Abstract
The rise of drug resistance and limitations of current antifungal treatments highlight the urgent need for innovative antifungal strategies. Here we present the development of cis-fumaramidmycin-derived analogs inhibiting the interactions of ribosome assembly factor Mrt4 with rRNA to combat fungal infections. Through antifungal screening, we identified a promising lead 20 with strong efficacy against various drug-resistant fungi, including notorious super-fungus Candida auris. A comprehensive approach combining active-and-inactive-based protein profiling (AIBPP), chemical-genetic profiling, and fluorescence polarization revealed that the antifungal activity of 20 is primarily due to selectively inhibiting essential CaMrt4-rRNA interaction by conjointly covalent engaging C96&C189 on CaMrt4 but inactive for HuMrt4-rRNA interaction, thereby disrupting fungal ribosomal assembly. Therapeutic efficacy of 20 in both Galleria mellonella larvae and murine candidiasis models validate this antifungal strategy. Collectively, our studies provide a potential and much needed therapeutic strategy to address the rapidly rising burden of drug-resistant fungal infections.
Similar content being viewed by others
Introduction
Invasive fungal infections (IFIs) affect over 6.55 million people and cause more than 3.75 million deaths annually1. In 2022, the WHO established a fungal priority pathogens list (FPPL) to classify fungal pathogens based on infection severity2. Over 90% of life-threatening infections are caused by Candida, Aspergillus, and Cryptococcus species3. Of note, C. auris has become a concern of health authorities around the world due to the increasing number of cases characterized by serious invasive and multidrug-resistant infections, which shows low susceptibility to the drugs commonly used to treat systemic fungal infections (azoles, echinocandins, and polyenes), resulting in outbreaks with high mortality4. Especially, the COVID-19 pandemic has further worsened the situation, increasing susceptibility to fungal infection5. Currently, treatment options are limited to polyenes, flucytosine, azoles, and echinocandins6, primarily targeting the fungal cell membrane and cell wall7 (Supplementary Fig. 1). The overuse of these drugs has led to an increasingly severe issue of fungal resistance8,9, particularly to azoles, now documented in over 200 countries, complicating clinical treatment efforts considerably8, underscoring the need for new antifungal target and treatment. However, unlike bacteria, eukaryotic fungal cells are more similar to mammalian cells, thus, identifying new targets of antifungals and discovering new anti-resistant fungal drugs remain enormous challenges7.
Fumarate (trans-butenedioic acid), a naturally occurring organic acid and a key intermediate in the citrate cycle10, has garnered attention for its physiological and pharmacological properties. Derivatives of maleate and fumarate have shown promise in the treatment of cancer11, infections12, and autoimmune disease13. The FDA’s 2013 approval of dimethyl fumarate (Tecfidera®) highlights its pharmacological promise in the treatment of multiple sclerosis13,14. Between 2015 and 2019, 9.6% of FDA-approved drugs used fumaric or maleic acid for drug salification15, including treatments for multiple sclerosis, breast cancer, and leukemia. Fumaramidmycin, a butenedioic acid-based antibiotic isolated in 1975 from S. kurssanovii, displayed activity against bacteria but not fungi16. In 2012, however, Shode et al. synthesized seven fumaramidmycin analogs with modest antifungal activity by conversion of a highly polar carboxylic group to an ester functional group17; nonetheless, the elucidation of their specific target remains a huge challenge.
In this work, we report the development of cis-fumaramidmycin-derived compounds as a chemotype with potent antifungal activity against azole-resistant fungi, including C. auris, based on the antibiotic fumaramidmycin. Using an integrated approach that combines active- and inactive-based chemical proteomics (AIBPP) with tailor-made cis-active and trans-inactive probes, together with chemical-genetic profiling, transcriptome assays, etc., we identified the inhibition of the ribosome assembly factor Mrt4 as an emerging antifungal strategy. Furthermore, we conducted site-directed mutagenesis, fluorescence polarization (FP) assays, and molecular dynamics (MD) simulations to elucidate the regulatory mechanism of lead compound 20 on Mrt4. The therapeutic efficacy in both Galleria mellonella larvae and murine candidiasis models validated this antifungal strategy. This study highlights that inhibiting Mrt4-rRNA interactions could provide a much-needed therapeutic strategy to address the rapidly increasing burden of drug-resistant fungal infection and that cis-fumaramidmycin-derived compounds are a class of chemical probes suitable for the assessment of Mrt4-dependent pharmacology in cells and in vivo.
Results
Antifungal activity screening of fumaramidmycin-based derivatives
Pioneering studies on fumaramidmycin16 and its analogs17 have established the fumarate scaffold as the active antifungal unit (Fig. 1a). To elucidate the structure activity relationship (SAR) and optimize the antifungal potency, we synthesized trans- and cis-isomers (Supplementary Fig. 2) and evaluated their inhibitory effects on azole-resistant C. albicans and C. auris (Supplementary Table 1). The results showed that the cis-compounds (maleate ester) had comparable antifungal activities against fungi, while the trans-compounds (fumarate esters) were inactive. For instance, cis-compounds 3 and 4 inhibited fungal growth with MIC80 values of 1–8 μg/mL, whereas trans-compounds 1 and 2 were inactive, prompting the further development of cis-compounds. Notably, compound 5, featuring an α, β-unsaturated ester group, demonstrated high activity against various drug-resistant fungi, including multidrug-resistant C. auris. In contrast, compound 10, containing an α, β-unsaturated amide group, was inactive. In addition, the substitution pattern at R1 significantly influenced antifungal activity, i.e., the hydrophobic alkyl group enhanced the potency (e.g., 3, 4, 5, 6), whereas the hydrogen atom rendered the compound inactive (7). Similarly, phenyl group substitutions at the R2 position also significantly influenced activity. Compound 3 exhibited high potency, whereas analogs 8 and 9 were inactive even at concentration of 64 µg/mL. This SAR analysis identified compound 3 as a simplified yet potent scaffold, that exhibited comparable antifungal activity against azole-resistant strains, with MIC80 values of 1–4 µg/mL.
a The antifungal activity of 20 against various drug-resistant fungi. b, c Time-killing kinetics of 20 against C. albicans SC5314 and C. auris CBS12774, expressed as the mean ± SD from three independent experiments. The data are presented as the mean values ± SD (n = 3 independent experiments). d Fluorescence micrograph images of propidium iodide (PI)-stained C. albicans treated with DMSO, 20 and amphotericin B (AmB). Scale bars: 100 µm. e Fluorescence microscopy images of C. albicans SC5314 cells treated with different concentrations of 20 and cultured in RPMI1640 medium for 6 h at 37 °C (scale bars: 50 µm). f The inhibition of biofilm formation by 20 was quantitatively assessed using the XTT reduction assay. The bars represent the means ± SDs, n = 3. g, h C. albicans SC5314 cells grown overnight and stained with the ROS probe 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) to determine basal intracellular ROS levels. Cells were treated with DMSO, fluconazole (FLC) or 20 for 9 h. The fluorescence intensity was measured by flow cytometry, with geometric mean (GMean) values calculated for each treatment group. The data are presented as the means ± SDs (n = 3 independent experiments). i Compound 20 has relatively low toxicity to the human cell line LO2 under effective antifungal concentration (MIC80). Statistical significance was determined using a two-tailed unpaired Student’s t-test for f, h. The data are presented as the means ± SDs (n = 3 independent experiments). Source data are provided as a Source data file for all the panels.
Starting with compound 3 as the parent scaffold, we next investigated the impact of benzene ring substituent on antifungal activity (Supplementary Fig. 3). As summarized in Supplementary Table 2, para-substituted analogs exhibited superior antifungal activity relative to their meta- and ortho-substituted counterparts. For example, the p-chloro derivative 13 showed greater potency than the corresponding m-chloro (14) and o-chloro (15) analogs. Notably, several compounds (11, 13, 14, 16 and 20) displayed strong activity against multiple fungal species, with MIC80 values in the range of 1–2 μg/mL. Among these, compound 13 and 20 were the most potent against wild-type C. albicans SC5314. Notably, compound 20 demonstrated reduced cytotoxicity toward human cell line LO2 and improved plasma stability in mice compared with compound 13 (Supplementary Fig. 5). On this basis, compound 20 was selected for further investigation. The data in Fig. 1b, c reveal that compound 20 exhibited time- and dose-dependent killing activity against two strains, with minimal fungicidal concentrations (MFCs) of 2 μg/mL. The damage to the cell membrane caused by 20 can also be observed by the absorption of propidium iodide (PI), which penetrates only damaged membranes, releasing red fluorescence (Fig. 1d). In addition, compound 20 effectively inhibited hyphal growth in liquid RPMI1640 medium culture in a dose-dependent manner (Fig. 1e) and blocked biofilm formation of C. albicans SC5314 cells in a dose-dependent manner (Fig. 1f and Supplementary Fig. 6). Given that filamentation and biofilm formation are critical virulence factors associated with drug resistance9, these results explain to some extent why compound 20 may potently inhibit drug-resistant fungi. Additionally, compound 20 induced reactive oxygen species (ROS) overproduction, as evidenced by flow cytometry analysis (Supplementary Fig. S7), which showed that ROS levels were elevated in cells treated with compound 20 or fluconazole (FLC) (Fig. 1g, h). This ROS accumulation likely contributed to oxidative damage increasing antifungal activity.
Toxicity assessments on the human cell lines LO2, 293T, B2B, and HUVECs revealed that 20 exhibited CC50 values in the range of 10.6–22.3 μg/mL (Fig. 1i and Supplementary Table 3), over tenfold higher than its MIC80 value (1 μg/mL) against C. albicans SC5314, indicating relatively low toxicity to human cells. To further evaluate its safety, we performed Ames tests and hemolytic assays. As shown in Supplementary Fig. 8a–h, the number of revertant colonies did not exceed twice (TA102 and TA98) or three times (TA1537 and TA1535) that of the vehicle control at any tested dose, indicating no detectable mutagenic activity in the tested strains. In the hemolytic assays (Supplementary Fig. 8i), 20 caused no marked hemolysis at concentrations of 2–8 μg/mL, and only weak hemolysis (<10%) was observed at 16 and 32 μg/mL, substantially lower than that of the FDA-approved antifungal drug miconazole (94% hemolysis at 16 μg/mL). Moreover, no adverse effects of 20 were observed in our mouse studies. Collectively, these results demonstrate the relatively low toxicity potential of compound 20 in the tested mouse.
Design and synthesis of active and inactive probes
Given the promising antifungal activity of cis-fumaramidmycin-derived compounds, elucidating their mode of action is essential. Activity-based proteome profiling (ABPP)18 has emerged as a pivotal tool in drug discovery, leveraging chemical probes to unearth molecular targets and mechanisms in complex biological systems. A critical requirement for ABPP is the synthesis of probes bearing functional groups that allow bioorthogonal conjugation, with terminal alkynes widely employed because of their rapid reactivity, well-established chemistry, excellent regioselectivity, and commercial availability19. Inspired by recent studies highlighting the potential of fragment-based covalent ligand screening of α, β-unsaturated methyl esters to target reactive cysteine residues in proteins20, we hypothesized that these compounds most likely exert their activity via Michael addition to nucleophilic residues on target proteins. To test this hypothesis, we strategically replaced the chlorine atom on the benzene ring of 20 with an alkynyl group, ensuring that the alkyne installation preserved the intrinsic reactivity of Michael acceptor. Additionally, because the length of the side-chain carboxylic ester could influence antifungal activity, we modified the ethyl ester to a methyl ester. The resulting active cis-probe 21 was successfully synthesized (Supplementary Fig. 4) for proteomic profiling assays. Compounds 20 and 21 exhibited comparable antifungal activity against C. albicans strains SC5314 and 103, with MIC80 values in the range of 1–2 μg/mL (Supplementary Table 4), suggesting that they act on the same antifungal-relevant target. To further explore the structural determinants of activity, we synthesized the trans-probe 22 (Supplementary Fig. 4), the trans-configuration of 21. As shown in Supplementary Table 4, probe 22 failed to inhibit the growth of C. albicans and C. auris (MIC80 > 64 μg/mL), indicating that it is an inactive probe.
Electrophilic reactivity to nucleophilic reagents
To determine whether both probes undergo covalent reactions, we evaluated the electrophilic reactivity of the active cis-probe 21, and the inactive trans-probe 22 using glutathione (GSH), a nucleophilic reagent commonly employed to assess the electrophilicity of covalent warheads21. Nuclear magnetic resonance (NMR) spectroscopy analysis (Supplementary Fig. 9a–c) revealed new peaks corresponding to the addition product in the GSH model reaction, indicating that both probes participate in Michael addition reactions.
Given the importance of warhead reactivity in covalent inhibitors—where excessive reactivity can cause off-target effects22—we thus compared the reactivity of compounds 21 and 22 with that of compound 20, the amide analog 10 and several FDA-approved covalent drugs by measuring their half-life (T1/2) values (Supplementary Fig. 9d–j). Compounds 20 and 21 reacted more readily with GSH, with T1/2 values of 155 min (Supplementary Fig. 9e) and 80.6 min (Supplementary Fig. 9f), respectively, whereas compounds 10 and 22 were markedly less reactive, with T1/2 values of 815 min (Supplementary Fig. 9d) and 2432 min (Supplementary Fig. 9g), respectively. For comparison, their reactivity was similar to afatinib (T1/2 = 108 min, Supplementary Fig. 9j), lower than that of disulfiram (T1/2 = 43 min, Supplementary Fig. 9i)23, and higher than that of nirmatrelvir (T1/2 > 6000 min, Supplementary Fig. 9h)24. These results indicate that 20 and 21 possess moderate electrophilic reactivity, making them promising candidates for further optimization. Importantly, compounds 20 and 21, with moderate GSH activity, displayed potent antifungal activity, whereas compounds 10 and 22, with weaker reactivity, were inactive (Supplementary Tables 1 and 2). This clear correlation demonstrates that electrophilic reactivity is essential for the antifungal efficacy of these fumaramidmycin-derived compounds, and further supports the utility of 21 as an activity-based probe for identifying covalently bound protein targets within the fungal proteome.
To rationalize the distinct reactivities of the cis- and trans-isomers and the inactive compound 10, we further performed quantum mechanical (QM) calculations of their reaction pathways. Supplementary Fig. 10 illustrated that the calculated reaction barriers increase in the following order: cis (31.0 kcal mol−1), cis-amide (106.7 kcal mol−1), and trans (277.0 kcal mol−1). This trend aligns with the Michael addition mechanism, where charge delocalization favors the cis-configuration over the trans-configuration. Therefore, delocalization partially disrupts the conjugate amide in compound 10 and increases the reaction barrier, making the cis-amide inactive.
Proteomic profiling using active cis-probe 21
Having established the potent effect on antifungal-resistant-fungus activity and balanced the electrophilic reactivity of active cis-probe 21, we conducted a global and in-depth analysis of target proteins in fungal cells to identify its antifungal target using ABPP combined with quantitative MS25. To evaluate the in vitro reactivity of probe 21, we performed gel-based ABPP in C. albicans SC5314 cell lysates treated with the probe at varying concentrations (2–16 μM). The lysates were subsequently reacted with an azide-biotin (N3-biotin) reporter tag under copper-catalyzed azide-alkyne cycloaddition (CuACC or click chemistry) conditions25, and probe-labeled proteins were visualized via western blotting. As shown in Supplementary Fig. 11a, probe 21 labeled proteins within the 30- to 130-kDa range in a concentration-dependent manner, demonstrating its ability to detect both on- and off-targets of covalent probes in fungal cell lysates. We selected 2 and 4 μM concentrations of 21 for pull-down experiments to enrich its binding proteins (Supplementary Fig. 11b). The enriched samples were subjected to protein identification using LC–MS/MS (Supplementary Data 1). To minimize the impact of nonspecific binding, we focused on the top 10 proteins with the highest abundance at each concentration (Supplementary Table 5 and 6). Among the consistently enriched proteins were translation elongation factor 1-α 2 (TEF2), alcohol dehydrogenase (ADH1), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), fructose-bisphosphate aldolase (FBA), phosphoglycerate kinase (PGK1), pyruvate decarboxylase (PDC11), and pyruvate kinase (PK). GAPDH, FBA, and PGK1 showing increased abundance ratios from 2 to 4 μM of probe 21. Given that GAPDH and FBA are key enzymes in glycolysis and have been reported to be potential antifungal targets26,27,28, we evaluated the inhibitory activity of compounds 20, 21 and 22 against GAPDH from C. albicans (CaGAPDH) and FBA from C. albicans (CaFBA). While none of the compounds inhibited CaFBA, both 20 and 21 potently inhibited CaGAPDH, with IC50 values of 0.11 μM. In contrast, compound 22 exhibited significantly weaker inhibition, with an IC50 value of 25.02 μM (Supplementary Table 7). These results suggest that CaGAPDH is likely the molecular target of these compounds.
To further investigate whether GAPDH (TDH3) is the antifungal-relevant target of compounds, we disrupted the TDH3 gene in C. albicans using double homologous recombination. As shown in Supplementary Table 8, the sensitivity of the TDH3-null strain to compound 20 was comparable to that of the wild-type strain, indicating that GAPDH is not the antifungal-relevant target of 20. In addition, this result suggests that general ABPP experiments often enrich abundant housekeeping proteins such as GAPDH, a ubiquitous internal reference protein. Nonetheless, this result alternatively highlights previous finding that GAPDH contains a highly nucleophilic cysteine (C150) essential for catalysis13, making it particularly susceptible to covalent modification, even by inhibitors with modest reactive electrophilicity. Indeed, several FDA-approved covalent drugs, i.g., disulfiram, afatinib, nirmatrelvir, are known to react with GAPDH and inhibit its catalytic activity (Supplementary Fig. 9k). However, this off-target activity does not preclude their approval as therapeutic agents.
Activity- and inactivity-based proteomic profiling (AIBPP) using cis-probe 21 and trans-probe 22
To identify the true molecular target of active probe 21, we improved ABPP approach by incorporating both active cis-probe 21 and inactive trans-probe 22, to create an activity- and inactivity-based proteomic profiling (AIBPP) strategy, as depicted in Fig. 2a. The active cis-probe 21-treated group displayed significant concentration-dependent labeling of proteins in 30–130 kDa range, whereas the inactive trans-probe 22 resulted in much weaker labeling (Fig. 2b), indicating a marked difference in target reactivity between the two probes. Although both probes labeled proteins in a concentration-dependent manner, labeling by 22 was considerably weaker, which is consistent with its lower target reactivity. Next, we performed pull-down experiments to separate target proteins based on affinity. After extensive washing to eliminate nonspecific binding, the captured proteins were analyzed by SDS–PAGE and Western blotting (Fig. 2c). The consistency of proteome labeling before and after pull-down confirmed the reliability of our approach. In-gel trypsin digestion followed by LC–MS/MS analysis (Supplementary Data 2) identified 12 quantitative proteins (Supplementary Table 9) with a p value < 0.05 and an abundance ratio >4 (Fig. 2d) when comparing 8 μM 21-treated lysates to 22-treated lysates in C. albicans. The abundance ratio was calculated as the difference in protein intensity between the active probe 21- and inactive probe 22-treated cell lysates. Furthermore, the binding energies (Supplementary Table 10) for probe 21 and the proteins identified in the AIBPP experiments were calculated using Cov_DOX29 methods, indicating that covalent binding between probe 21 and these proteins is feasible.
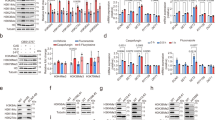
a Schematic illustration of the AIBPP strategy for identifying targets of fumaramidmycin-derived compounds. Created in BioRender. Wang, W. (2026) https://BioRender.com/l63dasz. b Dose-dependent labeling of C. albicans SC5314 cell lysates with 21 and 22. Coomassie blue staining (CBB) was used to confirm equivalent protein loading. c Streptavidin pull-down experiment showing enriched labeling of proteins from C. albicans cell lysate by 21 and 22. d Scatter plot of the 16 quantitatively identified proteins and specific targets from gel-free labeling experiments with 8 μM 21 versus 8 μM 22 in C. albicans. cell lysate (p value < 0.05, abundance ratio >4). Abundance ratios were calculated pairwise for replicate sets, with the median ratio reported for each protein. Statistical significance (p-value) was determined using a two-sample paired t-test. e, f Growth curves of wild-type and heterozygous deletion S. cerevisiae strains in response to 20 (2 µg/mL), as identified in the haploinsufficiency screen. g Relative growth rate plot of wild-type and heterozygous deletion strains of S. cerevisiae in response to 20 (2 µg/mL), as identified in the haploinsufficiency screen. Source data are provided as a Source data file for (e, f, g). Data are presented as the means ± SDs (n = 3 independent experiments).
To corroborate these findings, we further performed the competitive ABPP experiments. 20-pretreated Lysates displayed substantially reduced protein labeling compared with those pretreated with probe 21 alone, as shown by SDS–PAGE and Western blotting (Supplementary Fig. 12a), suggesting 20 and 21 have overlapping targets. LC–MS/MS analysis of enriched proteins confirmed this overlap, with volcano plots comparing 21 versus DMSO (Supplementary Fig. 12b and Supplementary Data 5) and 21 versus 20-pretreated samples (Supplementary Fig. 12c and Supplementary Data 6) consistently identifying Mrt4 as a significantly enriched protein (p < 0.05, abundance ratio >4). Collectively, these data establish Mrt4 as a key antifungal-relevant target of fumaramidmycin-derived compounds.
Identifying Mrt4 via 20-induced haploinsufficiency screening
Simple eukaryotes such as S. cerevisiae share significant biological similarities with C. albicans, and their genetic tractability facilitates chemogenomic profiling30. Several successful efforts have demonstrated the ability to determine drug modes of action through yeast genome-wide drug-induced haploinsufficiency screening31,32. To confirm the target of the cis-fumaramidmycin-derived compounds, we mapped orthologous proteins in S. cerevisiae corresponding to the differentially expressed proteins identified earlier. We monitored growth by measuring the optical density over time in liquid cultures of wild-type S. cerevisiae and the mutant strains (Fig. 2e). These strains were treated with 2 µg/mL lead compound 20, and real-time growth was recorded (Fig. 2f). Among the mutant strains, deletion of one copy of the MRT4 (mRNA turnover protein 4) gene in this diploid organism caused the most significant growth reduction upon treatment with 20, with a 30% decrease compared with wild-type growth (Fig. 2g). This supports the notion of Mrt4 as an antifungal-relevant target of 20, based on the principle of induced haploinsufficiency.
Target validation of Mrt4 as a direct antifungal target of cis-lead 20
To further confirm that compound 20 targets Mrt4 in C. albicans cells and inhibits growth, we generated an mrt4 null mutant with a single copy deletion. As shown in Fig. 3a, compound 20 induced haploinsufficiency, enhancing antifungal activity against MRT4/mrt4Δ strains, consistent with the results in S. cerevisiae mutant strains. A pioneering study33 revealed that at the early stages of ribosome assembly, Mrt4 forms a complex with Rpl12 and interacts with the GAR structural domain (residues 1217–1286) of 25S rRNA, which is anchored to the ribosomal stem loop and influences the assembly of the ribosomal stem. Subsequently, Yvh1 binds to the pre-60S subunit, which triggers Mrt4 release (Fig. 3b). This induces a series of biological events that ultimately lead to the translocation of the pre-60S subunit out of the nucleus and the formation of mature ribosomes34. To assess the potential effects of Mrt4 inhibition on ribosomes and other cellular processes, we performed the transcriptional analysis of C. albicans SC5314 under 20-treatment. GO enrichment analysis (Supplementary Data 4) revealed that ribosome-related pathways, including those associated with Mrt4-mediated ribosome assembly, were among the top enrichments for differentially expressed genes (DEGs) (Fig. 3c and Supplementary Fig. 13). The volcano plot (Fig. 3d and Supplementary Data 3) showed significant upregulation of MRT4, YVH1 and RPL12 mRNAs, encoding proteins involved in 60S ribosomal subunit assembly (Fig. 3b)34,35. These transcriptional variations of MRT4, YVH1, and RPL12 were further validated by quantitative real-time PCR (qPCR) (Fig. 3e).
a Sensitivity correlation of C. albicans strains to 20 and 21 based on the MRT4 gene dosage and mutation status. The growth of wild-type and mutant strains was monitored by measuring the optical density (OD600) over time in liquid cultures. The data represent the mean of three independent experiments. b Schematic representation of the Mrt4-associated pathway involved in the assembly of the ribosome stalk base in eukaryotes. Created in BioRender. Wang, W. (2026) https://BioRender.com/z9cwodr. c Top pathways affected by 20 treatment in C. albicans SC5314. The scatter plot shows the number of differentially expressed genes (DEGs) as a function of point size, while the color gradient (from blue to yellow) indicates increasing enrichment significance. d Global gene expression changes in C. albicans SC5314 following treatment of 20. Fold changes were determined by comparing normalized transcript counts between 20-treated samples and DMSO-treated controls. A one-sided version of Fisher’s exact test was performed, and the p-value was adjusted using the Benjamini–Hochberg method. e Relative transcriptional changes of MRT4 and related genes (YVH1 and RPL12) between the DMSO-treated group and 20-treated group. f Polysomal profiles of C. albicans SC5314 cells treated with DMSO (control) or 20. The data are representative of three independent experiments. g Quantitative comparison of the peak areas for ribosomal mono-subunits (40S, 60S, and 80S) between the DMSO-treated and 20-treated groups. The data are representative of three biological replicates. Data are presented as the means ± SDs (n = 3 independent experiments). Statistical significance was determined using a two-tailed unpaired Student’s t-test. Source data are provided as a Source data file for (e, f, g).
To further confirm that compound 20 affects ribosome assembly in C. albicans by targeting Mrt4, we performed sucrose density gradient centrifugation to extract ribosomal fractions from lysed cells pretreated with 20 or DMSO, followed by polysome profiles analysis. Figure 3f, g demonstrated significant differences between the polysome profiles of the 20-treated and control groups. Specifically, the 40S and 60S subunit peak were markedly elevated in the 20-treated group, whereas the 80S peak was significantly reduced. These results indicate that the accumulation of 40S and 60S subunits increased, whereas 80S subunit formation (resulting from the assembly of 40S and 60S subunits) decreased, possibly due to impaired ribosome assembly. Taken together, these findings further confirmed that Mrt4 is a valid target for cis-fumaramidmycin-derived compounds.
Inhibition of CaMrt4-rRNA interaction and ribosome assembly
To further elucidate the inhibitory mechanism of compound 20 against CaMrt4, recombinant CaMrt4 protein was expressed in E. coli. Dose-dependent labeling of recombinant CaMrt4 (Supplementary Fig. 14a) revealed that unlike inactive trans-probe 22, active cis-probe 21 selectively modified CaMrt4. Given the GTPase-associated region (GAR) structural domain in 25S rRNA (a component of the 60S subunit) serves as the binding site for Mrt4 from S. cerevisiae (ScMrt4)33 (Fig. 4a), with high (70%) Mrt4 sequence identity (Supplementary Fig. 15a) and 100% GAR domain identity (Supplementary Fig. 15b) in C. albicans and S. cerevisiae, we synthesized a single-stranded rRNA probe corresponding to the GAR structural domain sequence (1217–1286 nt of 25S RNA) and modified it with a 3′-fluoresceinamine group (FAM). The binding affinity of this RNA probe to recombinant CaMrt4, with or without 20 treatment, was analyzed using EMSA. The rRNA probe exhibited a relatively higher position in the gel upon the addition of CaMrt4 (Fig. 4b, lane 2), indicating the formation of larger complexes. However, with increasing concentrations of 20, the migration rate of the rRNA probe increased, and the band position shifted downwards (lanes 3–7), demonstrating that compound 20 can inhibit the binding of the RNA probe to CaMrt4 in a dose-dependent manner.
a The cryo-EM structure of the GAR domain-Mrt4 complex in the pre-60S ribosomal particle (PDB 8hfr). b Binding activity of a FAM-labeled ssRNA probe to CaMrt4 with or without 20 treatment, analyzed via Electrophoretic mobility shift assay (EMSA). The final concentrations of 20 in lanes 2–7 were 0, 0.63, 1.25, 2.5, 5, and 10 µM, respectively, while the FAM-labeled ssRNA probe concentration was fixed at 0.1 µM in all the lanes. The final CaMrt4 protein concentration was 5 µM in lanes 2–7, with lane 1 serving as the control (no CaMrt4 added). The RNA probe corresponds to the GAR structural domain of 25S RNA (1217–1286 nt) with 3′-FAM modification. B bound, U unbound. c Half maximal inhibitory concentration (IC50) of 20 for the interaction between the FAM-labeled ssRNA probe and wild-type CaMrt4. d Inhibition rates of cis-fumaramidmycin-based derivatives against the GAR domain-Mrt4 interaction at the concentration of 4 µM. The data are presented as the means ± SDs (n = 3 independent experiments). e–h Binding response curves and dissociation constants (Kd) of the FAM-labeled ssRNA probe with wild-type CaMrt4 cysteine mutants, as measured using FP assays. The protein was pre-incubated with or without 20 (4 µM) overnight at 4 °C. i Overlap of the 3D structures of CaMrt4 and HuMrt4 generated by AlphaFold3, highlighting the cysteine residues in both proteins. j Binding response curves and dissociation constants (Kd) of the FAM-labeled ssRNA probe with HuMrt4 at varying concentrations, with or without pretreatment with 20 (4 µM). All fluorescence polarization assays were performed under identical conditions. The data are presented as the means (n = 2 independent experiments) and the source data are provided as a Source data file.
Consistent results were obtained from fluorescence polarization (FP) assays. the dissociation constant (Kd) for CaMrt4 binding to the rRNA probe was determined to be 0.29 µM (Fig. 4e), which increased significantly (Kd = 7.49 µM, 25.8-fold) after treatment with compound 20. IC50 values were calculated by testing various concentrations of 20, revealing an IC50 value of 3.67 µM for the marked inhibition of the CaMrt4-rRNA interactions (Fig. 4c). Notably, a strong correlation was observed between the antifungal activity of the cis-fumaramidmycin analogs and their inhibitory activity on the CaMrt4-rRNA interaction (Fig. 4d and Supplementary Table 11), establishing that CaMrt4 is the principal antifungal-target of these compounds.
Liquid chromatography−mass spectrometry (LC−MS) analysis of CaMrt4-20 in Supplementary Fig. 14c, d revealed that 20 may covalently modify residues C96 and C189 in CaMrt4. To validate their functional relevance, we expressed and purified the single mutants C96A and C189A and the double mutant C96A/C189A. The binding affinities of CaMrt4-C96A/RNA-probe (Kd = 0.27 µM) and CaMrt4-C189A/RNA-probe (Kd = 0.39 µM) were similar to that of wild-type CaMrt4/RNA-probe (Kd = 0.29 µM) (Fig. 4f, g). However, C96A/C189A double mutant exhibited an ~6-fold decrease in binding affinity (Kd = 1.86 µM, Fig. 4h) with RNA-probe compared to wild-type (Kd = 0.29 µM). Upon pretreatment with 20, both the C96A and C189A mutants displayed reduced binding affinity (Kd = 1.94 µM for C96A and 1.24 µM for C189A) with the RNA-probe, whereas the C96A/C189A double mutant maintained its binding affinity with the RNA-probe. To confirm that covalent engagement underlies the observed inhibition, we also employed compound 24 (Supplementary Fig. 14e), a reduced analog of 13 lacking the α, β-unsaturated Michael acceptor, as a negative control. Compound 24 displayed no antifungal activity, and FP assays revealed negligible inhibition of the CaMrt4-rRNA interaction (Supplementary Fig. 14e), confirming that covalent engagement is essential for disrupting this interaction. These results demonstrate that both C96 and C189 are essential for the inhibition of 20 to CaMrt4/RNA interaction.
To investigate the species selectivity of Mrt4 as an antifungal target, HuMrt4 was heterologously expressed and purified, and its interaction with RNA was analyzed using the same FP assay. HuMrt4 bound RNA with a Kd of 0.26 µM, similar to that of CaMrt4. However, treatment with 20 caused only a modest 1.8-fold change in binding affinity (Kd = 0.48 µM; Fig. 4j), far less than the 25.8-fold effect observed for CaMrt4 (Fig. 4e). Structural comparison (Fig. 4i) of the CaMrt4 and HuMrt4 predicted by AlphaFold3 revealed that C189 is conserved, while C96 is absent in HuMrt4. This non-conserved cysteine (C96) represents a critical determinant for the selective inhibition of the CaMrt4-rRNA interaction by 20. Furthermore, analysis of residues within 5 Å of the conserved C189 (Supplementary Fig. 16b) revealed markedly different residue compositions (sequence identity <10%), highlighting distinct chemical microenvironment around C189 is fundamentally different in CaMrt4 and HuMrt4. Together, these results support the potential for designing covalent inhibitors that can selectively target fungal Mrt4.
Finally, comparative sequence analysis of Mrt4 across the key human-pathogenic fungal strains in our laboratory (C. albicans 5314, A. fumigatus 7544, F. solani 2478, R. oryzae 4549, C. glabrata 8535) revealed that C96 is unique to CaMrt4, whereas C189 is conserved across all the examined non-Candida pathogens (Supplementary Fig. 17). The overall sequence identities to CaMrt4 were 17.1% (RoMrt4), 66.1% (CgMrt4), 41.5% (AfMrt4), and 38.7% (FsMrt4). Consistent with these observations, compound 20 exhibited broad antifungal activity against all the tested pathogens, although its potency was consistently reduced in species lacking the C96 residue (Supplementary Table 12).
The inhibitory mechanism of lead compound 20 on CaMrt4-rRNA interaction
Sequence alignment revealed high conservation of Mrt4 (70% identity; Supplementary Fig. 15a) and GAR (100% identity; Supplementary Fig. 15b) between C. albicans and S. cerevisiae. Moreover, 20 potently inhibited the ScMrt4-rRNA interaction (Supplementary Fig. 15c). Thus, we performed the molecular dynamics (MD) simulations (Supplementary Data 7) using the cryo-EM structure of the Mrt4 from S. cerevisiae (ScMrt4)-rRNA complex (PDB ID: 7UQZ; 2.4 Å resolution) as initial structure to elucidate the mechanism by which lead compound 20 regulates CaMrt4-rRNA interactions.
Notably, the α, β-unsaturated ketone moiety of compound 20 contains two carbon atoms that are susceptible to covalent modification by cysteine thiols on CaMrt4. To clarify the preferred position of cysteine thiol addition, we used GAPDH from S. cerevisiae (ScGAPDH), a close homolog of CaGAPDH (Supplementary Fig. 18a), as a model system. We successfully resolved cocrystal structures of ScGAPDH in complex with cis-lead 20 (Supplementary Fig. 18c and Supplementary Table 13). Structural comparisons between ScGAPDH-20 and HuGAPDH-NAD+ revealed a conserved catalytic site centered on C15036 (Supplementary Fig. 18b). In the ScGAPDH-20 complex (PDB ID: 9IWD), a covalent bond can be observed between a carbon near the ester terminus of the α, β-unsaturated ketone in 20 and the sulfur of C150 on ScGAPDH. These findings suggest that the ester-side carbon is the favored electrophilic position for thiol addition, providing mechanistic insight into how the cysteine thiol in Mrt4 may covalently react with 20.
By incorporating insight from the ScGAPDH-20 cocrystal structure, we established an initial structure in which compound 20 covalently modified C189 and C96 on ScMrt4, followed by parallel MD simulations. The simulations revealed that the two domains of ScMrt4 form a large groove that accommodates the RNA substrate through extensive ionic interactions with phosphate groups. The two cysteine residues, C96 and C189, each covalently bound by compound 20, are in separate domains (Fig. 5a). The average RMSD of the two domains remain stable at less than 2 Å in the wild-type complexes, and the 20-modified complex exhibits a broader RMSD distribution, increasing by approximately 2 Å (Fig. 5d). Specifically, the covalent modification of C96 by compound 20 enables a new cation–π interaction with K69 and R70, disrupting a critical interface with rRNA (Fig. 5b); whereas the covalent bond of compound 20 with C189, which is situated deeper within domain II, induces a π–π stacking interaction with F135 and F206 altering the hydrophobic packing within this domain (Fig. 5c). MM-GBSA analysis (Fig. 5n) revealed that these structural rearrangements in the 20-modified complex further increased the RMSD of the RNA and dramatically reduced the binding affinity by over 40 kcal/mol relative to that of the wild-type. Detailed conformational clustering analyses (Supplementary Fig. 19) further revealed that the distributions of the R57–G1259 and K204–G1229 distances in the 20-modified complex across all four clusters predominantly shifted towards larger values from those in the wild-type (WT), which is consistent with the trends shown in Fig. 5d. Similarly, the distance distributions for K75–G1221 and R23–G1222 in WT are bimodal, whereas the corresponding cluster distributions shift slightly towards larger values, albeit to varying degrees. The structural perturbation induced by 20 attenuated key electrostatic interaction essential for the binding of Mrt4 to RNA. Prominent local structural changes are most pronounced around R57, K69, K75, and K23 near C96, where the average salt-bridge distance increases by over 1 Å (Fig. 5j–l), whereas the salt bridges of K168 and K204 in domain II are completely disrupted (Fig. 5j, m). Each individual modification by 20 independently reduces RNA-binding affinity, suggesting a complementary role in weakening the rRNA interaction and collectively contributing to the strong antifungal activity observed in the experiments.
a The Molecular Dynamics (MD) snapshots of 20-modified ScMrt4 superimposed on the wild-type ScMrt4-rRNA complex (PDB ID: 7UQZ). b Cation-pi interaction between 20 and ScMrt4 in domain I. c π-π stacking between 20 and ScMrt4 in domain II, altering the local hydrophobic packing with F135 and F206. d RMSD distribution comparison of the two protein domains and rRNA. e–i Validation of the predicted essential binding sites (K69, R70, F135 and F206) between 20 and CaMrt4 through mutational experiments. CaMrt4 protein was preincubated with or without 20 (4 µM) overnight at 4 °C. The response curves and dissociation constants (Kd) of FAM-labeled ssRNA probes against various concentrations of wild-type or mutant CaMrt4 were measured using FP assays. For all FP assays, the data are presented as the means ± SDs (n = 2 independent experiments) and the source data are provided as a Source data file. j Residue-specific analysis of the binding interaction between 20 and wild-type ScMrt4, highlighting a global reduction on interactions at both domains. k Attenuation of R57-ADE1259 interactions. l Attenuation of K75-ADE1221 interactions close to C96. m Attenuation of K204-GUA1229 interactions in proximity to C189.
To confirm the critical role of C96 in regulating fungal Mrt4-rRNA interactions, we examined the inhibitory effect of 20 on the interaction between Chaetomium thermophilum Mrt4 (CtMrt4) and rRNA. CtMrt4 shares 45% sequence identity (Supplementary Fig. 20a) with CaMrt4, and exhibits nearly complete conservation (94%) of the rRNA GAR domain (Supplementary Fig. 20b). Owing to the absence of the C96 residue in CtMrt4, FP assays revealed that compound 20 failed to inhibit CtMrt4 (WT)-rRNA interaction (Supplementary Fig. 20c). However, the introduction of a cysteine residue at position 96 rendered CtMrt4(L96C) sensitive to inhibition by compound 20 (Supplementary Fig. 20c), further underscoring the critical role of C96 in mediating the covalent inhibition of Mrt4-rRNA interactions. Notably, apo-CtMrt4(L96C) crystals were successfully obtained (Supplementary Fig. 20d), and structural analysis revealed moderate global similarity to CaMrt4 (RMSD = 3.86) with conservation of key cysteine residues.
To verify the role of K69, R70, F135, and F206 as essential binding sites, we further conducted protein mutation experiments on CaMrt4, altering these structurally conserved residues that occupy identical positions in S. cerevisiae and C. albicans orthologues. As shown in Fig. 5e–h, the binding affinity of single-point mutations (K69A, R70A, F135A and F206A) to the RNA-probe decreased by approximately 5–7 fold after pretreatment with compound 20, which was less than the reduction observed in wild-type CaMrt4 (where the Kd changed by 25-fold), indicating that these four residues are critical for the inhibitory effect of compound 20 on CaMrt4-rRNA interaction. Notably, for the K69A/R70A/F135A/F206A quadruple mutation, its binding affinity to RNA-probe decreased by only 2.3-fold (Fig. 5i) after pretreatment with 20, indicating that compound 20 almost completely loses its inhibitory activity in the absence of these four key residues, further validating the allosteric inhibitory mechanism of compound 20 on the CaMrt4-RNA interaction predicted by previous MD calculation.
To theoretically elucidate the inhibitory selectivity of compound 20 against HuMrt4-rRNA, we conducted MD simulations of the 20-adduct HuMrt4-rRNA complex. The initial structure of HuMrt4-rRNA was generated using AlphaFold3. Because HuMrt4 lacks the C96 residue (Fig. 4i), the adduct of compound 20 was modeled at C176 (corresponding to C189 in CaMrt4). As shown in Supplementary Fig. 21, the formation of the 20–C176 adduct stabilized rather than disrupted the HuMrt4-rRNA interaction. In contrast, the covalent modification of C189 in ScMrt4-rRNA by compound 20 introduced π–π stacking interactions with F135 and F206, thereby altering the local hydrophobic packing and weakening RNA binding (Fig. 5c). These results suggest that in addition to the reactive cysteines (C96 and C189), the surrounding chemical microenvironment—particularly residues such as F135 and F206 —plays a critical role in determining the inhibitory outcome of compound 20. Consequently, although compound 20 can covalently modify conserved cysteine C176 in HuMrt4, it does not impair the interaction of HuMRT4-rRNA, which is consistent with its experimentally observed selectivity and safety profile.
In vivo efficacy of 20 in azole-resistant Candida infection
Given the promising in vitro antifungal effects of 20, we next evaluated its therapeutic potential in vivo using G. mellonella larvae and a mouse-invasive candidiasis model. Owning to its efficient inoculum delivery and ease of handling, the G. mellonella larva model is a simple and valuable emerging model host for studying fungal pathogenesis and antifungal efficacy37. Therefore, we utilized this invertebrate model to investigate the in vivo efficacy of 20. Microscopic examination of periodic acid-Schiff (PAS)-stained tissue sections was conducted alongside fungal burden quantification. Selected fields containing fungal cells were imaged, and the results revealed that treatment with 20 significantly reduced the number of C. albicans cells (Fig. 6a). As shown in Fig. 6b, compound 20 had a therapeutic effect comparable to that of FLC in G. mellonella, significantly reducing the oral cavity fungal burden by more than 10-fold with that in the PBS control group. Additionally, both the 20-treated and FLC-treated groups significantly prolonged the survival time of G. mellonella larvae relative to controls (Fig. 6c).
a Photomicrographs of PAS-stained biopsies from G. mellonella larvae after the indicated treatments. Scale bar, 50 μm. b Fungal burden in the infected G. mellonella larvae treated with solvent, 20 or fluconazole (FLC) (n = 6 per group). c Survival curves of G. mellonella treated with vehicle, 20 and FLC following infection with C. albicans SC5314. The long-rank (Mantel-Cox) test was used to compare differences between groups. d Fungal burden in the kidneys of infected mice after treatment with 20 (n = 9 per group). Statistical comparisons between groups were performed using one-way ANOVA. e Stability and half-life (T1/2) of 20 in mouse plasma with and without JZL184 (n = 2 independent experiments). f In vivo antifungal potency of 20 in combination with JZL184 (n = 9 per group). g PAS (scale bar, 50 μm) and hematoxylin and eosin (H&E) staining (scale bar, 100 μm) of kidneys from infected mice treated with vehicle, 20, or 20 in combination with JZL184. Mice were infected with azole-resistant C. albicans strains 103 (6 × 104 cells per mouse) via tail vein injection to establish an invasive Candida infection model. The data are presented as the means ± SDs for (b, d, f).
Following the favorable antifungal effect observed in the G. mellonella larval model, we further evaluated the therapeutic efficacy of 20 in a mouse model of invasive candidiasis infection. Figure 6d displays that compound 20 reduced the fungal burden in the mouse kidney with limited efficacy by less than 3-fold compared to the control group in both 10 and 20 mg/kg treated group. This modest efficacy may be attributed to the rapid metabolism of compounds with carboxylester structures in vivo11. It has been reported that the carboxylesterase inhibitor JZL18411 can enhance the plasma stability of drugs containing such structures. Therefore, we tested whether the coadminstration of JZL184 improved the plasma stability and in vivo efficacy of compound 20. As shown in Fig. 6e, the half-life of 20 in mouse plasma was significantly extended in a dose-dependent manner with the addition of JZL184. On basis of these results, we selected a 1:4 molar ratio of 20 to JZL184 for further evaluation in a mouse infection model. As illustrated in Fig. 6f, the combination of 20 (20 mg/kg) and JZL184 (60 mg/kg) significantly increased the antifungal efficacy, reducing the fungal burden in the kidneys by approximately 6.6-fold compared with that in the control group. This improvement was markedly greater than that of 20 (20 mg/kg) treatment alone. In addition, treatment with JZL184 alone did not result in antifungal activity either in vitro (Supplementary Table 14) or in vivo (Fig. 6f). Histological analyses further corroborated these findings. PAS staining (Fig. 6g) revealed several filamentous fungal cells in the kidneys of mice treated with the vehicle, whereas fungal cells were rarely observed in the kidneys of mice treated with 20. H&E staining revealed significant renal damage, including glomerular shrinkage, the microtubular morphology in some tubules, and extensive inflammatory infiltration with nuclear fragmentation in the renal interstitial matrix in the vehicle-treated groups. PAS and H&E staining revealed a marked reduction in fungal counts and inflammatory infiltrates in the group treated with the combination of 20 and JZL184. Moreover, in the groups treated with 20 alone or 20 in combination with JZL184, the renal tissue was intact overall, the glomeruli were uniform in size without obvious atrophy and the local tubules were tightly connected. These findings support the potential of inhibiting CaMrt4-rRNA interaction as a promising antifungal strategy for treating drug-resistant candidiasis.
Discussion
This study identifies the ribosome assembly factor Mrt4 as a druggable vulnerability in fungal cells and establishes cis-fumaramidmycin–derived compounds as small-molecule modulators of Mrt4-dependent ribosome biogenesis. By integrating chemical proteomics with chemical–genetic and functional assays, we delineate a direct mechanistic link between compound binding, disruption of Mrt4–rRNA interactions, and antifungal activity.
A key conceptual advance of this work is the demonstration that lead compound 20 selectively perturbs the Mrt4-rRNA interaction rather than broadly inhibiting ribosome function. Notably, our data indicate that cysteine reactivity alone does not determine selective covalent modification of CaMrt4 by compound 20; instead, the chemical microenvironment surrounding C96 and C189 are equally critical. Previous studies38 have shown that catalytic cysteines often display enhanced nucleophilicity when paired with proximal histidine residues that function as acid-base catalyst. For instance, in GAPDH, the highly nucleophilic C150 is activated by neighboring H177, rendering it susceptible to covalent modification even by weak electrophiles, which explains why GAPDH was initially enriched by the active cis-probe 21. In contrast, C96 and C189 in CaMrt4 lack proximal histidine residues, resulting in markedly weaker intrinsic reactivity. MD simulation revealed that compound 20 compensates for this reduced cysteine reactivity by forming stabilizing non-covalent interactions within Mrt4 microenvironment. Specifically, 20 establishes a cation-π interaction with K69 and R70 adjacent to C96, disrupting a critical rRNA-binding interface, as well as π-π stacking interaction with F135 and F206 near C189, thereby altering local hydrophobic packing. Site-directed mutagenesis confirmed the functional importance of these residues in mediating compound-dependent inhibition. Collectively, these findings demonstrate that selective engagement of CaMrt4 arises from the combined contributions of covalent cysteine modification and precisely tuned non-covalent interactions, providing a mechanistic rationale for the high selectivity of compound 20 over other cysteine-containing proteins. These insights also establish a foundation for future structure-guided optimization of Mrt4-directed chemical probes.
Targeting ribosome-associated pathways has precedent in antifungal drug discovery, exemplified by sordarin, which selectively inhibits fungal eukaryotic elongation factor 2 (eEF2)39 by stabilizing the eEF2–GDP–ribosome complex in S. cerevisiae and blocking ribosomal translocation. Species selectivity arises from divergence within the sordarin specificity region of fungal eEF2, highlighting the feasibility of selectively modulating conserved ribosomal processes through exploitation of subtle structural differences40. Our findings extend this paradigm from translation to ribosome assembly, an earlier and mechanistically distinct stage of ribosome biogenesis.
The dysfunction of key proteins and mutation of RNAs in ribosome are usually associated with severe human diseases41,42, including cancer43, Alzheimer’s disease44, and congenital ribosomopathies. However, ribosome assembly is an exceptionally complex process involving numerous protein-protein and protein-RNA interactions33,45, which has historically limited its tractability to chemical intervention45,46, particularly in eukaryotic systems. Consequently, studies of ribosome biogenesis have relied predominantly on genetic and structural approaches33,34. While several antibacterial antibiotics targeting ribosome assembly or subunit formation have been identified47,48, selective chemical probes for fungal ribosome biogenesis remain scarce. By inhibiting the Mrt4-rRNA interaction, compound 20 disrupts ribosome assembly and maturation, resulting in impaired fungal protein synthesis. Beyond its therapeutic implications, compound 20 thus represents a valuable chemical probe for interrogating ribosome assembly dynamics in eukaryotic cells, and aids in the discovery of new antifungal drugs targeting the ribosome. Nevertheless, the precise spatiotemporal coordination of assembly factors during ribosome maturation, and how this coordination can be selectively perturbed by small molecules, remains an important direction for future studies.
The covalent warhead of these inhibitors is derived from fumaramidmycin, a well-known antibiotic isolated from Streptomyces. Notably, related electrophilic motifs are widely distributed among natural products, such as coumarin, canthin-6-one, piperitone, carvone, and car-3-ene-2,5-dione. Between 2015 and 2019, 9.6% of FDA-approved drugs used fumaric or maleic acid for drug salification15, including treatments for multiple sclerosis, breast cancer, and leukemia. Accordingly, fumaramidmycin and related natural products offer versatile scaffold bearing reactive warheads that may be broadly applicable to “undruggable” protein-protein or protein-RNA interactions. This highlights the power of natural product-inspired scaffolds in identifying unique protein microenvironments and in discovering covalent drugs that exploit subtle structural differences to achieve selectivity.
Despite their promising activity, the clinical potential of fumaradmycin derivatives as antifungal agents is limited by low plasma stability and a presumably narrow spectrum of activity. Further optimization efforts will therefore focus on improving metabolic stability through rational modification of aromatic substitutions, amide functionalities, scaffold cyclization, and sterically hindered esters, as well as exploring advanced drug delivery strategies such as liposomal nanocarriers (e.g., AmBisome®, DaunoXome®, and DepoDur®) to enhance bioavailability and in vivo efficacy.
Methods
Chemistry
Unless otherwise stated, all reagents and solvents were obtained from commercial suppliers, including Sigma-Aldrich and Aladdin, and used without additional purification. Reaction progress was monitored by thin-layer chromatography (TLC) using aluminum-precoated silica gel plate (Yantai Huiyou Co., China) and UV irradiation at 254 nm. Column chromatography was performed on silica gel 60 (F254, 200−300 mesh), with columns prepared as slurries in appropriate solvent/solvent systems, and samples applied as concentrated solutions.
1H NMR and 13C NMR spectra were recorded on a Bruker AMX 600 spectrometer in DMSO-d6 with tetramethylsilane (TMS) as an internal standard. Chemical shifts are reported in parts per million (ppm) relative to TMS, and coupling constants (J) are given in Hertz (Hz). Signal abbreviations are as follows: broad (br), singlet (s), doublet (d), triplet (t), quartet (q), and multiplet (m). High-resolution mass spectrometry (HRMS) data were acquired by electron ionization on a Waters GCT Premier instrument. Compound purity was confirmed by HPLC analysis using a DIONEX UltiMate 3000 system equipped with a Thermo Fisher Scientific 120 C18 column (4.6 × 250 mm2, 5 μm). The mobile phase employed a MeCN gradient, with detection at 254 nm. Purity was determined based on the average peak area, and all the compounds were verified to be >95% pure.
General Synthetic Procedure for methyl (E)-4-oxo-4-(phenylamino)but-2-enoate (1). Aniline (1b) (0.86 g, 5 mmol) was placed in a 100 mL round bottom flask, followed by the addition of THF (30 mL) and triethylamine (0.10 g, 10 mmol). Fumaroyl chloride (1a, 1.6 g, 11 mmol) was added dropwise with stirring in ice-water bath, and the reaction mixture was stirred for 3 h. Aniline (0.86 g, 5 mmol) was then added, and the reaction was continued for 6 h. The reaction was monitored by TLC and stopped when no further progress was observed. The reaction mixture was filtered, and the solvent was removed under reduced pressure. The residue was dissolved in ethyl acetate and washed with petroleum ether (500 mL). A large amount of white solid precipitated, which was collected by filtration and dried over anhydrous sodium sulfate to afford the intermediate 1c, which was used directly in the next step without further purification. Intermediate 1c was dissolved in 20 mL of methanol (1d-1), and the solution was heated to reflux. After complete dissolution, 0.125 mL of concentrated sulfuric acid was added dropwise, and the reaction was continued for 0.5 h. At the end of the reaction, the reaction was reduced to room temperature and 100 mL of water was added. The crude of the target product 1 was filtered, and the crude product was subjected to column chromatography (eluting with a solution of 25% ethyl acetate in petroleum ether) to obtain the final product of 1. White powder, Yield: 70%. 1H NMR (600 MHz, DMSO-d6) δ 10.56 (s, 1H), 7.68 (d, J = 7.9 Hz, 2H), 7.35 (t, J = 7.6 Hz, 2H), 7.24 (d, J = 15.4 Hz, 1H), 7.11 (t, J = 7.1 Hz, 1H), 6.73 (d, J = 15.4 Hz, 1H), 3.76 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 165.84, 161.62, 138.93, 138.33, 129.45, 129.32, 124.54, 119.80, 52.51. HRMS (ESI): m/z calculated for C11H11NO3 [M+Na]+: 228.0637, found 228.0637.
General Synthetic Procedure for ethyl (E)-4-oxo-4-(phenylamino)but-2-enoate (2). Compound 2 was synthesized via the same route as that used for 1 as a White powder, Yield: 70%. 1H NMR (600 MHz, DMSO-d6) δ 10.56 (s, 1H), 7.68 (d, J = 7.7 Hz, 2H), 7.35 (t, J = 7.1 Hz, 2H), 7.22 (d, J = 15.4 Hz, 1H), 7.11 (t, J = 7.1 Hz, 1H), 6.71 (d, J = 15.4 Hz, 1H), 4.29–4.15 (m, 2H), 1.27 (t, J = 7.0 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 165.34, 161.67, 138.96, 138.14, 129.83, 129.34, 124.55, 119.80, 61.27, 14.48. HRMS (ESI): m/z calculated for C12H13NO3 [M+Na]+: 242.0793, found 242.0795.
General Synthetic Procedure for compound 1 g. Maleic anhydride (1e, 0.98 g, 10 mmol) was placed in a 100 mL round bottom flask and dissolved with ethyl acetate (30 mL), aniline (0.93 g, 10 mmol) was added to it and stirred at room temperature for 6 h. After the reaction was monitored by TLC until the end of the reaction, the insoluble material was filtered to give reaction intermediate 1 g. It was used directly for the next step without purification.
General Synthetic Procedure for methyl (Z)-4-oxo-4-(phenylamino)but-2-enoate (3). Intermediate 1 g was dissolved in 20 mL of methanol, heated to reflux, after dissolving, 0.125 mL of concentrated sulfuric acid was added dropwise, and the reaction was continued for 0.5 h. At the end of the reaction, the reaction was reduced to room temperature and 100 mL of water was added, the crude of the target product 3 was filtered, and the crude product was subjected to column chromatography (ethyl acetate: petroleum ether = 1:4) to obtain the final product of 7. White powder, Yield: 77%. 1H NMR (600 MHz, DMSO-d6) δ 10.28 (s, 1H), 7.60 (d, J = 7.3 Hz, 2H), 7.32 (d, J = 6.9 Hz, 2H), 7.07 (d, J = 6.7 Hz, 1H), 6.51 (d, J = 11.7 Hz, 1H), 6.41 (d, J = 11.7 Hz, 1H), 3.67 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 167.04, 162.94, 139.07, 132.70, 129.22, 124.12, 119.81, 52.04. HRMS (ESI): m/z calculated for C11H11NO3 [M+Na]+: 228.0637, found 228.0641.
General Synthetic Procedure for ethyl (Z)-4-oxo-4-(phenylamino)but-2-enoate (4). Compound 4 was synthesized via the same route as that used for 3 as a Yellow powder, Yield: 74%. 1H NMR (600 MHz, DMSO-d6) δ 10.27 (s, 1H), 7.61 (d, J = 7.9 Hz, 2H), 7.32 (t, J = 7.6 Hz, 2H), 7.07 (t, J = 7.1 Hz, 1H), 6.52 (d, J = 11.7 Hz, 1H), 6.37 (d, J = 11.7 Hz, 1H), 4.13 (dd, J = 13.9, 6.9 Hz, 2H), 1.18 (t, J = 7.0 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 167.19, 163.92, 139.95, 133.82, 130.01, 129.78, 124.86, 120.56, 61.49, 15.13. HRMS (ESI): m/z calculated for C12H13NO3 [M+Na]+: 242.0793, found 242.0795.
General Synthetic Procedure for propyl (Z)-4-oxo-4-(phenylamino)but-2-enoate (5). Compound 5 was synthesized via the same route as that used for 3 as a Yellow powder, Yield: 70%. 1H NMR (600 MHz, DMSO-d6) δ 10.27 (s, 1H), 7.61 (t, J = 7.3 Hz, 2H), 7.32 (t, J = 7.3 Hz, 2H), 7.07 (t, J = 6.8 Hz, 1H), 6.53 (d, J = 11.7 Hz, 1H), 6.37 (d, J = 11.8 Hz, 1H), 4.04 (dd, J = 12.6, 6.1 Hz, 2H), 1.57 (dq, J = 14.1, 7.1 Hz, 2H), 0.86 (q, J = 6.9 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.28, 162.98, 138.99, 132.98, 129.02, 128.63, 123.86, 119.54, 66.07, 21.61, 10.57. HRMS (ESI): m/z calculated for C13H15NO3 [M+Na]+: 256.0944, found 256.0941.
General Synthetic Procedure for isopropyl (Z)-4-oxo-4-(phenylamino)but-2-enoate (6). Compound 6 was synthesized via the same route as that used for 3 as a White powder, Yield: 70%. 1H NMR (600 MHz, DMSO-d6) δ 10.24 (s, 1H), 7.61 (d, J = 7.7 Hz, 2H), 7.32 (t, J = 7.8 Hz, 2H), 7.07 (t, J = 7.2 Hz, 1H), 6.51 (d, J = 11.8 Hz, 1H), 6.32 (d, J = 11.8 Hz, 1H), 5.07–4.84 (m, 1H), 1.17 (d, J = 6.2 Hz, 6H). 13C NMR (151 MHz, DMSO-d6) δ 165.79, 163.21, 139.21, 133.02, 129.20, 129.01, 123.99, 119.68, 68.13, 21.86. HRMS (ESI): m/z calculated for C13H15NO3 [M+Na]+: 256.0950, found 256.0951.
General Synthetic Procedure for (Z)-4-oxo-4-(phenylamino)but-2-enoic acid (7). Compound 7 was synthesized via the same route as that used for 3 as a White powder, Yield: 95%. 1H NMR (600 MHz, DMSO-d6) δ 13.16 (s, 1H), 10.41 (s, 1H), 7.64 (d, J = 7.7 Hz, 2H), 7.34 (t, J = 7.0 Hz, 2H), 7.10 (t, J = 6.8 Hz, 1H), 6.49 (d, J = 11.9 Hz, 1H), 6.32 (d, J = 11.9 Hz, 1H). 13C NMR (151 MHz, DMSO- d6) δ 167.16, 163.50, 138.80, 131.94, 130.69, 129.08, 124.14, 119.78. HRMS (ESI): m/z calculated for C10H9NO3 [M+Na]+: 214.0475, found 214.0473.
General Synthetic Procedure for methyl (Z)-4-oxo-4-(thiazol-2-ylamino)but-2-enoate (8). Compound 8 was synthesized via the same route as that used for 3 as a White powder, Yield: 70%. 1H NMR (600 MHz, DMSO-d6) δ 12.49 (s, 1H), 7.52 (s, 1H), 7.28 (s, 1H), 6.58 (d, J = 6.9 Hz, 2H), 3.70 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.65, 162.35, 157.72, 138.21, 131.49, 129.58, 114.34, 52.06. HRMS (ESI): m/z calculated for C8H8N2O3S [M+Na]+: 235.0153, found 235.0157.
General Synthetic Procedure for methyl (Z)-4-(ethylamino)-4-oxobut-2-enoate (9). Compound 9 was synthesized via the same route as that used for 3 as a Faint yellow liquid, Yield: 62%. 1H NMR (600 MHz, DMSO-d6) δ 8.17 (s, 1H), 6.25 (d, J = 11.8 Hz, 1H), 6.20 (d, J = 11.8 Hz, 1H), 3.61 (s, 3H), 3.13–3.03 (m, 2H), 1.01 (t, J = 7.1 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 171.90, 168.70, 137.50, 132.67, 56.57, 38.49,19.61. HRMS (ESI): m/z calculated for C7H11NO3 [M+Na]+: 180.0637, found 180.0639.
General Synthetic Procedure for N1-phenyl-N4-propylmaleamide (10). Compound 10 was synthesized first by the same route as compound 7 for compound 1j. Subsequently, compound 1j (1.15 g, 6 mmol) was dissolved in dry DCM, followed by the addition of EDCI (1.15 g, 6 mmol), the reaction was carried out for about 15 min followed by the addition of 0.30 g (5 mmol) of aniline (1b) reaction and the reaction was carried out for 1 h. At the end of the reaction, the solution was concentrated in vacuo, and then 100 mL of saturated saline was added and the product was extracted with ethyl acetate (3 × 50 mL). The combined organics were first washed with 2 N hydrochloric acid, water, 5 wt% sodium bicarbonate and then water. The organics were dried over magnesium sulfate, filtered, and concentrated in vacuum to give the crude product, which was subjected to column chromatography (ethyl acetate: petroleum ether = 1:4) to final product. White powder, Yield: 75%. 1H NMR (600 MHz, DMSO-d6) δ 11.39 (s, 1H), 8.72 (s, 1H), 7.62 (d, J = 7.8 Hz, 2H), 7.32 (t, J = 7.7 Hz, 2H), 7.07 (t, J = 7.2 Hz, 1H), 6.27 (s, 2H), 3.11 (dd, J = 12.7, 6.4 Hz, 2H), 1.47 (dd, J = 14.3, 7.2 Hz, 2H), 0.88 (t, J = 7.3 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 165.20, 163.58, 139.23, 133.28, 131.46, 129.23, 123.94, 119.68, 40.98, 22.48, 11.94. HRMS (ESI): m/z calculated for C13H16N2O2 [M+Na]+: 255.1109, found 255.1107.
General Synthetic Procedure for methyl (Z)-4-([1,1’-biphenyl]-4-ylamino)-4-oxobut-2-enoate (11). Compound 11 was synthesized via the same route as that used for 3 as White powder. 1H NMR (600 MHz, DMSO-d6) δ 10.40 (s, 1H), 7.72 (d, J = 8.0 Hz, 2H), 7.65 (d, J = 7.5 Hz, 4H), 7.45 (t, J = 7.0 Hz, 2H), 7.33 (t, J = 6.9 Hz, 1H), 6.55 (d, J = 11.7 Hz, 1H), 6.44 (d, J = 11.7 Hz, 1H), 3.68 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 167.04, 162.98, 140.06, 138.55, 135.79, 132.66, 129.34, 129.28, 127.53, 127.43, 126.71, 120.19, 52.06. HRMS (ESI): m/z calculated for C17H15NO3 [M+Na]+: 304.0954, found 304.0950.
General Synthetic Procedure for methyl (Z)-4-((4-methoxyphenyl)amino)-4-oxobut-2-enoate (12). Compound 12 was synthesized via the same route as that used for 3 as a Silver solid, Yield: 75%. 1H NMR (400 MHz, DMSO-d6) δ 10.14 (s, 1H), 7.57–7.48 (m, 2H), 6.93–6.85 (m, 2H), 6.47 (d, J = 11.8 Hz, 1H), 6.38 (d, J = 11.7 Hz, 1H), 3.72 (d, J = 1.3 Hz, 3H), 3.66 (d, J = 1.2 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 167.18, 162.37, 155.93, 132.48, 132.20, 129.08, 121.30, 114.34, 55.60, 52.01. HRMS (ESI): m/z calculated for C12H13NO4 [M+Na]+: 258.0742, found 258.0746.
General Synthetic Procedure for methyl (Z)-4-((4-chlorophenyl)amino)-4-oxobut-2-enoate (13). Compound 13 was synthesized via the same route as that used for 3 as a White powder, Yield: 55%. 1H NMR (600 MHz, DMSO-d6) δ 10.41 10.41 (s, 1H), 7.64 (s, 2H), 7.38 (s, 2H), 6.52 (d, J = 11.5 Hz, 1H), 6.42 (d, J = 11.3 Hz, 1H), 3.67 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 171.63, 167.87, 142.77, 137.45, 134.02, 133.89, 132.46, 126.11, 56.81. HRMS (ESI): m/z calculated for C11H10ClNO3 [M+Na]+: 262.0247, found 262.0251.
General Synthetic Procedure for methyl (Z)-4-((3-chlorophenyl)amino)-4-oxobut-2-enoate (14). Compound 14 was synthesized via the same route as that used for 3 as a White powder, Yield: 72%. 1H NMR (600 MHz, DMSO-d6) δ 10.49 (s, 1H), 7.84 (s, 1H), 7.47 (d, J = 7.7 Hz, 1H), 7.36 (t, J = 7.8 Hz, 1H), 7.15 (d, J = 7.4 Hz, 1H), 6.54 (d, J = 11.7 Hz, 1H), 6.44 (d, J = 11.7 Hz, 1H), 3.68 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.81, 163.39, 140.49, 133.54, 132.73, 130.95, 129.35, 123.84, 119.22, 118.19, 52.11. HRMS (ESI): m/z calculated for C11H10ClNO3 [M+Na]+: 262.0247, found 262.0251.
General Synthetic Procedure for methyl (Z)-4-((2-chlorophenyl)amino)-4-oxobut-2-enoate (15). Compound 15 was synthesized via the same route as that used for 3 as a White powder, Yield: 65%. 1H NMR (600 MHz, DMSO-d6) δ 9.94 (s, 1H), 7.74 (d, J = 7.5 Hz, 1H), 7.51 (d, J = 7.9 Hz, 1H), 7.35 (t, J = 7.5 Hz, 1H), 7.22 (t, J = 7.3 Hz, 1H), 6.65 (d, J = 11.7 Hz, 1H), 6.46 (d, J = 11.6 Hz, 1H), 3.66 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 167.06, 163.31, 135.52, 134.69, 131.94, 129.94, 127.82, 127.08, 126.92, 126.69, 52.04. HRMS (ESI): m/z calculated for C11H10ClNO3 [M+Na]+: 262.0247, found 262.0252.
General Synthetic Procedure for methyl (Z)-4-((4-bromophenyl)amino)-4-oxobut-2-enoate (16). Compound 16 was synthesized via the same route as that used for 3 as a White powder, Yield: 75%. 1H NMR (600 MHz, DMSO-d6) δ 10.43 (s, 1H), 7.60 (d, J = 8.1 Hz, 2H), 7.51 (d, J = 8.2 Hz, 2H), 6.52 (d, J = 11.7 Hz, 1H), 6.43 (d, J = 11.7 Hz, 1H), 3.67 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.90, 163.14, 138.44, 132.66, 132.06, 129.33, 121.71, 115.77, 52.08. HRMS (ESI): m/z calculated for C11H10BrNO3 [M+Na]+: 305.9742, found 305.9746.
General Synthetic Procedure for methyl (Z)-4-((4-cyanophenyl)amino)-4-oxobut-2-enoate (17). Compound 17 was synthesized via the same route as that used for 3 as a White powder, Yield: 80%. 1H NMR (600 MHz, DMSO-d6) δ 10.71 (s, 1H), 7.79 (s, 4H), 6.57 (d, J = 11.8 Hz, 1H), 6.46 (d, J = 11.8 Hz, 1H), 3.67 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.65, 163.84, 143.25, 133.76, 132.88, 129.49, 119.85, 119.41, 105.85, 52.14. HRMS (ESI): m/z calculated for C12H10N2O3 [M+Na]+: 253.0589, found 253.0590.
General Synthetic Procedure for methyl (Z)-4-oxo-4-((4-(trifluoromethyl)phenyl)amino)but-2-enoate (18). Compound 18 was synthesized via the same route as that used for 3 as a White powder, Yield: 70%. 1H NMR (600 MHz, DMSO-d6) δ 10.66 (s, 1H), 7.83 (d, J = 6.4 Hz, 2H), 7.70 (d, J = 8.3 Hz, 2H), 6.58 (d, J = 11.7 Hz, 1H), 6.47 (d, J = 11.8 Hz, 1H), 3.68 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.76, 163.63, 142.62, 132.75, 129.46, 127.43, 126.54, 125.63, 124.38, 124.17, 123.96, 123.83, 123.75, 122.03, 119.69, 117.44, 52.08. HRMS (ESI): m/z calculated for C12H10NO3F3 [M+Na]+: 296.0510, found 296.0516.
General Synthetic Procedure for methyl (Z)-4-((4-nitrophenyl)amino)-4-oxobut-2-enoate (19). Compound 19 was synthesized via the same route as that used for 3 as a White powder, Yield: 71%. 1H NMR (600 MHz, DMSO-d6) δ 10.87 (s, 1H), 8.22 (d, J = 5.2 Hz, 2H), 7.83 (d, J = 2.9 Hz, 2H), 6.57 (d, J = 10.2 Hz, 1H), 6.46 (d, J = 10.6 Hz, 1H), 3.65 (s, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.63, 163.96, 145.18, 142.90, 132.75, 129.72, 125.44, 119.52, 52.15. HRMS (ESI): m/z calculated for C11H10N2O3 [M+Na]+: 273.0487, found 273.0490.
General Synthetic Procedure for ethyl (Z)-4-((4-chlorophenyl)amino)-4-oxobut-2-enoate (20). Compound 20 was synthesized via the same route as that used for 3 as a White powder, Yield: 55%. 1H NMR (600 MHz, DMSO-d6) δ 10.41 (s, 1H), 7.65 (d, J = 8.6 Hz, 2H), 7.38 (d, J = 8.6 Hz, 2H), 6.53 (d, J = 11.8 Hz, 1H), 6.38 (d, J = 11.8 Hz, 1H), 4.13 (dd, J = 13.9, 6.9 Hz, 2H), 1.17 (t, J = 7.0 Hz, 3H). 13C NMR (151 MHz, DMSO-d6) δ 166.21, 163.29, 138.08, 132.98, 129.12, 129.05, 127.60, 121.25, 60.70, 14.29. HRMS (ESI): m/z calculated for C12H12ClNO3 [M+Na]+: 276.0403, found 276.0406.
General Synthetic Procedure for methyl (Z)-4-((4-ethynylphenyl)amino)-4-oxobut-2-enoate (21). Compound 21 was synthesized via the same route as that used for 3 as a White powder, Yield: 73%. 1H NMR (600 MHz, DMSO-d6) δ 10.45 (s, 1H), 7.63 (d, J = 8.3 Hz, 2H), 7.44 (d, J = 8.3 Hz, 2H), 6.52 (d, J = 11.7 Hz, 1H), 6.43 (d, J = 11.7 Hz, 1H), 4.11 (s, 1H), 3.67 (s, 3H).13C NMR (151 MHz, DMSO-d6) δ 166.91, 163.19, 139.60, 132.86, 132.66, 129.39, 119.64, 117.01, 83.91, 80.58, 52.09. HRMS (ESI): m/z calculated for C13H11NO3 [M+Na]+: 252.0637, found 252.0640.
General Synthetic Procedure for methyl (E)-4-((4-ethynylphenyl)amino)-4-oxobut-2-enoate (22). Compound 22 was synthesized via the same route as that used for 1 as a Faint yellow powder, Yield: 75%.1H NMR (600 MHz, DMSO-d6) δ 10.74 (s, 1H), 7.71 (d, J = 8.4 Hz, 2H), 7.48 (d, J = 8.4 Hz, 2H), 7.23 (d, J = 15.4 Hz, 1H), 6.75 (d, J = 15.4 Hz, 1H), 4.15 (s, 1H), 3.77 (s, 3H).13C NMR (151 MHz, DMSO-d6) δ 165.76, 161.81, 139.40, 137.98, 132.93, 129.77, 119.69, 117.45, 83.80, 80.79, 52.55. HRMS (ESI): m/z calculated for C13H11NO3 [M + H]+: 230.0817, found 230.0816.
General Synthetic Procedure for ethyl (Z)-4-((4-ethynylphenyl)amino)-4-oxobut-2-enoate (23). Compound 23 was synthesized via the same route as that used for 3 as a White powder, Yield: 59%. 1H NMR (600 MHz, DMSO-d6) δ 10.44 (s, 1H), 7.64 (d, J = 7.8 Hz, 2H), 7.44 (d, J = 7.7 Hz, 2H), 6.54 (d, J = 11.7 Hz, 1H), 6.38 (d, J = 11.7 Hz, 1H), 4.22 – 4.04 (m, 3H), 1.17 (t, J = 6.6 Hz, 3H).13C NMR (151 MHz, DMSO-d6) δ 166.20, 163.38, 139.66, 133.03, 132.84, 129.09, 119.59, 116.95, 83.92, 80.46, 60.73, 14.31. HRMS (ESI): m/z calculated for C14H13NO3 [M+Na]+: 266.0793, found 266.0796.
General Synthetic Procedure for methyl 4-((4-ethynylphenyl)amino)-4-oxobutanoate (24). Compound 24 was synthesized via the same route as that used for 3 as a White powder, Yield: 47%. 1H NMR (400 MHz, DMSO-d6) δ 10.13 (s, 1H), 7.61 (d, J = 8.9 Hz, 2H), 7.40 – 7.26 (m, 2H), 3.60 (s, 3H), 2.61 (s, 4H). 13C NMR (101 MHz, DMSO-d6) δ 173.32, 170.47, 138.65, 129.09, 126.97, 120.91, 51.87, 31.32, 28.86. HRMS (ESI): m/z calculated for C14H13NO3 [M+Na]+: 264.0398, found 264.0396.
NMR method to determine the half-life
An 18 μM solution of compound was prepared in an NMR tube using 500 μL of DMSO‑d6 as solvent. To this solution, 54 μM of reduced glutathione (GSH) dissolved in 100 μL of freshly prepared phosphate-buffered saline (PBS, pH 7.4) was added, followed by thorough mixing via rapid agitation. ¹H NMR spectra were acquired on a nuclear magnetic resonance spectrometer, with TMS in DMSO‑ d6 serving as the internal reference. The relative concentration of the sample was quantitatively determined by comparing the integrated signal area of characteristic proton resonances of compound to that of TMS. Let A0 represent the initial concentration of compound and A denote its concentration at time t. Temporal changes in concentration revealed first-order reaction kinetics between compound and GSH. The reaction rate constant k was derived from the linearized first-order rate equation: ln(A0/A) = kt. The half-life (t1/2) was subsequently calculated as: t1/2 = = ln(2)/k. For comparative purposes, control compounds including Nirmatrelvir, afatinib and disulfiram were subjected to the same reaction conditions with GSH, and their respective first-order rate constants and half-lives were determined. Additionally, the stability of all compounds was evaluated in both buffer and DMSO solutions under identical experimental conditions.
Protein expression and purification
The wild-type and mutant CaMrt4 proteins were expressed and purified using previously reported methods49. Briefly, the cDNA of Mrt4 (NCBI Number: XM_714020.2) was cloned into pET-28a vector (catalog no. P3110-20, Solarbio). Mutant constructs were generated using a two-step PCR method by introducing specific base changes into a double-stranded DNA plasmid. An eight-amino-acid sequence (SYYHHHHHH) was added to the N-terminus of Mrt4 proteins to facilitate purification. The plasmid was transformed into E. coli BL21 (DE3) cells for protein expression. The expressed proteins were purified using a HisTrap_FF_5 mL column (Cytiva) on an ÄKTA Pure chromatography system. Protein samples were diluted 3-5 times with 20 mM Tris-Cl, pH 7.5, and further purified using a Resource S cation exchange column (Cytiva). Final purification was achieved using a Superdex 75 size-exclusion chromatography column (Cytiva) equilibrated with 20 mM Tris-Cl, pH 7.5, and 150 mM NaCl. The eluted protein samples were stored in 25 mM Tris-HCl, pH 7.4, for subsequent experiments. For S. cerevisiae GAPDH (ScGAPDH), the GAPDH gene was cloned into the pET-28a expression vector. The enzyme was expressed in E. coli BL21 (DE3) cells and purified to homogeneity as previously described50. The eluted protein samples were stored in 25 mM Tris-HCl, pH 7.4, for enzymatic inhibition assays and protein crystallization.
In vitro inhibitory activity evaluation
The activity of recombinant ScGAPDH was measured spectrophotometrically with slight modifications to the method described by Cheng Luo51. Assays were performed in an assay buffer containing 100 mM HEPES (pH 7.4), 100 mM NaCl, 2 mM MgCl2, 0.1 mM DTT, 0.5 mM EDTA, and 1 mg/mL BSA. Enzymatic activity was monitored in kinetics mode using a 96-well plate format with a total reaction volume of 200 µL, and was determined using a coupled assay with water-soluble tetrazolium-8 (WST-8) and 1-methoxy-5-methylphenazium methyl sulfate (1-mPMS) reagent. Initial reaction velocities in the presence of test compounds were compared to those with DMSO (Control) to generate IC50 curves. IC50 or enzyme kinetic constant values were calculated using logistic equations (nonlinear analysis) or Hill kinetics equations in the Origin 7.5 program package.
Protein crystallization and structure determination
To obtain complex structures, the WT ScGAPDH protein (0.2 mg/mL at a final concentration) was incubated with 5 μM inhibitor for 40 min. The sample was then concentrated by ultrafiltration to 8 mg/mL (measured by absorbance at 280 nm). Crystals were grown at 18 °C using the hanging-drop vapor diffusion method. Each drop consisted of 1 µL of protein solution mixed with 1 µL well solution containing 0.2 M NH4Ac, 0.1 M Bis-Tris (pH = 7.0), and 24% v/v PEG3350. Crystals were cryoprotected using well solution supplemented with 20% glycerol and flash-frozen in liquid nitrogen.
X-ray diffraction data were collected at the BL19U1 and BL10U2 beamlines at the Shanghai Synchrotron Radiation Facility. The data were indexed, integrated, and scaled using XDS software packages52. Molecular replacement was performed with Phaser, and the structure was built using Coot and refined with PHENIX. Atomic restraints for the inhibitor were generated using eLBOW, and the final model was validated using MolProbity. Structural figures were created using PyMOL.
Preparation of cell lysates
C. albicans cells were grown in YEPD medium at 35 °C for 24 h. The cells were washed three times with PBS before adding 200 µL of RIPA lysate (P0013B, Beyotime) per 0.4 g of wet fungus. The samples were then transferred to a mortar and ground three times with liquid nitrogen. After grinding, the lysates were centrifuged at 12,000 rpm for 10 min at 4 °C. The resulting supernatant was collected as the total fungal protein lysate. The protein concentration was measured using a BCA protein assay kit (Beyotime) and adjusted to 2 mg/mL by dilution with PBS.
Pull-down experiment
C. albicans cells were cultured until reaching 70–80% confluence. Cells were lysed using RIPA buffer, and sonication on ice was performed to ensure complete lysis. Total protein concentration was determined by the Bradford method and adjusted to 2 mg/mL with RIPA lysate after centrifugation. Cell lysates (2 mg/mL at a final concentration) were incubated with probe for 2 h. To initiate the reaction, 12 μL of sodium ascorbate solution (10 mM), 12 μL of biotin-PEG3-azide (0.5 mM) and 36 μL of premixed BTTAA (2 mM) /CuSO4 (1 mM) solution were added to 300 μL of the supernatant. Following the reaction, 1.5 mL of pre-cooled acetone was added to precipitate the proteins, which were separated by centrifugation at 6500 × g for 5 min, air-dried, and resuspended in 1 mL of RIRA lysate using ultrasound. The samples were washed three times with 1% SDS/PBS, 6 M urea, and PBS. The final pellet containing specifically bound protein was collected and analyzed by 12% SDS-PAGE for subsequent western blot analysis.
Western blot analysis
Denatured protein samples were resolved on 12% SDS-PAGE gels, and then the separated proteins were transferred to PVDF membranes. Membranes were washed three times with Tris-buffered saline containing 0.1% Tween 20 (TBST), and then blocked with 5% nonfat dried milk in TBST for 1.5 h. For specific detection of biotin-labeled proteins, the membrane was incubated overnight at 4 °C with streptavidin−horseradish peroxidase (HRP, Beyotime) diluted 1:5000 in 5% nonfat dried milk TBST. After incubation, the membrane was washed with PBST, and signals were visualized using enhanced chemiluminescence (ECL) reagents (Beyotime) and detected with a ChemiDoc XRS + system (Bio-Rad). Uncropped and unprocessed scans of the most important blots (Fig. 2b, c, and Supplementary Figs. 11a, b, 12a, 14a, b) are provided in the Source data file.
Transcriptome analysis and qPCR
C. albicans SC5314 cells (2 × 105 cells/mL) were cultured in RPMI-1640 medium with 4 µg/mL 20; cells cultured without 20 served as control. After 6 h, the cells were harvested and washed three times with PBS. Total RNA was extracted using the RiboPure-Yeast Kit (Roth) following the manufacturer’s instructions. RNA integrity was assessed using an Agilent 2100 Bioanalyzer (Agilent), and samples with RNA integrity number (RIN) > 8.0 were used for sequencing. cDNA libraries were prepared and sequenced by Novogene Corporation (Tianjin, China) using the Illumina HiSeqTM X platform. The RNA-Seq data were deposited in the Gene Expression Omnibus (GEO) under accession number GSE209608. For quantitative PCR (qPCR), purified RNA was reverse-transcribed into cDNA using the PrimeScriptTM RT reagent Kit (Takara). Gene expression analysis was performed with TB Green® Premix Ex TaqTM (Takara) in an Eppendorf Mastercycler Real-Time PCR System. The PCR protocol included an initial denaturation step at 95 °C for 20 m, followed by 40 cycles of denaturation at 95 °C for 20 s, annealing at 60 °C for 20 s, and primer extension at 72 °C for 10 s. The transcript levels of hyphal formation-related genes were calculated using the 2−ΔΔCT method.
In vitro antifungal assays
The broth microdilution method, as outlined in the CLSI M27A353 and M38A2 guidelines, was used to test the fungicidal activity (MIC80) of various Candida species. MIC80 is defined as the minimum concentration of a compound required to inhibit fungal growth by more than 80% compared to untreated controls. The C. albicans strain SC5314, C. auris CBS12774, and fluconazole-resistant Candida strains (103, GU5) were obtained from the Second Military Medical University. Fluconazole was used as the quality control compound, and C. parapsilosis (ATCC22019) was used as the quality control strain to ensure the accuracy and reproducibility.
Test compounds were dissolved in DMSO and serially diluted across a concentration range of 0.125 to 64.0 μL/mL for checkerboard microdilution assays. DMSO served as a negative control, FCZ was used as a positive control and wells without Candida species were used as blank controls. Microtiter plates containing Candida species were incubated at 35 °C for 48 h, then the OD values at 630 nm were measured. MIC80 was defined as the lowest concentration of a compound resulting in an OD630 reduction of more than 80% compared to the negative control. When the MIC80 value exceeded the tested concentration range, the following calculation methods were used: if MIC80 was higher than 64 mg/L, it was recorded as “ >64 mg/L”; if MIC80 was at or below the lowest concentration, it was recorded as “≤0.125 mg/L”.
Ames tests and hemolysis assays
The mutagenic potential of compound 20 was assessed using the standard bacterial reverse mutation assay (Ames test) with Salmonella typhimurium strains TA1537, TA98, TA102, and TA1535, in compliance with NMPA, ICH, and OECD guidelines. Tests were performed both in the presence (+S9) and absence (−S9) of a metabolic activation system (Aroclor 1254-induced rat liver S9). Compound 20, dissolved in DMSO, was evaluated over multiple doses. Concurrent controls included the vehicle (DMSO) groups and established positive mutagens (e.g., 2-AA, 2-AF with S9; MMS, 4-NO2-OPD without S9). Revertant colony counts on minimal glucose agar plates supplemented with trace histidine/biotin quantified mutagenesis for each strain/dose combination.
An in vitro hemolysis assay was performed using rabbit red blood cells (RBCs) in accordance with National Medical Products Administration (NMPA)guidelines. Fresh blood (5 mL) was collected from New Zealand rabbit jugular veins, defibrinated by shaking with glass beads, and washed three times with 0.9% saline. A 2% (v/v) RBC suspension was prepared in saline. Seven test tubes were allocated: five contained graded volumes of test compound 20 (0.02–0.32 mL; final concentration 0.5 mg/mL), one negative control (0.9% saline), and one positive control (deionized water). All tubes received 2.5 mL of RBC suspension and saline to equalize volumes (total volume 5 mL). Hemolysis were monitored visually after incubation at 37 °C for 3 h. Hemolysis was defined as transparent red supernatant and the test was valid only if negative controls showed no hemolysis/aggregation and positive controls exhibited complete hemolysis.
Preparation for ABPP and AIBPP-based LC-MS
All pull-down enriched samples were prepared for mass spectrometry by gel electrophoresis. Electrophoresis was performed in a specialized tank designed for mass spectrometry applications. The process was stopped once the Marker enters the separation gel. The gel was washed with high-purity water and stain with Caulmers Brilliant Blue. The entire lane was excised, and peptide samples were prepared through a series of steps, including decolourisation, reduction, alkylation and trypsin digestion.
LC-MS/MS measurement and analysis
Above spin-dried peptide samples were resuspended in an aqueous solution containing 0.1% (v/v) formic acid and analyzed by LC-MS/MS, LC was separated by a low pH mobile phase using an EASY-nLC 1200 liquid phase tandem QE HF-X mass spectrometer (Thermo Fisher Scientific). EASY-nLC 1200 UPLC system, mobile phase liquid A: 0.1% (v/v) formic acid aqueous solution, liquid B: 0.1% formic acid acetonitrile solution. The columns were: reversed-phase analytical column (25 cm, 75 μm id.). The column was a reversed-phase column (25 cm, 75 μm id.).
The gradient was set as follows: 5 s, Liquid B: 3–5%; 5 s–53 min 36 s, Liquid B: 5–15%; 53 min 36 s–100 min 15 s, Liquid B: 15–28%; 100 min 16 s–116 min 55 s, Liquid B: 28–38%; Liquid B was increased to 100% within 5 s and maintained for 8 min. The constant flow rate was 300 nL/min. The mass spectrometry acquisition conditions were as follows: positive ion mode, Orbitrap mass analyser, primary spectrum scanning range of 350 to 1800 Da, resolution of 60,000, profile format, secondary spectrum data acquisition in data-dependent mode, centroid format, and the 20 ion peaks with the highest intensities were taken for the secondary HCD (high-energy collision induced). The resolution of the secondary spectrum is 15,000. Other parameters to be set include isolation window, 1.6 m/z units; default charge, 2+; normalized collision energy, 28%; maximum IT, 50 ms; dynamic exclusion, 30.0 s. Quantitative analysis of the spectral data was performed using Thermo Proteome Discoverer 2.5 with library search capabilities. The mass spectrometry experiments comprised four independent sets. One set, analyzed in Supplementary Fig. 11, Supplementary Tables 5 and 6, included three samples (one DMSO-treated control) and was performed with one biological replicate (n = 1). The other three sets were each conducted with two biological replicates (n = 2): one set of four samples (two replicates of 22-treated samples as control) was analyzed in Fig. 3d; another set of four samples (two DMSO-treated controls) was analyzed in Supplementary Fig. 12b; and a final set of four samples (controls pretreated with 20 followed by 21 treatment) was analyzed in Supplementary Fig. 12c.
Binding energy calculation
Since the crystal structures of those proteins from C. albicans have yet to be determined, we used AlphaFold3 for the prediction of 3D protein structures. Gold docking calculation was performed for the determination of covalent binding structure. Several binding poses were obtained for each protein. In order to improve the accuracy, we further used the Cov_DOX method previously developed by our group29 to calculate the binding energy of each binding poses. The Cov_DOX method is a sophisticated combination of multiple potential energy surface theories including semi-empirical quantum chemistry theory and first principle density functional theory which provides reliable electronic structure description for the covalent ligand binding. Specifically, ωB97xD/6-311 + G**(high level)/PM7(low level) level of XO2 calculation upon PM7 optimized protein-ligand complex structure was performed for binding energy calculation.
Construction of heterozygous mutant strains
PCR amplification was used to generate a ~3700 bp fusion fragment containing upstream and downstream regions of the C. albicans MRT4 gene and the HIS1 selection marker (~2300 bp). A single colony of C. albicans SN152 (genotypes: LEU2/leu2Δ, HIS1/his1Δ and ARG4/arg4Δ) was inoculated into 3 mL YPD medium and incubated overnight at 30 °C with oscillation at 200 rpm. The fusion fragments were mixed with the activated fungal strains and electroporated using an electroporation cup. Forty eight transformants were selected on histidine dropout plates, purified, and enriched. Genomic DNA was extracted from the transformants, and successful integration of construct was validated using two pairs of primers. All primer sequences have been documented in Supplementary Table 15. The S. cerevisiae mutant strains were acquired from the Yeast Heterozygous Collection (Horizon Discovery). Gene deletions in this collection were constructed using a PCR-based strategy that precisely replaces each open reading frame from the start to stop codon with a KanMX module. The KanMX module contains the KanMX selectable marker and is flanked by unique 20-nucleotide molecular barcodes (UPTAG and DNTAG) for strain-specific identification. Each deletion was verified by polymerase chain reaction (PCR) amplification and analysis. An overview of the Saccharomyces Genome Deletion Project can be found at: http://sequence-www.stanford.edu/group/yeast_deletion_project/deletions3.html.
Oligonucleotide synthesis
The ssRNA probe (corresponding to 25S rRNA positions 1217–1286) with a 3′-terminal 5-carboxyfluorescein (5-FAM) modification, as well as all primers listed in Supplementary Table 15, were synthesized by General Biol Co., Ltd using solid-phase phosphoramidite chemistry. The procedure consisted of the following key steps: oligonucleotide chain assembly on an automated synthesizer, cleavage from the solid support and deprotection by ammonolysis, purification via HPLC, and quality control by LC-MS to confirm molecular weight and purity (Supplementary Fig. 22). The sequence is 5′-UCUAGACAGCCGGACGGUGGCCAUGGAAGUCGGAAUCCGCUAAGGAGUGUGUAACAACUCACCGGCCGAA-3′. The synthesized RNA probe was utilized for subsequent EMSA and FP experiments. All oligonucleotides described in this study are available to the research community upon request.
Electrophoretic mobility shift assay (EMSA)
Five micromolar proteins were incubated with or without test compounds pretreated for 1 h at 4 °C. subsequently, the protein samples were incubated with 100 nM FAM-labeled ssRNA probe in binding reactions containing 25 mM Tris-HCl, pH 8.0, 5 mM MgCl2 and 40 mM NaCl for 30 min at 25 °C. The binding reactions were resolved on 6% native acrylamide gels (37.5: 1 acrylamide: bis-acrylamide) in 0.5 × Tris-glycine buffer at 15 V/cm for 1.5 h. Gels were visualized using a Sapphire RGB screen on a Sapphire Biomolecular Imager (Azure Biosystems).
Fluorescence polarization assay
The FAM-labeled ssRNA probe of the GAR structural domain (1217-1286 nt) was used in the FP assay. To determine the binding affinities of RNA-FAM for CaMrt4 protein, serial dilutions of CaMrt4 was incubated with each compound at the indicated final concentrations for 1 h at 37 °C for 30 min. Then, RNA-FAM at a final concentration of 5 nM was added to initiate the reaction. Fluorescence polarization was measured for each reaction and plotted using GraphPad Prism software to deduce dissociation constants (Kd) after fitting to a one-site binding equation.
To determine the IC50 of compounds inhibiting the RNA-CaMrt4 interaction, serial dilution compounds were added to 3 µM CaMrt4 with incubation at 37 °C for 30 min. After that, a reaction mixture containing 50 mM HEPES buffer (pH = 7.5), and 5 nM ssRNA probe was added with a final volume of 60 μL. Subsequently, FP was measured on a Microplate Reader at the wavelength of 480 nm for excitation and 520 nm for emission, respectively. The FP values in the presence of the compound was normalized according to the equation percentage inhibition = (Max – Signal)/(Max – Min). The data were fit in GraphPad Prism 9.0 to obtain IC50 values with nonlinear regression analysis using equation, Y = 100/ (1 + 10^ ((logIC50 – X)*Hill Slope)), where Y stands for normalized FP value and X stands for compound concentration. The IC50 values were calculated by GraphPad Prism 9.0.
Sucrose gradient analysis
For polysome profile assays, cultures were collected at an OD600 of 0.5–0.6. C. albicans SC5314 (5 × 106 cells/mL) cells were prepared in RPMI 1640 medium and incubated with 4 μg/mL compound 20 at 30 °C for 3 h. At the end of incubation, cycloheximide (0.1 mg/mL final concentration) was added in growth media for 5 min on ice. Then, the cells were washed twice with ice-cold PBS supplemented with 0.1 mg/mL of cycloheximide, lysed with lysis buffer (QingZebio, Guangzhou, China) and centrifuged. Extracts were loaded onto linear 10–45% sucrose gradients in polysome lysis buffer. After a 3 h spin at 36,000 rpm in a rotor, gradient fractions were collected on a density gradient fractionator (Biocomp, Canada), continuously measuring absorbance at 260 nm.
Molecular dynamics simulations
The protein complex of wild type and the Cys96 covalent modification were set up via the CHARMM-GUI webserver based on the Cryo-EM structure (PDB ID: 7UQZ), respectively. Briefly, the protein complex was solvated in a cubic water box that extends at least 12.0 Å from protein surface, and K+ and Cl− were added to obtain a salinity of roughly 0.1 M and maintain neutral total charge. This yields a total of around 92,550 atoms in each simulation box. The lead compound parameters were parameterized via CHARMM-GUI in accordance with the CGENFF36 force fields, and the related topologies were manually edited. Parallel simulations are performed simultaneously using GROMACS 202154 in CHARMM36 force fields, with water molecules described by the TIP3P model. All protein covalent H-bonds were constrained with the LINCS algorithm. and long-range electrostatic interactions are treated with the particle-mesh Ewald method with a real-space cutoff of 12.0 Å. The system was first energy-minimized by 125,000 steps by using the steepest descent method to make the system reach the initial equilibrium structure. Then a 1 ns of NVT heating process was conducted with temperature control by Langevin dynamics with a friction coefficient of 1 ps−1. The system is equilibrated for 10 ns in the NPT ensemble at 1 atm and 300 K, and the production run continues for another 200 ns with snapshot saved every 50 ps.
Inter-residue distances and clustering were calculated using GROMACS 2021. we performed MMGBSA analysis on the last 100 ns MD trajectories (2000 snapshots) via gmx-MMPBSA. The contribution of all amino acids within 10 angstroms from the binding interface to the binding energy was collected, and any residue with absolute contribution large than 0.5 kcal/mol was reported.
Anti-biofilm assay
The overnight cultured C. albicans SC5314 cells were diluted to 1 × 106 CFU/mL with RPMI 1640 medium. A 100 μL aliquot of the cell suspension was added to each well of a 96-well plate, and 20 were tested at concentrations of 16, 8, 4, 2, 1 μg/mL, with a blank control group included. The plates were incubated at 37 °C for 24 h. The upper layer of medium was discarded and the bottom of the plate was washed three times gently with 100 μL of PBS. Following washing, 100 μL of PBS was added to each well, and the growth of periplasm was photographed with an inverted microscope. Subsequently, 10 μL of CCK-8 reagent was added into each well, and OD450 values were measured using a microplate reader 1 h of incubation at at 37 °C. The staining results were also photographed.
Cell cytotoxicity assay
The cytotoxic activity of 20 was tested by CCK-8 assay using human normal hepatocytes cell line (LO2). Cells were seeded in 96-well plates, at a density of 4 × 103 cells/well. After overnight incubation at 37 °C to allow adherence, varying concentrations of 20 or DMSO were added to the wells for 24 h. Detection reagent was then added, and the plates were incubated for 4 h at 37 °C. Absorbance was measured at 450 nm using a microplate reader (Molecular Devices, CA, USA), and the CC50 value was calculated. 293T (Product number: STCC10301P), HUVEC (Product number: STCC12103P), BEAS-2B (Product number: STCC10202P) were obtained from Servicebio Co., Ltd., and LO2 (Product number: SNL-141) was obtained from SUNCELL Technology Co., Ltd.
Plasma stability assay
Mouse plasma was treated with both compound 20 (10 μM) and JZL184 (0-40 μM) at 37 °C with shaking for 0–120 min. After incubation, samples were mixed with four times their volume of ice-cold acetonitrile. The remaining concentration of each compound was determined by LC-MS/MS (Triple Quad 6500+, SCIEX) and expressed as the percentage remaining, calculated by comparing the sample/IS peak area ratio at each time point to that at 0 min. The percent remaining at individual time points was transformed as a natural log and plotted as In(% remaining) vs. time. half-life was determined as half- life = In(2)/-slope.
In vivo antifungal activity assay
The G. mellonella were randomly divided into a growth control group (no drug administration after fungal infection), a positive drug control group (fluconazole administration after fungal infection), and a compound 20 administration group, with 12 G. Mellonella in each group. The overnight cultured C. albicans SC5314 was diluted with PBS to 2 × 107 CFU/mL. Ten microliters was injected into the G. mellonella at the ventral peduncle with a microfuge, and the infection was successful when the body of the G. mellonella was black or a distinct black line appeared on the back. The drug to be tested was diluted with PBS to the required concentration, and 10 μL of the drug solution was injected into the other abdominal legs of the G. mellonella using a microsampler, while the growth control group was injected with the same volume of PBS solution. After all the G. mellonella were treated, they were placed in an incubator at 30 °C for observation and recording. Growth was observed and recorded every 24 h. G. mellonella were considered dead if they were black and did not respond to external stimuli, and survival curves were plotted with GraphPad Prism 8 and analyzed using the Mantel-Cox model.
All animal experiments were approved by the Committee on the Ethics of Medicine of the Second Military Medical University (Certificate: SCXK-2024-0004). Female ICR mice (Mus musculus; outbred stock; 4–6 weeks old; 18–22 g) were purchased from the Chinese Center for Type Culture Collection (CCTC; Certificate No. SYXK-2022-1248-0011). Drug-resistant C. albicans cells 0304103 at the late stage of exponential growth were resuspended in saline for injection to achieve a fungal concentration of 3.0 × 105 CFU/mL. 0.2 mL of the above fungal suspension was inoculated into mice by tail vein injection to establish an invasive Candida infection model. After infection, mice were randomly divided into 10 groups: blank solvent group (Control), compound group (20 and 21), set at three different dose groups of 5.0, 10, and 20 mg/kg, respectively), and FLC group (set at three different dose groups of 5.0, 10, and 20 mg/kg), with five mice in each group. The drug was administered by intraperitoneal injection of 0.2 mL per mouse per time at 2.0 h after infection for 5 consecutive days, and then on day 6, the mice were executed and the left kidney was removed and placed in a 2.0 mL tissue grinder tube with 1.0 mL of saline. Use a tissue grinder to uniformly grind the kidney tissue, take an appropriate amount of tissue homogenate and dilute it in blank saline (10,000-fold dilution), mix it well, and then aspirate 50 µL of the diluted tissue homogenate and evenly spread it on SDA solid medium containing chloramphenicol. The petri dishes were incubated in a constant temperature incubator at 35 °C for 48 h and counted after complete colony formation in the medium, thus calculating the number of C. albicans per unit mass of kidney tissue. Data were graphically processed using GraphPad Prism, and the variability between groups of data was analyzed by One-way ANOVA.
For mouse kidney pathology experiments, we obtained the right kidney before fixation by 4% paraformaldehyde solution. It was then processed according to conventional methods, including dehydration, paraffin embedding, making 4.0 μm thick sections and HE staining. Observations were made under a light microscope and photographs were recorded.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The X-ray crystallographic coordinates and structure factors are available from the Protein Data Bank with accession numbers 9IWD and 9XB4, respectively. The RNA-seq data for the transcriptomic analysis generated in this study have been deposited in the Gene Expression Omnibus (GEO) under the accession code GSE302228. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD068797 and PXD073550. The data that support the findings of this study are available within the paper and its Supplementary Information files. Source data are provided with this paper.
Code availability
The molecular dynamics simulations in this work utilized the following publicly available software packages: GROMACS, Plumed (https://www.plumed.org), gmx_MMPBSA (https://valdes-tresancoms.github.io/gmx_MMPBSA), and Qbics (https://www.qbics.info/).
References
Denning, D. W. Global incidence and mortality of severe fungal disease. Lancet Infect. Dis. 24, e428–e438 (2024).
Say, R. F. & Fuchs, G. Fructose 1,6-bisphosphate aldolase/phosphatase may be an ancestral gluconeogenic enzyme. Nature 464, 1077–1081 (2010).
Cowen, L. E. The evolution of fungal drug resistance: modulating the trajectory from genotype to phenotype. Nat. Rev. Microbiol. 6, 187–198 (2008).
Sarma, S. & Upadhyay, S. Current perspective on emergence, diagnosis and drug resistance in Candida auris. Infect. Drug Resist. 10, 155–165 (2017).
Hoenigl, M. et al. COVID-19-associated fungal infections. Nat. Microbiol. 7, 1127–1140 (2022).
Perfect, J. R. The antifungal pipeline: a reality check. Nat. Rev. Drug Discov. 16, 603–616 (2017).
Liu, N., Tu, J., Dong, G., Wang, Y. & Sheng, C. Emerging new targets for the treatment of resistant fungal infections. J. Med. Chem. 61, 5484–5511 (2018).
Fisher, M. C., Hawkins, N. J., Sanglard, D. & Gurr, S. J. Worldwide emergence of resistance to antifungal drugs challenges human health and food security. Science 360, 739–742 (2018).
Lee, Y., Puumala, E., Robbins, N. & Cowen, L. E. Antifungal drug resistance: molecular mechanisms in candida albicans and beyond. Chem. Rev. 121, 3390–3411 (2021).
Noda, S., Shirai, T., Mori, Y., Oyama, S. & Kondo, A. Engineering a synthetic pathway for maleate in Escherichia coli. Nat. Commun. 8, 1153 (2017).
Zaro, B. W., Whitby, L. R., Lum, K. M. & Cravatt, B. F. Metabolically labile fumarate esters impart kinetic selectivity to irreversible inhibitors. J. Am. Chem. Soc. 138, 15841–15844 (2016).
Becker, D. et al. Irreversible inhibitors of the 3C protease of Coxsackie virus through templated assembly of protein-binding fragments. Nat. Commun. 7, 12761 (2016).
Kornberg, M. D. et al. Dimethyl fumarate targets GAPDH and aerobic glycolysis to modulate immunity. Science 360, 449–453 (2018).
Blair, H. A. Dimethyl fumarate: a review in relapsing-remitting MS. Drugs 79, 1965–1976 (2019).
Bharate, S. S. Recent developments in pharmaceutical salts: FDA approvals from 2015 to 2019. Drug Discov. Today 26, 384–398 (2021).
Maruyama, H. B. et al. New antibiotic, fumaramidmycin .1. Production, biological properties and characterization of producer strain. J. Antibiot. 28, 636–647 (1975).
Bello, I. A., Ndukwe, G. I., Amupitan, J. O., Ayo, R. G. & Shode, F. O. Syntheses and biological activity of some derivatives of C-9154 antibiotic. Int. J. Med. Chem. 2012, 148235 (2012).
Ma, T. et al. Low-dose metformin targets the lysosomal AMPK pathway through PEN2. Nature 603, 159–165 (2022).
Bird, R. E., Lemmel, S. A., Yu, X. & Zhou, Q. A. Bioorthogonal chemistry and its applications. Bioconjug. Chem. 32, 2457–2479 (2021).
Johansson, H. et al. Fragment-based covalent ligand screening enables rapid discovery of inhibitors for the RBR E3 ubiquitin ligase HOIP. J. Am. Chem. Soc. 141, 2703–2712 (2019).
Borsari, C. et al. Covalent proximity scanning of a distal cysteine to target PI3Kalpha. J. Am. Chem. Soc. 144, 6326–6342 (2022).
Abdeldayem, A., Raouf, Y. S., Constantinescu, S. N., Moriggl, R. & Gunning, P. T. Advances in covalent kinase inhibitors. Chem. Soc. Rev. 49, 2617–2687 (2020).
Wen, W. et al. N-acylamino saccharin as an emerging cysteine-directed covalent warhead and its application in the identification of novel fbpase inhibitors toward glucose reduction. J. Med. Chem. 65, 9126–9143 (2022).
Owen, D. R. et al. An oral SARS-CoV-2 M(pro) inhibitor clinical candidate for the treatment of COVID-19. Science 374, 1586–1593 (2021).
Lanning, B. R. et al. A road map to evaluate the proteome-wide selectivity of covalent kinase inhibitors. Nat. Chem. Biol. 10, 760–767 (2014).
Wen, W. et al. Structure-guided discovery of the novel covalent allosteric site and covalent inhibitors of fructose-1,6-bisphosphate aldolase to overcome the azole resistance of candidiasis. J. Med. Chem. 65, 2656–2674 (2022).
Han, X. et al. Structure-based rational design of novel inhibitors against fructose-1,6-bisphosphate aldolase from Candida albicans. J. Chem. Inf. Model. 57, 1426–1438 (2017).
Chen, X. et al. Potential antifungal targets based on glucose metabolism pathways of Candida albicans. Front. Microbiol. 11, 296 (2020).
Wei, L. et al. Cov_DOX: a method for structure prediction of covalent protein-ligand bindings. J. Med. Chem. 65, 5528–5538 (2022).
Hoepfner, D. et al. High-resolution chemical dissection of a model eukaryote reveals targets, pathways and gene functions. Microbiol. Res. 169, 107–120 (2014).
Baetz, K. et al. Yeast genome-wide drug-induced haploinsufficiency screen to determine drug mode of action. Proc. Natl. Acad Sci. USA 101, 4525–4530 (2004).
McLellan, C. A. et al. Inhibiting mitochondrial phosphate transport as an unexploited antifungal strategy. Nat. Chem. Biol. 14, 135–141 (2018).
Li, Z. et al. Nuclear export of pre-60S particles through the nuclear pore complex. Nature 618, 411–418 (2023).
Sarkar, A., Pech, M., Thoms, M., Beckmann, R. & Hurt, E. Ribosome-stalk biogenesis is coupled with recruitment of nuclear-export factor to the nascent 60S subunit. Nat. Struct. Mol. Biol. 23, 1074–1082 (2016).
Lo, K. Y., Li, Z., Wang, F., Marcotte, E. M. & Johnson, A. W. Ribosome stalk assembly requires the dual-specificity phosphatase Yvh1 for the exchange of Mrt4 with P0. J. Cell Biol. 186, 849–862 (2009).
Laserna, V. et al. Dichloro butenediamides as irreversible site-selective protein conjugation reagent. Angew. Chem. Int. Ed. Engl. 60, 23750–23755 (2021).
Fuchs, B. B., O’Brien, E., El Khoury, J. B. & Mylonakis, E. Methods for usingGalleria mellonellaas a model host to study fungal pathogenesis. Virulence 1, 475–482 (2014).
Galbiati, A., Zana, A. & Conti, P. Covalent inhibitors of GAPDH: From unspecific warheads to selective compounds. Eur. J. Med. Chem. 207, 112740 (2020).
Odds, F. C., Brown, A. J. P. & Gow, N. A. R. Antifungal agents: mechanisms of action. Trends Microbiol. 11, 272–279 (2003).
Shastry, M. et al. Species-specific inhibition of fungal protein synthesis by sordarin: identification of a sordarin-specificity region in eukaryotic elongation factor 2The GenBank accession numbers for the sequences reported in this manuscript are AF107286–AF107291, AF292693 and AF248644. Microbiology 147, 383–390 (2001).
Ruggero, D. et al. Dyskeratosis congenita and cancer in mice deficient in ribosomal RNA modification. Science 299, 259–262 (2003).
Cao, X. et al. Nascent alt-protein chemoproteomics reveals a pre-60S assembly checkpoint inhibitor. Nat. Chem. Biol. 18, 643–651 (2022).
Pelletier, J., Thomas, G. & Volarevic, S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat. Rev. Cancer 18, 51–63 (2018).
Nyhus, C., Pihl, M., Hyttel, P. & Hall, V. J. Evidence for nucleolar dysfunction in Alzheimer’s disease. Rev. Neurosci. 30, 685–700 (2019).
Kawashima, S. A. et al. Potent, reversible, and specific chemical inhibitors of eukaryotic ribosome biogenesis. Cell 167, 512–524 e14 (2016).
Thomson, E., Ferreira-Cerca, S. & Hurt, E. Eukaryotic ribosome biogenesis at a glance. J. Cell Sci. 126, 4815–4821 (2013).
Nikolay, R., Schmidt, S., Schlömer, R., Deuerling, E. & Nierhaus, K. Ribosome assembly as antimicrobial target. Antibiotics 5, 18 (2016).
Stokes, J. M. & Brown, E. D. Chemical modulators of ribosome biogenesis as biological probes. Nat. Chem. Biol. 11, 924–932 (2015).
Kashif, M. et al. Recombinant expression and biophysical characterization of Mrt4 protein that involved in mRNA turnover and ribosome assembly from Saccharomyces cerevisiae. Bioengineered 13, 9103–9113 (2022).
Park, J. B., Park, H., Son, J., Ha, S. J. & Cho, H. S. Structural study of monomethyl fumarate-bound human GAPDH. Mol. Cells 42, 597–603 (2019).
Li, W. et al. Natural product 1,2,3,4,6-penta-O-galloyl-beta-D-glucopyranose is a reversible inhibitor of glyceraldehyde 3-phosphate dehydrogenase. Acta Pharmacol. Sin. B 43, 470–482 (2022).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Merkulova, M. et al. Aldolase directly interacts with ARNO and modulates cell morphology and acidic vesicle distribution. Am. J. Physiol. Cell Physiol. 300, C1442–C1455 (2011).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Acknowledgements
This work was supported by the National Key R&D Program of China (2023YFE0110100, Y.R.), Natural Science Foundation of China (Nos. 22377030, Y.R., 82330109, C.S., 22177036, J.W., 22373039, L.R., 92269102, Y.W. and 82273975, W.C.), the Program for the PCSIRT (No. IRT0953, J.W.). Additional support was provided by Guangdong Pearl River Talent Program (2021QN02Y618, Y.W.) and the Program of Introducing Talents of Discipline to Universities of China (111 Program, B17019, J.W.). We thank the staff members of Mass Spectrometry System at the National Facility for Protein Science in Shanghai (NFPS), Zhangjiang Lab, China for providing technical support and assistance in data collection and analysis. We also acknowledge the staff of beamline BL10U2 at the Shanghai Synchrotron Radiation Facility (SSRF) and BL19U1 beamlines at NFPS, for technical support in X-ray diffraction data collection and analysis. We are grateful to ChomiX biotech (Nanjing, China) for proteomics support. Y.W. acknowledges computational works carried out at the Shenzhen Bay Laboratory Supercomputing Centre.
Author information
Authors and Affiliations
Contributions
Y.R., C.S., W.C., and J.W. conceived and designed the experiments. All authors contributed to this work as follows: H.C., J.T., J.C., B.C., Z.H., and Y.W. were contributed equally. Y.R. and H.C. edit the manuscript. H.C. performed the target identification, co-crystal structure determination; Z.H. and J.B. performed the synthesis of hit compounds; J.T. contributed the murine candidiasis models assays; J.C. and W.C. contributed to the chemical-genetic profiling, transcriptional analysis, and Galleria mellonella larvae assays; B.C. and Y. W. performed the molecular dynamic simulation; L.R. and Z.L. performed the binding energy calculations; J.S. contributed to the Mrt4 gene dosage and mutation assays in C. albicans; X.H. performed the Mrt4-rRNA interaction analysis assays, C.S. performed the LC/MS-MS experiments; N.P.M. and M.S.H. contributed to the review and editing of the manuscript. All authors have given approval to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Olaf Kniemeyer, Sandra Macedo-Ribeiro, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cao, H., Tu, J., Chen, J. et al. Inhibiting Mrt4-rRNA interaction with fumaramidmycin-based derivatives as an antifungal strategy. Nat Commun 17, 3422 (2026). https://doi.org/10.1038/s41467-026-70226-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70226-4