Abstract
Single-cell spatial transcriptomes have demonstrated molecular and cellular diversity in the brain, but gene regulatory mechanisms underlying transcriptomic profiles and disease pathogenesis remain largely unknown in primates. Here we performed single-nucleus Assay for Transposase-Accessible Chromatin followed by sequencing (snATAC-seq) for ~1.6 million cells from 142 cortical regions of two male cynomolgus monkeys (Macaca fascicularis), and identified distinct chromatin accessibility profiles of cis-regulatory elements (CREs) for various cell types. By integrative analysis with large-scale spatial transcriptome data, we found that these CREs showed laminar and regional preferences, with their regional accessibility exhibiting striking dependence on the region’s hierarchical level. Cross-species comparison of snATAC-seq data revealed human/macaque-enriched layer-4 glutamatergic neurons and LAMP5/LHX6-expressing GABAergic neurons as well as human/macaque-biased CREs for genes related to neurodevelopment and psychiatric diseases. Importantly, risk single-nucleotide polymorphisms for many brain disorders strongly associated with human/macaque-biased CREs in glutamatergic neuronal types and those for Alzheimer’s disease strongly associated with CREs exclusively in microglia. Our results provided the basis for understanding the spatial gene regulatory mechanisms underlying cellular diversity and disease pathogenesis in the primate cortex.
Similar content being viewed by others
Introduction
Single-cell transcriptome analyses have facilitated our understanding of the diversity of cell types and their spatial distributions in the brain. Hundreds of glutamatergic, GABAergic, and glial cell types have now been identified1,2,3,4,5,6,7,8. However, the gene regulatory mechanisms underlying their identity and function remain largely unclear. The spatiotemporal pattern of gene expression depends on the chromatin accessibility of cis-regulatory elements (CREs), which are binding sites for sequence-specific transcription factors (TFs) that recruit proteins for transcriptional initiation of their target genes. A comprehensive characterization of CREs at the single-cell level will help to elucidate gene regulatory mechanisms in diverse cell types and the molecular basis of brain development, function and diseases.
Recent studies using the snATAC-seq method have enabled the analysis of cell type-specific CREs in the adult mouse brain9,10,11,12. Previous bulk sequencing assays have revealed that enhancer elements, the main component of CREs, exhibit rapid changes during mammalian evolution13 and primate-specific enhancer elements were found to be involved in brain diseases14. Thus, primate-specific CREs could appear during primate evolution and be specifically responsible for human diseases. Indeed, comparative studies of single-cell chromatin accessibility datasets have shown conserved chromatin accessibility and CREs in human, monkey and mouse brains15,16. However, whether primate-specific CREs underlie transcriptomic profiles of primate-enriched cell types and human disease pathogenesis remains largely unknown. Moreover, chromatin accessibility in bulk tissues was found to exhibit region-selective patterns in the primate brain17 and single-cell chromatin accessibility has been examined for a limited number of primate brain regions15,18,19. Thus, a comprehensive characterization of single-cell chromatin accessibility covering the entire macaque cortex will help to advance our understanding of CREs underlying the diversity of cell types and their regional distribution in the primate brain.
In this study, we generated a single-nucleus chromatin accessibility dataset for ~1.6 million single nuclei from 142 cortical regions of the adult macaque brain using the snATAC-seq method. We defined 230 clusters of cells based on this chromatin accessibility atlas and identified 615,873 candidate CREs associated with various cell types in the macaque brain. Mapping of CREs with the spatial transcriptome showed their laminar and regional preferences. Further cross-species comparison of single-cell chromatin accessibility data from macaques, mice, and humans revealed human/macaque-biased CREs for psychiatric disorder-associated genes in layer-4 (L4) glutamatergic cell types and for neurogenesis-related genes in GABAergic neurons. Notably, we also found a strong association between human/macaque-biased CREs in specific cell types and risk single-nucleotide polymorphisms (SNPs) for various brain diseases. These results provide a comprehensive database for studying gene regulatory programs at the single-cell level and for understanding pathogenic mechanisms underlying brain disorders. An interactive website for accessing our dataset is available through https://macaque.digital-brain.cn/chromatin-accessibility.
Results
Characterization of chromatin accessibility in various macaque cortical cell types
To understand gene regulatory programs underlying cellular diversity in the primate cortex, we performed chromatin accessibility profiling using a previously reported droplet-based snATAC-seq method20 for tissue samples from 142 cortical regions (128 from Macaque #1 and 132 from Macaque #2) covering the entire cortex of two adult male cynomolgus monkeys (Macaca fascicularis) (Fig. 1a and Supplementary Data 1). The data reliability was confirmed by the high numbers of uniquely mapped reads, high proportion of properly paired reads, high numbers of unique fragments, low doublet ratio, and consistent quality metrics between biological replicates (Supplementary Fig. 1a-e). A total of 1,615,430 nuclei passed quality control criteria including the number of unique fragments per cell, TSS enrichment score, promoter ratio and doublet removal (Methods and Supplementary Fig. 1f, g). The remaining cells exhibited an average of 20,534 chromatin fragments in each nucleus and 57.6% of reads in peak regions. These data were used for unsupervised clustering analysis, and assisted by a previously reported spatial transcriptome dataset21, our chromatin accessibility data were used for further spatial mapping of CREs and gene regulatory networks (GRNs; Fig. 1a).
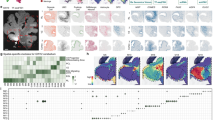
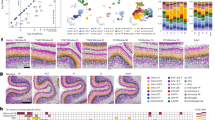
a Schematics of experimental procedure and integration of snATAC-seq with snRNA-seq and spatial transcriptome data. Single nuclei from 142 cortical regions of two monkeys were isolated for snATAC-seq and snRNA-seq analysis, and co-embedded with spatial transcriptome data from a previous study21. b Upper, UMAP embedding and clustering analysis based on chromatin accessibility profiles of snATAC-seq data. Below, UMAP layout showing single nuclei from different cortical lobes. Single nuclei are colored by cell clusters or cortical lobes. A full list and description of cell clusters and subclusters are provided in Supplementary Data 2. L2/3, layer 2/3, CR, Cajal-Retzius, PV, Parvalbumin, CHC, chandelier cell, SST, Somatostatin, ASC, astrocyte, OPC, oligodendrocyte precursor cell, OLG, oligodendrocyte, MG, microglia, EC, endothelial cells, PC, pericytes, VLMC, vascular leptomeningeal cells. c Marker peaks of aggregated chromatin accessibility profiles for each snATAC-seq-based cell cluster at selected marker gene loci that were used for cell annotation. d Left, Heatmap showing the chromatin accessibility of CREs across various cell clusters. Right, Heatmap showing the enriched TF motifs for various cell clusters. Representative TFs and their corresponding motif logos were shown on the right. e Venn plots showing the overlap between distal CRE linked genes and differentially expressed genes in the same cell type. P values were calculated by hypergeometric test.
Unsupervised clustering based on the representations after iterative latent semantic indexing (LSI) of chromatin accessibility in each cell resulted in three major types, glutamatergic (726,611; 45%), GABAergic (275,073; 17%) and non-neuronal (613,746; 38%) cells. Further iterative clustering of each major type revealed 91 subclusters (14 clusters) of glutamatergic neurons, 105 subclusters (8 clusters) of GABAergic neurons, and 34 subclusters (7 clusters) of non-neuronal cells (Fig. 1b and Supplementary Data 2). The major clusters and subclusters were annotated by the peak accessibility at the promoter region of their canonical marker genes6,22,23. For examples, high chromatin accessibility was observed at the promoter region of SLC17A7 for all glutamatergic neurons and GAD2 for GABAergic neurons. Clusters associated with astrocytes (ASCs), oligodendrocyte progenitor cells (OPCs), oligodendrocytes (OLGs) and microglia cells (MGs) showed open chromatin peaks at the promoter of GFAP, PDGFRA, PLP1 and PLAC8, respectively (Fig. 1c). Cells from multiple cortical areas of two monkeys were present in the majority of cell types (Fig. 1b), and high consistency in the proportions of various clusters was found between biological replicates (Supplementary Fig. 1h-k), demonstrating the reliability of the batch correction. Hierarchical clustering of the averaged LSI representations in each cluster resulted in a taxonomy tree (Supplementary Fig. 1l). Integrative analysis of snATAC-seq and snRNA-seq data showed that the clusters of chromatin accessibility profiles identified by snATAC-seq corresponded well with snRNA-seq-defined cell types (Supplementary Fig. 2a, b and Supplementary Data 2).
We further applied the MACS2 method24 to identify open chromatin regions for each cell type by aggregating chromatin accessibility profiles (Methods). After filtering out those with low confidence, we detected an average of 138,477 open chromatin regions per cell type and a total of 615,873 open chromatin regions in all cell types (Supplementary Fig. 2c). Among these open chromatin regions, more than 90% were located outside ±1 kb window around known transcriptional start site (TSS) of protein-coding and long noncoding RNA genes, consistent with previous data for human and mouse brains11,15,18,19 (Supplementary Fig. 2d). Interestingly, with comparable numbers of chromatin fragments among different cell types, the proportions of open chromatin peaks enriched in the TSS or promoter regions (more adjacent cis-regions) in non-neuronal cells and GABAergic cells were much higher than those in glutamatergic cells (Supplementary Fig. 2e). Our analysis of previously published datasets of human, macaques and mice11,15,18,19 showed the same phenomenon (Supplementary Fig. 2e). The lower TSS enrichment in glutamatergic neurons might be induced by higher activity of glutamatergic neurons or more distant regulatory elements from the TSS in glutamatergic cells. As a previous study25 has shown that neuronal activity induces a major reconfiguration of the chromatin state, with most of the activity induced open regions located in the intergenic and intronic regions.
We thus defined these distal open chromatin regions as cis-regulatory elements (CREs) and linked them with putative target genes by calculating the co-accessibility of each distal CRE and open regions within ±1 kb from the TSS of targeted genes in each cell type using the Cicero method26, resulting in 1,872,039 CRE-gene pairs (Supplementary Data 3). Over 70% of CREs (436,615) showed differential chromatin accessibility across different cell subclasses, and a large proportion of CRE-linked genes in each cell subclass overlapped with marker genes in the corresponding snRNA-seq-defined cell subclasses (Fig. 1e, Supplementary Fig. 2f and Supplementary Data 4). For examples, L2/3 glutamatergic cell subclass showed high chromatin accessibility in CREs linked to GPR83 and PDZD2 genes, which were also highly expressed in snRNA-seq-defined L2/3 neurons. Moreover, these cell subclass-restricted CREs were also enriched for distinct sets of DNA motifs recognized by known transcriptional regulators (Fig. 1d). For instances, the motif binding activity of the nuclear receptor gene NR1D2, which encodes the TF involved in several brain disorders, showed the highest level in L4_2 glutamatergic neurons. The activity of binding motifs by SOX family TFs SOX9 and SOX10, which are involved in oligodendrocyte specification and myelination27, was found to be highest in OPCs (Fig. 1d). These findings establish a basis for understanding gene regulatory mechanisms underlying various transcriptome-defined cell types in the macaque cortex.
Spatial mapping of chromatin accessibility revealed by snATAC-seq data
To further examine the spatial accessibility of cell type-specific CREs, we integrated the snATAC-seq data with a single-cell spatial transcriptome map of the macaque cortex21 for one-to-one pairing. Specifically, we down-sampled the “gene activity score” matrix of snATAC-seq data to mimic the gene capture rate of Stereo-seq data, and then used autoencoder and generative adversarial network to transfer snATAC-seq-identified clusters onto the spatial map (Fig. 2a). The optimal transport method was used at the same time to achieve one-to-one mapping, in which chromatin accessibility profiles from snATAC-seq data were assigned to single cells on the spatial transcriptome map (Methods). The accuracy of this cell mapping was evidenced by the high consistency in the relative percentage of different cell types and marker genes between the snATAC-seq and spatial transcriptome data for the same cell types (Supplementary Fig. 3a, b).
a Diagram showing the integration and one-to-one pairing of snATAC-seq and spatial transcriptome data. b Genome browser track view showing differential accessible distal CREs linked to GPR83, ADCYAP1, RORB and TLE4 in various glutamatergic neurons. c Spatial maps and bar plots showing the chromatin accessibility of four example CREs and expression levels of these CREs-linked genes across cortical layers in a representative coronal section (EBZ + 10). The quantification was calculated from chromatin accessibility of all single nuclei in each layer, and the sample sizes are listed in Supplementary Data 10. Error bars represent standard error of the mean (SEM). d Gene regulatory networks of the V1-specific L4_2 glutamatergic neurons. Red and green colors represent TFs and their target genes, respectively. The directed edges indicate the regulation. e Genome browser track view showing a differential accessible distal CRE linked to CPA6 in various glutamatergic neurons. f Spatial maps of expression patterns of MEF2C and CPA6, and chromatin accessibility of the CRE linked to CPA6 at two representative sections (EBZ −6.3 and −6.8). g RNAscope assay showing the expression pattern of the CPA6 gene in V1 and V2 regions of the macaque cortex. Probes against CPA6 (ACD, cat#1843691-C1) was list in Supplementary Data 9. DAPI was used for nuclear counterstaining. One of two biological replicates was shown from a single experiment. Independent replicates showed consistent patterns. h Co-localization of RNAscope labeling of the CPA6 gene and the MEF2C gene in the V1 region, a magnified view in (g). i Heatmap showing the correlation of chromatin accessibility and the hierarchy levels of various brain regions in the visual and somatosensory system. Correlation coefficients and P values were calculated by Spearman correlation analysis. * P < 0.05, ** P < 0.01, *** P < 0.001. j Dot plots showing the correlation of chromatin accessibility and the hierarchy levels of various brain regions in the visual system listed in (i). k Genome browser tracks showing the chromatin accessibility of two CREs linked to CBLN2 among L2/3, L3/4 and other cell subclasses.
To explore the spatial distribution pattern of chromatin accessibility for cell type-specific CREs, we then calculated the relative percentage of each cell type in each of the six cortical layers and found that glutamatergic subclusters showed clear laminar preference (Supplementary Fig. 3c). The cell type-specific CREs also showed laminar-specific chromatin accessibility in the spatial map, consistent with the preferential laminar expression of their targeted genes. For example, the CRE linked to the marker gene GPR83 of L2/3 glutamatergic neurons showed open chromatin accessibility peaks only in L2/3 (Fig. 2b). Spatial mapping of this CRE also showed significantly higher chromatin accessibility in L2 and L3, consistent with the spatial expression preference of GPR83 (Fig. 2c and Supplementary Fig. 3d). In addition, distal CREs linked to ADCYAP1, RORB, and TLE4 also showed consistent preferences in spatial chromatin accessibility with the expression pattern of corresponding marker genes for L2/3, L4 and L6 glutamatergic neurons, respectively (Figs. 2b, c and Supplementary Fig. 3d).
In addition to layer-specific distribution, we also found cortical region-specific distribution of some cell clusters based on their relative density among nine cortical regions (Supplementary Fig. 3c). For example, the L4_2 glutamatergic subclass exhibiting high chromatin accessibility at the promoter of MYOC was enriched in the V1 region of the occipital lobe quantified from both snATAC-seq and spatial transcriptome data (Fig. 1c and Supplementary Fig. 3e, f). This cluster corresponded well to snRNA-seq-defined L4.4 and L4.8 cell types that were also specifically distributed in V121 (Supplementary Fig. 3g). The DNA sequences of CREs enriched in L4_2 glutamatergic cluster were then scanned for TF-binding motifs with the CellOracle method28, generating a gene regulatory network (GRN) for all potential regulatory interactions (Fig. 2d). One marker CRE linked to the CPA6 gene was significantly enriched with the binding motif for TF MEF2C (Fig. 2e), which is involved in normal neuronal development, cell distribution, and electrical activity of the neocortex as well as neuropsychiatric disorders29. Spatial mapping of this CRE also showed that its chromatin accessibility was V1-specific (Fig. 2f). Furthermore, the RNAscope analysis validated the V1-specific expression of CPA6 and higher expression of MEF2C in V1 than V2 (Fig. 2g and Supplementary Fig. 3h). In addition, co-localization of the TF MEF2C and its target gene CPA6 were found in the V1 region (Fig. 2h), suggesting their V1-specific regulation. Similarly, both the chromatin accessibility of CRE linked to LAMA4 and the expression of LAMA4 showed V1 specificity (Supplementary Fig. 3i, j). These results indicate that the laminar- and regional-specific accessibility of CREs might underlie the specific expression patterns of their targeted genes.
As the evolutionarily ancient part of the cerebral cortex in the brain, the allocortex has a simpler structure than the more common neocortex30. To understand the regulatory differences between the allocortex and neocortex, we next performed differential CRE analysis between the allocortical piriform cortex and all the other neocortical regions. A total of 19,376 CREs showed significantly differential chromatin accessibility between the two regions (Supplementary Fig. 4a). The up-regulated CREs in the piriform cortex were enriched in oligodendrocytes (OLGs) and other glial cells, whereas the down-regulated CREs were mainly contributed by deep-layer glutamatergic neurons (Supplementary Fig. 4b). Moreover, the piriform cortex included a higher proportion of glial cells (especially OLGs) and lacked L4 neuronal subtypes (L4_1, L4_2 and L4_3) (Supplementary Fig. 4c), in line with the anatomical absence of layer 4 in the piriform cortex. In addition, GO functional enrichment analysis revealed that genes linked by up-regulated CREs (CSF1R, P2RY1, CX3CR1) were significantly enriched in the “gliogenesis” term, whereas genes linked by down-regulated CREs (KCNIP4, RGS7, SLC24A2) were enriched in “potassium ion transport” term (Supplementary Fig. 4d, e). Spatial mapping of these pathways showed that the “gliogenesis” pathway was significantly enriched in the piriform cortex, whereas the “potassium ion transport” showed a lower abundance than other neocortical lobes (Supplementary Fig. 4f, g). As examples, the CRE linked to CSF1R showed high chromatin accessibility in the piriform cortex (Supplementary Fig. 4h). In contrast, the CREs linked to KCNIP4 showed high chromatin accessibility in the other neocortex (Supplementary Fig. 4i). Therefore, these results reveal different regulatory programs between the piriform and neocortical regions.
Given the laminar and regional preferences of CRE accessibility, we next wondered whether the accessibility of cell type-specific CREs showed intracortical gradients along the neocortex. As an example, the CBLN2 is known to exhibit an anterior-posterior gradient in expression enriched in the prefrontal cortex during both development and adulthood in macaques and humans, and its enhancer has been identified to be active in the prefrontal cortex31. In order to assess the potential gradients of CREs associated with CBLN2 along the macaque cortical hierarchy, we mapped the CREs linked to CBLN2 gene in our spatial transcriptome map and found their higher chromatin accessibility in L2 and L3 (Supplementary Fig. 4j, k). We further performed Spearman correlation analysis of chromatin accessibility and the hierarchy levels of various brain regions in the visual system and the somatosensory system. The results showed that the accessibility of most of these CREs in L3 exhibited a significantly positive correlation with the hierarchy level of brain regions in both the visual system and the somatosensory system (Fig. 2i). Two example CREs (chr18:7561002-7561502 and chr18:7597050-7597550) showed open chromatin peaks in L2/3 and L3/4 and exhibited increasing accessibility along the hierarchy of both visual and somatosensory systems in L3 (Fig. 2j, k and Supplementary Fig. 4l). Taken together, these results reveal differences in CRE accessibility across cortical layers and regions, supporting the role of different regulatory programs in the spatial organization of various cell types in the primate neocortex.
Human/macaque-biased CREs in glutamatergic neurons
To determine whether the gene regulation landscape in the cortex are conserved among humans, macaques and mice, we retrieved previously published snATAC-seq data for human and mouse cortices11,15. Joint clustering analysis was performed using gene activity scores of homologous genes in snATAC-seq datasets for cortical tissues from all three species. We found generally conserved snATAC-seq-based clusters of GABAergic and non-neuronal cells across the three species, except for L4_1, L4_3 and L4/5/6_2 glutamatergic neurons that existed only in humans and macaques, as shown by the extracted maps of clusters for each species (Fig. 3a, Supplementary Fig. 5a-b and Supplementary Data 5). Further quantitative analysis among the three species using MetaNeighbor32 showed that macaque L4_1, L4_3 and L4/5/6_2 neurons showed high similarity with human L3/4 IT, L4 IT and L4/5 IT cell clusters, respectively, but no correspondence to any cluster in mice (Fig. 3b). Further comparison of all chromatin accessibility clusters revealed 12 clusters (belonging to L4_1, L4_3 and L4/5/6_2 subclasses) that were exclusively identified in primates but not in mice (Supplementary Fig. 5c). The human/macaque-biased clusters also corresponded well to snRNA-seq-based L4 and L4/5/6 glutamatergic cell types (Supplementary Fig. 3g), which were previously shown to be primate-specific21.
a UMAP co-embedding of snATAC-seq data from macaques, humans and mice. Colored dots at the bottom indicate human/macaque-biased glutamatergic neurons derived from snATAC-seq data. b Heatmap showing the similarity of snATAC-seq derived glutamatergic cell clusters across macaques, humans and mice using MetaNeighbor. c The outer box represents the number of CREs showing higher chromatin accessibility in human/macaque-biased than other cell types of the macaque cortex. The inner box represents CREs with high chromatin accessibility in the corresponding human cell types. The inner circle indicates CREs showing no homologous sequence in the mouse genome. d GO enrichment for genes linked to human/macaque-biased CREs (inner circle in c). e Genome browser tracks showing the chromatin accessibility of human/macaque-biased CREs located on the DCC gene in the human/macaque-biased and other cell subclasses. f Spatial map showing the chromatin accessibility of the human/macaque-biased CREs shown in e in a representative Stereo-seq section at indicated coordinate. The right panel showing the quantitative chromatin accessibility level across six cortical layers. The sample sizes are listed in Supplementary Data 10. Error bars represent SEM. g RNAscope results showing the layer 4-specific expression of DCC gene in TPO (left) and V4t (right) regions of the macaque cortex. Probe against DCC (ACD, cat#1843701-C1) was listed in Supplementary Data 9. DAPI was used for nuclear counterstaining. One of two biological replicates was shown from a single experiment. Independent replicates showed consistent patterns. h Scatterplots showing the expression of layer 4 marker gene RORB and DCC gene in human MTG of the previously published MERFISH dataset34. i Bar plots showing 7 categories of CREs in various cell subclasses from macaques, humans and mice. j Boxplots showing the percentage of CREs colocalized with TE in different categories (N = 20 subclasses). P value was calculated using a two-sided Wilcoxon rank-sum test, adjusted by Benjamini-Hochberg FDR correction. k Boxplots showing the percentage of CREs located in different types of TEs (N = 20 subclasses). Statistical testing was performed analogous to j.
Further differential CRE analysis between human/macaque-biased cell types and all other glutamatergic cell types identified 17,546 CREs showing higher chromatin accessibility in the human/macaque-biased than other cell types (Supplementary Fig. 5d). Among these CREs, 4,333 CREs also showed high open chromatin in the corresponding human cell types, and 1,834 CREs had no homologous sequences in mice (Fig. 3c, Supplementary Fig. 5e and Supplementary Data 5). These human/macaque-biased CREs linked genes were significantly enriched in “glutamatergic synaptic transmission”, “postsynaptic specialization”, “regulation of trans-synaptic signaling” and “postsynaptic density membrane” (Fig. 3d). Specifically, we found that the chromatin accessibility of two CREs located in the intron of the neurodevelopmental disorder-associated gene DCC (netrin 1 receptor)33 were much higher in these human/macaque-biased neurons than those in other cell types (Fig. 3e), consistent with the higher expression level of DCC in L4, L3/4, L3/4/5 and L4/5 glutamatergic neurons (Supplementary Fig. 5f). These CREs showed much higher accessibility in L4 for most coronal regions and in L3 and L5 for regions without L4 quantified by spatial mapping (Fig. 3f and Supplementary Fig. 5g), consistent with the expression pattern of DCC. RNAscope assay verified that the expression of DCC was enriched in layer 4 (Fig. 3g). Analysis using the previously published MERFISH dataset from the human MTG region34 further validated the layer 4-specific expression of DCC gene, which is consistent with the expression pattern of RORB, a canonical marker gene of layer 434 (Fig. 3h). Another example is a distal CRE that linked to TRDN with higher chromatin accessibility in these human/macaque-biased glutamatergic neurons (Supplementary Fig. 5h). The decreased expression of TRDN was reported to be associated with apoptosis of dopaminergic cells in Parkinson’s disease35. Spatial mapping showed the higher chromatin accessibility of this CRE and higher expression level of TRDN in deeper layers (Supplementary Fig. 5i, j). These results suggest the unique roles of human/macaque-biased CREs in regulating the spatial expression of disease-associated genes in the primate cortex.
Furthermore, we also examined whether CREs identified in macaque cell types that were conserved across three species were present in human or mouse cortices. For this analysis, we lifted CREs identified in each species to the human genome by sequence homology, and compared the sequence similarity and chromatin accessibility of CREs in each conserved cell subclass. Only CREs showing both sequence similarity and open chromatin between species were considered as conserved (Methods). We found that in comparison with species-conserved CREs, a higher proportion of CREs showed open chromatin in a species-specific pattern (Fig. 3i). Interestingly, we found that ~60% of human/macaque-biased CREs were colocalized with known transposable elements (TEs) (Fig. 3j), consistent with the notion that TEs as a driver of genomic diversity36. In comparison with CREs conserved between primates and mice, human/macaque-biased or species-specific CREs showed significantly higher frequency in co-localization with TE elements including LINE, SINE, DNA and long terminal repeats. In addition, human/macaque-biased CREs showed comparable enrichment with LINE and SINE, whereas conserved CREs between primates and mice showed much higher co-localization with SINE element (Fig. 3k). Thus, we have identified human/macaque-biased CREs that may determine layer- and region-specific glutamatergic cell types and may be responsible for gene regulation underlying brain disorders.
Characterization of CREs in GABAergic cell types
The integrative clustering analysis also showed clear correspondence of snATAC-seq identified clusters and snRNA-seq-defined GABAergic cell types (Supplementary Fig. 6a). However, based on chromatin accessibility profiles, we found that transcriptome-defined LAMP5 + GABA cell subclass could be further divided into LAMP5 and LAMP5/LHX6 cell types that exhibited low and high chromatin accessibility of the LHX6 gene, respectively. We identified 3,829 and 1,989 differentially accessible (DA) chromatin peaks in LAMP5/LHX6 and LAMP5 cells, respectively. GO analysis of genes linked to these distal DA peaks revealed enrichment in trans-synaptic signaling and postsynaptic membrane regulation for LAMP5/LHX6 cells, consistent with fast and precise synaptic inhibition37,38,39 In contrast, LAMP5 cell–specific peaks were associated with pathways related to long-term synaptic depression, reflecting features of sustained inhibition in CGE-derived neurons (Supplementary Fig. 6b). Interestingly, the enrichment of ZIC4, USF1, and MEF2B motifs in primate LAMP5/LHX6 cells, but not in mice (Fig. 4b), suggests an evolutionary divergence in the regulatory programs governing interneuron specialization. These transcription factors are known to regulate telencephalic regionalization, neuronal differentiation, and activity-dependent plasticity40,41,42. Their primate-specific motif enrichment therefore implies adaptive cis-regulatory innovations contributing to cortical expansion and refined inhibitory circuit modulation in primates. Moreover, the pronounced regional variation in the ratio of LAMP5/LHX6 to LAMP5 cells, with the highest abundance in the cingulate cortex (Fig. 4c), indicates potential region-specific functions. The preferential enrichment of LAMP5/LHX6 cells in association cortices may reflect their role in higher-order integrative and cognitive processing43,44, whereas the broader distribution of LAMP5 cells across cortical areas likely supports more generalized inhibitory balance. By examining the shared CREs in LAMP5/LHX6 cells between macaques and humans, we found enrichment of CREs for genes in the pathways of monoatomic and inorganic cation transmembrane transport, such as KCNQ3, SLC6A1 and CACNA1G (Supplementary Fig. 6c and 6d), suggesting the distinct role of this enriched cell type in the primate brain.
a The UMAP visualization (left) and relative percentage (right) of GABAergic LAMP5 and LAMP5/LHX6 cell subclasses among humans, macaques and mice. b Dot plot showing the transcription factor binding site (TFBS) activities in LAMP5/LHX6 GABAergic neurons across species. c Relative ratio of LAMP5/LHX6 vs. LAMP5 cell subclasses in various macaque cortical regions. d UMAP embedding of LAMP5/LHX6 neurons (left) and normalized gene activity score of PROX1 and NOS1 (right). e Volcano plot showing the differentially enriched DORCs in PROX1+ and NOS1+ populations of LAMP5/LHX6 neurons. Differential enrichment was defined as adjusted P-value < 1×10⁻⁵ and |log₂FC | > 0.75. f The number of distal CREs linked to TSS region of PROX1. g Genome browser track view showing two primate-specific distal CREs linked to TSS region of NOS1 in LAMP5/LHX6 cell clusters. h UMAP embedding of GABAergic SST neurons labeled with their layer specificity. i Gene regulatory networks for layer-specific SST neuron groups. j Genome browser track showing two differential accessible distal CREs linked to KIRREL3 in L2/3 SST neurons. The two CREs harbor binding motifs of TF KLF7. k Spatial visualization of chromatin accessibility of two CREs linked to KIRREL3 in SST neurons in representative sections at different coordinates in EBZ numbers (left). The bar plot illustrates the mean chromatin accessibility of CREs across different layers (right). The sample sizes are listed in Supplementary Data 10. Error bars represent SEM. l Boxplot showing chromatin accessibility of two CREs linked to KIRREL3 (upper/lower correspond to left/right in k) in SST neurons across cortical lobes. P values were calculated by Kruskal-Wallis test. The sample sizes are listed in Supplementary Data 10. Center line, median; box limits, 25th–75th percentiles; whiskers, 1.5 × IQR; points beyond, outliers.
Examination of chromatin accessibility in LAMP5/LHX6 cells showed that these cells could be divided into two populations with differential accessibility of the PROX1 gene (Fig. 4d), in line with our finding based on a previous transcriptomic dataset on primate cortical cells21 (Supplementary Fig. 6e). Moreover, we also found differential chromatin accessibility for the marker gene NOS1 in macaque LAMP5/LHX6 cells (Fig. 4d), suggesting complementary roles of PROX1 and NOS1. To validate the presence of PROX1⁺ and NOS1⁺ subpopulations within LAMP5/LHX6 cells, we analyzed a published MERFISH dataset from human dlPFC45, which confirmed the existence and spatial distribution of these two subgroups at single-cell resolution (Supplementary Fig. 6f-k). We then identified the domains of regulatory chromatin (DORC) in each of the LAMP5/LHX6 cell populations and found that DORC of SEC22A, ADARB2 and PROX1 genes were upregulated in PROX1+ cells, and DORC of ETS1, NOS1 and EML6 genes were upregulated in NOS1+ cells (Fig. 4e and Supplementary Fig. 6l). This result further supports the importance of PROX1 and NOS1 in determining the heterogeneity of primate cortical LAMP5/LHX6 cells. Among the CREs associated with the PROX1 and NOS1 genes in LAMP5/LHX6 cells, 6 out of 21 CREs linked to PROX1 were human/macaque-biased (Fig. 4f), whereas 2 out of 5 distal CREs linked to NOS1 were human/macaque-biased (Fig. 4g). Differential spatial distributions were also observed, with the PROX1+ population being enriched in layers 3, 5, and 6, while the NOS1+ population was enriched in layers 2, 3, and 5 (Supplementary Fig. 6n). A previous study of human embryonic ganglionic eminence development46 identified an MGE-derived LHX6⁺CRABP1- population proposed as progenitors of adult LAMP5/LHX6 neurons, which we found also co-express PROX1 and NOS1. Consistently, in macaque, adult LAMP5/LHX6 neurons from our previously snRNA-seq dataset21 clustered with an embryonic LHX6⁺/CCK⁺ population (CRABP1-) reported as their putative progenitors42, likewise marked by PROX1 and NOS1 expression (Supplementary Fig.6n, o). These findings support a developmental continuity from MGE-derived embryonic progenitors to adult PROX1⁺ and NOS1⁺ LAMP5/LHX6 subpopulations.
SST cells were resolved into 27 transcriptionally distinct clusters characterized by differential chromatin accessibility (Supplementary Fig. 7a), as shown by the top differentially accessible (DA) peaks (Supplementary Fig. 7b). Genes linked to these DA peaks indicated cell type-specific epigenomic regulation. Notably, clusters enriched for high HTR2A/HTR2B gene activity (SST.1, SST.2 and SST.3) corresponded to serotonergic-responsive interneurons (Supplementary Fig. 7c). To further characterize functional differences, we performed rGREAT analysis on DA peaks from these three clusters and found that distinct SST subgroups were enriched in different biological processes: SST.1 in neuropeptide signaling pathway, SST.2 in energy homeostasis, and SST.3 in synapse organization (Supplementary Fig. 7d), underscoring the biological relevance of this fine-grained classification.
Previous study has revealed the layer-specific distribution of SST neurons that contributed to various anatomical and electrophysiological features of the human cortex47. Layer enrichment of various SST cell types was also found in a single-cell spatial transcriptome map of the macaque cortex21. We then further focused on the layer-specific regulatory programs of various SST cell types. We classified layer-specific SST neurons based on clustering analysis of their chromatin accessibility profiles and annotated them based on their layer preferences (from L2/3 to L5/6, Fig. 4h), using gene activity scores of previously identified layer marker genes21 (Supplementary Fig. 7e). Further GRN analysis revealed that different layer-enriched SST cell types showed layer-enriched activity scores for genes targeted by different TFs, such as KLF7 and FOXO3 for L2/3 SST cells, ETS1 and BCL11B for L3 SST cells, BATF and POU1F1 for L3/5 SST cells, NR2F2 and MITF for L5 cells, and BACH2 and NFE2L2 for L5/6 SST cells (Fig. 4i and Supplementary Data 6). As an example, we demonstrated that the preferential accessibility of two CREs associated with KIRREL3 for KLF7 binding in L2/3 on the spatial transcriptome map (Fig. 4j, k). Interestingly, we also noticed that the accessibility of the two CREs showed regional differences, with the highest levels in parietal and prefrontal lobes, respectively (Fig. 4l). In another example, we found the preferential L5/6 accessibility of CREs associated with DCC for BACH2 binding (Supplementary Fig. 7f, g) and the highest level of this CRE accessibility in the prefrontal and cingulate cortical regions (Supplementary Fig. 7h). These findings support the notion that binding of specific TFs to accessible CREs for genes expressed in SST cells may determine their layer-specific and region-enriched distribution.
Characterization of CREs for cortical non-neuronal cells
We have further explored the diversity of gene regulatory mechanisms that may be responsible for generating distinct non-neuronal cell types and their distribution in various macaque brain regions, as revealed by the previous spatial transcriptome analysis21. Cross-species comparison of our macaque data with those from humans and mice11,15 showed that these non-neuronal cell types are all present in the cortices of the three species, with the percentage of various non-neuronal cells varied among the three species. While the astrocytes were found to be present at similar percentages, the percentages of oligodendrocytes (OLGs) increased gradually from mice to macaques and humans, and that of microglia in the human cortex was much higher than that found in both macaque and mouse cortices (Supplementary Fig. 8a, b). Notably, we found that the percentages of cortical oligodendrocyte precursor cells (OPCs) were similar between macaques and humans, but much higher than that in mice (Supplementary Fig. 8a, b). This finding is in line with the prolonged myelination and altered gene regulation in the oligodendrocyte lineage of the primate brain2,48,49,50.
We next examined the abundance of OPCs vs. OLGs across the 143 cortical regions and found the opposite abundance of OPCs and OLGs in both snATAC-seq and spatial transcriptome data (Fig. 5a and Supplementary Fig. 8c). Interestingly, we observed a strong negative correlation (R = -0.9, P < 2.2×10-16) between the percentages of OPCs and OLGs among various cortical regions in the visual system51 (Fig. 5b). Specifically, higher abundances of OPCs and OLGs were found in cortical regions of high and low hierarchy levels, respectively. The high potential for oligodendrocyte proliferation due to OPCs in regions of higher hierarchy suggests higher plasticity in the regulation of axon myelination in these regions. To further understand the differences in regulatory elements for OPCs and OLGs among different brain regions, we performed Spearman correlation analysis of chromatin accessibility and the hierarchy levels of various brain regions in the visual system. The results showed a consistent higher number of CREs exhibiting chromatin accessibility negatively correlated with the hierarchy level in OPCs regardless of thresholds for coefficients and P values (Supplementary Fig. 8d). Interestingly, more than 30% of these negatively correlated CREs were human/macaque-biased (Supplementary Fig. 8e, f), and their linked genes were significantly involved in GO terms including glutamatergic synapse, visual system development and positive regulation of kinase activity (Supplementary Fig. 8g and Supplementary Data 7). For examples, one mammal-conserved CRE showing open chromatin peaks only in OPCs was linked to PDGFRA, a typical marker gene for OPCs (Fig. 5c). Another example is a human/macaque-biased CRE with open chromatin only in OPCs that linked to FBLN2, which is an extracellular matrix inhibitor of OPC differentiation52. These results suggest the reciprocal regulation between OPCs and OLGs and the importance of OPCs in regulating higher cortical functions.
a Heatmap showing the percentages of OPCs and OLGs among all glial cells in various brain regions of the macaque cortex after spatial mapping. The brain regions are ordered by the decreasing percentage of OPCs. b Correlation between the percentage of OPCs and OLGs in various cortical regions in the visual system. Brain regions are colored by their hierarchical levels. Correlation coefficients and P values were calculated by Spearman correlation analysis. c Genome browser track view showing one mammal-conserved CRE linked to the TSS region of PDGFRA in OPCs. d Genome browser track view showing one human/macaque-biased CRE linked to the TSS region of FBLN2 in OPCs. e Heatmap showing the chromatin accessibility of CREs in protoplasmic and interlaminar astrocytes. f Dot plot showing the hub score of enriched TFs in the GRNs of interlaminar and protoplasmic astrocytes. The x-axis is ordered by the decreasing hub scores of TFs in the GRNs. g Gene regulatory network of TFs and targeted genes in interlaminar astrocytes. h Genome browser tracks showing the chromatin accessibility of CRE linked to MMP28 between protoplasmic and interlaminar astrocytes. i Spatial maps of expression patterns of ID3 and MMP28, and chromatin accessibility of the CRE linked to MMP28 at a representative section (EBZ + 10). j Bar plots showing quantitative levels of ID3 and MMP28 expression as well as chromatin accessibility of the CRE linked to MMP28 across six cortical layers. The sample sizes are listed in Supplementary Data 10. Error bars represent SEM.
Previous spatial transcriptome studies have shown laminar preferences of astrocytes (ASCs)21,45. Our snATAC-seq-based cell clustering analysis further revealed differential chromatin accessibility of CREs associated with interlaminar ASCs (enriched in L1) and protoplasmic ASCs (enriched in L2-L6) (Supplementary Fig. 8h). For example, higher chromatin accessibility was found for CREs linked to ID3, FABP7, VCAN and MMP28 and those linked to LUZP2, GRID1, KCNH5 and KCNMA1 in interlaminar and protoplasmic ASCs, respectively (Fig. 5e). Based on GO analysis, CREs found in the interlaminar ASCs were enriched in genes associated with “cell adhesion” and “cell projection” pathways, whereas those in the protoplasmic ASCs were associated with “intracellular sodium ion homeostasis”, “GABAergic synapse”, and “positive regulation of calcium ion transport” (Supplementary Fig. 8i). Moreover, we constructed GRNs for the two groups of ASCs and found that interlaminar ASCs showed high enrichment of hub TF genes ID3 and PBX3 (Fig. 5f). The targeted genes of ID3 (including GATA2, MSX1, TFAP2C and PBX3) were also involved in the central nervous system development (Fig. 5g). On the other hand, the protoplasmic ASCs were enriched in the hub TF gene NPAS3 (Fig. 5f), which contributes to neurodevelopmental transcription factor networks and the regulation of brain glucose metabolism53 (Supplementary Fig. 8j). As examples, the CRE linked to MMP28 showed high chromatin accessibility in interlaminar but not protoplasmic ASCs (Fig. 5h). Spatial mapping of this CRE also showed higher chromatin accessibility in L1, in line with the high expression of its linked gene MMP28 and its binding TF ID3 in L1 (Figs. 5i, j and Supplementary Fig. 8k). In contrast, one of the NPAS3-targeted CRE that linked to LUZP2 showed significantly higher chromatin accessibility in protoplasmic ASCs, and spatial mapping of this CRE also showed higher chromatin accessibility across cortical layers, consistent with spatial expression pattern of LUZP2 (Supplementary Fig. 8l, m). Polymorphic variants in LUZP2 have been reported to be associated with Alzheimer’s disease54, schizophrenia55, intelligence56, and verbal memory57. Taken together, these results provide insights into regulatory programs underlying the diversity and spatial organization of non-neuronal cells in the primate brain.
Association of cell type-specific CREs with brain disease risk SNPs
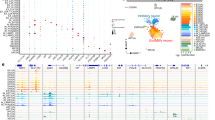
To explore the contribution of cell type-specific cis-regulatory elements (CREs) to the genetic architecture of brain diseases, we applied linkage disequilibrium score regression (LDSC) to test the enrichment of GWAS-identified risk SNPs from 28 brain disorders and human traits in our cell type-specific CREs (Supplementary Data 8). Hierarchical clustering based on enrichment similarity revealed strong associations between neuronal CREs, particularly those in glutamatergic neurons, and multiple brain disorders (Figs. 6a, b and Supplementary Fig. 9a; Full list of cell types in Supplementary Data 8). For examples, schizophrenia (SCZ) and bipolar disorder (BP) risk SNPs were enriched in CREs of superficial-layer glutamatergic neurons, whereas BP SNPs were also present in PVALB and SST neurons. Generalized epilepsy (GE) SNPs were enriched in RELN neurons, and Alzheimer’s disease (AD) SNPs were concentrated in microglia (Fig. 6a). Furthermore, SNPs related to cognitive traits such as cognitive function (CGF), intelligence quotient (IQ), reaction time (RT), and years of education (YED) showed the highest enrichment in L4 glutamatergic neurons (Fig. 6a). These results highlight the selective localization of disease-associated variants in CREs of specific neuronal subtypes, underscoring their potential role in disease susceptibility.
a Heatmap showing the enrichment of cell type-specific CREs containing risk SNPs of various brain disorders and human traits for all neuronal and non-neuronal cell types from snATAC-seq data of the macaque cortex. Intensity of the red color codes LDSC enrichment score (defined as -log10 P values, with P representing the probability of CREs containing risk SNPs). Black triangles mark the two brain disorders with most prominent enrichment probabilities. AjD: adjustment disorder, AUD: alcohol dependence, AD: Alzheimer disease, AMN: amnesia, GAD: anxiety disorder, ADHD: Attention deficit hyperactivity disorder, ASD: Autism spectrum disorder, BP: bipolar disorder, MDD: Major depression disorder, CD: cognitive disorder, GE: generalized epilepsy, ID: Insomnia, LBD: Lewy body dementia, OUD: opioid dependence, PaD: panic disorder, PD: Parkinson Disease, PE: partial epilepsy, PS: psychosis, SCZ: Schizophrenia, TS: Tourette syndrome, CGF, IQ: Intelligence, cognitive function, MTP: Morning Person, RT: reaction time, SWB: subjective well-being, YED: Years of Education, BMI: Body mass index, CAD: coronary artery disease. b Boxplot showing the relative enrichment levels of 223 cell types across different diseases in a. Significant difference was observed among different diseases (P < 1e-20, Kruskal-Wallis test). Center line, median; box limits, 25th–75th percentiles; whiskers, 1.5 × IQR; points beyond, outliers. c Heatmap showing the Pearson correlation coefficients for the association of cell type-specific CREs with disease risk SNPs loci for various brain disorders. Star marked the high similarity in disease risk SNP-associated cell type-specific CREs between schizophrenia and bipolar disorder. d Genome browser tracks showing chromatin accessibility of CREs for gene ADNP that contain risk SNPs (marked by dots) of both SCZ (schizophrenia) and BP (bipolar disorder). The arrow line indicates the targeted gene ADNP. e Spatial maps of cell type-enriched CREs containing SCZ- and BP-associated SNPs loci shown in d at three representative sections along the coronal coordinates, indicating the layer-specific distribution of the CRE for ADNP. The sample sizes are listed in Supplementary Data 10. Error bars represent SEM.
We next examined the genomic context and evolutionary properties of CREs colocalized with disease-associated SNPs. Approximately 40% of these CREs were intronic, and ~50% were located more than 1 kb from transcription start sites (Supplementary Fig. 9b). Compared to background peaks, they exhibited significantly lower phyloP scores58,59 (Supplementary Fig. 9c), suggesting rapid evolutionary turnover. Enrichment patterns across diseases showed the highest correlation between SCZ and BP (Fig. 6c and Supplementary Fig. 9d), consistent with their known genetic overlap60. Notably, we identified two shared SNPs (rs8118905 and rs1040711) within a CRE linked to the ADNP promoter61,62, with accessibility peaking in L2/3 glutamatergic neurons (Fig. 6d). Spatial mapping of this CRE also showed the highest level of accessibility in L2 and L3 of the cortex (Figs. 6d and e). In addition, we found two other SNPs rs2051307 and rs7753616 were linked to SETBP1 gene in BP and MDGA1 gene in SCZ, respectively (Supplementary Fig. 9e). Both genes were related to neurodevelopment63,64, and the spatial distributions of SNP-associated CREs for the two genes were also enriched in the L2/3 glutamatergic neurons (Supplementary Fig. 9f). Together, these findings indicate that SCZ- and BP-associated SNPs preferentially disrupt regulatory elements active in L2/3 glutamatergic neurons, highlighting a shared neurodevelopmental basis for disease risk.
To assess the evolutionary specificity of disease-associated CREs, we classified SNP-containing CREs as human/macaque-biased or mammal-conserved. Glutamatergic neurons, especially the L4_3 subclass, harbored significantly more human/macaque-biased disease-associated CREs than other cell types (Fig. 7a). In L4_3 subclass, human/macaque-biased CREs carrying SCZ SNPs displayed higher accessibility than mammal-conserved CREs (Fig. 7b). Among them, genes linked to human/macaque-biased CREs (181 gene; including ABCA765, ADORA166 and CHGA67) and mammal-conserved CREs (293 genes, including CHRM468, CXXC169 and OPCML70) showed only minimal overlap (12 genes, including PDYN)71 (Fig. 7c and Supplementary Fig. 9g). Spatial mapping showed cell type- and region-specific accessibility patterns of these CREs, with L4_3.11 neurons in the occipital lobe exhibiting the highest accessibility (Fig. 7d). Thus, SCZ-associated regulatory mechanisms appeared to be highly cell type- and brain region-specific. Gene regulatory network analysis of L4_3 neurons identified transcription factors CTCF72, NEUROD673, SREBF174, HIF1A75 and CXXC169 as central regulators (Fig. 7e). Furthermore, the human/macaque-biased CRE linked to GFRA276 and the mammal-conserved CRE linked to NEFH77, containing binding sites for regulators of SREBF1 and HIF1A, respectively, both exhibited the highest chromatin accessibility in L4 of the spatial maps (Fig. 7f and Supplementary Fig. 9h). Collectively, these results indicate that primate-evolved regulatory networks may play a pivotal role in SCZ pathology.
a Dot plot showing the ranked distribution of human/macaque-biased vs. mammal-conserved CRE ratios (with disease-risk SNPs) across cell subclasses. Cell types with fold changes below the background are shown with reduced transparency. The violin plot (upper left) illustrates comparisons between glutamatergic neurons, GABAergic neurons, and non-neuronal cells. P values were calculated using a two-sided Wilcoxon rank-sum test (*** P < 0.001). For Glut. Vs GABA., P < 2.22 × 10-16; Glut vs non-neurons, P = 2 × 10-8. b Ridge plot depicting the chromatin accessibility of human/macaque-biased and mammal-conserved CREs (colocalized with SCZ risk SNPs) in the L4_3 glutamatergic cluster. Significant difference was calculated by a two-sided Student’s t-test. c Venn Diagram showing the overlap of genes associated CREs in b. d Bar plot illustrating the relative chromVAR deviations of CREs in b. The sample sizes are listed in Supplementary Data 10. Error bars represent SEM. e Gene regulatory network (GRN) showing the key transcription factors (TFs) for targeted genes linked to CREs associated with SCZ risk SNPs in L4_3 neuron. SCZ ontology genes were labeled. f Genome browser tracks showing chromatin accessibility in L4_3 neurons at SCZ risk SNP-associated CREs linking to GFRA2 and NEFH, respectively. g Box plot showing the ranked distribution of the fold change in the relative composition of human/macaque-biased versus mammal-conserved CREs across various brain disorders. The sample sizes are listed in Supplementary Data 10. Significant difference was observed by two-sided Wilcoxon rank-sum test. Center line, median; box limits, 25th–75th percentiles; whiskers, 1.5 × IQR; points beyond, outliers. h Venn Diagram illustrating the overlap of genes associated with human/macaque-biased and mammal-conserved CREs in microglia cells. i Pathways enriched by genes linking to AD risk SNP associated CREs in microglia. j Genome browser tracks plot showing chromatin accessibility in microglia at the AD risk SNP associated CREs linked genes in h. k Dot plot illustrating the relative chromVAR deviations of human/macaque-biased and mammal-conserved CREs (colocalized with AD risk SNPs) across brain lobes in microglia.
We also investigated the distribution and functional properties of human/macaque-biased and mammal-conserved CREs carrying AD risk SNPs in microglia. AD SNP–associated CREs were exclusively localized to microglia (Fig. 6a), with the highest ratio of human/macaque-biased to conserved CREs observed among all brain diseases examined (Fig. 7g). These two CRE classes were largely distinct in their target genes (127 vs. 207 genes with 17 shared) (Fig. 7h). Human/macaque-biased CREs carrying AD risk SNPs were linked to genes involved in pathways such as amyloid-beta clearance, LDL response, and small molecule metabolism—absent from those associated with conserved CREs (Fig. 7i). Notable AD-related genes linked to human/macaque-biased CREs included C378, TREM279, NFYA80, CDNF81 and IDE82. For instance, one human/macaque-biased CRE in microglia was linked to TREM2 and NFYA. On the other hand, both mammal-conserved and human/macaque-biased CREs were linked to AD-related genes83,84,85 APOE, TOMM40 and TRAPPC6A in microglia (Fig. 7j). Spatial accessibility analysis revealed that human/macaque-biased CREs were most accessible in occipital and somatosensory cortices, whereas conserved CREs peaked in insular and auditory cortices (Fig. 7k). These results suggest that human/macaque-biased microglial CREs regulate distinct gene programs relevant to AD pathogenesis, highlighting their potential as species-specific therapeutic targets.
Given that LAMP5/LHX6 neurons are enriched in both primates (Fig. 4a), we next assessed whether disease risk SNPs are differentially enriched between human-specific and macaque-specific CREs in this cell type. We focused on three brain disorders—attention deficit hyperactivity disorder (ADHD), cognitive disorder (CD), and generalized epilepsy (GE)—which exhibited significant heritability enrichment in our LDSC analysis (Fig. 6a). For each trait, genes linked to CREs harboring risk SNPs showed limited overlap between the two species (human vs. macaque, ADHD: 919 vs. 1246 genes, 65 shared; CD: 771 vs. 1055 genes, 70 shared; GE: 470 vs. 634 genes, 33 shared). Notably, within conserved genomic regions, disease-associated SNPs mapped to distinct CREs that were either human- or macaque-specific (Supplementary Fig. 10a-c). For examples, an ADHD risk SNP (rs34376065) was co-localized with a human-biased CRE linked to the NTNG1 gene. In contrast, open chromatin peaks were only found in the macaque genome for CREs harboring several ADHD risk SNPs (Supplementary Fig. 10b). These results highlight species-specific divergence in regulatory architectures underlying disease heritability, even within consensus primate cell types such as LAMP5/LHX6 neurons.
Discussion
We have performed a comprehensive study of chromatin accessibility at the single-cell level across 142 brain regions of the macaque cortex and identified specific CREs for various cell types. Leveraging the newly developed method for integrating chromatin accessibility and spatial transcriptome data, we found that many cell type-specific CREs showed prominent laminar and regional preferences in their chromatin accessibility, thus providing gene regulatory mechanisms underlying the laminar and regional diversification of cell types. Cross-species comparison with human and mouse snATAC-seq data further identified CREs associated with brain disease-related genes in human/macaque-enriched L4 glutamatergic and GABAergic cell types. Further comprehensive association analysis between cell type-specific CREs and brain disease risk SNPs revealed potential contribution of human/macaque-biased CREs in various brain diseases. Notably, human/macaque-biased CREs in L4 glutamatergic neurons are much more prominently associated with risk SNPs for schizophrenia. Furthermore, we also identified human/macaque-biased CREs that regulate AD pathogenesis-related genes (C3, TREM2, NFYA, CDNF and IDE) exclusively in microglia. This study offers a comprehensive database for further investigating gene regulatory mechanisms underlying cell type diversity in the primate brain as well as brain disease pathogenesis.
Our analysis of the chromatin accessibility landscape in primate and mouse cortices revealed the role of human/macaque-biased CREs in the emergence of human/macaque-biased cell types as well as their potential involvement in psychiatric diseases. For example, in human/macaque-biased L4 glutamatergic neurons, we identified CREs localized in the regulatory region of DCC, a gene involved in neurodevelopment and psychiatric disorders86,87,88. We also found that human trait-related SNPs largely resided in CREs of human/macaque-biased glutamatergic cell types, suggesting potential contributions of these neuronal types to cortical development underlying human intellectual traits. The discovery of distinct presence of human/macaque-biased CREs related to some brain disorders such as schizophrenia and AD supports the notion that these brain diseases are due to cortical pathologies that are largely primate-specific.
Previous studies have shown that cortical LHX6+LAMP5+ GABAergic neurons are more abundant in primates4, but their molecular diversity and regulatory features remain poorly characterized. Here, we further subdivided the LAMP5+LHX6+ population into two transcriptionally and epigenetically distinct subtypes—PROX1⁺ and NOS1⁺—characterized by mutually exclusive chromatin accessibility at their respective lineage-defining genes (PROX1 for CGE origin46, NOS1 for MGE origin89), as well as distinct laminar distributions within the cortex. Beyond previous transcriptomic observations, our analysis revealed subtype-specific DORCs and identified multiple human/macaque-biased CREs associated with PROX1 and NOS1, suggesting evolutionary innovation in the regulatory architecture of these interneurons. These findings highlight previously unrecognized epigenetic and spatial heterogeneity within LAMP5⁺LHX6⁺neurons and provide a resource for understanding their developmental origin and primate-specific specialization.
Our comprehensive analysis of chromatin accessibility and spatial transcriptome allowed us to identify differences in chromatin accessibility across various cortical regions. This information is critical for understanding the gene regulation underlying the regional differentiation of cell types. Our findings of the reciprocal relationship between the abundance of OPCs and OLGs across all brain regions suggest that OLGs may provide negative regulation of OPC proliferation. The opposite trend in the abundance of OPCs and OLGs at different hierarchy levels of the visual system51 further suggests the functional role of axon myelination in neural properties at various hierarchy levels, and OPC biogenesis and myelination are tightly regulated across cortical hierarchy. The higher abundance of OPCs in cortices of higher hierarchy suggests their higher capability of generating OLGs, contributing to the plasticity of myelin formation and regeneration in these cortices.
Elucidation of regulatory mechanisms underlying diversity of cortical cell types and their distribution is important for understanding not only the structure and function of the brain but also pathogenic mechanisms underlying brain disorders. Through association analysis between cell-type specific CREs and brain disease risk SNPs, we uncovered hundreds of disease-related CREs in the primate brain, including those regulating genes involved in SCZ, BP and AD. Our finding on AD risk SNP-associated CREs exclusively in microglia provides strong support to the notion that microglia play a critical role in the AD etiology90,91. Notably, prominent AD risk SNP alleles were found in CREs linked to TREM2 and APOE genes in microglia, and CREs for TREM2 were found to be human/macaque-biased. In addition, SCZ- and BP-associated CREs were generally enriched only in the primate L2/3 glutamatergic neurons. Importantly, we found a large increase in the frequency of transposon element insertion in human/macaque-biased CREs. This suggests that the high frequency of brain disorder risk SNPs that are linked to human/macaque-biased CREs might be attributed to the extensive proliferation of cortical neurons during primate brain development.
Methods
Tissue collection and nucleus extraction
We sampled 128 cortical regions from Macaque #1 and 132 regions from Macaque #2 for snATAC-seq data generation. For Macaque #1, samples were collected from thick adjacent sections to the ones used for Stereo-seq. For Macaque #2, we dissected brain samples from block-face cortical regions of the hemisphere opposite to that used for Stereo-seq. Using tissue punchers (2.5 - 4 mm in diameter), we segmented the cortical areas at a depth of 1 – 2 mm in the cryostat. After dissection, the samples were promptly frozen in liquid nitrogen and then stored in dry ice or a -80 °C refrigerator. Throughout the sampling process, great care was taken to transfer the tissues to pre-chilled tubes without allowing them to thaw. Single nuclei were isolated, stained with NeuN antibody and DAPI, sorted by FACS (DAPI⁺/NeuN⁺), and stored at -80 °C, in accordance with the method described previously92. The isolated single nuclei were evenly split for snRNA-seq and snATAC-seq library preparation, respectively. In brief, frozen sections of monkey brain tissue were placed in a Dounce homogenizer containing 2 ml of pre-chilled homogenization buffer (containing 20 mM Tris pH8.0, 500 mM sucrose, 0.1% NP-40, 0.2U/μL RNase inhibitor, 1x protease inhibitor cocktail, 1% bovine serum albumin (BSA), and 0.1 mM DTT), and the Dounce homogenizer was kept on ice during the grinding process. The tissue was homogenized with 10-15 strokes of pestle A, followed by 10-15 strokes of pestle B. Subsequently, 2 ml of homogenization buffer was added to the Dounce homogenizer, and the homogenate was filtered through 30 μm MACS SmartStrainers (Miltenyi Biotech, #130-110-915) into a 15 ml conical tube. The tube was then centrifuged at 500 g for 5 minutes at 4°C to pellet the nuclei. The pellet was resuspended in 1.5 ml of blocking buffer (containing 1% BSA and 0.2U/μL RNase inhibitor in 1x PBS) and centrifuged again at 500 g for 5 minutes at 4°C to pellet the nuclei. The nuclei were finally resuspended in cell resuspension buffer (PBS containing 1% BSA) for subsequent snATAC-seq library preparation.
Library preparation for single-nucleus ATAC-seq
For the preparation of single-nucleus ATAC-seq libraries, we employed the DNBelab C Series Single-Cell ATAC Library Prep Set (BGI-research) with the procedure of step transposed, droplet encapsulation, pre-amplification, emulsion breakage, then the captured beads were collected. After DNA amplification and purification, the indexed sequencing snATAC library was already to sequence. Finally, we used the Qubit ssDNA Assay Kit (Thermo Fisher Scientific) to measure the concentrations of the sequencing libraries. All libraries were sequenced using a paired-end sequencing scheme by the ultra-high-throughput DIPSEQ T1 sequencer platform using the following read lengths: 70 bp for read 1, 50 bp for read 2, and 20 bp for the sample index sequencing scheme of the China National GeneBank (CNGB).
Data processing and quality control of snATAC-seq data
We used the newest pipeline to process the snATAC-seq data of DNBelab C4 (https://github.com/MGI-tech-bioinformatics). First, the raw reads were separated into insertions and barcodes and then filtered and demultiplexed using PISA (v1.1) with a minimum sequencing quality of 20. Next, the filtered reads were aligned to the Macaca fascicularis genome (5.0.91) using Chromap (v0.2.3_r407) and a mismatched barcode was not allowed. Reads with mapping quality less than 10 and reads mapped to the mitochondria or genome scaffold (chrAQ*, chrU*, chr*_random*, and chrK*) were filtered out and PCR duplicates were removed according to the cell barcode and mapping loci. The resulting BAM files were processed using d2c (v1.4.4) to identify barcodes from the same cell. The retained fragments of each library were used for further analysis.
Basic processing of the snATAC-seq data was performed using ArchR93 (v1.0.2) package in R (v4.2.2). Genome annotation file of Macaca fascicularis was download from Ensembl (release 91). Fragments files were loaded by “createArrowFiles” function, with chrM and chrY chromosomes excluded. Each sample was separately computed for counts for each tile per cell via “addTileMatrix” function with tileSize = 500. Doublets were identified by “addDoubletScores” function with Doublet score <0.15. A total number of 1,615,430 cells from 405 snATAC-seq libraries was kept with threshold of TSS enrichment >= 5 and <=30 and Promoter ratio >= 0.05 and <= 0.6.
Dimension reduction and clustering of snATAC-seq data
First, we used “addGroupCoverages” function and “addReproduciblePeakSet” function in MACS224 (v2.2.7.1) to create a reproducible merged peak set for each library. A new matrix was created within the new merged peak set with “addPeakMatrix” function for downstream analyses. Next, we further performed an iterative LSI dimensionality reduction via the “addIterativeLSI” function with varFeatures = 50000. Harmony was applied to remove the batch effects across libraries. In order to better cluster the data, we applied a three-step clustering approach to refine cell type identification based on chromatin accessibility. This approach resulted in 3 classes, 29 subclasses and 230 cell types overall. We first carried out unsupervised clustering for all cells with resolution = 3, categorized the initial cell clusters into 3 categories based on classical marker genes, including glutamatergic neurons (SLC17A6, SLC17A7), GABAergic neurons (GAD1, GAD2) and non-neuronal cells (GFAP, PDGFRA, PLP1, PLAC8, CLDN5 and COL1A1), then repeated the above analysis workflow with higher resolution for each category.
A total of 230 cell clusters were kept after removing clusters with the number of cells less than 100, or ambiguous clusters (expressing both glutamatergic (SLC17A6, SLC17A7) and GABAergic (GAD1, GAD2) marker genes or non-neuronal marker genes). We have provided a comprehensive list of marker genes used for the identification of ambiguous clusters in Supplementary Data 2. Previous study showed that most doublets would either form separate clusters, or would tend to end up at the fringes of other clusters (in graph embeddings)94. Therefore, we also applied the same procedure to further remove doublets from the clustering results. Specifically, cells were marked as doublets with their distances to the cluster center lower or higher than 1.5*interquartile range (25th and 75th percentile).
Correspondence of cell types defined by snATAC-seq and snRNA-seq data
We calculated gene activity score for each gene by summing up the counts within the gene body + 2 kb upstream of our snATAC-seq data using ArchR “addGeneScoreMatrix” function, and compared with the snRNA-seq data derived from a recently published macaque cortex study21. We randomly selected 200 cells from each cell cluster for both snATAC-seq and snRNA-seq data, and then performed data normalization using “NormalizeData” function. Integration analysis was conducted with Seurat “IntegrateData” function with top 3000 repeatedly variable genes as features. The proportion of nuclei overlap between omics represents the number of cells clustered in the same clusters of the integrative clustering result divided by the total number of cells for specific cell types from the two omics datasets, using the “compare_cl” function of the published pipeline6.
Identification of reproducible peak sets
We performed peak calling according to the ENCODE ATAC-seq pipeline (https://www.encodeproject.org/atac-seq/). Across each cell type, we pooled all appropriately paired reads to produce a pseudo-bulk ATAC-seq dataset for each biological replicate using “getFrags” function. In addition, we created two pseudo-replicates by dividing the reads from each biological replicate in half. We called peaks for each of the four datasets and a pool of both replicates independently. Peak calling was performed on the Tn5- corrected single-base insertions using MACS2 (v2.2.7.1) with these parameters: ––shift -75 --extsize 150 --nomodel --call-summits --SPMR -q 0.01. Finally, we extended peak summits by 250 bp on each side, resulting in a final width of 501 bp, for merging and downstream analysis. To identify a list of reproducible peaks, we filtered peaks with the following two criteria as in previous studies15, (1) detected in the pooled dataset and overlapped ≥50% of peak length with a peak in both individual replicates; or (2) detected in the pooled dataset and overlapped ≥50% of peak length with a peak in both pseudo-replicates.
Due to the nature of the Poisson distribution test in MACS2, the MACS2 score was influenced by the number of nuclei or the read depth. Thus, we utilized the adjusted normalization method which converted the MACS2 scores (−log10 (q-value)) to “score per million (SPM)”, and filtered reproducible peaks by choosing a SPM cut-off of 2. Finally, we only kept reproducible peaks on chromosome 1–19 and both sex chromosomes, and filtered blacklist regions. A union peak list for the whole dataset was obtained by merging peak sets from all cell types using Bedtools (v2.29.1).
Chromatin co-accessibility identification
We used R package Cicero26 (v1.20.0) to compute co-accessible regions for all open regions. In each cell type, we randomly sampled 5,000 nuclei (All cells were retained for cell types with less than 5,000 nuclei). We set parameters of Cicero as: k = 50, window size = 500 kb, distance constraint = 250 kb. To avoid using an empirical cut-off, we also generated a random shuffled CRE-by-cell matrix and computed co-accessible regions on this shuffled matrix. Then, we fitted the distribution of co-accessibility scores from randomly shuffled background into a normal distribution model by using the R package fitdistrplus. Next, we tested each co-accessibility pair and set the cut-off at co-accessibility score with significance threshold of FDR < 0.01 to filter out co-accessible pairs.
We defined the CREs outside of ±1 kb of the gene transcriptional start site (TSS) as distal and the others as proximal. Then, we divided all co-accessible pairs into three groups: proximal-to-proximal, distal-to-distal, and distal-to-proximal. We focused only on distal-to-proximal pairs because they represent potential gene expression regulatory relationships. We classified all distal CREs in the above distal-to-proximal co-accessible pairs as final CREs.
We annotated the genome distribution of a total of 615873 CREs by “annotatePeak” function from R package ChIPseeker95 (v.1.34.0), setting 1 kb upstream and 1 kb downstream of the TSS as promoter.
Differentially accessible CREs among different cell types and transcription factor (TF) binding motif analysis
We applied Wilcoxon rank sum test using the “getMarkerFeatures” function for identification of marker peaks with useMatrix = “PeakMatrix” at the cluster level. The test was performed as a two-sided test, and p-values were adjusted using the Benjamini-Hochberg false discovery rate (FDR) correction. The CREs with log2 fold-change > 1 and FDR < 0.05 were identified as significant marker features for each cell type. Then, we removed the duplicated CREs and calculated the differential motifs across cell types with gimme maelstrom96. The “gimme.vertebrate.v5.0” was used as reference database.
We assessed TF binding motif enrichment in accessible peaks using chromVAR97 (v1.24.0). Briefly, we initially employed the “addMotifAnnotations” function in ArchR to determine motif presence in the peak set. Subsequently, GC bias was corrected using “addBgdPeaks”, followed by calculation of deviation z-scores for each TF motif in each cell using the “addDeviationsMatrix” function. The cell-by-motif matrix was then stored in ArchR arrow files.
Gene regulatory network inference
We used Python package CellOracle28 (v0.14.0) to infer GRNs by integrating with our previously published snRNA-seq data28. The snRNA-seq data were processed by Seurat with default parameters, using the ‘FindVariableFeatures’ to select the top 3,000 highly variable genes in each corresponding cell subclass, and randomly sampled 5,000 RNA nuclei for each cell subclass. After that, we predicted GRNs by three steps. First, we defined the co-accessible distal-to-proximal pairs as described above. Second, we mapped distal CREs to known motifs by the “scan” function of GimmeMotifs, using the macaque genome macFas5 and the “gimme.vertebrate.v5.0” motif database as references. Finally, we identified the regulatory relationships between TFs and the potential target genes by fitting a regularized linear regression model using processed snRNA-seq data. We evaluated key TFs potentially involved in regulatory functions across different cell types based on “eigenvector_centrality” values. Subsequently, we filtered downstream target genes highly regulated according to their “coef_abs” values.
To identify hub TFs in the putative gene regulatory networks, we then calculated the hub score to infer the importance of different TFs. The hub score was calculated by “hub_score” function from R package igraph (v 2.1.1), with “coef_abs” value as a weight edge attribute.
Integration of snATAC-seq and Stereo-seq data
Data preprocessing
To enable the integration of the query modality (Stereo-seq) and the reference modality (snATAC-seq), a systematic data preprocessing pipeline was designed. First, gene names from both modalities were extracted, and their intersection was computed to generate a shared gene set. The data were subsequently trimmed to a unified gene feature space. For snATAC-seq data, the gene score matrix was subjected to normalization and logarithmic transformation for each cell. Subsequently, the top 3000 highly variable genes were selected to construct a high-information gene subset, which reduced data dimensionality while retaining critical biological features.
To simulate the complexities and uncertainties inherent in cross-modal data integration and to create a more challenging and generalizable data distribution, the gene score matrix of snATAC-seq was intentionally perturbed with added noise. During the perturbation process, a disturbance matrix \({p}_{{ik}}\) was constructed based on the proportional calculation and normalization of gene expression values. This matrix comprised two components: gene-level perturbation and cell-level perturbation.
Gene-level perturbation was calculated based on the ratio of average gene expression values between the Stereo-seq and snATAC-seq modalities. Let \({X}_{{ik}}\) denote the gene activity score matrix of the snATAC-seq modality and \({Y}_{{jk}}\) denote the gene expression matrix of the Stereo-seq modality. The average expression value for each gene k in the two modalities is defined as:
where \({n}_{x}\) and \({n}_{y}\) represent the number of cells in the snATAC-seq and Stereo-seq modalities, respectively.
The average capture rate, \({\bar{{{{\rm{p}}}}}}(p_{{{{\rm{mean}}}}})\), is defined as the ratio of the average gene expression values between the two modalities and calculated as:
Based on this, the gene-specific perturbation factor \({r}_{k}\) is defined as:
The cell-specific average capture rate \({p}_{i}\) is calculated by matching the i-th quantiles of the library sizes between the query modality (\({{{\mathrm{ln}}}}_{{{{\rm{RNA}}}},j}\)) and the reference modality (\({{{\mathrm{ln}}}}_{{{{\rm{ATAC}}}},i}\)). This ratio is used to adjust the gene expression distribution of the reference modality. The formula is as follows:
The cell-specific perturbation factor \({r}_{i}\) is obtained by dividing by \({p}_{{{{\rm{mean}}}}}\):
Finally, by combining the cell-specific perturbation factor and the gene-specific perturbation factor, the gene expression matrix is adjusted to generate the gene-cell perturbation ratio:
Sampling of the original snATAC-seq data is performed through the perturbation matrix, simulating a distribution that is closer to the target modality.
Data perturbed \({X^{\prime} }_{{ik}}\) in this manner serves as a reference, enhancing the model’s robustness to noise in real biological data and providing a complex scenario simulation for experimental design and biological hypothesis testing.
Next, we similarly trim the Stereo-seq data based on the set of highly variable genes derived from the reference modality, ensuring that its features are consistent with those of the snATAC-seq data for seamless cross-modal analysis. Additionally, the trimmed Stereo-seq data undergoes normalization and logarithmic transformation, using the same strategy applied to the snATAC-seq data.
Encoder and decoder
The autoencoder (AE) module consists of an encoder and a decoder, both utilizing fully connected layers to achieve dimensionality reduction and reconstruction of high-dimensional gene features. The encoder maps the input high-dimensional gene expression data (with dimensions corresponding to the number of genes) to a lower-dimensional latent space (with dimensions corresponding to the number of cell types) using a single fully connected layer, with the bias set to False. In the latent space, \({U}_{{ic}}\) \({{{\rm{and}}}}\) \({V}_{{jc}}\) represent cell-type-specific metagene expressions for snATAC-seq and Stereo-seq, which are weighted combinations of different genes, \({W}_{0}\) denotes the weight matrix of the encoder as follows:
As the key component for dimensionality reduction, the encoder’s fully connected layer extracts cell-specific features related to cell types through linear mapping, while incorporating Dropout (dropout rate = 0.8) during the mapping process to enhance the model’s generalization ability and reduce the risk of overfitting.
The decoder employs a symmetric structure, with a fully connected layer that maps the latent variables from the low-dimensional metagene features back to the original high-dimensional space. The decoder’s task is to supervise the encoder by reconstructing the input reference modality gene expression matrix, ensuring that the latent variables extracted by the encoder preserve the major biological information. \({{{{\rm{W}}}}}_{1}\) denotes the weight matrix of the decoder as,
The use of fully connected layers not only simplifies the model design but also enhances the ability to adapt to the dimensionality reduction characteristics of different modalities, enabling the encoder and decoder to exhibit strong robustness in handling high-dimensional sparse data.
Discriminator
The design of the discriminator module is based on the low-dimensional metagene features generated by the encoder, serving as an integral part of the adversarial mechanism. Its task is to distinguish the modality source of the metagene features. The input to the discriminator is the latent variables generated by the encoder, and the output is the probability of modality labels. During adversarial training, the encoder attempts to generate modality-agnostic metagene features to “fool” the discriminator, making it difficult to accurately determine the modality source, thereby achieving cross-modal feature alignment in the latent variable space. \({W}_{2}\) denotes the weight matrix of the discriminator:
Loss function design
Discriminator Loss
The discriminator loss is used to measure the discriminator’s ability to distinguish between two modalities. The goal is to maximize the probability of correct classification while ensuring that the contributions of different modalities are balanced. The formula for calculating the discriminator loss is as follows:
Where:
Cross-entropy loss
This loss is used to evaluate the difference between the model’s predicted categories and the true categories. The formula for calculating the cross-entropy loss is as follows:
Where \({T}_{{ic}}\) denotes the true label indicating whether the observed sample j belongs to category c. To address the class imbalance issue, we utilize class weights \({W}_{c}\) to adjust the contribution of each class to the loss. The class weights are calculated as the inverse of the frequency \({n}_{c}\) of each class:
Reconstruction loss
The reconstruction loss is used to ensure that the output of the autoencoder is as close as possible to the input reference data, thereby preserving key information and minimizing information loss. The basic form of the reconstruction loss is the mean squared error (MSE), and its formula is as follows:
Adversarial loss
The adversarial loss is used to optimize the feature extractor in adversarial learning, making its generated features difficult for the discriminator to distinguish between different modalities. This is achieved by training the feature extractor to “fool” the discriminator, producing modality-agnostic features. The formula is as follows:
Entropy regularization loss
Entropy regularization is used to encourage a more uniform distribution of the model’s output categories, thereby preventing overly confident predictions. The formula for calculating the entropy regularization loss is as follows:
L2 regularization loss
L2 regularization is used to constrain the magnitude of the model parameters, preventing overfitting. The formula is as follows:
Where w represents the model parameters, which are the training parameters of the autoencoder.
Model Training
In the training process, a two-step training approach is employed, optimizing the discriminator and the feature extractor separately. This training strategy aims to achieve effective integration of the feature spaces from the two modalities (snATAC-seq and Stereo-seq) through adversarial learning, while simultaneously performing cell type classification and the reconstruction of input gene expression data. The training parameters set for the entire process include 1000 epochs, the Adam optimizer, and a learning rate of 0.01. Regularization parameters such as the L2 regularization coefficient are set to 0.01, and the dropout rate is set to 0.8.
In the first step, the parameters of the feature extractor (autoencoder module) are fixed, and only the discriminator is optimized. The task of the discriminator is to learn to distinguish the features of data from different modalities, with the input being the features extracted by the autoencoder from both modalities (e.g., snATAC-seq and Stereo-seq). The discriminator’s loss function is based on binary cross-entropy, aiming to maximize the accuracy of modality discrimination, while balancing the class imbalances caused by unequal sample sizes through a weighted loss. Through optimization, the discriminator gradually learns how to distinguish the features of the two modalities, providing a clear optimization target for the adversarial optimization of the feature extractor in the next step. The loss function for the first step is as follows:
In the second step, the parameters of the discriminator are fixed, and only the feature extractor is optimized. The task of the feature extractor is to generate modality-agnostic shared features, while simultaneously performing cell type classification and high-fidelity reconstruction of the input data. To achieve this, the optimization of the feature extractor includes a weighted combination of multiple loss functions:
These two steps are carried out sequentially. In each training round, the discriminator is first updated to enhance its ability to distinguish between modalities, followed by the update of the feature extractor to make the modalities indistinguishable. After multiple rounds of alternative training, the model reaches a balance: the shared features generated by the feature extractor both preserve biological information and are modality-agnostic; the discriminator can no longer improve its classification accuracy, indicating that modality alignment has been achieved.
One-to-one pairing between omics data
The metagene generated \({U}_{{ic}}\),\({V}_{{jc}}\) by the encoder is used for cell type prediction and cell matching. We employ the optimal transport algorithm for cross-modal mapping. The goal of optimal transport is to minimize the distance between cells. The key difference from traditional optimal transport algorithms lies in the computation of the cost matrix. Based on \({U}_{{ic}}\), \({V}_{{jc}}\), we rank the different cell types for each cell according to their probabilities, with the cell type showing the highest probability first and the lowest last, resulting in a type ranking vector s\({U}_{{ic}}\), s\({V}_{{jc}}\) for each cell. For two cells, \(i\) and \(j\), from different modalities, if their top \({B}_{{ij}}\) most likely cell types align, then the cost matrix is defined as:
Next, to solve the optimal transport problem, the following objective function is constructed:
\({T}_{{ij}}\) represents the transport amount from cell i to cell j. We use Earth Mover’s Distance (EMD) to solve this. For each cell j in Stereo-seq, we search for the cell i in snATAC-seq with the highest transport amount. In this way, each cell on the spatial transcriptome map was paired with chromatin accessibility from the snATAC-seq data. The chromatin accessibility of CREs was thus quantified in different layers and regions based on this mapping result.
Layer and regional preference of various cell types and corresponding CREs
Based on one-to-one cell pairing result between snATAC-seq data and Stereo-seq data, we obtained the spatial distribution of 230 cell types among six cortical layers and 143 brain regions, classified into 9 cortical lobes, including prefrontal, frontal, cingulate, somatosensory, insular, auditory, temporal, parietal, and occipital lobes. The cell density per cell cluster in six layers was calculated as the number of cells in the layer divided by the area size of a certain layer. The preference of each cell cluster in different brain regions was calculated as follows:
Here, I represented the specificity index among cortical lobes, \({P}_{j}\) is the proportion in a single brain region j, S is the number of brain regions the cell cluster was distributed.
Cross-species analysis of various cell types
Human and mouse snATAC-seq data were retrieved from previously published studies11,15 and downloaded from the chromatin accessibility database (http://catlas.org/humanbrain and http://catlas.org/mousebrain) and the GEO database11 (accession number: GSE246791). We applied the subsampling method to ensure similar numbers of nuclei across species at the subclass level based on previous studies6. Specifically, the species with the largest number of clusters under a given subclass was allowed a maximum of 200 nuclei per cluster. The remaining species then split this theoretical maximum (200 nuclei multiplied by the maximum number of clusters under the subclass) evenly across their clusters. The intersection of homologous genes among the three species according to the Ensemble ortholog database (version 91) were used for cross-species analysis, and gene activities of the 14,382 orthologous genes were quantified from chromatin accessibility of snATAC-seq data using “addGeneScoreMatrix” and “getMatrixFromArrow” function of ArchR. We then performed data normalization using “NormalizeData” function from Seurat (v4.0.2) package. Further integration analysis was conducted with Seurat “IntegrateData” function with top 2000 repeatedly variable genes as features. The proportion of nuclei overlap between species was calculated by the number of cells clustered in the same clusters of the integrative clustering result divided by the total number of cells for specific cell types from two species, using the “compare_cl” function from the published pipeline6. MetaNeighbor analysis was run to systematically assess the similarity between cell types within species. The data slot of the integrated assay was reformatted and used as input with the “SummarizedExperiment” function of SummarizedExperiment (v.1.28.0) package. We calculated the AUROC score by the “MetaNeighborUS” function of MetaNeighbor (v.1.18.0) package with the parameter “one_vs_best=TRUE”.
Cross-species analysis of CREs
The human and mouse CRE lists were download from http://catlas.org/humanbrain and https://decoder-genetics.wustl.edu/catlasv1/catlas_wholemousebrain/. First, we converted macaque and mouse CREs to human coordinates using the liftOver method with minMatch = 0.3 and UCSC chain files (“macFas5.hg38.all.chain”, “hg38.MacFas5.all.chain”, “mm10.hg38.all.chain”, and “hg38.mm10.all.chain”). Next, we reciprocally lifted the accessible elements back to raw genomes and only kept the regions that mapped to their original loci. We further removed converted regions with length > 1 kb or in chrM, chr_random, chr_Un, and chr_alt chromosomes. Finally, we compared the homologous CREs using intervene98 (v.0.6.5) with f = 0.1. CREs showing both sequence similarity and open chromatin between two species were considered as conserved, whereas CREs with no homologous sequence or no open chromatin peaks in the other species were considered as species-specific. In this way, we classified CREs into 7 categories: Human/Macaque/Mouse-conserved (HMm, mammal-conserved), Human/Macaque-biased (HM, Human/Macaque-biased), Human/Mouse-specific (Hm), Macaque/Mouse-specific (Mm), Human-specific, Macaque-specific, Mouse-specific. We then calculated the proportion of these 7 categories of CREs among all CREs in each cell subclass.
In addition, we download the known transposable elements annotation file from UCSC. Only 4 TE types were kept for further analysis, including “DNA”, “SINE”, “LINE”, “LTR”. We calculated the percentage of CREs located in TE in these 7 categories, and performed pairwise Wilcoxon rank sum test with “bonferroni” correction for multiple testing among various categories.
Comparison of LAMP5 and LAMP5/LHX6 between species
Human and mouse snATAC-seq data were retrieved from previously published studies11,15, and we filtered to retain cells originating from the cortex. Despite some differences in annotation between datasets, we identified the most comparable subclass groups using gene activity scores of classical markers while preserving the original labels from the previous studies for visualization.
In order to compare the cell composition of LAMP5 and LAMP5/LHX6, we merged these two subclasses to calculate their proportions. To facilitate downstream computations, 5,000 cells were randomly sampled from the merged data for each species. Gene activity scores were normalized using a log2 transformation, and we used Seurat and Signac R package to perform integration.
To identify the primate-enriched motifs, we utilized Jaspar motifs (JASPAR2020, all vertebrate) to generate a motif matrix and motif object that was added to the Seurat object using Signac “AddMotifs” function. For motif activity scores, chromVAR was carried out according to default parameters with the Signac “RunChromVAR” function on the respective peak count matrices. The motif matrices of human, macaque, and mouse were merged, and differential enrichment was identified using the “FindMarkers” function with the mean.fxn parameter set to rowMeans.
Characteristics of SST layer and GRN construction
We re-clustered the SST cell subclass as described above to identify cell groups with layer preferences. The previously published study had identified the different layer-specific markers of SST neurons21, so we examined the gene activity score of these layer markers and assign different cell types to the respective layers.
The GRN (gene regulatory network) construction was performed as described above. Briefly, we used CellOracle to calculate the transcription factor (TF)-target gene correlations in different SST groups with layer preferences. For visualization, we ranked the TF by their “eigenvector_centralit” and selected the top 5 TFs. At the same time, we also ranked the target genes by “coef_abs” and selected the top 5 genes. In the network, the target genes which were identified as layer markers in the previous study21 was highlighted.
Correlation of OPC-specific and OLG-specific CREs with cortical hierarchy levels
Hierarchy levels of cortical regions in the visual system were derived from Felleman and Van Essen51. Based on the one-to-one cell pairing result between snATAC-seq data and Stereo-seq data, we extracted the chromatin accessibility matrix of OPC-specific CREs in OPC cells and OLG-specific CREs in OLG cells. Then, we calculated the average chromatin accessibility for each OPC-specific CRE among all OPC cells and OLG-specific CRE among all OLG cells in various cortical regions. We applied the Spearman analysis to calculate the correlation between averaged chromatin accessibility and the hierarchy level for each CRE using the “cor.test” function. The CREs showing positive or negative correlation with the hierarchy levels were identified with cutoffs of P value < 0.01, R > 0.4 or R < -0.4. GO enrichment analysis was performed using these CREs associated genes by clusterProfiler (v.4.6.0).
LDSC cell type-specific analysis
To enable comparison with GWAS data of human phenotypes, we utilized the ‘liftOver‘ with default parameters to convert cell type-specific CREs from “Macaca_fascicularis_5.0” to hg19 genomic coordinates via a chain file “macFas5ToHg19.over.chain” download from UCSC. We further filtered out converted regions exceeding 1 kb and retained only coordinates on each standard chromosome (chromosomes 1-22).
We used Linkage Disequilibrium Score Regression Analysis (LDSC) to assess the extent of genetic polymorphism enrichment associated with chromatin accessible regions differing among various cell types and traits. Summary statistics for GWAS of brain disorders, neurological traits, and control traits were obtained from the GWAS Catalog and publications (Supplementary Data 8). We prepared the summary statistics in the standard format for linkage disequilibrium score regression and downloaded the relevant genotype resource from the pipeline vignettes at https://github.com/bulik/ldsc. Using genotype data from the “1000G_Phase3_plinkfiles.tgz” and haplotype information from ‘hapmap3_snps.tgz’ as reference panels, we generated binary annotations for the homologous DNA sequences of each cell type and estimated LD scores on each standard chromosome. For each trait, we used the baseline model “1000G_Phase3_baseline_ldscores.tgz” along with the standardized regression weights “weights_hm3_no_hla.tgz” to partition phenotypic heritability into cell type-specific annotations, using the superset of all candidate regulatory peaks as background controls.
Variant to CRE mapping
To broadly map the genomic features of disease risk signals across species, we first extracted all potential pathogenic variants from the GWAS summary data that passed the statistical test (P-value ≤ 0.05). Next, we used the R package “SNPlocs.Hsapiens.dbSNP144.GRCh37” along with their rsid information to retrieve the coordinates of these signals in the hg19 reference genome from the “dbSNP144” database. Subsequently, we performed intersection analysis between these coordinates and Cis-Regulatory Elements (CREs) in the Macaca fascicularis genome to identify disease risk-related CREs that may have similar functions across different species.
Gene set related to brain disorder
The gene set related to brain disorders was obtained from Disease Ontology (DO). In R, we used the getHgDisease function from the geneset package to load the DO database by specifying source = “do”. We then queried the DOID (Supplementary Data 8) and employed the transId method from the genekitr package, setting transTo = “symbol” and unique = TRUE, to retrieve the gene symbols.
Sequence evolutionary conservation analysis
The phyloP score is a conservation score that quantifies the evolutionary conservation of each nucleotide by comparing the aligned sequences across multiple species58,59. By evaluating phyloP scores, we assessed the evolutionary conservation of CREs associated with disease risk loci and those not associated with disease, in the context of cell-type genetic identity across species, relative to a shuffle background.
To focus the nucleotide evolutionary conservation assessment on comparisons between the macaque and human and mouse genomes, we first downloaded whole-genome masked sequences for three species (macFas5, mm10, and hg38) from UCSC, retaining the standard chromosome assemblies (sequences prefixed with “chr”) for alignment analysis. Next, we performed reference-based alignment using the ‘LASTZ‘ software, with parameter settings according to UCSC guidelines. For the closely related species macFas5 and hg38, we used the parameters “M = 254 K = 3000 L = 3000 Y = 3400 E = 30 H = 2000 O = 400 T = 2”; for the more distantly related macFas5 and mm10, we used the parameters “M = 0 K = 3000 L = 3000 Y = 9400 E = 30 H = 2000 O = 400 T = 1”. Subsequently, we applied ‘MULTIZ‘ to assemble the pairwise ‘LASTZ‘ alignment results into a multi-genome alignment output and set the phylogenetic tree for the three species to “((macfas5, hg38), mm10)” using the query from https://lifemap-ncbi.univ-lyon1.fr.
For evolutionary model generation, we used ‘phyloFit‘ with the parameters ‘-i MAF --tree “((macfas5, hg38), mm10)” --subst-mod REV‘; then, we used ‘phyloP‘ to generate per-base phyloP score data, with the parameters ‘--mode CONACC --method LRT --wig-scores‘ to compare the evolutionary conservation of the macaque genome against human and mouse genomes. Finally, we obtained the average phyloP scores for all CREs using ‘bigWigAverageOverBed‘.
In the statistical analysis, for each cell type’s diff-CREs that co-localized with disease risk signals, we calculated its average phyloP score. To ensure a fair comparison of the evolutionary conservation of these Risk CREs, we performed 1000 bootstrap resampling on other diff-elements not belonging to Risk CREs and the supercluster of peaks, each time sampling an equal number of elements as the Risk CREs. We then aggregated the average scores from the 1000 re-samplings for statistical analysis.
Gene ontology (GO) enrichment
Gene Ontology annotation of interested CREs was performed using R package rGREAT99 (v.2.2.0). We downloaded AH111915 (Macaca fascicularis) database as a reference using AnnotationHub (v. 3.8.0). For the pathway gene sets of Gene Ontology database, we extended the genes from their transcription start site with default parameters of “basal_upstream = 5000” and “basal_downstream = 1000”. The total CREs of corresponding cell clusters were used as the background peak set for each. GREAT analysis was performed with “great” function of “min_gene_set_size = 5”.
RNA scope
Macaque #3 was transcardially perfused with diethyl pyrocarbonate-treated PBS (DEPC-PBS), and then followed by 4% DEPC-PFA. The brain was removed and immediately embedded in optimum cutting temperature compound (OCT, SAKURA, cat#4583). The brain samples were stored at -80 °C. During samples processing, the OCT blocks were sectioned at 15 μm using a RWD freezing microtome, and brain sections were fixed onto adhesion microscope slides. The slides were baked at 60 °C overnight. Then, RNA Scope Pretreatment Kit (Advanced Cell Diagnostics, cat#322381 and cat#322000) was used to unmask target RNA and permeabilize cells. Subsequently, RNA Scope Target Probe are hybridized to target RNA molecules. RNA Scope Detection Reagents (ACD, cat#322310) amplify the hybridization signals via sequential hybridization of amplifiers and color label probes. The RNA Scope procedure was performed according to the manufacturer’s instructions (https://acdbio.com). The details about the probes employed in the experiments are as follows: Probes against CPA6 (ACD, cat#1843691-C1) and MEF2C (ACD, cat#1843711-C2) were used to identify V1-specific L4 neurons in macaque monkeys; Probes against DCC (ACD, cat#1843701-C1) were used to identify layer4 neurons in macaque monkeys.
Data visualization
For bigwig files, presented by Signac package “BigwigTrack” function. For the putative gene regulatory networks, presented with Cytoscape (v.3.10.1) tools. The residual graphs were plotted with R package ggplot2 (v.3.4.1) and pheatmap (v.1.0.12).
Ethics statement
Animal protocol was reviewed and approved by the Biomedical Research Ethics Committee of CAS Center for Excellence in Brain Science and Intelligence Technology, Chinese Academy of Sciences (ION-2019011). Animal care complied with the guideline of this committee. Adult macaque monkeys (M. fascicularis; #1, 6-year-old, 4.2 kg, male; #2, 4-year-old, 3.7 kg, male; #3, 12-year-old, 5.1 kg, female) were used in this study.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
For snATAC-seq data in this paper, all raw data are available in CNGB Nucleotide Sequence Archive under accession code CNP0005530. The processed data ready for exploration could be accessed and downloaded via https://macaque.digital-brain.cn/chromatin-accessibility. The stereo-seq data are from a previous study and available via https://macaque.digital-brain.cn/spatial-omics/singleCellData?project=neocortex. The public snATAC-seq data of human and mouse are accessible via https://catlas.org/catlas/. Source data are provided with this paper.
Code availability
All data were analyzed with standard programs and packages, as detailed above, and the corresponding codes were accessible through https://github.com/JuneMeng/snATAC_MacaqueCTX_Atlas and at the Zenodo (https://doi.org/10.5281/zenodo.18431761)100. Custom code of cell-cell pairing of snATAC-seq and spatial transcriptome data is available at https://github.com/sunyk740/RMGE/ and at the Zenodo (https://doi.org/10.5281/zenodo.18432489)101. Additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
References
Lein, E., Borm, L. E. & Linnarsson, S. The promise of spatial transcriptomics for neuroscience in the era of molecular cell typing. Science 358, 64–69 (2017).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, (2018).
Khrameeva, E. et al. Single-cell-resolution transcriptome map of human, chimpanzee, bonobo, and macaque brains. Genome Res 30, 776–789 (2020).
Krienen, F. M. et al. Innovations present in the primate interneuron repertoire. Nature 586, 262–269 (2020).
Berg, J. et al. Human neocortical expansion involves glutamatergic neuron diversification. Nature 598, 151–158 (2021).
Bakken, T. E. et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 598, 111–119 (2021).
(BICCN) BICCN. A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 598, 86-102 (2021).
Yao, Z. et al. A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex. Nature 598, 103–110 (2021).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat. Neurosci. 21, 432–439 (2018).
Cusanovich, D. A. et al. A Single-Cell Atlas of In Vivo Mammalian Chromatin Accessibility. Cell 174, 1309–1324 e1318 (2018).
Zu, S. et al. Single-cell analysis of chromatin accessibility in the adult mouse brain. Nature 624, 378–389 (2023).
Li, Y. E. et al. An atlas of gene regulatory elements in adult mouse cerebrum. Nature 598, 129–136 (2021).
Villar, D. et al. Enhancer evolution across 20 mammalian species. Cell 160, 554–566 (2015).
Won, H. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016).
Li, Y. E. et al. A comparative atlas of single-cell chromatin accessibility in the human brain. Science 382, eadf7044 (2023).
Zemke, N. R. et al. Conserved and divergent gene regulatory programs of the mammalian neocortex. Nature 624, 390–402 (2023).
Yin, S. et al. Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain. Nat. Commun. 11, 474 (2020).
Chiou, K. L. et al. A single-cell multi-omic atlas spanning the adult rhesus macaque brain. Sci. Adv. 9, eadh1914 (2023).
Lei, Y. et al. Spatially resolved gene regulatory and disease-related vulnerability map of the adult Macaque cortex. Nat. Commun. 13, 6747 (2022).
Han, L. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature 604, 723–731 (2022).
Chen, A. et al. Single-cell spatial transcriptome reveals cell-type organization in the macaque cortex. Cell 186, 3726–3743.e3724 (2023).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Yao, Z. et al. A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell 184, 3222–3241.e3226 (2021).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Su, Y. et al. Neuronal activity modifies the chromatin accessibility landscape in the adult brain. Nat. Neurosci. 20, 476–483 (2017).
Pliner, H. A. et al. Cicero Predicts cis-Regulatory DNA Interactions from Single-Cell Chromatin Accessibility Data. Mol. Cell 71, 858–871.e858 (2018).
Reiprich, S. et al. Transcription factor Sox10 regulates oligodendroglial Sox9 levels via microRNAs. Glia 65, 1089–1102 (2017).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Basu, S. et al. The Mef2c Gene Dose-Dependently Controls Hippocampal Neurogenesis and the Expression of Autism-Like Behaviors. J Neurosci 44, (2024).
Pandya, D., Seltzer, B., Petrides, M. & Cipolloni, P. B. Cerebral Cortex: Architecture, Connections, and the Dual Origin Concept. (Oxford University Press, 2015).
Shibata, M. et al. Hominini-specific regulation of CBLN2 increases prefrontal spinogenesis. Nature 598, 489–494 (2021).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Manitt, C. et al. dcc orchestrates the development of the prefrontal cortex during adolescence and is altered in psychiatric patients. Transl. Psychiatry 3, e338 (2013).
Gabitto, M. I. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. 27, 2366–2383 (2024).
Seo, M. H., Lim, S. & Yeo, S. Association of decreased triadin expression level with apoptosis of dopaminergic cells in Parkinson’s disease mouse model. BMC Neurosci. 22, 65 (2021).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory activities of transposable elements: from conflicts to benefits. Nat. Rev. Genet 18, 71–86 (2017).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Gouwens, N. W. et al. Integrated Morphoelectric and Transcriptomic Classification of Cortical GABAergic Cells. Cell 183, 935–953.e919 (2020).
Wei, J. R. et al. Identification of visual cortex cell types and species differences using single-cell RNA sequencing. Nat. Commun. 13, 6902 (2022).
Chen, W. G. et al. Upstream stimulatory factors are mediators of Ca2+-responsive transcription in neurons. J. Neurosci. 23, 2572–2581 (2003).
Flavell, S. W. et al. Genome-wide analysis of MEF2 transcriptional program reveals synaptic target genes and neuronal activity-dependent polyadenylation site selection. Neuron 60, 1022–1038 (2008).
Micali, N. et al. Molecular programs of regional specification and neural stem cell fate progression in macaque telencephalon. Science 382, eadf3786 (2023).
Jahangir, M., Shah, S. M., Zhou, J. S., Lang, B. & Wang, X. P. Parvalbumin interneurons in the anterior cingulate cortex exhibit distinct processing patterns for fear and memory in rats. Heliyon 11, e41218 (2025).
Qi, C. et al. Anterior cingulate cortex parvalbumin and somatostatin interneurons shape social behavior in male mice. Nat. Commun. 16, 4156 (2025).
Fang, R. et al. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 377, 56–62 (2022).
Shi, Y. et al. Mouse and human share conserved transcriptional programs for interneuron development. Science 374, eabj6641 (2021).
Urban-Ciecko, J. & Barth, A. L. Somatostatin-expressing neurons in cortical networks. Nat. Rev. Neurosci. 17, 401–409 (2016).
Berto, S. et al. Accelerated evolution of oligodendrocytes in the human brain. Proc. Natl. Acad. Sci. USA 116, 24334–24342 (2019).
Miller, D. J. et al. Prolonged myelination in human neocortical evolution. Proc. Natl. Acad. Sci. USA 109, 16480–16485 (2012).
Caglayan, E. et al. Molecular features driving cellular complexity of human brain evolution. Nature 620, 145–153 (2023).
Felleman, D.J. & Van Essen, D.C. Distributed hierarchical processing in the primate cerebral cortex. Cerebral cortex (New York, NY: 1991) 1, 1–47 (1991).
Ghorbani, S. et al. Fibulin-2 is an extracellular matrix inhibitor of oligodendrocytes relevant to multiple sclerosis. J Clin Invest 134, (2024).
Li, Y. et al. Npas3 deficiency impairs cortical astrogenesis and induces autistic-like behaviors. Cell Rep. 40, 111289 (2022).
Cummings, A. C. et al. Genome-wide association and linkage study in the Amish detects a novel candidate late-onset Alzheimer disease gene. Ann. Hum. Genet 76, 342–351 (2012).
Schizophrenia Working Group of the Psychiatric Genomics, C. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Loo, S. K. et al. Genome-wide association study of intelligence: additive effects of novel brain expressed genes. J. Am. Acad. Child Adolesc. Psychiatry 51, 432–440.e432 (2012).
Seshadri, S. et al. Genetic correlates of brain aging on MRI and cognitive test measures: a genome-wide association and linkage analysis in the Framingham Study. BMC Med Genet 8, S15 (2007).
Zhang, G. et al. Construction of single-cell cross-species chromatin accessibility landscapes with combinatorial-hybridization-based ATAC-seq. Dev. Cell 59, 793–811.e798 (2024).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res 20, 110–121 (2010).
Bipolar, D. Schizophrenia Working Group of the Psychiatric Genomics Consortium. Electronic address drve, Bipolar D, Schizophrenia Working Group of the Psychiatric Genomics C. Genomic Dissection of Bipolar Disorder and Schizophrenia, Including 28 Subphenotypes. Cell 173, e1716 (2018).
Merenlender-Wagner, A. et al. Autophagy has a key role in the pathophysiology of schizophrenia. Mol. Psychiatry 20, 126–132 (2015).
Liu, X., Bipolar Genome Study (BiGS), Kelsoe, J. R. & Greenwood, T. A. A genome-wide association study of bipolar disorder with comorbid eating disorder replicates the SOX2-OT region. J. Affect Disord. 189, 141–149 (2016).
Kahler, A. K. et al. Association analysis of schizophrenia on 18 genes involved in neuronal migration: MDGA1 as a new susceptibility gene. Am. J. Med Genet B Neuropsychiatr. Genet 147B, 1089–1100 (2008).
Banfi, F. et al. SETBP1 accumulation induces P53 inhibition and genotoxic stress in neural progenitors underlying neurodegeneration in Schinzel-Giedion syndrome. Nat. Commun. 12, 4050 (2021).
Yamazaki, K. et al. ABCA7 Gene Expression and Genetic Association Study in Schizophrenia. Neuropsychiatr. Dis. Treat. 16, 441–446 (2020).
Gotoh, L. et al. Association analysis of adenosine A1 receptor gene (ADORA1) polymorphisms with schizophrenia in a Japanese population. Psychiatr. Genet 19, 328–335 (2009).
Takahashi, N. et al. Association between chromogranin A gene polymorphism and schizophrenia in the Japanese population. Schizophr. Res 83, 179–183 (2006).
Pozhidaev, I. V. et al. Association of cholinergic muscarinic M4 receptor gene polymorphism with schizophrenia. Appl Clin. Genet 13, 97–105 (2020).
Li, M. et al. Novel genetic susceptibility loci identified by family based whole exome sequencing in Han Chinese schizophrenia patients. Transl. Psychiatry 10, 5 (2020).
Zhang, Z. et al. The Schizophrenia Susceptibility Gene OPCML Regulates Spine Maturation and Cognitive Behaviors through Eph-Cofilin Signaling. Cell Rep. 29, 49–61.e47 (2019).
Ventriglia, M. et al. Allelic variation in the human prodynorphin gene promoter and schizophrenia. Neuropsychobiology 46, 17–21 (2002).
Juraeva, D. et al. Integrated pathway-based approach identifies association between genomic regions at CTCF and CACNB2 and schizophrenia. PLoS Genet 10, e1004345 (2014).
Tiihonen, J. et al. Molecular signaling pathways underlying schizophrenia. Schizophr. Res 232, 33–41 (2021).
Le Hellard, S. et al. Polymorphisms in SREBF1 and SREBF2, two antipsychotic-activated transcription factors controlling cellular lipogenesis, are associated with schizophrenia in German and Scandinavian samples. Mol. Psychiatry 15, 463–472 (2010).
Aberg, K. A. et al. Methylome-wide association study of schizophrenia: identifying blood biomarker signatures of environmental insults. JAMA Psychiatry 71, 255–264 (2014).
Souza, R. P. et al. Genetic association of the GDNF alpha-receptor genes with schizophrenia and clozapine response. J. Psychiatr. Res 44, 700–706 (2010).
Pinacho, R. et al. Altered CSNK1E, FABP4 and NEFH protein levels in the dorsolateral prefrontal cortex in schizophrenia. Schizophr. Res 177, 88–97 (2016).
Shim, K. H. et al. Subsequent correlated changes in complement component 3 and amyloid beta oligomers in the blood of patients with Alzheimer’s disease. Alzheimers Dement 20, 2731–2741 (2024).
Colonna, M. & Wang, Y. TREM2 variants: new keys to decipher Alzheimer disease pathogenesis. Nat. Rev. Neurosci. 17, 201–207 (2016).
Wang, Y. et al. Key variants via the Alzheimer’s Disease Sequencing Project whole genome sequence data. Alzheimers Dement 20, 3290–3304 (2024).
Joshi, H. et al. Decreased Expression of Cerebral Dopamine Neurotrophic Factor in Platelets of Probable Alzheimer Patients. Alzheimer Dis. Assoc. Disord. 36, 269–271 (2022).
Corraliza-Gomez, M. et al. Insulin-degrading enzyme (IDE) as a modulator of microglial phenotypes in the context of Alzheimer’s disease and brain aging. J. Neuroinflammation 20, 233 (2023).
Chang, J. Y. & Chang, N. S. WWOX dysfunction induces sequential aggregation of TRAPPC6ADelta, TIAF1, tau and amyloid beta, and causes apoptosis. Cell Death Discov. 1, 15003 (2015).
Chiba-Falek, O., Gottschalk, W. K. & Lutz, M. W. The effects of the TOMM40 poly-T alleles on Alzheimer’s disease phenotypes. Alzheimers Dement 14, 692–698 (2018).
Jackson, R. J., Hyman, B. T. & Serrano-Pozo, A. Multifaceted roles of APOE in Alzheimer disease. Nat. Rev. Neurol. 20, 457–474 (2024).
Reynolds, L. M. et al. DCC Receptors Drive Prefrontal Cortex Maturation by Determining Dopamine Axon Targeting in Adolescence. Biol. Psychiatry 83, 181–192 (2018).
Li, H. J. et al. Further confirmation of netrin 1 receptor (DCC) as a depression risk gene via integrations of multi-omics data. Transl. Psychiatry 10, 98 (2020).
Vosberg, D. E., Leyton, M. & Flores, C. The Netrin-1/DCC guidance system: dopamine pathway maturation and psychiatric disorders emerging in adolescence. Mol. Psychiatry 25, 297–307 (2020).
Tricoire, L. et al. Common origins of hippocampal Ivy and nitric oxide synthase expressing neurogliaform cells. J. Neurosci. 30, 2165–2176 (2010).
Kosoy, R. et al. Genetics of the human microglia regulome refines Alzheimer’s disease risk loci. Nat. Genet 54, 1145–1154 (2022).
Mathys, H. et al. Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186, 4365–4385.e4327 (2023).
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PloS one 13, e0209648 (2018).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Zeisel, A. et al. Molecular Architecture of the Mouse Nervous System. Cell 174, 999–1014.e1022 (2018).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
van Heeringen, S. J. & Veenstra, G. J. GimmeMotifs: a de novo motif prediction pipeline for ChIP-sequencing experiments. Bioinformatics 27, 270–271 (2011).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Gu, Z. & Hübschmann, D. rGREAT: an R/bioconductor package for functional enrichment on genomic regions. Bioinformatics 39, (2023).
JuneMeng & chenc. JuneMeng/snATAC_MacaqueCTX_Atlas: Single-cell spatial map of cis-regulatory elements for disease-related genes in the macaque cortex (v1.0). Zenodo. https://doi.org/10.5281/zenodo.18431762 (2026).
sunyk740. sunyk740/RMGE: RMGE V1.0(v1.0.2). Zenodo. https://doi.org/10.5281/zenodo.18432489 (2026).
Acknowledgements
The project was supported by National Key Research and Development Program of China (No. 2024YFC3408000), National Science and Technology Innovation 2030 Major Program (STI2030-2021ZD0200101 and 2021ZD0200103), National Natural Science Foundation of China (NSFC) Outstanding Youth Foundation (No. T2422026), National Key R&D Program of China No.2022YEF0203200, National Natural Science Foundation of China (No. U23A6010), Lingang Laboratory (No. LGL-6672-07) and Shanghai Science and Technology Development Funds (23QA1410400) to Y-D.S.
Author information
Authors and Affiliations
Contributions
Methodology, J.M., C.C., Z.Z., Y.S., K.H. and T.F.; Investigation, J.M., Y.L., C.C., Z.Z., Y.B.; Formal Analysis, J.M., C.C., Z.Z., Y.S. and Y.H.; Validation, J.F., L.W., J.M. and Li.L.; Writing – Original Draft, Y.L. and Y-D.S.; Writing – Review & Editing, J.M., T.F., Z.L., Ch.L., Z.S., L-Q.L., C-Y.L., T.S., C-R.L., and M.P.; Conceptualization and supervision, S.L., Y.L. and Y-D.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yang Yu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Meng, J., Chen, C., Zhu, Z. et al. Single-cell spatial map of cis-regulatory elements for disease-related genes in the macaque cortex. Nat Commun 17, 4041 (2026). https://doi.org/10.1038/s41467-026-70497-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70497-x