Abstract
High-grade serous ovarian cancer (HGSOC) is characterized by a complex, immunosuppressive tumor microenvironment (TME) that contributes to poor clinical outcomes and resistance to therapy. To replicate the human TME in vitro, we develop a 3D pentaculture model incorporating five human cell types: malignant HGSOC cells and primary fibroblasts, mesothelial cells, adipocytes, monocytes. Monocytes differentiate into macrophages without exogenous cytokines in the pentacultures. Bulk RNA sequencing and deconvolution with single-cell RNA data from patient biopsies reveal that macrophage clusters within the pentacultures replicate those found in human HGSOC metastases, and proportions of individual clusters vary according to the malignant cell line. The pentacultures enable detailed analysis of malignant cell–macrophage interactions and highlight the influence of malignant cell genomic, transcriptomic and proteomic heterogeneity on the TME. Furthermore, targeting of “do-not-eat-me” signals with anti-CD47 and anti-CD24 monoclonal antibodies demonstrate differential effects on macrophage activity and cancer cell viability, again depending on the individual malignant cell line. Real-time microscopic monitoring of the pentacultures confirms dynamic modulation of macrophage behavior. We conclude that this pentaculture model offers a platform to study malignant cell control of TME elements and in particular their interactions with myeloid cells.
Similar content being viewed by others
Introduction
High-grade serous ovarian cancer (HGSOC) is the most diagnosed and deadliest type of ovarian cancer. The lack of specific symptoms accounts for a late diagnosis, when the disease is already in an advanced state, rendering it challenging to treat1,2. Although most of the patients respond well to first line platinum-taxane based chemotherapy, more than 80% of patients with advanced disease will relapse and eventually develop chemoresistance3,4. In recent years, the addition of anti-angiogenic and targeted therapies has improved outcomes for some patients, although the five-year survival rate remains low5,6,7. Genomic and transcriptomic analysis of HGSOC tumors has shed light on the heterogeneity of this disease and how tumor intrinsic characteristics can affect response to treatment. Several studies have identified distinct molecular subtypes by clustering tumors with similar transcriptomic profiles8,9,10,11. The HGSOC subtypes identified correlate with different clinical and/or pathological characteristics10, and reflect variations in immune and stromal content of the tumor microenvironment (TME)11. Moreover, single-cell and spatial transcriptomics studies have further highlighted the intratumoral heterogeneity of HGSOC tumors. These analyses have also revealed heterogeneity in the ovarian tumor microenvironment which can be influenced by subclone-specific signaling to stromal and immune cells12.
Until recently, only few in vitro or in vivo models of HGSOC reflected its complex TME and malignant cell genetics13,14,15,16. This may have contributed to the limited success of recent clinical trials e.g., with immune checkpoint blockade or chimeric antigen receptor (CAR) T cell technology17. Apart from therapies that aim to stimulate T cell cytotoxicity, macrophage targeted therapies are under investigation for the treatment of many different malignancies18. Current clinical trials in various malignancies aim to inhibit the recruitment of immunosuppressive macrophages by targeting chemokine pathways such as the C-C motif chemokine ligand 2 (CCL2) axis or colony stimulating factor 1 receptor (CSF1R). Alternatively, they focus on reprogramming myeloid cells from a pro-tumor to an anti-tumor state by modulating specific receptors or signaling pathways19,20. Macrophages are highly abundant in the TME of HGSOC, and they are known to contribute to the immunosuppressive microenvironment9,21,22,23. Macrophages exhibit an innate ability to kill cancer cells through mechanisms such as phagocytosis and the activation of adaptive immune responses. However, in advanced tumors, macrophages often support cancer progression.
Macrophage-targeting agents can be studied in mouse models24,25, but many of the potential agents are humanized antibodies for which mouse phenocopies might not be available or may not reflect the mechanism of action on human cells. Furthermore, establishing mouse models that replicate the TME of advanced disease for the evaluation of novel treatments can be time-consuming compared to in vitro models. Pre-clinical experiments that test new therapies in more relevant mouse models can take many months e.g.24. In vitro experiments can be more efficient, but plastic can polarize macrophages, and this may lead to biased results. Thus, to better study the potential of myeloid-targeted drugs, researchers need complex in vitro TME models that move away from 2D culture14,26,27,28.
In this work, building on our previous work developing 3D in vitro human cell models of HGSOC29,30, we incorporate human monocytes into our established tetraculture model, creating a pentaculture (i.e., five cell types) which aims to mimic later stage of metastasis dissemination. In this 3D in vitro model, monocytes rapidly differentiate to macrophages. We use this model to investigate how malignant HGSOC cells influence the TME, particularly in driving macrophage polarization and activity. Monocytes differentiate to macrophage subsets previously identified by single cell RNAseq (scRNAseq) in human HGSOC omental metastases24, and these subsets vary according to the malignant cell used. Genomic, transcriptomic, and proteomic differences between three HGSOC malignant cell lines shape the TME and modulate macrophage polarization. Cancer cells can coerce macrophages to immunosuppressive phenotypes and inhibit phagocytosis through different signaling pathways, such as the “do-not-eat-me” signals31,32. Using the pentaculture models, we study therapeutic drugs targeting the “do-not-eat-me” signals and investigate characteristics of malignant cells that affect these processes. We conclude that the 3D in vitro pentaculture models described in this article provide insights into mechanisms by which malignant cells can drive myeloid cell differentiation that are relevant to the HGSOC TME in patient samples and may provide a medium-throughput platform for assessing cancer therapies that target malignant cell-macrophage interactions.
Results
3D pentaculture models reveal monocyte-to-macrophage differentiation and malignant cell-driven TME architecture
In an attempt to reconstruct the TME of HGSOC in vitro, we isolated primary fibroblasts, mesothelial cells and adipocytes from macroscopically normal omental tissue from patients undergoing gynae-oncological surgery either chemo naïve or post three/six cycle of chemotherapy, following a previously established protocol33.
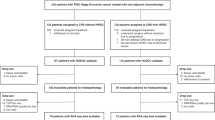
To explore the role of macrophages in the TME of HGSOC, we introduced CD14+ peripheral blood monocytes from healthy donors into our previously published tetraculture model30 which also included one of three HGSOC malignant cell lines, AOCS1, G164 and OvCAR334. Briefly, adipocyte gels were layered with primary fibroblasts and mesothelial cells to recreate a ‘normal’ omental environment. Malignant cells and monocytes were subsequently added, and the resulting model was maintained in culture for 7 days (Fig. 1A). After 7 days of culture, we observed a significantly different proportion of cancer cells between the pentacultures, whereas the proportions of fibroblasts, mesothelial cells, and macrophages were similar (Supplementary Fig. 1A). Overall cell viability remained high (Supplementary Fig. 1B). While cancer and stromal cells remained highly viable, macrophage viability was lower at day 7. However, no significant differences were observed between the different pentacultures in the viability of the primary cells (Supplementary Fig. 1C).
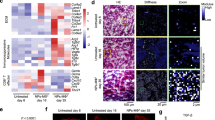
A Schematic of the cells isolated from the omental tissue and of the pentaculture setup. Created in BioRender. Malacrida, B. (2026) https://BioRender.com/g8aynuf. B H&E and CD45, CD163, CD206 IHC images of the pentaculture models. Scale bars 50 µm (representative images of n = 3 biological replicates). Flow cytometry analysis of the surface expression of CD45+ (C). Data presented as mean ± SEM (n = 5 biological replicates using 3 different donors of the primary cells), One-Way ANOVA, Tukey’s multiple comparison test. Flow cytometry analysis of the surface expression of CD163+, CD206+, and HLADR+ (D) of viable CD45+ cells. Data presented as mean ± SEM (n = 5 biological replicates using 4 different monocyte donors), Two-Way ANOVA, Tukey’s multiple comparison test. Flow cytometry analysis of the expression of CD163+, CD206+ and HLA-DR+ monocytes at day 0 (E) and representative overlayed graphs showing macrophages differentiation at day 3 (F, light color) and day 7 (F, dark color). Data presented as mean ± SEM (n = 3, three different donors). Each plot represents the expression of the macrophage marker on the x-axis versus side scatter area (SSC-A) on the y-axis. Arrows in F indicate the center of the population, grey for day 3 and black for day 7. Source data are provided as a Source Data file.
The pentaculture self-organized in a manner that resembled the micro-metastases seen in HGSOC metastases in human omentum (Fig. 1B and Supplementary Fig. 1D). However, the morphology of the pentacultures varied, depending on the malignant cell line that was added (Fig. 1B). AOCS1 and OvCAR3 cancer cells were located close to the stromal cells in the pentacultures, while G164 cancer cells generally formed clusters away from the stromal cells (Fig. 1B, Supplementary Fig. 1E and 1F). Immunohistochemistry (IHC) and flow cytometry analyses showed that AOCS1, G164, and OvCAR3 pentacultures retained a substantial population of CD45+ cells (Fig. 1B, C). To evaluate the potential monocyte to macrophage differentiation, we measured the expression of macrophage surface markers CD163, CD206, and HLA-DR after 7 days of culture (Fig. 1B, D and Supplementary Fig. 1G). We also examined macrophage polarization dynamics by assessing CD163+, CD206+, and HLA-DR+ macrophage markers at day 0, which represents the monocytic state, (Fig. 1E), day 3, and day 7 of culture (Fig. 1F). We observed a progressive increase in marker expression as the culture developed, as visualized by representative overlaid flow cytometry contour plots in Fig. 1F. These profiles show a clear shift toward higher membrane marker expression at day 7 (dark lines) compared to day 3 (light lines).
Thus, our 3D HGSOC in vitro model incorporated five human cell types, four of which were primary cells, including monocytes which differentiated to macrophages within the culture, without the addition of exogenous cytokines or any growth factors to the culture medium. Furthermore, the results suggested that malignant cells might have a role in influencing the morphology of these models.
Bulk RNA-sequencing analysis reveals malignant cells as main drivers of the TME in the pentaculture models
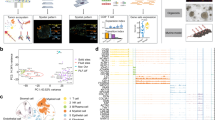
Next, we investigated the role of malignant cells in modulating the TME in a bulk RNA sequencing (RNAseq) experiment. For this experiment, we constructed three omental microenvironments by incorporating primary adipocytes, fibroblasts, and mesothelial cells isolated from three different chemo naïve donors used as biological replicates. Monocytes from a single donor were used in all experiments. This setup ensured that the only variable across the different models was the malignant cells (Fig. 2A). Additionally, we extracted RNA from 2D monocultures of fibroblasts, mesothelial cells, monocytes, malignant cells and from omental biopsies free of malignant tissue. Unsupervised clustering revealed that each pentaculture model segregated according to the malignant cell line rather than the donor of normal cells (Fig. 2B). Monocultures of malignant cells in 2D clustered close to each other and separately from the corresponding cell line pentaculture, highlighting the effect of the microenvironment on the pentaculture transcriptomes. Fibroblasts, mesothelial cells and monocyte monoculture RNA profiles were in distinct clusters as was the adjacent non-malignant omental tissue. We also observed that subclustering within the pentaculture models highlighted transcriptome variations that occurred within the context of the multicellular system, extending beyond the baseline differences observed in the malignant cell line controls (Fig. 2C, Supplementary Data 1). Together these data suggest that the malignant cells played a pivotal role in orchestrating the TME of the pentacultures.
A Schematic of RNAseq experiment setup. Created in BioRender. Malacrida, B. (2026) https://BioRender.com/g8aynuf. B Unsupervised clustering analysis of pentacultures as well as 2D cultured malignant cells, fibroblasts, monocytes and adjacent normal non-malignant omental tissue. D1 donor 1, D2 donor 2, D3 donor 3. C Heatmap illustrating normalised log2RPKM gene expression of differentially expressed genes with a significant change in at least one of the pentacultures vs the other pentacultures (generalized linear model, glm, with Benjamini-Hochberg (BH) adjusted p-value (adjp) <0.05 and logFC > |0.5 | ) and a non-significant change in any of the malignant cell monoculture pairwise contrasts (BH adjp > 0.1) clustered with k-means clustering (D, E) Heatmaps illustrating GSVA enrichment scores of (D) macrophage-related Gene Ontology Biological processes (GOBPs), (E) Hallmark terms for malignant and microenvironmental processes. D, E Two-sided Welch t-test, BH adjp <= 0.05 for penta-cultures vs single cell-types. Source data are provided as a Source Data file.
Consistent with the monocyte-to-macrophage differentiation shown in Fig. 1, we found that pathways for macrophage chemotaxis, migration, and differentiation were upregulated in the pentacultures and non-malignant omental tissue samples when compared to pathways in monocytes, primary cell monocultures (fibroblasts and mesothelial cells) and malignant cell line monocultures (Fig. 2D and Supplementary Fig. 2A). Moreover, hallmark pathways related to cell cycle regulation were highly represented in the 2D malignant cell cultures, whereas pathways resembling a more tissue-like environment appeared in the pentacultures, but not in the 2D cell cultures (Fig. 2E). To further investigate the relevance of our models, we performed a direct correlation of GSVA enrichment scores of Hallmark pathways of the AOCS1 pentacultures to publicly available RNAseq data from the original AOCS1 tumor35. This analysis showed that each of the three AOCS1 pentacultures positively correlated with the original tumor’s Hallmark signatures, whereas the AOCS1 cell line cultured alone did not significantly correlate (Supplementary Fig. 2B). This finding suggested that our pentaculture system induces elements of the TME that resemble the original human tumor from which the malignant cell was derived.
Taken together, these results suggested malignant cells were the primary drivers of the TME in our models. Notably, some macrophage-related pathways were only found in the 3D models and tissue samples, underscoring the differentiation of monocytes to macrophages in these conditions.
Macrophage clusters identified in human diseased ovarian cancer tissues are maintained in the pentaculture models
We next asked if there were transcriptional similarities between the macrophages in the pentacultures and in biopsies from HGSOC omental metastases. We used our recently published scRNAseq data from diseased HGSOC omental tissue24 to deconvolute the bulk RNAseq data from the models and quantify the proportions of the different cell populations within them. CIBERSORTx analysis revealed that fibroblast and mesothelial cell gene signatures were present at similar abundance across the three different pentacultures, whereas cancer cells and macrophage signatures varied among the models (Fig. 3A). The monocyte RNAseq data served as a control for the analysis and showed only monocyte lineage transcripts. Examining the cancer cell abundance, we observed a reduced percentage of malignant cells in the AOCS1 and OvCAR3 pentacultures compared to the G164 pentaculture. To determine if this difference in RNA abundance was reflected in the malignant cell proportions in the models, we assessed the percentage of viable cancer cells after 7 days of culture by flow cytometry. Results showed the percentage of G164 cancer cells was significantly higher than AOCS1 and OvCAR3 cells after 7 days of culture (Fig. 3B), also confirming the data shown previously in Supplementary Fig. 1A.
A CIBERSORTx analysis of macropopulation in pentacultures. B Flow cytometry analysis of the % viable cancer cells present in the models after 7 days. Data are represented as mean ± SEM (n = 4 biological replicates using 3 different donors of the primary cells). Unpaired two-tailed t-test. C CIBERSORTx analysis of the macrophage clusters in the pentacultures. D Table of the clusters and related genes selected for subsequent protein validation. Flow cytometry analysis of the expression CD169+ (E) and GPNMB+ (F) macrophages in the pentacultures. Data presented as mean ± SEM (n = 3 biological replicates using 3 different donors of the primary cells). Welch two-tailed t-test. ELISA assay results showing the release of CCL4 (G) and CXCL10 (H) of the pentacultures after 7 days. Data presented as mean ± SEM (n = 4 and n = 6 biological replicates using 4 different donors of the primary cells). Unpaired two-tailed t-test. MSD results showing the release of cytokines and chemokines by the pentacultures (I) and the malignant cell lines cultured in 2D (J). Heatmaps created using the mean values of three independent experiments (n = 3). Source data are provided as a Source Data file.
Data from our group24 has revealed 10 distinct clusters of macrophages in human HGSOC omental biopsies. As we had evidence of macrophage differentiation in the cultures, we used our published scRNAseq data to deconvolute the bulk RNAseq data from the pentacultures to see if similar macrophage clusters were seen. CIBERSORTx analysis demonstrated the macrophage clusters identified in human omental diseased tissues were also present in the models, with cluster proportions varying based on the malignant cell line used (Fig. 3C).
To validate the scRNAseq/bulk RNAseq data, we selected four different macrophage clusters: M0_Siglec1, M1_GPNMB, M4_CCL3/CCL4 and M5_CXCL9 clusters (Fig. 3D) and assessed protein expression of a representative gene within the top five expressed from each cluster using either flow cytometry or ELISA. Surface expression of CD169 (sialic acid binding immunoglobulins (Ig)-like lectin-1 or Siglec1) on macrophages was higher in the OvCAR3 pentaculture compared to AOCS1 and G164, consistent with cluster data (Fig. 3E). Macrophages GPNMB (Glycoprotein Nmb) protein expression was also significantly higher in AOCS1 and OvCAR3 pentacultures relative to G164, aligning with the corresponding RNA cluster data (Fig. 3F). Furthermore, the release of CCL4 (chemokine (C-C motif) ligands 4), IL1β (interleukin 1β), and CXCL10 (C-X-C motif chemokine ligand 10) by the pentacultures was significantly higher in G164 pentacultures than in AOCS1 and OvCAR3, again reflecting the original cluster data (Fig. 3G, H and Supplementary Fig. 3A).
To determine if the presence of cancer cells was necessary for initial monocyte-to-macrophage differentiation, we analyzed the surface expression of CD45, CD163, CD206, and HLA-DR in cultures of adipocytes, fibroblasts, mesothelial cells and monocytes (i.e. tetraculture, Supplementary Fig. 3B). Consistent with the presence of macrophages in normal omental tissue, these markers were also expressed in the tetraculture model. This finding confirms that the presence of cancer cells is not required for the initial differentiation of monocytes into macrophages.
To confirm the need for cancer cells for macrophages to acquire specific phenotypes, we analyzed the presence or the secretion of the same markers in the tetraculture model described above. We found that, unlike the initial monocyte-to-macrophage differentiation which could be promoted by adipocyte, fibroblasts and mesothelial cells alone, the expression of CD169 and GPNMB by macrophages required the presence of cancer cells (Supplementary Fig. 3C), while the secretion of CXCL10 and CCL4, predominantly produced by the macrophages, varies in the pentacultures but not in cultures without malignant cells (Supplementary Fig. 3D).
We next assayed the cytokine and chemokine secretome. The pentacultures secreted a broad spectrum of cytokines and chemokines, with patterns influenced by the presence of macrophages and malignant cells in the model (Fig. 3I). Notably, these cytokine and chemokine levels were elevated compared to those released from the 2D cancer cell cultures (Fig. 3J) and the other components, such as fibroblasts, mesothelial cells, and adipocytes grown as monocultures (Supplementary Fig. 3E). Interestingly, stromal cells, such fibroblasts and mesothelial cells, were found to secrete the highest levels of CCL2 and M-CSF (Supplementary Fig. 3E and 3F). Since these cytokines are required to promote macrophage differentiation and viability, this finding confirms that the stromal cells alone are sufficient to drive the initial monocyte-to-macrophage differentiation (Supplementary Fig. 3B) and shows why external supplementation of these cytokines was not necessary in the culture medium.
Lastly, we investigated whether malignant cells could also influence fibroblast phenotypes in our pentaculture models. Our results indicated that malignant cells might also drive fibroblast activation and differentiation transcriptional programmes (Supplementary Fig. 3G). We looked at two different fibroblasts markers, αSMA (alpha smooth muscle actin) and FAP (fibroblast activation protein), using either immunofluorescence or flow cytometry. We found that malignant cells might influence the fibroblast phenotype, though this effect was less pronounced than the effects observed on macrophages (Supplementary Fig. 3H and 3I). Specifically, a significant increase in αSMA expression was observed in the AOCS1 pentaculture fibroblasts compared to their OvCAR3 counterparts, with a non-significant trend toward an increase also noted in the G164 comparison to OvCAR3 (Supplementary Fig. 3G).
In conclusion, we observed distinct proportions of malignant cell line populations in the pentacultures models at the RNA level, which were corroborated by measuring the amount of viable cancer cells in the models after 7 days of culture. We found that macrophage clusters identified by scRNAseq in HGSOC omental diseased tissues were also present in our pentacultures in varying proportions, depending on the malignant cell line used. This finding further suggests that malignant cells influence the TME and drive macrophage polarization in the pentacultures in a similar manner to that seen in human biopsies.
Transcriptomic and genomic diversity among HGSOC malignant cells
To understand the mechanisms by which malignant cells influence the TME of the pentacultures, we explored the genome and transcriptome of AOCS1, G164 and OvCAR3 cells prior to their incorporation in the 3D pentaculture. At genomic level, we confirmed TP53 mutations consistent with HGSOC in all three cell lines (Fig. 4A), along with mutations in other genes commonly associated with HGSOC (Fig. 4B, Supplementary Data 2). No somatic BRCA1/2 or homologous recombination deficiency (HRD)-related gene mutations were detected but there was copy number loss of BRCA2, RAD51D, FANCA, CDK12 and RB1 in the G164 cell line (Fig. 4C). In line with the copy number loss of HRD genes, there was low activity of homologous recombination and mismatch repair functionality in G164 cells at transcriptome level (Supplementary Fig. 4A). All cell lines had extensive copy number alterations which reflect the chromosomal instability observed in HGSOC driven by loss of TP53 function (Supplementary Data 3)36. AOCS1 exhibited a tetraploid karyotype, G164 a diploid state, while OvCAR3 was triploid (Fig. 4D).
A Schematic of TP53 protein domains with marking on the position of non-synonymous mutations identified in AOCS1, G164 and OvCAR3 prior addition to the pentaculture models. B Oncoplot illustrating non-synonymous mutations in oncogenic drivers in the three cell lines. Light grey—no mutation detected, black—multi hit mutation, red—nonsense mutation, green—missense mutation, dark grey—pathway affected by mutation. C Copy number heatmap of genes involved in DNA homologous recombination repair. D Copy number profiles of the three cell lines. E Stacked bar plot of copy number signature activities in the three cell lines. F CCL2, IL6 and CXCL10 copy number (CN) barplots. AOCS1 are denoted in purple, G164 in pink, and OvCAR3 in red. G RNAseq analysis heatmap of normalized expression of genes maximally segregating the three cell lines identified using SAM and FDR < 0.05. AOCS1 are denoted in purple, G164 in pink, and OvCAR3 in red. cluster A denotes genes upregulated in AOCS1, Cluster G denotes genes upregulated in G164, and Cluster O denotes genes upregulated in OvCAR3. Further annotation of genes of interest present in the three clusters has been included. H Enrichment dotplot illustrating Gene Ontology Biological Processes (GOBPs) significantly enriched in the three clusters of genes, BH adjp <= 0.05; for clusterA (cA) and clusterG (cG) the top ten significant processes are displayed; for clusterO (cO) the two processes that reached significance are displayed. Source data are provided as a Source Data file.
We then explored previously developed HGSOC copy number (CN) signatures that represent different putative causes of CIN (chromosomal instability)37, and found distinct profiles of these CN signatures across our cell lines (Fig. 4E). G164 presented high activity of signature 3 (s3), in line with HRD-related genomic lesions observed in these cells; AOCS1 and OvCAR3 presented high activity of signature 4 confirming whole genome duplication events and in line with a higher amplification of CCNE1. OvCAR3 exhibited activity of signature 2, a signature of replication stress-induced tandem duplication. Taken together these data indicate that our cell lines capture a diversity of chromosomal instability features that are also observed in HGSOC tumors and in HGSOC cell organoids37. Furthermore, in addition to our analysis of HGSOC CN signatures on the three cell lines, we have conducted an additional exploration using a pan-cancer study defining 17 CN signatures38. Again, we observed that the cell lines exhibit profiles of signature activities capturing a diversity of CIN features that are also observed in human tumors. (Supplementary Fig. 4B). We also observed the expression of several genes, including CCL2 and IL6, that may influence the behavior of these cells and the pentacultures (Fig. 4F and Supplementary Fig. 4C).
We then focused on differentially expressed genes that maximally segregated the three cell lines and observed that G164 was highly enriched in wound healing factors and inflammatory response mediators including IL6, TNF and IL8 (Fig. 4G, Supplementary Data 4 and 5). AOCS1 had the highest expression of the monocyte/macrophage chemoattractant CCL2 along with high expression of CXCL12, IL33 and extracellular matrix organization genes ITGB3 COL14A1, COL27A1, MMP10, VCAN. OvCAR3 cells had the lowest expression of inflammatory mediators and had elevated expression of the metabolic regulators AKT2, ALDH1A1 (Fig. 4G, H and Supplementary Data 4 and 5). As shown in Fig. 3J, the levels of secreted CCL2, IL6, IL1β, TNFα, IL8 cytokines matched their transcript abundance in the respective cell lines. Moreover, consistent with other published studies demonstrating how tumor-intrinsic genomic and epigenetic alterations can define a specific tumor microenvironment (TME)39, we found that the G164 cell line had increased cGAS-STING pathway expression compared to the other two malignant cell lines (Supplementary Fig. 4E). This profile is associated with a more inflammatory TME, which is consistent with the profiles of both the G164 malignant cell line and its corresponding pentacultures, as shown previously in Fig. 3I, J.
In conclusion, the malignant cell lines used in our pentacultures share genomic, transcriptomic and proteomic features with HGSOC patient biopsies and organoids. HGSOC malignant cells exhibited significant genomic and transcriptomic heterogeneity. Despite sharing TP53 mutations and common CN alterations typical of HGSOC tumors, the malignant cells displayed considerable differences in CN alterations and transcriptomic profiles that are consistent with distinct proteomic characteristics and might explain their different influence in our models.
Pentaculture models display distinct phagocytosis mechanisms through the expression of ‘do-not-eat-me’ signals CD47 and CD24
Our results so far showed that the different histology and macrophage subtypes generated by the three HGSOC malignant cell lines may be influenced by transcriptomic and genomic differences that alter cytokine profiles. We also found that the number of malignant cells varied in the pentacultures. This could have been due to variation in their proliferation, but we wondered whether this was influenced by malignant cell interactions with the macrophages. To investigate potential mechanisms underlying the higher proportion of G164 cancer cells in pentacultures compared to AOCS1 and OvCAR3, as shown in Fig. 3B and previously in Supplementary Fig. 1A, we examined the expression of phagocytosis pathways. We observed that phagocytosis pathways were enriched in the pentaculture transcriptomes but were absent in the 2D malignant cell lines (Fig. 5A). Of note, some of these pathways were expressed at higher levels in the AOCS1 and OvCAR3 pentacultures compared to the G164 model. To investigate this further, we used red-labeled malignant cells and examined the proportion of macrophages positive for red cell fragments using flow cytometry. The results showed that OvCAR3 and AOCS1 cells were significantly more phagocytosed than G164 cells in the pentaculture models after seven days of culture (Fig. 5B). Of note, in models without macrophages (i.e. tetracultures) the number of viable malignant cells was similar for all three cell lines after 7 days culture (Supplementary Fig. 5A).
A Heatmap showing the expression of different phagocytosis pathways in the pentacultures vs malignant cells cultured in 2D (one-way anova p < 0.05). B Flow cytometry analysis of the percentage of viable macrophage positive for red cancer cells after 7 days of culture. Data shown as mean ± SEM (n = 3 biological replicates for AOCS1 pentaculture, n = 4 for G164 and OvCAR3 biological replicates pentacultures using 3 different donors of the primary cells). Welch two-tailed t-test (AOCS1 vs G164 p = 0.0328, AOCS1 vs OvCAR3 p = 0.0137, G164 vs OvCAR3 p < 0.0001). C IF snapshots from live recordings of the pentacultures. CD45/CD14 in green, FN1 in blue, CD90 in purple, malignant cells in red. Scale bar 50 µm. D Average percentage of macrophages at a distance less than 20 µm from malignant cells. Data presented as mean ± SEM of 5 frames for three independent videos (n = 3). Unpaired two-tailed t-test. E Overall trajectory length in µm and trajectory speed µm/sec (F) of the different pentacultures. Data shown as mean ± SEM (n = 4 biological replicates using 3 different donors of the primary cells). G Representative western blot of CD47 (50 kDa) protein and housekeeping protein Actin (42 kDa) in the 2D culture (Representative blot of n = 2 independent replicates). H CD47 log2RPKM value of the 2D malignant cell lines. Data shown as mean ± SEM (n = 2 biological replicates). I CD47 copy number alteration in the three different malignant cell lines. J CD47 and K SIRPα RNA and protein expressions in the different pentacultures. Data shown as mean ± SEM (n = 3 and n = 5 biological replicates pentacultures using 3 different donors of the primary cells). One-way ANOVA with Tukey’s multiple comparison test and Welch’s two-tailed t-test. L GSEA enrichment plots illustrate a significant increase in phagocytosis post chemotherapy compared to matched pre-chemotherapy samples in patients of late disease recurrence in the Adzibolosu et al. human HGSOC dataset41, GSE227100 (p = 2.07E-06, NES = 1.73). No significant enrichment in post versus pre-chemotherapy samples was observed in early recurrence patients (p = 0.58, NES = 0.97). Source data are provided as a Source Data file.
To visualize and quantify the spatial relationship between macrophages and malignant cells in our models, we used a technique developed by us on mouse and human ex vivo tumor slices40. Spinning disk fluorescence imaging of entire pentacultures showed that significantly more macrophages, stained in green for CD45+/CD14+, were located less than 20 µm from malignant cells in the AOCS1 and OvCAR3 pentacultures compared to G164 pentacultures (Fig. 5C, D). Time-lapse recordings of live cultures also enabled the assessment of trajectory length, speed, and macrophage behavior based on movement metrics such as maximum displacement and trajectory length (Supplementary videos 1–3). Using these parameters, we were previously able to identify four different types of immune cell behavior: static, wobbling (defined as visually moving cells that present either with non-directional migration or dynamic cytoplasmic extensions over time), migrating and long migrating40. Analysis of macrophage trajectory length, speed, and straightness (Fig. 5E, F and Supplementary Fig. 5B), as well as the different proportions of macrophages classified by behavior (Supplementary Fig. 5C), revealed similar types of cell behavior in the pentacultures. Moreover, macrophages in the pentacultures had similar behavior characteristics to macrophages in HGSOC patient biopsies and mouse HGSOC models40. This suggests that macrophage behavior is consistent across the models, but their functions may be influenced by spatial localization within the microenvironment.
Given the role of “do-not-eat-me” signals, such as CD47 and CD2432, in modulating phagocytosis, we examined their membrane expression on the malignant cells. We found that CD47 protein (Fig. 5G) and RNA (Fig. 5H) were significantly higher in OvCAR3 and G164 cells grown in 2D compared to AOCS1, consistent with CD47 copy number alterations (Fig. 5I). Similar results were observed for CD24 expression in 2D malignant cell lines (Supplementary Fig. 5D). We then looked at the expression of these markers in the pentacultures and found similar results: CD47 and CD24 remained highly expressed at both RNA and protein levels in OvCAR3 and G164 pentacultures compared to AOCS1 (Fig. 5J and Supplementary Fig. 5E). As phagocytosis inhibition by CD47 and CD24 requires interaction with their respective macrophage receptors, signal regulatory protein α (SIRPα) and sialic acid-binding Ig-like lectin 10 (SIGLEC10), we looked at the expression of both macrophage membrane markers in our models. We found that SIRPα was highly expressed at both RNA and protein levels across all the pentacultures (Fig. 5K), while SIGLEC10 was significantly expressed only by the OvCAR3 associated macrophages (Supplementary Fig. 5F). The low CD47 levels in AOCS1 explained the high phagocytosis observed, while the elevated CD47 expression in G164 might account for less phagocytosis by macrophages. However, the high CD47 levels and increased phagocytosis in the OvCAR3 pentaculture were unexpected. We looked back at CD47 expression in OvCAR3 cells using flow cytometry and found that these cells exhibit heterogeneous CD47 expression with 33.5% low and 68.8% high expressing cells (Supplementary Fig. 5G). We hypothesized that low-CD47 OvCAR3 cells were preferentially phagocytosed in the model, while the cancer cells remaining at the end of the culture have higher level of CD47.
CD47 expression in human tumors has been widely studied, but we wanted to investigate if the patterns of expression in omental biopsies of HGSOC were also variable. We performed CD47 staining on omental biopsies from 43 patients and observed variable staining patterns. Tumor cells in the biopsies exhibited either uniform CD47 positivity, complete absence of staining, or a heterogeneous (patchy) pattern characterized by regions of both high and low CD47 expression (Supplementary Fig. 5H), suggesting that similar mechanisms may be at play in patients. Moreover, to investigate potential clinical outcomes when macrophage activity is engaged, we analyzed a previously published ovarian cancer dataset of matched pre- and post-chemotherapy samples41. We found a significant increase in phagocytic activity in post-chemotherapy samples from patients who had a good response to chemotherapy (late relapse patients) compared with samples from patients who relapsed within six months from the end of treatment (Fig. 5L). This observation suggests that successful treatment or post-treatment engagement of macrophage phagocytic capacity might provide more sustained response to therapy.
Overall, we observed significantly higher macrophage uptake of red-labeled cancer cells as an indicator of phagocytosis in OvCAR3 and AOCS1 pentacultures compared to G164. These findings, supported by analyses of macrophage-cancer cell interactions and localization, suggest that the CD47-SIRPα and CD24-SIGLEC10 pathways may contribute to these differences.
Anti-CD47 and anti-CD24 mAbs enhance macrophage phagocytosis and reduce cancer cell number in OvCAR3 but not G164 models
Given that targeting “do-not-eat-me” signals can effectively modulate macrophage function42, we next tested monoclonal antibodies (mAbs) against CD47 and CD24 in our pentaculture models. Treatments with an anti-CD47 mAb or anti-CD24 mAb did not affect the overall percentage of viable cells in the pentacultures (Supplementary Fig. 6A). As expected, in AOCS1 pentacultures, which express very low levels of CD47 and CD24, treatment with either mAb had no effect on cancer cell proportion and did not increase phagocytosis (Fig. 6A, B and Supplementary Fig. 6B). The G164 pentaculture models showed no significant changes in cancer cell amount or phagocytosis following mAb treatment (Fig. 6C and Supplementary Fig. 6C), although we observed a reduction in CD47 and CD24 staining following the treatments indicating that the antibodies have successfully bound to the target (Supplementary Fig. 6D). To investigate this further, we examined macrophage localization in the G164 pentaculture models using immunofluorescence. We found that in both control and anti-CD47 treated G164 pentacultures, macrophages resided primarily in stromal areas (Fig. 6D). We hypothesized that the absence of phagocytosis in this model might reflect the localization of the macrophages or additional mechanisms of phagocytosis resistance in G164 malignant cells that prevent macrophage recruitment to, or recognition of, the cancer cells.
A Schematic of the “do-not-eat-me” signal pathways. Created in BioRender. Malacrida, B. (2026) https://BioRender.com/ur79c5g. Percentage of remaining viable cancer cells and of macrophage positive for cancer cells in (B) AOCS1 and (C) G164 pentacultures treated with anti-CD47 mAb. Data presented as mean ± SEM (n = 3 biological replicates using 3 different donors of the primary cells). D IF snapshots from live recordings of the G164 pentaculture. CD45/CD14 in green, FN1 in blue, CD90 in purple, malignant cells in red. Scale bar 50 µm. E Percentage of remaining viable cancer cells and of macrophage positive for cancer cells in OvCAR3 pentaculture. Data presented as mean ± SEM (n = 3 biological replicates using 3 different donors of the primary cells). Paired two-tailed t-test. F IF snapshots from live recordings of the OvCAR3 pentaculture. CD45/CD14 in green, FN1 in blue, CD90 in purple, malignant cells in red. Scale bar 50 µm. G Average percentage of macrophages at a distance less than 20 µm from malignant cells. Data presented as mean ± SEM of 5 frames. Unpaired two-tailed t-test (n = 2 in Supplementary Fig. 6 G, biological replicates using 2 different donors of the primary cells). H Proportions of the different types of behavior (blue, static; beige, wobbling; light grey, migrating; light blue, long migrating), I percentage of the wobbling macrophages, J Trajectory straightness and speed of the wobbling macrophages in the OvCAR3 pentacultures. Data shown as mean ± SEM (n = 2). K Percentage of remaining cancer cells and of macrophage positive for cancer cells in G164 pentacultures control and treated with TGFβRi. Data shown as mean ± SEM (n = 4 biological replicates using 3 different donors of the primary cells). Paired two-tailed t-test. L Percentage of viable cancer cells on G164 tetraculture after 7 days. Data shown as mean ± SEM (n = 3 biological replicates using 3 different donors of the primary cells). M ELISA assay results showing the release of TNFα in the G164 pentacultures control and after TGFβRi treatment. Data shown as mean ± SEM (n = 3 biological replicates using 3 different donors of the primary cells). Paired two-tailed t-test. Source data are provided as a Source Data file.
In contrast, treatment of OvCAR3 pentacultures with either anti-CD47 or anti-CD24 antibodies led to a significant reduction in numbers of cancer cells, accompanied by a marked increase in phagocytosis (Fig. 6E and Supplementary Fig. 6E). Live imaging analysis further revealed an increased number of macrophages in proximity to cancer cells following anti-CD47 mAb treatment compared to control (Fig. 6F, G and Supplementary Fig. 6F, Supplementary Videos 4, 5). Additionally, together with the increase in phagocytosis, we observed an increase in ‘wobbling’ macrophages following treatment, accompanied by a decrease in their trajectory straightness and speed (Fig. 6H, I and J), suggesting that the wobbling behavior might be indicative of an increased activity of the macrophages.
One hypothesized mechanism of resistance to anti-CD47 treatment in our G164 pentaculture model is the failure of macrophages to move from the stromal areas. As the inhibition of TGFβ has been shown to enhance immune cell motility in in vitro and in vivo43 and TGFβ pathway expression is elevated in our pentaculture models and in biopsies from HGSOC patients44, we utilized a TGFβ receptor inhibitor (TGFβRi), previously tested in our models with promising results in terms of enhanced CAR-T cell motility and reduction of poor prognosis-associated extracellular matrix (ECM) components30,45. Treatment of G164 pentacultures with TGFβRi led to a significant reduction in the number of viable cancer cells (Fig. 6K) without affecting the overall viability of the cultures (Supplementary Fig. 6G). However, the reduction in cancer cell number did not correspond to an increase in macrophage-mediated phagocytosis (Fig. 6K). To determine whether this effect was due to direct toxicity of the TGFβRi on G164 cancer cells, we treated a G164 tetraculture (lacking macrophages) with the TGFβRi and observed no significant change in viable cancer cell number (Fig. 6L). This suggests that the observed reduction in cancer cells in the pentaculture requires the presence of macrophages, which are not actively phagocytosing cancer cells, but could contribute to cancer cell death through alternative mechanisms, such as the release of cytokines or chemokines. Indeed, there was an increase TNFα release in G164 pentacultures following TGFβRi treatment suggesting that TNFα could be one of the cytokines mediating this effect (Fig. 6M).
In summary, our results demonstrated that the pentaculture model may be useful for studying macrophage-targeted therapies and macrophage-malignant cell interactions. We found that anti- CD47 and anti-CD24 mAb treatments significantly reduced cancer cell numbers and enhanced phagocytosis in the OvCAR3 model. However, no such effects were observed in the G164 model, possibly due to an alternative resistance mechanism and/or macrophage localization, or in the AOCS1 model because of its low expression of both CD47 and CD24. However, treatment with TGFβRi reduced the number of viable cancer cells in the G164 pentaculture but not in the G164 tetraculture, suggesting that macrophages may contribute to cancer cell death through alternative mechanisms in this model.
Discussion
The limited treatment options for HGSOC emphasize the need for a more comprehensive understanding of the TME and the mechanisms by which resistant malignant cells drive tumor relapse, often rendering subsequent tumors untreatable46. Investigating the TME and testing new therapeutic targets in vivo remains challenging and time-consuming, underscoring the demand for in vitro models. Here, we presented a multicellular in vitro model of one of the primary sites of metastasis of ovarian cancer, omental metastases, that integrates five cell types to better mimic the complex human TME. Different from other published multi-cellular cultures28, our model uses human primary cells for the TME components and enables an in-depth study of monocyte-to-macrophage differentiation. This approach also avoids the use of external cytokine and growth factor supplements.
Our analysis, including bulk RNAseq and deconvolution with scRNAseq data24, identified macrophage clusters consistent with those found in human diseased tissue, with cluster proportions varying depending on the HGSOC malignant cell line used. The high degree of heterogeneity in HGSOC tumors contributes to treatment resistance, complicating efforts to identify broadly effective therapeutic targets47,48. Our findings revealed that the malignant cells dictated the TME morphology pattern, cytokine profile, and macrophage phenotype, showing how intrinsic features of the malignant cells influence TME morphology and macrophage differentiation.
While our results highlighted the predominant role of malignant cells in directing macrophage differentiation, we acknowledge the potential contribution of other TME components, such as fibroblasts, mesothelial cells, and adipocytes, to the immunosuppressive state of the TME, as previously49,50,51,52,53,54. Additionally, recent genomic and transcriptomic analyses of HGSOC patients have attempted to classify patient subtypes11,12,55,56,57, but our work shows that, beyond common HGSOC mutations (e.g., TP53, BRCA1/2 mutations), copy number alterations in genes encoding for cytokines, chemokines, and “do-not-eat-me” signals, which are reflected at transcriptomic and proteomic state, might drive key changes in the TME and macrophage polarization.
Given the prevalence of macrophages in the TME of solid tumors and their general correlation with poor prognosis, targeting macrophages has emerged as a novel therapeutic strategy18,19,58,59,60. Traditional approaches targeting monocyte recruitment (e.g., anti-CCL2 antibodies)61,62 or differentiation (e.g., anti-CSF1R antibodies)63,64,65 have so far shown limited efficacy. This suggests strategies aimed at reprogramming existing macrophages within the TME from a pro-tumoral to an anti-tumoral state may offer greater therapeutic potential. scRNAseq analyses across primary and metastatic tumors could aid in the identification of novel macrophage targets11,24, which could be tested in our in vitro models. For example, targeting “do-not-eat-me” pathways such as CD47-SIRPα and CD24-SIGLEC10 hold promise66,67,68,69,70, despite challenges observed in the latest clinical trials42.
Here, we explored the potential of anti-CD47 and anti-CD24 antibodies to modulate macrophage behavior, with encouraging results in at least one model. Using live imaging techniques on our models, we could visualize macrophage interactions with malignant cells and categorize their movement. As the increased phagocytosis was accompanied by an increase in the wobbling macrophage population, we suggest that wobbling macrophages might be more active and phagocytic. We recognize, however, that a restriction in their movement could also be attributed to variable ECM stiffness or the presence of cytokines/chemokines. This further enhances the knowledge on macrophages localization and function and could be potentially exploited in an ex-vivo setting as previously described by us and others71,72,73,74. Additionally, we propose that alternative macrophage-driven mechanisms of cancer cell death can be investigated using our models, as demonstrated by the effective treatment of the G164 pentaculture with a well-characterized TGFβRi previously used in various variants of our in vitro models30,45. By contrast, our experiments targeting cytokines and chemokines (i.e. CCL2, IL8, TNFα and IL6) with neutralizing antibodies in these models has not influenced macrophage activity, suggesting that a more effective treatment approach might involve targeting either cytokine receptors or multiple cytokines simultaneously.
While we believe that the pentaculture models may be useful for studying myeloid cells in the TME, we acknowledge their limitations for studying adaptive immunity in vitro. However, in line with our published work45 and as part of our ongoing projects, such models may be useful to study the influence of other TME cells on CAR-T cell activity in vitro in a medium-throughput manner. We also suggest that further adaptation of these models, i.e. using both myeloid and CAR-T cells from the same donor would allow for the study of more complex TMEs.
In conclusion, this pentaculture model integrates five cell types using a combination of primary human cells and malignant cell lines and allows us to study malignant cell interactions with stromal cells such as macrophages. Multi-cellular models, such as we have described here, may be complementary to, and reduce the use of, mouse cancer models for understanding and targeting the human tumor microenvironment.
Methods
Patient samples and study approval
Human samples used in this research were obtained from Royal London Hospital under SIGNPOST (Systematic GeNetic Testing for Personalised Ovarian Cancer Therapy) study ethics (REC 17/LO/0405) and from Royal Marsden Hospital (REC 20/LO/0529). All ethics used for this study were approved by a UK review board. Experiments were conducted in accordance with the Declaration of Helsinki and International Ethical Guidelines for Biomedical Research Involving Human Subjects. Omental tissues deemed by a lead surgeon to be surplus to diagnostic requirement were collected after the provision of written informed consent. Macroscopically normal omental tissues were kindly donated by patients undergoing gynecological debulking surgery, either primary debulking surgery (chemo naïve) or interval debulking surgery (after three to six cycles of chemotherapy). For the RNAseq experiment of Fig. 2, three chemo naïve samples have been used to avoid any bias.
Primary cell line isolation
Human omental mesothelial cells, fibroblasts and adipocytes were isolated from macroscopic normal omentum through a series of digestion and centrifugation following the protocol described previously33. Briefly, 10 cm3 of the normal omentum was incubated in a solution of DPBS and Trypsin-EDTA 10X (Gibco, Cat. 15400054) 1:1 for 20 minutes at 37 C. After the incubation, tissue was removed and equivalent amount of DMEM:F12 Glutamax with 1% pen/strep, 5% FBS was added to the solution to neutralize the trypsin. The solution was spun down at 300 g for 5 min. The pellet, recognized as mesothelial cells, was then plated in mesothelial cell medium. The tissue was then minced in small pieces and digested using Liberase TL 1 mg/mL (Roche, Cat. 5401020001) in 20 ml of DMEM:F12 Glutamax with 1% pen/strep, 5% FBS for 30-45 min at 37 C with gentle shaking. Digested tissue was then filtered using 250 µm filters (ThermoFisher, Cat. 87791). The filtered tissue was spun down at 300 g for 3 min to separate the adipocyte content (top layer) from the pellet, stromal vascular fraction, which represents the fibroblasts. Isolated mesothelial cells and fibroblasts were cultured in DMEM:F12 Glutamax medium, supplemented either with 20% FBS for the mesothelial cells or 10% FBS for the fibroblasts, and 1% pen/strep until use. Mesothelial cells and fibroblasts were previously characterized using different markers, such as calretinin, α smooth muscle actin (αSMA) and/or fibroblast activation protein (FAP)29. Adipocytes were cast immediately in a collagen I based gel (ThermoFisher, Cat. A1048301) following the protocol previously described33 and kept in M199 medium (Gibco, Cat. 22350029) supplemented with 10% FBS and 1% pen/strep until further use.
Cell culture
AOCS1 cells were kindly gifted from Prof Bowtell from the Australian Ovarian Cancer Study. G164 cells were established in our laboratory from omental HGSOC tumors and are well characterized34. HGSOC cell line, OvCAR3 (ATCC, Cat. HTB-161), was obtained from ATCC. AOCS1 and OvCAR3 cell lines were maintained in RPMI 1640 (Gibco, Cat. 21875034) supplemented with 10% fetal bovine serum (FBS, Gibco, Cat.), 1% L-glutamine (Sigma, G7513-100ML) and 1% penicillin/streptomycin (pen/strep)(Sigma, Cat. P4333). G164 cell line was maintained in DMEM:F12 medium (Gibco, Cat. 31331093) with the addition of 4% human serum (Sigma, Cat. H4522) and 1% pen/strep. For all the experiments, 2D and pentaculture, all the cells and pentaculture were maintained in the G164 medium to standardize the culture conditions. To better visualize the malignant cells, we have also used m-Cherry lentiviral transfected AOCS1, G164 and OvCAR3 (Sartorius, Incucyte® Nuclight Red Lentivirus (EF1a, Puro), Cat. 4476; Cambridge Bioscience, Cat. BPS-78932-P; Puromycin, Gibco, Cat. A1113802). STR sequencing (ATCC, 135-XV) and mycoplasma testing (InvivoGen) were carried out routinely.
Isolation of CD14+ monocytes from peripheral blood samples
Blood was collected from healthy donors and purchased in the form of leukocyte cones. CD14+ monocytes were positively selected using CD14 MicroBeads (Milteny Biotec, Cat. 130-050-201) and LS columns (Milteny Biotec, Cat. 130-042-401) following the manufacturer protocol. CD14+ isolated monocytes were then subjected to purity check using flow cytometry. All the monocyte donors used throughout this work (n = 8) were 93-98% pure.
Three-dimensional models setup
For the pentaculture model, adipocyte gels were anchored to the bottom of the well of a 96-well plate using a layer of activated collagen as described before33. 40,000 human primary fibroblasts were plated on the adipocyte gels, followed after 2 hours by 200,000 mesothelial cells per gel. After 24 hours, 40,000 malignant cells and 400,000 CD14+ monocytes per gel were added to the model. After 24 hours, gels were scooped out and placed in G164 culture medium in a 24-well plate, 3 gels per well.
Enzyme-linked immunosorbent assay (ELISA) and Meso scale discovery (MSD)
Supernatant subjected to ELISA and MSD analysis were collected at the selected time point. Media were spun down 8000 g for 5 minutes to remove cellular debris and subsequently stored at -80C until use. ELISA was used to quantify CXCL10 (Bio-Techne, Cat. DIP100) and CCL4 (Bio-Techne, Cat. DMB00) according to manufacturer’s protocol. ELISA plates were read using FLUOstar optima microplate reader (BMG Labtech). MSD was performed using the V-PLEX Proinflammatory Panel 1 human kit, the V-PLEX Cytokine Panel 1 human kit and the V-PLEX Chemokine Panel 1 Human Kit (MSD, Cat. K15054D-1) following the manufacturer’s instructions. Plates were read on the MSD sector 600 imager (MSD).
Flow cytometry
Pentaculture gels were digested using 1 mg/ml Liberase TL (Merck, Cat. 05401020001) for 10 min and then spun down at 5,000 g for 5 minutes. Cells were then resuspended and incubated in 1X PBS with 1:100 Human TruStain FcX (BioLegend, Cat. 422301) and 1:1000 fix viability dye eFlour 780 (ThermoFisher, Cat. 65-0865-14). Cells were spun down and resuspended in FACS buffer (1X DPBS + 2 mM EDTA + 2.5% BSA) with the fluorescent antibody mix. Each fluorescent antibody was diluted 1:200 in FACS buffer. Cells were then fixed using a solution 1:1 of 10% formalin buffer solution and 1X PBS. Data was the acquired on LSR fortessa cell analyzer (BD Bioscience) and analyzed using FlowJo v10.1.5 (BD Bioscience). A table with the antibodies used, catalogue numbers and dilution is provided in Supplementary Data 6.
Bulk RNA-sequencing (RNAseq)
RNA isolation was performed using RNeasy microkit (Qiagen, Cat. 74004) according to manufacturer’s instructions. RNA sequencing was performed by the Wellcome Trust Centre for Human Genetic (Oxford, UK) to an average of 62 million reads per sample. Library preparation was performed using ribosomal depletion and a strand-specific protocol and sequencing was carried out generating 150 base pairs (bp) paired-end reads on the NovaSeq600 platform. FastQC v0.11.5 was performed and quality trimming was applied with trimgalore v0.6.5. Alignment to the reference genome GRCh37 was performed using STAR v2.7.9a (PMID: 26334920). Counting of the reads was performed with RSEM v1.3.1 using the Ensembl annotation GRCh37.87. Only genes that achieved at least one count per million (CPM) in at least 25% of the samples were kept and a log2 RPKM normalized gene expression matrix was generated. To identify clusters of differentially expressed genes the sam function from R package siggenes was used followed by kmeans clustering. Gene set enrichment analysis for Gene Ontology Biological processes on gene clusters was performed using R package ClusterProfiler. Gene set variation analysis (GSVA) was performed with R package gsva. Gene set enrichment analysis for phagocytosis on the Adzibolosu et al. dataset, GSE227100, was performed using R package fgsea on the ranked log fold-changes using the GOBP_PHAGOCYTOSIS.v2024.1.Hs.gmt gene-list. RNA-seq data have been deposited in Gene Expression Omnibus under the accession number GSE293887.
Whole Genome Sequencing
Genomic DNA was isolated and sequenced at Novogene. Whole genome sequencing was performed on the NovaSeq X Plus Series, with 150 bp, paired-end reads, yielding 90 G of raw data per sample ( ~ 300 million sequencing reads per sample, 30x coverage). After initial quality check using fastqc v0.11.5, reads were quality trimmed using trimgalore v0.6.5 and aligned to the human reference genome GRCh37 (hg19) using bwa v0.7.17. Picard v2.25.7 was used to mark read duplicates and assess insert size distributions. Base quality score recalibration was performed on known sites of variation using gatk v4.2.1.0 and Homo_sapiens_assembly19.dbsnp138.vcf, Homo_sapiens_assembly19.known_indels.vcf Mills_and_1000G_gold_standard.indels.b37.vcf, downloaded from the GATK bundle. Variant calling was performed using mutect275 on tumor only mode using gnomad.exomes.r2.1.1.sites.vcf.bgz as a germline resource. VCF files were further filtered using gatk FilterMutectCalls followed by filtering out annovar of variants from 1000 genomes (1000g2015aug_all and the exome sequencing project esp6500siv2_all with maf <0.01. Annotation with refgene, avsnp150, cosmic92_coding and cytoband was performed in annovar. Further filtering was subsequently applied to obtain variants with sequencing depth DP > 10 and AF > 0.1. Non synonymous mutations with cosmic ID were used for further downstream analysis and graphical representations. CNV analysis was performed with R package ACE. Ploidy of OvCAR3 is reported in ATCC, ploidy of AOCS1 and G164 cells has been determined previously34. Activity of the ovarian CIN signatures was calculated using britroc.cnsignatures functions from37. Activity of the 17 pan-cancer CIV signatures was calculated using R package CINSignatureQuantification (ref. 38).
Immunohistochemistry (IHC)
Pentaculture gels were fixed in 10% formalin buffer solution (Sigma, Cat. HT501128) and subsequently embedded in 2% agarose (Thermo Fisher, Cat. 10264544) in PBS prior to paraffin embedding and sectioning. Sections were then deparaffinized by incubating the sections at 60 C for 30 minutes followed by two washes in xylene. Rehydration was performed using a sequence of washing in 100%, 90%, 70% and 50% ethanol solutions. Antigen unmasking was performed using a 1x antigen retrieval Tris-EDTA buffer, pH 9.0 (abcam, Cat. ab93684) depending on the antibody used. Peroxidase enzymes were blocked with 0.6% hydrogen peroxide (H2O2; Fisher Scientific, Cat. H/1800/15) in methanol. Sections were then blocked using 2.5% goat serum (Life Technologies, Cat. 16210064) + 2.5% BSA in 1X PBS and subsequently incubated with primary antibody for 1 hour. Impress goat anti-rabbit or anti-mouse kit (Vector Laboratories, Cat. MP273 7451-50 or MP-7452-15) followed by DAB (Dako, Cat. K346811-2) chromogen staining was used for antibody detection. Sections were counterstained in Gill’s I hematoxylin (Merck, Cat. CI75290) and dehydrated with washes in 50%, 70%, 90 % and 100% ethanol solutions, followed by xylene. Sections were then mounted in DPX resin (Sigma-Aldrich, Cat. 06522) and imaged using a Pannoramic 250 high throughput scanner (3DHistech). A table with the antibodies used, catalogue numbers and dilution is provided at the end of the manuscript (Supplementary Data 6).
Treatments
At day 3 and 5, cells or gels were treated with 10 µg/ml anti-CD47 mAb, magrolimab (Cambridge Bioscience, Cat. HY-P99029) or 10 µg/ml anti-CD24, mAb, ATG-031 (Cambridge Bioscience, Cat. HY-P99176) diluted in G164 culture medium. Human IgG4κ isotype control was used as control in these experiments (Cambridge Bioscience, Cat. HY-P99003). G164 pentaculture were also treated with 20 μmol/L of TGFβ receptor inhibitor SB431542 (Sigma-Aldrich, Cat. S417). Conditioned media were collected at 7 days, unless otherwise indicated and stored at -80C until further analysis.
Western blot (WB)
Cells were lysed with RIPA buffer (Sigma, Cat. R0278) containing 1:100 protease inhibitors (Roche, Cat. 11836153001) and 1:1000 phosphatase inhibitors (Sigma, Cat. P0044). 40 μg of protein were loaded into a 4-12% NuPAGE Bis-Tris gels (Invitrogen, Cat. NP0321BOX). Samples were run in 1x NuPAGE running buffer (Invitrogen, Cat. NP0001) and transferred onto a nitrocellulose membrane (Immobilon, Cat. IPVH00010) in 1x NuPAGE transfer buffer (Invitrogen, NP0006-1). The membrane was blocked in 5% BSA in TBS-T (0,05%) for 1 hour at room temperature. The membrane was incubated with primary antibodies antibody at 4 °C overnight, followed by incubation with secondary horseradish peroxide (HPR) conjugated antibodies (Dako, Cat. P0160) for 1 hour at room temperature. HRP activity was visualized with Amersham Imager 600 (GE Healthcare).
Pentaculture live imaging
At day 7, pentacultures were washed in 1X PBS and placed in a 96-well plate in 100 µl of Human TruStain FcX (BioLegend, Cat. 422301) for 15 min at 37 °C. After blocking non-specific binding, antibodies diluted 1:10 in phenol red–free RPMI 1640 medium were incubated with the pentacultures for 15 minutes at 37 °C and subsequently washed twice with phenol red-free RPMI1640 for 2 min at 37 °C. Gels were then transferred to a glass bottom dish, covered with phenol red–free RPMI 1640 medium and secured with a tissue slice anchor.
Gels were imaged using Nikon TE Eclipse confocal microscope equipped with a spinning disk and a temperature-controlled chamber set at 37 °C, with 5% CO2. Slices were imaged every 15 seconds for 30 minutes. Images were analyzed using Imaris v9.9 (Oxford Instruments) and migration metrics were aggregated in R v4.2.2 and RStudio 2022.12.0.353, using a custom built R script, as previously described71.
Immunofluorescence
At day 7, pentaculture gels were washed with 1X PBS and fixed in 10% neutral-buffered formalin for 30 min at room temperature (RT). Samples were then permeabilized and blocked for 2 h at RT in blocking buffer consisting of 1% bovine serum albumin (BSA), 0.5% Triton X-100, 0.05% Tween-20, 2% goat serum, and 0.01% sodium deoxycholate in PBS. Unconjugated primary antibodies were diluted in blocking buffer and incubated with the gels overnight (o/n) at 4 C on a shaker. The following day, gels were washed three times for 10 min in PBS containing 0.05% Tween-20 (PBS-T). Fluorescent secondary antibodies were diluted 1:500 in blocking buffer and incubated with the gels for 4 h at RT on a shaker, followed by three 10-min washes in PBS-T. Where applicable, fluorophore-conjugated primary antibodies were subsequently added (diluted in blocking buffer) and incubated o/n at 4 °C. On day three, gels were washed three times for 10 min in PBS-T. Nuclei were stained with DAPI (1:5000 in PBS) for 10 min at RT, rinsed with PBS, and imaged without mounting using glass-bottom dishes. Images were acquired using an Evident FV4000 confocal microscope. Image analysis was performed using Imaris v9.9 (Oxford Instruments) and Fiji/ImageJ v2.17.0.
Real-time qPCR (RT-qPCR)
RNA isolation was performed using RNeasy microkit (Qiagen, Cat. 74004) according to manufacturer’s instructions. RNA purity and quality was checked evaluating the 260/280 nm ratio. cDNA was then synthesized from 800 ng of total RNA using the Vilo cDNA kit (Invitrogen, Cat. 11754-050).
Applied Biosystems® QuantStudio™ 7 Flex Real-Time PCR was used to analyze CD24 gene expression. Taqman Gene Expression Assay Kits (Hs02379687_s1, Invitrogen) were purchased to examine CD24 gene expression and HRPT1 and GADPH as housekeeping genes.
Statistical analysis
All statistical analysis were performed with the software Graphpad Prism (San Diego, CA, version 9.0.0). Kolmogorov–Smirnov test was used to determine whether the data was normally distributed. Differences between two groups were analyzed using an unpaired student t-test. Mann–Whitney test was used for nonparametric data. Paired samples were analyzed using the Wilcoxon test. Statistical differences between multiple groups were evaluated using multiple unpaired t-test, one-way or two-way analysis of variance (ANOVA). p values, along with n numbers are displayed in the figures.
Data availability
Bulk RNA sequencing data have been deposited in Gene Expression Omnibus under the accession number GSE293887. All data will be publicly available as of the date of publication. Genomic somatic mutation and copy-number variation data are available in the supplementary data files. Other RNAseq data were previously published under the following accession numbers and re-analyzed, GEO accession GSE22439224 and GSE22710041, and European Genome-phenome Archive (EGA) repository under the accession code EGAD0000100087735. The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Bowtell, D. D. et al. Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat. Rev. Cancer 15, 668–679 (2015).
Lheureux, S., Gourley, C., Vergote, I. & Oza, A. M. Epithelial ovarian cancer. Lancet (Lond., Engl.) 393, 1240–1253 (2019).
Gockley, A. et al. Outcomes of Women With High-Grade and Low-Grade Advanced-Stage Serous Epithelial Ovarian Cancer. Obstet. Gynecol. 129, 439–447 (2017).
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA: a cancer J. clinicians 74, 12–49 (2024).
Tew, W. P., Lacchetti, C. & Kohn, E. C. Poly(ADP-Ribose) Polymerase Inhibitors in the Management of Ovarian Cancer: ASCO Guideline Rapid Recommendation Update. J. Clin. Oncol. 40, 3878–3881 (2022).
Monk, B. J. et al. Niraparib first-line maintenance therapy in patients with newly diagnosed advanced ovarian cancer: final overall survival results from the PRIMA/ENGOT-OV26/GOG-3012 trial. Ann. Oncol. 35, 981–992 (2024).
Burger, R. A. et al. Incorporation of Bevacizumab in the Primary Treatment of Ovarian Cancer. N. Engl. J. Med. 365, 2473–2483 (2011).
Izar, B. et al. A single-cell landscape of high-grade serous ovarian cancer. Nat. Med. 26, 1271–1279 (2020).
Xu, J. et al. Single-Cell RNA Sequencing Reveals the Tissue Architecture in Human High-Grade Serous Ovarian Cancer. Clin. Cancer Res 28, 3590–3602 (2022).
Olbrecht, S. et al. High-grade serous tubo-ovarian cancer refined with single-cell RNA sequencing: specific cell subtypes influence survival and determine molecular subtype classification. Genome Med. 13, 111 (2021).
Vázquez-García, I. et al. Ovarian cancer mutational processes drive site-specific immune evasion. Nature 612, 778–786 (2022).
Denisenko, E. et al. Spatial transcriptomics reveals discrete tumour microenvironments and autocrine loops within ovarian cancer subclones. Nat. Commun. 15, 2860 (2024).
Teng, K. et al. Modeling High-Grade Serous Ovarian Carcinoma Using a Combination of In Vivo Fallopian Tube Electroporation and CRISPR-Cas9-Mediated Genome Editing. Cancer Res. 81, 5147–5160 (2021).
Lopez, E. et al. In Vitro Models of Ovarian Cancer: Bridging the Gap between Pathophysiology and Mechanistic Models. Biomolecules 13 https://doi.org/10.3390/biom13010103 (2023).
Battistini, C. et al. Tumor microenvironment-induced FOXM1 regulates ovarian cancer stemness. Cell Death Dis. 15, 370 (2024).
White, E. A., Kenny, H. A. & Lengyel, E. Three-dimensional modeling of ovarian cancer. Adv. Drug Deliv. Rev. 79-80, 184–192 (2014).
Pujade-Lauraine, E. et al. Avelumab alone or in combination with chemotherapy versus chemotherapy alone in platinum-resistant or platinum-refractory ovarian cancer (JAVELIN Ovarian 200): an open-label, three-arm, randomised, phase 3 study. Lancet Oncol. 22, 1034–1046 (2021).
Mantovani, A., Allavena, P., Marchesi, F. & Garlanda, C. Macrophages as tools and targets in cancer therapy. Nat. Rev. Drug Discov. 21, 799–820 (2022).
Cassetta, L. & Pollard, J. W. Targeting macrophages: therapeutic approaches in cancer. Nat. Rev. Drug Discov. 17, 887–904 (2018).
Mantovani, A., Marchesi, F., Di Mitri, D. & Garlanda, C. Macrophage diversity in cancer dissemination and metastasis. Cell Mol. Immunol. 21, 1201–1214 (2024).
Heath, O. et al. Chemotherapy Induces Tumor-Associated Macrophages that Aid Adaptive Immune Responses in Ovarian Cancer. Cancer Immunol. Res 9, 665–681 (2021).
Osborn, G. et al. Macrophages in ovarian cancer and their interactions with monoclonal antibody therapies. Clin. Exp. Immunol. 209, 4–21 (2022).
Gupta, V., Yull, F. & Khabele, D. Bipolar Tumor-Associated Macrophages in Ovarian Cancer as Targets for Therapy. Cancers (Basel) 10 https://doi.org/10.3390/cancers10100366 (2018).
Elorbany, S. et al. Immunotherapy that improves response to chemotherapy in high-grade serous ovarian cancer. Nat. Commun. 15, 10144 (2024).
Wattenberg, M. M. et al. Cancer immunotherapy via synergistic coactivation of myeloid receptors CD40 and Dectin-1. Sci. Immunol. 8, eadj5097 (2023).
Boucherit, N., Gorvel, L. & Olive, D. 3D Tumor Models and Their Use for the Testing of Immunotherapies. Front. Immunol. 11, 603640 (2020).
Habanjar, O., Diab-Assaf, M., Caldefie-Chezet, F. & Delort, L. 3D Cell Culture Systems: Tumor Application, Advantages, and Disadvantages. Int. J. Mol. Sci. 22 https://doi.org/10.3390/ijms222212200 (2021).
Estermann, M. et al. A 3D multi-cellular tissue model of the human omentum to study the formation of ovarian cancer metastasis. Biomaterials 294, 121996 (2023).
Delaine-Smith, R. M. et al. Modelling TGFβR and Hh pathway regulation of prognostic matrisome molecules in ovarian cancer. iScience 24, 102674 (2021).
Malacrida, B. et al. A human multi-cellular model shows how platelets drive production of diseased extracellular matrix and tissue invasion. iScience 24, 102676 (2021).
Brown, G. C. Cell death by phagocytosis. Nat. Rev. Immunol. 24, 91–102 (2024).
Kelley, S. M. & Ravichandran, K. S. Putting the brakes on phagocytosis: “don’t-eat-me” signaling in physiology and disease. EMBO Rep. 22, e52564 (2021).
Malacrida, B., Pearce, O. M. T. & Balkwill, F. R. Building invitro 3D human multicellular models of high-grade serous ovarian cancer. STAR Protoc. 3, 101086 (2022).
Tamura, N. et al. Specific Mechanisms of Chromosomal Instability Indicate Therapeutic Sensitivities in High-Grade Serous Ovarian Carcinoma. Cancer Res. 80, 4946–4959 (2020).
Patch, A. M. et al. Whole-genome characterization of chemoresistant ovarian cancer. Nature 521, 489–494 (2015).
Bell, D. et al. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Macintyre, G. et al. Copy number signatures and mutational processes in ovarian carcinoma. Nat. Genet 50, 1262–1270 (2018).
Drews, R. M. et al. A pan-cancer compendium of chromosomal instability. Nature 606, 976–983 (2022).
van Weverwijk, A. & de Visser, K. E. Mechanisms driving the immunoregulatory function of cancer cells. Nat. Rev. Cancer 23, 193–215 (2023).
Laforêts, F. et al. Semi-supervised analysis of myeloid and T cell behavior in ex vivo ovarian tumor slices reveals changes in cell motility after treatments. iScience 26, 106514 (2023).
Adzibolosu, N. et al. Immunological modifications following chemotherapy are associated with delayed recurrence of ovarian cancer. Front. Immunol. 14, 1204148 (2023).
Jiang, C. et al. Targeting the CD47/SIRPα pathway in malignancies: recent progress, difficulties and future perspectives. Front Oncol. 14, 1378647 (2024).
Sutherland, T. E., Dyer, D. P. & Allen, J. E. The extracellular matrix and the immune system: A mutually dependent relationship. Science 379, eabp8964 (2023).
Pearce, O. M. T. et al. Deconstruction of a metastatic tumor microenvironment reveals a common matrix response in human cancers. Cancer Discov. CD-17-0284 (2017).
Joy, J. D. et al. Human 3D Ovarian Cancer Models Reveal Malignant Cell-Intrinsic and -Extrinsic Factors That Influence CAR T-cell Activity. Cancer Res. 84, 2432–2449 (2024).
Ledermann, J. A. et al. Newly diagnosed and relapsed epithelial ovarian carcinoma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 24, vi24–vi32 (2013).
Lambrechts, S. et al. Genetic heterogeneity after first-line chemotherapy in high-grade serous ovarian cancer. Eur. J. Cancer 53, 51–64 (2016).
Shang, S. et al. Chemotherapy-Induced Distal Enhancers Drive Transcriptional Programs to Maintain the Chemoresistant State in Ovarian Cancer. Cancer Res. 79, 4599–4611 (2019).
Nieman, K. M. et al. Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nat. Med. 17, 1498–1503 (2011).
Wang, T. et al. Effect of fibroblast heterogeneity on prognosis and drug resistance in high-grade serous ovarian cancer. Sci. Rep. 14, 26617 (2024).
Qian, J. et al. Cancer-associated mesothelial cells promote ovarian cancer chemoresistance through paracrine osteopontin signaling. J. Clin. Investig. 131 https://doi.org/10.1172/jci146186 (2021).
Natarajan, S. et al. Collagen Remodeling in the Hypoxic Tumor-Mesothelial Niche Promotes Ovarian Cancer Metastasis. Cancer Res. 79, 2271–2284 (2019).
Chauvin, M. et al. Cancer-associated mesothelial cells are regulated by the anti-Müllerian hormone axis. Cell Rep. 42, 112730 (2023).
Mukherjee, A. et al. Adipocyte-Induced FABP4 Expression in Ovarian Cancer Cells Promotes Metastasis and Mediates Carboplatin Resistance. Cancer Res. 80, 1748–1761 (2020).
Murakami, R. et al. Establishment of a Novel Histopathological Classification of High-Grade Serous Ovarian Carcinoma Correlated with Prognostically Distinct Gene Expression Subtypes. Am. J. Pathol. 186, 1103–1113 (2016).
Geistlinger, L. et al. Multiomic Analysis of Subtype Evolution and Heterogeneity in High-Grade Serous Ovarian Carcinoma. Cancer Res. 80, 4335–4345 (2020).
Millstein, J. et al. Prognostic gene expression signature for high-grade serous ovarian cancer. Ann. Oncol. 31, 1240–1250 (2020).
Engblom, C., Pfirschke, C. & Pittet, M. J. The role of myeloid cells in cancer therapies. Nat. Rev. Cancer 16, 447–462 (2016).
Coussens, L. M., Zitvogel, L. & Palucka, A. K. Neutralizing tumor-promoting chronic inflammation: a magic bullet. Science 339, 286–291 (2013).
Ruffell, B. & Coussens, L. M. Macrophages and therapeutic resistance in cancer. Cancer Cell 27, 462–472 (2015).
Pienta, K. J. et al. Phase 2 study of carlumab (CNTO 888), a human monoclonal antibody against CC-chemokine ligand 2 (CCL2), in metastatic castration-resistant prostate cancer. Investig. N. Drugs 31, 760–768 (2013).
Nywening, T. M. et al. Targeting tumour-associated macrophages with CCR2 inhibition in combination with FOLFIRINOX in patients with borderline resectable and locally advanced pancreatic cancer: a single-centre, open-label, dose-finding, non-randomised, phase 1b trial. Lancet Oncol. 17, 651–662 (2016).
Razak, A. R. et al. Safety and efficacy of AMG 820, an anti-colony-stimulating factor 1 receptor antibody, in combination with pembrolizumab in adults with advanced solid tumors. J. Immunother Cancer 8 https://doi.org/10.1136/jitc-2020-001006 (2020).
Lin, C.-C. et al. Abstract CT171: Phase I study of BLZ945 alone and with spartalizumab (PDR001) in patients (pts) with advanced solid tumors. Cancer Res. 80, CT171–CT171 (2020).
Watson, S. S. et al. Fibrotic response to anti-CSF-1R therapy potentiates glioblastoma recurrence. Cancer cell 42, 1507–1527.e1511 (2024).
Barkal, A. A. et al. CD24 signalling through macrophage Siglec-10 is a target for cancer immunotherapy. Nature 572, 392–396 (2019).
Li, S. et al. IMM47, a humanized monoclonal antibody that targets CD24, exhibits exceptional anti-tumor efficacy by blocking the CD24/Siglec-10 interaction and can be used as monotherapy or in combination with anti-PD1 antibodies for cancer immunotherapy. Antib. Therapeutics 6, 240–252 (2023).
Aroldi, A. et al. CD24/Siglec-10 “Don’t Eat Me” Signal Blockade Is a Potential Immunotherapeutic Target in Mantle-Cell Lymphoma. Blood 138, 2276 (2021).
Al-Sudani, H. et al. Targeting CD47-SIRPa axis shows potent preclinical anti-tumor activity as monotherapy and synergizes with PARP inhibition. npj Precis. Oncol. 7, 69 (2023).
Jia, Y. et al. Combined Blockade of CD47-Sirpa Interaction By 5F9 (Magrolimab) and Azacitidine/Venetoclax Therapy Facilitates Macrophage-Mediated Anti-Leukemia Efficacy in AML Pre-Clinical Models. Blood 138, 510 (2021).
Laforêts, F., Donnadieu, E. & Balkwill, F. Protocol for real-time monitoring of CD8(+) T and myeloid cell behavior in human high-grade serous ovarian cancer slices. STAR Protoc. 5, 103102 (2024).
Sivakumar, R. et al. Organotypic tumor slice cultures provide a versatile platform for immuno-oncology and drug discovery. Oncoimmunology 8, e1670019 (2019).
Espie, D., Barrin, S., Rajnpreht, I., Vimeux, L. & Donnadieu, E. Live Imaging of CAR T Cell Ca(2+) Signals in Tumor Slices Using Confocal Microscopy. Methods Mol. Biol. (Clifton, N. J.) 2654, 453–462 (2023).
Lheureux, S. et al. Identification of predictive factors of response to the BH3-mimetic molecule ABT-737: an ex vivo experiment in human serous ovarian carcinoma. Int J. Cancer 136, E340–350 (2015).
Benjamin, D. et al. Calling Somatic SNVs and Indels with Mutect2. bioRxiv, 861054 (2019).
Acknowledgements
The authors would like to thank the gynaecological oncology surgeons of Barts NHS Trust for their help in supplying human tumors. The authors would also like to thank the members of the Centre for Tumour Microenvironment at Barts Cancer Institute, and Dr Chiara Battistini, Prof Ugo Cavallaro, Dr Hilary Kenny and Dr Ernst Lengyel for productive scientific discussions. We wish to acknowledge Nadia Rahman and the BCI pathology core, Sam Wallis and the BCI microscopy core and Manuela Terranova Barberio and the BCI Flow Cytometry Facility core (CRUK core service grant at Barts Cancer Institute (CTRQQR-2021\100004)). This work was funded by a UK Research and Innovation (UKRI) under the UK government’s Horizon Europe funding guarantee (EPSRC EP X028704, to B.M., R.C.B.R., J.D.J., E.M. and F.B.), by a Cancer Research UK program grant (A25714, to F.L., P.K., E.M. and F.R.B.), by an Ovarian Cancer Research Alliance grant (648516 to B.M., S.L.P.S. and F.R.B.), by Wellbeing of Women grants (ELS906 and RTF1103 to S.E.), by Barts Charity (G-002595 to S.E.) and CRUK City of London Centre Award (CTRQQR-2021\100004, to S.E.). R.M. reports support for this work by a grant from Barts Charity (MGU0295). Finally, we are extremely grateful to the patients who donated samples and made this study possible.
Author information
Authors and Affiliations
Contributions
B.M., S.E., E.M. and F.R.B. conceived the project and planned the research. B.M. designed, performed, collected and analyzed the data of most of the experiments. S.E., M.S. and E.M. designed and performed bioinformatic analysis. F.L., F.K., R.C.B.R., S.L.P.S. and J.D.J. provided technical support and performed some experiments. P.K. provided intellectual input. F.K., M.S., O.M.H. and R.M. provided patient material and provided intellectual input. B.M., E.M. and F.R.B. drafted the paper. All authors contributed to drafts and the final version.
Corresponding author
Ethics declarations
Competing interests
B.M. and F.R.B. declare research funding from AstraZeneca. F.R.B. declares being on the Scientific Advisory Board of iOmx Therapeutics AG. R.M. reports research funding grants from GSK, Yorkshire Cancer Research, Rosetrees Trust outside the submitted work, and honoraria for advisory board membership from Astrazeneca/MSD/GSK/EGL. The remaining authors declare declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Benjamin Neel, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Malacrida, B., Elorbany, S., Laforêts, F. et al. 3D pentaculture model unveils malignant cell-driven macrophage polarization in high-grade serous ovarian cancer. Nat Commun 17, 2451 (2026). https://doi.org/10.1038/s41467-026-70398-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70398-z