting post-transcriptional imbalance in DS.
Methods
Postmortem tissue acquisition and DNA testing
All tissue samples were collected at the University Hospital of Obstetrics and Gynecology, Sofia, in collaboration between the Lieber Institute for Brain Development (LIBD) and the Medical University of Sofia, Bulgaria. The postmortem brain tissues were collected after written informed consent was obtained from the mothers and stored fresh frozen at the biobank of the Molecular Medicine Center, Medical University of Sofia, before transport to LIBD for further processing. Altogether, fetal brain tissues from 20 T21 cases (14 male, 6 female) and 27 euploid controls (17 male, 10 female) between 13–22 gestational weeks were used in the study. All 20 T21 cases analyzed in this study were confirmed as full trisomy 21 by clinical karyotyping reports obtained from the source biobank. No partial or mosaic cases were included. The fetal material was collected after termination of pregnancy for medical reasons (severe congenital anomalies or genetic disorder). Invasive prenatal aneuploidy testing using Aneufast™ QF-PCR Kit (Molgentix Sl) was performed at the University Hospital of Obstetrics and Gynecology, and microarray testing of DNA from the abortive tissue was done using SurePrint G3 Human CGH Microarray Kit, 4 × 180 K (Agilent Technologies Inc.) at the Molecular Medicine Center.
All research involving human fetal tissue was conducted in accordance with institutional and national ethical guidelines and regulations. Written informed consent was obtained from all donors prior to tissue collection. No embryos or gametes were created or used for research purposes. The research project “Genetic and epigenetic research of human fetal brain development” was reviewed and approved by the Institutional Ethics Review Board of the Medical University of Sofia (Protocol No. 2981/06.07.2012).
Portions of the fetal brain tissue samples were consumed during RNA extraction and downstream molecular analyses. Any remaining unused tissue material is conserved at the biobank of the Molecular Medicine Center, Medical University of Sofia. Due to ethical and regulatory constraints associated with human fetal tissue, remaining biological material is not broadly distributable; however, access requests may be considered on a case-by-case basis, subject to local ethical approval and material transfer agreements. Inquiries regarding potential access to remaining tissue material should be directed to the Molecular Medicine Center, Medical University of Sofia, in coordination with the Lieber Institute for Brain Development.
RNA-sequencing data generation
We assembled a discovery cohort consisting of paired prefrontal cortex (PFC) and hippocampal formation tissue samples (referred to as “hippocampus” throughout this body of work) from 20 individuals with trisomy 21 (T21) and 27 euploid controls, collected between 13 and 22 post-conception weeks (PCW), a critical period for human brain development. At these early developmental stages, the hippocampal formation represents the broader precursor region giving rise to the mature hippocampus. Donor metadata are detailed in Supplemental Data 1. The cohort included 16 females and 31 males, with a mean gestational age of 18.4 ± 3.2 weeks. All samples were from individuals of Caucasian ancestry with no reported major brain malformations. Total RNA was extracted from frozen tissue using the AllPrep DNA/RNA Mini Kit (Qiagen, Cat No. 80204). RNA quality was assessed using the Agilent 2100 Bioanalyzer; all included samples had RNA Integrity Numbers (RIN) ≥ 9.0 (9.8 ± 0.3). RNA-seq libraries were prepared from 300 ng of total RNA using the TruSeq Stranded Total RNA Library Prep Kit with Ribo-Zero Gold (Illumina), which depletes both cytoplasmic and mitochondrial ribosomal RNA. Libraries were sequenced on an Illumina HiSeq 3000 platform at the Promega sequencing core, generating 200-bp paired-end reads at an average depth of ~34.5 million reads per sample.
RNA-sequencing data processing and quality control
Sequencing reads were processed using the RAPiD-nf pipeline (https://github.com/CommonMindConsortium/RAPiD-nf). Reads were aligned to the GRCh38 human reference genome (Ensembl release 95) using STAR v2.7.3a48. Alignment quality was assessed using RSeQC and Picard tools, including metrics for GC content, duplication rates, insert size, gene body coverage, and library complexity. PCR duplicates were marked but retained during quantification. Gene-level quantification was performed using featureCounts49 (Subread v2.0.1) based on GENCODE v32 annotations. Genes with non-zero counts in at least one-third of all samples were retained. Counts were normalized using the VOOM transformation from the limma package (v3.48.3)50 to form region-specific expression matrices (PFC and hippocampus). A total of 16,472 genes were analyzed in the PFC, and 17,035 genes were analyzed in the hippocampus. Principal component analysis (PCA) did not identify any expression outliers using a 95% confidence interval across the top three PCs.
Cellular deconvolution of bulk tissues
Cell-type composition was estimated using non-negative least squares (NNLS) implemented in the est_frac function of the bMIND R package (v1.2.0)51. This approach applies Bayesian inference to estimate cell-type–specific expression profiles while constraining coefficients to non-negative fractions. Prior information for the model was derived from a reference single-cell expression signature matrix from Pei et al.52. This matrix includes transcriptional profiles for 11 fetal brain cell types: astrocytes, microglia, oligodendrocytes, oligodendrocyte precursor cells (OPCs), endothelial cells, pericytes, intermediate progenitor cells (IPCs), neuroepithelial cells (NEPs), and three neuronal subtypes (quiescent, replicating, transitional). Log2-transformed counts per million (CPM) were used as input53. Deconvolution results were used to adjust for cellular heterogeneity in downstream modeling. This approach enabled us to assess and adjust for potential shifts in cell type composition while minimizing confounding due to cellular heterogeneity, consistent with prior studies of fetal brain transcriptomics.
Partitioning gene expression variance
We used the variancePartition R package54 to fit linear mixed models quantifying the proportion of expression variance attributable to technical and biological factors. Covariates included neuronal proportion (quiescent + replicating + transitional neurons), % mRNA reads, RIN, gestational age, sex, library depth, and genetic diagnosis. Donor identity was modeled as a random effect. These estimates informed covariate selection for differential expression and network modeling.
Differential gene expression analysis
Differential expression was performed using the limma-voom pipeline53. An initial analysis compared PFC versus hippocampus, adjusting for diagnosis, percentage of mRNA reads, gestational age, sex, and RIN, with donor modeled via duplicateCorrelation. T21-specific effects were then assessed separately for each region using region-specific VOOM-normalized matrices (16,471 genes in PFC; 17,035 in hippocampus). Models included covariates for neuronal proportion, percentage of mRNA reads, age, sex, and RIN. DEGs were defined as those with Benjamini-Hochberg FDR < 0.05.
Weighted Gene Co-expression Network Analysis (WGCNA)
To identify coordinated transcriptional programs disrupted in T21, we performed weighted gene co-expression network analysis (WGCNA)55 separately for the PFC and hippocampus using VOOM-normalized expression matrices. A total of 16,471 genes (PFC) and 17,035 genes (hippocampus) were retained. Signed co-expression networks were constructed by computing absolute Pearson correlation coefficients between all gene pairs, followed by soft-thresholding to approximate scale-free topology (target R² > 0.80). Power parameters (β) of 5 and 9 were selected for the PFC and hippocampus, respectively, based on scale-free fit and mean connectivity criteria. Networks were clustered using the dynamic tree cut algorithm with a minimum module size of 50 and a cut height of 0.999. Modules were summarized by their eigengenes (first principal component of each module’s expression matrix) and tested for association with T21 diagnosis using linear models in limma, adjusting for the same covariates as in the differential gene expression analyses (neuronal proportion, % mRNA, age, sex, and RIN). Module significance was assessed via moderated t-tests. Qualitative inspection of module eigengene expression across diagnostic groups was used to confirm biologically interpretable patterns. These analyses enabled the identification of gene networks associated with T21 that may not be captured through single-gene differential expression alone.
Gene Ontology and enrichment analysis
To interpret the functional significance of differentially expressed genes, we performed gene set enrichment analysis using the CAMERA method56, a correlation-adjusted competitive test that accounts for inter-gene correlation within gene sets. Differential expression t-statistics were input from the limma-based comparisons between T21 and control samples for each brain region. CAMERA ranks genes based on differential expression and tests whether predefined gene sets are overrepresented at the extremes of this ranked list relative to random expectation. Gene sets were derived from Gene Ontology (GO) biological processes, molecular functions, and MSigDB hallmark gene sets. Variance inflation due to inter-gene correlation was corrected using the default inter-gene correlation parameter (set to 0.01), and resulting enrichment p-values were adjusted for multiple testing using the Benjamini–Hochberg method. Enrichment results were used to identify biological processes and pathways disproportionately affected in T21 fetal brain tissues.
Permutation-based gene set preservation in T21
We assessed whether co-regulatory structures within predefined gene sets were disrupted in T21 using the Zsummary preservation statistic from WGCNA57, computed with 200 permutations. This statistic integrates measures of intramodular density and connectivity across conditions. Zsummary scores <2 indicate no preservation, 2–10 suggest weak to moderate preservation, and >10 reflect strong preservation. Because unsupervised gene co-expression network structure appeared preserved between T21 and controls, we focused on smaller curated gene sets to detect more targeted disruptions, including GO biological processes, GO molecular functions, and MSigDB hallmark gene sets. To correct for the known dependency of preservation scores on module size, we modeled Zsummary of a random geneset matched for geneset size, as a comparative for each tested geneset, as a function of module size and derived an expected preservation score (Zrandom_predicted) for each gene set. We then computed an adjusted rank score incorporating (1) the observed Zsummary, (2) its FDR-adjusted p-value, and (3) the deviation of random module preservation from expectation. Gene sets meeting strict criteria (Zsummary<2, p.adj > 0.1, Zsummary ≤ Zrandom_predicted− 3, and p.random.adj <0.05) were classified as non-preserved. These modules exhibited significantly lower preservation than size-matched random sets (Wilcoxon p < 0.05, PFC and hippocampus), supporting loss of co-regulatory structure in biological pathways.
Cross-Study Meta-Analysis of RNA-Seq Datasets
To validate and generalize transcriptomic findings across diverse developmental contexts, we performed a harmonized meta-analysis of our data with nine additional publicly available RNA-seq datasets profiling T21 and euploid control samples (Table 1). These datasets spanned a range of early gestational tissues and in vitro cell types, including induced pluripotent stem cells (iPSCs), neural progenitor cells (NPCs), cortical neurons, fetal fibroblasts and immune cells. In total, the meta-analysis encompassed 108 T21 and 117 control samples across the following GEO accession numbers: GSE52249, GSE185192, PRJNA751601, GSE101942, GSE144857, GSE55504, GSE79842, GSE84531, and GSE128621. For each dataset, raw FASTQ files were reprocessed through a standardized RNA-seq pipeline (RAPiD-nf; https://github.com/CommonMindConsortium/RAPiD-nf) to ensure uniformity in alignment, quantification, and normalization. Reads were aligned to the human genome (hg38), and gene-level expression was quantified using featureCounts49. Differential gene expression within each dataset was computed using limma with voom transformation44, adjusting for relevant covariates such as age, sex, batch, and neuronal fraction where available.
For each gene, log₂ fold-change and standard error estimates were extracted from limma-voom analyses of each dataset. Genes with low expression (not expressed in at least one-third of samples per dataset) were excluded. Meta-analysis was performed using a random-effects model implemented via the rma.uni() function in the metafor R package (REML estimator) was used to account for between-study heterogeneity58. Only genes expressed in ≥4 datasets and with consistent directionality in at least 70% of datasets were retained in the final result set. Genes with Benjamini–Hochberg adjusted p-values < 0.05 were considered significantly dysregulated. Meta-analysis summary statistics, including forest plots for key dosage-sensitive genes, are publicly available (https://andyyang.shinyapps.io/Ts21-dashboard/). This approach enabled robust quantification of consistent transcriptional changes across diverse developmental contexts and experimental platforms.
T21 microarray data reprocessing and differential expression analysis
To evaluate the comparability of our RNA-seq findings with prior transcriptomic data, we reprocessed and analyzed a publicly available fetal brain microarray dataset profiling Trisomy 21 (T21) and control samples (GEO accession: GSE59630)3. This dataset was generated using the Affymetrix Human Exon 1.0 ST Array platform and includes 58 paired tissue samples from 15 T21 and 15 euploid control brains, ranging in age from 14 post-conception weeks (PCW) to 42 years old. For consistency with our fetal RNA-seq dataset, we focused our analysis on the cerebral cortex and hippocampus, corresponding to the neocortical (CBC) and hippocampal (HIP) samples in the original study.
Raw.CEL files were downloaded and imported using the oligo package in R59. Robust Multi-array Average (RMA) normalization60 was applied to compute expression values at the exon level, followed by log₂ transformation. Metadata for all samples was manually curated and aligned to expression profiles, with key covariates including sex, post-mortem interval (PMI), RNA integrity number (RIN), ethnicity, anatomical region, and developmental stage (converted to numeric age in years, accounting for fetal and postnatal time points). Genes were filtered to retain those expressed above the sample-wise median in at least one-third of samples, resulting in a matrix of robustly expressed transcripts. Gene expression variance was decomposed using the variancePartition R package54, employing a linear mixed model that included fixed effects for age, RIN, PMI, and developmental period, and random intercepts for sex, ethnicity, and diagnosis. Differential expression was assessed using the limma framework. A design matrix was constructed to model expression as a function of T21 diagnosis, RIN, PMI, age (in years), and developmental period. Duplicate samples per subject were accounted for by controlling for correlated measurements. A contrast was specified to test for differential expression between DS and control samples. Empirical Bayes moderation was applied to stabilize variance estimates. Significance was defined using a Benjamini-Hochberg false discovery rate (FDR) threshold of <1%.
Fold-changes were directionally aligned to match RNA-seq comparisons, with log₂ fold-changes sign-inverted such that positive values indicate higher expression in T21. Differentially expressed genes (DEGs) were classified as upregulated in T21, upregulated in controls, or non-significant. The resulting gene-level summary statistics were exported for comparison with RNA-seq–derived DEG lists.
RNA editing site detection and annotation
To quantify RNA editing across fetal brain samples, we employed a supervised approach that focused on known A-to-I editing sites. Specifically, we queried nucleotide positions curated from high-confidence databases: (i) REDIportal61, (ii) previously annotated sites from human brain cell types28, and (iii) a comprehensive list of evolutionarily conserved A-to-I recoding sites62. Using samtools mpileup, we extracted reference and edited read counts for these known sites directly from STAR-aligned BAM files, following our previously validated pipelines16,28,29,34. We then applied stringent quality control criteria to ensure the inclusion of bona fide A-to-I editing events. Supervised sites with <5 total reads or <3 edited reads were excluded. The resulting set of high-confidence editing sites was functionally annotated using ANNOVAR63, assigning gene symbols based on RefGene, repetitive elements based on RepeatMasker v4.1.164, and known editing annotations via the latest REDIportal database. Evolutionary conservation scores were incorporated using phastCons scores from the PHAST package65 to highlight conserved editing events across mammals.
Alu Editing Index (AEI)
To measure global RNA editing activity, we calculated the Alu Editing Index (AEI)66, a quantitative metric reflecting A-to-I editing levels in repetitive SINE/Alu elements, where the majority of non-coding editing occurs. The AEI was computed using AEI v1.0, which takes STAR-aligned BAM files as input. Specifically, the AEI is calculated as the percentage of A-to-G mismatches relative to total adenosine coverage within annotated Alu elements (from UCSC RepeatMasker annotations, hg38). Known polymorphic sites were masked using coordinates from the UCSC CommonGenomicSNPs150 track to reduce false positives due to DNA variation. The AEI method preserves signal even in low-coverage regions and has demonstrated robustness across hundreds of postmortem human brain RNA-seq samples generated with varied protocols. AEI values were multiplied by 100 to report editing levels as percentages. This metric served as a transcriptome-wide surrogate for ADAR-mediated editing activity and was used to assess global dysregulation in T21 across brain regions and external replication datasets.
Quantifying and normalizing RNA recoding dynamics in typical and trisomy 21 fetal brain
To assess RNA recoding dynamics during fetal brain development and in trisomy 21 (T21), we analyzed A-to-I editing levels at canonical recoding sites across two datasets: (1) neurotypical mid-fetal cortical samples (Periods 5 and 6) from the BrainVar project chromosome 21, and (2) fetal PFC samples from individuals with T21 and matched controls (this study). Editing levels were computed as the ratio of guanosine (G) reads to total adenosine + guanosine (A + G) reads at each site, based on samtools pileups. To contextualize editing disruptions in T21, we visualized recoding site editing levels in T21 and control samples against the backdrop of typical developmental trajectories. Specifically, for each recoding site, we generated boxplots using BrainVar Period 5–6 PFC samples (n = 67) and overlaid individual T21 (n = 20) and euploid control (n = 27) fetal brain samples as points. This approach allowed for direct visual comparison of editing levels relative to normative developmental distributions.
To formally quantify deviation from typical editing patterns, we computed z-scores for each recoding site and individual sample. Z-scores were calculated using the mean (μ) and standard deviation (σ) of editing levels in typical brain development (TBD) BrainVar samples (Periods 5–6, PFC only), according to:
This yielded a normalized deviation score for each site per sample, allowing quantitative comparison across groups. Z-scores were visualized separately by diagnosis (T21 vs. control) and brain region. Group differences in z-scores were assessed using two-sided Wilcoxon rank-sum tests. To account for multiple comparisons across recoding sites, p-values were adjusted using the Benjamini-Hochberg procedure (FDR < 0.05 threshold). This approach enabled the identification of recoding events where editing in T21 significantly deviated from neurotypical developmental trajectories.
RNA editing meta-analysis across developmental datasets
For each editing site, the mean difference in editing rate (T21 – control) and corresponding standard error were computed from each dataset using linear models adjusted for relevant covariates when provided in the corresponding metadata files (e.g., age, batch, and RNA quality). Sites with low coverage (<10 total reads in at least one-third of samples) were excluded to ensure reliability. Editing site meta-analysis was performed using a random-effects model implemented via the rma.uni() function in the metafor R package, using the restricted maximum likelihood (REML) estimator to account for between-study heterogeneity. Only sites detected in ≥4 datasets and exhibiting consistent directionality of effect (same sign in at least 70% of datasets) were retained. Sites with Benjamini–Hochberg adjusted p-values < 0.05 were considered significantly differentially edited. This analysis identified high-confidence A-to-I sites consistently over-edited in T21 across fetal brain and in vitro models.
miRNA–mRNA interactions
miRNA-mRNA interactions were studied using miRANDA67 as previously described29,68. In brief, to assess the impact of RNA editing on miRNA binding, we compared the minimum free energy (ΔG) of miRNA-3′UTR interactions at edited versus unedited sites. Next, we extracted matched 101 bp sequences surrounding each editing site (with either A or G), aligned them to all mature miRNAs using miRANDA, and calculated ΔG for each interaction. focusing especially on seed region matches using strict parameters. Statistical differences in ΔG were evaluated using the Mann–Whitney U test.
Secondary T21 peripheral blood RNA-sequencing data analysis
To examine ADAR expression and A-to-I RNA editing profiles in peripheral blood, we analyzed bulk RNA-seq data from 304 individuals with T21 and 96 euploid controls obtained from GEO (GSE190125)20. Raw FASTQ files were processed using the same standardized pipeline as for the fetal brain and in vitro datasets to ensure analytic consistency. Reads were aligned to the GRCh38 reference genome using STAR48, and gene-level expression quantification was performed with featureCounts49. ADAR, ADARB1, and ADARB2 expression levels were extracted from VOOM normalized44 values for downstream analysis. The AEI66 was calculated for each sample as a transcriptome-wide proxy for global editing activity. To investigate subgroup-specific effects, we stratified individuals by previously defined transcriptional subtypes from the original study (e.g., MS1, MS2, and MS3). Differences in gene expression between T21 and controls, and among molecular subgroups, were performed using limma. All analyses were conducted in R (v4.2.2).
Representative brain diagrams
Brain diagrams found in Fig. 1 and Fig. 2A, B were constructed in the cerebroViz R package (v1)69.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-sequencing data generated in this study have been deposited in the NCBI Gene Expression Omnibus (GEO) database under accession code GSE301886. These data are publicly available without restriction. The GEO accession includes raw sequencing data (FASTQ files), processed gene-level expression matrices, RNA editing summaries, and associated sample metadata required to reproduce all analyses reported in this study. RNA-sequencing datasets used for cross-study gene expression and RNA editing meta-analyses are also publicly available from GEO or the European Nucleotide Archive under accession codes GSE52249, GSE185192, PRJNA751601, GSE101942, GSE144857, GSE55504, GSE79842, GSE84531, GSE128621, and GSE190125. All source data underlying the main and Supplementary Figs., including differential expression statistics, RNA editing results, and meta-analysis summary statistics, are provided in the Supplementary Data files and Source Data accompanying this paper. To enable gene-by-gene inspection of between-study effect sizes and heterogeneity, we developed an interactive RShiny application (https://andyyang.shinyapps.io/Ts21-dashboard/) that generates meta-analytic forest plots for any gene of interest across the harmonized datasets. For each gene, the application displays study-specific log fold changes, standard errors, standardized mean differences, confidence intervals, and random-effects model estimates, as well as expression boxplots and developmental expression trajectories. This RShiny application also hosts all processed gene-level expression matrices, associated sample metadata, and summary files generated in the current study. Source data are provided with this paper.
Code availability
Computational code is provided on GitHub (https://github.com/BreenMS/Trisomy21) and includes R code for: RNA-sequencing quality control, short-read mapping, counting, and normalization, differential gene expression, cell type estimation, A-to-I editing quantification, differential A-to-I editing comparisons, and meta-analyses. Contact the lead contact (M.S.B) for any questions.
References
Antonarakis, S. E. et al. Down syndrome. Nat. Rev. Dis. Prim. 6, 9 (2020).
Klein, J. A. et al. Sonic hedgehog pathway modulation normalizes expression of Olig2 in rostrally patterned NPCs with trisomy 21. Front. Cell. Neurosci. 15, 794675 (2022).
Olmos-Serrano, J. L. et al. Down syndrome developmental brain transcriptome reveals defective oligodendrocyte differentiation and myelination. Neuron. 89, 1208–1222 (2016).
Palmer, C. R., Liu, C. S., Romanow, W. J., Lee, M. H. & Chun, J. Altered cell and RNA isoform diversity in aging Down syndrome brains. Proc. Natl. Acad. Sci. USA 118, e2114326118 (2021).
Head, E. et al. Aging in Down syndrome and the development of Alzheimer’s disease neuropathology. Curr. Alzheimer Res. 13, 18–29 (2016).
Lott, I. T. & Head, E. Down syndrome and Alzheimer’s disease: a link between development and aging. Ment. Rretard. Dev. Disabil. Res. Rev. 7, 172–178 (2001).
Lockstone, H. et al. Gene expression profiling in the adult Down syndrome brain. Genomics. 90, 647–660 (2007).
Rosenthal, J. J. & Seeburg, P. H. A-to-I RNA editing: effects on proteins key to neural excitability. Neuron. 74, 432–439 (2012).
Behm, M. & Öhman, M. RNA editing: a contributor to neuronal dynamics in the mammalian brain. Trends Genet. 32, 165–175 (2016).
Susco, S. G. et al. Molecular convergence between Down syndrome and fragile X syndrome identified using human pluripotent stem cell models. Cell Rep. 40, 111312 (2022).
Ponroy Bally, B. et al. Human iPSC-derived Down syndrome astrocytes display genome-wide perturbations in gene expression, an altered adhesion profile, and increased cellular dynamics. Hum. Mol. Genet. 29, 785–802 (2020).
Kawahara, Y. et al. Regulation of glutamate receptor RNA editing and ADAR mRNA expression in developing human normal and Down’s syndrome brains. Dev. Brain Res. 148, 151–155 (2004).
Gonzales, P. K. et al. Transcriptome analysis of genetically matched human induced pluripotent stem cells disomic or trisomic for chromosome 21. PloS ONE 13, e0194581 (2018).
Ma, Y. et al. Atlas of RNA editing events affecting protein expression in aged and Alzheimer’s disease human brain tissue. Nat. Commun. 12, 7035 (2021).
Tran, S. S. et al. Widespread RNA editing dysregulation in brains from autistic individuals. Nat. Neurosci. 22, 25–36 (2019).
Breen, M. S. et al. Global landscape and genetic regulation of RNA editing in cortical samples from individuals with schizophrenia. Nat. Neurosci. 22, 1402–1412 (2019).
Waugh, K. A. et al. Mass cytometry reveals global immune remodeling with multi-lineage hypersensitivity to type I interferon in Down syndrome. Cell Rep. 29, 1893–1908 (2019).
Sullivan, K. D. et al. Trisomy 21 consistently activates the interferon response. elife 5, e16220 (2016).
Araya, P. et al. Trisomy 21 dysregulates T cell lineages toward an autoimmunity-prone state associated with interferon hyperactivity. Proc. Natl. Acad. Sci. USA 116, 24231–24241 (2019).
Donovan, M. G. et al. Variegated overexpression of chromosome 21 genes reveals molecular and immune subtypes of Down syndrome. Nat. Commun. 15, 5473 (2024).
Eisenberg, E. & Levanon, E. Y. A-to-I RNA editing—immune protector and transcriptome diversifier. Nat. Rev. Genet. 19, 473–490 (2018).
Hu, S. B. & Li, J. B. RNA editing and immune control: from mechanism to therapy. Curr. Opin. Genet. Dev. 86, 102195 (2024).
Rodriguez de los Santos, M. et al. Divergent landscapes of A-to-I editing in postmortem and living human brain. Nat. Commun. 15, 5366 (2024).
Mao, R., Zielke, C. L., Zielke, H. R. & Pevsner, J. Global up-regulation of chromosome 21 gene expression in the developing Down syndrome brain. Genomics 81, 457–467 (2003).
Nishikura, K. A-to-I editing of coding and non-coding RNAs by ADARs. Nat. Rev. Mol. Cell Biol. 17, 83–96 (2016).
Patterson, J. B. & Samuel, C. E. Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: evidence for two forms of the deaminase. Mol. Cell. Biol. 15, 5376–5388 (1995).
Wahlstedt, H., Daniel, C., Enstero, M. & Öhman, M. Large-scale mRNA sequencing determines global regulation of RNA editing during brain development. Genome Res. 19, 978–986 (2009).
Chen, C. X. et al. A third member of the RNA-specific adenosine deaminase gene family, ADAR3, contains both single- and double-stranded RNA binding domains. RNA. 6, 755–767 (2000).
Cuddleston, W. H. et al. Spatiotemporal and genetic regulation of A-to-I editing throughout human brain development. Cell Rep. 41, 111784 (2022).
Hunter, S., Hendrix, J., Freeman, J., Dowell, R. D. & Allen, M. A. Transcription dosage compensation does not occur in Down syndrome. BMC Biol. 21, 228 (2023).
Letourneau, A. et al. Domains of genome-wide gene expression dysregulation in Down’s syndrome. Nature 508, 345–350 (2014).
Busciglio, J. & Yankner, B. A. Apoptosis and increased generation of reactive oxygen species in Down’s syndrome neurons in vitro. Nature 378, 776–779 (1995).
Helguera, P. et al. Adaptive downregulation of mitochondrial function in Down syndrome. Cell Metab. 17, 132–140 (2013).
Cuddleston, W. H. et al. Cellular and genetic drivers of RNA editing variation in the human brain. Nat. Commun. 13, 2997 (2022).
Lomeli, H. et al. Control of kinetic properties of AMPA receptor channels by nuclear RNA editing. Science 266, 1709–1713 (1994).
Krestel, H. E. et al. A genetic switch for epilepsy in adult mice. J. Neurosci. 24, 10568–10578 (2004).
Tanaka, H., Grooms, S. Y., Bennett, M. V. & Zukin, R. S. The AMPAR subunit GluR2: still front and center-stage. Brain Res. 886, 190–207 (2000).
Horsch, M. et al. Requirement of the RNA-editing enzyme ADAR2 for normal physiology in mice. J. Biol. Chem. 286, 18614–18622 (2011).
Jain, M. et al. RNA editing of Filamin A pre-mRNA regulates vascular contraction and diastolic blood pressure. EMBO J. 37, e94813 (2018).
Vissel, B. et al. The role of RNA editing of kainate receptors in synaptic plasticity and seizures. Neuron 29, 217–227 (2001).
Vollmar, W. et al. RNA editing (R/G site) and flip–flop splicing of the AMPA receptor subunit GluR2 in nervous tissue of epilepsy patients. Neurobiol. Dis. 15, 371–379 (2004).
Caracciolo, L. et al. Altered mRNA editing and expression of ionotropic glutamate receptors after kainic acid exposure in cyclooxygenase-2 deficient mice. PLoS ONE 6, e19398 (2011).
Singh, M. et al. Hyperphagia-mediated obesity in transgenic mice misexpressing the RNA-editing enzyme ADAR2. J. Biol. Chem. 282, 22448–22459 (2007).
Costa Cruz, P. H. & Kawahara, Y. RNA editing in neurological and neurodegenerative disorders. RNA Edit. 2181, 309–330 (2020).
Sharma, S. D. et al. Astrocytes mediate cell non-autonomous correction of aberrant firing in human FXS neurons. Cell Rep. 42, 112019 (2023).
Araujo, B. H. et al. Down syndrome iPSC-derived astrocytes impair neuronal synaptogenesis and the mTOR pathway in vitro. Mol. Neurobiol. 55, 5962–5975 (2018).
Samuel, C. E. Adenosine deaminase acting on RNA (ADAR1), a suppressor of double-stranded RNA–triggered innate immune responses. J. Biol. Chem. 294, 1710–1720 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013). Jan.
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, 1–7 (2014).
Wang, J., Roeder, K. & Devlin, B. Bayesian estimation of cell type–specific gene expression with prior derived from single-cell data. Genome Res. 31, 1807–1818 (2021).
Pei, G. et al. Gene expression imputation and cell-type deconvolution in human brain with spatiotemporal precision and its implications for brain-related disorders. Genome Res. 31, 146–158 (2021).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Hoffman, G. E. & Schadt, E. E. variancePartition: interpreting drivers of variation in complex gene expression studies. BMC Bioinforma. 17, 1–3 (2016).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genetics Mol. Biol. 4, 17 (2005).
Wu, D. & Smyth, G. K. Camera: a competitive gene set test accounting for inter-gene correlation. Nucleic Acids Res. 40, e133 (2012). Sep 1.
Langfelder, P., Luo, R., Oldham, M. C. & Horvath, S. Is my network module preserved and reproducible? PLoS Comput. Biol. 7, e1001057 (2011). Jan 20.
Viechtbauer, W. Conducting meta-analyses in R with the metafor package. J. Stat. Softw. 36, 1–48 (2010).
Carvalho, B. et al. Package ‘oligo’ (2013).
Kim, C. S., Hwang, S. & Zhang, S. D. RMA with quantile normalization mixes biological signals between different sample groups in microarray data analysis. In 2014 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), pp 139–143. (IEEE, 2014).
Picardi, E., D’Erchia, A. M., Lo Giudice, C. & Pesole, G. REDIportal: a comprehensive database of A-to-I RNA editing events in humans. Nucleic Acids Res. 45, D750–D757 (2017).
Gabay, O. et al. Landscape of adenosine-to-inosine RNA recoding across human tissues. Nat. Commun. 13, 1184 (2022).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164–e164 (2010).
Chen, N. Using Repeat Masker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinforma. 5, 4–10 (2004).
Hubisz, M. J., Pollard, K. S. & Siepel, A. PHAST and RPHAST: phylogenetic analysis with space/time models. Brief. Bioinforma. 12, 41–51 (2011).
Roth, S. H., Levanon, E. Y. & Eisenberg, E. Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity. Nat. Methods 16, 1131–1138 (2019).
Betel, D., Koppal, A., Agius, P., Sander, C. & Leslie, C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol. 11, 1–14 (2010).
Hwang, T. et al. Dynamic regulation of RNA editing in human brain development and disease. Nat. Neurosci. 19, 1093–1099 (2016).
Bahl, E., Koomar, T. & Michaelson, J. J. cerebroViz: an R package for anatomical visualization of spatiotemporal brain data. Bioinformatics 33, 762–763 (2017).
Acknowledgements
We sincerely thank the families of the brain donors for their invaluable contributions to this research. Without their generosity and willingness to support scientific discovery, this work would not have been possible. We also acknowledge the Lieber Institute for Brain Development Human Brain Repository and collaborating institutions for providing access to these precious samples. This work was supported by the Beatrice and Samuel A. Seaver Foundation. M.S.B. is a Seaver Foundation Faculty Scholar. Additional support was provided by the National Institute on Aging (NIA) through grant 5R01AG087324-02 awarded to M.S.B. This work was also supported by the Lieber Institute for Brain Development with funding provided by the families of Constance and Steven Lieber and Milton and Tamar Maltz.
Author information
Authors and Affiliations
Contributions
K.M., G.S., S.S., R.K., V.D., and S.Sa. (Sylvia Savova) were responsible for human fetal tissue ascertainment, clinical annotation, and sample procurement. R.T., D.R.W., J.E.K., T.M.H., and V.V. oversaw tissue processing, RNA isolation, and sample quality control at the Lieber Institute and Johns Hopkins University. Data generation was performed jointly by teams at the Icahn School of Medicine at Mount Sinai and Johns Hopkins University. M.S.B. led the computational analyses, with major contributions from A.Y., X.W., and M.R.d.l.S. Interpretation of results was led by M.S.B.; all figures and tables were generated from A.Y., with input from J.D.B., T.M.H., D.R.W., J.E.K., and V.V. M.S.B. drafted the manuscript, with critical revisions from J.D.B., T.M.H., D.R.W., and J.E.K. All authors reviewed and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
DRW serves on the advisory board of Pasithea Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xinshu Xiao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Breen, M.S., Yang, A., Wang, X. et al. Trisomy 21 Drives ADARB1 Overexpression and Premature RNA Recoding in the Developing Fetal Brain. Nat Commun 17, 2797 (2026). https://doi.org/10.1038/s41467-026-70217-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70217-5