Abstract
Brain metastasis, a common complication of advanced non-small cell lung cancer (NSCLC), leads to poor prognosis and reduces the efficacy of immune checkpoint inhibitors (ICIs). However, the mechanisms underlying this diminished ICI response remain unclear. Here we perform single-cell RNA sequencing on 101,959 tumor-infiltrating immune cells from primary tumors and brain metastases to delineate their distinct immune landscapes. Brain metastases display profound immunosuppressive reprogramming, characterized by enrichment of HSP70-high stress-responsive T cells and PLTP⁺ tumor-associated macrophages, and depletion of memory T cells and cDC2-like dendritic cells, which show opposing associations with ICI outcomes. Integrating these findings, we derive a seven-gene brain metastasis–derived immune signature (BMIS) that is associated with ICI response and survival in NSCLC and metastatic urothelial carcinoma and provides information complementary to tumor mutational burden. These results highlight immune features specific to brain metastatic NSCLC and suggest candidate biomarkers and therapeutic avenues for improving immunotherapy in this high-risk population.
Similar content being viewed by others
Introduction
Non-small cell lung cancer (NSCLC) is the leading cause of cancer-related deaths worldwide1. Brain metastases (BMs) are common in patients with advanced NSCLC and are associated with poor prognosis2,3. Therapeutic options for patients with BMs are limited, as systemic treatments such as chemotherapy are less effective owing to the protective nature of the blood–brain barrier (BBB). Immunotherapy with immune checkpoint inhibitors (ICIs) that target the PD-1/PD-L1 axis has become the standard first-line treatment for advanced NSCLC. This treatment has shown promise for managing BMs4,5,6. However, many patients do not respond to this therapy, and inconsistent responses between primary tumors and brain metastases are frequently observed7. This suggests significant heterogeneity in the tumor immune microenvironment (TIME) between primary and metastatic sites.
The brain is an immune-privileged site with a complex immune landscape. The development of BMs disrupts the BBB. This allows immune cells, such as T cells and monocyte-derived macrophages, to infiltrate affected regions and potentially alter the TIME8,9. BMs have lower CD8⁺ T cell infiltration and higher PD-1 and PD-L1 expression than primary tumors (PTs)10,11,12,13. Recent studies have leveraged single-cell RNA sequencing (scRNA-seq) to elucidate the immune microenvironment in brain metastatic NSCLC and other cancers. These studies have demonstrated the broad immunosuppressive mechanisms in BMs, including enrichment of regulatory T cells, T-cell exhaustion, and macrophage recruitment14,15,16,17,18,19. However, these studies have predominantly focused on general cellular heterogeneity rather than specific immune profiles linked to therapeutic resistance. In addition, only a few studies have mapped precise immune cell populations that might contribute to the diminished efficacy of ICIs. The key aspects of immune dysfunction in the BM, including the comprehensive characterization of immune cell subtypes and their specific roles in ICI resistance, remain unclear.
In this study, we aimed to create a comprehensive, high-resolution map of the TIME to elucidate the immune landscape specific to BM NSCLC. Our findings provide a resource and generate hypotheses for the development of tailored strategies that enhance the efficacy of ICIs in patients with BM. Furthermore, the present study highlights immune features unique to BM NSCLC.
Results
Single-cell transcriptomic analyses reveal distinct immune landscapes in primary and brain metastatic NSCLC
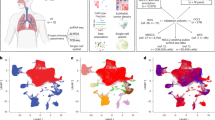
To investigate the differing immune microenvironments of PTs and BMs in NSCLC, we performed scRNA-seq on live, immune-related (CD45⁺) cells isolated from both sites using fluorescence-activated cell sorting (FACS) (Fig. 1A). We also integrated scRNA-seq data from previous studies by selecting BM samples containing more than 2,000 cells14. Together, these constituted our discovery cohort of 19 samples (8 PTs and 11 BMs). After stringent quality control and doublet removal, our final dataset comprised 117,752 cells with an average of 1532 genes and 5519 unique molecular identifiers (UMIs) per cell (Figure S1A–C, Supplementary Data 1). Removal of batch effects between our inhouse and integrated samples were performed using Harmony algorithm (Figure S1D).
A Schematic overview of the study design. Created in BioRender. Bai, M. (2026) https://BioRender.com/b4nlzov. B–D UMAP plots of 117,752 cells colored by cell type (B), sample origin—primary or brain metastasis (C), and individual samples (D). E A dot plot showing expression levels of marker genes across the indicated cell types. F UMAP plots displaying marker gene expression used for cell type identification.
We used unsupervised clustering based on gene expression profiles to identify seven major cell types characterized by lineage-specific markers: 15,019 epithelial cells (EPCAM, CDH1), 774 cancer-associated fibroblasts (COL1A2, SPARC), 70,895 T and natural killer (NK) cells (CD2, CD3D), 7097 B cells (CD19, MS4A1), 3484 plasma cells (MZB1, IGHG3), 2798 mast cells (KIT, CPA3), and 17,685 myeloid cells (LYZ, ITGAX) (Fig. 1B–F). Data on epithelial cells and cancer-associated fibroblasts were obtained from previous studies and were excluded from further analyses.
HSP70-High T cells enriched in BMs are associated with poor immunotherapy outcomes in NSCLC
Given the pivotal role of tumor-infiltrating lymphocytes in the tumor microenvironment and their impact on patient outcomes, we first evaluated T cell populations. We compared the transcriptional profiles of CD4⁺ and CD8⁺ T cells from PTs and BMs to investigate the unique transcriptional state of T cells in BMs. The heat shock proteins HSPA1A and HSPA1B were among the most upregulated genes in both CD4⁺ and CD8⁺ T cells in BMs (Fig. 2A; Figure S2A, B, and Supplementary Data 2, 3). To determine whether this upregulation extended to other HSP70 family members, we examined the expression of additional isoforms (HSPA5, HSPA8, HSPA9 etc.). However, these isoforms showed no significant differences between PTs and BMs (Figure S2C), indicating that the observed upregulation was specific to the inducible HSP70 isoforms HSPA1A and HSPA1B. Sample-level analysis of HSP70 module scores based on expression of HSPA1A and HSPA1B confirmed significantly elevated expression in both CD4⁺ (p = 0.027) and CD8⁺ (p = 0.0018) T cells from BMs compared to PTs (Fig. 2B). Based on this score, we categorized CD4⁺ and CD8⁺ T cells into HSP70-High, HSP70-Medium, and HSP70-Low groups. scRNA-seq data revealed an increased proportion of HSP70-High CD4⁺ and CD8⁺ T cells in BMs (Figure S2D).
A Volcano plots showing differentially expressed genes between primary and brain metastatic NSCLC in CD4⁺ and CD8⁺ T cells. B Boxplots comparing the mean HSP70 score between primary and brain metastatic samples. P-values from two-sided Wilcoxon Rank Sum tests are shown. CD4⁺ T cells: Primary n = 8 and Brain n = 10; CD8⁺ T cells: Primary n = 8 and Brain n = 11. C Representative multiplex immunofluorescent images of NSCLC sections stained for CD4 (green), CD8 (red), and HSP70 (yellow); PTs and BMs are compared. Scale bars, 50 μm (main panels) and 10 μm (insets). Images are representative of n = 34 PT and n = 26 BM patients. D, E Box plots quantifying HSP70⁺ CD4⁺ T (D) and HSP70⁺ CD8⁺ T E cells from multiplex immunofluorescence images. (n = 34 Primary and n = 26 Brain samples). P-values from two-sided Wilcoxon Rank Sum tests are shown. F A dotplot illustrating average score of curated gene signatures in HSP70-High, HSP70-Medium, and HSP70-Low CD4+ (left) and CD8+ (right) T cells. G, H Enriched Gene Ontology (GO) terms in HSP70-High CD8⁺ T (G) and HSP70-High CD4⁺ T (H) cells. I, J Kaplan–Meier survival curves for patients with NSCLC in the Ravi (n = 122 patients) and Kim cohorts (n = 27 patients), stratified by HSP70-High CD8⁺ T cell (I) and HSP70-High CD4⁺ T cell (J) signature scores. Optimal cutoffs were determined using “surv_cutpoint” within each cohort. Hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (B, D, E), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
To address potential confounding effects of disease stage, we assembled a stage-matched validation dataset by integrating publicly available scRNA-seq data of T cells from stage III-IV NSCLC patients with both primary tumors and brain metastases14,20,21,22 (Supplementary Data 4, Figure S2D, E). The upregulation of HSPA1A and HSPA1B in T cells showed a consistent trend in this stage-matched validation cohort (Figure S2E, F), suggesting that elevated HSP70 expression may be intrinsic to the brain metastatic microenvironment rather than solely a consequence of disease stage differences. In our in-house SDZL cohort (Supplementary Data 5), comprised of treatment naïve PT (n = 34) and BM (n = 26) samples form surgical resection or diagnostic biopsy, multiplex immunofluorescence staining revealed elevated HSP70 expression across multiple cell types in BMs (Fig. 2C). While HSP70 upregulation was not restricted to T cells, reflecting a broader stress response in the metastatic microenvironment, we focused detailed quantification on T cells given their central role as effector cells and pharmacological targets of immune checkpoint blockade. This analysis showed a significantly higher proportion of HSP70⁺ CD4⁺ T cells (p = 0.002, Fig. 2D) and a trend toward higher HSP70⁺ CD8⁺ T cells (p = 0.081, Fig. 2E) in BMs compared to PTs. Consistently, when we restricted the mIF analysis to stage IV patients (10 PT vs. 26 BM), BMs again showed higher proportions of HSP70⁺ CD4⁺ (p = 0.019) and a trend toward higher HSP70+ CD8⁺ (p = 0.073) T cells than extracranial PTs, indicating that this pattern persists within advanced-stage disease (Figure S2G).
The elevated expression of HSPA1A or HSPA1B in CD4⁺ and CD8⁺ T cells is associated with a heightened cellular response state linked to immunotherapy resistance23. Thus, we analyzed curated T cell signatures to further explore the functional characteristics of HSP70-High T cells (Fig. 2F). Both CD4⁺ and CD8⁺ HSP70-High T cells displayed elevated stress response signatures and reduced naïve signatures compared to their HSP70-Low counterparts. Beyond these shared features, HSP70-High CD4⁺ T cells exhibited increased exhaustion signatures, whereas HSP70-High CD8⁺ T cells showed elevated cytotoxicity signatures accompanied by attenuated NF-κB signaling. Pathway enrichment analysis based on sub-cluster-specific differentially expressed genes revealed that both HSP70-High CD4⁺ and CD8⁺ T cells were significantly enriched for biological processes related to protein folding, response to topologically incorrect protein, and chaperone-mediated protein folding (Fig. 2G, H; Supplementary Data 6, 7), consistent with an active cellular stress response program.
To assess the clinical relevance of HSP70-High T cells, we analyzed their association with patient outcomes in two independent NSCLC immunotherapy cohorts24,25. Higher signature scores for HSP70-High CD8⁺ T cells were significantly associated with shorter progression-free survival (PFS) in both the Ravi cohort (p = 0.015) and Kim cohort (p = 0.032), indicating association with poor immunotherapy outcome (Fig. 2I). For HSP70-High CD4⁺ T cells, higher signature scores were similarly associated with worse outcomes in the Ravi cohort (p = 0.038), with a consistent trend observed in the Kim cohort (p = 0.081) (Fig. 2J).
To elucidate the mechanisms underlying the upregulation of HSP70 expression in T cells, we performed NicheNet analysis to predict the ligands driving transcriptomic changes in these cells. PTGS2, encoding the COX-2 enzyme, was prioritized as a top candidate upstream regulator for both HSP70-high CD4⁺ and CD8⁺ T cells (Figure S2H), which we interpret as implicating activation of a COX-2–PGE₂ axis rather than PTGS2 acting as a classical secreted ligand. Additionally, the signatures of HSP70-High CD4+ and CD8+ T cells were significantly correlated with PTGS2 expression in the TCGA-LUAD cohort (Figure S2I).
In summary, our analysis revealed that T cells in brain metastases are characterized by elevated expression of HSPA1A and HSPA1B, which encode inducible HSP70 proteins. These HSP70-high T cells exhibit heightened stress responses, and their enrichment is associated with poor outcomes following immunotherapy, suggesting they may contribute to the immunosuppressive microenvironment in brain metastatic NSCLC.
Depletion of memory T cells in brain metastases is associated with immunotherapy resistance in NSCLC
Further re-clustering of T/NK cells in PTs and BMs based on transcriptional profiles and curated gene signatures revealed 15 distinct clusters. These included two NK cell clusters—NK_FGFBP2 and NK_XCL1—characterized by high expression of natural killer markers, and 13 T cell clusters marked by CD3D⁺, CD4⁺, and CD8⁺ expression (Fig. 3A, B, Figure S3, Figure S4A, Supplementary Data 8).
A A UMAP plot of all T and NK cells, colored by identified cell clusters. B A heat map illustrating normalized expression of canonical T and NK cell marker genes across clusters; TRM denotes tissue-resident memory cells. C A dotplot showing average score of curated gene signatures among T cell clusters. D Box plots comparing the proportions of T and NK cell subclusters between primary (n = 8) and brain metastatic (n = 11) NSCLC samples. P-values from two-sided Wilcoxon Rank Sum tests are shown. E, F Enriched GO terms in CD4_IL7R (E) and cycling (F) cells. P-values were calculated using one-sided hypergeometric tests with Benjamini–Hochberg FDR correction. G, H Kaplan–Meier survival curves for patients with NSCLC in the Ravi (n = 122 patients) and Kim (n = 27 patients) cohorts, as stratified by high and low levels of CD4_IL7R (G) and cycling (H) cells signature score; optimal cutoff values were determined using “survminer.” Hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (D), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
Among CD4⁺ T cells, we identified seven functionally distinct subtypes. The central memory-like T cell cluster (Tcm-like, CD4_IL7R) exhibited elevated expression of canonical central memory markers including CCR7, SELL, IL7R, TCF7, and KLF2, alongside early activation markers CD69 and CD40LG (Fig. 3B, Supplementary Data 8). While these cells are present in the tumor microenvironment and undergo clonal expansion due to antigen stimulation, as indicated by CD69 expression, they exhibit gene expression patterns associated with long-term survival and self-renewal, similar to central memory T cells. The TIMP1⁺ memory T cell cluster (TIMP1+ Tn, CD4_TIMP1) showed high expression of IL7R, TIMP1, FTH1, and CD40LG. The interferon-stimulated gene (ISG)-enriched helper T cell cluster (ISG⁺ Th, CD4_ISG) was characterized by elevated expression of interferon-response genes and displayed Th1-associated features including CXCR3 and TBX21 expression (Fig. 3C, Figure S4B, C). The stress-response T cell cluster (Tstr, CD4_NR4A1) exhibited elevated expression of heat shock proteins (HSPA1A, HSPA1B, HSP90AA1) and showed high stress response signature scores among CD4⁺ T cells. The follicular helper T cell cluster (Tfh, CD4_CXCL13) was marked by high expression of CXCL13, CTLA4, PDCD1, TOX, and TIGIT, with elevated exhaustion signatures. The regulatory T cell cluster (Treg, CD4_FOXP3) displayed classical Treg markers including TNFRSF4, IL2RA, FOXP3, and CTLA4. An additional CD4⁺ cluster (CD4_XIST) was also identified. Assessment of classical Th1/Th2/Th17 lineage markers revealed that while CD4_ISG exhibited Th1-associated features, no clusters showed strong Th2 (CCR4⁺GATA3⁺) or Th17 (CCR6⁺RORC⁺IL17A⁺) polarization (Figure S4B, C).
Among CD8⁺ T cells, we identified six functionally distinct subtypes. The effector memory T cell cluster (Tem, CD8_GZMK) displayed high expression of effector molecules GZMK, GZMA, KLRG1, and CD44, with elevated activation signature scores. The tissue-resident memory T cell cluster (Trm, CD8_ZNF683) exhibited characteristic residency markers, including CD69, ITGAE, and ZNF683, with downregulated tissue egress markers CCR7, SELL, and S1PR1. The interferon-response T cell cluster (Tisg, CD8_ISG) showed elevated expression of interferon-stimulated genes and high interferon-response signature scores. The proliferating T cell cluster (CD8_Cycling) was characterized by high expression of proliferation markers MKI67, TOP2A, and STMN1, alongside elevated metabolic signatures. The exhausted T cell cluster (Tex, CD8_HAVCR2) displayed high expression of exhaustion markers HAVCR2, LAG3, TIGIT, PDCD1, and CTLA4. Consistent with previous findings, this cluster also expressed cytotoxicity-associated genes including CXCL13, ENTPD1, and TNFRSF9, suggesting these are antigen-experienced cells. The precursor exhausted T cell cluster (Tpex, CD8_SPRY1) occupied an intermediate state between effector and exhausted T cells, showing relatively lower expression of TOX and checkpoint inhibitors, while maintaining higher expression of TCF7, IL7R, and CD28 (Fig. 3A–C, Figure S4D, Supplementary Data 8). The annotation of these T cell clusters was supported by gene signatures from published studies23,26,27.
Comparative analysis of these sub-clusters between PTs and BMs, using both Wilcoxon sum rank test and scCODA analysis28, a Bayesian compositional data analysis framework, revealed notable differences (Fig. 3D, Figure S4E). BMs exhibited a lower fraction of CD4⁺ Tcm-like cells (CD4_IL7R) and CD8+ Trm cells (CD8_ZNF683) than the PTs. In contrast, BMs showed a trend of enrichment proliferating CD8⁺ T cells (CD8_Cycling). Given the imbalanced EGFR mutation distribution between PTs and BMs in our cohort (Supplementary Data 1), we performed logistic regression analysis adjusting for EGFR status. Both CD4_IL7R and CD8_ZNF683 depletion in BMs remained statistically significant after adjustment (CD4_IL7R: adjusted P = 0.0002, OR change = 5.1%; CD8_ZNF683: adjusted P = 0.00046, OR change = 31.7%), demonstrating that memory T cell depletion is largely independent of EGFR mutation status, though CD8_ZNF683 shows potential EGFR-associated variation (Supplementary Data 9). In the stage-matched validation cohort, we identified 12 major T/NK cell clusters (Figure S4F, G, Supplementary Data 10). Consistent with our discovery cohort, we identified CD4_IL7R as central memory-like T cells characterized by elevated IL7R and CD40LG expression pattern (Figure S4H). Further, we found CD4_IL7R were significantly depleted in BMs compared to PTs in validation cohort (p = 0.012, Figure S4I).
Pathway enrichment analysis of subcluster-specific differentially expressed genes revealed that CD4_IL7R cells were enriched in cellular responses to extracellular stimuli, including oxygen levels, hormonal signals, and mechanical cues, alongside lymphocyte differentiation pathways (Fig. 3E). CD8_ZNF683 cells displayed elevated expression of genes associated with lymphocyte-mediated immunity and cytotoxicity, including T cell receptor signaling, antigen receptor-mediated signaling, and cell killing pathways (Fig. 3F). These findings suggest that both CD4_IL7R and CD8_ZNF683 cells contribute to active immune surveillance and anti-tumor responses.
Pseudotime trajectory analysis using Monocle 2 with latent-dynamics validation by scTour confirmed the developmental relationships among T cell subclusters (Figure S4J, K). Both CD4_IL7R and CD8_ZNF683 occupied early positions in their respective developmental trajectories, consistent with their memory phenotypes and capacity for differentiation into more specialized effector states.
To assess the clinical relevance of these T cell populations, we analyzed bulk RNA-seq data from patients with NSCLC undergoing immunotherapy. Higher abundance of CD4_IL7R cells was significantly associated with improved PFS in the Ravi cohort (p = 0.0078) and showed a consistent trend in the Kim cohort (p = 0.057) (Fig. 3G). Similarly, elevated CD8_ZNF683 levels correlated with better prognosis in the Kim cohort (p = 0.00073), although this association did not reach statistical significance in the Ravi cohort (p = 0.28) (Fig. 3H). In contrast, increased abundance of proliferating CD8⁺ T cells (CD8_Cycling) were associated with poor outcomes in both cohorts (Ravi: p = 0.0028; Kim: p = 0.057) (Figure S4L). Additionally, we examined the clinical relevance of the precursor exhausted T cell cluster CD8_SPRY1 (Tpex), which represents a critical target population for PD-1 inhibitors. Higher abundance of CD8_SPRY1 cells was associated with improved prognosis in patients receiving immunotherapy (Ravi: p = 0.0014; Kim: p = 0.0065) (Figure S4M), consistent with their role as a renewable reservoir for anti-tumor immune responses.
These findings revealed distinct T cell populations with contrasting prognostic roles: CD4_IL7R central memory-like T cells and CD8_ZNF683 tissue-resident memory T cells, which are enriched in PTs and associated with favorable clinical outcomes; and proliferating CD8_Cycling cells, which are enriched in BMs and linked to poor prognosis following immunotherapy.
Anti-inflammatory cDC2-like dendritic cells are abundant in primary NSCLC and enhance immunotherapeutic efficacy
Owing to the crucial regulatory role of myeloid cells in the TIME, particularly in BMs, we analyzed this cell lineage. Unsupervised clustering revealed 12 myeloid cell subsets, including four tumor-associated macrophage clusters (TAM_MARCO, TAM_PLTP, TAM_SPP1, and TAM_MKI67), two monocyte clusters (Mono_FCN1 and Mono_TIMP1), four dendritic cell (DC) clusters (DC_CLEC10A, DC_CLEC9A, DC_CD207, and DC_LAMP3), and one neutrophil cluster (Fig. 4A; Figure S5). These clusters exhibited distinct gene expression patterns and transcription factor profiles (Fig. 4B, C; Supplementary Data 11).
A A UMAP plot of all myeloid cells, colored by identified cell clusters. B–D Heat maps showing normalized expression of differentially expressed genes (B), transcription factors (C), and canonical dendritic cell marker genes (D) among clusters; TLRs denote toll-like receptors. E Box plots comparing the proportions of dendritic cell subclusters between primary (n = 8) and brain metastatic (n = 11) NSCLC samples. P values from two-sided Wilcoxon Rank Sum tests are shown. F, G Enriched GO terms in DC_CLEC10A (F) and DC_CD207 (G) cells. P-values were calculated using one-sided hypergeometric tests with Benjamini–Hochberg FDR correction. H, I Trajectory analysis of dendritic cells inferred by Monocle 2, colored by cell cluster (H) and pseudotime (I). J Kaplan–Meier survival curve for patients with NSCLC in the Ravi cohort (n = 122 patients), as stratified by high and low levels of DC_CLEC10A cells; the optimal cutoff value was determined using “survminer.” Hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (E), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
Examination of DC populations (Figure S6A) revealed distinct transcriptional profiles between DCs from PTs and BMs. Differential expression analysis showed that DCs in PTs upregulated genes associated with antigen presentation and immune activation, while DCs in BMs showed upregulation of genes related to metabolic reprogramming and stress response (Figure S6B; Supplementary Data 12). Pathway enrichment analysis further confirmed these differences: DCs from PTs were enriched for pathways related to antigen processing and presentation, peptide antigen assembly with MHC class II, and adaptive immune response, whereas DCs from BMs showed enrichment in ATP metabolic processes and oxidative phosphorylation (Figure S6C). Among the four DC clusters, DC_CLEC9A displayed high expression of cDC1 markers (XCR1, CLEC9A, CADM1); DC_CD207 and DC_CLEC10A were associated with cDC2 markers (CD1C, FCER1A, CLEC10A); and DC_LAMP3 was classified as activated DCs, characterized by high expression of activation and co-stimulatory genes (LAMP3, CD80, CD86) (Fig. 4D).
Comparative analysis of DC subcluster proportions between PTs and BMs, using both Wilcoxon rank-sum tests and scCODA, revealed that the two cDC2 clusters, DC_CLEC10A and DC_CD207, were significantly enriched in PTs (p = 0.00099 and p = 0.028, respectively, scCODA FDR < 0.05) (Fig. 4E, Figure S6D). Pathway enrichment analysis of cluster-specific differentially expressed genes revealed that DC_CLEC10A was enriched for antigen presentation of exogenous antigen, antigen presentation via MHC class II, peptide antigen assembly with MHC class II, and MHC protein complex assembly (Fig. 4F). Similarly, DC_CD207 displayed enrichment in antigen processing and presentation, peptide antigen assembly with MHC class II, and lymphocyte-mediated immunity (Fig. 4G). These findings indicate that both cDC2 populations contribute to antigen presentation and T cell activation in primary tumors. Pseudotime trajectory analysis using Monocle 2, validated by scTour, revealed the developmental relationships among DC subclusters (Fig. 4H, I; Figure S6E). Both DC_CD207 and DC_CLEC10A occupied early to intermediate positions along the developmental trajectory, consistent with their roles as antigen-presenting cells capable of T cell priming.
To assess the clinical relevance of these DC populations, we analyzed their association with patient outcomes in NSCLC patients receiving immunotherapy. The abundance of DC_CLEC10A was significantly correlated with improved overall survival (p = 0.035, Fig. 4J), whereas DC_CD207 showed a trend toward worse survival that did not reach statistical significance (p = 0.073, Figure S6F). These findings suggest a distinct distribution pattern of DC subclusters between PTs and BMs in NSCLC, with DC_CLEC10A representing a prognostically favorable population associated with enhanced immunotherapy efficacy.
Distinct monocyte subsets in NSCLC with differential associations to immunotherapy response
We identified two monocyte subclusters characterized by elevated expression of monocyte markers, such as FCN1, VCAN, S100A8, and S100A9 (Fig. 5A; Figure S7A). Comparative analysis of monocyte proportions between PTs and BMs revealed significant differences by Wilcoxon rank-sum test (Mono_FCN1 enriched in PTs, p = 0.011; Mono_TIMP1 enriched in BMs, p = 2.6 × 10⁻⁵). scCODA compositional analysis, which accounts for the interdependence of cell type proportions, confirmed Mono_FCN1 depletion in BMs as the primary compositional shift (Fig. 5B, Figure S6D). This distribution pattern was validated by multiplex immunofluorescence, which demonstrated significantly higher levels of CD14⁺FCN1⁺ cells in primary tumors (p = 7.5 × 10⁻⁵), with a trend toward increased CD14⁺TIMP1⁺ cells in brain metastases, though this did not reach statistical significance (p = 0.15) (Fig. 5C, D). In a subset analysis restricted to stage IV disease (10 PT vs. 26 BM), PTs again exhibited higher proportions of CD14⁺FCN1⁺ cells (p = 0.0085), whereas BMs showed a similar trend toward higher CD14⁺TIMP1⁺ cells (p = 0.17), indicating that this pattern persists within advanced-stage disease and is more consistent with tissue-specific rather than purely stage-driven differences (Figure S7B).
A UMAP plots showing two monocyte subclusters. B Box plots comparing the proportions of monocyte subclusters between primary (n = 8) and brain metastatic (n = 11) NSCLC samples. P values from two-sided Wilcoxon Rank Sum tests are shown. C Representative multiplex immunofluorescent images of NSCLC sections stained for TIMP1 (green), FCN1 (red), and CD14 (yellow), comparing PTs and BMs. Scale bars, 50 μm. Images are representative of n = 34 PT and n = 26 BM patients. D Box plots quantifying FCN1⁺ CD14⁺ and TIMP1⁺ CD14⁺ cells from the immunofluorescence images (n = 34 PT and n = 26 BM patients). P values from two-sided Wilcoxon Rank Sum tests are shown. E Violin plots showing pseudobulk expression (log2 CPM) of representative genes in Mono_FCN1 (green) and Mono_TIMP1 (orange) (n = 8 PT and n = 11 BM samples). P-values calculated using two-sided quasi-likelihood F-tests in edgeR with FDR correction. F Enriched GO terms in Mono_FCN1 and Mono_TIMP1 cells. P-values were calculated using one-sided hypergeometric tests with Benjamini–Hochberg FDR correction. G Trajectory analysis of monocytes and tumor-associated macrophages inferred by Monocle 2, colored by cell cluster. H, I Kaplan–Meier survival curves for NSCLC patients (n = 122 patients) in the Ravi cohort, as stratified by high and low levels of Mono_FCN1 (H) and Mono_TIMP1 (I) cells; optimal cutoff values were determined using “survminer.” Hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (B, D), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
Transcriptional profiling using pseudobulk analysis revealed distinct functional programs between these monocyte subsets (Figure S7C). Mono_FCN1 was characterized by elevated expression of immediate early transcriptional regulators (FOS, FOSB, and NR4A2) together with immune activation markers (NLRP3 and CD83), suggesting that this subset represents an activated monocyte population undergoing transcriptional reprogramming associated with differentiation and immune activation. In contrast, Mono_TIMP1 exhibited higher expression of genes involved in immunometabolic reprogramming including MIF, LDHA, ENO1, TPI1 and GSTO1, indicative of a metabolically altered state that may support immunosuppressive functions (Fig. 5E, Figure S5C, Supplementary Data 13).
Pathway enrichment analysis further confirmed these functional distinctions (Fig. 5F). Mono_FCN1 was enriched for pathways related to response to corticosteroid, positive regulation of leukocyte-mediated cytotoxicity, chronic inflammatory response, and regulation of B cell activation, consistent with an activated, pro-inflammatory phenotype. In contrast, Mono_TIMP1 showed enrichment in ribonucleoside metabolic processes, ATP metabolic processes, and purine nucleoside triphosphate metabolic processes, underscoring its metabolically reprogrammed state. Trajectory analysis using Monocle 2 and scTour confirmed that Mono_TIMP1 represents naïve-like monocytes, while Mono_FCN1 represents an intermediate state between monocytes and macrophages (Fig. 5G, Figure S7D).
We analyzed bulk RNA-seq data from patients with NSCLC to evaluate the potential clinical relevance of these monocyte subclusters. While Mono_FCN1 abundance showed no significant association with PFS (p = 0.25, Fig. 5H), higher abundance of Mono_TIMP1 was associated with poorer prognosis (p = 0.02, Fig. 5I). These findings suggest that Mono_FCN1 is associated with an activated, pro-inflammatory state in primary tumors, whereas Mono_TIMP1 exhibits metabolic reprogramming in brain metastases that may contribute to an immunosuppressive microenvironment and reduced immunotherapy efficacy.
TAM subclusters in brain metastatic NSCLC modulate the immunosuppressive microenvironment and correlate with immunotherapeutic efficacy
Tumor-associated macrophages (TAMs) mediate antitumor immunity in PTs and BMs. We identified four distinct TAM sub-clusters in the present study (Fig. 6A). Of these subclusters, TAM_MARCO and TAM_PLTP exhibited elevated expression of immunosuppressive genes such as C1QA, C1QB, GPNMB, and APOE. Similarly, the expression of M2-like signature genes such as CD163, CCL18, MSR1, and that of MHC class II genes was elevated. This expression profile suggests an involvement in antigen presentation and the modulation of the immune microenvironment. Furthermore, TAM_SPP1 was associated with tumor angiogenesis and immune evasion based on elevated levels of VEGFA, CD163 and TGFB1. Contrastingly, TAM_MKI67 was characterized by high expression of proliferation markers MKI67 and TOP2A, indicating its role in the proliferation of TAMs (Figure S7A).
A UMAP plots showing four TAM subclusters. B Box plots comparing the proportions of TAM subclusters between primary (n = 8) and brain metastatic (n = 11) NSCLC samples. P-values from two-sided Wilcoxon Rank Sum tests are shown. C Representative multiplex immunofluorescent images of NSCLC sections stained for CD68 (green) and PLTP (red), comparing PTs and BMs. Scale bars, 50 μm. Images are representative of n = 34 PT and n = 26 BM patients. D Box plots quantifying PLTP⁺ CD68⁺ cells from the immunofluorescence images (n = 34 PT and n = 26 BM patients). P-values from two-sided Wilcoxon rank-sum tests are shown. E Volcano plot of pseudobulk differential gene expression between TAM_MARCO and TAM_PLTP. Significantly upregulated genes (FDR < 0.05, |log2FC | > 0.5) are colored: TAM_MARCO-enriched genes in dark grey, TAM_PLTP-enriched genes in red. F Violin plots showing pseudobulk expression (log2 CPM) of representative genes in TAM_MARCO (dark grey) and TAM_PLTP (blue) (n = 8 PT and n = 11 BM samples). P-values calculated using two-sided quasi-likelihood F-tests in edgeR with FDR correction. G Bar plots showing enriched GO terms in TAM_MARCO and TAM_PLTP cells. P-values were calculated using one-sided hypergeometric tests with Benjamini–Hochberg FDR correction. H, I Kaplan–Meier survival curves for NSCLC patients (n = 122 patients) in the Ravi cohort, as stratified by high and low levels of TAM_MARCO (H) and TAM_PLTP (I) cells; optimal cutoff values were determined using “survminer.” Hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (B, D), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
Comparative analysis of TAM subcluster proportions between PTs and BMs, using both Wilcoxon rank-sum tests and scCODA, revealed significant differences (Fig. 6B, Figure S6D). TAM_MARCO was significantly enriched in PTs (p = 0.0053), whereas TAM_PLTP was predominantly found in BMs (p = 2.6 × 10⁻⁵), indicating site-specific functional specialization of macrophage populations. This differential distribution was validated by multiplex immunofluorescence staining, which demonstrated significantly higher levels of CD68⁺PLTP⁺ cells in BMs compared to PTs (p = 6.8 × 10⁻⁸) (Fig. 6C, D). A subset analysis restricted to stage IV disease (10 PT vs. 26 BM) likewise confirmed enrichment of CD68⁺PLTP⁺ cells in BMs (p = 0.0009), indicating that this pattern persists within advanced-stage disease and is more consistent with tissue-specific rather than purely stage-driven differences (Figure S7E).
To characterize the functional differences between TAM_MARCO and TAM_PLTP, we evaluated macrophage polarization signatures and performed pseudobulk differential expression analysis. TAM_MARCO exhibited elevated M0 and M1 signature scores and higher expression of interferon-stimulated genes (MX1, IFIT3, ISG15) alongside immune activation markers (CD44, TRIM22). In contrast, TAM_PLTP showed elevated M2 signature scores and higher expression of genes associated with immunoregulatory functions, including PLTP, FOLR2, and CD300A, as well as complement components C1QA and C1QB (Fig. 6E, F; Figure S7F, G; Supplementary Data 14).
Pathway enrichment analysis of differentially expressed genes between two subclusters revealed that TAM_MARCO and TAM_PLTP engage distinct immune activation programs (Fig. 6G). TAM_MARCO was enriched for interferon-mediated signaling and antiviral defense pathways, characteristic of anti-tumor macrophages. TAM_PLTP was enriched for complement activation (classical pathway, humoral immune response by immunoglobulin, B cell-mediated immunity) and adaptive immune responses. While both populations display immune activation signatures, complement-mediated immunity in the tumor microenvironment has been linked to tumor promotion through angiogenesis and recruitment of immunosuppressive myeloid cells29,30,31, consistent with the divergent clinical outcomes associated with these TAM subtypes.
To assess the clinical relevance of these TAM populations, we analyzed their association with patient outcomes in NSCLC patients receiving immunotherapy. Higher abundance of TAM_MARCO showed a trend toward improved PFS (p = 0.071, Fig. 6H), whereas elevated TAM_PLTP levels were significantly associated with poor prognosis (p = 0.016, Fig. 6I). These findings indicate that TAM_PLTP may contribute to the immunosuppressive microenvironment characteristic of brain metastatic NSCLC.
Characteristics of B and plasma cell subclusters in PTs and BMs
B and plasma cells are actively involved in antitumor immunity and are associated with the efficacy of immunotherapy. We re-clustered these cells into 14 distinct sub-clusters to assess their landscape in the PTs and BMs of NSCLC. These included five memory B-cell clusters (Bm_MS4A1, Bm_CD83, Bm_CD74, Bm_ALDOA, and Bm_FCRL4), two naïve B-cell clusters (Bn_IGHD and Bn_SPP1), a germinal center B-cell cluster (Bgc_LMO2), and six plasma cell clusters (PC_IGHG1, PC_FKBP2, PC_XBP1, PC_JSRP1, PC_MKI67, and PC_IGHM) (Fig. 7A, Figure S8A–E, Supplementary Data 15).
A UMAP plots showing B and plasma cell subclusters. B Box plots comparing the proportions of B and plasma cell subclusters between primary (n = 8) and brain metastatic (n = 11) NSCLC samples. P-values from two-sided Wilcoxon Rank Sum tests are shown. C Violin plots showing marker gene expression of Bn_IGHD cells across clusters. D A heat map displaying potential ligands driving the phenotype of Bn_IGHD cells. E Bar plots showing enriched GO terms in Bn_IGHD cells. P-values were calculated using one-sided hypergeometric tests with Benjamini–Hochberg FDR correction. F Kaplan–Meier survival curve for NSCLC patients (n = 122 patients) in the Ravi cohort, as stratified by high and low levels of Bn_IGHD cells; the optimal cutoff value was determined using “survminer”. Hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (B), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
We compared the relative abundance of these clusters between PTs and BMs to characterize their functions. We observed a decreased proportion of the naïve B cell cluster Bn_IGHD in BMs in both Wilcoxon and scCODA analysis (Fig. 7B, Figure S8F). This cluster was characterized by the elevated expression of IGHD, FCER2, IL4R, and YBX3 (Fig. 7C). B cells expressing IL4R and FCER2 are associated with B cell activation and increased IgE production32. Ligand prediction using NicheNet further identified interleukin-4 as a specific ligand for Bn_IGHD (Fig. 7D). Pathway enrichment analysis revealed that this cluster was enriched for B cell activation, leukocyte cell-cell adhesion, and antigen presentation pathways, including peptide antigen assembly with MHC class II and MHC class II protein complex assembly (Fig. 7E). These findings suggest that Bn_IGHD cells contribute to B cell-mediated immune responses and antigen presentation in primary tumors.
We explored the clinical relevance of Bn_IGHD by assessing its association with patient outcomes. Enrichment of this naïve B cell cluster showed a trend toward improved PFS (p = 0.06, Fig. 7F). These findings suggest that Bn_IGHD may contributes to antitumor immunity and serves as a potential biomarker for therapeutic responses.
Coordinated immune cell networks differ between primary tumors and brain metastases
To assess whether the immune cell populations identified above vary independently or form coordinated modules, we next quantified the correlations between the relative abundances of all immune subclusters across PT and BM samples. Spearman correlation analysis revealed groups of positively correlated clusters as well as pairs with strong inverse relationships, indicating that patients tend to segregate into distinct immune compositions rather than showing isolated changes in single subsets (Figure S9A). We then compared correlation coefficients between PT and BM samples and observed substantial remodeling of these associations, including both strengthened and weakened correlations, with multiple cluster pairs showing large shifts in correlation strength (|Δr | > 1.0), consistent with rewiring of the immune ecosystem in brain metastases (Figure S9B).
To probe potential functional links between key clusters, we further applied CellChat to model ligand–receptor–mediated communication among Mono_FCN1, DC_CLEC10A, CD4_IL7R, CD8_TRM, and TAM_PLTP. This analysis identified several interaction pathways, including adhesion and antigen-presentation related axes (e.g., ADGRE5–CD55, ANXA1–FPR1, APP–CD74, ICAM–ITGB2 and LGALS9–HAVCR2), that connect Mono_FCN1 and DC_CLEC10A with CD4_IL7R and CD8_TRM, many of which showed altered communication probabilities between PT and BM, and highlighted additional signals from TAM_PLTP to CD4_IL7R (Figure S9C). Together, these data indicate that the favorable PT-enriched populations (Mono_FCN1, DC_CLEC10A, CD4_IL7R, CD8_TRM) and the BM-enriched immunosuppressive TAM_PLTP cluster participate in a coordinated and site-specific interaction network, rather than acting in isolation.
BM-derived Immune Signature (BMIS) associates with immunotherapy efficacy across multiple cancers
To validate the clinical relevance of the cell subpopulations identified through scRNA-seq, we initially performed unsupervised clustering of the Ravi cohort based on signature scores of six key cell populations differentially abundant between primary tumors and brain metastases (CD4_IL7R, DC_CLEC10A, TAM_MARCO, HSP70-High CD8, HSP70-High CD4, TAM_PLTP). This analysis identified four distinct patient clusters characterized by differing immune infiltration profiles (Fig. 8A). Cluster 2, marked by elevated infiltration of brain metastasis-enriched populations (HSP70-High T cells and TAM_PLTP), showed a trend toward worse overall survival in the Ravi cohort, though this did not reach statistical significance (p = 0.054, Fig. 8B, Figure S10A). A similar pattern was observed in the mUC cohort (p = 0.065, Fig. 8C, Figure S10B, C).
A Heatmap showing unsupervised clustering of NSCLC patients in the Ravi cohort (n = 122 patients) based on signature scores of six cell populations. Clinical characteristics (gender, age, initial stage) and signature scores are annotated. B, C Kaplan-Meier survival curves comparing overall survival of Cluster 2 (high brain metastasis-enriched populations) versus other clusters in the Ravi cohort (n = 122 NSCLC patients) (B) and mUC cohort (n = 298 mUC patients) (C). P-values from two-sided log-rank tests are shown. D Workflow for BMIS development. Differentially expressed genes (DEGs) from brain metastasis-enriched cell populations were subjected to feature selection using Variable Selection Using Random Forests (VSURF), yielding a 7-gene signature used to construct a logistic regression model. Created in BioRender. Bai, M. (2026) https://BioRender.com/xfyb5zp. E Kaplan-Meier curves for progression-free survival (left) and overall survival (right) stratified by BMIS in the Ravi cohort (n = 122 NSCLC patients). Tick marks indicate censored observations. F Distribution of BMIS scores across immunotherapy response categories (PD, n = 41 patients; SD, n = 32 patients; PR, n = 42 patients; CR, n = 7 patients) in the Ravi cohort. G Proportion of patients achieving objective response (PR/CR) or progressive/stable disease (PD/SD) stratified by BMIS group in the Ravi cohort. Two-sided Fisher’s exact test P-value is shown. H Progression-free survival curves (left) and response rates (right) stratified by BMIS in the Kim cohort (n = 27 NSCLC patients). Two-sided Fisher’s exact test P value is shown. I Overall survival curves (left) and response rates (right) stratified by BMIS in the mUC cohort (n = 298 mUC patients). Tick marks indicate censored observations. Two-sided Fisher’s exact test P-value is shown. J, K Receiver operating characteristic (ROC) curves (left) and forest plots (right) comparing the performance of BMIS with established immunotherapy biomarkers, including tumor mutational burden (TMB), gene expression profile score (GEP), and cytolytic activity score (CYT), in the Ravi (n = 122 NSCLC patients) (J) and mUC (n = 298 mUC patients). Area under the curve (AUC) with 95% confidence intervals are shown. In survival analysis, hazard ratios (HRs) with 95% confidence intervals (CIs) and P-values from two-sided log-rank tests are shown. Survival curves represent Kaplan–Meier estimates; +, censored observations. In box plots (F), center line represents the median, box bounds indicate the 25th and 75th percentiles (IQR), whiskers extend to 1.5× IQR from box bounds, and outliers are shown as individual points.
Given the association between brain metastasis-enriched cell populations and immunotherapy resistance, we developed a quantitative prognostic model by extracting differentially expressed genes from cell populations associated with poor outcomes (HSP70-High CD4, HSP70-High CD8, TAM_PLTP). Using Variable Selection Using Random Forests (VSURF) for feature selection (Fig. 8D), we identified a robust 7-gene signature (PLK3, VAMP8, CKS2, TSPYL4, CDKN1A, LAMTOR1, IFNG). We then trained a logistic regression model to predict therapeutic response (CR/PR vs PD/SD) and used the model-derived predicted probabilities as continuous BMIS scores (Figure S10D–F).
We first evaluated BMIS performance in the discovery cohort (Ravi, n = 122). Using a prespecified cutoff (BMIS = 0.5; the standard decision boundary for predicted probabilities), patients with high BMIS scores exhibited significantly shorter PFS (log-rank p < 0.0001) and OS (log-rank p = 0.0019) (Fig. 8E). Multivariate Cox regression analysis, adjusting for age, sex, and initial tumor stage, confirmed that BMIS served as an independent prognostic factor for both PFS (p < 0.001) and OS (p = 0.016) (Figure S10G, H). Furthermore, BMIS scores were significantly elevated in patients with no therapeutic benefit (PD/SD) compared to responders (PR/CR) (Fig. 8F). Correspondingly, the objective response rate (PR/CR) in the high BMIS group was only 27.2%, substantially lower than 65.9% in the low BMIS group (Fisher’s exact test p = 7.03 × 10⁻⁵) (Fig. 8G).
To assess the generalizability of BMIS, we performed validation in three independent cohorts. In the Kim cohort (n = 27 NSCLC patients), the high BMIS group showed a consistent trend toward shorter PFS (log-rank p = 0.1), though this did not reach statistical significance, likely due to limited sample size (Fig. 8H). In the larger metastatic urothelial carcinoma (mUC) cohort (n = 298), BMIS demonstrated robust prognostic value, with high BMIS scores significantly associated with worse OS (log-rank p < 0.0001) and substantially lower objective response rates (14.4% vs 38.8%, Fisher’s exact test p < 0.0001) (Fig. 8I). However, in the melanoma cohort (n = 121), we did not observe significant associations between BMIS and either PFS or OS (PFS: p = 0.28; OS: p = 0.14) (Figure S10I–K). This suggests that the NSCLC brain metastasis-derived signature may not be directly applicable to melanoma, likely reflecting fundamental differences in tumor biology and immune microenvironments between cancer types.
Finally, we compared the predictive performance of BMIS with existing biomarkers, including tumor mutational burden (TMB), a widely used biomarker for immunotherapy response. Receiver operating characteristic (ROC) analysis in the Ravi cohort showed that BMIS (AUC = 0.773) exhibited comparable predictive power to TMB (AUC = 0.715). Importantly, a combined model integrating BMIS and TMB achieved superior performance (AUC = 0.805) (Fig. 8J). This complementary predictive value was confirmed in the mUC cohort, where the combined BMIS + TMB model (AUC = 0.764) outperformed either BMIS (AUC = 0.682) or TMB (AUC = 0.730) alone (Fig. 8K). These findings indicate that BMIS may captures immune microenvironment features distinct from tumor mutational landscape and that their integration enhances prediction of immunotherapy outcomes.
Discussion
Patients with BMs caused by NSCLC experience significantly reduced efficacy of ICIs. However, the mechanisms underlying this resistance remain unclear. Although previous bulk RNA-seq and immunohistochemical studies have identified a more immunosuppressive environment in BMs than in PTs, they lack the resolution to uncover the specific cellular contributions to this phenotype. More recent scRNA-seq efforts in metastatic NSCLC have described immune heterogeneity but have not consistently linked specific immune states to therapeutic resistance. Here, we employ a discovery-to-validation framework: we first characterize the immune landscape of brain metastases at single cell resolution to identify cell states enriched in this therapeutically resistant niche, and then test whether analogous BM-enriched programs, when present in primary tumors, are associated with ICI outcomes in independent cohorts. This strategy highlights immunosuppressive subpopulations, including HSP70-high and cycling T cells, TIMP1⁺ monocytes and PLTP⁺ TAMs, together with a loss of key memory and antigen-presenting populations, as major contributors to immune evasion in BMs. Collectively, these findings provide hypotheses regarding mechanisms of ICI resistance and nominate candidate biomarkers and therapeutic avenues for improving patient outcomes.
The expression of the HSP70-encoding genes HSPA1A and HSPA1B was elevated in both CD4⁺ and CD8⁺ T cells from BM samples. This enrichment was reproduced in an independent stage-matched cohort and confirmed by multiplex immunofluorescence. Consistent with a broadly stressed immune microenvironment in BMs, increased HSP70 expression was also observed in other immune cell lineages; however, given the central role of T cells in immune checkpoint blockade, we focused our mechanistic and clinical analyses on the T-cell compartment. Although HSP70 is associated with T-cell activation and cytotoxicity in the tumor microenvironment, its role in T cells remains incompletely defined33,34,35. In the present study, the elevated HSP70 expression in BM-derived T cells correlated with poor prognosis in patients receiving immunotherapy, suggesting that HSP70-High T cells contribute to ICI resistance. Further analysis revealed upregulation of exhaustion and interferon response pathways in HSP70⁺ T cells, implying a more complex biological mechanism underlying this phenotype. We predicted ligand-receptor interactions to identify potential drivers of the upregulation of HSP70 in T cells; we identified PTGS2 (COX-2) as a potential mediator. Given the established role of PTGS2 in BM formation36,37 and recent evidence that its downstream product PGE2 suppresses T-cell stemness38,39, our data raise the possibility that the COX-2–PGE2 axis promotes the accumulation of stressed, dysfunctional T cells in BMs and thereby contributes to ICI resistance, a hypothesis that warrants further experimental testing.
Beyond HSP70-high T cells, our data highlight a reorganization of the CD4⁺ and CD8⁺ T-cell compartments along a memory–exhaustion continuum in BMs. Central memory-like CD4_IL7R T cells and tissue-resident memory CD8_ZNF683 T cells were enriched in PTs and associated with improved PFS in patients treated with ICIs, independent of EGFR mutation status and disease stage. Memory T cells are characterized by their self-renewal capacity, longevity, and ability to rapidly differentiate into effector cells upon antigen re-encounter40,41. Recent studies have shown that CD4+ T cells with memory or stem-like phenotypes exhibit superior persistence and therapeutic efficacy compared to terminally differentiated effector cells in adoptive cell transfer and checkpoint blockade settings42,43. We also identified a progenitor-exhausted CD8_SPRY1 population, similar to previously described stem-like T cells, that was linked to favorable responses to PD-1 blockade27,44. In contrast, BMs harbored increased proportions of cycling CD8⁺ T cells, a terminally differentiated proliferative population with elevated stress and metabolic activity. Although such dysfunctional proliferative CD8⁺ T cells can still recognize tumors in some settings45,46, in our study cycling and HSP70-high T cells were negatively associated with immunotherapy responses, indicating a state of stressed proliferation that impairs effective antitumor immunity. Together, these findings suggest that the loss of memory and progenitor-exhausted T-cell pools, coupled with the expansion of stressed proliferating T cells, is a central feature of the BM immune microenvironment.
Among the DCs, cDC2-like cells (DC_CLEC10A) were significantly enriched in PTs and correlated with a better prognosis in patients undergoing immunotherapy. Consistent with their role as MHC class II antigen-presenting cells, loss of DC_CLEC10A may contribute to the immunosuppressive environment in BMs47,48. We identified two TAM subclusters: TAM_MARCO and TAM_PLTP. These subclusters were specifically enriched in PTs and BMs, respectively. Although MARCO is often considered an M2-TAM marker that is negatively associated with immunotherapy efficacy49,50,51, our study revealed an association between TAM_MARCO, interferon-gamma signaling, and improved prognosis in patients undergoing immunotherapy, suggesting functional heterogeneity among MARCO⁺ macrophages in NSCLC. Conversely, TAM_PLTP was negatively associated with patient prognosis and linked to increased lipid metabolism and reduced inflammatory responses30,52.
We developed the Brain Metastasis-derived Immune Signature (BMIS), an immune-derived gene signature that is associated with ICI efficacy, to translate our findings into clinical practice. Prognostic models based on scRNA-seq datasets have shown great potential for patient stratification53,54,55, and our scoring scheme integrates the transcriptional programs of HSP70-high CD4⁺ and CD8⁺ T cells and TAM_PLTP macrophages to enhance predictive accuracy. The BMIS model integrates genes with opposing prognostic associations, with negative coefficients (IFNG, VAMP8, CKS2, TSPYL4) indicating better outcomes when highly expressed and positive coefficients (PLK3, CDKN1A, LAMTOR1) associated with worse survival. The former encodes proteins involved in interferon-γ signaling (IFNG), cytotoxic vesicle trafficking (VAMP8) and cell-cycle control (CKS2, TSPYL4), whereas the latter participate in stress-activated kinase signaling (PLK3), cyclin-dependent kinase inhibition (CDKN1A) and lysosomal–mTORC1–dependent metabolic reprogramming (LAMTOR1). This bidirectional pattern is consistent with the idea that BMIS is associated with interactions between protective effector programs and detrimental stress responses in a microenvironment shaped by chronic antigenic stimulation, metabolic stress, and impaired effector function. BMIS demonstrated prognostic value across several independent immunotherapy cohorts and improved risk stratification when combined with tumor mutational burden, indicating that it captures aspects of the immune microenvironment that are complementary to existing biomarkers. The strength and direction of association varied between tumor types, suggesting that BMIS may be most informative in cancers that share immune microenvironmental features with NSCLC, and underscoring the need for prospective, histology-specific validation before clinical implementation.
Despite its comprehensive scope, our study has several limitations. The discovery scRNA-seq cohort is modest in size, lacks paired PT–BM specimens, and integrates in-house with public datasets, which may limit generalizability despite cross-cohort consistency and pseudo-bulk analyses. scRNA-seq also lacks spatial context, precluding localization of key cell states within BMs. In addition, our dataset does not include BM samples collected from patients who had received immune checkpoint inhibitors, which limits the assessment of treatment-induced immune remodeling. Although we applied Harmony-based integration, differences in tissue processing (e.g., dissociation and enrichment protocols) may introduce residual batch or protocol effects. Finally, neutrophils are underrepresented in 10x Genomics data due to low RNA content and fragility, preventing detailed characterization of this compartment. Future studies combining spatial transcriptomics and larger, ideally paired and ICI-treated BM cohorts, as well as protocols optimized for granulocytes, will be needed to refine and extend these findings.
In conclusion, our study provides a comprehensive overview of the immune microenvironment in primary and brain metastatic NSCLC at the single-cell level. We delineated the distinct molecular profiles and relative abundances of various cell clusters at these sites to identify potential therapeutic targets. Finally, we introduce BMIS as a brain metastasis–derived immune signature that may help refine immunotherapy stratification.
Methods
Patient enrolment and sample collection
This study was approved by the Ethical Committee of Shandong Cancer Hospital and Institute (SDTHEC2024006025) and complies with all relevant ethical regulations. We enrolled 12 patients (7 males and 5 females; median age 56.5 years, range 41–70) with pathologically confirmed NSCLC for scRNA-seq. Of these patients, eight and four had PTs and BMs, respectively. All patients underwent surgical resection at our institution prior to any systemic therapy. Informed consent was obtained from all participants. To augment our cohort of brain metastatic samples, we incorporated seven additional treatment-naïve specimens—curated from the GSE131907 dataset by Kim et al., based on cell count ( > 2000), resulting in eleven BM samples (Supplementary Data 1). Furthermore, we retrospectively collected 34 PT and 26 BM samples from treatment-naïve patients who were pathologically diagnosed with NSCLC and underwent surgery or diagnostic biopsy at Shandong Cancer Hospital and Institute between 2021 and 2023, prior to receiving any systemic therapy (Supplementary Data 5). Detailed patient characteristics, including sex, age, histology, and mutational status, are provided in Supplementary Data 1, 5.
Sample processing and cell sorting
Fresh tumor tissues were placed in MACS Tissue Storage Solution (130-100-008, Miltenyi Biotec) and stored on ice immediately after surgical resection. The samples were washed with phosphate-buffered saline (PBS), minced into fragments smaller than 1 mm3, and digested in 5 mL of a buffer containing collagenase IV (2 mg/mL; C4-22, Sigma-Aldrich) and deoxyribonuclease I (1 mg/mL; DN25, Sigma-Aldrich) at 37 °C for 30 min, as described previously54,56. The resulting cell suspension was mixed with 5 mL PBS supplemented with 2% fetal bovine serum (FBS) and centrifuged at 300 × g for 5 min at 4 °C to pellet the cells. Red blood cells were lysed using a red blood cell lysis buffer (420301, BioLegend) for 10 min at room temperature, followed by centrifugation at 300 × g for 5 min at 4 °C. The remaining cells were stained with antibodies against CD45 (clone HI30, 304031, Biolegend) for 30 min at 4 °C. Next, 7-aminoactinomycin D (7-AAD; 00-6993-50, eBioscience) was added to exclude the dead cells. Live immune cells (CD45⁺)—identified as 7-AAD-negative—were sorted using a BD FACSAria II cell sorter (BD Biosciences) and collected for single-cell RNA-seq. The sequential gating strategy is shown in Figure S11.
Single-cell RNA-seq and data pre-processing
Single-cell transcriptomes were amplified using a Chromium Single Cell 3’ Library Kit v3 (PN-1000075, 10x Genomics) according to the manufacturer’s instructions. The sorted cells were loaded onto a microfluidic chip to generate cDNA libraries. Sequencing was performed using an Illumina NovaSeq 6000 system (Illumina Inc.) across six lanes. Raw sequencing data were processed with the Cell Ranger software (version 4) using default parameters. The reads were aligned to the GRCh38 human reference genome, and we generated gene expression matrices using the STAR aligner.
To augment our brain metastasis cohort, we integrated seven treatment-naïve BM samples from the GSE131907 dataset, which were generated using compatible 10x Chromium technology (Single Cell 3′ v2) and showed comparable sequencing depth (median UMIs per cell 3500–6000 vs. our in-house median of 5,428). Raw gene expression matrices from GSE131907 were obtained and processed together with our in-house data using the same downstream quality-control and normalization pipeline. BM samples from GSE131907 with >2,000 cells after quality control were included (n = 7; Supplementary Data 1).
Data preprocessing was conducted using the Seurat R package (version 3.1.4). Genes expressed in at least 10 cells per sample were analyzed. Low-quality cells were excluded based on the following criteria: fewer than 2,000 UMIs, more than 6000 or fewer than 500 detected genes, or > 20% mitochondrial gene content. Potential doublets were identified and removed using the DoubletFinder (v2.0.3) package in R. Gene expression was normalized using Seurat’s LogNormalize function (scale factor = 10,000), then scaled using ScaleData while regressing out total UMI count and mitochondrial gene percentage. The expression matrices were integrated using the Harmony (v0.1.0) R package with dataset origin as the batch covariate to correct for technical variation between in-house and GSE131907 samples. Batch correction was validated by confirming intermixed cell populations from both datasets within each major cell lineage on UMAP embeddings and consistent expression of canonical marker genes across batches (Figure S1D). Highly variable genes (n = 2000) were identified using Seurat’s FindVariableFeatures function (selection.method = ‘vst’) and used for principal component analysis (PCA). The top 30 principal components computed from the integrated data were used as input for Harmony to correct for batch effects. The resulting Harmony embeddings were then used for Uniform Manifold Approximation and Projection (UMAP) to reduce dimensionality and visualize gene expression patterns.
Integration of the stage-matched validation cohort
To address potential confounding by disease stage, we assembled an independent validation cohort from publicly available scRNA-seq datasets of treatment-naïve stage III–IV NSCLC primary tumors (GSE131907, GSE148071, GSE179994) and brain metastases (HRA006468). Raw count matrices were processed using the same quality control pipeline as the discovery cohort, retaining only samples with more than 800 high-quality cells after filtration.
Because brain metastasis samples were derived from a single dataset (HRA006468), which could confound batch effects with biological differences, we included brain metastasis samples from GSE131907 during batch correction to serve as a “bridge” between tissue types. These GSE131907 brain metastasis samples were subsequently excluded from downstream analyses, as they had been used in the discovery cohort.
Following initial integration of all cell types, T cells were identified based on canonical marker expression (CD3D, CD3E, and CD2) and subset for lineage-specific analysis. T cells from GSE131907 (both primary tumor and brain metastasis samples) were merged with T cells from stage-matched validation datasets (GSE148071, GSE179994, HRA006468) using Seurat’s merge function. The merged dataset was normalized using LogNormalize (scale factor = 10,000), followed by identification of 3000 highly variable genes and scaling with ScaleData. Principal component analysis was performed on 50 PCs, and Harmony integration was applied to the top 30 PCs using dataset origin as the batch covariate (max.iter.harmony = 20). UMAP dimensionality reduction and clustering were performed on Harmony-corrected embeddings.
Cell type annotation
We identified cellular subclusters using the “FindAllMarkers” function in the Seurat R package with default parameters. Cell types and subtypes were annotated based on the expression of established canonical marker genes. Subclusters exhibiting similar gene expression profiles were grouped under the same cell type.
T cell stratification based on HSP70 expression
To stratify T cells based on HSP70 expression levels, we calculated a composite HSP70 module score for each cell using HSPA1A and HSPA1B as a gene set with Seurat’s AddModuleScore function. CD4+ and CD8+ T cells were then classified into three groups based on quartile distribution of the module scores: HSP70-High (top 25%), HSP70-Medium (middle 50%), and HSP70-Low (bottom 25%). This quartile-based classification was consistently applied across all datasets (Discovery cohort, and stage-matched validation cohort) to ensure reproducibility. All downstream analyses, including gene signature scoring, pathway enrichment, and survival analyses, were performed using this classification scheme.
Compositional analysis of cell type abundances
We compared cell type abundances between PT and BM samples using Wilcoxon rank-sum test for initial assessment. To account for the compositional nature of single-cell data and variable cell numbers across samples, the results were further validated using scCODA (v0.1.9), a Bayesian framework specifically designed for compositional data analysis. The CompositionalAnalysis model was applied separately for each major cell lineage (T/NK cells, myeloid cells, B/plasma cells) with tissue origin (PT vs. BM) as covariate, automatic reference cell type selection, and Hamiltonian Monte Carlo sampling (4 chains, 6,000 iterations including 3000 warmup). Cell types were considered differentially abundant only if they showed concordant significance in both analyses (Wilcoxon P < 0.05 and scCODA 95% highest density interval excluding zero).
To assess whether EGFR mutation status, classified as Major (19DEL or L858R), wild-type, or uncommon mutations, influenced site-associated differences, we performed Firth’s penalized logistic regression using the logistf R package (v1.26.1) on sample-level cell proportions, comparing unadjusted and EGFR-adjusted models. EGFR was considered a potential confounder if odds ratios changed by more than 10% after adjustment.
Pseudotime trajectory and cellular dynamics inference
We conducted pseudo-time analysis using Monocle 2 (version 2.14.0) to explore the developmental trajectories of T and myeloid cells57. Raw UMI counts for the cell subsets of interest were modeled with the “negbinomial.size” distribution. Standard normalization and variance estimation were performed using the “estimateSizeFactors” and “estimateDispersions” functions. Ordering genes were defined as the top 50 differentially expressed markers between the clusters of interest. The trajectory was constructed using the ‘DDRTree’ algorithm, and visualized with the “plot_cell_trajectory” function.
To validate the inferred directionality, the deep learning tool scTour (v0.1.3) was applied to infer cellular dynamics58. The scTour model was trained with default parameters to infer the differentiation trajectory based on raw UMI counts from cell clusters of interest. The resulting vector field was visualized by plotting its streamlines on scTour’s latent space-based UMAP embedding.
Pathway analysis
Differentially expressed genes with an absolute log fold change > 0.5 and an adjusted P-value < 0.05 were selected for Gene Ontology (GO) enrichment analysis. We employed the “compareCluster” function in the clusterProfiler (v4.2.2) R package to identify enriched GO terms across different subclusters59,60. Detailed results of GO enrichment were listed in Supplementary Data 16. To assess the functional status of T cells, we calculated gene signature scores using curated gene sets from published studies23 via the “AddModuleScore” function in Seurat.
Survival analysis
Bulk RNA-seq data and clinical information from patients in the TCGA-LUAD cohort were obtained from the UCSC Xena database (https://xenabrowser.net/datapages/) as previously described in ref. 61. We curated transcriptomic and clinical data from several published immunotherapy cohorts to validate our findings. Specifically, the Ravi cohort included 122 advanced NSCLC patients treated with ICIs with detailed survival and treatment response data, sourced from Ravi et al.24. The Kim cohort included 27 NSCLC patients treated with ICIs; transcriptomic data were obtained from GEO database (accession number GSE135222), and the corresponding survival data were retrieved from supplementary information of original publication62. The mUC cohort consisted of 298 metastatic urothelial carcinoma patients with detailed survival and treatment response data, curated from the IMvigor210CoreBiologies R package63. The Melanoma cohort comprised 121 patients, and their transcriptome and survival data were retrieved from the study by Liu et al.21. To estimate the abundance of different sub-clusters in the bulk RNA-seq data, we used the CIBERSORTx (https://cibersortx.stanford.edu/) algorithm64 for myeloid cells, employing the top 50 differentially expressed genes of each sub-cluster. For T cells, we applied Gene Set Variation Analysis (GSVA R package, v1.40.1) to calculate the combined expression values of cell type–specific signatures (the top 20 differentially expressed genes) because CIBERSORTx yielded markedly low proportions of sub-clusters. Optimal cutoffs for high and low expression were determined using the “surv_cutpoint” function in the survminer (v0.4.8) R package. Kaplan–Meier survival curves were generated using the Survminer package. Hazard ratios with 95% confidence intervals were calculated using Cox proportional hazards models. Univariate Cox models were applied for cell population survival analyses. For BMIS validation, multivariate Cox models adjusting for age, sex, and initial tumor stage were used to assess independent prognostic value.
Pseudo-bulk Differential Expression Analysis
For differential expression analysis between monocyte and TAM subpopulations, we employed a pseudobulk approach to properly account for patient-to-patient variability. Raw UMI counts from all cells within each cluster were aggregated per patient using Seurat’s AggregateExpression function, treating each patient as an independent biological replicate. Differential expression testing was performed using the edgeR package (v3.40.2). Genes with counts per million (CPM) > 1 in at least 3 samples were retained. Library size normalization was performed using the trimmed mean of M-values (TMM) method. Quasi-likelihood F-tests were used to identify differentially expressed genes, with FDR < 0.05 considered statistically significant.
NicheNet analysis
We used NicheNet analysis to identify potential ligand–receptor interactions that may drive the distinct immune phenotypes observed in primary and metastatic tumors65. Focusing on HSP70-positive T cells, which showed differential abundance between the two sites, we quantified the differentially expressed genes between HSP70-positive and HSP70-negative T cells. Genes with a log2 fold change > 0.2 and an adjusted P-value < 0.05 were considered significant. Using all genes expressed in HSP70-positive T cells as the background, we constructed ligand–receptor interactions based on the expressed receptors in these cells. Ligand activity was predicted using the “predict_ligand_activity” function of NicheNet (v1.0.0) R package.
Cell–cell communication analysis
Cell–cell interactions between PT and BM were inferred using CellChat v1.6.166. Immune subsets with ≥ 100 cells in both PT and BM were analyzed using normalized RNA assay data and the CellChatDB.human database. Communication probabilities were computed with the triMean method and filtered for interactions supported by ≥ 10 cells. PT and BM CellChat objects were then compared to identify differential signaling pathways and interaction strengths between conditions.
Construction of the Brain Metastasis-derived Immune Signature (BMIS)
BMIS was developed using the Ravi NSCLC immunotherapy cohort (n = 122) as the discovery dataset. We first identified differentially expressed genes (log₂ fold change > 0.5) from cell populations enriched in brain metastases and associated with poor outcomes: HSP70-High CD4⁺ T cells, HSP70-High CD8⁺ T cells, and TAM_PLTP macrophages. For each patient with documented immunotherapy response, we extracted normalized bulk RNA-seq expression values for these candidate genes. Patients were classified as responders (partial or complete response) or non-responders (progressive or stable disease) based on RECIST criteria. Variable selection was performed using the VSURF (v1.1.0) R package (ntree = 10,000), which applies random forest-based variable importance to identify genes discriminating responders from non-responders as previously described67,68. The seven genes retained at the interpretation step (PLK3, VAMP8, CKS2, TSPYL4, CDKN1A, LAMTOR1, IFNG) were entered into a logistic regression model with response status as the outcome. For each sample, the predicted probability of non-response from this model was used as the continuous BMIS score, with higher values indicating lack of clinical benefit. A fixed cutoff of 0.5 was used to stratify patients into high-BMIS and low-BMIS groups across all cohorts, ensuring consistent classification criteria. In validation cohorts, BMIS scores were computed by applying the trained model coefficients to expression data for the seven signature genes.
Calculation of established immunotherapy biomarkers
For comparison with BMIS, we calculated tumor mutational burden (TMB), gene expression profile (GEP) score, and cytolytic activity (CYT) score in the Ravi and mUC cohorts. TMB was defined as the total number of nonsynonymous mutations per megabase. The T cell-inflamed GEP score was calculated as the weighted mean expression of 18 interferon-γ-responsive genes related to antigen presentation, chemokine expression, cytotoxic activity, and adaptive immune resistance, as previously described in refs. 69,70. The CYT score was computed as the geometric mean of granzyme A (GZMA) and perforin 1 (PRF1) expression levels71. All scores were standardized within each cohort for comparative analyses.
Multiplex immunofluorescence staining
Multiplex immunofluorescence staining was performed on formalin-fixed paraffin-embedded tissue sections from 60 samples (34 primary tumors and 26 brain metastases) using Opal fluorophores (Akoya Biosciences; Opal 520: FP1487001KT, Opal 620: FP1495001KT, Opal 690: FP1497001KT). Three antibody panels were employed: (1) anti-CD4 (clone EPR6855, ab133616, Abcam; 1:300)/Opal690, anti-CD8 (clone EPR21769, ab217344, Abcam; 1:500)/Opal520, anti-HSP70 (clone EPR16892, ab181606, Abcam; 1:1000)/Opal620; (2) anti-CD14 (clone 1H5D8, ab181470, Abcam; 1:1000)/Opal620, anti-FCN1 (polyclonal, ab223712, Abcam; 1:400)/Opal690, anti-TIMP1 (clone EPR18352, ab211926, Abcam; 1:1000)/Opal520; (3) anti-CD68 (clone EPR20545, ab213363, Abcam; 1:500)/Opal690, anti-PLTP (clone EPR24448-21, ab282456, Abcam; 1:500)/Opal620. All slides within each panel were processed in a single batch following antibody validation and protocol optimization. Slides were counterstained with spectral DAPI (FP1490, Akoya Biosciences) and mounted with ProLong Gold antifade mountant (P36930, Thermo Fisher). Detailed antibody information, including clone names, catalogue numbers, and dilutions, is provided in Supplementary Data 17.
Whole-slide imaging was performed using automated multispectral imaging at 200× magnification (Vectra Polaris, Akoya Biosciences). Spectral unmixing was conducted using inForm software (version 2.4, Akoya Biosciences) with spectral libraries generated from single-stain control slides for each fluorophore. Quantitative image analysis was performed using QuPath version 0.4.3 (Queen’s University)72. Tumor regions were delineated using an automated tissue classifier trained on pathologist-annotated examples and verified by a pathologist blinded to clinical information. Non-tumor areas, including normal lung parenchyma, necrosis, hemorrhage, and tissue artifacts, were excluded from analysis. Within tumor-annotated regions, cell segmentation was performed using DAPI-based nuclear detection. Detection parameters were optimized for immune cells (expected cell diameter 10–20 μm, minimum cell area 50 μm²), with cell boundaries extended 2–3 μm beyond nuclei to capture cytoplasmic marker expression. Marker-positive cells were identified using intensity thresholds established from positive and negative control regions within each slide. The proportion of marker-positive cells was calculated as (number of marker-positive cells / total tumor cells) × 100%. The accuracy of the resulting cell segmentation and phenotyping was further confirmed by qualitative visual inspection by a pathologist on five randomly selected slides per panel to ensure appropriate cell identification and classification. A median of 597,132 tumor cells per sample (interquartile range: 120,818-910,273) were analyzed.
Statistical analysis
All statistical analyses were performed in R (version 4.2.0). For differential expression analyses, cell type marker genes were identified using Seurat’s FindAllMarkers function with Bonferroni correction, while pseudobulk differential expression was performed using edgeR with Benjamini-Hochberg FDR correction. Gene set enrichment analyses were conducted using the Benjamini-Hochberg method for FDR control. Statistical comparisons of immune cell frequencies between PTs and BMs in mIF staining were performed using two-sided Wilcoxon rank-sum tests. P-values < 0.05 were considered statistically significant. Survival analyses were performed using Kaplan-Meier curves with log-rank tests. All statistical tests were two-sided unless otherwise mentioned.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw scRNA-seq data generated in this study have been deposited in the Genome Sequence Archive for Human (GSA-Human) under accession code HRA011002. The processed data matrices have been deposited in the OMIX database of the National Genomics Data Center under accession code OMIX015258. The publicly available datasets used in this study are available under accession codes GSE131907, GSE135222, GSE148071, GSE179994, and HRA006468. The TCGA-LUAD data are available at https://xenabrowser.net/datapages/. The IMvigor210 cohort data are available through the IMvigor210CoreBiologies R package. All other data supporting the findings of this study are provided in the Supplementary Information and Source Data files or from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. Ca. Cancer J. Clin. 72, 7–33 (2022).
Peters, S., Bexelius, C., Munk, V. & Leighl, N. The impact of brain metastasis on quality of life, resource utilization and survival in patients with non-small-cell lung cancer. Cancer Treat. Rev. 45, 139–162 (2016).
Cagney, D. N. et al. Incidence and prognosis of patients with brain metastases at diagnosis of systemic malignancy: A population-based study. Neuro. Oncol. 19, 1511–1521 (2017).
Reck, M. et al. Pembrolizumab versus Chemotherapy for PD-L1–Positive Non–Small-Cell Lung Cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Mansfield, A. S. et al. Outcomes with pembrolizumab (pembro) monotherapy in patients (pts) with PD-L1–positive NSCLC with brain metastases: Pooled analysis of KEYNOTE-001, −010, −024, and −042. Ann. Oncol. 30, v604–v606 (2019).
Lau, S. C. M. et al. Durability of CNS disease control in NSCLC patients with brain metastases treated with immune checkpoint inhibitors plus cranial radiotherapy. Lung Cancer 156, 76–81 (2021).
Goldberg, S. B. et al. Pembrolizumab for management of patients with NSCLC and brain metastases: long-term results and biomarker analysis from a non-randomised, open-label, phase 2 trial. Lancet Oncol. 21, 655–663 (2020).
Zhou, S. et al. Anti-PD-(L)1 immunotherapy for brain metastases in non-small cell lung cancer: Mechanisms, advances, and challenges. Cancer Lett. 502, 166–179 (2021).
Quail, D. F. & Joyce, J. A. The Microenvironmental Landscape of Brain Tumors. Cancer Cell 31, 326–341 (2017).
Berghoff, A. S. et al. Tumor-Infiltrating Lymphocytes (Tils) and Pd-L1 Expression in Non- Small Cell Lung Cancer Brain Metastases (Bm) and Matched Primary Tumors (Pt). Ann. Oncol. 25, iv465 (2014).
Kudo, Y. et al. Suppressed immune microenvironment and repertoire in brain metastases from patients with resected non-small-cell lung cancer. Ann. Oncol. 30, 1521–1530 (2019).
Song, S. G. et al. Comparative analysis of the tumor immune-microenvironment of primary and brain metastases of non-small-cell lung cancer reveals organ-specific and EGFR mutation-dependent unique immune landscape. Cancer Immunol. Immunother. 70, 2035–2048 (2021).
Jiang, T. et al. Characterization of evolution trajectory and immune profiling of brain metastasis in lung adenocarcinoma. npj Precis. Oncol. 5, 6 (2021).
Kim, N. et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat. Commun. 11, 2285 (2020).
Gonzalez, H. et al. Cellular architecture of human brain metastases. Cell 185, 729–745.e20 (2022).
Karimi, E. et al. Single-cell spatial immune landscapes of primary and metastatic brain tumours. Nature 614, 555–563 (2023).
Zou, Y. et al. The Single-Cell Landscape of Intratumoral Heterogeneity and The Immunosuppressive Microenvironment in Liver and Brain Metastases of Breast Cancer. Adv. Sci. 10, 2203699 (2023).
Song, Q. et al. Single-cell sequencing reveals the landscape of the human brain metastatic microenvironment. Commun. Biol. 6, 1–13 (2023).
Laughney, A. M. et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat. Med. 26, 259–269 (2020).
Xing, X. et al. Pan-cancer human brain metastases atlas at single-cell resolution. Cancer Cell 1–19 https://doi.org/10.1016/j.ccell.2025.03.025 (2025).
Wu, F. et al. Single-cell profiling of tumor heterogeneity and the microenvironment in advanced non-small cell lung cancer. Nat. Commun. 12, 2540 (2021).
Liu, B. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2021).
Chu, Y. et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Ravi, A. et al. Genomic and transcriptomic analysis of checkpoint blockade response in advanced non-small cell lung cancer. Nat. Genet. 55, 807–819 (2023).
Jung, H. et al. DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load. Nat. Commun. 10, 1–12 (2019).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Liu, Z. et al. Progenitor-like exhausted SPRY1+CD8+ T cells potentiate responsiveness to neoadjuvant PD-1 blockade in esophageal squamous cell carcinoma. Cancer Cell 41, 1852–1870.e9 (2023).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. J. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, 1–10 (2021).
Markiewski, M. M. et al. Modulation of the antitumor immune response by complement. Nat. Immunol. 9, 1225–1235 (2008).
Zhang, L. et al. Single-Cell Analyses Inform Mechanisms of Myeloid-Targeted Therapies in Colon Cancer. Cell 181, 442–459.e29 (2020).
Zhang, H. et al. Spatially resolved C1QC+ macrophage-CD4+ T cell niche in colorectal cancer microenvironment: implications for immunotherapy response. Cell Discov. 11, 60 (2025).
Aranda, C. J. et al. IgG memory B cells expressing IL4R and FCER2 are associated with atopic diseases. Allergy Eur. J. Allergy Clin. Immunol. 78, 752–766 (2023).
Millar, D. G. et al. Hsp70 promotes antigen-presenting cell function and converts T-cell tolerance to autoimmunity in vivo. Nat. Med. 9, 1469–1476 (2003).
Figueiredo, C. et al. Heat shock protein 70(HSP70) induces cytotoxicity of T-helper cells. Blood 113, 3008–3016 (2009).
Giri, B. et al. Hsp70 modulates immune response in pancreatic cancer through dendritic cells. Oncoimmunology 10, 1976952 (2021).
Allen, J. E. et al. COX-2 drives metastatic breast cells from brain lesions into the cerebrospinal fluid and systemic circulation. Cancer Res. 74, 2385–2390 (2014).
Bos, P. D. et al. Genes that mediate breast cancer metastasis to the brain. Nature 459, 1005–1009 (2009).
Lacher, S. B. et al. PGE2 limits effector expansion of tumour-infiltrating stem-like CD8+ T cells. Nature 629, 417–425 (2024).
Morotti, M. et al. PGE2 inhibits TIL expansion by disrupting IL-2 signalling and mitochondrial function. Nature 629, 426–434 (2024).
Speiser, D. E., Chijioke, O., Schaeuble, K. & Münz, C. CD4+ T cells in cancer. Nat. Cancer 4, 317–329 (2023).
Liu, Q., Sun, Z. & Chen, L. Memory T cells: strategies for optimizing tumor immunotherapy. Protein Cell 11, 549–564 (2020).
Borst, J., Ahrends, T., Bąbała, N., Melief, C. J. M. & Kastenmüller, W. CD4+ T cell help in cancer immunology and immunotherapy. Nat. Rev. Immunol. 18, 635–647 (2018).
Brightman, S. E. et al. Neoantigen-specific stem cell memory-like CD4+ T cells mediate CD8+ T cell-dependent immunotherapy of MHC class II-negative solid tumors. Nat. Immunol. 24, 1345–1357 (2023).
Galluzzi, L., Smith, K. N., Liston, A. & Garg, A. D. The diversity of CD8 + T cell dysfunction in cancer and viral infection. Nat. Rev. Immunol. 25, 662–679 (2025).
Guo, X. et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 24, 978–985 (2018).
Yang, Y. et al. Genomic and the tumor microenvironment heterogeneity in multifocal hepatocellular carcinoma. Hepatology 82, 582–598 (2025).
Cheng, S. et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell 184, 792–809.e23 (2021).
Saito, Y., Komori, S., Kotani, T., Murata, Y. & Matozaki, T. The Role of Type-2 Conventional Dendritic Cells in the Regulation of Tumor Immunity. Cancers (Basel) 14, 1976 (2022).
Azizi, E. et al. Single-Cell Map of Diverse Immune Phenotypes in the Breast Tumor Microenvironment. Cell 174, 1293–1308.e36 (2018).
Yang, Y. S. et al. Superenhancer drives a tumor-specific splicing variant of MARCO to promote triple-negative breast cancer progression. Proc. Natl. Acad. Sci. Usa. 119, 1–12 (2022).
Dong, Q. et al. International Immunopharmacology MARCO is a potential prognostic and immunotherapy biomarker. Int. Immunopharmacol. 116, 109783 (2023).
Vikstedt, R. et al. Macrophage phospholipid transfer protein contributes significantly to total plasma phospholipid transfer activity and its deficiency leads to diminished atherosclerotic lesion development. Arterioscler. Thromb. Vasc. Biol. 27, 578–586 (2007).
Bill, R. et al. CXCL9:SPP1 macrophage polarity identifies a network of cellular programs that control human cancers. Science. 381, 515–524 (2023).
Liu, C. et al. Single-cell RNA-sequencing reveals radiochemotherapy-induced innate immune activation and MHC-II upregulation in cervical cancer. Signal Transduct. Target. Ther. 8, 44 (2023).
Salcher, S. et al. High-resolution single-cell atlas reveals diversity and plasticity of tissue-resident neutrophils in non-small cell lung cancer. Cancer Cell 40, 1503–1520.e8 (2022).
Liu, C. et al. Single-cell dissection of cellular and molecular features underlying human cervical squamous cell carcinoma initiation and progression. Sci. Adv. 9, 1–20 (2023).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Li, Q. scTour: a deep learning architecture for robust inference and accurate prediction of cellular dynamics. Genome Biol. 24, 1–33 (2023).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: An R package for comparing biological themes among gene clusters. Omi. A J. Integr. Biol. 16, 284–287 (2012).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov 2, 100141 (2021).
Bai, M. et al. Integrated analysis of EGFR mutated non-small cell lung cancer reveals two distinct molecular subtypes. Clin. Transl. Med. 13, 1–7 (2023).
Kim, J. Y., Choi, J. K. & Jung, H. Genome-wide methylation patterns predict clinical benefit of immunotherapy in lung cancer. Clin. Epigenetics 12, 1–10 (2020).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1–20 (2021).
Chen, H. et al. Molecular characterization and clinical relevance of metabolic signature subtypes in gastric cancer. Cell Rep. 43, 114424 (2024).
Liu, Z. et al. A proteomic and phosphoproteomic landscape of KRAS mutant cancers identifies combination therapies. Mol. Cell 81, 4076–4090.e8 (2021).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Chen, H. et al. Deciphering the tumor microenvironment cell-infiltrating landscape reveals microenvironment subtypes and therapeutic potentials for nonsquamous NSCLC. JCI Insight 7, e152815 (2022).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 1–7 (2017).
Acknowledgements
We thank all patients enrolled in this study. We thank Professor Xiaolu Zhang from Shandong University for her guidance through data analysis. We thank Professor Nu Zhang from Sun Yat-sen University and his team for permission of access to dataset HRA006468. We thank Berry Genomics Co. Ltd. for sequencing support and Shandong PhenoScience Biotechnology Ltd. for assistance with multiplex immunofluorescence staining. We would like to thank Editage (www.editage.com) for English language editing. This study was supported by the National Natural Science Foundation of China (82473296 to F.T., 82403773 to C.L.), China Postdoctoral Science Foundation (2025M771937 to M.B., 2025T180600 to C.L., 2024M760140 to C.L.), Natural Science Foundation of Shandong Province (ZR2025QC1758 to M.B.), Beijing Science and Technology Innovation Medical Development Foundation (KC2023-JX-0288-PM32 to F.T.), and Young Elite Scientists Sponsorship Program of the Beijing High Innovation Plan (to C.L.).
Author information
Authors and Affiliations
Contributions
Conceptualization: F.T., M.B., J.Y., C.L. Methodology: M.B., T.Y., X.L., P.W., X.S., Y.L., B.Z., C.L., F.T., P.J., H.G., X.G., F.L. Investigation: M.B., T.L., X.L., X.G. F.L., P.J., C.L. Visualization: M.B., X.L., T.Y., P.J., C.L. Supervision: F.T., C.L., J.Y., X.S. Writing—original draft: M.B., T.Y., F.T., C.L. Writing—review & editing: J.Y., C.L., F.T., M.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hiroshi Kagamu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bai, M., Yin, T., Li, X. et al. Identification of altered immune landscape at single-cell resolution in NSCLC brain metastasis and its association with poor immune checkpoint inhibitor responses. Nat Commun 17, 2370 (2026). https://doi.org/10.1038/s41467-026-70715-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70715-6