Abstract
Entamoeba histolytica causes amoebiasis and damages intestinal epithelium, but how individual parasite factors coordinate parasite gene regulation with host remodeling is unclear. We investigate lysine- and glutamic acid-rich protein 2, a factor linked to the host brush border. Sequence analysis, imaging, and functional assays show that lysine- and glutamic acid-rich protein 2 accumulates in the parasite nucleus, binds AT-rich DNA, and modulates transcriptional programs related to Amoebiasis, including cysteine protease and sulfur metabolism. During parasite-epithelium contact, the protein enters host epithelial cells and is associated with increased deoxyribonucleic acid synthesis, altered cytoskeletal regulators, actin remodeling, and reduced barrier integrity. Here, we propose a working model in which lysine- and glutamic acid-rich protein 2 links parasite chromatin-associated regulation with host epithelial remodeling during contact. Notably, our data support host cytoskeletal and junctional phenotypes and do not yet establish a direct role for it in host chromatin regulation. Together, these observations suggest a potentially broader mechanism by which extracellular pathogens deploy effectors to optimize virulence and adapt to diverse host environments.
Similar content being viewed by others
Introduction
Pathogenic microbes have evolved sophisticated mechanisms to hijack host cellular functions, enabling colonization and persistence by remodeling host biology. Intracellular pathogens, such as Mycobacterium tuberculosis, Listeria monocytogenes, Plasmodium falciparum, and Toxoplasma gondii invade host cells to escape immune surveillance and establish specialized intracellular niches, subverting vesicular trafficking, immune signaling, and cytoskeletal dynamics within host cells1,2. In contrast, extracellular pathogens remain outside host cells but still engage in complex interactions with host tissues to reprogram membrane-proximal signaling and epithelial barrier function, thereby creating permissive niches3,4. Unlike intracellular microbes that directly access host machinery, extracellular pathogens must deploy surface- and secretome-mediated mechanisms to persist. For example, Trypanosoma brucei evades immune recognition via antigenic variation of variant surface glycoproteins (VSGs)5, and Giardia perturbs epithelial integrity through ventral disc-mediated attachment—a microtubule-based structure—and by disrupting tight-junction architecture, increasing paracellular permeability3. Consequently, extracellular protozoa provide tractable systems for dissecting such outside-in mechanisms.
Entamoeba histolytica is the etiologic agent of amoebiasis, a disease of significant global health concern that ranges from asymptomatic colonization to invasive colitis and extra-intestinal disease such as liver abscesses6. Within the intestine, E. histolytica induces severe inflammation, tissue perforation, and fulminant amoebic colitis, leading to potentially fatal outcomes7. Parasite survival at the mucosal surface depends on adhesion, signaling at the epithelial surface, and remodeling of junctional complexes. Tissue damage involves contact-dependent processes—notably trogocytosis8, a process in which the parasite nibbles portions of live host cell membranes—and secreted effectors that affect extracellular matrix and disrupt tight-junctional architecture9. Prominent among these are the Gal/GalNAc lectin (a key membrane-associated factor) and cysteine proteases, which compromise epithelial integrity10,11. Despite these mechanistic insights, how E. histolytica coordinates additional effector proteins with host epithelial responses during mucosal colonization remains incompletely understood12.
Recently, affinity-based isolation of E. histolytica membrane-biotinylated proteins that bind to Caco-2 brush-border components recovered the KERP (lysine- and glutamic acid-rich protein) family together with the Gal/GalNAc lectin intermediate subunit and an alpha-actinin-like protein, indicating association with brush-border components13. KERP1 was identified in membrane fractions, associates with the brush border of Caco-2 cells, and has been linked to liver abscess formation as a virulence factor14, whereas KERP2 is conserved across Entamoeba species, also associates with the brush border directly or indirectly, but remains mechanistically underdefined. Population genetics suggest that polymorphisms in the kerp2 locus associate with disease outcome, indicating potential clinical relevance and evolutionary pressure on this locus15. Interestingly, KERP2 contains a predicted nuclear localization signal (NLS) and shows DEK-like annotation in InterPro but otherwise lacks conventional annotated globular domains in standard profile searches16. Together with its recovery from membrane-enriched fractions, these observations frame an apparent conflict between nuclear targeting and membrane in which a protein with nuclear-targeting potential is detected in membrane compartments and at the host interface—raising the possibility of non-canonical trafficking between nuclear and plasma-membrane-proximal sites during host interaction.
Only a limited set of nuclear chromatin-associated proteins are known to traffic beyond the nucleus and even participate in cell-surface or extracellular functions. DEK, a conserved chromatin factor harboring a SAP domain that modulates DNA topology and chromatin accessibility, illustrates this duality17. DEK can be released to the extracellular milieu and signal in a cytokine-like fashion via CXCR2 with heparan sulfate proteoglycans as co-receptors, regulating hematopoietic stem/progenitor compartments and engaging MAPK/AKT pathways18. Although SAP domain was long thought to be restricted to higher eukaryotes, recent work identified a SAP-domain-containing protein in T. brucei (TbSAP) with a similar role in gene repression19. The DEK paradigm, a protein with chromatin-engagement capacity that can also access the cell surface/extracellular space, provides a conceptual framework for considering how KERP2, as a parasitic protein, might bridge parasite nuclear processes with host-interacting communication.
We therefore focus on the intestinal epithelial interface—the earliest host surface encountered by Entamoeba—to elucidate contact-dependent mechanisms using barrier-specific readouts under controlled conditions. Specifically, we aim to: (i) elucidate the intracellular trafficking of KERP2 and its role in regulating Entamoeba cellular functions; (ii) assess whether KERP2 contacts and is internalized by intestinal epithelial cells during trophozoites-enterocyte interaction; and (iii) identify host cellular processes most plausibly influenced by KERP2 at epithelial interface that are central to mucosal defense and colonization. Although innate immune cells are key determinants of lesion formation in vivo, here we deliberately use epithelial-only models as reductionist systems to define epithelial barrier-interface mechanisms in isolation from immune-cell-mediated effects.
In this work, we show that KERP2 localizes to the parasite nucleus, associates with DNA, and modulates transcriptional programs that include cysteine proteases and sulfur metabolism, and that KERP2 is detected in host epithelial cells in association with changes in DNA synthesis, cytoskeletal regulation, actin remodeling, and compromised epithelial barrier function, supporting a model in which a traceable parasite factor links chromatin regulation in the parasite to remodeling of host barrier integrity.
Results
In silico analysis of KERP2
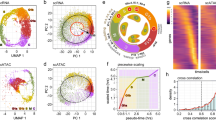
To investigate the evolutionary relationships among KERPs, we compared their primary structures and found that KERP2 (EHI_065630) shares 67.5% sequence identity across ~67% of its length with KERP3 (EHI_198680), whereas both are distinct from KERP1 (EHI_098210). Using KERP2 as a query, BLASTp and HMMER searches identified regional conservation with DEK-like proteins, primarily within residues 8-233 (HMMER E-value 1.60E-13; bit score 61.9) (Fig. S1a). To further examine the evolutionary history of KERP2, we performed maximum-likelihood phylogenetic reconstruction with IQ-TREE using homologous sequences from a broad range of eukaryotic taxa (Fig. 1a). The results revealed that KERP2 and KERP3 are restricted to Entamoeba species, forming a well-supported monophyletic group, suggesting lineage-specific conservation, whereas relationships to more distantly-related eukaryotes, including fungi, algae, plants, and metazoans, remain poorly supported (Fig. 1a). To assess selection pressure acting on KERP2 within Entamoeba, a codon-based dN/dS (ω) analysis yielded ω = 0.07979, consistent with strong purifying selection and functional constraint. Notably, a tBLASTn search within Amoebozoa identified short, low-scoring segments in Polysphondylium pallidum and Dictyostelium purpureum (Fig. S1b), suggesting limited, non-orthologous similarity outside Entamoeba.
a Maximum likelihood phylogenetic tree of KERP2 (EHI_065630) and homologous sequences across eukaryotic taxa, constructed using IQ-TREE and visualized with iTOL. The circular display represents bootstrap values with color gradients, while the rectangular display provides a simplified view for clarity, focusing on Entamoeba species. Bootstrap values are indicated at major branch points. b Multiple sequence alignment of the KERP2 SAP domain (amino acids 104–141) with representative SAP domains from other organisms, constructed as described in ref. 17. Conserved residues associated with DNA binding are highlighted by residue properties: hydrophobic (yellow), small (green), polar (purple), and bulky (gray). c Predicted secondary structures, shown schematically, feature two amphipathic helices characteristic of the SAP domain. Structural model of KERP2 generated by I-TASSER (white) showing conserved architecture resembling the N-terminal region of DEK proteins, including the embedded SAP domain (green; PDB ID: 2JX3). d Sequence analysis identifies a coiled-coil domain in the C-terminal region (amino acids 178–216) predicted by PCOILS and MARCOIL. Source data are provided as a Source Data file. e Schematic representation of the domain architecture of KERP2, highlighting the SAP domain (green), coiled-coil domain (purple), and nuclear localization signal (NLS, red line).
Given that KERP2 is restricted to Entamoeba and shares resemblance with DEK proteins, we examined its structural organization. Despite limited overall sequence conservation, manual alignment using SAP/SAF domain criteria identified a putative SAP module in KERP2 (residues 104–141) (Fig. 1b). Secondary-structure predictions using PHD identified two amphipathic helices (110–119 and 130–141), and I-TASSER modeling indicated structural similarity to the N-terminal SAP domain of DEK (PDB: 2JX3) (Fig. 1c). Unlike canonical DEK proteins, which carry a C-terminal DNA-binding domain, KERP2 lacks this feature (Fig. S1c). Instead, its C-terminus harbors a coiled-coil domain (a.a. 178–216), as predicted by Marcoil and PCOILS (Fig. 1d). KERP2 is highly basic (pI = 9.75) with a positively charged surface (Fig. S1d). An NLS spanning residues 192–223, located within the coiled-coil domain, was identified by NLS Mapper (score = 6.4), suggesting potential nuclear-cytosolic shuttling (Fig. 1e).
Intracellular localization of KERP2 in E. histolytica
To investigate the intracellular localization of KERP2 in E. histolytica and based on in silico predictions suggesting a basic, DEK-like protein with a C-terminal coiled-coil containing an NLS (Fig. 1e), we generated an amebic line expressing HA-tagged full-length KERP2 and a truncated variant, HA-KERP2∆185−239, which lacks the coiled-coil/NLS region (Fig. 2a). Immunofluorescence analysis (IFA) with line-intensity profiling revealed distinct localization patterns (Fig. 2b): wild-type HA-KERP2 predominantly localized to the nuclear periphery and nucleoplasm, with punctate signals in the cytosol, whereas HA-KERP2∆185−239 showed a marked reduction in nuclear signal and redistribution to the cytosol.
a Schematic representation of HA-KERP2 constructs used in this study: full-length HA-KERP2 and truncated HA-KERP2∆185−239 lacking the coiled-coil domain. b Immunofluorescence of E. histolytica trophozoites expressing HA-KERP2 variants. Cells were stained with anti-HA (green; Alexa 488), anti-cysteine synthase 1 (red; Alexa 594), and Hoechst 33342 (blue). CS1 is shown as a qualitative cytosolic marker in IFA. Single optical sections are displayed. White arrowheads indicate nuclear localization; white arrows highlight cytosolic puncta/vesicular structures; scale bar as indicated. Line-intensity profiles quantify nuclear enrichment versus cytosolic redistribution along the indicated lines. CS1 immunofluorescence is shown as a qualitative cytosolic marker and was not quantified. c Native EMSA with purified His-GFP-KERP2 and 5’-biotinylated DNA probes. KERP2 binds an AT-rich double-stranded DNA duplex (dsAT) and an AT-rich single-stranded DNA (ssAT) over 4–40 fmol protein, but not a GC-rich double-stranded DNA duplex (dsGC); unlabeled ssAT competes binding. A faster-migrating complex is observed on AT probes. His-GFP alone does not bind. (Positions of ssDNA and dsDNA are indicated at left.) Source data are provided as a Source Data file. d EMSA as in (c) with His-GFP–KERP2Δ185–239. The truncation retains AT-selective binding and the characteristic fast-migrating complex, indicating that DNA engagement resides in the N-terminal SAP-like region. Source data are provided as a Source Data file. e Size-exclusion chromatography (SEC) of His-GFP–KERP2. Coomassie-stained SDS–PAGE of fractions (F6–F21) shows a dominant peak in the monomer window with an earlier shoulder consistent with a minor higher-molecular-weight population. Source data are provided as a Source Data file. f Size-exclusion chromatography of His-GFP–KERP2Δ185–239. Elution shifts to later fractions and the early shoulder is lost, consistent with reduced self-association upon coiled-coil deletion. Source data are provided as a Source Data file.
To validate these observations, we performed live-cell imaging using GFP-HA-tagged constructs. The results were consistent with the IFA data, with GFP-HA-KERP2 being predominantly nuclear (Supplementary Movie 1), while GFP-HA-KERP2∆185−239 remained cytosolic (Supplementary Movie 2). Immuno-electron microscopy (EM) further demonstrated HA-KERP2 enrichment in electron-dense nuclear regions typically associated with chromatin-rich areas, and also detected HA-KERP2 in small electron-dense granule (EDG)-like puncta, larger vesicles, and diffusely distributed in the cytosol (Fig. S2a, b).
To complement the imaging data, we conducted subcellular fractionation to assess KERP2 distribution biochemically. Cellular material was separated into a 14,000 × g pellet (crude nuclear/organellar, including nuclei and heavy membranes) and supernatant (cytosol and small vesicles) (Fig. S2c-d). HA-KERP2 was enriched in the pellet fraction, consistent with nuclear and membrane association, whereas HA-KERP2∆185−239 was recovered predominantly in the supernatant, aligning with its cytosolic localization.
Domain characterization of KERP2 reveals AT-rich DNA binding and multimerization
Because KERP2 contains a predicted N-terminal SAP-like DNA-binding module and accumulates in the nucleus of the parasite, we investigated its DNA binding affinity using electrophoretic mobility shift assay (EMSA) with biotinylated probes. Recombinant His-GFP–KERP2 produced a dose-dependent mobility shift on an AT-rich DNA duplex and an AT-rich single-strand DNA, whereas it showed no binding to a GC-rich duplex at any concentration (Fig. 2c). Excess unlabeled AT-rich single-stranded DNA competed away the shifted complex, confirming sequence selectivity. The Δ185–239 truncation behaved similarly, producing robust shifts on AT-rich duplex and single-stranded AT-rich DNA, showing no binding to GC-rich duplex, and exhibiting full competition by unlabeled AT-rich DNA—indicating that AT-DNA recognition is preserved in the absence of the coiled-coil domain (Fig. 2d). His-GFP alone did not bind. Laddering of shifted bands on AT probes is consistent with multi-site occupancy. These data establish intrinsic, SAP-consistent, AT-selective DNA binding in vitro.
To assess sequence features of KERP2 binding in the cellular context, we performed ChIP–seq for GFP-HA–KERP2 and called peaks using the Model-based Analysis of ChIP-Seq 2 (MACS2) in narrow mode to capture summit-centric signals. The resulting peak-width distributions indicated punctate occupancy: in all three biological replicates, signals extended only a few hundred base pairs, accompanied by a long, low-intensity tail (Fig. S3a). Global ChIP/Input enrichment was moderate (3.89×, 1.86×, 1.75× across replicates), and, together with limited inter-replicate concordance, suggested that overall peak strength was weak (Supplementary Data 2). We therefore restricted downstream analyses to reproducible peaks defined by triple overlap. Genomic annotation of these reproducible peaks revealed a strong promoter-proximal bias (Fig. S3b). De novo motif discovery on this high-confidence set identified A/T-rich motifs that are significantly enriched relative to matched background controls (Fig. S3c). Background-aware motif logos showed short GC-anchored positions embedded within predominantly A/T-rich contexts, arguing against the presence of a single, high-information motif. Instead, these features are consistent with a model in which KERP2 functions as a chromatin co-regulator with weak, shape-biased sequence preferences rather than as a canonical transcription factor with strong sequence specificity.
Given the predicted C-terminal coiled-coil and reduced nuclear enrichment of Δ185–239, we asked whether this region supports self-association and contributes to nuclear entry. Using a calibrated size exclusion chromatography (SEC) column (Ve = 32.92–8.172 x log10(MWkDa), R2 = 0.976), the monomeric elution range corresponded to the fractions F18–F21 ( ~ 45–60 kDa), with F10 ( ~ 130 kDa), F16 ( ~ 58 kDa), and F17 ( ~ 51 kDa) serving as calibration references (Fig. S4a-b). His-GFP–KERP2 (58.2 kDa) eluted with a dominant peak at F16 with a minor earlier-eluting shoulder (F9–F14), indicating the presence of a small higher-molecular-weight population (Fig. 2e, Fig. S4c). By contrast, His-GFP–KERP2Δ185–239 (52 kDa) displayed a later-eluting peak at F17 and lacked the early-eluting shoulder, consistent with reduced self-association in vitro (Fig. 2f, Fig. S4d). In E. histolytica lysates resolved by SEC and probed with anti-HA antibody, HA-KERP2 eluted predominantly within the monomeric range in both cytosolic (F17–F19 with peak at F18–F19) and nuclear fractions (F18–F20 with peak at F19–F20), with no detectable early-eluting species in nuclear samples (Fig. S4e). HA-KERP2Δ185–239 eluted even later in both compartments (F19–F21, peak F20), again consistent with a predominantly monomeric population (Fig. S4f). Together, these data show that the C-terminal coiled-coil promotes self-association in the purified protein and supports a minor higher-molecular-weight cytosolic population in vivo. However, under our SEC conditions, the nuclear pool of both constructs is largely monomeric. Combined with the reduced nuclear enrichment of Δ185–239 observed by imaging and biochemical fractionation, these data indicate that the coiled-coil is required for efficient nuclear localization and contributes to self-association, whereas any stable higher-order assemblies are either low in abundance or not retained during SEC.
Gene silencing reveals KERP2 as a suppressor of virulence functions
To investigate the functional role of KERP2, we generated a KERP2-knockdown strain (psAP-KERP2gs), using small interfering RNAs expressed from the psAP-2-Gunma plasmid. Quantitative reverse transcription PCR (qRT-PCR) confirmed a near-complete reduction of KERP2 transcript levels (Fig. 3a, Fig. S5a). Cell-growth assays over 96 h showed no significant differences between psAP-KERP2gs and psAP-mock control (Fig. 3b, Fig. S5b-c).
a qRT-PCR confirming efficient knockdown of KERP2 in independent psAP-KERP2gs clones relative to psAP-mock. RNA polymerase II served as an endogenous control. Violin plots show data distribution with all individual technical replicates overlaid (n = 3 per clone). Adjusted P: #1 < 0.0001, #2 < 0.0001, #3 < 0.0001. Statistical analyses are described in the Methods (Statistics and reproducibility). b Proliferation over 96 h shows no growth defect upon KERP2 silencing. Cell density (log ml⁻¹) was measured at 24-h intervals. Points represent mean ± SD (n = 3 technical replicates). Biological replicates are provided in Fig. S5b-c. Adjusted P (24, 48, 72, 96 h): 0.6023, 0.9657, 0.0288, 0.9114. Statistical analyses are described in the Methods (Statistics and reproducibility). Source data are provided as a Source Data file. c KEGG analysis of RNA-Seq data from psAP-KERP2gs versus psAP-mock (up-regulated genes). Bubble size denotes the number of genes per term, color encodes –log₁₀(FDR), and the x-axis shows fold enrichment. Enriched categories include sulfur/cysteine–methionine metabolism, ubiquitin-mediated proteolysis/proteasome, carbon metabolism, ribosome, and amoebiasis. d Representative differentially expressed genes in psAP-KERP2gs (vs psAP-mock) grouped by KEGG category, with fold-change values. Notable increases include cysteine synthase paralogs, a cysteine protease, and amoebapore precursors, along with proteasome components. e Intracellular cysteine protease activity. f released cysteine protease activity into the medium. Values in (e, f) are normalized to psAP-mock ( = 100%). Violin plots show data distribution with all individual technical replicates overlaid (n = 8). Biological replicates are shown in Fig. S5f–i. Adjusted P in (e): HA-KERP2_G3 vs HA-mock_G3 0.0012; psAP-KERP2gs vs psAP-mock <0.0001. Adjusted P in (f): HA-KERP2_G3 vs HA-mock_G3 0.0135; psAP-KERP2gs vs psAP-mock <0.0001. Statistical analyses are described in the “Methods” (Statistics and reproducibility). Source data are provided as a Source Data file.
To assess transcriptional changes resulting from KERP2 knockdown, we performed RNA-Seq. Principal component analysis showed clear separation between psAP-KERP2gs and psAP-mock strains (Fig. S5d). Unless stated, differentially expressed genes (DEGs) are defined as padj <0.05 (Benjamini–Hochberg) and |log2FC | ≥ 1. Under this criterion, we identified 71 DEGs (47 up, 24 down; Supplementary Data 3a). For transparency, we also report genes with raw fold change ≥3 (|log2FC | ≥1.585) regardless of FDR (Supplementary Data 3b), along with the unfiltered counts table (Supplementary Data 3c); many entries in the ≥ 3-fold list do not meet multiple-testing correction and are not interpreted as significant.
KEGG over-representation of up-regulated gene set yielded several FDR-significant pathways (Fig. 3c). Amoebiasis (q = 2.1 × 10–4, n = 5) and Ribosome (q = 6.5 × 10–4, n = 7) were the strongest terms, followed by Sulfur metabolism (q = 3.6 × 10–3, n = 2), Ubiquitin-mediated proteolysis (q = 3.6 × 10–3, n = 3), and Cysteine & methionine metabolism (q = 1.0 × 10–2, n = 2). Additional lower-ranking but still significant categories included Carbon metabolism, Biosynthesis of amino acids, Sulfur cycle, and Proteasome (each q = 2.8 × 10–2, n = 1–2). Because several terms are supported by only 1–3 genes, we prioritize interpretation of the top-ranked Amoebiasis enrichment and consider the remaining terms as small-n secondary signal. Representative induced genes included cysteine synthases A paralogs ( ~ 7.74-fold; ~4.68-fold), a cysteine protease ( ~ 2.11-fold), amoebapore/pore-forming peptide precursors ( ~ 2.2–4.26-fold), the Gal/GalNAc lectin subunit precursor ( ~ 4.85-fold), and proteostasis components such as a 26S proteasome regulatory subunit ( ~ 10.35-fold) and ubiquitin-RPL40 fusion proteins ( ~ 2.3–2.4-fold) (Fig. 3d). No KEGG pathway passed FDR among down-regulated genes (n = 24), consistent with limited statistical power and the sparse annotation of Entamoeba.
Given the central role of cysteine proteases (CPs) in amoebic virulence, we quantified CP activity in knockdown and overexpression contexts and report all values as % of psAP-mock (set to 100%). Because Cl6 and G3 are known to differ in virulence, we first benchmarked CP activity and confirmed higher intracellular CP activity in Cl6 than G3 (p < 0.001), whereas released CP activity was comparable (n.s.) (Fig. S5e). To minimize strain-background effects in downstream comparisons, HA-KERP2 and the mock control were expressed in G3 background for experiments requiring pairing with gene-silencing strains. In this framework, psAP-KERP2gs displayed increased intracellular CP activity (129.0 ± 2.8%) and markedly elevated released CP activity (663.8 ± 4.5%) relative to psAP-mock (both p < 0.0001) (Fig. 3e, f; Fig. S5f-i; Supplementary Data 4). Conversely, HA-KERP2 overexpression in G3 decreased intracellular CP activity to 87.8 ± 1.1% versus 97.1 ± 1.0% in HA-mock_G3 (p = 0.0031) and reduced released CP activity to 117.1 ± 10.5% versus 155.3 ± 14.1% in HA-mock_G3 (p = 0.0324). We note that CP transcripts change modestly ( ~ 2-fold); the enzyme assays serve as an orthogonal functional readout of virulence rather than indicating extensive transcriptional regulation of individual protease genes.
Nuclear co-immunoprecipitation reveals KERP2 involvement in nucleic-acid binding and vesicle-trafficking networks in E. histolytica
To elucidate KERP2 function through protein-protein interactions, we performed anti-HA co-immunoprecipitation followed by MS on four biological replicates using HM-1:IMSS Cl6 transformants to maximize expression efficiency. HA-KERP2, HA-KERP2∆185−239, and HA-mock lines were fractionated into nuclear (nuclear/organellar-enriched) and cytosolic (cytosolic/light-vesicular) fractions before independent co-IPs. Western blotting confirmed bait recovery, and the HA constructs were identified by MS in inputs and eluates (Fig. S6a–d; Supplementary Data 5a-b). One nuclear HA-mock sample in Trial 3 showed non-specific HA signal and was excluded from analysis a priori.
Hit calling required ≥2‑fold higher quantitative value (QV) in HA-KERP2 or HA-KERP2∆185−239 versus the matched HA-mock fraction and detection in ≥2/4 replicates. Under these criteria, the nuclear HA-KERP2 co-IP yielded 75 double-hit and 4 triple-hit interactors, whereas the cytosolic HA-KERP2 co-IP yielded 14 double-hit and 5 triple-hit interactors (Fig. 4a, b, Supplementary Data 5c-d). Gene Ontology enrichment among nuclear interactors included vesicle-related and non-membrane-bound organelle components (including ribosomes), with functional associations in nucleic-acid/protein binding, metabolism, and ribosome biogenesis (Fig. 4c; Fig. S6e-f). In the cytosolic fraction, KEGG enrichment highlighted involvement in ribosome and protein processing in the endoplasmic reticulum (ER) (Fig. 4d).
a Venn diagram summarizing significant nuclear preys detected by anti-HA co-IP/MS across four biological replicates. Proteins were classified as double or triple hits based on quantitative values (QV) compared to HA-mock controls across replicates with adjusted p-value < 0.05. b Venn diagram of HA-KERP2 cytosolic proteome, highlighting double and triple hits as described above. c GO Cellular Component enrichment for the nuclear hit set (bubble size, number of genes; color, –log10 FDR; x-axis, fold enrichment). Enriched terms include vesicles/endocytic vesicle, non-membrane-bounded organelles, and ribosome/cytosolic ribosome. d KEGG pathway enrichment analysis for the cytosolic interactome hit set, highlighting glycolysis/gluconeogenesis, carbon metabolism, biosynthesis of amino acids, ribosome, and protein processing in the ER. e Heat map of nuclear co-IP/MS (centered and scaled per protein) across HA-KERP2, HA-KERP2∆185−239, and HA-mock pulldowns in each replicate. Bars annotate hit class (double/triple) and functional grouping (protein transport, protein folding, nucleic-acid binding, ribosome biogenesis). f Heat map as in E for cytosolic interactome (functional groupings: protein transport, protein folding, and nucleic-acid binding). g Immunofluorescence with anti-EhRab11B (red) and anti-HA (green) shows HA-KERP2 puncta partially co-localizing with EhRab11B-positive vesicles (arrows) in addition to nuclear signal (arrowheads). Single optical sections of the higher magnification image were acquired on an Olympus FV3000 confocal microscope (IX83) using a UPLSAPO 100×/1.40 oil-immersion objective. Images were recorded as 12-bit, 3-channel frames at 1024 × 1024 pixels (pixel size 0.124 × 0.124 µm). Laser settings (AOTF transmissivity, imaging phase) were: 405 nm 2.3% (CH1), 488 nm 7.4% (CH2), and 594 nm 86.9% (CH3). Representative line-scan intensity (grey value) profiles across the indicated lines are on the right. Scale bars: top, 10 µm; bottom, 5 µm. Source data are provided as a Source Data file. A 2D intensity histogram shows the correlation of EhRab11B (red) and HA (green) signals.
To further characterize the nuclear interactome, proteins were grouped by function beyond KEGG—protein transport, protein folding, nucleic-acid binding, and ribosome biogenesis—and visualized as a QV heat map (Fig. 4e, Table 1). Among the nuclear double/triple-hit interactors, predicted nucleic-acid-binding factors included Transcription factor/Nuclear export subunit protein 2 (EHI_052850; mean FC ≈ 60.9), Cell-division protein kinase, putative (EHI_105300; ≈110.1), Enhancer-binding protein-1 (EHI_121780; ≈100.0), HMG-box protein (EHI_179340; ≈58.5), RRM-domain protein (EHI_026440; ≈90.1), and a Histone deacetylase, putative (EHI_119320; ≈22.4) (Table 1). Enrichment versus HA-mock across ≥2/4 replicates is consistent with KERP2’s DEK-like, chromatin-engaging behavior.
Comparing full-length KERP2 with ∆185 − 239 revealed domain-dependent differences in the interactome. Proteins annotated for nucleic-acid binding and ribosome biogenesis were more enriched with full-length HA-KERP2, whereas ∆185 − 239 showed higher representation of protein-folding factors and selected transport components (Fig. 4e; Table 1; Supplementary Data 5c). The cytosolic interactome also revealed interactions with Granin1, Granin2, and Bip, consistent with nuclear interactors, along with EhC2B (Fig. 4f; Supplementary Data 5d). These patterns are consistent with the C-terminal coiled-coil contributing to interaction specificity in vivo.
Using the intersection of MS-quantified nuclear proteins and DESeq2-filtered genes from RNA-seq as the background, we evaluated the high-confidence nuclear interactors (double/triple hits; N = 79, Supplementary Data 5c) against KERP2gs vs psAP-mock RNA-seq (Supplementary Data 5e). The overlap was minimal: KERP2 (bait) and one non-bait interactor, the 26S proteasome regulatory subunit 7 (EHI_080890; RNA log2FC = 3.37, padj = 2.9 × 10–11; IP mean FC = 58.5). Excluding the bait, this corresponds to 1/79 (1.3%) overlap. Given the very small overlap set, an IP-RNA correlation cannot be interpreted meaningfully. These data indicate that KERP2’s physical interaction network is not explained by bulk expression shifts, consistent with an architectural chromatin co-regulator producing modest, distributed changes rather than large gene-specific transcriptional swings.
Among trafficking-related candidates captured with nuclear HA-KERP2 were EhRab11B, EhVPS45A, and a clathrin-coat assembly protein. Cytoplasmic ROI-restricted colocalization (nuclei excluded; Fiji/Coloc2, Costes thresholds) showed a moderate correlation between HA-KERP2 and EhRab11B in low-magnification fields (Pearson’s r ≈ 0.62; Costes P = 1.0; ~20 cells), while Manders’ coefficients were low (tM1 ≈ 0.000, tM2 ≈ 0.006), indicating sparse but reproducible co-occurrence of puncta in E. histolytica (Fig. 4g). Consistent with this, a reciprocal Myc-EhRab11B IP reproducibly recovered a weak HA-KERP2 signal absent from Myc-only control (Fig. S5j), supporting a low-abundance, transient association. These data do not establish direct binding or transport directionality, but support a model in which a subset of KERP2 engages EhRab11B-linked vesicular pathways implicated in cysteine-protease secretion in Entamoeba20.
Translocation of KERP2 from E. histolytica to Host Epithelial Cells
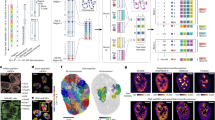
KERP1 and KERP2 were originally identified by their presence in the interactomes from the brush border of Caco-2 cells, but whether KERP2 merely associated with or is actively transferred into host cells has not been tested. To address this gap, we co-cultured HA-KERP2- or HA-mock-expressing trophozoites with Caco-2 monolayers. After 2 h, immunofluorescence revealed punctate HA-KERP2 signals within both the cytosol and nucleus of Caco-2 cells (Fig. 5a), whereas no signal was detected with HA-mock trophozoites. While nuclear-associated signal is detectable by imaging, we do not yet have mechanistic evidence for nuclear import routes or nuclear function of KERP2 in host cells. Interestingly, in contrast to the predominantly nuclear distribution of HA-KERP2 in the amoebic trophozoites at rest, a minor subset of interacting trophozoites displayed redistribution of HA-KERP2 to the cytosol and membrane (Fig. 5a). In a targeted search across six fields enriched for such events, 8 of 44 interacting trophozoites showed redistribution; however, this proportion should not be interpreted as an unbiased frequency estimate. These observations do not establish whether bulk uptake is required to elicit the host phenotypes reported below. One plausible explanation is that local enrichment at parasite–epithelial contact sites (microvilli/early endosomes) could be sufficient to trigger downstream signaling, even if net uptake across the monolayer is low.
a Caco-2 monolayers co-cultured 2 h with HA-KERP2–overexpressing trophozoites show punctate HA-KERP2 signal (anti-HA, green) with partial nuclear localization (DRAQ5, far red) alongside host cytoplasm (CellTracker™ Blue) and trophozoites (CellTracker™ Red). Right: representative line-scan intensity profiles across the indicated lines (profiles 1–3). Source data are provided as a Source Data file. Host-ROI colocalization analysis (amoebae excluded): Pearson’s r = 0.19 (no threshold), above-threshold Pearson’s r = −0.28; Manders’ tM1 = 0.302, tM2 = 0.432; Costes P = 1.00. Metrics and auto-threshold behavior were consistent with limited colocalization of HA-KERP2 signal in host cells. Single optical section shown unless indicated. b Live imaging of GFP-HA-KERP2 (no immunostaining) trophozoites interacting with a Caco-2 monolayer shows the appearance of GFP-positive puncta in host cells within ~30–45 min (DIC overlays; boxed regions mark Zones). Scale bar, 10 µm. c Orthogonal Z-slices from Zone 1 at the end of the movie (GFP-HA-KERP2, green; Hoechst 33342, blue) confirm intracellular puncta; 3D projection and DIC are shown at right. Scale bars, 5 µm. d Schematic of the human intestinal 3D crypt model (stem-cell–derived crypts differentiated to enterocytes). Created in BioRender. Santos, H. (2026) https://BioRender.com/w10n355e Crypts were exposed for 2 h to GFP–HA–KERP2 (top) or GFP–HA control (bottom). Anti-HA (green), Hoechst (blue), and CellTracker Deep Red labeled amoebae (magenta) imaged by spinning-disk confocal microscopy with orthogonal views. Scale bars, 20 µm. f Fractionation and immunoblot showing transfer of HA-KERP2 to host cells after co-culture. Parasite (E. histolytica) and host (Caco-2) lysates were separated and probed with anti-HA. Lane order (left to right): E. histolytica lysates—1 h; 1 h + galactose (adhesion inhibition); 2 h; 3 h. Caco-2 lysates—1 h; 1 h + galactose; 2 h; 3 h; unexposed Caco-2 control. Fractionation controls: CS-1 (amoeba marker) and GAPDH (host marker). HA-KERP2 is detectable in host fractions by 1 h, increases at 2–3 h, and is reduced by galactose, consistent with adhesion-dependent translocation. Equal total protein was loaded per lane; molecular-weight standards indicated at left. Source data are provided as a Source Data file.
To distinguish true host-cell signal from parasite signal, we masked CellTracker-Red–positive amoebae and computed colocalization within the remaining host-cell ROI. Across these ROIs, Pearson’s r (no threshold) between HA-KERP2 and the host channel was 0.19, while the above-threshold Pearson’s was −0.28; Manders’ coefficients were tM1 = 0.302 and tM2 = 0.432. Costes randomization yielded P = 1.00. The analysis also flagged that automatic thresholds were set above channel means and that the regression intercept was far from zero (b/mean = −3.27), indicating background/offset effects that can bias above-threshold overlap. Taken together, these metrics support detectable but limited colocalization in host cells under these conditions. Extra-amoebic HA-KERP2 fluorescence was extremely faint—consistent with high dilution near the detection limit—but line-profile analyses across the amoeba–host interface nevertheless demonstrated a shallow and reproducible shift of HA-KERP2 into adjacent Caco-2 cytoplasm (Fig. 5a).
To minimize projection-induced background, live imaging of GFP-HA-KERP2 trophozoites moving over Caco-2 monolayers is presented as single optical sections; one maximum-intensity projection is included for morphology only (Fig. 5b; Supplementary Movie 3-4). Under these conditions, GFP signal appeared within Caco-2 cells between 30 and 45 min of contact, distributed in discrete zones; higher-magnification single sections again showed punctate structures resembling those observed by IFA (Fig. 5c). No transfer was observed with trophozoites expressing GFP-RtcB2, a cytosolic tRNA ligase, which serves as an irrelevant control (Fig. S7a-b).
In a biopsy-derived 3D crypt model with differentiated enterocytes (Fig. 5d), we used a 3D reconstruction to visualize tissue architecture by labeling enterocyte nuclei with Hoechst and E. histolytica with CellTracker Deep Red. Compared with planar Caco-2 monolayers, this model preserves epithelial polarization and crypt–villus topology, providing a native apical interface for parasite contact. Under these conditions, GFP-HA-KERP2 puncta were detected inside host cells after 2 h of contact, whereas GFP-HA alone remained within trophozoites (Fig. 5e). As above, the 3D views are provided to illustrate spatial context; quantitative conclusions rely on nucleus-excluded, single-section ROI colocalization metrics (Pearson/Manders) and biochemical fractionation/immunoblotting below; line scans are shown only as representative visualizations.
Fractionation and immunoblotting again confirmed transfer. After co-culture of HA-KERP2 trophozoites with Caco-2 monolayers, physical separation of host and parasite fractions was validated by GAPDH and CS1, which serves as a host or parasite marker, respectively. Immunoblot analysis of these fractions revealed HA-KERP2 in host-cell lysates within 1 h, with signal increasing over 2–3 h (Fig. 5f). Galactose treatment, which inhibits E. histolytica adhesion, significantly reduced HA-KERP2 detection in host fractions, indicating that adhesion is required for translocation. These observations demonstrate that KERP2 can be transferred to host cells under our experimental conditions but do not establish how frequently such transfer occurs during infection.
KERP2 trafficking mechanism in E. histolytica and Caco-2 cells
To investigate KERP2 trafficking during E. histolytica interaction with epithelial cells, we performed immuno-EM to visualize the localization of HA-KERP2 (Fig. 6a). E. histolytica trophozoites expressing HA-KERP2 were co-cultured with Caco-2 for 1 h. In E. histolytica, HA-KERP2 signals were predominantly detected near the contact site with Caco-2 cells, in addition to the nuclear pool. In Caco-2 cells, HA-KERP2 was observed at the microvillus-rich brush border, within the cytosol, and in endosome-like structures. This spatial pattern is consistent with a model in which contact-site microdomains, rather than uniform cytosolic accumulation, are the functionally relevant sites of KERP2 action in host cells.
a Immuno-EM of parasite-host interfaces. E. histolytica trophozoites were co-cultured with Caco-2 monolayers for 1 h co-culture for anti-HA immunogold EM. Overview images (left) and corresponding high-magnification boxes (right; orange and green) show HA-KERP2 signals (yellow circles in trophozoites; light blue circles in Caco-2. Label is detected at the parasite plasma membrane, in granule-like compartments, and on/near host microvilli and endosome-like profiles (arrows). Scale bars as indicated. b Phospholipid binding. Lipid overlay (PIP-strip) probed with recombinant His-KERP2 or His-GFP (negative control). Spots contain 100 pmol per lipid. His-KERP2 binds mono- and bis-phosphorylated phosphatidylinositols [PI(3)P, PI(4)P, PI(5)P, PI(4,5)P₂]; negligible signal is seen with His-GFP. c Time course of uptake by flow cytometry. Representative FSC-A vs FITC-A plots of Caco-2 cells trypsinized after treatment to 3 μM His-GFP-KERP2 (upper row) or His-GFP (bottom row) for indicated times (0-24 h). d Quantification of uptake. Median FITC fluorescence intensity (MFI) from (c) over time. Violin plots show data distribution with all individual biological replicates overlaid (n = 3). Adjusted P (0, 1, 2, 4, 6, 24 h): >0.9999, 0.6650, 0.3627, 0.0643, 0.0010, <0.0001. Statistical analyses are described in the Methods (Statistics and reproducibility). Source data are provided as a Source Data file. e Energy dependence of internalization. Representative FACS plots after 4 h incubation at 35.5 °C or 4 °C. Top row: 3 µM His-GFP-KERP2 (FITC-A). Bottom row: RITC-dextran endocytic tracer (PE-A) run in parallel. Reduced signals at 4 °C indicate an energy-dependent process. f Persistence after washout. Caco-2 cells were exposed to 3 µM His-GFP-KERP2 (or His-GFP), washed after 24 h, and analyzed immediately (24 h), 24 h post-wash, and 48 h post-wash. Flow plots show retention of intracellular KERP2 signal compared with GFP control.
We next asked whether KERP2 is secreted in extracellular vesicles (EVs) or associated directly with membranes. Under our conditions, HA-KERP2 was undetectable in EV preparations, whereas HA-KERP2Δ185–239 and GFP-HA were enriched (Fig. S7c), suggesting that EV-mediated export is not a major route for the native full-length KERP2 protein. Given co-IP data hinting at the involvement of EhRab11B-dependent recycling endosomes in KERP2 trafficking in E. histolytica, we directly examined KERP2’s lipid interactions using a lipid overlay assay. The lipid overlay assay showed that recombinant His-KERP2, but not His-GFP control, bound to anionic phospholipids, including PI(3)P, PI(4)P, PI(5)P, PI(4,5)P2, and PA (Fig. 6b; Fig. S8a-b).
To further investigate the mechanism of KERP2 uptake by host cells, His-GFP-KERP2 and His-GFP expressed in E. coli were applied to Caco-2 monolayers at a final protein concentration of 3 μM in complete EMEM. After incubation, Caco-2 cells were trypsinized to remove surface-bound protein, and GFP uptake was quantified by flow cytometry (Fig. 6c, d; Fig. S8c-d). Within 1 h, 4.33% of Caco-2 cells were GFP-positive when treated with His-GFP-KERP2, rising to 25.2% by 24 h, accompanied by an increase in median fluorescence intensity (MFI). In contrast, cells exposed to His-GFP alone showed minimal uptake.
To determine whether endocytosis plays a role in KERP2 internalization, we compared uptake at 35.5 °C vs 4 °C using RITC-dextran (endocytosis marker) and His-GFP-KERP2 (Fig. 6e). RITC-dextran uptake decreased from 53.7% to 1.92% at 4 °C, and His-GFP-KERP2 uptake reduced from 16.5 to 2.25%, supporting an energy-dependent endocytic process.
Immunofluorescence analysis of His-GFP-KERP2–treated Caco-2 cells revealed punctate, endosome-like structures with partial nuclear localization, mirroring the pattern observed when HA-KERP2 is deposited by trophozoites (Fig. S8e). After a 24 h pulse with His-GFP-KERP2 followed by extensive washing and medium replacement, GFP signals persisted in 11.7% and 7.2% of Caco-2 cells at 24 h and 48 h post-wash, respectively, suggesting that a fraction of internalized His-GFP-KERP2 is retained (Fig. 6f).
Functional impact of KERP2 on host protein networks and gene expression
To investigate the impact of KERP2 on host epithelial cells, we performed co-IP coupled with MS on three biological replicates to identify KERP2-interacting proteins in Caco-2 cells. Caco-2 monolayers were co-cultured with HA-KERP2- or HA-mock-expressing trophozoites, followed by separation, lysate collection, and co-IP. Candidate interactors were called primarily by replicate reproducibility: proteins showing ≥2-fold enrichment over in HA-mock in ≥2/3 replicates were designated as “double-hit” and those enriched in 3/3 replicates were labeled “triple-hit”, resulting in 78 double-hit and 5 triple-hit proteins (Fig. 7a; Supplementary Data 6a-b). For transparency, we also report per-protein log2 fold-change and Welch’s two-sample p values (left-censored log scale imputation), and BH-FDR (Supplementary Data 6a). Because many interactors are bait-specific or near the detection limit in the mock control, these statistical measures are conservative; thus, our primary criterion is double-/triple-hit reproducibility, with p/FDR values provided as supplementary evidence.
a Anti-HA IP-MS of HA-KERP2-exposed in Caco-2 cells (three biological replicates). Venn shows significant interactors: double hits ( ≥ 2-fold vs HA-mock in ≥2/3 replicates, padj <0.05) and triple-hits (meeting the criterion in all three). b GO Molecular Function enrichment of the combined double-/triple-hit list. Bubble plot: x-axis = GeneRatio; bubble size = gene count; color = FDR. c Heat map of representative interactors (columns) across HA-KERP2, HA-KERP2Δ185–239, and HA-mock IPs (rows). Values are QV intensities. Annotation bars indicate hit type (double/triple) and manual functional grouping (cell-cycle & proliferation; cytoskeletal dynamics; cell–cell/ECM adhesion; intracellular trafficking & vesicle transport; immune/stress; gene regulation & transcription). d Contrast-focused PCA. Genes from the HA-KERP2 vs psAP-KERP2gs contrast (DESeq2, adjusted p < 0.05, |log2FC | ≥ 1) were passed to PCA analysis, showing condition-specific clustering. e Gene-wise comparison of differential expression for HA-KERP2 (x-axis, log2FC) versus psAP-KERP2gs (y-axis, log2FC). Points are colored by significance (padj <0.05): in HA-KERP2, in KERP2gs, or significant in both; exemplar genes are labeled. f, g Preranked GSEA for the HA-KERP2 vs KERP2gs contrast. Genes were ranked by signed –log10(padj) (sign = sign of log2FC); positive NES indicates enrichment in HA-KERP2 and negative NES indicates enrichment in KERP2gs. Only sets with 10–500 genes and FDR q < 0.25 are plotted. f Hallmark collection. g GO Biological Process. Bubble size = gene-set size; color = –log10(FDR).
GO enrichment revealed associations with cadherin binding, actin binding, and transcription co-activator binding (Fig. 7b). Functional categorization (Fig. 7c) highlighted triple-hit interactors PPP6C, PPP6R, and RANGAP1, and double-hit proteins CUL5, UBR5, and MIOS, which are linked to signaling and cell-cycle control. Cytoskeletal/adhesion components were prominent (PDLIM1, PFN1, ARPC3, CNN3, LASP1, CLDN4, ITGB1, and CTNNB1), alongside trafficking factors (EEA1, SNX5, and DCTN1).
We next compared the HA-KERP2 interactome in Caco-2 cells with previously characterized amoebic HA-KERP2 interactomes (nuclear and cytosolic fractions; Fig. 4c, d). No shared protein identifiers were detected, confirming that the host pulldown captured host rather than parasite proteins. Nonetheless, functional overlap emerged at the pathway level. Amoebic nuclear and cytosolic interactomes were enriched for endocytic vesicle organization, glycolysis/carbon metabolism, and ribosome components, while the Caco-2 interactome showed enrichment for cadherin binding, actin filament organization, transcriptional co-activation, and vesicle trafficking (Fig. 7b, c). These results suggest that KERP2 targets analogous structural and trafficking frameworks in both parasite and host contexts, despite differences in the specific proteins involved.
To evaluate the effects of KERP2 on host gene expression, RNA-seq analysis was performed after co-culture of Caco-2 cells with wild-type G3, HA-KERP2, and psAP-KERP2gs, all of which have the identical genetic background of G3—alongside unexposed Caco-2 controls (n = 3 per condition; Supplementary Data 7a). PCA and volcano plots confirmed clear condition-specific clustering (Fig. S9a–d). Focusing on KERP2-dependent effects, a contrast-focused PCA on differentially expressed genes from HA-KERP2 vs KERP2gs again showed clear separation (adjusted p < 0.05, |log2FC | ≥ 1; Fig. 7d; Supplementary Data 7b). In HA-KERP2-exposed Caco-2 cells, representative upregulated genes included IL1B, IL36G, SLA, HRG, whereas CYTIP, MYOCD, NUPR1, SLC15A3 were downregulated (Fig. 7e). Conversely, psAP-KERP2gs-exposed cells showed increased TNC, MYL7, CYP4B1 and decreased CFAP43, SLC2A12, TKTL1 gene expression.
Consistent with the previous Caco‑2 3D intestinal model studies21, our monolayer data show a UPR/stress response (CHOP), chemokine induction (IL1B and CXCL8), and a cell‑cycle restraint signature (CDKN1A) (Supplementary Data 7c). We also observed a modest decrease in STAT1, compatible with early interferon‑pathway dampening. Notably, transcripts for IL6, TNF, and CSF2 were not robustly induced, and no global down‑shift of ribosome biogenesis was detected—likely reflecting the absence of non‑epithelial compartments and paracrine amplification in our monolayer system. Collectively, these datasets suggest a shared epithelial program characterized by stress response, inflammatory chemokine induction, and proliferation restraint, whereas broader immune and translational suppression is primarily observed in more complex 3D tissue models.
We next performed preranked GSEA on the HA-KERP2 vs KERP2gs contrast (following the convention that positive NES indicates enrichment in HA-KERP2 and negative NES indicates enrichment in KERP2gs). In the Hallmark collection, HA-KERP2 showed enrichment for glycolysis, fatty-acid/bile-acid/heme metabolism, peroxisome, hypoxia/mTORC1 signaling, and protein secretion (FDR q < 0.25), while KERP2gs showed enrichment for apical junction, apoptosis/ROS, E2F targets/G2M checkpoint, and MYC/WNT programs (Fig. 7f). In GO Biological Process, HA-KERP2 was enriched for phosphatidylinositol/phospholipid/glycerolipid metabolism, organic-acid catabolism/biosynthesis, and macroautophagy, whereas KERP2gs was enriched for actin/cytoskeleton organization, integrin-mediated signaling, actomyosin structure organization, and nucleosome organization (Fig. 7g). In GO Molecular Function, HA-KERP2 showed enrichment in oxidoreductase and enzyme-adaptor activities, whereas KERP2gs was enriched for cadherin/integrin binding, actin-filament binding, and structural constituent of chromatin (Fig. S9e).
Effects of KERP2 on DNA synthesis and cell-cycle progression in epithelial cells
To evaluate the effects of KERP2 on host cell-cycle progression, EdU incorporation assays were performed in Caco-2 cells following a 2-h interaction with E. histolytica strains in the G3 background (HA-KERP2, HA-mock, KERP2gs, and psAP-mock) or with no exposure. After co-culture, parasites were removed by 2% galactose washes and Caco-2 cells were pulsed with EdU for 6 h in complete EMEM. Confocal imaging showed group-wise differences in EdU-positive nuclei (Fig. 8a).
a EdU incorporation in Caco-2 monolayers after brief co-culture with E. histolytica strains. Cells were exposed for 2 h to HA-KERP2, HA-mock, KERP2gs, or psAP-mock (or left unexposed), then pulsed with EdU for 6 h and processed with the Click-iT kit (Alexa Fluor 488, green); nuclei were counterstained with Hoechst 33342 (blue). Images are single optical sections acquired on an Olympus FV3000 (IX83) with a UPLSAPO 20×/0.75 air objective; pinhole 120 µm; 12-bit. Laser settings (AOTF): 488 nm 7% (EdU), 405 nm 0.6% (Hoechst). Scale bars, as indicated. b Quantification of EdU-positive nuclei for the condition in (a). Violin plots show data distribution with all individual fields overlaid (n = 5 fields per condition). Statistics: ordinary one-way ANOVA (two-sided; F(4,20) = 12.55, P < 0.0001), followed by Dunnett’s multiple comparisons vs unexposed Caco-2 (multiplicity-adjusted P values, family-wise α = 0.05). Adjusted P values: HA-KERP2, 0.0006; HA-mock, 0.0030; KERP2gs, 0.8711; psAP-mock, 0.0036. Source data are provided as a Source Data file. c Dose–response with recombinant protein. Caco-2 cells were incubated for 2 h with His-GFP-KERP2 (1, 3, or 5 µM) or His-GFP control, followed by a 6 h EdU pulse and Click-iT detection (Alexa Fluor 594, red). Nuclei were counterstained with Hoechst 33342 (blue). Images acquired on an Olympus FV3000 (IX83) with PLAPON 60×/1.40 oil; pinhole 213 µm, 12-bit. Laser settings (AOTF transmission): 594 nm 98.7% (EdU) and 405 nm 52.5% (Hoechst). Scale bars, as indicated. d Quantification of EdU-positive nuclei for the conditions in (c). Violin plots show data distribution with all individual fields overlaid (n = 5 fields per condition). Adjusted P: His-GFP 1 µM > 0.9999; His-GFP 3 µM 0.9711; His-GFP 5 µM > 0.9999; His-GFP–KERP2 1 µM 0.0059; 3 µM 0.0011; 5 µM < 0.0001. Statistical analyses are described in the “Methods” (Statistics and reproducibility). Source data are provided as a Source Data file.
Quantification of EdU-positive nuclei, counterstained with Hoechst 33342, showed higher EdU-positive fractions in HA-KERP2-, HA-mock-, and psAP-mock-exposed cells (48 ± 4.0%, 46 ± 5.3%, 45 ± 3.3%) relative to unexposed Caco-2 (35 ± 4.2%), whereas psAP-KERP2gs exposure resulted in 33 ± 4.3% (Fig. 8b). To test sufficiency, recombinant His‑GFP–KERP2, but not His‑GFP, increased EdU labeling in a dose‑responsive manner, with significant effects at higher concentrations that recapitulated the co‑culture phenotype (Fig. 8c, d). For context, the recombinant KERP2 concentrations used (1–5 µM) correspond to ~6 × 1014–3 × 1015 molecules per mL of medium. Because we did not measure absolute intracellular copy number or uptake, and exposures were performed on confluent Caco-2 monolayers in Millicell EZ 8 well chamber slides with 200 μL of medium (growth area 1.0 cm² per well), these values should be interpreted only as an external exposure scale, not a quantified uptake per cell. Although absolute EdU fractions varied across experimental formats, statistical comparisons within each format were internally consistent. Because EdU detects both replicative and repair DNA synthesis, these findings support a KERP2-dependent increase in host DNA synthesis during early contact, without resolving a specific cell-cycle transition involved.
Effects of KERP2 on the cytoskeleton and monolayer motility
To examine the impact of KERP2 on cytoskeletal organization, Caco-2 monolayers were stained for E-cadherin (adherens junctions), F-actin (phalloidin 594), and nuclei (Hoechst 33342) after a 2-h interaction (Fig. 9a). Morphometric analysis revealed a decrease in cellular circularity (form factor) in HA-KERP2-exposed cells versus unexposed controls (0.53 ± 0.091 vs 0.59 ± 0.073), whereas KERP2gs-exposed cells (0.57 ± 0.09) were similar to controls (Fig. 9b). Consistent with elongation, aspect ratio increased modestly but significantly with HA-KERP2 (1.7 ± 0.5) compared with unexposed controls (1.6 ± 0.38), while KERP2gs remained unchanged (1.6 ± 0.39) (Fig. 9c). HA-KERP2-exposed cells displayed disrupted basal stress fibers and a thinner, less continuous cortical F-actin belt at cell edges, whereas KERP2gs exposure was associated with prominent stress fiber formation (Fig. 9d). Qualitatively similar morphology was observed after short-term exposure to recombinant His-GFP–KERP2 compared with His-GFP (Fig. S8e-f). These patterns indicate actin cytoskeletal remodeling, with KERP2 favoring loss/weakening of contractile bundles and KERP2gs favoring stress-fiber assembly.
a Apical confocal images of Caco-2 monolayers after 2 h co-culture with the indicated E. histolytica strains. E-cadherin (green), F-actin (phalloidin, red), nuclei (Hoechst, blue). scale bars as indicated. b, c Cell-shape quantification from apical planes in (a), pooled from 5 imaging fields per condition (each dot = one cell; total cell n in Source Data). b Form factor (FF = 4π·area/perimeter²). Adjusted P: HA-KERP2 < 0.0001; KERP2gs 0.0012. c Aspect ratio (major/minor axis). Adjusted P: HA-KERP2 0.0005; KERP2gs 0.1130. d Basal-plane phalloidin staining after 2 h co-culture highlighting cortical actin remodeling; scale bars as indicated. e Wound-healing assay with recombinant proteins. Relative wound area (%) over time after 2 h treatment with His-GFP-KERP2 (1–5 µM), His-GFP, or untreated. n = 2 technical replicates (imaging fields); biological replicates in Fig. S10d. Points show mean ± SD; linear fits (slope annotated). Exact P values (vs untreated): His-GFP (5 µM) 0.7941; His-GFP–KERP2 (3 µM) < 0.0001; His-GFP–KERP2 (5 µM) < 0.0001. f TEER during co-culture with live parasites. Values normalized per well to pre-addition TEER and expressed as % of mean t = 0 (set to 100%). Curves show mean ± SD (n = 3 technical replicates); biological replicates in Fig. S11a–d. g Summary of (f). Violin plots; n = 3 technical replicates. Exact P values in Supplementary Data 8a (and on plot). h Recombinant proteins: TEER normalized to per-well baseline (%), mean t = 0 set to 100%. Points show mean ± SD (n = 3 technical replicates); biological replicates in Fig. S11e–f. Significance: ns, P ≥ 0.05; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Exact P values in Supplementary Data 8b. i Size-selective permeability (10-kDa RITC–dextran flux; final time point). Violin plots; n = 4 technical replicates, normalized to unexposed Caco-2; biological replicates in Fig. S11g–h. P: HAKERP2 vs HA 0.0002; KERP2gs vs psAP 0.0003. j Proposed working model of KERP2 linking parasite nuclear regulation to epithelial cytoskeletal remodeling and barrier disruption. Created in BioRender. Santos, H. (2026) https://BioRender.com/efd7t8x. All statistical analyses are described in the “Methods” (Statistics and reproducibility). Source data are provided as a Source Data file.
Because changes in cell shape and stress fibers often translate into altered collective motility, a wound healing assay was performed under low serum to limit proliferation. Monolayers were exposed for 2 h to HA-KERP2, HA-mock, KERP2gs, or psAP-mock, washed with 2% galactose, and monitored for gap recovery (Fig. S10a). During the first 6 h, HA-KERP2-exposed monolayers tended to close more slowly than KERP2gs-exposed monolayers, but differences were not significant across conditions (Fig. S10b). To enable longer treatments with more uniform uptake, we next used recombinant proteins and further reduced serum to 0.5% to minimize proliferation. Under these conditions, recombinant His-GFP–KERP2 reproducibly and significantly reduced the wound closure compared to His-GFP and untreated controls (Fig. 9e; Fig. S10c-d). Thus, KERP2 exposure is sufficient to reduce wound closure under reduced-proliferation conditions, whereas co-culture with KERP2gs yielded an upward trend but no significant effect in our short assay window. Notably, EdU labels S-phase at a time point and does not by itself quantify net proliferation during the closure window; accordingly, we report closure phenotypes without assigning them to migration in the absence of a mitotic block.
Effects of KERP2 on epithelial barrier integrity
To evaluate the impact of KERP2 on epithelial monolayer integrity, transepithelial electrical resistance (TEER) was measured in Caco-2 cells cultured on transwell inserts and exposed to different E. histolytica strains. Because trophozoites deteriorate under aerobic conditions, TEER measurement during live co-culture was limited to a 4-h window (Fig. 9f, g; Fig. S11a–d). TEER declined for all strains, with the largest loss for KERP2gs, dropping from 101.7 ± 5.7% to 49.7 ± 1.2% at 1 h (–52.0 ± 4.5%) and to 28.7 ± 1.2% at 2 h (-73.0 ± 4.5%). HA-KERP2 decreased from 98.7 ± 4.0% to 62.7 ± 0.6% at 1 h (-36.0 ± 3.5%) and to 33.0 ± 1.0% at 2 h (-65.7 ± 3.5%). HA-mock decreased to 71.0 ± 1.0% (-32.3 ± 3.2% at 1 h) and 49.3 ± 1.5% (-54.0 ± 3.2% at 2 h), whereas psAP-mock to 79.0 ± 0.0% (–25.3 ± 4.5%) and 68.0 ± 1.0% (–36.3 ± 4.5%) over the same interval. Thus, the magnitude of TEER loss ranked KERP2gs > HA-KERP2 > HA-mock > psAP-mock. Because KERP2gs increases cysteine-protease activity, its larger TEER loss is likely secondary to upregulated virulence factors, rather than a direct effect of KERP2 on host cells.
To test sufficiency and to examine kinetics beyond the 4-h co-culture limit, purified His-GFP–KERP2 (1–5 µM) or His-GFP control was applied to the apical medium of Caco-2 cell cultures, and TEER was followed at 0–7 h (extended kinetics) and 12–24 h (recovery/steady-state phase) (Fig. 9h; Fig. S11e, g). KERP2 reduced TEER in a dose- and time-dependent manner: by 7 h, TEER reached 89% ( − 11%) at 1 µM, 87% ( − 13%) at 3 µM, and 79% ( − 21%) at 5 µM, whereas untreated controls increased to 141%, yielding deficits of 52–62% vs untreated. At 12 h, TEER values were 72% ( − 28%), 69% ( − 31%), and 83% ( − 17%); at 24 h, 68% ( − 32%), 57% ( − 43%; largest sustained loss), and 65% ( − 35%), respectively at each concentration. His-GFP controls remained at 100–124%, and EGTA (5 mM) stabilized ~75% at 24 h.
Finally, RITC–dextran permeability diverged from TEER rankings (Fig. 9i; Fig. S11f, h). HA-KERP2 produced the highest flux (166.8 ± 2.2% of unexposed), exceeding HA (154.0 ± 5.4%) and psAP (118.8 ± 2.1%). KERP2gs was near baseline (106.8 ± 4.2%) and lower than psAP. EGTA reduced TEER but did not increase 10-kDa flux (103.8 ± 1.3%; n.s. vs unexposed 99.8 ± 4.3%,).
Both live E. histolytica strain and purified KERP2 depressed TEER, with live KERP2gs strain and recombinant KERP2 showing the strongest sustained reductions. However, increased macromolecular permeability was most pronounced with HA-KERP2-expressing strain, and EGTA lowers TEER without increasing 10-kDa flux. Thus, TEER reduction alone does not uniformly predict 10-kDa permeability, indicating distinct effects on ion conductance versus macromolecular passage. These relationships between KERP2-dependent TEER depression, macromolecular flux, and parasite versus recombinant protein exposure are integrated in a conceptual model of epithelial barrier modulation (Fig. 9j).
Discussion
E. histolytica heavily relies on adhesion to epithelial cells to initiate colonization and infection, a process mediated by interactions between the trophozoite plasma membrane and host brush border microvilli13. KERP2, identified alongside KERP1, was initially predicted to localize to the extracellular milieu despite lacking a transmembrane domain or GPI anchor, and its high isoelectric point and positively charged surface suggested potential interactions with negatively charged molecules at the host-pathogen interface.
In light of our current findings, KERP2 is better interpreted as a lineage-specific, chromatin-engaging factor that Entamoeba has retained under purifying selection for essential cellular functions. Phylogenetically, KERP2 (with its paralog KERP3) forms an Entamoeba-restricted clade; homolog searches across multiple kingdoms reveal no confidently assignable orthologs outside Entamoeba and only weakly conserved KERP2-like sequences in P. pallidum and D. purpureum, underscoring a specialized role within Amoebozoa.
Its domain features including a DEK-like SAP module compatible with AT-rich DNA engagement, an overall basic surface, and a C-terminal coiled-coil region overlapping a nuclear localization signal, are more consistent with a chromatin-associated architectural adaptor that modulates nucleosome topology and nucleocytoplasmic routing than with a sequence-specific transcription factor22,23,24,25. This interpretation is also supported by its nuclear accumulation and interactome, which includes other architectural and chromatin-associated factors. Together, these observations argue that KERP2 is a conserved, Entamoeba-specific chromatin regulator that has secondarily been co-opted as host interaction.
The localization of KERP2 in E. histolytica was confirmed using multiple complementary approaches, including IFA, live-cell imaging, immuno-EM, and co-IP. HA-KERP2 predominantly resides at the nuclear periphery and within the nucleoplasm, with additional puncta observed in the cytosol, whereas the truncated HA-KERP2∆185−239 variant lacking the coiled-coil region is largely cytosolic, underscoring the requirement of the C terminus for efficient nuclear enrichment. Immuno-EM further reveals HA-KERP2 within electron-dense nuclear regions typically associated with chromatin-rich areas, and nuclear co-IP recovers an interactome enriched for transcription-associated proteins and nuclear transport factors.
In vitro, KERP2 binds AT-rich DNA and displays a distinctive EMSA phenotype in which KERP2–DNA complexes migrate faster than free probe under native conditions. A parsimonious explanation is that KERP2 induces bending or compaction of AT-rich DNA, thereby reducing hydrodynamic drag—an effect reminiscent of the faster migration of supercoiled DNA relative to relaxed topoisomeric forms26. Consistent with a compaction-driven mechanism, the “fast shift” is competitively attenuated by unlabeled AT-rich DNA, implying mass-action sequestration of KERP2 and reduced compaction of the labeled probe.
In vivo, chromatin immunoprecipitation demonstrates that GFP-HA-KERP2, but not GFP-HA-mock, recovers parasite genomic DNA. Although enrichment is modest and no strict consensus motif emerges, enriched regions show a clear bias toward A/T-rich, promoter-proximal intervals, and lack discrete sequence motifs. This pattern argues that KERP2, like DEK proteins, recognizes high-order chromatin structures rather than a specific base-pair word. Transcriptomic profiling of KERP2 knockdown parasites reveals coordinated downregulation of genes involved in sulfur amino acid metabolism genes, including cysteine synthases, as well as virulence-associated factors such as cysteine protease and pore-forming peptides.
Taken together, these observations support a model in which KERP2 functions as a broad chromatin-associated co-regulator that buffers multiple transcriptional modules, rather than acting as a canonical, sequence-specific transcription factor restricted to sulfur metabolism. These findings suggest that KERP2 may function as a chromatin-associated co-regulator that modulates virulence factor expression. Sulfur metabolism pathway genes represent one responsive node within a larger regulatory network, not an exclusive target. This regulation is particularly critical for E. histolytica as an anaerobic parasite, where sulfur-containing amino acids are essential for maintaining the antioxidant defenses required for survival and pathogenesis. Loss of KERP2 therefore appears to destabilize chromatin states governing both metabolic homeostasis and virulence outputs, shifting the parasite toward a transcriptional profile associated with heightened aggressiveness.
Available data link KERP2’s domains to separable functions. The N-terminal SAP-like domain provides a DNA-binding “handle”: it engages AT-rich DNA duplex (and single-stranded DNA) and produces the characteristic fast EMSA behavior, a profile more consistent with compaction than simple mass loading. In contrast, the C-terminal coiled-coil region, which overlaps a predicted NLS, is required for efficient nuclear enrichment and promotes self-association27,28,29. Removing this region reduces nuclear accumulation and shifts purified protein toward a monomer-range elution profile, supporting the view that the coiled-coil increases effective avidity through weak oligomerization or through docking to partner proteins30.
Under native conditions, nuclear KERP2 remains largely monomeric, indicating that any higher-order assemblies are transient, low-abundance, or nucleated only upon chromatin engagement. In combination, these features support a model in which the SAP-like N-terminus anchors KERP2 at AT-rich chromatin, while the coiled-coil/NLS module tunes nuclear access and interaction valency—properties that align with a chromatin co-regulator that restrains inappropriate activation of virulence- and metabolism-linked genes.
KERP2 exhibits a dynamic intracellular distribution in E. histolytica, with immuno-EM localization revealing its presence in the nucleus, vesicular structures, and electron-dense granule (EDG)-like compartments within the cytosol. Co-IP experiments identify EhRab11B, EhVPS45, and clathrin-coat assembly protein as interactors, placing KERP2 on a recycling endosomes-exocytic axis consistent with EhRab11B-mediated trafficking31,32,33. In E. histolytica, EhRab11B regulates vesicle transport and secretion of cysteine proteases20, suggesting that KERP2 may traverse similar routes to reach membrane-proximal compartments.
One possible explanation for the selective secretion or surface access of KERP2 is its preferential association with anionic membrane interfaces, potentially driven by its strongly positively charged surface34,35. Lipid overlay assays show that recombinant His-GFP-KERP2 strongly binds to phosphoinositides, including PI3P, PI4P, and PI(4,5)P2, which are enriched in endosomal and plasma membranes36. These interactions support a model in which electrostatic docking to phosphoinositide-rich microdomains facilitate KERP2 positioning at host-parasite contact sites.
Despite its presence in EhRab11B-positive vesicles, KERP2 was not detected in crude EV fractions under our conditions, nor in previously published EV proteomes that intermittently detect KERP137. This pattern suggests that, in our system, KERP2 reaches the parasite surface via targeted recycling/endosomal pathways, rather than bulk exosome-like release. However, we cannot exclude the possibility of low-abundance EV incorporation; surface-labeling assays under non-permeabilizing conditions, EV co-isolation with canonical vesicle markers when available, targeted MS of surface- and EV-enriched fractions, and quantitative lipid binding will be required to resolve these routes more definitively.
To answer a critical question whether KERP2 can be translocated into host cells during E. histolytica-epithelial interactions, IFA, live-cell imaging, and immunoblot analysis of E. histolytica-Caco-2 co-cultures were performed. KERP2 signals were detected inside Caco-2 cells and in a 3D enterocyte crypt model as punctate cytosolic structures. Blocking parasite adhesion abolished detectable KERP2 transfer, and Immuno-EM localized KERP2 to microvilli, the host cytosol, and endosome-like vesicles, consistent with contact-dependent endocytic uptake. Uptake of recombinant KERP2 persisted in internal structures for at least 48 h, arguing against rapid lysosomal turnover. To minimize artefacts from parasite deterioration, we deliberately restricted co-culture assays to up to 2 h and confirmed that trophozoites remained highly motile and morphologically intact in our imaging fields under these conditions, making widespread lysis unlikely. Nonetheless, we cannot absolutely exclude rare trophozoite damage or early rupture, which could contribute a minor background of passively released KERP2 superimposed on the contact-dependent uptake described here.
Host co-IP and transcriptomic analysis identify multiple pathways impacted by KERP2 exposure. Two functional modules emerge as most prominently affected: cell cycle regulation and cytoskeletal dynamics. On the signaling side, interaction with PPP6C and PPP6R suggest potential interference with PP6-linked control of DNA-damage responses and inflammatory signaling, processes with direct implications for cell cycle progression38,39. In parallel, interactions with PFN1, ARPC3, LASP1 and PDLIM1, together with associated transcriptional changes, support a role for KERP2 in modulating actin polymerization, focal adhesion organization, and epithelial junctional stability.
Collectively, these findings indicate that internalized KERP2 primarily functions within the host cytosolic and endosomal compartments to modulate cytoskeletal architecture, junctional organization, and signaling pathways, with co-enriched candidates requiring orthogonal validation. They do not yet provide evidence for a direct role in host chromatin regulation, which remains an open question requiring further investigation.
Functional assays demonstrate that KERP2 disrupts host cytoskeletal dynamics. When KERP2 is present and transferable, the dominant signature is epithelial retuning: cells become more elongated, cortical F-actin weakens, wound closure is impaired under reduced-proliferation conditions, and barrier properties shift in a way that affects both TEER and macromolecular flux. This constellation of phenotypes is consistent with KERP2 acting at the membrane–cytoskeleton–junction interface to reshape actin architecture and tight junction organization, rather than functioning primarily as a cytolytic factor. Association with candidate junctional and actin-associated partners reinforce this view and provide a mechanistic route by which a soluble parasite effector can reprogram epithelial mechanics and ion-selective barrier function.
In contrast, loss of KERP2 pushes the system into a protease-dominant regime. KERP2gs parasites show elevated cysteine-protease activity and cause the largest TEER loss, yet without a corresponding rise in 10-kDa dextran permeability. At the same time, host cells exposed to KERP2gs reinforce stress fibers and close wounds at least as efficiently as controls within the time window examined. This combination suggests a state of junctional and matrix stress driven by enhanced protease activity, overlaid with actin-based compensation that preserves size-selective exclusion of larger solutes despite substantial electrical leakage.
Within this framework, TEER alone is not a sufficient surrogate for barrier integrity: similar electrical readouts can reflect very different balances of ion conductance, macromolecular permeability, and cytoskeletal compensation. KERP2 therefore appears to bias E. histolytica toward a cytoskeleton- and junction-focused mode of barrier modulation when present, whereas its loss favors a more classical protease-driven insult that the epithelium partially counteracts through contractile remodeling.
Together, these findings support a working model in which changes in KERP2 availability and localization may couple parasite chromatin state to the choice of epithelial damage strategy in our experimental systems. In conditions where KERP2 is predominantly nuclear, virulence genes appear buffered and the parasite may rely more on KERP2 transfer to tune epithelial mechanics and ion transport. Conversely, when KERP2 is reduced or redistributed, this buffering may relax, protease output increases, and junctional degradation is a major contributor to TEER loss. Because our current host-model systems lack innate immune compartments, a next step will be to incorporate defined innate immune cell lineages into 3D crypt co-cultures and then test trophozoites in established models of intestinal amoebiasis and liver abscess. Future work (e.g., separation-of-function/rescue) dissecting how nuclear–cytoplasmic partitioning of KERP2 is regulated, and how this intersects with cysteine protease expression and host junctional targets, will be essential for determining how often each route is deployed in vivo and whether shifting this balance can mitigate tissue injury without compromising colonization. More broadly, the role of KERP2 in trophozoites development warrants further investigation, including whether KERP2 localization/expression fluctuates in a periodic manner during trophozoite development in vivo; our current asynchronous trophozoite assays do not resolve stage-dependent dynamics or the frequency of KERP2 transfer during infection.
Methods
Structural modeling, domain identification, and biochemical property analysis
The three-dimensional structure of KERP2 was modeled using I-TASSER40,41,42. Structural visualization and refinement were performed using PyMOL (The PyMOL Molecular Graphics System, Version 3.0 Schrödinger, LLC.). Secondary structural motifs, including a coiled-coil domain spanning amino acids 178–216, were predicted using Marcoil and PCOILS43,44,45,46. The SAP domain of KERP2 was manually aligned based on residue conservation patterns described in ref. 17. Automated alignment attempts using tools such as CLC Viewer (QIAGEN Bioinformatics, Aarhus, Denmark) and PSI-BLAST did not yield satisfactory results due to the weak sequence conservation of the SAP domain. To address this, residues in KERP2 were manually examined and aligned to known SAP domains following the consensus criteria outlined in the reference study. Key features used for the alignment included: hydrophobic or aliphatic residues (YFWLIVMA), small residues (SAGTVPNHD), polar residues (STQNEDRKH), and bulky residues (KREQWFYLMI). The alignment process was further verified by predicted secondary structure features by PHD program47. Nuclear localization signals (NLS) were mapped using the NLS Mapper tool48. The isoelectric point (pI) of KERP2 was calculated using ExPASy’s Compute pI/Mw tool49. Structural models from I-TASSER were converted to PQR with PDB2PQR and used as input to APBS to compute electrostatic potentials; APBS potentials are reported in kT/e50. ChimeraX-1.10.1 was used for visualization51.
Phylogenetic analysis
BLASTp and HMMER searches were performed to identify KERP2 orthologs across a broad range of taxa using KERP2 as a query. The obtained sequences were aligned using MAFFT v7.47552 and trimmed by TrimAl v1.453. Phylogenetic reconstruction was conducted using the maximum likelihood method in IQ-TREE v2.3.654 with default parameters. The resulting phylogenetic tree was visualized using iTOL v7.055. The ratio of nonsynonymous (dN) to synonymous (dS) substitutions (ω = dN/dS) was calculated using the codeml module in PAML v4.9.
Plasmid construction
Total RNA from E. histolytica trophozoites was extracted using the TRIzol reagent (Invitrogen) following the manufacturer’s protocol. Briefly, cells were lysed directly in TRIzol reagent, and the homogenized lysate was incubated at room temperature for 5 min to allow complete dissociation of nucleoprotein complexes. Chloroform (0.2 mL per 1 mL of TRIzol) was added to the lysate, vortexed vigorously for 15 s, and incubated at room temperature for 3 min. The mixture was centrifuged at 12,000 × g for 15 min at 4 °C to separate phases. The aqueous phase containing RNA was transferred to a new tube, and RNA was precipitated by adding isopropanol (0.5 mL per 1 mL of TRIzol), incubated at room temperature for 10 min, and centrifuged at 12,000 × g for 10 min at 4 °C. The RNA pellet was washed with 75% ethanol (1 mL per 1 mL of TRIzol) and centrifuged at 7500 × g for 5 min at 4 °C. After air-drying, the pellet was dissolved in RNase-free water and quantified using a NanoDrop spectrophotometer (ThermoFisher). Messenger RNA (mRNA) was purified, and cDNA synthesis was performed using the SuperScript III First-Strand Synthesis System (Invitrogen, ThermoFisher) according to the manufacturer’s protocol.
The protein-coding region of KERP2 was amplified by PCR from E. histolytica cDNA using specific oligonucleotides summarized in Supplementary Data 1. To express KERP2 fused with an HA tag at the amino terminus in E. histolytica trophozoites, PCR fragments were digested with XmaI and XhoI, purified, and ligated into XmaI- and XhoI-digested pEhExHA vector56 using cohesive ends, producing the plasmid pEhEx-HA-KERP2.
To generate deletion and fusion constructs:
pEhEx-HA-KERP2∆185−239: Reverse PCR was performed on pEhEx-HA-KERP2 using paired primers, followed by blunt-end ligation with T4 Polynucleotide Kinase and T4 DNA Ligase.
pEhEx-GFP-HA-KERP2: GFP was amplified from the pEhEx-GFP vector56 with BglII end, then ligated into pEhEx-HA-KERP2 digested with BglII.
pEhEx-GFP-HA-KERP2∆185−239: GFP was amplified from pEhEx-GFP and inserted into pEhEx-HA-KERP2∆185–239 same as described above.
Hygtet-HA-KERP2: the HA-KERP2 coding sequence was PCR-amplified from pEhEx-HA-KERP2 using primers bearing KpnI and BamHI sites and subcloned into pEhHygtetR-O-Cass that had been double-digested with KpnI/BamHI, yielding Hygtet-HA-KERP2. The pEhHygtetR-O-Cass vector was a kind gift from Dr. William A. Petri Jr..
To silence KERP2 in E. histolytica using antisense small RNA, a 420 base-pair fragment corresponding to the 5’ sequence of KERP2 was amplified by PCR from cDNA using appropriate primers. The amplified fragment was digested with StuI and SacI and cloned into StuI/SacI-digested psAP2-Gunma vector (Mi-Ichi et al., 2011), generating the plasmid psAP2-KERP2gs.
To express His-tagged KERP2 variants in E. coli, the KERP2 coding sequence was first subcloned into pEhEx-GFP using primers bearing XmaI and XhoI sites, following the same strategy used for pEhEx-HA-KERP2. The resulting pEhEx-GFP-KERP2 served as template to PCR-amplify the GFP–KERP2 open reading frame, which was inserted into pET-151 (N-terminal 6×His tag) using In-Fusion HD to generate pET-151-GFP-KERP2. A (Gly-Ala)_5 flexible linker (5×GA; XmaI-flanked; synthesized by FASMAC) was placed between GFP and KERP2 to reduce steric constraints. To obtain a GFP-only control, reverse (inverse) PCR on pET-151-GFP-KERP2 was used to excise the KERP2 insert, and the linear product was re-circularized by In-Fusion, yielding pET-151-GFP. The GFP–KERP2Δ185–239 mutant (deletion of amino acids 185–239) was generated from pET-151-GFP-KERP2 by removing the internal fragment between XmaI and XhoI and ligating the flanking regions to create the in-frame deletion; equivalently, the same junction was produced by inverse PCR with primers spanning the Δ185–239 boundary followed by T4 ligation.
All constructs were validated by Sanger sequencing to ensure accurate cloning. Primer sequences and synthetic oligonucleotides are listed in Supplementary Data 1.
Cell culture and transfection
Trophozoites of the E. histolytica strain HM-1:IMSS Cl657 and G358 were maintained axenically in Diamond’s BI-S-33 medium57 at 35.5 °C. Plasmids encoding HA-tagged and GFP-HA-fused KERP2 or its variants, generated as described above, were introduced into HM-1 or G3 trophozoites via lipofection, following the protocol established by ref. 59. Transfected trophozoites were selected and maintained in medium containing 10 μg/mL of G418 (#11811031, Gibco/Life Technologies, Waltham, MA, USA). Plasmids for gene silencing experiments were introduced into the G3 strain by the same lipofection method, and the transfectants were also maintained in medium supplemented with 10 μg/mL of G418. Protein expression was verified by western blotting as described below. For co-transfection experiments, Myc-EhRab11B (G418-selectable) was co-introduced with the Hygtet-HA-KERP2 construct; cells were selected and maintained under dual selection with G418, 10 µg/mL, plus hygromycin (Invitrogen, San Diego, CA, USA), 20 µg/mL. Protein expression was confirmed by western blotting.
Caco-2 cells (ATCC, HTB-37) were cultured in Eagle’s Minimum Essential Medium (EMEM; ATCC 30-2003) supplemented with 20% heat-inactivated fetal bovine serum (FBS; Sigma-Aldrich, St. Louis, MO, USA.) as complete EMEM. The medium was replaced every two days, and cells were incubated under standard cell culture conditions at 37 °C in a humidified atmosphere with 5% CO₂. The Caco-2 cell line (ATCC, HTB-37) is not listed in the ICLAC register of misidentified cell lines; no commonly misidentified cell lines were used in this study.
Establishment of 3D crypt model
Primary human colonic crypts were isolated from sigmoid colon biopsy tissue collected under an approved protocol (University of Tokyo IRB 2022115NI; informed consent obtained). Although biopsies were obtained from three consenting adult patients, only one donor-derived sample was used to generate the 3D crypt cultures analyzed in this study. Crypt isolation and culture were performed as described previously60 with minor modifications. Biopsy tissue was incubated in crypt isolation buffer containing EDTA and dithiothreitol (DTT) to release crypts. The suspension was centrifuged (600 × g, 1 min), and the pellet was resuspended in pre-warmed maintenance medium (MM) at 240 crypts/mL. Crypts were seeded into 6-well plates pre-coated with neutralized collagen hydrogels (pH 7.4) and allowed to polymerize for 1 h at 37 °C in a humidified CO₂ incubator. Cultures were maintained at 37 °C with MM changes every 48 h until ~80% confluency, then passaged by mechanical fragmentation using a double–pipette-tip method and reseeded into fresh collagen-coated scaffolds. To generate 3D crypt arrays, micromolded collagen scaffolds were fabricated using PDMS molds to recapitulate intestinal crypt architecture. Crypts were seeded into the scaffolds in differentiation medium supplemented with 10 µM Y-27632. Media were changed every 48 h. Differentiation was induced by establishing a growth-factor gradient across the scaffold: WRN medium was supplied to the basal reservoir and differentiation medium to the luminal reservoir for 4 days to promote formation of a crypt–villus axis. Differentiated 3D crypts were then used for imaging and functional assays.
Preparation of recombinant proteins
Plasmids pET-KERP2, pET151-GFP-KERP2, and pET151-GFP, designed to express recombinant His-GFP-KERP2 and His-GFP proteins, respectively, were introduced into E. coli BL21 (DE3) cells by heat shock at 42 °C for 1 min. Transformed E. coli cells were initially grown in 100 mL of Luria Bertani (LB) medium supplemented with 100 μg/mL carbenicillin at 37 °C. The overnight culture was then used to inoculate 500 mL of fresh LB medium, and the cells were grown at 37 °C with shaking at 200 rpm. When the optical density at 600 nm (A600) reached 0.4, the cultures were chilled on ice, and 2% ethanol and 1 mM isopropyl β-D-thiogalactopyranoside (IPTG) were added to induce protein expression. Cultivation was continued for an additional 16 h at 18 °C. Induced E. coli cells were harvested by centrifugation at 8000 × g for 10 min at 4 °C. The cell pellets were washed with Tris-saline buffer (10 mM Tris-HCl, pH 7.6, and 150 mM NaCl) and resuspended in 40 mL of lysis buffer containing 50 mM Tris-HCl (pH 7.6), 250 mM NaCl, 10% (w/v) sucrose, 100 μg/mL lysozyme, 0.5 μg/mL E64, 1× cOmplete Mini Protease Inhibitor Cocktail (Roche), and 1 mM phenylmethylsulfonyl fluoride (PMSF). The suspension was incubated at room temperature for 20 min and lysed using a French press. Triton X-100 was added to a final concentration of 0.1% (v/v), and the lysate was centrifuged at 25,000 × g for 30 min at 4 °C to remove cell debris. The clarified supernatant was incubated with 1 mL of 50% Ni²⁺-NTA His-Bind resin slurry at 4 °C for 1 h with gentle rotation. The resin was transferred to a column and washed three times with wash buffer containing 50 mM Tris-HCl (pH 7.6) and 2 M KCl. Bound proteins were eluted using elution buffer containing 50 mM Tris-HCl (pH 7.6), 150 mM NaCl, 10% (v/v) glycerol, and 10–750 mM imidazole. The integrity and purity of the recombinant proteins were confirmed by 12% SDS-PAGE, followed by Coomassie Brilliant Blue staining or western blot analysis.
Electrophoretic mobility shift assay (EMSA)
EMSA was performed using the Gelshift Chemiluminescent EMSA Assay Kit (#37341, Active Motif) according to the manufacturer’s instructions with minor adaptations. Native separations were carried out on Novex DNA Retardation Gels (6%, 1.0-mm, 15-well; Thermo Fisher) in 0.5× TBE. The following 60-mer oligonucleotides were synthesized; the sense strands for the AT-rich and GC-rich probes were 5’-biotinylated:
-
AT-rich sense (biotinylated): 5’-TTAATTAAATAAATTAAAAAATTTTTTAATTAAATTAATTAATAAATTAAAATAAATTTTTTAATTAAATTTTAA-3’
-
AT-rich antisense (for duplex): 5’-TTAAAATTTAATTAAAAAATTTATTTTAATTTATTAATTAATTTAATTAAAAAATTTTTTAATTTATTTAATTAA-3’
-
GC-rich sense (biotinylated):
5’-CGGGGCGCCGCGGGGCCCCCGGCCCGCCCCGCGGGGCGCCGGGCCCCGCC-3’
-
GC-rich antisense (for duplex): 5’-GGCGGGGCCCGGCGCCCCGCGGGGCGGGCCGGGGGCCCCGCGGCGCCCCG-3’
For double-stranded (ds) probes, same estimated amount of sense and antisense oligos were mixed, heated to 95 °C for 5 min, and slow-cooled to room temperature. Single-stranded (ss) probes were the biotinylated sense strands alone. Probe input per reaction was typically 60 fmol. Recombinant proteins were buffer exchanged to remove imidazole. Reactions (total 20 µl) were assembled on ice in this order: 2 µl 10× Binding Buffer (kit; final 1×), 1 µl 50% glycerol (final 2.5%), 1 µl 100 mM MgCl₂ (final 5 mM), 1 µl poly(dI·dC) (1 µg/µl stock; final 50 ng/µl), biotinylated DNA probe, recombinant protein (His-GFP-KERP2 or His-GFP-KERP2Δ185–239 at 4, 10, 20, or 40 fmol), then nuclease-free water to 20 µl. For competition EMSA, unlabeled AT competitor (ss) was added at 1× the labeled probe and pre-incubated with protein for 10 min at room temperature before adding the probe. All reactions were incubated 20 min at room temperature. Samples were then added with loading buffer and loaded directly onto 6% DNA Retardation Gels and electrophoresed at 120 V (room temperature) in 0.5× TBE. DNA was transferred to a Biodyne B Nylon Membrane (0.45 μm; Thermo Fisher), UV-crosslinked, and detected by streptavidin–HRP with chemiluminescence according to the Active Motif protocol.
Chromatin immunoprecipitation (ChIP) and ChIP–seq analysis
Chromatin immunoprecipitation (ChIP) was performed on E. histolytica expressing GFP–HA–KERP2. Cells from ten 10-cm dishes were cross-linked in PBS with 0.5% formaldehyde for 10 min at room temperature (RT) and quenched with 125 mM glycine for 5 min. Cells were chilled on ice, collected (500 × g, 10 min, 4 °C), washed once in ice-cold PBS, and nuclei were prepared by incubation in nuclei isolation buffer (10 mM HEPES–KOH pH 7.9, 1.5 mM MgCl₂, 10 mM KCl, protease inhibitors: PMSF 0.5 mM, E-64 1 μM, Complete Mini 1×) for 20 min on ice, followed by addition of NP-40 to 2% (v/v) and rotation for 20 min at 4 °C. Nuclei were pelleted (1,200 rpm), washed in nuclei washing buffer (10 mM Tris-HCl pH 8.0, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, + protease inhibitors), and resuspended in sonication buffer (50 mM Tris-HCl pH 8.0, 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate, 5 mM DTT, + protease inhibitors); chromatin was sheared by Covaris S2 at intensity 4 for 15 cycles to enrich ~200–500 bp. An input aliquot was reserved. For immunoprecipitation, clarified chromatin (16,000 × g, 20 min, 4 °C) was incubated overnight at 4 °C with pre-conjugated Protein A Dynabeads (Thermo Fisher Scientific, Waltham, MA, USA) carrying anti-GFP (ab290, Abcam, Cambridge, UK, 6 μL per sample) in ChIP Dilution Buffer (16.7 mM Tris-HCl pH 8.0, 167 mM NaCl, 1.2 mM EDTA, 1% Triton X-100, 0.01% SDS; supplemented with protease inhibitors and BSA). Beads were washed five times in cold low-salt wash buffer (20 mM Tris-HCl pH 8.0, 150 mM NaCl, 2 mM EDTA, 0.1% SDS, 1% Triton X-100) and twice in high-salt wash buffer (20 mM Tris-HCl pH 8.0, 500 mM NaCl, 2 mM EDTA, 0.1% SDS, 1% Triton X-100), and DNA–protein complexes were eluted in ChIP elution buffer (ChIP EB) (10 mM Tris-HCl pH 8.0, 300 mM NaCl, 5 mM EDTA, 1% SDS) for 20 min at RT. Cross-links (ChIP and input) were reversed at 65 °C for 8 h, followed by RNase (37 °C, 1 h) and Proteinase K (55 °C, 2 h) treatments. DNA was purified by UltraPure phenol:chloroform:isoamyl alcohol (Thermo Fisher Scientific) with back-extraction in TE/NaCl (TE + 200 mM NaCl), ethanol-precipitated ( − 30 °C, 6 h), washed in 80% ethanol, and resuspended (ChIP 20 μL; input 40 μL); DNA was quantified by Qubit dsDNA HS (Thermo Fisher Scientific). For ChIP-seq, reads were adapter-trimmed and aligned to the Entamoeba reference with Bowtie2/BWA; uniquely mapped reads (MAPQ ≥ 30) were retained, PCR duplicates were removed, and peaks were called with MACS2 using matched input. Reproducible binding sites were defined as the intersection of peaks across three biological trials and used for downstream analyses. De novo motif discovery on ±200-bp windows around reproducible peaks was performed with HOMER (repeat-masked, default GC/AT-matched background), and background-aware sequence logos were generated from PWMs using genome-wide base frequencies qb with letter heigh \({p}_{b}{\log }_{2}\left(\frac{{p}_{b}}{{q}_{b}}\right)\,\); positional specificity was evaluated by summit-centered motif density.
Multimerization
Size-exclusion chromatography (SEC) was performed on a Cytiva ÄKTA go fitted with a Superdex 200 Increase 10/300 GL column (24-mL bed volume) at RT using 50 mM Tris-HCl (pH 7.6), 150 mM NaCl as running buffer (0.22 µm filtered, degassed). The column was equilibrated with ≥2 column volumes and run at 0.50 mL min⁻¹, collecting 0.5-mL fractions across 1.2 CV. For calibration, a gel-filtration standard (BioRad) was run on the same column/buffer. The elution volume (Ve, mL) of each standard was plotted directly against log10(MW in kDa) and fit by linear regression, yielding the relationship.
Purified recombinant proteins (His-GFP–KERP2 and His-GFP–KERP2Δ185–239) were exchanged into running buffer overnight (Slide-A-Lyzer Dialysis Cassette; 3.5 K MWCO; Thermo Fisher), clarified at 14,000 × g for 10 min at 4 °C, and 0.22 µm filtered prior to injection. Typical loads were 100–200 µL at 3.5–4 mg mL⁻¹ (500 µg total). Elution was monitored at A280; fractions were analyzed by SDS-PAGE/Coomassie.
For amebic inputs (HA-KERP2 and HA-KERP2Δ185–239), crude nuclear and cytosolic fractions were prepared as follows: trophozoites from 15 dishes were collected in PBS and pelleted; the pellet was weighed and resuspended in HEPES + + buffer (10 mM HEPES, pH 7.6; 10 mM KCl; 1.5 mM MgCl₂; 1× cOmplete Mini protease inhibitors; 0.14 mM E64 protease inhibitor) at 0.75 mL per 0.1 g cells. Suspensions were incubated on ice for 1 h with gentle mixing every 20 min until translucent, then NP-40 was added to 2% final and incubated 6 min on ice. The sample was centrifuged (150 x g, 10 min; 4 °C) to yield supernatant (cytosol) and pellet (nuclei). Nuclear pellets were washed 3× in PBS at the same volume used for HEPES + + (each wash 2400 rpm, 5 min, 4 °C), then resuspended in lysis buffer at 50 µL per 0.1 g starting pellet to generate the nuclear fraction. Both fractions were exchanged into SEC running buffer by Amicon Ultra Centrifugal Filter (10 kDa MWCO; MERCK) and 0.22 µm-filtered before injection. Elution was monitored at A280; fractions were analyzed by SDS-PAGE/Coomassie.
Immunofluorescence assays (IFA)
Immunofluorescence assays were performed to analyze the localization of KERP2 and its variants in E. histolytica or mammalian cells.
E. histolytica cells were fixed in 4% paraformaldehyde (PFA) in phosphate-buffered saline (PBS) for 10 min at room temperature, followed by permeabilization and blocking with 0.2% saponin and 1% bovine serum albumin (BSA) in PBS for 10 min. Cells were incubated with primary antibodies, including anti-HA (1:200 dilution, clone 16B12; AB_2820200, BioLegend), anti-CS1 (1:500 dilution, In-house antibody), or anti-EhRab11B (1:500 dilution, In-house antibody), for 1 h at room temperature. After PBS washes, samples were stained with Alexa Fluor™ 488 Goat anti-Mouse IgG (H + L) (AB_141607, Thermo Fisher Scientific, Waltham, MA, USA) and Alexa Fluor™ 594 Goat anti-Rabbit IgG (H + L) (AB_141637, Thermo Fisher Scientific, Waltham, MA, USA) at 1:1000 dilution, along with Hoechst 33342 (1:8000 dilution; Thermo Fisher Scientific) for 1 h at room temperature in the dark.
Caco-2 monolayers, cultured for 21 days on Millicell EZ slides (Merck Millipore, Burlington, MA, USA), were pre-labeled with CellTracker™ Blue (Thermo Fisher Scientific) prior to co-culture, and E. histolytica HA-KERP2-expressing strains were pre-labeled with CellTracker™ Red (Thermo Fisher Scientific), according to the manufacturers’ instructions. Labeled Caco-2 monolayers were then co-cultured with the labeled parasites for 2 h. Following interaction, cells were fixed with 4% PFA in PBS for 20 min, then permeabilized and blocked with 0.2% saponin, 1% BSA, and 50 mM glycine in PBS for 2 h. Samples were incubated overnight at 4 °C with anti-HA (1:200 dilution, 16B12; AB_2820200, BioLegend). After washing, cells were stained with Alexa Fluor™ 488 Goat anti-Mouse IgG (H + L) (AB_141607, Thermo Fisher Scientific) at 1:1000 dilution and DRAQ5™ (#65-0880-92, Invitrogen) for nuclear staining for 1 h at room temperature in the dark.
E. histolytica GFP-HA-KERP2 strains were pre-stained with CellTracker Deep Red (Thermo Fisher Scientific) in Opti-MEM (Thermo Fisher Scientific) for 1 h. Parasites were seeded onto the 3D-crypt enterocyte model and allowed to interact for 2 h. Immunofluorescence staining followed the same protocol as for Caco-2 cells.
For Caco-2 morphology observation after treatment with E. histolytica transformants, the same IFA process was used. Primary antibody anti-E-cadherin (1:1000 dilution, clone 3195; AB_2291550, Cell Signaling Technology, Danvers, MA, USA) and secondary antibody Alexa Fluor™ 488 Goat anti-Rabbit IgG (H + L) (AB_143165, Thermo Fisher Scientific) were used together with Alexa Fluor™ 594 Phalloidin (1:400 dilution; Thermo Fisher Scientific) and Hoechst 33342 (1:8000 dilution; Thermo Fisher Scientific).
Imaging was performed using an FV3000 confocal microscope (EVIDENT) for E. histolytica and Caco-2 cells, and a Dragonfly High-Speed Confocal Microscope System (Oxford Instruments, Abingdon, Oxfordshire, UK) for 3D-crypt enterocytes. Image processing and line intensity profile analyses were conducted using Imaris 10.2 and Fiji software61. For colocalization analysis, nuclei were segmented to create a nuclear mask. This mask was added to the ROI Manager and inverted (inverse selection) to produce a nucleus-excluded ROI encompassing the cytoplasm. Colocalization was run in Fiji/Coloc2 (Costes auto-thresholds; 100 randomizations; PSF = 2 px) on the nucleus-excluded ROI.
Live imaging
Live-cell imaging was conducted to observe the localization and dynamics of GFP-HA-tagged KERP2 constructs in real time. For imaging, cells were pre-stained with Hoechst 33342 (Thermo Fisher Scientific) in Opti-MEM (Thermo Fisher Scientific) for 40 min and maintained in complete BI-S33 medium at 35.5 °C. Live cells were plated on glass-bottom dishes and allowed to settle for 30 min before imaging. Imaging was performed using an LSM 780 confocal microscope (Carl Zeiss Microscopy, White Plains, NY, USA) equipped with a temperature-controlled chamber set to 35.5 °C and a 63× oil-immersion objective. Time-lapse images were captured continuously for up to 5 min.
For KERP2 transfer studies, 21-day cultured Caco-2 monolayers established on glass-bottom dishes were pre-stained with Hoechst 33342 (Thermo Fisher Scientific) in EMEM (ATCC) for 40 min. GFP-HA-KERP2-expressing E. histolytica was added to the Caco-2 monolayers and incubated for 30 min. Live imaging was performed as described above, with time-lapse images captured continuously for up to 30 min. Z-stack scanning was performed following time-lapse imaging.
Image acquisition and analysis were conducted using Imaris 10.2 (Oxford Instruments) and Fiji software.
Immuno-electron microscopy
Trophozoites overexpressing HA-KERP2 and HA-mock were cultured in BIS medium and incubated with gold disks at 35.5 °C overnight to facilitate attachment. The disks carrying attached amoebae were rapidly frozen in liquid propane at −175 °C. Freeze substitution was carried out in 2% tannic acid in ethanol with 2% distilled water at −80 °C for 48 h. The samples were then gradually warmed to −20 °C for 4 h and subsequently to 4 °C for 1 h. Dehydration was performed with three changes of anhydrous ethanol at 4 °C for 30 min each. Dehydrated samples were infiltrated with a 50:50 mixture of ethanol and resin (LR White; London Resin Co. Ltd., Berkshire, UK) at 4 °C for 30 min, followed by three changes of 100% LR White resin at 4 °C for 30 min each. Samples were transferred to fresh 100% resin and polymerized at 50 °C overnight. Polymerized samples were sectioned into ultra-thin slices (70 nm) using a diamond knife on an ultramicrotome (Ultracut UCT; Leica Microsystems, Vienna, Austria). Sections were mounted on nickel grids and incubated overnight at 4 °C with a primary antibody, anti-HA (1:500 dilution, clone 16B12; AB_2820200, BioLegend), diluted in 1% BSA/PBS. Grids were washed three times with 1% BSA/PBS for 1 min each, then incubated for 2 h at room temperature with a secondary antibody conjugated to 15 nm gold particles (goat anti-mouse IgG pAb). After secondary antibody incubation, grids were washed with PBS and fixed with 2% glutaraldehyde in 0.1 M phosphate buffer. Finally, grids were dried and stained with 2% uranyl acetate for 10 min, followed by lead stain solution (Sigma-Aldrich Co., Tokyo, Japan) at room temperature for 3 min. Samples were observed using a transmission electron microscope (JEM-1400Plus; JEOL Ltd., Tokyo, Japan) operating at an acceleration voltage of 100 kV. Digital images (3296 × 2472 pixels) were captured using a CCD camera (EM-14830RUBY2; JEOL Ltd., Tokyo, Japan).
qRT-PCR and RNA-seq
Total RNA was extracted from KERP2gs or psAP-mock trophozoites, and complementary DNA (cDNA) was synthesized as described in previous methods. The efficiency of KERP2 silencing in trophozoites was validated using quantitative real-time PCR (qRT-PCR). RNA polymerase II (EHI_056690) was used as the internal control. Primers were designed to amplify a 493 bp segment of KERP2 and a 204 bp segment of RNA polymerase II. Twenty-fold diluted cDNA from each strain was used as a template in reactions performed with the Fast SYBR Green Master Mix (Applied Biosystems, Foster City, CA, USA), following the manufacturer’s protocol. qRT-PCR was conducted on a StepOne Plus Real-Time PCR System (Applied Biosystems), and data analysis was performed using the DataAssist software (Thermo Fisher Scientific). RNA-seq was performed on an Illumina NovaSeq 6000 platform (Fujifilm, Japan) similarly as described below.
For co-culture experiments, total RNA was extracted from Caco-2 cells interacting with HA-KERP2-expressing E. histolytica G3 strain, psAP-KERP2gs trophozoites, or wild-type G3 trophozoites after 4 h of co-culture in complete EMEM medium. RNA integrity was assessed using an Agilent 2100 Bioanalyzer, and sequencing libraries were prepared using a strand-specific RNA-seq library preparation kit. Libraries were sequenced on an Illumina NovaSeq X Plus (Novogene, China), generating paired end reads. Sequence reads were quality-checked using FastQC (v0.11.8) and trimmed using Trimmomatic (v0.39)62. Trimmed reads were aligned to the E. histolytica reference genome assembly (AmoebaDB-68) or the human reference genome (GRCh38.p14) using STAR (v2.7.10a)63 with default parameters and gene annotation provided in the corresponding GTF file. Mapped reads were indexed and sorted using Samtools (v1.15.1)64. Read count quantification at the gene level was performed using FeatureCounts (v2.0.1)65 with strand-specific parameters. Differential gene expression analysis was conducted using the DESeq2 R package (v1.36.0)66, with three biological replicates per condition. Gene expression normalization was performed using DESeq2’s median-of-ratios method. Log2 fold changes and adjusted P-values were computed using the Benjamini-Hochberg method to control the false discovery rate (FDR). Genes with an adjusted P-value (padj) ≤ 0.05 and an absolute log2 fold change ≥1 were classified as differentially expressed. Principal component analysis (PCA) and hierarchical clustering were performed to assess sample variance. Volcano plots and heatmaps were generated using ggplot2 (v3.5.1), ggrepel (v0.9.5), and pheatmap (v1.0.12) to visualize differentially expressed genes across conditions. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were conducted using clusterProfiler (v4.2.2)67 or ShinyGO (v0.82)68 to identify functional categories associated with differentially expressed genes. For each contrast, genes were ranked by the DESeq2 Wald statistic (signed, continuous; universe = genes expressed in Caco-2 under any condition, e.g., baseMean > 10). Pre-ranked GSEA was performed in clusterProfiler (v4.2.2) using GO Molecular Function, GO Biological Process, and KEGG gene sets (human; annotations from org.Hs.eg.db and KEGG on the analysis date). Parameters were nPerm = 1000, minGSSize = 10, maxGSSize = 500, and pAdjustMethod = “BH”. Enrichment was summarized as ES and NES with FDR q-values; gene sets with FDR q ≤ 0.25 (GSEA convention) were considered enriched, and where indicated we additionally report FDR q ≤ 0.05. Leading-edge subsets were extracted from the running-sum statistic. Visualization used enrichplot (ridgeplots, running-sum curves, cnetplots) and ggplot2/ggrepel for dot plots and labeling, with all plots generated from the same preranked lists to ensure comparability across contrasts.
Growth kinetics
Approximately 3 × 10⁴ exponentially growing trophozoites of the E. histolytica G3 strain, transformed with either psAP-KERP2gs or psAP-mock (control), were inoculated into 6 mL of fresh BI-S-33 medium supplemented with 10 μg/mL G418. The cultures were incubated under standard conditions, and trophozoites were counted every 24 h using a hemocytometer to assess growth over time.
Cysteine protease (CP) activity assay
Approximately 1.2 × 10⁶ E. histolytica transformants were cultured in 500 μL of Transfection Medium (Opti-MEM supplemented with 5 mg/mL L-cysteine and 1 mg/mL ascorbic acid) at 35.5 °C for 1 h. After incubation, cells were aliquoted into 50 μL samples and centrifuged at 3000 × g for 3 min. The supernatant was subsequently centrifuged at 13,000 × g for 5 min to collect the released form of cysteine protease (CP) activity. The cell pellet was lysed in 100/200 μL (among trials) of PBS using three freeze-thaw cycles, and cellular debris was removed by centrifugation at 15,000 × g for 5 min. The resulting supernatant from this step was collected as the intracellular form of CP activity. Both released and intracellular samples were pre-incubated in assay buffer (0.1 M KH₂PO₄, pH 6.1, 1 mM EDTA, 2 mM DTT) at room temperature for 15 min to activate pro-forms of CP. Following activation, 75 μL of the enzyme mixture was combined with Benzyloxycarbonyl-L-arginyl-L-arginine 4-methylcoumaryl-7-amide (#3123-v, Peptide Institute, Osaka, Japan) to achieve a final substrate concentration of 10 mM. Fluorescence emission at 460 nm (excitation at 355 nm) was measured for 1 h using a SpectraMax Paradigm multimode microplate reader (Molecular Devices, San Jose, CA, USA). 7-amino-4-methylcoumarin (#3099-v, Peptide Institute) served as the calibration standard to quantify specific cysteine protease (CP) activity. Lysate protein concentration was determined and used to normalize raw fluorescence. For reporting, normalized CP activities were expressed relative to the psAP-mock control (psAP-mock set to 100% within each experiment).
Nuclear fractionation and co-immunoprecipitation
Nuclear protein fractions were prepared from approximately 2 × 10⁷ cells of E. histolytica HA-KERP2, HA-KERP2∆185−239, and HA-mock strains. Prior to nuclear fractionation, cells were cross-linked with 0.8 mg/mL Pierce dithiobis (succinimidyl propionate) (DSP) (Thermo Fisher Scientific) in PBS to stabilize protein-protein interactions. Nuclear protein extraction was conducted using the Nuclear Complex Co-IP Kit (#54001, Active Motif, Carlsbad, CA, USA) according to the manufacturer’s protocol. Extracted nuclear fractions were treated with the enzymatic cocktail reagent included in the kit, followed by nucleic acid digestion to degrade DNA and RNA, ensuring a focus on protein-protein interactions. The nuclear extracts were diluted in the IP low buffer provided in the kit, supplemented with protease and phosphatase inhibitors. Cytosolic fractions were also prepared for comparative analysis. Diluted nuclear and cytosolic fractions were incubated separately with anti-HA antibody (clone 16B12; AB_2820200, BioLegend) overnight at 4°C. Antibody-bound complexes were captured by incubating the samples with 45 μL of pre-washed Dynabeads™ Protein A (Thermo Fisher Scientific) for 1 h at 4 °C. To minimize non-specific interactions, beads were washed three times with 500 μL of IP low buffer containing BSA, followed by three additional washes with 500 μL of IP low buffer without BSA. Bound proteins were eluted by incubating the beads with 0.3 mg/mL HA peptide (#I2149, Sigma-Aldrich) overnight at 4 °C. Eluted samples were collected for downstream phosphorylation validation, western blot analysis, or mass spectrometry-based identification. For reciprocal IP, total lysates were prepared from the corresponding trophozoites (no subcellular fractionation), clarified by centrifugation, and incubated overnight at 4 °C with anti-Myc (9E10, #ab32, AB_303599, Abcam; dilution 1:1000). Immune complexes were captured on Dynabeads Protein A for 1 h at 4 °C, washed as above, and eluted in SDS sample buffer. Eluted material was analyzed by immunoblot using anti-HA (for co-precipitated HA-KERP2) and anti-Myc ab32 (for IP verification).
Extracellular vesicle isolation
Approximately 50 mL of serum-deprived culture medium from E. histolytica transformants was collected and centrifuged at 1000 × g for 10 min at 4 °C to remove intact cells. The resulting supernatant was filtered through a 0.22 μm membrane using a 50 mL syringe to eliminate larger debris. The filtered medium was then concentrated using a 100 kDa MWCO Amicon filter (Merck Millipore) at 4000 × g for 15 min at 4 °C. The concentrated cell-free medium was transferred to an ultracentrifuge tube and subjected to ultracentrifugation at 100,000 × g for 75 min at 4 °C to pellet extracellular vesicles (EVs), including exosomes. The EV pellet was washed with 1× PBS, followed by a repeat of the ultracentrifugation step under identical conditions. After discarding the supernatant, the final EV pellet was resuspended in 50 μL of 1× PBS. For EV lysis, 1% Triton X-100 was added to the resuspended pellet, and the mixture was incubated on ice for 30 min.
Protein–lipid overlay assays
The soluble fraction of recombinant His-KERP2 or His-GFP was obtained as described above and used to probe a P-6001 phospholipid membrane strip (Echelon Biosciences, Salt Lake City, UT, USA). The membrane was blocked with 3% bovine serum albumin (BSA) in phosphate-buffered saline (PBS) for 2 h at room temperature (RT) before incubation with 1 μg/mL of recombinant protein in PBS containing 3% BSA for 16 h at 4 °C. Following incubation, the membrane was washed three times with PBS containing 0.1% Tween-20 (PBS-T) and then incubated with an anti-His primary antibody (His-Tag (27E8) Mouse mAb #2366, Cell Signaling Technology, 1:1000) in PBS with 3% BSA for 2 h at RT. After additional washes, the membrane was incubated with HRP-conjugated goat anti-mouse IgG secondary antibody (Cat# 32430, Invitrogen, Thermo Fisher Scientific, 1:5000) in PBS with 3% BSA for 1 h at RT. The blot was washed, developed using a chemiluminescent HRP substrate (Merck Millipore), and visualized using a ChemiDoc™ Imaging System (Bio-Rad Laboratories, Hercules, CA, USA).
Sample preparation for KERP2 translocation in Caco-2 cells
Confluent Caco-2 cells cultured in T75 flasks ( ~ 5 × 10⁶ cells per flask) were co-cultured with E. histolytica HA-KERP2 strains at a 5:1 ratio (Caco-2: E. histolytica) for 1, 2, and 3 h in complete EMEM (ATCC). For control experiments, E. histolytica HA-mock strains were co-cultured with Caco-2 cells for 1 h. An additional experimental group involved pretreatment of E. histolytica HA-KERP2 strains with 2% galactose in complete Diamond’s BI-S-33 medium for 30 min prior to co-culture. During the interaction for this group, 2% galactose was also added to the complete EMEM for a 1-h incubation. After co-culture, the mixtures were washed three times with pre-cooled 2% galactose in PBS (Thermo Fisher Scientific) to remove E. histolytica cells. The remaining Caco-2 cells were collected, while E. histolytica cells were processed separately. Both cell types were lysed using M-PER Mammalian Protein Extraction Reagent (Thermo Fisher Scientific), supplemented with cOmplete™ Protease Inhibitor Cocktail (Roche). The lysates were subjected to western blotting for the detection of KERP2 translocation. Anti-HA antibody (ab9110, Abcam, Cambridge, UK; dilution 1:1000) was used to enhance detection sensitivity during western blot analysis.
Caco2 co-immunoprecipitation
Confluent Caco-2 cells grown in T75 flasks ( ~ 5 × 10⁶ cells per flask) were used to interact with E. histolytica HA-KERP2 or HA-mock strains at a 5:1 ratio (Caco-2: E. histolytica) for 1 h. Following interaction, six flasks of co-cultures were washed three times with pre-cooled 2% galactose in PBS to remove E. histolytica cells. The remaining Caco-2 cells were harvested using a cell scraper. The collected Caco-2 cells were cross-linked with 0.8 mg/mL Pierce DSP (ThermoFisher Scientific) in PBS for stabilization of protein-protein interactions. After cross-linking, cells were lysed using M-PER Mammalian Protein Extraction Reagent (ThermoFisher Scientific) supplemented with a cOmplete protease inhibitor cocktail (Roche). Lysates were incubated with anti-HA antibody (clone 11MO; Covance) overnight at 4 °C. The antibody-bound complexes were captured by incubation with 40 μL of pre-washed Dynabeads Protein A (Invitrogen, ThermoFisher) for 1 h at 4 °C. Beads were washed three times with Washing Buffer (50 mM Tris-HCl, 150 mM NaCl, 0.1% Triton X-100, 0.1% BSA) and three additional times with Washing Buffer lacking BSA to minimize non-specific interactions. Bound proteins were eluted by incubating the beads with 0.3 mg/mL HA peptide (Sigma-Aldrich) overnight at 4 °C. Eluted samples were collected and prepared for downstream analyses.
Flamingo gel staining
Co-IP samples were separated by SDS-PAGE as described in the relevant methods sections. Following electrophoresis, gels were subjected to Flamingo fluorescent gel staining (Bio-Rad Laboratories) to visualize protein elution in nuclear co-IP and Caco-2 co-IP pull-down samples. The staining was performed according to the manufacturer’s protocol. Stained gels were imaged using a ChemiDoc imaging system (Bio-Rad Laboratories) to detect protein bands.
Western blotting
Protein samples were separated by SDS-PAGE or Phos-tag™ SDS-PAGE (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) and transferred onto PVDF membranes (Merck Millipore) using a semi-dry transfer system. Membranes were blocked with 5% non-fat milk in TBST (20 mM Tris-HCl, pH 7.4, 150 mM NaCl, 0.1% Tween-20) for 30 min at room temperature. Blocked membranes were incubated overnight at 4°C with primary antibodies, including anti-HA (1:1000, 16B12; AB_2820200, BioLegend), anti-CS1 (1:1000, in-house source), anti-GAPDH (1:5000, AM4300, Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA), anti-Histone H3 (1:1000, ab1791, Abcam), Anti-Myc (1:1000, 9E10, #ab32, AB_303599, Abcam) or anti-His (1:1000, #2366, Cell Signaling Technology, Danvers, MA, USA). After primary antibody incubation, membranes were washed three times with TBST and then incubated with HRP-conjugated secondary antibodies for 1 h at room temperature. Secondary antibodies used included Donkey anti-Rabbit IgG (H + L) Cross-Adsorbed, HRP (31458, Invitrogen, Thermo Fisher Scientific) and Goat anti-Mouse IgG (H + L), HRP (32430, Invitrogen, Thermo Fisher Scientific) in a dilution 1:10000. Following secondary antibody incubation, membranes were washed three more times with TBST. Signals were detected using a chemiluminescence HRP substrate (Merck Millipore) and visualized using a ChemiDoc™ Imaging System (Bio-Rad Laboratories).
Mass spectrometry
Protein extracts were buffer-exchanged using SP3 paramagnetic beads (Cytiva, Marlborough, MA, USA). Samples were rehydrated in 100 μL of 10 mM triethylammonium bicarbonate (TEAB) containing 1% SDS. Disulfide bonds were reduced with 10 μL of 50 mM dithiothreitol (DTT) at 60°C for 1 h. After cooling to room temperature (RT), the pH was adjusted to approximately 7.5, and alkylation was performed by adding 10 μL of 100 mM iodoacetamide (Sigma-Aldrich) and incubating in the dark at RT for 15 min. SP3 beads (100 μg; 2 μL of 50 μg/μL) were added to the samples, followed by the addition of 120 μL of 100% ethanol. Samples were incubated at RT with shaking for 5 min to bind proteins to the beads. Beads were then washed three times with 180 μL of 80% ethanol. On-bead protein digestion was performed overnight at 37 °C using trypsin (Pierce Trypsin, Thermo Fisher Scientific, Waltham, MA, USA; 1 μg enzyme per sample). After digestion, the supernatant was removed, dried, and rehydrated in 2% acetonitrile/0.1% formic acid for mass spectrometry analysis. Peptide analysis was performed by reverse-phase chromatography-tandem mass spectrometry (LC-MS/MS) using an EasyLC 1100 UPLC system interfaced with an Orbitrap Fusion Lumos mass spectrometer (Thermo Fisher Scientific). Peptides were separated on a 75 μm × 15 cm column (PicoFrit Self-pack emitter, New Objective, Woburn, MA, USA) packed in-house with ReproSIL-Pur-120-C18-AQ resin (3 μm, 120 Å bulk phase, Dr. Maisch GmbH, Ammerbuch, Germany). Separation was achieved using a 0–100% acetonitrile gradient in 0.1% formic acid over 90 min at a flow rate of 300 nL/minute. Survey scans of precursor ions were acquired over a range of 350–1800 m/z with a resolution of 120,000 at 200 m/z, an automatic gain control (AGC) target of 4 × 10⁵, and an RF lens setting of 45%. Internal mass calibration was used for accuracy. Precursor ions were individually isolated within a 1.5 m/z window using data-dependent acquisition with a 15-s dynamic exclusion. Fragmentation was performed using higher-energy collision dissociation (HCD) at a collision energy of 30. Fragment ions were analyzed at a resolution of 30,000 with an AGC target of 1 × 10⁵. The raw data files were analyzed using Mascot software (Matrix Science, London, UK; version 2.8.2) with the UniProt E. histolytica database (taxon ID: 294381; 20,546 entries). Searches assumed trypsin as the digestion enzyme, with a fragment ion mass tolerance of 10.0 ppm and a parent ion mass tolerance of 5.0 ppm. Carbamidomethylation of cysteine was specified as a fixed modification, while deamidation of asparagine and glutamine, oxidation of methionine, phosphorylation of serine, threonine, and lysine, and formylation of lysine and the protein N-terminus were defined as variable modifications. Peptide identifications were validated and processed using Scaffold software (Proteome Software Inc., Portland, OR, USA). PeptideProphet was used for peptide validation with a 1% false discovery rate (FDR), while ProteinProphet was used for protein inference with a confidence threshold of 95%. The overlap was defined as the set of EHI IDs that are both high-confidence nuclear interactors and differentially expressed in RNA-seq (padj <0.05 and |log2FC | ≥ 1).
EdU incorporation assay
Four-day-cultured Caco-2 cells were co-cultured with 1 × 10⁴ E. histolytica transformants ( ≈ 10:1 Caco-2:parasite) for 2 h, then washed 3x with pre-cooled 2% (w/v) galactose in PBS to remove parasites. Removal of adherent trophozoites was verified by phase-contrast microscopy although cannot be categorically excluded. In a parallel recombinant-protein arm, monolayers were treated apically for 2 h with His-GFP–KERP2 (1–5 µM) or His-GFP control and then the medium was replaced to remove protein. For both arms, Caco-2 DNA synthesis was assessed by adding EdU (15 µM) for 6 h, followed by staining with the Click-iT™ EdU Cell Proliferation Kit, Alexa Fluor™ 488/594 dye (Thermo Fisher Scientific), according to the manufacturer’s instructions. EdU staining was analyzed using CellProfiler 4.2.769 to quantify DNA-synthesis labeling (EdU positivity) in Caco-2 cells cocultured with E. histolytica transformants. Images of EdU-labeled nuclei (Alexa Fluor™ 488) and Hoechst 33342-stained nuclei were processed to identify EdU-positive cells. Hoechst-stained nuclei were segmented as primary objects based on intensity thresholds, with object size constrained to 30–100 pixels to exclude artifacts. EdU-positive regions were identified as secondary objects using propagation-based segmentation linked to the primary nuclei. Illumination correction was applied to normalize intensity across EdU-stained images. Morphological measurements were extracted, including the proportion of EdU-positive cells relative to the total nuclei. Data from five images per condition were exported and statistically analyzed. One-way ANOVA followed by Dunnett’s multiple comparisons test was performed using GraphPad Prism (GraphPad Software, USA).
Morphological analysis
Caco-2 cells cocultured with E. histolytica transformants were imaged as described in the IFA method above. Morphological analysis was conducted using CellProfiler 4.2.769. For each treatment, 5 images of Hoechst 33342-stained nuclei, E-cadherin-stained cell boundaries, and Phalloidin-stained F-actin were analyzed to quantify changes in cellular morphology. Nuclei were identified as primary objects using intensity-based segmentation of DNA-stained images, with a size range of 30–100 pixels. Cell boundaries were segmented as secondary objects using propagation-based methods, expanding from the nuclei and leveraging both E-cadherin and Phalloidin images for boundary definition. Illumination correction was applied to E-cadherin and Phalloidin images to normalize intensity variations. Data were exported as.csv files for statistical analysis, with parameters for segmentation and measurement optimized to ensure accurate object identification and quantification. One-way ANOVA followed by Dunnett’s multiple comparisons test was performed using GraphPad Prism (version 10.2.0 MacOS, GraphPad Software).
Morphological measurements were calculated, including:
Form Factor, defined as \(\text{Form Factor}=\frac{4{\rm{\pi }}A}{{P}^{2}}\), where A is the area and P is the perimeter of the cell, used to assess circularity.
Aspect Ratio, defined as \(\text{Aspect Ratio}=\frac{\text{Major Axis}}{\text{Minor Axis}}\), representing the elongation of cells.
Transepithelial electrical resistance (TEER) measurement
TEER was measured to assess epithelial barrier integrity using the EVOM™ Manual for TEER Measurement system (World Precision Instruments, Sarasota, FL, USA). Caco-2 cells were cultured on 12 mm Transwell inserts with a 0.4 μm pore size (3460, Corning Inc., Corning, NY, USA) as described in the cell culture method. Cells were maintained for 21 days until the TEER value stabilized at approximately 1300 Ω, indicating the formation of a confluent monolayer. Caco-2 cells were interacted with E. histolytica transformants at a ratio of 10:1 (Caco-2: E. histolytica) or different concentration of recombinant proteins. TEER measurements were recorded before the interaction and at 1-h intervals for up to 4 h. For the recombinant-protein assays, monolayers were treated apically with His-GFP–KERP2 (1–5 µM), His-GFP control, or EGTA (5 mM) for up to 24 h. All measurements were performed using pre-warmed media and pre-equilibrated electrodes, and plates were kept outside the incubator for <5 min per reading to minimize temperature-induced fluctuations. For each well, TEER was measured three times at each time point and averaged. Values were first normalized to each well’s pre-addition TEER, then normalized to the mean t = 0 value (set to 100%). Statistical significance at each time point was determined using a one/two-way ANOVA followed by Dunnett’s multiple comparisons test on GraphPad Prism (GraphPad Software).
RITC-dextran influx assay
For paracellular flux, Caco-2 monolayers were cultured to confluence on Transwell inserts (24-well format) and used only after a stable baseline TEER was recorded. For parasite-exposure assays, E. histolytica strains or EGTA (5 mM) were added to the apical chamber for 2 h, removed, and both chambers were replaced with fresh completed EMEM. Immediately thereafter, Rhodamine B isothiocyanate–Dextran ( ~ 10 kDa; MERCK) was added to the apical chamber at 1 mg mL⁻¹ and plates were incubated at 37 °C in 5% CO₂. After 4 h, medium from the basolateral (lower) chamber was collected, clarified by a brief spin, transferred to a black 96-well plate, and fluorescence was measured on a plate reader using rhodamine settings; blank medium without dextran was used for background subtraction. Unexposed Caco-2 monolayers served as negative controls, and apical treatment with EGTA served as a control. Replicates were biological (independent monolayers), and fluorescence values from the lower chamber were recorded for subsequent statistical analysis in GraphPad Prism (v10.2.0). Three independent trials were performed; within each trial, n = 4 inserts per condition.
Wound healing assay
For parasite-exposure assays, wounds were created with a Culture-Insert 2 Well in µ-Dish 35 mm (81176, ibidi). Caco-2 cells were grown 4 days to confluence; the insert was removed to form a defined gap and rinsed with warm PBS. Removal of adherent trophozoites was verified by phase-contrast microscopy. For recombinant-protein assays, gaps were created with a Culture-Insert 2 Well 24 (80242, ibidi) placed in a standard 24-well plate; seeding, growth to confluence, and insert removal were as above. E. histolytica transformants (2 × 10³) were added to both sides of the gap for 2 h. Parasites were removed by 3× washes with pre-cooled 2% (w/v) galactose in PBS. Monolayers were then maintained in EMEM + 2% FBS and imaged for 0–6 h. To enable longer, uniform treatments, monolayers were switched to EMEM + 0.5% FBS and treated with His-GFP–KERP2 (1–5 µM; typically 3 or 5 µM), His-GFP (1–5 µM; 5 µM shown), or vehicle immediately after insert removal. Proteins remained in the medium throughout imaging; no parasites were added. Phase-contrast images were taken hourly (0–6 h, parasite-exposure) and every 1–2 h (0–24 h, recombinant) at 2 fields per well. Wound size was measured in Fiji (Manual Wound Healing tool) as gap area/width (pixels) and converted to relative wound area (% of 0 h)70. For each biological replicate, simple linear regression of relative wound area versus time yielded the closure rate (slope, %/h). Slope comparisons were performed in GraphPad Prism v10.2.0 (two-sided).
Flow cytometry
Flow cytometry was performed to analyze Caco-2 cells treated with recombinant His-GFP-KERP2 or His-GFP proteins. Caco-2 cells were cultured for 21 days as described above and treated with 3 μM of His-GFP-KERP2 or His-GFP under various conditions. After treatment, cells were washed three times with 1× PBS and incubated with 0.5% trypsin for 5 min to remove excess recombinant protein and isolate the Caco-2 cells. The trypsinized cells were directly loaded into a BD Accuri™ C6 Plus flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA) at a low flow rate. Data acquisition was performed using the instrument’s standard settings, and subsequent analysis and plotting were carried out using FlowJo software (BD Biosciences) and GraphPad Prism.
Microscopy and image acquisition
Imaging for Figs. 2b, 5a–e, 8a,c, and 9a, d was performed by confocal microscopy. Figure 2b single optical sections were acquired on an Olympus FV3000 (IX83) with a PLAPON 60×/1.40 NA oil objective (pinhole ~213 µm) using 405/488/594 nm excitation (AOTF: 405 nm 0.5%, 488 nm 1%, 594 nm 23%). Figure 5a confocal images were acquired on an Olympus FV3000 (IX83) with a PLAPON 60×/1.40 NA oil objective using line-sequential 405/488/561/640 nm excitation; Fig. 5b live-cell time-lapse imaging used a Zeiss laser-scanning confocal with a Plan-Apochromat 63×/1.40 NA oil objective; Fig. 5c Z-stacks/orthogonal views were acquired on a Zeiss LSM 780 (Plan-Apochromat 63×/1.40 NA oil); Fig. 5e crypt imaging used spinning-disk confocal microscopy with a 40×/0.95 NA objective. Figure 8a EdU images were acquired as single optical sections on an Olympus FV3000 (IX83) with a UPLSAPO 20×/0.75 NA air objective (pinhole 120 µm; 12-bit) using 488/405 nm excitation (AOTF: 488 nm 7%, 405 nm 0.6%). Figure 8c images were acquired on an Olympus FV3000 (IX83) with a PLAPON 60×/1.40 NA oil objective (pinhole 213 µm; 12-bit) using 594/405 nm excitation (AOTF: 594 nm 98.7%, 405 nm 52.5%). Figure 9a Z-stacks were acquired on an Olympus FV3000 (IX83) with a PLAPON 60×/1.40 NA oil objective (19 planes, 1.0 µm spacing; pinhole 213 µm; 12-bit) using 488/594/405 nm excitation (AOTF: 488 nm 20%, 594 nm 82.3%, 405 nm 2%) and displayed as en-face views at the apical plane. Scale bars are indicated in the figures.
Statistics and reproducibility
All statistical analyses were performed in GraphPad Prism unless otherwise noted; tests were two-sided. Exact P values are reported in figure panels/legends or in the indicated Supplementary Data tables; multiplicity-adjusted P values are reported when post hoc testing was performed. Data are presented as stated in each figure (mean ± SD).
qRT–PCR and single–factor comparisons (one-way ANOVA): For Fig. 3a, Fig. 8b,d, Fig. 9b, c, i, and cysteine protease activity in Fig. 3e,f, group differences were tested by ordinary one-way ANOVA. Post hoc testing used Dunnett’s test when comparing multiple groups to a single control (Fig. 8b, d; Fig. 9b,c) and Šídák’s test for pre-specified contrasts (Fig. 3e,f; Fig. 9i). Family-wise α = 0.05.
Time-course comparisons (two-way repeated-measures ANOVA): For growth and barrier time courses (Fig. 3b, Fig. 6d, Fig. 9g, Fig. 9h), effects of time, condition/group, and time×condition interaction were evaluated by two-way RM ANOVA. Post hoc comparisons were performed within each time point using Šídák’s test (Fig. 3b; Fig. 6d; Fig. 9g) or Dunnett’s test versus the designated control (Fig. 9h), with multiplicity-adjusted P values (family-wise α = 0.05). Complete time-point–resolved adjusted P values for Fig. 9g–h are provided in Supplementary Data 8a–b.
Wound-healing migration rate (regression slope comparison): For Fig. 9e, migration rates were quantified by simple linear regression (Y = m·x + b), and slopes were compared using an extra sum-of-squares F test; 95% confidence intervals were calculated for fitted parameters.
Replication and n: The unit of study and the nature of replication (biological vs technical) are defined in each legend. Where technical replicates are shown in main figures, corresponding biological replicates are provided in the indicated Supplementary Figs. Representative imaging/phenotypic observations were reproduced in independent experiments with consistent outcomes. Specifically, Fig. 2b was performed >3 times with similar results; Fig. 2c–f were performed twice with similar results; Fig. 4g was performed three times with consistent observations; Fig. 5a was performed >3 times with consistent observations; Fig. 5f was performed twice with similar results; Fig. 6a was performed once, with multiple cells imaged showing consistent labeling patterns; and Fig. 9d was performed twice with the same phenotype consistently observed across imaged areas and between trials.
Ethics statement
Human colonic biopsy specimens were obtained from three consenting adult patients under a protocol approved by the University of Tokyo Institutional Review Board (Approval Number: 2022115NI). All procedures were performed in accordance with institutional ethical guidelines and informed consent requirements. Primary colonic crypts were isolated from these biopsies and used to establish the 3D crypt model as described below. Sex was recorded at sample collection; sex and/or gender were not considered as variables in the study design and no sex- or gender-based analyses were performed because the 3D crypt experiments used tissue from a single donor and the study does not assess participant-level outcomes.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data generated in this study have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession codes GSE290785 and GSE290901. The ChIP-seq data generated in this study have been deposited in GEO under accession code GSE292714. The proteomics data generated in this study have been deposited in the ProteomeXchange Consortium under project identifiers PXD062553 and PXD062502. Source data are provided with this paper. No datasets generated in this study are available under restricted access. Newly generated materials used in this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Koch, A. & Mizrahi, V. Mycobacterium tuberculosis. Trends Microbiol. 26, 555–556 (2018).
Disson, O., Moura, A. & Lecuit, M. Making sense of the biodiversity and virulence of listeria monocytogenes. Trends Microbiol 29, 811–822 (2021).
Einarsson, E., Ma’ayeh, S. & Svard, S. G. An up-date on Giardia and giardiasis. Curr. Opin. Microbiol 34, 47–52 (2016).
Begum, S., Gorman, H., Chadha, A. & Chadee, K. Entamoeba histolytica. Trends Parasitol. 37, 676–677 (2021).
Mugnier, M. R., Stebbins, C. E. & Papavasiliou, F. N. Masters of Disguise: Antigenic Variation and the VSG Coat in Trypanosoma brucei. PLoS Pathog. 12, e1005784 (2016).
Kantor, M. et al. Entamoeba Histolytica: Updates in Clinical Manifestation, Pathogenesis, and Vaccine Development. Can. J. Gastroenterol. Hepatol. 2018, 4601420 (2018).
Zulfiqar, H., Mathew, G. & Horrall, S. Amebiasis. In StatPearls (StatPearls Publishing, Treasure Island, FL, 2025).
Ralston, K. S. et al. Trogocytosis by Entamoeba histolytica contributes to cell killing and tissue invasion. Nature 508, 526–530 (2014).
Stanley, S. L. Jr Amoebiasis. Lancet 361, 1025–1034 (2003).
Blazquez, S., Rigothier, M. C., Huerre, M. & Guillen, N. Initiation of inflammation and cell death during liver abscess formation by Entamoeba histolytica depends on activity of the galactose/N-acetyl-D-galactosamine lectin. Int J. Parasitol. 37, 425–433 (2007).
Lidell, M. E., Moncada, D. M., Chadee, K. & Hansson, G. C. Entamoeba histolytica cysteine proteases cleave the MUC2 mucin in its C-terminal domain and dissolve the protective colonic mucus gel. Proc. Natl. Acad. Sci. USA 103, 9298–9303 (2006).
Faust, D. M. & Guillen, N. Virulence and virulence factors in Entamoeba histolytica, the agent of human amoebiasis. Microbes Infect. 14, 1428–1441 (2012).
Seigneur, M., Mounier, J., Prevost, M. C. & Guillen, N. A lysine- and glutamic acid-rich protein, KERP1, from Entamoeba histolytica binds to human enterocytes. Cell Microbiol 7, 569–579 (2005).
Santi-Rocca, J. et al. The lysine- and glutamic acid-rich protein KERP1 plays a role in Entamoeba histolytica liver abscess pathogenesis. Cell Microbiol. 10, 202–217 (2008).
Das, K. et al. Multilocus sequence typing (MLST) of Entamoeba histolytica identifies kerp2 as a genetic marker associated with disease outcomes. Parasitol. Int. 83, 102370 (2021).
Blum, M. et al. InterPro: the protein sequence classification resource in 2025. Nucleic Acids Res. 53, D444–D456 (2025).
Aravind, L. & Koonin, E. V. SAP - a putative DNA-binding motif involved in chromosomal organization. Trends Biochem Sci. 25, 112–114 (2000).
Capitano, M. L. et al. Secreted nuclear protein DEK regulates hematopoiesis through CXCR2 signaling. J. Clin. Invest 129, 2555–2570 (2019).
Davies, C. et al. TbSAP is a novel chromatin protein repressing metacyclic variant surface glycoprotein expression sites in bloodstream form Trypanosoma brucei. Nucleic Acids Res. 49, 3242–3262 (2021).
Mitra, B. N., Saito-Nakano, Y., Nakada-Tsukui, K., Sato, D. & Nozaki, T. Rab11B small GTPase regulates secretion of cysteine proteases in the enteric protozoan parasite Entamoeba histolytica. Cell Microbiol 9, 2112–2125 (2007).
Aguilar-Rojas, A. et al. Insights into amebiasis using a human 3D-intestinal model. Cell Microbiol 22, e13203 (2020).
Miyaji, M. et al. Selective DNA-binding of SP120 (rat ortholog of human hnRNP U) is mediated by arginine-glycine rich domain and modulated by RNA. PLoS One 18, e0289599 (2023).
Waldmann, T., Baack, M., Richter, N. & Gruss, C. Structure-specific binding of the proto-oncogene protein DEK to DNA. Nucleic Acids Res. 31, 7003–7010 (2003).
Berger, S. L., Kouzarides, T., Shiekhattar, R. & Shilatifard, A. An operational definition of epigenetics. Genes Dev. 23, 781–783 (2009).
Cheeseman, K. & Weitzman, J. B. Host-parasite interactions: an intimate epigenetic relationship. Cell Microbiol 17, 1121–1132 (2015).
Waldmann, T., Eckerich, C., Baack, M. & Gruss, C. The ubiquitous chromatin protein DEK alters the structure of DNA by introducing positive supercoils. J. Biol. Chem. 277, 24988–24994 (2002).
Crick, F. H. C. The packing of α-helices: Simple coiled-coils. Acta Crystallogr. 6, 689–697 (1953).
Kohn, W. D., Mant, C. T. & Hodges, R. S. Alpha-helical protein assembly motifs. J. Biol. Chem. 272, 2583–2586 (1997).
Lupas, A. N. & Bassler, J. Coiled coils - a model system for the 21st century. Trends Biochem Sci. 42, 130–140 (2017).
Zhu, J. et al. Intramolecular masking of nuclear import signal on NF-AT4 by casein kinase I and MEKK1. Cell 93, 851–861 (1998).
Zulkefli, K. L., Houghton, F. J., Gosavi, P. & Gleeson, P. A. A role for Rab11 in the homeostasis of the endosome-lysosomal pathway. Exp. Cell Res 380, 55–68 (2019).
Wilson, B. et al. Proximity labelling identifies pro-migratory endocytic recycling cargo and machinery of the Rab4 and Rab11 families. J. Cell Sci.136. https://doi.org/10.1242/jcs.260468 (2023).
Welz, T., Wellbourne-Wood, J. & Kerkhoff, E. Orchestration of cell surface proteins by Rab11. Trends Cell Biol. 24, 407–415 (2014).
Goldenberg, N. M. & Steinberg, B. E. Surface charge: a key determinant of protein localization and function. Cancer Res 70, 1277–1280 (2010).
Woolfson, D. N. Understanding a protein fold: The physics, chemistry, and biology of alpha-helical coiled coils. J. Biol. Chem. 299, 104579 (2023).
Posor, Y., Jang, W. & Haucke, V. Phosphoinositides as membrane organizers. Nat. Rev. Mol. Cell Biol. 23, 797–816 (2022).
Sharma, M. et al. Characterization of extracellular vesicles from entamoeba histolytica identifies roles in intercellular communication that regulates parasite growth and development. Infect Immun 88. https://doi.org/10.1128/IAI.00349-20 (2020).
Ohama, T., Wang, L., Griner, E. M. & Brautigan, D. L. Protein Ser/Thr phosphatase-6 is required for maintenance of E-cadherin at adherens junctions. BMC Cell Biol. 14, 42 (2013).
Ohama, T. The multiple functions of protein phosphatase 6. Biochim Biophys. Acta Mol. Cell Res 1866, 74–82 (2019).
Zhou, X. et al. I-TASSER-MTD: A deep-learning-based platform for multi-domain protein structure and function prediction. Nat. Protoc. 17, 2326–2353 (2022).
Zheng, W. et al. Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations. Cell Rep. Methods 1 https://doi.org/10.1016/j.crmeth.2021.100014 (2021).
Yang, J. & Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 43, W174–W181 (2015).
Gabler, F. et al. Protein sequence analysis using the MPI bioinformatics Toolkit. Curr. Protoc. Bioinforma. 72, e108 (2020).
Delorenzi, M. & Speed, T. An HMM model for coiled-coil domains and a comparison with PSSM-based predictions. Bioinformatics 18, 617–625 (2002).
Lupas, A. Prediction and analysis of coiled-coil structures. Methods Enzymol. 266, 513–525 (1996).
Lupas, A., Van Dyke, M. & Stock, J. Predicting coiled coils from protein sequences. Science 252, 1162–1164 (1991).
Rost, B. & Sander, C. Combining evolutionary information and neural networks to predict protein secondary structure. Proteins 19, 55–72 (1994).
Kosugi, S., Hasebe, M., Tomita, M. & Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 106, 10171–10176 (2009).
Gasteiger, E. et al. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 31, 3784–3788 (2003).
Jurrus, E. et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 27, 112–128 (2018).
Meng, E. C. et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics. 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Letunic, I. & Bork, P. Interactive tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Nakada-Tsukui, K., Okada, H., Mitra, B. N. & Nozaki, T. Phosphatidylinositol-phosphates mediate cytoskeletal reorganization during phagocytosis via a unique modular protein consisting of RhoGEF/DH and FYVE domains in the parasitic protozoon Entamoeba histolytica. Cell Microbiol. 11, 1471–1491 (2009).
Diamond, L. S., Harlow, D. R. & Cunnick, C. C. A new medium for the axenic cultivation of Entamoeba histolytica and other Entamoeba. Trans. R. Soc. Trop. Med Hyg. 72, 431–432 (1978).
Bracha, R., Nuchamowitz, Y., Anbar, M. & Mirelman, D. Transcriptional silencing of multiple genes in trophozoites of Entamoeba histolytica. PLoS Pathog. 2, e48 (2006).
Nozaki, T. et al. Characterization of the gene encoding serine acetyltransferase, a regulated enzyme of cysteine biosynthesis from the protist parasites Entamoeba histolytica and Entamoeba dispar. Regulation and possible function of the cysteine biosynthetic pathway in Entamoeba. J. Biol. Chem. 274, 32445–32452 (1999).
Hinman, S. S., Wang, Y., Kim, R. & Allbritton, N. L. In vitro generation of self-renewing human intestinal epithelia over planar and shaped collagen hydrogels. Nat. Protoc. 16, 352–382 (2021).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G. et al. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Stirling, D. R. et al. CellProfiler 4: Improvements in speed, utility and usability. BMC Bioinforma. 22, 433 (2021).
Suarez-Arnedo, A. et al. An image J plugin for the high throughput image analysis of in vitro scratch wound healing assays. PLoS One 15, e0232565 (2020).
Acknowledgements
This work was supported partly by Grants-in-Aid for Scientific Research (B) from the Japan Society for the Promotion of Science (JSPS) (JP21H02723 to T.N.), Fostering Joint International Research (B) from JSPS (JP21KK135 to T.N.), and Grant for Science and Technology Research Partnership for Sustainable Development (SATREPS) from Japan Agency for Medical Research and Development (AMED) and Japan International Cooperation Agency (JICA) (JP24jm0110022) to T.N., Grant for research on emerging and re-emerging infectious diseases from AMED (JP24fk0108680 to T.N.), and support from the University of Tokyo Pandemic Preparedness, Infection, and Advanced Research Center (UTOPIA) and AMED (JP243fa627001) to T.N. This work was also partly supported by JSPS Grants-in-Aid for Scientific Research (23K06514 to H.J.S.), JSPS Bilateral Joint Research Grant (JPJSBP120223203 to H.J.S.), and JST SPRING (Grant Number JPMJSP2108 to R.P.). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. We thank Dr. Cecilia Villegas Novoa, Dr. Yuli Wang, and Prof. Nancy Allbritton from the Department of Bioengineering, University of Washington and Prof. Soichiro Ishihara and Dr. Yuzo Nagai from Department of Surgery, The University of Tokyo Hospital for their help in constructing 3D-crypt model. We thank Prof. Tomoko Ishino and Dr. Naoki Shinzawa from the Department of Parasitology and Tropical Medicine, Science Tokyo for their help in optimizing ChIP protocol.
Author information
Authors and Affiliations
Contributions
R.P., H.S. and T.N. contributed to conceptualization; R.P., H.S. contributed to methodology; R.P. contributed to investigation and experiments; R.P., H.S. and T.N. contributed to manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jesús Valdés, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Peng, R., Santos, H.J. & Nozaki, T. A multifaceted model of Entamoeba histolytica KERP2 regulating gene expression and host cell responses. Nat Commun 17, 4433 (2026). https://doi.org/10.1038/s41467-026-70847-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70847-9