Abstract
Limited efficacy of immunotherapy in oral squamous cell carcinoma (OSCC) is driven by an immunosuppressive tumor microenvironment, yet the role of intratumoral spatial heterogeneity in immune responses remains unclear. Here, we employ single-cell and spatial transcriptomics to dissect the cellular composition and spatial organization of OSCC. We find CD8+ T cells are spatially localized yet functionally suppressed in late-stage OSCC, while myeloid-derived suppressor cells (MDSCs) transition from tumor core infiltration in early-stage OSCC to marginal localization with CD8+ T cells in advanced stages. ANXA1-FPR2 signaling mediates tumor-MDSCs communications, sustaining MDSCs recruitment and immune suppression. Disrupting ANXA1-FPR2 with an antagonist enhances the efficacy of immune checkpoint blockade therapy in OSCC mouse models. These findings reveal the spatial dynamics of MDSCs as key modulators of immune suppression and therapeutic resistance, offering a promising target to improve immunotherapy outcomes in OSCC.
Similar content being viewed by others
Introduction
Oral squamous cell carcinoma (OSCC) is a prevalent malignancy with poor prognostic outcomes, accounting for approximately 377,713 new cases and 177,757 deaths globally in 20201. Despite early-stage OSCC being associated with lower mortality risk and often amenable to surgical intervention, traditional therapies, including surgery and chemotherapy, show limited efficacy in advanced disease stages. Alarmingly, nearly 60% of OSCC cases are diagnosed at an advanced stage2, which is not suitable for surgery and underscoring the urgent need for more effective treatment strategies.
Immunotherapy, which harnesses innate and adaptive immune cells to target and eliminate tumor cells, has emerged as a promising therapeutic approach, frequently outperforming conventional radiotherapy and chemotherapy3. However, the efficacy of immunotherapy in OSCC remains highly variable among patients, largely due to the profound heterogeneity of the tumor microenvironment (TME). The TME plays a pivotal role in shaping immune responses4, and its dynamic changes during OSCC progression remain poorly understood.
Emerging evidence highlights significant alterations in cellular composition and spatial organization within the TME, even during the precancerous and early stages of OSCC5. Myeloid-derived suppressor cells (MDSCs) are key players in modulating the immune landscape of the TME6. By expressing immunosuppressive factors such as arginase-1 and PD-L1, MDSCs suppress the cytotoxic activity of CD8+ T cells7,8. Furthermore, MDSCs exacerbate immune suppression by recruiting other immunosuppressive cell types, including M2 macrophages and regulatory T cells (Tregs)9, thereby amplifying tumor immune evasion. Tumor cells actively recruit MDSCs to the TME, fostering a suppressive immune milieu that promotes tumor progression10.
Despite these insights, our understanding of the intricate cellular interactions within the OSCC immune microenvironment remains incomplete. Key gaps persist regarding the mechanisms of cell-cell communication among tumor cells, immunosuppressive cells, effector immune cells, and stromal components, as well as the spatial and temporal dynamics of these interactions during OSCC progression. Addressing these gaps is critical for developing effective therapeutic strategies. A deeper understanding of the interplay between immunosuppressive and effector cells, and their spatial organization, may unlock novel approaches to reprogram the immune landscape and enhance anti-tumor immunity. Annexin A1 (ANXA1), a recently discovered secretory calcium-dependent phospholipid-binding protein, plays a significant role in the regulation of the tumor microenvironment. Studies have shown that ANXA1 exhibits high affinity for the formyl peptide receptor family (FPR1/2/3), and the specific binding between them plays an important role in regulatory processes such as the recruitment of anti-inflammatory monocytes, directional migration of neutrophils, and polarization of immunosuppressive macrophages11,12,13. FPR2, belongs to the FPR family, plays a dual role in immunoregulation14. Recent study reveals that FPR2 leads exhaustion of T cells and drives immune escape tumor environment15.
In this study, to investigate the dynamic development of TME with OSCC progression and its underlying events contributing to immune tolerance, we performed single-cell transcriptome and spatial transcriptome analyses on primary OSCC specimens, including early-stage (stage I-II) and late-stage (stage III-IV) tumors. Our findings reveal that in early-stage OSCC, MDSCs predominantly accumulate in the tumor region, driven by Annexin A1 (ANXA1) expressed on tumor cells. As OSCC progresses, tumor cell ANXA1 expression decreases, leading to MDSCs redistribution to tumor margin regions enriched with CD8+ T cells. This redistribution enhances immune suppression by directly inhibiting CD8+ T cell cytotoxicity, thereby exacerbating immune evasion. ANXA1-FPR2 signaling as a critical axis mediating MDSCs recruitment and immunosuppressive activity. This study elucidates a novel ANXA1-FPR2-mediated mechanism of immune suppression in OSCC, offering a promising therapeutic target.
Results
Single-cell transcriptome analysis reveals the immunosuppressive tumor microenvironment in advanced OSCC
To investigate the cellular composition and dynamics of the TME in OSCC, we performed single-cell RNA sequencing (SC-seq) on stage I–II (n = 7 patients) and stage III–IV (n = 9 patients) OSCC specimens (Fig. 1a). The characteristics of the OSCC specimen were shown in Supplementary Table 1. Pathological staging was validated through hematoxylin and eosin (H&E) staining (Fig. 1b). High-quality single cells were defined by mitochondrial RNA content of less than 10% and fewer than 2500 detected RNA features. Batch effects were mitigated using the harmony algorithm. Dimensionality reduction and clustering via UMAP revealed 10 distinct cell clusters within OSCC tumors (Fig. 1c). Heatmap analysis of the top 10 highly expressed genes in each cluster confirmed unique transcriptional profiles across clusters (Supplementary Fig. 1a).
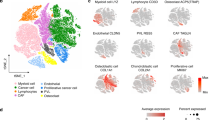
a Schematic of the high-throughput single-cell transcriptome analysis workflow for tissues collected from early-stage and late-stage OSCC. Created in BioRender. Li, F. (2026) https://BioRender.com/p0lqjyu. b Representative hematoxylin and eosin (H&E)-stained images of OSCC tumor tissue sections. Scale bar, 1000 μm. c Dimensionality reduction plot visualizing the single-cell RNA-seq data, depicting cell populations within the OSCC microenvironment (n = 102,681 cells). d Dot plot illustrating the expression of key marker genes in corresponding cell types, providing cell type identification (n = 102,681 cells). e Relative abundance of each cell type in early-stage and late-stage OSCC (left panel). Dotplot was generated by foldchange and −log10 (p value) between early-stage (n = 7 biologically independent samples) and late-stage (n = 9 biologically independent samples) OSCC (middle panel). Dimensionality reduction plot was grouped by disease stage (right panel). Significance was calculated by two-sided Student’s t test. Exact p values are provided in the Source Data file. f, g Gene set variation analysis (GSVA) evaluated the enrichment of T cell receptor signaling pathways in CD8+ T cells from early-stage and late-stage OSCC. Fold change referred to the change of the average score (late-stage vs early-stage). Data were shown as mean ± SD. n = 18,815 cells for early-stage and n = 12,226 cells for late-stage. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. h, i GSVA assessed the enrichment of immunosuppressive signaling pathways in myeloid cells during early-stage and late-stage OSCC. Fold change referred to the change of the average score (late-stage vs early-stage). Data were shown as mean ± SD. n = 5210 cells for early-stage and n = 5781 cells for late-stage. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file.
Cluster-specific markers identified various cell types: squamous epithelial cells (KRT5, KRT14), CD8+ T cells (CD3D, CD8A), CD4+ T cells (CD3D, CD4), B cells (CD79A, CD19), myeloid cells (CD14, LYZ), fibroblasts (COL1A1, TAGLN, DCN), pericytes/smooth muscle cells (RGS5, TAGLN), mast cells (KIT, TPSAB1), and nerve-like cells (CNTNAP2) (Fig. 1d). Early-stage OSCC exhibited fewer pericytes but more CD8+ T cells with significance change comparing to late-stage OSCC (Fig. 1e and Supplementary Fig. 1b), indicating that the number of T cells and their activity changed during tumor progression.
To further assess T-cell activity in OSCC, gene set variation analysis (GSVA) was performed on T-cell receptor (TCR) signaling pathways. Both signatures of TCR signaling showed diminished enrichment in late-stage tumors (Fig. 1f, g). Interestingly, immunosuppressive signaling in myeloid cells increased across tumor stages (Fig. 1h). Antigen presentation pathways were significantly reduced in myeloid cells from late-stage OSCC, indicating an impaired immune response as tumors progress (Fig. 1i).
We further evaluated the chromosomal alternation of individual cell clusters using inferCNV analysis, employing CD8+ T cells, CD4+ T cells, and pericytes as reference normal cells. Chromosomal abnormalities were predominantly observed in epithelial cells, affecting loci across multiple chromosomes (Supplementary Fig. 1c). Epithelial cells exhibited significantly higher inferCNV scores compared to other cell types (Supplementary Fig. 1 d), with malignancy increasing in late-stage OSCC (Supplementary Fig. 1e) by using the GSVA to examine the enrichment of EMT signaling16. These findings suggest that late-stage OSCC is characterized by an immunosuppressive microenvironment.
Cellular communication between epithelial tumor cells and MDSCs declines in advanced OSCC
To elucidate the mechanisms underlying the formation of an immunosuppressive tumor microenvironment in OSCC, we analyzed cell-cell communications among the 10 cell types identified via SC-seq. Interactions between epithelial tumor cells and myeloid cells were notably elevated in early-stage OSCC compared to advanced OSCC (Fig. 2a and Supplementary Fig. 2a). Moreover, epithelial tumor cells exhibited reduced communication with other cell clusters, particularly endothelial cells, in late-stage OSCC (Supplementary Fig. 2a). This observation aligns with the notion that epithelial tumor cell junctions deteriorate during OSCC progression17.
a The ratio of interaction strength between epithelial cells and myeloid cells in all significant interactions among all cell subclusters from early-stage (n = 7 biologically independent samples) and late-stage (n = 9 biologically independent samples) OSCC. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p value was provided in the Source Data file. b The interaction strength of ANXA1-FPR1 defined as significance by ‘CellphoneDB’ between epithelial cells and myeloid cells from early-stage (n = 5 biologically independent samples) and late-stage (n = 8 biologically independent samples) OSCC was shown. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t-test. Exact p value was provided in the Source Data file. c The interaction strength of ANXA1-FPR3 defined as significance by ‘CellphoneDB’ between epithelial cells and myeloid cells from early-stage (n = 7 biologically independent samples) and late-stage (n = 7 biologically independent samples) OSCC was shown. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p value was provided in the Source Data file. d The occurrence of ANXA1-FPR2 pair defined as significant interaction between epithelial cells and myeloid cells from early-stage (n = 7 biologically independent samples) and late-stage (n = 9 biologically independent samples) OSCC. Significance was calculated by Fisher test. Exact p value was provided in the Source Data file. (0: no; 1: yes) e Myeloid cell populations were clustered using the “Harmony” method, and the reduction plot was shown, with dots colored to indicate different sub-clusters. f Dot plot presenting the expression of key myeloid cell markers in the two subclusters of myeloid cells. g Dot plot showing the expression of FPR1/2/3 in myeloid cells, indicating their potential role in MDSCs-tumor interactions. n = 4973 cells in MDSCs and 6018 cells in TAM. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. h Dimensionality reduction plot visualizing the FPR2high subclusters of myeloid cells. i Dot plot showing the expression of FPR1/2/3 in FPR2high subclusters of myeloid cells. j–l The expression of FPR1/2/3 in FPR1high, FPR2high, FPR3high myeloid cells respectively grouped by MDSCs and TAM from the 16 OSCC samples. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (cells) were provided in the Source Data file. m–o The expression of FPR1/2/3 in FPR1high, FPR2high, FPR3high myeloid cells respectively grouped by early-stage (n = 7 biologically independent samples) and late-stage (n = 9 biologically independent samples) OSCC. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (cells) were provided in the Source Data file. ns, non-significant. p The cell portion of FPR2high myeloid cells (MDSCs and TAM) in total MDSCs or myeloid cells from early-stage (n = 7 biologically independent samples) and late-stage (n = 9 biologically independent samples) OSCC. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. ns non-significant.
In early-stage OSCC, epithelial tumor cell-to-myeloid cell communication was predominantly mediated by ANXA1-FPR1 and ANXA1-FPR3 ligand-receptor pairs, which diminished in advanced disease stages (Fig. 2b, c and Supplementary Fig. 2b). Interestingly, ANXA1-FPR2 pairs defined as significant interactions emerged in 5/9 late-stage OSCC samples, suggesting a compensatory role for ANXA1-FPR2 in maintaining communication between epithelial tumor cells and myeloid cells when ANXA1-FPR1 and ANXA1-FPR3 interactions were diminished (Fig. 2d and Supplementary Fig. 2c). ANXA1 was expressed across multiple cell types, whereas FPR1, FPR2, and FPR3 expression was restricted to myeloid cells (Supplementary Fig. 2d-e).
The myeloid cell compartment consisted of tumor-associated macrophages (TAM) and MDSCs, distinguished by their respective marker expression profiles (Fig. 2e, f). These subpopulations exhibited distinct transcriptional patterns (Supplementary Fig. 2f). And the cell portion of these two subclusters in all cells had no significant change with OSCC progression (Supplementary Fig. 2g).
Notably, FPR1/2/3 expression was heterogeneous among these myeloid subclusters: FPR3 was highly expressed in TAM, while FPR1 and FPR2 were predominantly expressed in MDSCs (Fig. 2g). FPR2 expression, however, was limited to approximately 10% of myeloid cells, and was higher expressed in myeloid cells from late-stage OSCC (Fig. 2g and Supplementary Fig. 2h). To identify specific myeloid cells expressing FPR2, we focused on myeloid cells with FPR2 expression levels exceeding a defined threshold (average expression level). Within this subset, both TAM and MDSCs were present (Fig. 2h, i).
Remarkably, in FPR1high/FPR2high/FPR3high myeloid cells, the expression of these three receptors was significantly higher in MDSCs than TAM (Fig. 2j-l). This suggests that MDSCs are the principal mediators of ANXA1-FPRs communication with epithelial tumor cells in OSCC. The expression of FPR3 in MDSCs was polarized, with many showing low expression. Therefore, when viewed overall, FPR3 appeared to have higher expression in TAMs. However, when filtering by mean values was applied, it effectively excluded those with low expression, resulting in the appearance of higher expression in MDSCs. Importantly, FPR1 and FPR3 expression in MDSCs and TAM, respectively, was independent of FPR2 expression (Supplementary Fig. 2i-j). This indicates that ANXA1-FPR1-mediated communication predominantly involves MDSCs, whereas ANXA1-FPR3 interactions are mainly facilitated by TAM.
In advanced OSCC, FPR1 and FPR2 expression were significantly increased, while FPR3 expression in myeloid cells had not significant difference across disease progression (Fig. 2m-o). Furthermore, the proportion of FPR2high MDSCs increased in late-stage OSCC compared to early-stage disease, whereas the proportion of FPR2high TAM had no significant change (Fig. 2p). In contrast, the ratios of MDSCs and TAM in total or FPR1high/FPR2low/FPR3high myeloid cells remained unchanged between early and late-stage OSCC (Supplementary Fig. 2k-m). In addition, the communications between epithelial cells and myeloid cells had no significant difference in the groups with different age or gender (Supplementary Fig. 2n). Thus, the change in early- and late- stage was demonstrated as the major factor to cause the different phenomenon. These findings underscore the compensatory role of FPR2high MDSCs in sustaining the immunosuppressive tumor microenvironment in advanced OSCC.
MDSCs predominantly reside in non-tumor regions in advanced OSCC
SC-seq analysis revealed exacerbation of immunosuppressive features and dynamic changes in cell-cell communication during OSCC progression. However, spatial context, which provides critical insights into the tumor microenvironment, was absent in SC-seq. To address this, spatial RNA-seq (SP-seq) was performed on tissues from three early-stage OSCC and three late-stage OSCC, in conjunction with SC-seq (Fig. 1b). A total of 3591 spots (early-stage OSCC) and 5864 spots (late-stage OSCC) were analyzed, with an average of 46,677 reads and a median of 7132 genes per spot (Supplementary Fig. 3a, b). SP-seq identified cell types consistent with SC-seq (Fig. 3a).
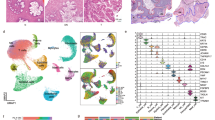
Spatial transcriptome analysis combined with SC-seq and RCTD deconvolution to elucidate the spatial distribution of specific cell types in OSCC. a The probability distribution calculated by RCTD deconvolution of each cell cluster across early-stage (n = 3 biologically independent samples) and late-stage (n = 3 biologically independent samples) OSCC was shown as scatter pies mapping to the SP-seq spots. Scale bar, 500 μm. b The ratio of each cell type in R1 (the spots with ratio of epithelial cells minor than 3/4 quartile value) and R2 (the spots with ratio of epithelial cells larger than 3/4 quartile value) from early-stage and late-stage OSCC samples was quantified. Data were presented as mean ± SD. Significance was calculated by one-way ANOVA with Bonferroni’s multiple comparisons test. Exact p values and sample amount (spots) were provided in the Source Data file.
Using Robust Cell Type Decomposition (RCTD) for deconvolution, cell type distributions across spatial spots were quantified (Fig. 3b). The tissues were divided into two regions: epithelial tumor cell-poor regions (region I, outside the ROI with black line) and epithelial tumor cell-rich regions (region II, within the ROI with black line) (Fig. 3b). In early-stage OSCC, myeloid cells were enriched in region II, while in late-stage OSCC, they predominantly resided in region I (Fig. 3b). Notably, this spatial redistribution was specific to myeloid cells, as other cell types did not exhibit similar changes.
Using the neighborhood analysis of spatial spots with reference to epithelial cells, we found that myeloid cells were enriched in intra-spots in early-stage OSCC, while in late-stage OSCC, they predominantly resided in inter- and distal-spots (Fig. 4a–b). Similarly to the results from RCTD, this spatial redistribution was specific to myeloid cells, as other cell types did not exhibit similar changes (Fig. 4b and Supplementary Fig. 3c, d). Additionally, CD8+ T cells infiltrated to intra-, inter- or distal-spots of epithelial tumor cells with similar degree in early-stage OSCC, but CD8+ T cells rarely infiltrated to intra-spots while dominantly located in inter- and distal-spots of epithelial tumor cells in late-stage OSCC (Fig. 4c and Supplementary Fig. 3d).
a Neighborhood analysis of epithelial tumor cells in SP-seq, displaying the distribution of intra, inter, and distal spots in early-stage and late-stage OSCC. Scale bar, 500 μm. b, c The relative abundance of CD8+ T cells and myeloid cells within each of the three spot categories (intra, inter, distal) in early-stage (n = 3 biologically independent samples) and late-stage (n = 3 biologically independent samples) OSCC. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (spots) were provided in the Source Data file. ns, non-significant. d–i Quantification of the ratio of each cell type within intra, inter, and distal spots in early-stage (n = 3 biologically independent samples) and late-stage (n = 3 biologically independent samples) OSCC. Data were presented as mean ± SD. Significance was calculated by one-way ANOVA with Bonferroni’s multiple comparisons test. Exact p values and sample amount (spots) were provided in the Source Data file. ns, non-significant. j Spatial expression of ANXA1 visualized using the “SpatialFeaturePlot” function from the Seurat R package, highlighting its expression across the tissue sections (Early-stage, n = 3 biologically independent samples; Late-stage, n = 3 biologically independent samples). Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p value and sample amount (spots) were provided in the Source Data file. Scale bar, 500 μm. k Epithelial tumor cell weights stratified by high/low MDSC or TAM weights (defined by the upper quartile) (Early-stage, n = 3 biologically independent samples; Late-stage, n = 3 biologically independent samples). Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p value and sample amount (spots) were provided in the Source Data file. Scale bar, 500 μm.
Neighborhood analysis of spatial spots revealed distinct interactions in early and late-stage OSCC. In early-stage OSCC, intra-spot ratios of myeloid cells had the highest ratio, which was decreased in inter- and distal-spots (Fig. 4d-f). In late-stage OSCC, fibroblasts dominated intra-spot regions, while endothelial cells and fibroblasts were more prominent in inter- and distal-spots (Fig. 4g-i), accompanied by increased myeloid cell ratios (Fig. 4b and Supplementary Fig. 3d). Further analysis demonstrated that MDSCs from early-stage group preferred to locate in tumor cells enrich region (Fig. 4k). However, MDSCs from late-stage group and TAM from the both groups mainly distributed in tumor cells poor region (Fig. 4k), suggesting that MDSCs were the dominant component of myeloid cells mediating immune-suppressive tumor environment in OSCC. Although ANXA1 was expressed across most cell types in SC-seq, SP-seq revealed its elevated expression in epithelial tumor cell-enriched regions (Fig. 4j).
Immunofluorescence (IF) validated SP-seq findings, showing CD11b+ MDSCs predominantly localized to tumor regions in early-stage OSCC but distributed mainly in non-tumor regions in late-stage OSCC, with a small subset still present in tumor regions (Fig. 5a-c). These spatial patterns were corroborated in a transgenic OSCC mouse model (Tgfbr1f/f; Ptenf/f; K14-CreERT2) (Fig. 5d, e).
a Immunofluorescence staining of epithelial tumor cells in OSCC tissue sections using anti-PanCK antibody (red) to assess their spatial distribution. Data was a representative of three independent experiments. b Immunofluorescence analysis of MDSCs distribution using anti-CD11b antibody (red) in consecutive tissue sections from early-stage and late-stage OSCC samples. c Quantification of CD11b positive cells. The bar graph showed the ratio of CD11b positive cells in R1 or R2 region. Data were a representative of three independent experiments. Data from three biological replicates in each group were presented as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. d Analysis of tumor tissues from a spontaneous OSCC mouse model (Tgfbr1f/f; Ptenf/f; K14-CreERT2) using mIHC with anti-PanCK (red), anti-CD11b (purple), and anti-CD8 (green) antibodies; nuclei were counterstained with DAPI (blue). e Quantification of CD11b positive cells. The bar graph showed the ratio of CD11b positive cells in R1 or R2 region. Data were a representative of three independent experiments. Data from two biological replicates in each group were presented as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. f Representative mIHC images of human OSCC tissue arrays showing protein expression of CD11b (purple), PanCK (red), and CD8 (green). Split images display individual markers, and merged images illustrate co-localization patterns. Data were a representative of three independent experiments. g Quantification of the percentage of the overlay area between CD11b-positive and PanCK-positive regions relative to the total CD11b-positive area in stage I (n = 22) and stage IV (n = 37) OSCC specimens. Data were presented as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. h Kaplan-Meier survival curves for overall survival (OS) of OSCC patients, stratified by the percentage of overlay between CD11b-positive and PanCK-positive regions in the total CD11b-positive area. Significance was calculated by Gehan-Breslow-Wilcoxon test. Exact p values were provided in the Source Data file.
Multiplex immunohistochemistry (mIHC) analysis of human OSCC tissues further confirmed that MDSCs were preferentially localized to epithelial tumor cell-enriched regions in stage I OSCC but not in stage IV OSCC (Fig. 5f). The percentage overlap of MDSCs and epithelial tumor cell regions was significantly higher in stage I OSCC than in stage IV OSCC (Fig. 5g). Importantly, lower overlap between MDSCs and epithelial tumor cell regions correlated with poor overall survival (Fig. 5h).
These findings strongly suggest that the redistribution of MDSCs to non-tumor regions in advanced OSCC is closely associated with disease prognosis, underscoring the critical role of spatial MDSCs dynamics in shaping the tumor microenvironment.
FPR1+ or FPR2+ myeloid cells have alternative contact with ANXA1+ epithelial cells across progression
Above SC-seq results demonstrated that the cell-cell communication pair of ANXA1-FPR1 between epithelial tumor cells and myeloid cells was elevated in early-stage OSCC, while ANXA1-FPR2 was increased in late-stage OSCC. To further validate this observation, we used mIHC to assess the three proteins in corresponding samples to SC-seq. As shown in Fig. 6a, b and Supplementary Fig. 4a, b, FPR1 positive myeloid cells, which were near to the ANXA1 positively expressed epithelial tumor cells, significantly reduced in the late-stage OSCC, while FPR2 positive myeloid cells significantly increased. Notably, the protein levels of ANXA1 decreased in late-stage OSCC, whereas protein levels of FPR2 increased with no significant change of FPR1 across disease progression (Fig. 6c, d), which was consistent to the results from SC-seq. These results demonstrated that the decreased expression of ANXA1 may contributed to the reduced interactions with FPR1 between epithelial tumor cells and myeloid cells, and increased expression of FPR2 attempted to remedy these cell-cell communications.
a The distribution of FPR1 (purple), FPR2 (green) or ANXA1 (red) positive cells (right panel) in the sections on the border between epithelial tumor cells and myeloid cells determined by SP-seq as indicated by the white rectangular box in the left panel from early-stage (n = 3 biologically independent samples) and late-stage (n = 3 biologically independent samples) OSCC specimen. b The ratio of FPR1 or FPR2 positive cells was quantified grouped by early- and late-stage. Data was a representative of three independent experiments. Data from three biological replicates in each group were shown as means ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. Scale bar in a, 500 μm; scale bar in b, 50 μm. c, d The representative photos showing the expression of ANXA1 (red), FPR1 (purple) and FPR2 (green) (c), which was quantified by average optical density (AOD) (d). Data were from seven biologically independent samples in early-stage OSCC and nine biologically independent samples in late-stage OSCC. Data were a representative of three independent experiments. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. ns non-significant. Scale bar = 50 μm.
ANXA1high epithelial tumor cells are identified as the major sub-cluster communicated to MDSCs
To delineate the epithelial tumor cell sub-clusters involved in communication with MDSCs via ANXA1-FPR1/2/3 signaling, epithelial tumor cells were subjected to dimensionality reduction clustering. Two distinct sub-clusters were identified: ANXA1 highly expressing epithelial tumor cells (ANXA1high_Epi) and ANXA1 lowly expressing epithelial tumor cells (ANXA1low_Epi) (Fig. 7a and Supplementary Fig. 5a, b). Notably, ANXA1 expression was enriched in ANXA1high_Epi but nearly absent in ANXA1low_Epi (Fig. 7b), suggesting that ANXA1high_Epi predominantly engages MDSCs through ANXA1-FPR interactions. Notably, the expression of ANXA1 from late-stage OSCC in both subclusters significantly reduced (Fig.7c). However, the cell portion of these two subclusters of epithelial tumor cells had no significant difference (Fig. 7d). Interestingly, although ANXA1 was predominantly expressed in ANXA1high_Epi, its expression decreased during OSCC progression (Fig. 7c). This reduction likely contributed to the weakened interaction between epithelial tumor cells and MDSCs via ANXA1-FPRs in advanced OSCC.
a Sub-clustering of epithelial tumor cells into two distinct sub-clusters, ANXA1high_Epi and ANXA1low_Epi, visualized in the reduction plot. b ANXA1 expression levels in the two subclusters from the 16 OSCC samples. ANXA1 was significantly upregulated in ANXA1high_Epi cells. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (cells) were provided in the Source Data file. c The expression of ANXA1 in ANXA1high_Epi or ANXA1low_Epi cells grouped by early-stage (n = 7 biologically independent samples) or late-stage (n = 7 biologically independent samples) OSCC. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (cells) were provided in the Source Data file. d The cell portion of ANXA1high_Epi and ANXA1low_Epi cells in total epithelial tumor cells from each sample (n = 16 biologically independent samples). Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values were provided in the Source Data file. ns, non-significant. e Stemness of epithelial tumor cell sub-clusters was analyzed using CytoTRACE from the 16 OSCC samples. ANXA1high_Epi exhibited higher stemness compared to ANXA1low_Epi. f In CytoTRACE output boxplots, the center line represented the median, the box bounds denoted the interquartile range (25th to 75th percentiles), the whiskers extended to the largest/smallest values within 1.5 times the interquartile range, and any points beyond the whiskers represented outliers. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (cells) were provided in the Source Data file. GSVA analysis assessing the activity of g tumor stemness and h antigen presentation pathways in epithelial tumor cells from the 16 OSCC samples. Data were shown as mean ± SD. Significance was calculated by two-sided Student’s t test. Exact p values and sample amount (cells) were provided in the Source Data file.
The stemness of the two subclusters was further analyzed using CytoTRACE. ANXA1high_Epi subcluster exhibited higher stemness than ANXA1low_Epi subcluster (Fig. 7e, f), and the tumor stemness signaling was also enriched in epithelial tumor cells from early-stage OSCC (Fig. 7g), indicating that ANXA1-FPRs-mediated communication occurs primarily during early tumorigenesis. Further analysis revealed a dramatic downregulation of interferon (IFN-γ and IFN-α) response signaling and antigen presentation in epithelial tumor cells from late-stage OSCC (Fig. 7h and Supplementary Fig. 5c-e).
Gene enrichment analysis highlighted distinct functional profiles of these sub-clusters. ANXA1high_Epi cells showed reduced Myc targets, G2M checkpoints and E2F targets pathways (Supplementary Fig. 5f), indicating that ANXA1high_Epi cells had lower proliferation activity than ANXA1low_Epi cells.
Together, these results demonstrated that OSCC epithelial tumor cells exhibited suppressed IFN response signaling, facilitating immune evasion with disease progression. The progressive reduction of ANXA1 expression in epithelial tumor cells underscored its critical role in tumor-immune cell communication and highlighted a potential mechanism for diminished immune engagement during OSCC progression.
Treatment of FPR2 antagonist enhances the response to anti-PD-1 antibody in OSCC mice
As the above data suggested, cell communication among epithelial tumor cells and MDSCs through ANXA1-FPR2 played a complementary signaling for decreased ANXA1-FPR1/3 signaling, keeping the recruitment of MDSCs by epithelial tumor cells in late-stage OSCC. Therefore, the disruption of FPR2 from MDSCs may enhance the effect of ICB treatment. To establish the OSCC model, we injected the advanced OSCC cell line SCC7 into C3H/HeJ mice subcutaneously. Once the tumor volume reached 50 mm3, the mice were randomly divided into four groups, receiving IgG isotype plus placebo, IgG isotype plus WRW4 (FPR2 inhibitor)18, anti-PD-1 antibody plus placebo and anti-PD-1 antibody plus WRW4, respectively (Fig. 8a). The tumor growth was significantly inhibited by combination therapy of WRW4 with anti-PD-1 antibody, while the single therapy groups had little effect on the tumor growth (Fig. 8b).
a Schematic of the OSCC syngeneic mouse model. SCC7 cells were subcutaneously injected into C3H/HeJ mice. Treatment schedules with anti-PD-1 antibody and/or WRW4 are shown. b The volume of tumors from mice received indicated agents (n = 5 biologically independent animals per group) was measured to generate the growth curves. Data were shown as mean ± SEM. Significance was calculated by two-way ANOVA with Tukey’s multiple comparisons test. Exact p values were provided in the Source Data file. ns non-significant. c Kaplan-Meier survival analysis of SCC7 syngeneic mice. Mice were considered deceased when tumor volume exceeded 800 mm³. Significance was assessed using the Log-rank (Mantel-Cox) test (n = 5 per group). Exact p values were provided in the Source Data file. ns, non-significant. d Tumor volumes and weight measured at the endpoint. Statistical significance was assessed. Data were shown as mean ± SEM. Significance was calculated by one-way ANOVA with Bonferroni’s multiple comparisons test. Exact p values were provided in the Source Data file. ns, non-significant. e Tumor sections from syngeneic mice were processed for multiplex immunohistochemistry (mIHC) with antibodies targeting PanCK (red), CD8 (green), and CD11b (purple). DAPI (blue) was used for nuclear staining. Data were a representative of two independent experiments. Scale bar, 50 μm. f, g Ratios of CD11b+ MDSCs and CD8+ T cells were analyzed in region I (R1) and region II (R2) of the syngeneic tumors based on mIHC results. Data from three biological replicates over two independent experiments in each group were shown as mean ± SEM. Significance was calculated by one-way ANOVA with Bonferroni’s multiple comparisons test. Exact p values were provided in Source Data file.
Survival analysis demonstrated that single treatment of anti-PD-1 or WRW4 barely affected the survival period, whereas combination treatment of anti-PD-1 with WRW4 significantly extended the survival prognosis (Fig. 8c). In addition, anti-PD-1 or WRW4 treatment alone hardly inhibited tumor growth, which was significantly inhibited by combination treatment (Fig. 8d).
Furthermore, tumors from mice received vehicle and isotype control IgG were infiltrated with large amount of CD11b+ MDSCs, but the CD8+ T cells were hardly distributed in the PanCK abundant region (Fig. 8e-g). Even though anti-PD-1 single treatment could increase the infiltration of CD8+ T cells within the squamous tumor cells enriched region, extensive CD11b+ MDSCs distributed in the whole microenvironment (Fig. 8e-g). Although CD11b+ MDSCs were eliminated and the CD8+ T cells were abundant in tumor microenvironment by WRW4, the growth of tumors was not inhibited (Fig. 8e-g). Combination of anti-PD-1 with WRW4 treatment led to significantly decreased CD11b+ MDSCs infiltration and increased CD8+ T cells infiltration, which had impressive inhibitory effect on tumor growth (Fig. 8d). Although both inhibition of FPR2 and PD-1 increased the anti-tumor effect in the mice, exhausted signaling in CD8+ T cells reduced in late-stage OSCC (Supplementary Fig. 4c, d), which aligns with the opinion that persistent T cell receptor (TCR) stimulation can effectively induce T cell exhaustion19 as our current findings showing a significant enhancement of TCR signaling in early-stage OSCC (Fig. 1f, g).
These results demonstrated that recruitment of CD11b+ MDSCs was predominantly maintained by FPR2 signaling. Both FPR2 inhibitor and ICB were necessary to inhibit immunosuppressive actions within the tumor microenvironment of advanced OSCC. Altogether, ANXA1 is a predominant ligand to FPRs which is responsible for cell communications among epithelial tumor cells and MDSCs. Enhanced communication of epithelial tumor cells ANXA1 and MDSCs FPR1 contributes to recruitment of MDSCs, which is partially compensated by ANXA1-FPR2 pairing accompanied with reduced expression of ANXA1 in advanced OSCC tumor tissues. Weakened cell talking among epithelial tumor cells and MDSCs via ANXA1-FPRs with tumor progression leads to diffuse of MDSCs to cytotoxic T cells, enhancing the immunosuppressive tumor microenvironment. Blocking of ANXA1-FPR2 by FPR2 inhibitor promoted the anti-tumor effect with anti-PD-1 antibody.
Discussion
TME of OSCC remains incompletely understood, despite its critical association with immune therapy response rates. In precancerous lesions, CAFs, monocytes, and Tregs contribute to creating a pro-tumorigenic microenvironment, facilitating tumor initiation5. Cellular communication between cancer cells and MDSCs significantly influences disease progression in various cancers. MDSCs secrete factors such as IL-6, NO, and TGFβ, enhancing tumor cell stemness and epithelial-mesenchymal transition (EMT)20. Conversely, tumor cells amplify the immunosuppressive functions of MDSCs via pathways including NF-κB, STAT, and ER stress responses21.
In this study, the ANXA1high_Epi sub-cluster of epithelial tumor cells emerged as a major contributor to interactions with MDSCs, characterized by a low differentiation state consistent with prior findings21. However, whether MDSCs directly drive the stemness of ANXA1high_Epi cells warrant further investigation. Sub-cluster differentiation within tumor cells plays pivotal roles in tumor initiation, therapy resistance, metastasis, and modulation of immune cell composition22. We demonstrate that ANXA1high_Epi tumor cells act as a “bait” for MDSCs via ANXA1-FPR signaling, leading to their accumulation. As OSCC progresses, reduced ANXA1 expression in ANXA1high_Epi tumor cells promote MDSCs migration from tumor centers to the tumor periphery, inhibiting CD8+ T-cell activity.
The functional role of FPRs in immune regulation is complex and context-dependent. Consistent with our findings, FPRs are predominantly expressed in myeloid cells and exhibit diverse immune-regulatory functions. FPR2, for instance, promotes dendritic cell maturation and monocyte polarization to M1 macrophages while also inducing IL-10-secreting neutrophils, which suppress CD8+ T-cell proliferation23,24,25. In OSCC, myeloid cells primarily comprise TAM and MDSCs, which was increased in OSCC tumors26. In the present study, the elevated number of FPR2high MDSCs in late-stage OSCC suggests that FPR2 expression undergoes dynamic regulation and indicates that the complex role of the subclusters of MDSCs in OSCC. Pharmacological inhibition of FPR2 with WRW4 significantly reduced MDSCs infiltration and enhanced CD8+ T-cell-mediated anti-tumor immunity, highlighting the therapeutic potential of FPR2 antagonists for advanced OSCC.
Tumor-derived factors such as GM-CSF, M-CSF, IL-6, and TNF, along with chemokines like CXCR2 and CCR2 ligands, are well-documented recruiters of MDSCs21. Our study underscores the importance of the ANXA1-FPR2 axis in mediating interactions between tumor cells and MDSCs, contributing to the immunosuppressive microenvironment. ANXA1-FPR2 interactions have been implicated in diverse immune regulatory processes, including anti-inflammatory monocyte recruitment and inhibition of leukocyte migration11,12,27. While ANXA1 is expressed across various cell types, including tumor cells, fibroblasts, and mast cells, FPR2 expression in OSCC is restricted to myeloid cells. This tissue-specific expression pattern contrasts with ESCC, where FPR2 is found in fibroblasts responding to ANXA128.
Our findings reveal that MDSCs redistribution during OSCC progression is a critical driver of the immunosuppressive TME. Accumulating evidence links the spatial organization of CAFs, epithelial cells, and immune cells within the TME to tumor initiation and immune responses5,29,30. Here, we observed a unique translocation of MDSCs from the tumor center to the periphery in late-stage OSCC, correlating with poor survival outcomes. Reduced ANXA1 expression in S100A9_Epi cells during tumor progression diminishes intrinsic epithelial-MDSCs communication, facilitating MDSCs migration to regions enriched with CD8+ T cells.
The role of ANXA1 in cancer progression remains controversial. While ANXA1 promotes invasion in breast cancer, it inhibits invasion in lung cancer31,32. Loss of ANXA1 expression facilitates the transition of normal fibroblasts to cancer-associated fibroblasts, driving ESCC progression28. In OSCC, we identify ANXA1-FPR2 signaling as a central pathway promoting immunosuppressive TME formation. Inhibition of FPR2 with WRW4 significantly enhanced anti-PD-1 therapy, reducing MDSCs abundance and increasing CD8+ T-cell activity.
These findings provide novel insights into the mechanisms underlying immunosuppression in OSCC and emphasize the potential of targeting the ANXA1-FPR2 axis to improve immune checkpoint blockade efficacy.
Methods
Ethics statement and collection of patient samples
To investigate the cellular composition and dynamics of the TME in OSCC, primary human OSCC tumor specimens were collected from patients who underwent surgical resection at the First Affiliated Hospital of Chengdu Medical college. The procedures were reviewed and approved by the Institutional Review Board (IRB) at the First Affiliated Hospital of Chengdu Medical college (Ethics Approval No. 2022CYFYIRB-BA-Aug11). Samples were collected according to the institutional regulation of the Institutional Review Board (IRB) and were examined by pathologists. All procedures were performed with signed informed consent from the patients, and the donation of tumor tissues was entirely voluntary and uncompensated.
Samples preparation for single-cell transcriptome analysis
Fresh OSCC tissue samples were collected and divided into two portions: one for single-cell dissociation and the other stored temporarily in tissue preservation solution (MACS, 130-100-008, Miltenyi Biotec) at 4 °C for nuclear preparation. To synchronize the preparation of single-cell and nuclear suspensions, single-cell dissociation was performed first. For single-cell dissociation, tissues were washed twice with pre-cooled DMEM medium (11965092, Gibco) supplemented with 0.04% BSA (30066575, Gibco), minced into small pieces, and enzymatically digested in a solution containing DMEM, 0.04% BSA, 0.2% collagenase II (17101015, Gibco), and 100 μg/L DNase I (A3778, AppliChem) at 37 °C with gentle agitation (22 rpm) for ~30 min. This was followed by the addition of 0.25% trypsin (2062476, Gibco) for an additional 5 min of digestion. The resulting cell suspension was passed through a 40 μm cell strainer, centrifuged at 4 °C at 300 g for 7 min, and treated with Red Blood Cell Lysis Solution (130-094-183, Miltenyi Biotec) to remove erythrocytes. After washing with medium, the cell pellet was resuspended in PBS containing 1% BSA, and cell count and viability were assessed using a Luna Fluorometer (LUNA-FLTM, Logos Biosystems). The final cell concentration was adjusted to 700–1200 cells/μL.
In parallel, nuclear suspension preparation was initiated once single cells were centrifuged post-erythrocyte lysis. Tissue for nuclear isolation was washed twice with pre-cooled medium, minced into pieces ~millet-sized, and lysed in 1 mL pre-cooled NST buffer (0.1% NP-40, 10 mM Tris-HCl, 146 mM NaCl, 1 mM CaCl₂, 21 mM MgCl₂, and 1 U/μL RNase inhibitor) for 3–5 min on ice. The nuclear suspension was filtered through a 40 μm cell strainer, centrifuged at 4 °C at 500 g for 10 min, and washed with 2 mL wash buffer (10 mM Tris-HCl, 146 mM NaCl, 1 mM CaCl₂, 21 mM MgCl₂, 0.01% BSA, and 40 U/mL RNase inhibitor). After a second centrifugation at 4 °C at 500 g for 5 min, the nuclei were resuspended in PBS containing 1% BSA and 1 U/μL RNase inhibitor. Nuclei were stained with Trypan blue and counted under a microscope. The final nuclear concentration was adjusted to 700–1200 nuclei/μL and maintained on ice until use.
Single-cell RNA-seq and analysis
SC-seq was performed using the MobiCube High-throughput Single Cell 3’ Transcriptome Set V2.1 (PN-S050200301) and the MobiNova-100 microfluidic platform at OEbiotech company. Single-cell suspensions were adjusted to a concentration of 700–1200 cells/μL and immediately loaded into a chip for micro-droplet formation on the MobiNova-100 system33. Reverse transcription, cDNA amplification, and DNA library construction were performed following the manufacturer’s protocol. High-throughput sequencing was conducted using paired-end 150 bp (PE-150) mode.
Sample quality control was performed using MobiVision, which incorporates the STARSolo software for data processing. Sequencing reads were aligned to the reference genome (GRCh38), and metrics such as cell count, gene count, and genome alignment rate were obtained from the raw data to evaluate sample quality. For downstream analysis, the ‘Seurat’ R package (version 5.1.0)34 was used to create analysis objects. Cells with mitochondrial RNA content below 10%, gene counts (nFeature_RNA) between 200 and 2500, and high-quality transcriptomes were retained for further analysis. Batch effects were corrected using the ‘Harmony’ R package to ensure robust integration across samples.
Copy number variations analysis
To investigate copy number variations (CNVs) across different cell populations within OSCC tissue, we performed inferCNV analysis35, a computational approach designed to detect large-scale chromosomal alterations in SC-seq data. SC-seq data from OSCC tissue samples were preprocessed to remove low-quality cells, normalize gene expression levels, and annotate cells by type using clustering or marker gene-based methods. A reference set comprising CD8+ T cells, CD4+ T cells, and pericytes was included to serve as a baseline for CNV comparison. CNV analysis was conducted using the inferCNV R package on a high-performance computing cluster. The processed SC-seq gene expression matrix was input into the inferCNV pipeline, which calculates relative gene expression changes to infer chromosomal alterations. The results were visualized using the ggplot2 R package, generating heatmaps and plots to illustrate CNV patterns across cell populations.
Cell-cell communication analysis
To investigate the interactions between different cell types in the tissue microenvironment, we performed cell-cell communication analysis using the CellPhoneDB method36. CellPhoneDB analysis was carried out using the CellPhoneDB Python package, which was run on a computational cluster. The gene expression data was processed and formatted as required by the tool, and ligand-receptor interaction predictions were made using default parameters unless otherwise specified. The resulting interaction networks were visualized using ggplot2 in R for heatmaps and plots.
Spatial transcriptome analysis
Freshly collected OSCC tissue samples were divided into appropriately sized pieces and fixed in 4% formaldehyde, followed by paraffin embedding. Tissue sections (5 μm thick) were prepared from the embedded samples and processed for hematoxylin-eosin (HE) staining, imaging, and destaining, following the protocol (CG000614) recommended by 10× Genomics. Subsequent steps, including probe hybridization, probe release, and transfer to the 10× Visium CytAssist slide, were performed according to the 10x Genomics workflow (CG000495). Library construction was carried out using the Visium CytAssist Spatial Gene Expression for FFPE kit (PN-1000520, 10× Genomics). The resulting DNA libraries were subjected to high-throughput sequencing using paired-end 150 bp (PE-150) mode to ensure comprehensive spatial transcriptome profiling.
SP-seq data preprocessing
FASTQ files generated from spatial RNA sequencing were processed and aligned to the GRCh38 human reference genome using Space Ranger software (version 2.0.1) from 10x Genomics37. Unique molecular identifier (UMI) counts were summarized for each barcode. Tissue-overlaying spots were identified based on the accompanying histological images to distinguish tissue-associated signals from background noise. The filtered UMI count matrix was analyzed using the Seurat R package (version 5.1.0). Data normalization and identification of the top 3000 highly variable genes (HVGs) were performed using the SCTransform function38. Dimensionality reduction was conducted using principal component analysis (PCA) on the log-transformed gene-barcode matrices of the HVGs. Graph-based clustering was applied to classify cells based on their gene expression profiles using the FindClusters function. Data visualization was achieved with the Uniform Manifold Approximation and Projection (UMAP) algorithm, implemented through the RunUMAP function. Marker genes for each cluster were identified using the FindAllMarkers function with the test.use = “bimod” parameter to ensure robust detection of cluster-specific genes.
Deconvolution of spatial transcriptome data
To investigate the cellular composition of each spatial transcriptomics spot, we employed the Robust Cell-Type Deconvolution (RCTD, version 1.1.0) method39. RCTD is a computational framework designed to infer the proportions of different cell types in complex tissues based on gene expression data. Spatial transcriptomics data, providing gene expression profiles for individual tissue locations, were paired with a reference cell-type gene expression matrix derived from SC-seq data of corresponding OSCC samples (early-stage and late-stage respectively). The reference matrix encompassed gene expression profiles for various cell types identified within the tissue, serving as signatures for the deconvolution process. This approach enabled the precise estimation of cell-type proportions across spatial locations, facilitating the exploration of spatial cellular heterogeneity and its role in OSCC progression.
Neighborhood analysis of epithelial tumor cells
Neighborhood analysis40 was conducted to examine the spatial interactions of epithelial tumor cells with surrounding cell types, based on the results from the RCTD deconvolution assay applied to spatial transcriptomics data. Epithelial tumor cells were identified as intra-spots, while other cell types within two layers of proximity were categorized as inter-spots. Cells located beyond this range were classified as distal-spots. The relative proportions of each cell type in intra, inter, and distal spots were calculated for both early-stage and late-stage OSCC samples. The results were visualized through comparative plots to illustrate the spatial organization and dynamics of cellular neighborhoods around epithelial tumor cells during OSCC progression. The code for this analysis, which was developed by a company and author, is available from the corresponding author on reasonable request.
Multiplex immunohistochemical (mIHC) assay
Paraffin-embedded tissue sections were heated in an oven at 65 °C for 2 h, and the subsequent dewaxing process included two rounds of 10-min xylene treatment. Next, the sections were rehydrated in 100% (for twice), 95%, and 70% alcohol in sequence, followed by heat-induced antigen retrieval with Tris-EDTA buffer (pH 9.0), followed by natural cooling to room temperature. 3% H2O2 was added for endogenous peroxidase blocking. 3% BSA was used to block unspecific binding at room temperature for 1 h. After the removal of the 3% BSA, the sections were incubated rabbit anti-pan-Keration (1: 200, 26411-1-AP, Proteintech) at 4 °C for overnight. The sections were then washed with PBS three times, and incubated with HRP-labeled secondary antibodies for 10 min at room temperature. Signal amplification was performed using Cy3-labled tyramide signal amplification working solution (1: 500, G1223, Servicebio) and incubated at room temperature for 10 min, followed by the treatment of microwave heat. Repeat with rabbit anti-CD8 (1:200, ab316778, Abcam), rabbit anti-CD11b (1:200, ab133357, Abcam) combined with FITC-labeled tyramide (1:200, G1231, Servicebio) or Cy5-labled tyramide (1:200, G1232, Servicebio) signal amplification working solution, respectively. Nuclei were then stained with DAPI (G1012, Servicebio). Antibodies against to ANXA1(1: 200, ab214486, Abcam), FRP1 (1:200, ab113531, Abcam) and FRP2 (1:200, ab203129, Abcam) were obtained from Abcam. The stained slides were scanned with Olympus VS200 and analyzed with OlyVIA analysis software (Version 3.3).
Cell culture
SCC7 cells were acquired from WANWUSHENGWU (Delf-17084) and authenticated via short tandem repeat (STR) profiling, and mycoplasma contamination was routinely tested and confirmed to be negative. SCC7 cells were cultured in RPMI 1640 (Gibco, Shanghai, China) supplemented with 10% FBS (10099141, Gibco), penicillin (100 U/mL) and streptomycin (100 mg/mL) (15140122, Gibco) at 37 °C in a humidified incubator under 5% CO2.
Animal care and mouse models
Animal care and experiments in this study were approved by the Institutional Animal Care and Use Committee (IACUC) of Chengdu Medical College (approval No. CMC-IACUC-2024088), and were conducted in accordance with the institutional guidelines under the official animal license (SYXK(Sichuan) 2020-0196) issued by the Sichuan Provincial Department of Science and Technology. All the mice were bred and maintained in a specific pathogen-free (SPF) barrier facility at Laboratory Animal Center of Chengdu Medical College. The mice were maintained on a 12/12-h light/dark cycle (lights on at 09:00 a.m.), 22 ± 2 °C with a standard and sterile chow diet and water ad libitum.
For generation of spontaneous OSCC genetic mice model, Pten/Tgfbr1 double conditional knockout (2cKO) mice are prone to spontaneously developing OSCC41. Ptenflox/flox, Tgfbr1flox/flox and K14-CreERT2 mouse line (genetic strain FVB/N) were purchased from The Jackson Laboratory and crossed to generate Tgfbr1/Pten 2cKO mice (Tgfbr1f/f; Ptenf/f; K14-CreERT2). Littermates were genotyped at 1 week of age and grouped based on genotypes for the experiments. Tamoxifen (200 μL of 10 μg/μL in corn oil) was administrated in the oral cavity of 3-month-old Tgfbr1/Pten 2cKO mice for 5 consecutive days to induce deletion of Tgfbr1 and Pten in epithelia to trigger the initiation of OSCC. When tumors initiated in the oral cavity, mice were switched to soft food and monitored daily. Histological slides were prepared to identify primary tumors.
For in vivo tumor growth, syngeneic mouse model was generated. Six-week-old male C3H/HeJ mice were purchased from Charles River (Beijing, China). The 1 × 106 SCC7 oral cancer cells were subcutaneously injected into flank of mice. Tumor size was measured with a caliper every three days. Tumor volume was calculated by width2 × length × 1/2. Once the tumor volume reached to 50 mm3, 20 mice were randomly divided into four groups (5 mice per group), receiving IgG (A2123, Selleck, China), IgG plus WRW4 (S9818, Selleck, China), anti-PD-1 antibody (A2122, Selleck, China) or WRW4 plus anti-PD-1 antibody respectively by intraperitoneal injection. The dose of IgG isotype control was 10 μg/injection diluted in 100 μL PBS. The dose of anti-PD-1 antibody was 200 μg/injection diluted in 100 μL PBS. The dose of WRW4 was 10 μg/injection diluted in 20 μL saline. Treatment was administrated once every two days for 6 times, then the tumors were dissociated and the weight as well as volume of tumors were measured. The maximal tumor size permitted by the IACUC of Chengdu Medical College is 20 mm at the largest diameter in mice. The maximal tumor size in our animal experiments did not exceed the permitted maximal tumor size. The mice were euthanized by using carbon dioxide when tumor volume reached 1500 mm³ or on day 20, whichever came first. 3 days after fixed in 4 % paraformaldehyde, tumors were embedded with paraffin for mIHC assay.
Statistics analysis
For experiments containing multiple groups, one/two-way ANOVA was used to statistical analysis following appropriate methods to compare two groups (Tukey/Bonferroni’s multiple comparisons test). Unpaired two-tailed student’s t-test was employed for comparing two groups. Studies involving animals displayed data from specified sample sizes. More detail information for statistics analysis can be found in the legend of each panel of figures. Statistical analyses were carried out using GraphPad Prism V8 software. Significance levels were denoted as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, and ‘ns’ indicating nonsignificant results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The processed expression data have been deposited in GEO dataset under accession number (GSE310797, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE310797). The original sequence data have been deposited in SRA under accession number (PRJNA1348242, https://www.ncbi.nlm.nih.gov/Traces/study/?acc=PRJNA1348242). All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
Code availability
The code used in this study is available on the Github dataset (https://doi.org/10.5281/zenodo.18804677). The implementation code used for neighborhood analysis is developed by a company. Detailed information can be obtained by contacting the corresponding author.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Flügge, T. et al. Detection of oral squamous cell carcinoma in clinical photographs using a vision transformer. Sci. Rep. 13, 2296 (2023).
Tan, Y. et al. Oral squamous cell carcinomas: state of the field and emerging directions. Int. J. oral. Sci. 15, 44 (2023).
Zhang, L. et al. Targets of tumor microenvironment for potential drug development. MedComm—Oncol. 3, e68 (2024).
Sun, L. et al. Single-cell and spatial dissection of precancerous lesions underlying the initiation process of oral squamous cell carcinoma. Cell Discov. 9, 28 (2023).
Schol, P. et al. Myeloid effector cells in cancer. Cancer Cell 42, 1997–2014 (2024).
Li, K. et al. Myeloid-derived suppressor cells as immunosuppressive regulators and therapeutic targets in cancer. Signal Transduct. Target. Ther. 6, 362 (2021).
Jiang, Y. et al. Prognostic role of CD11b(+) myeloid-derived suppressor cells in oral squamous cell carcinoma. Arch. Med. Sci. 19, 171–179 (2023).
Murciano-Goroff, Y. R., Warner, A. B. & Wolchok, J. D. The future of cancer immunotherapy: microenvironment-targeting combinations. Cell Res. 30, 507–519 (2020).
Zhang, R. et al. PMN-MDSCs modulated by CCL20 from cancer cells promoted breast cancer cell stemness through CXCL2-CXCR2 pathway. Signal Transduct. Target. Ther. 8, 97 (2023).
McArthur, S. et al. Definition of a novel pathway centered on lysophosphatidic acid to recruit monocytes during the resolution phase of tissue inflammation. J. Immunol. 195, 1139–1151 (2015).
McArthur, S. et al. Annexin A1 drives macrophage skewing to accelerate muscle regeneration through AMPK activation. J. Clin. Investig. 130, 1156–1167 (2020).
Erickson, H. L. et al. Cancer stem cells release interleukin-33 within large oncosomes to promote immunosuppressive differentiation of macrophage precursors. Immunity 57, 1908–1922.e1906 (2024).
Lee, C., Han, J. & Jung, Y. Formyl peptide receptor 2 is an emerging modulator of inflammation in the liver. Exp. Mol. Med. 55, 325–332 (2023).
He, F. et al. FPR2 shapes an immune-excluded pancreatic tumor microenvironment and drives t-cell exhaustion in a sex-dependent manner. Cancer Res. 83, 1628–1645 (2023).
Gavish, A. et al. Hallmarks of transcriptional intratumour heterogeneity across a thousand tumours. Nature 618, 598–606 (2023).
Liu, W. et al. Gap junction-mediated cell-to-cell communication in oral development and oral diseases: a concise review of research progress. Int. J. Oral. Sci. 12, 17 (2020).
Bae, Y. S. et al. Identification of peptides that antagonize formyl peptide receptor-like 1-mediated signaling. J. Immunol. 173, 607–614 (2004).
Wang, Y. et al. Proteotoxic stress response drives T cell exhaustion and immune evasion. Nature 647, 1025–1035 (2025).
Cui, C. et al. WNT2 blockade augments antitumor immunity by attenuating myeloid-derived suppressor cells in colorectal cancer. MedComm – Oncol. 3, e70004 (2024).
Lasser, S. A., Ozbay Kurt, F. G., Arkhypov, I., Utikal, J. & Umansky, V. Myeloid-derived suppressor cells in cancer and cancer therapy. Nat. Rev. Clin. Oncol. 21, 147–164 (2024).
Jones, C. L., Inguva, A. & Jordan, C. T. Targeting Energy Metabolism in Cancer Stem Cells: Progress and Challenges in Leukemia and Solid Tumors. Cell Stem Cell 28, 378–393 (2021).
Chen, K. et al. The formylpeptide receptor 2 (Fpr2) and its endogenous ligand cathelin-related antimicrobial peptide (CRAMP) promote dendritic cell maturation. J. Biol. Chem. 289, 17553–17563 (2014).
Liu, Y. et al. Cell surface receptor FPR2 promotes antitumor host defense by limiting M2 polarization of macrophages. Cancer Res. 73, 550–560 (2013).
De Santo, C. et al. Invariant NKT cells modulate the suppressive activity of IL-10-secreting neutrophils differentiated with serum amyloid A. Nat. Immunol. 11, 1039–1046 (2010).
Pang, X. et al. Myeloid-derived suppressor cells contribute to the malignant progression of oral squamous cell carcinoma. PLoS One 15, e0229089 (2020).
Ansari, J. et al. Targeting the AnxA1/Fpr2/ALX pathway regulates neutrophil function, promoting thromboinflammation resolution in sickle cell disease. Blood 137, 1538–1549 (2021).
Chen, Y. et al. Epithelial cells activate fibroblasts to promote esophageal cancer development. Cancer Cell 41, 903–918.e908 (2023).
Kiviaho, A. et al. Single cell and spatial transcriptomics highlight the interaction of club-like cells with immunosuppressive myeloid cells in prostate cancer. Nat. Commun. 15, 9949 (2024).
Li, M. et al. Spatial proteomics of immune microenvironment in nonalcoholic steatohepatitis-associated hepatocellular carcinoma. Hepatology 79, 560–574 (2024).
Song, L. et al. E2F1-initiated transcription of PRSS22 promotes breast cancer metastasis by cleaving ANXA1 and activating FPR2/ERK signaling pathway. Cell Death Dis. 13, 982 (2022).
Lee, S. et al. Cancer-associated fibroblasts activated by miR-196a promote the migration and invasion of lung cancer cells. Cancer Lett. 508, 92–103 (2021).
Wen, L. et al. Single-cell technologies: from research to application. Innovation 3, 100342 (2022).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Method of the Year 2020: spatially resolved transcriptomics. Nat. Methods 18, 1, https://doi.org/10.1038/s41592-020-01042-x (2021).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Gao, S. et al. Identification of HSC/MPP expansion units in fetal liver by single-cell spatiotemporal transcriptomics. Cell Res. 32, 38–53 (2022).
Bian, Y. et al. Loss of TGF-β signaling and PTEN promotes head and neck squamous cell carcinoma through cellular senescence evasion and cancer-related inflammation. Oncogene 31, 3322–3332 (2012).
Acknowledgements
We thank all the funders who supported the study and the investigators contributing to this work. This work was supported in part by National Natural Science Foundation of China (81903018, 82272825, 82472148). CMC Excellent-talent Program (2024qnGzn01), High-level Talent Cultivation Program of Chengdu Medical College (KYPY23-09), Key Clinical Specialty Construction Project of Sichuan Province (2024GXWKP002). Chengdu Medical College Clinical Joint Fund (24LHFYSZ1-30, 25LHGJ2-01). Chengdu Technological Innovation R&D Project (2024-YF05-01984-SN). Sichuan Science and Technology Education Joint Fund (2025NSFSC2105). Sichuan Provincial Natural Science Foundation (2022NSFSC1333, 2024NSFSC0759). The Key Project of Sichuan Science and Technology Program (2022YFS0622-A6, 2023YFS0178). The Applied Basic Research Project of Luzhou Science and Technology and Talent Work Bureau (2022-JYJ-123).
Author information
Authors and Affiliations
Contributions
F.L. and H.C. conceptualized. F.L. and Y.H. performed formal analysis, data visualization and drafted manuscript. F.O. were engaged in sample collection and data curation. L.D., H.L., X.Y., Y.Y. and R.M. performed the functional experiments. H.C., Z.Y. and Z.W. acquired funding for the study, reviewed and prepared the final manuscript. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, F., Han, Y., Ou, F. et al. Spatial heterogeneity of MDSCs mediated by ANXA1-FPRs signaling drives immune suppression in OSCC progression. Nat Commun 17, 2535 (2026). https://doi.org/10.1038/s41467-026-70861-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70861-x