Abstract
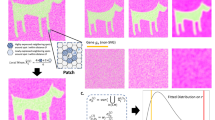
In single-cell experiments spanning diverse conditions, distinguishing variation specific to one condition (e.g., treatment) from shared or background variation (e.g., control) is critical for uncovering treatment-specific molecular responses. However, these studies typically yield ultra-high-dimensional data, necessitating effective dimension reduction for reliable biological interpretation. Contrastive dimension reduction methods address this challenge by identifying low-dimensional features enriched in a target dataset relative to a background dataset that captures shared variation. Despite their growing utility, the success of such methods critically depends on the choice of background, yet no formal criterion exists for evaluating or selecting backgrounds. To address this gap, we introduce BasCoD, a statistical testing framework based on spectral subspace inclusion theory, that enables rigorous evaluation and systematic selection of background datasets. Applying BasCoD across a range of single-cell datasets, we show that it effectively identifies suitable backgrounds, substantially improving the contrast and interpretability of the resulting target representations. We further demonstrate how BasCoD can guide the design of contrastive analyses in large-scale single-cell experiments conducted under heterogeneous conditions and elucidate potential interaction effects in perturbation studies.
Similar content being viewed by others
Data availability
• Mouse protein expression data. The processed mouse protein expression data used in this study are available at the cPCA GitHub repository. • Perturb-seq data. The processed Perturb-seq data used in this study are available at Figshare through the ContrastiveVI tutorial. • Mouse intestinal single-cell RNA-seq data. The processed mouse intestinal single-cell RNA-seq data used in this study are available at the ContrastiveVI tutorial. • Human Cell Atlas bone marrow (HCA-BM) data. The processed HCA-BM data used in this study are available at the Lamian GitHub repository. • Population-scale single-cell RNA-seq data with ROT treatment. The processed population-scale single-cell RNA-seq data used in this study are available in the Zenodo database under accession code 4333872. • Single-cell RNA-seq data with inflammation treatment. The processed single-cell RNA-seq data used in this study are available at Zenodo: https://zenodo.org/records/18776758. Source data are provided with this paper.
Code availability
The code used to develop the model, perform the analyses and generate results in this study is publicly available and has been deposited in the GitHub repository at https://github.com/keleslab/BasCoD, under the MIT license. The specific version of the code associated with this publication is archived in Zenodo and is accessible via https://doi.org/10.5281/zenodo.1829118331.
References
Jerber, J. et al. Population-scale single-cell rna-seq profiling across dopaminergic neuron differentiation. Nat. Genet. 53, 304–312 (2021).
Soskic, B. et al. Immune disease risk variants regulate gene expression dynamics during cd4+ t cell activation. Nat. Genet. 54, 817–826 (2022).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell rna profiling of pooled genetic screens. cell 167, 1853–1866 (2016).
Abid, A., Zhang, M. J., Bagaria, V. K. & Zou, J. Exploring patterns enriched in a dataset with contrastive principal component analysis. Nat. Commun. 9, 2134 (2018).
Abid, A., Zou, J. Contrastive variational autoencoder enhances salient features. arXiv preprint arXiv:1902.04601 (2019).
Severson, K.A., Ghosh, S., Ng, K. Unsupervised learning with contrastive latent variable models. In: Proceedings of the AAAI Conference on Artificial Intelligence. Volume 33. 4862–4869 (2019).
Weinberger, E., Lin, C. & Lee, S. I. Isolating salient variations of interest in single-cell data with contrastivevi. Nat. Methods 20, 1336–1345 (2023).
Weinberger, E., Covert, I., & Lee, S. I. Feature selection in the contrastive analysis setting. Adv. Neural. Inf. Process. Syst. 36, 66102–66126 (2023).
Zhang, B., Nyquist, S., Jones, A., Engelhardt, B. E. & Li, D. Contrastive linear regression. Ann. Appl. Stat. 19, 1868 (2025).
Ebrahimi, A., Siahpirani, A. F. & Montazeri, H. scin: a contrastive learning framework for single-cell multi-omics data integration. Brief. Bioinforma. 26, bbaf411 (2025).
Li, W., Murtaza, G. & Singh, R. sccontrast: A contrastive learning based approach for encoding single-cell gene expression data. bioRxiv 2025–04 (2025).
He, K., Fan, H., Wu, Y., Xie, S. & Girshick, R. Momentum contrast for unsupervised visual representation learning. In: Proc. of the IEEE/CVF conference on computer vision and pattern recognition 9729–9738 (2020).
Hawke, S., Zhang, E., Chen, J. & Li, D. Contrastive dimension reduction: A systematic review. arXiv preprint arXiv:2510.11847 (2025).
Townes, F. W., Hicks, S. C., Aryee, M. J. & Irizarry, R. A. Feature selection and dimension reduction for single-cell rna-seq based on a multinomial model. Genome Biol. 20, 295 (2019).
Ahmed, M. M. et al. Protein dynamics associated with failed and rescued learning in the ts65dn mouse model of down syndrome. PloS one 10, e0119491 (2015).
Higuera, C., Gardiner, K. J. & Cios, K. J. Self-organizing feature maps identify proteins critical to learning in a mouse model of down syndrome. PloS one 10, e0129126 (2015).
Norman, T. M. et al. Exploring genetic interaction manifolds constructed from rich single-cell phenotypes. Science 365, 786–793 (2019).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. methods 15, 1053–1058 (2018).
Wilson, D. J. The harmonic mean p-value for combining dependent tests. Proc. Natl. Acad. Sci. 116, 1195–1200 (2019).
Trapnell, C. et al. Pseudo-temporal ordering of individual cells reveals dynamics and regulators of cell fate decisions. Nat. Biotechnol. 32, 381 (2014).
Ji, Z. & Ji, H. Tscan: Pseudo-time reconstruction and evaluation in single-cell rna-seq analysis. Nucleic acids Res. 44, e117–e117 (2016).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC genomics 19, 1–16 (2018).
Hou, W. et al. A statistical framework for differential pseudotime analysis with multiple single-cell rna-seq samples. Nat. Commun. 14, 7286 (2023).
DeTomaso, D. & Yosef, N. Hotspot identifies informative gene modules across modalities of single-cell genomics. Cell Syst. 12, 446–456 (2021).
Ficara, F. et al. Pbx1 restrains myeloid maturation while preserving lymphoid potential in hematopoietic progenitors. J. cell Sci. 126, 3181–3191 (2013).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterprofiler: an r package for comparing biological themes among gene clusters. Omics: a J. Integr. Biol. 16, 284–287 (2012).
Murray, C. W. et al. LKB1 drives stasis and C/EBP-mediated reprogramming to an alveolar type II fate in lung cancer. Nat. Commun. 13, 1090 (2022).
Lara-Astiaso, D. et al. In vivo screening characterizes chromatin factor functions during normal and malignant hematopoiesis. Nat. Genet. 55, 1542–1554 (2023).
Hawke, S., Ma, Y. & Li, D. Contrastive dimension reduction: when and how?. Adv. Neural Inf. Process. Syst. 37, 74034–74057 (2024).
Liu, Y. & Xie, J. Cauchy combination test: a powerful test with analytic p-value calculation under arbitrary dependency structures. J. Am. Stat. Assoc. 115, 393–402 (2020).
Park, K., Sun, Z., Liao, R., Bresnick, E.H. & Keleş, S. Systematic background selection with bascod enhances contrastive dimension reduction in single cell genomics. GitHub Repository: BasCoDhttps://doi.org/10.5281/zenodo.18291183 (2026).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Acknowledgements
We thank Shuchen Yan from the University of Wisconsin–Madison for sharing the processed scRNA-seq dataset from Jerber et al.1. We thank Dr. Siqi Shen (Fred Hutchinson Cancer Center) and Coleman Breen (University of Wisconsin–Madison) for insightful discussions. This work was supported by NIH grants R01HG003747 (S.K.) and R21HG012881 (S.K.), and a Chan Zuckerberg Initiative Data Insights Award (S.K.).
Author information
Authors and Affiliations
Contributions
K.P. and S.K. conceived the project. K.P. and S.K. designed the research and developed the method. K.P. performed the experiments and simulation studies. K.P. and S.K. contributed to the preparation of the manuscript. R.L. and E.B. generated the single-cell RNA-seq dataset with inflammation treatment. Z.S. processed the dataset and designed experimental ideas involving double-perturbed cells.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chaojie Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, K., Sun, Z., Liao, R. et al. Systematic background selection with BasCoD enhances contrastive dimension reduction in single cell genomics. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70652-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70652-4