Abstract
Sequential inflammatory stages characterizing early tuberculosis (TB) disease and reports of differentially culturable M. tuberculosis have compounded existing gaps in the detection of paucibacillary TB disease, threatening global elimination goals. Here we report unanticipated results we encountered while conducting early development work for an ultrasensitive molecular TB assay that has been validated in various cohorts of patients with suspected TB disease. Detection of M. tuberculosis DNA (TB-DNA) was confirmed by an alternate molecular target and sequencing. Over a six-year period, we conducted three separate clinical studies (N = 297) that tested two sets of anonymized respiratory samples from patients hospitalized in two Boston hospitals, and a longitudinal observational study to determine clinical associations and outcomes. We found an unexpectedly high prevalence of TB-DNA in US-born patients and a potential association with acute chest syndrome in patients with sickle cell disease. These results are preliminary and will require further study in prospective studies that include clinical, radiological, immunological, and microbiological correlation.
Similar content being viewed by others
Introduction
Tuberculosis (TB) remains one of the greatest threats to global health, productivity, and socioeconomic development, and recently superseded HIV and SARS-CoV-2 as the leading cause of death by an infectious agent worldwide1. The World Health Organization (WHO) has proposed an ambitious strategy to end the TB epidemic by 20352. In a comprehensive review of requirements for a TB-free world, improved diagnostics were identified as a top priority3. In 2022, the United States (US) Centers for Disease Control (CDC) reported 8331 notified cases of TB, over 600 TB-related deaths, and an estimated 13 million people with Mycobacterium tuberculosis infection4. While US disease incidence has steadily decreased, the rate of decline is too slow for elimination in this century5.
Despite recent advances2,3, improved understanding of pathogenesis and technological breakthroughs have highlighted several gaps in providing microbiological confirmation for the full range of TB disease clinical presentations2,6,7. The enduring deficiencies in diagnosis are largely due to our reliance on mycobacterial cultures to both detect and define clinically relevant TB disease3,8. Whereas cultures remain the most sensitive diagnostic method available, there is abundant evidence, both old and new, detailing their suboptimal performance for diagnosing several patient groups requiring antituberculous treatment. Most glaringly, there is an urgent need to increase the detection of paucibacillary TB disease in people living with HIV (PLHIV), children, and patients with extra-pulmonary TB. More recently, the recognition of sequential inflammatory stages that characterize early TB disease7,9,10,11 is concerning because these universally culture-negative conditions are usually undetected due to unconventional clinical presentations, and insensitive diagnostics12,13,14. Lastly, evidence of active TB disease caused by differentially culturable M. tuberculosis has been expanded using advanced imaging techniques8 and enriched mycobacterial culture media15. Taken together, these findings have complicated the diagnostic landscape and prompted calls to revisit TB disease case definitions to incorporate these “pre-diagnostic” stages14,16,17,18.
In this study, we report unanticipated results encountered while conducting early development work for the “Totally Optimized PCR (TOP) TB assay”—an ultrasensitive molecular assay with sensitivity superior to mycobacterial cultures that has been validated in various cohorts of patients with presumptive TB disease19,20. By interrogating respiratory samples for the presence of M. tuberculosis at a level of detection below that of currently available technologies, we detected M. tuberculosis DNA (TB-DNA) in an unexpectedly high proportion of hospitalized patients at a safety net hospital in Boston, Massachusetts. Although preliminary at this stage, we posit these data may represent the initial description of a previously unsuspected variant of paucibacillary TB disease that is of yet uncertain clinical and public health relevance.
Results
Anonymized studies
From May to September 2013 (Fig. 1), we obtained 226 anonymized respiratory specimens. After discarding low volume (<1 mL) and frankly watery samples, we stored 146 specimens in four batches of 42, 32, 43, and 29 samples each. Of these, 18 (12·3%) were TB-DNA+, and sequencing demonstrated assorted M. tuberculosis 2-ponA variants (Table S1). Of the 50 anonymized samples from the control population collected between May and July 2014, 1 (2%) was TB-DNA+ (p = 0.048).
We first collected anonimized respiratory samples from BMC (2013). Results from this study prompted an anonimized study in a control population (2014), and a second collection of coded samples from BMC (2014). The latter patients were followed longidudinaly (2015–2019). BMC = Boston Medical Center.
Longitudinal clinical study
From February to June 2014, we collected 104 coded respiratory specimens from 101 patients at BMC; duplicate samples were excluded. At the time of sample collection, all patients were hospitalized except one. The specimens were variable in terms of volume: 46% low (<1 mL) and 24% high (>15 mL) volume, and visual appearance: mucoid (29%), salivary (22%), purulent (15%), and 32% were blood-stained.
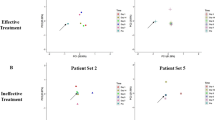
Table 1 and Table S2 show characteristics of the study cohort. Of the 101 samples tested, 16 (15.8%) were TB-DNA+, and sequencing results confirmed the presence of M. tuberculosis DNA in all except one (unable to sequence). The relative distribution of 2ponA variants was similar in both sets of BMC samples that were collected almost 1 year apart (Fig. 2). The mycobacterial culture requested retrospectively for TB-DNA+ cases was negative in all but one that resulted Mycobacterium avium. When clinical data were linked to TOP results (Fig. 3), age at admission separated TB-DNA+ results into one large group (n = 12, mean age 62), one small group (n = 3, mean age 24), and one teenage patient. The latter individual had previously presented to BMC over a decade earlier with fever and upper lobe infiltrates on chest radiography. The clinical summaries for the 16 TB-DNA+ individuals are shown in Table S3.
In addition to those shown in the graph, other discharge diagnoses of TOP-positive patients include brain hemorrhage (1), peripheral vascular disease (1), gastrointestinal hemorrhage (1), cocaine overdose (1), and cardiac device complication (1).
In unadjusted analyses, TB-DNA positivity was associated with sickle cell disease (p = 0.003), asthma (p = 0.05), and a longer period since first attending BMC (p = 0.002). Also, TB-DNA+ patients were more likely to undergo testing for TB infection before or during hospital admission (50% vs. 22%, p = 0.03) and were more likely to have a negative TST or IGRA result (44% vs. 5%, p < 0.001).
TB-DNA+ individuals were more likely to have an admission diagnosis for sickle cell disease pain crisis (p = 0.02) or chest pain (p = 0.06). During the index hospitalization, TB-DNA+ subjects were more likely to have anemia (p = 0.002), elevated direct bilirubin (p = 0.01), and leukocytosis (p = 0.06). At discharge, the only three subjects diagnosed with acute chest syndrome during the study period were all TB-DNA+ (3/16 vs. 0/85, p = 0.003).
Clinical outcomes
Clinical outcomes are shown in Table 2. During a median follow-up time of 1819 days (interquartile range [IQR] 1658–1842), none of the 16 TB-DNA+ subjects developed a clinical or microbiological diagnosis of TB disease. One TOP-negative patient was diagnosed with pulmonary TB disease (Xpert MTB/RIF positive) in January 2016.
The proportion of patients that died during the index admission was similar between TB-DNA+ and negative groups (p = 0.51). During follow-up, mortality among TB-DNA+ and TB-DNA– patients was 37.5% and 24.7%, respectively (p = 0.29), with death occurring at a median of 390 vs 26 days after hospital discharge, respectively. In an unadjusted Cox regression analysis (Fig. 4), there was no significant difference between the hazard of death in the first 650 days (Hazard Ratio [HR]: 1.2, 95% CI: 0.4–3.6); although the difference in hazards increased after 650 days, it did not achieve statistical significance (HR: 3.3, 95% CI: 0.4–26.7). Gender, alcohol use, and homelessness were not related to death, so our final multivariable Cox model adjusted for age and time since first hospital registration. These adjusted results were similarly non-significant: TB-DNA+ patients had a 10% increased hazard of death in the first 650 days, but this difference was not statistically significant (adjusted HR: 1.1, 95% CI: 1.0–1.2). After 650 days, the hazard increased, but still was not statistically significant (adjusted HR: 3.7, 95% CI: 0.5–30.3). Overall, TB-DNA+ patients were more likely to die from septicemia (67% vs. 14%; p = 0.02).
CI Confidence interval. All comparisons are two-sided.
Discussion
In this study, we detected an unexpectedly high prevalence of M. tuberculosis DNA in 12 to 16% of respiratory samples from predominantly US-born patients hospitalized in the principal safety net hospital in Boston with a variety of infectious and non-infectious syndromes. While unforeseen and preliminary, these results are plausible given: (1) TOP assay’s established diagnostic accuracy against a composite reference standard in over 400 patients with presumptive pulmonary TB from Uganda, Brazil and the US19,20; (2) the strength of our laboratory methods that demonstrate robust analytical sensitivity, and analytical specificity supported by DNA amplicon sequencing in both in-vitro conditions and clinical studies; (3) relatively high TOP optic density values (median 0.520) using a conservative cut-off, making false positive readings unlikely; (4) evidence of reproducibility by closely replicating both the point prevalence estimate, and the genetic distribution of M. tuberculosis 2ponA variants in two independent study samples obtained almost 1 year apart; (5) evidence of specificity by detecting a markedly lower positivity rate (2%) in a comparable cohort with a presumably lesser TB burden; (6) finding a clear bimodal distribution of positive results according to age—a key epidemiologic parameter of TB disease in the US4, and; (7) presence of known social, clinical and laboratory indicators of TB disease in TB-DNA+ individuals. Pending confirmation from larger multicenter studies, these findings open the possibility of an association—etiological, mechanistic, or non-causal- between M. tuberculosis and a variety of clinical syndromes in patients with a low pre-test probability of TB in the US.
The natural history of TB disease is a continuum that begins with the establishment of M. tuberculosis infection following an infectious exposure6,11. In individuals that are unable to contain bacterial replication, early disease progression occurs through consecutive asymptomatic or pauci-symptomatic inflammatory stages that are mostly culture-negative, eventually culminating in symptomatic illness that is usually confirmed by mycobacterial cultures6,7,9,11,12,13,14,16,21,22. However, bacterial replication kinetics are variable, often following one of several possible paucibacillary trajectories over time, including persistently low bacterial load, delayed progression, and even self-cure9,12,14,16. Recently developed molecular assays approved by the WHO (e.g., Xpert MTB/RIF Ultra and TrueNat) have a sensitivity comparable to culture and have significantly shorten the “sample-to-treatment” cascade of care, but they are still best suited for the detection of TB disease at later stages and have had limited impact in improving treatment outcomes23. Responding to a need for more sensitive tools to identify early stages of TB disease2,5,21, recent development efforts have focused on blood-based gene signatures and inflammatory biomarkers that have been tightly calibrated against culture-confirmed TB disease. However, these biosignatures have demonstrated sub-optimal diagnostic accuracy and reproducibility problems across populations2,10,21,22. In contrast, molecular detection methods with sensitivity superior to culture, such as the TOP assay, have a long track record of delivering clinically useful results, often replacing culture as the preferred diagnostic modality for multiple organisms and sample types24. This study provides further impetus to reconsidering the standard practice of anchoring new TB diagnostic studies to a microbiological yardstick defined by culture, as such evaluations may be flawed by design3,20,21. This focus on demonstrating bacterial viability with cultures (or other means) as a prerequisite to defining clinically relevant TB disease also limits the interpretation of our results within the framework of the recent “International Delphi Consensus Classification of Early Tuberculosis States”17,18.
While certainly unforeseen, several results from the present study align well with currently available knowledge. First, 75% of TB-DNA+ patients were 50 years or older, a finding that is consistent with the epidemiology of TB in the US, where the rate of TB disease is highest in older individuals, and 85% of new TB cases are related to reactivation from a remote exposure4,25,26. This result may be best understood by applying the “iceberg principle,” where this ultrasensitive assay is simply detecting additional TB cases that are currently hidden below the level of detection of current technologies. Similarly, the clinical description of the TB-DNA+ adolescent is consistent with unstable TB infection or early TB disease6,11, as he first presented to BMC when he was <5 years of age with a classic presentation of pediatric TB (e.g., high-grade fever and upper lobe infiltrate on chest radiograph) while living in homeless shelters. Further, certain characteristics of TB-DNA+ patients have long been associated with TB risk, such as previous or repeated testing for TB infection, and anemia27.
Consequently, upon careful consideration, the most unexpected finding of this study is the detection of M. tuberculosis DNA in the sputum of the only three patients with sickle cell disease that shared a diagnosis of acute chest syndrome (3/16 vs. 0/85, p = 0.003), a finding that should be considered preliminary given the small sample size despite strong statistical significance. The increased risk of infectious complications in patients with sickle cell is well established, particularly when complicated by acute chest syndrome28,29. In their seminal study, Vichinsky et al. found that among 671 episodes of acute chest syndrome, 216 (32%) were caused by infections from 27 different microorganisms, predominantly atypical bacteria and viruses, but 2 (0.3%) cases were attributed to M. tuberculosis28. Importantly, in 46% of patients, the etiology of acute chest syndrome was undetermined (e.g., no evidence of infection, fat embolism, or infarction); in the present study, all three patients had a negative concomitant microbiological work-up. Although US populations at risk for TB and those at risk for sickle cell disease may have overlapping demographic and social characteristics, a definitive link between these two illnesses has not been established28. If confirmed in larger studies, this previously unsuspected clinical association may be particularly relevant to settings where both maladies are more prevalent, such as in sub-Saharan Africa29.
After a dramatic decline in the number of TB cases in the US over the last 25 years, there is growing evidence the epidemic has entered a period of non-declining disease rates4, particularly following the COVID period4. Complicating elimination, 87% of new cases arise from the large reservoir of individuals with untreated TB infection4. When individuals with TB infection escape latency, establishing a diagnosis of TB disease is often challenging because of evolving risk factors, atypical clinical presentations following immunotherapy, and a growing need for detection of culture-negative disease2,9,13,16. In 2022, 21% of TB cases reported to CDC were not microbiologically confirmed (e.g., clinical TB case definition with no M. tuberculosis viability demonstrated), and this proportion has slowly increased over time due to epidemiological and biological factors4,13,19. Interestingly, most TB-DNA+ patients in this study were TST or IGRA-negative, a poorly understood condition that has been associated with advanced age, low peripheral lymphocyte counts, and poor clinical outcomes in microbiologically confirmed TB cases in the U.S. and elsewhere30,31. This observation is also frequently reported in presumptive TB patients that are Xpert MTB/RIF Ultra trace-positive, a semi-quantitative diagnostic category measuring very low bacillary loads that are often culture-negative32.
Currently, isolated detection of M. tuberculosis DNA is not universally accepted as evidence of TB disease20 but this opinion is evolving with the growing body of evidence provided by presumptive TB patients that are Ultra trace-positive33. Although preliminary, the results of this study lead to several mechanistic, clinical, and infection control questions that will require further investigation to elucidate. First, detection of M. tuberculosis DNA in certain patients may be clinically inconsequential, as it may be capturing non-viable bacteria or represent shedding from a latent focus8. Yet, disregarding evidence of M. tuberculosis DNA in respiratory samples without further study is surely unwise. Second, if linear disease progression culminating in culture-positive illness is indeed the common final stage of TB, our findings may suffer from insufficient follow-up time to culture-positive conversion. An alternative explanation is that disease progression was interrupted or modified by partial treatment with beta-lactam antibiotics (81% of TB-DNA+ patients received them), given their well-described antituberculous activity34. Finally, we hypothesize that this new diagnostic tool may have permitted the discovery of a previously undetectable variant of paucibacillary TB disease that stands as a polar opposite to the traditional, culture-positive multibacillary form of the disease. This presumed bipolar TB pathogenesis model that has been proposed by others35, would have a strong biological precedent, as shown by other closely related mycobacteria such as M. leprae35 and certain nontuberculous mycobacteria36. One possible mechanistic explanation for this bipolar spectrum of TB disease is our household contact study from Brazil, showing that less transmissible M. tuberculosis clinical isolates cause diffuse inflammation in C3HeB/FeJ mice lung pathology when compared to highly transmissible strains that exhibited more caseating granulomas, a lesion type with high potential to cavitate and therefore cause multibacillary TB disease37.
Our study has limitations. The TOP TB assay is a research-use-only tool that currently lacks regulatory clearance. Also, our results are challenged by the absence of a confirmatory independent test, as currently available diagnostic methods do not have the requisite sensitivity to detect the very low bacterial loads ascertained by the TOP assay (i.e., culture, other molecular assays and metagenomic sequencing), or because they depend on M. tuberculosis growth in culture to sufficiently enrich the molecular target (i.e., whole-genome sequencing). The use of discarded respiratory samples in all three studies is not ideal because it may have affected the quality and volume of specimens, and risk sample contamination. Whereas the likelihood that cross-contamination or insufficient assay specificity may have resulted in false-positive results is low, it cannot be completely excluded. Because of the current diagnostic uncertainty of detecting M. tuberculosis DNA in culture-negative samples, we cannot certify M. tuberculosis directly contributed to symptoms, pathology, or death in the absence of tissue or autopsy results. The latter limitation also needs to be considered when interpreting the positive predictive value of the TOP assay, given its heightened sensitivity and low pre-test probability of disease in the study cohort. Finally, while passive follow-up through serial chart review provided detailed and reliable information, some additional characteristics might not have been identified. Future prospective clinical studies with dedicated respiratory specimens will be necessary to confirm and expand our findings.
In summary, we detected a higher-than-expected prevalence of M. tuberculosis DNA in respiratory samples of predominantly US-born patients hospitalized in a Boston safety-net hospital. Whereas most TB-DNA+ patients were older and presented with a variety of infectious and non-infectious syndromes commonly encountered in hospitalized individuals, there was also a group of four young patients, three of which shared the diagnosis of acute chest syndrome—a striking and potentially consequential clinical association. While preliminary, we hypothesize that our findings indicate the existence of a paucibacillary form of TB that remains unrecognized and is not detectable using current diagnostic tools. These results will require confirmation in larger prospective studies that include clinical, radiological, immunological, and microbiological correlation. Yet, even without complete knowledge of biological mechanisms, impact on clinical outcomes, or transmissibility, the evidence presented supports dissemination given the potential implications for medical care and public health in the US, and elsewhere.
Methods
Ethical approvals
The studies were approved (protocols H-32288 and H-32655) by the Boston University Medical Center Institutional Review Board (IRB) as “Exempt” (e.g., did not meet the definition of Human Subjects Research). The IRB waived the need to obtain informed consent. The IRB at St Elizabeth Medical Center approved the study without further review.
Study population
This report includes three separate studies conducted over 6 years at two separate hospitals in Boston, Massachusetts (Fig. 1). Two of the studies were performed at Boston Medical Center (BMC), a private 496-bed academic medical center that serves as the principal safety net hospital for the greater Boston metropolitan area (57% under-served populations). The Clinical Microbiology Laboratory at BMC is state-of-the-art, providing a full range of clinical diagnostic infectious diseases testing, performing about 2·8 million tests per annum. The laboratory has a successful record of supporting pre-clinical and clinical research, including the provision of clinical samples and microbial isolates, and performing comparative assay evaluations.
Over the 3 years preceding this study (2011–2013)38, the Boston Public Health Commission (BPHC) registered 40–45 new TB cases per year (incidence rate 6·5–7·1 per 100,000 population). Of these, 65% had pulmonary TB, 80% were non-US born, 8–9% were in PLHIV, and 65–77% were diagnosed while hospitalized38. BMC reports an average of 2–3 new TB cases monthly, corresponding to 50–65% of notified cases in Boston.
Control population
To understand the specificity of results at BMC, we sourced a parallel set of anonymized discarded respiratory samples with identical inclusion criteria from patients hospitalized at St Elizabeth’s Medical Center, a private 272-bed community hospital located in Boston’s Brighton neighborhood. Within the hospital’s primary service area, residents are predominantly white (79%), followed by Asian (11%) and Black (5%). The age distribution reflected the large student population, as 24% were 20–29 years of age, and 12% were ≥65 years. Per BHPC records, from 2009 to 2013 St Elizabeth’s Medical Center reported 11 hospitalizations for evaluation of TB disease compared to BMC that reported 141 during the same 5-year period preceding this study.
TOP TB assay
A detailed description of the TOP TB assay has been reported19,20. Briefly, the assay targets a gene (ponA1) involved in the assembly of peptidoglycans in the M. tuberculosis bacterial wall. The assay’s diagnostic primer set (3-ponA-F/R) targets sequences unique to all species in the M. tuberculosis complex. Amplicons generated by 3-ponA are detected using a capture-probe colorimetric assay, and the resultant Optical Densities (OD) provide a semi-quantitative measurement of bacillary load19. TOP TB assay 3-ponA primers and probe have demonstrated no cross-reactivity with respiratory pathogens and nontuberculous mycobacteria, and the signal amplified is specific to M. tuberculosis19,20. Specimens that are positive using the diagnostic primer set were tested with a second primer set (2-ponA-F/R) that serves both as an alternate molecular target39, and to distinguish among five possible genetic variants of M. tuberculosis (e.g., 0T, 1T/1, 2, 3, and 4), as previously described19.
Anonymized respiratory specimens
During early assay development, we sourced anonymized discarded respiratory samples sourced in sterile containers that were submitted for routine clinical testing to the BMC laboratory. These samples were originally intended as a real-world clinical specimen matrix for spiking M. tuberculosis in decreasing concentrations (e.g., serial dilutions)39. To minimize the prevalence of M tuberculosis, we included only samples without a mycobacterial laboratory request by the ordering clinician. After routine microbiology work that included pipetting of specimens to inoculate standard bacterial cultures was completed, the remnant aliquot was refrigerated unprocessed for up to 1 week to enable add-on testing as requested by clinicians. Specimens were then transported weekly to the research laboratory in a temperature-controlled container where they were frozen unprocessed for ~3 months in four batches, each containing 29–43 samples. Before performing serial dilution experiments, specimens were thawed, processed, and tested with the TOP TB assay to ensure a negative baseline result. As an additional measure to minimize cross-contamination, all PCR and DNA extraction reagents were discarded, and new reagents utilized each week. Detection of TB-DNA in blank study samples was verified by repeat testing, and M. tuberculosis sequencing (Genewiz, NJ, USA), as described19. We followed the same laboratory procedures for a parallel set of anonymized samples sourced from patients hospitalized at St Elizabeth’s Medical Center (e.g., control population).
Longitudinal clinical study
Because of the unexpected results in anonymized samples at BMC, we then conducted an observational longitudinal study to determine pre-specified clinical outcomes, including incident microbiological or clinical TB diagnosis, hospital admissions, and vital status. For this purpose, we sourced a second set of respiratory specimens with identical inclusion criteria as the first anonymized BMC study, but this time assigning each sample a study identifier. We followed the same sample selection, transport, and storage methods as in the anonymized study. However, in this study, selected specimens underwent TOP testing each week to retrospectively perform mycobacterial cultures in TB-DNA+ samples. Coded study results were later linked to relevant patient data, including any previous TB-related history (i.e., exposures, BCG vaccination, TST, IGRA, TB disease, and treatment), as well as social, clinical, microbiological, and radiographic data after independent chart review by personnel unrelated to the research study. An independent clinician conducted annual reviews of BMC hospital charts and BPHC records to capture a subsequent clinical or microbiological diagnosis of TB disease, hospital readmissions, and to determine vital status. Date and cause of death data were extracted from the medical chart or death certificate when available.
Analytical strategy
This is an observational study reporting unanticipated results in a population with a presumably low pre-test probability of pulmonary TB disease; as such, we considered it inappropriate to report results according to the Standards for Reporting of Diagnostic Accuracy guidelines40. We calculated the diagnostic cut-off for TOP OD using a value of three standard deviations above the mean of the OD values of negative controls (e.g., laboratory cut-off), after validating the latter method using a Receiver Operating Curve analysis19. We compared patient characteristics according to TOP results using either the Wilcoxon test (continuous data) or Fisher’s exact test (categorical data) all two-sided. We followed patients for clinical outcomes (incident TB, readmission, or death) until April 2019. We show Kaplan Meier curves and use an unadjusted stratified Cox proportional hazards model to determine if there are differences in the time to death for TB-DNA+ and negative patients before and after 650 days. We also fit a Cox proportional hazards model, considering adjustment for homelessness, alcohol use, age, biological sex, and time since first hospital visit, and use backwards selection to select a parsimonious model. All comparisons are two-sided, and we did not adjust for multiple comparisons
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for Fig. 2 is included in Table S1 and Table S3. Source data for the longitudinal clinical study pertains to individual clinical information that was extracted by the Boston Medical Center data warehouse into Excel sheets using study codes to maintain patient confidentiality and fulfill Health Insurance Portability and Accountability Act (HIPAA) requirements. Therefore, widespread access to this data is restricted due to patient privacy. Upon appropriately justified requests, coded clinical data may be made available within 60 days after approval by the Boston University Medical Center Institutional Review Board. Please contact the corresponding author (EJ-L) for any data requests. The DNA sequences for the 3ponA and 2ponA primer sets are available in Supplementary Fig. III and Supplementary Table IV in the Appendix of reference #19 (Madico et al., PLoS One, 2016).
References
World Health Organization. Global tuberculosis report 2023. License: CC BY-NC-SA 3.0 IGO (World Health Organization, Geneva, 2023).
Keshavjee, S. et al. Moving toward tuberculosis elimination. Critical issues for research in diagnostics and therapeutics for tuberculosis infection. Am. J. Respir. Crit. Care Med. 199, 564–571 (2019).
Reid, M. J. A. et al. Building a tuberculosis-free world: the Lancet Commission on tuberculosis. Lancet https://doi.org/10.1016/S0140-6736(19)30024-8 (2019).
Schildknecht, K. R., Pratt, R. H., Feng, P. I., Price, S. F. & Self, J. L. Tuberculosis - United States, 2022. MMWR Morb. Mortal. Wkly. Rep. 72, 297–303 (2023).
Bayer, R. & Castro, K. G. Tuberculosis elimination in the United States—the need for renewed action. N. Engl. J. Med. 377, 1109–1111 (2017).
Salgame, P., Geadas, C., Collins, L., Jones-Lopez, E. & Ellner, J. J. Latent tuberculosis infection—revisiting and revising concepts. Tuberculosis 95, 373–384 (2015).
Scriba, T. J. et al. Sequential inflammatory processes define human progression from M. tuberculosis infection to tuberculosis disease. PLoS Pathog. 13, e1006687 (2017).
Dartois, V., Saito, K., Warrier, T. & Nathan, C. New evidence for the complexity of the population structure of Mycobacterium tuberculosis increases the diagnostic and biologic challenges. Am. J. Respir. Crit. Care Med. 194, 1448–1451 (2016).
Drain, P. K. et al. Incipient and subclinical tuberculosis: a clinical review of early stages and progression of infection. Clin. Microbiol. Rev. 31, https://doi.org/10.1128/CMR.00021-18 (2018).
Weiner, J. et al. Metabolite changes in blood predict the onset of tuberculosis. Nat. Commun. 9, 5208 (2018).
Esmail, H., Barry, C. E. III, Young, D. B. & Wilkinson, R. J. The ongoing challenge of latent tuberculosis. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 369, 20130437 (2014).
Achkar, J. M. & Jenny-Avital, E. R. Incipient and subclinical tuberculosis: defining early disease states in the context of host immune response. J. Infect. Dis. 204, S1179–S1186 (2011).
Nguyen, M. H. et al. Factors associated with sputum culture-negative vs culture-positive diagnosis of pulmonary tuberculosis. JAMA Netw. Open 2, e187617 (2019).
Dowdy, D. W., Basu, S. & Andrews, J. R. Is passive diagnosis enough? The impact of subclinical disease on diagnostic strategies for tuberculosis. Am. J. Respir. Crit. Care Med. 187, 543–551 (2013).
Gap-Gaupool, B. et al. Nitric oxide induces the distinct invisibility phenotype of Mycobacterium tuberculosis. Commun. Biol. 7, 1206 (2024).
Houben, R. et al. Spotting the old foe-revisiting the case definition for TB. Lancet Respir. Med. 7, 199–201 (2019).
Coussens, A. K. et al. Classification of early tuberculosis states to guide research for improved care and prevention: an international Delphi consensus exercise. Lancet Respir. Med. 12, 484–498 (2024).
Dheda, K., Perumal, T. & Fox, G. J. Asymptomatic tuberculosis: undetected and underestimated, but not unimportant. Lancet 405, 1797–1800 (2025).
Madico, G. et al. Detection and quantification of Mycobacterium tuberculosis in the sputum of culture-negative HIV-infected pulmonary tuberculosis suspects: a proof-of-concept study. PLoS ONE 11, e0158371 (2016).
Madico, G. et al. Further evidence of Mycobacterium tuberculosis in the sputum of culture-negative pulmonary tuberculosis suspects using an ultrasensitive molecular assay. Tuberculosis 116, 1–7 (2019).
Walzl, G. et al. Tuberculosis: advances and challenges in development of new diagnostics and biomarkers. Lancet Infect. Dis. 18, e199–e210 (2018).
Warsinske, H., Vashisht, R. & Khatri, P. Host-response-based gene signatures for tuberculosis diagnosis: a systematic comparison of 16 signatures. PLoS Med. 16, e1002786 (2019).
Bonnet, M. Xpert MTB/RIF Ultra: what is the real impact?. Lancet Respir. Med. 8, 325–326 (2020).
Miller, J. M. et al. A guide to utilization of the microbiology laboratory for diagnosis of infectious diseases: 2018 update by the Infectious Diseases Society of America and the American Society for Microbiology. Clin. Infect. Dis. 67, 813–816 (2018).
Hochberg, N. S. & Horsburgh, C. R. Jr Prevention of tuberculosis in older adults in the United States: obstacles and opportunities. Clin. Infect. Dis. Publ. Infect. Dis. Soc. Am. 56, 1240–1247 (2013).
Whalen, C. C. The replacement principle of tuberculosis. Why prevention matters. Am. J. Respir. Crit. Care Med. 194, 400–401 (2016).
Gil-Santana, L. et al. Tuberculosis-associated anemia is linked to a distinct inflammatory profile that persists after initiation of antitubercular therapy. Sci. Rep. 9, 1381 (2019).
Vichinsky, E. P. et al. Causes and outcomes of the acute chest syndrome in sickle cell disease. National Acute Chest Syndrome Study Group. N. Engl. J. Med. 342, 1855–1865 (2000).
Uyoga, S. et al. The epidemiology of sickle cell disease in children recruited in infancy in Kilifi, Kenya: a prospective cohort study. Lancet Glob. Health 7, e1458–e1466 (2019).
Nguyen, D. T., Teeter, L. D., Graves, J. & Graviss, E. A. Characteristics associated with negative interferon-gamma release assay results in culture-confirmed tuberculosis patients, Texas, USA, 2013-2015. Emerg. Infect. Dis. 24, 534–540 (2018).
Yamasue, M. et al. Factors associated with false negative interferon-gamma release assay results in patients with tuberculosis: a systematic review with meta-analysis. Sci. Rep. 10, 1607 (2020).
Sung, J. et al. Evidence for tuberculosis in individuals with Xpert Ultra “Trace” sputum during screening of high-burden communities. Clin. Infect. Dis. 78, 723–729 (2024).
Sung, J. et al. The long-term risk of tuberculosis among individuals with Xpert Ultra “trace” screening results: a longitudinal follow-up study. medRxiv https://doi.org/10.1101/2025.03.20.25324205 (2025).
Diacon, A. H. et al. Beta-Lactams against Tuberculosis–New Trick for an Old Dog?. N. Engl. J. Med. 375, 393–394 (2016).
Modlin, R. L. & Bloom, B. R. TB or not TB: that is no longer the question. Sci. Transl. Med. 5, 213sr216 (2013).
Pan, S. W. et al. Microbiological persistence in patients with Mycobacterium avium complex lung disease: the predictors and the impact on radiographic progression. Clin. Infect. Dis. 65, 927–934 (2017).
Verma, S. et al. Transmission phenotype of Mycobacterium tuberculosis strains is mechanistically linked to induction of distinct pulmonary pathology. PLoS Pathog. 15, e1007613 (2019).
Boston Public Health Commission, Research and Evaluation Office. Health of Boston 2012–2013: A Neighborhood Focus. (Boston, MA, 2013).
Clinical and Laboratory Standards Institute. Establishing Molecular Testing in Clinical Laboratory Environments; Proposed Guideline. CLSI document MM19-P. (Clinical and Laboratory Standards Institute, Wayne, PA, 2011).
Bossuyt, P. M. et al. Towards complete and accurate reporting of studies of diagnostic accuracy: the STARD initiative. Ann. Intern. Med. 138, 40–44 (2003).
Acknowledgements
This study was funded with internal funds. LFW was funded by the Providence/Boston Center for AIDS Research (P30AI042853). The authors acknowledge the invaluable contributions made by Carol Sulis, MD for independently reviewing patient charts at Boston Medical Center. We also thank Claudia Nader, MD, and Lisa Zenkin, PhD for providing samples from St Elizabeth’s Medical Center, and Mr. Albert Magode for help with data management. We thank the personnel at Boston Medical Center’s Office of Development (Kirsten Hinsdale and Hugh Keeping) and Boston’s University Office of Technology Development (Michael Pratt, Lee Tien) for their continued support, and Mr. Joe Straight who was instrumental in establishing an assay development strategy. We also wish to acknowledge Michael P. Dube, MD, Leo Martinez, PhD, Jerrold J. Ellner, MD, Mariana J. Kaplan, MD, and Carlos Acuña-Villaorduña, MD for critically reviewing the manuscript, and Ms. Camila Jones-Martinez for assistance in editing.
Author information
Authors and Affiliations
Contributions
Conception and Design: E.J.-L., N.M., and G.M.; Acquisition of Data: G.M., B.O., N.M., and E.J.-L.; Analysis and Interpretation: E.J.-L., L.W., and G.M. Also, S.V., M.M., P.O., J.M.-A., M.P., R.D., and Y.B. participated in early development of the TOP TB assay. All authors contributed to either drafting or revising this manuscript and gave final approval.
Corresponding author
Ethics declarations
Competing interests
Dr. Jones-López and Dr. Madico are co-inventors of TOP TB assay’s primers and probes. The invention is owned by Boston Medical Center/Boston University Medical School, which has obtained an international patent application (US Patent and Trademark Office No. 10,190,176 B2). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tobias Broger and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jones-López, E.C., Miller, N.S., Orr, B. et al. Unexpected detection of Mycobacterium tuberculosis DNA in US-born patients in putative association with clinical syndromes. Nat Commun 17, 2709 (2026). https://doi.org/10.1038/s41467-026-70890-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70890-6



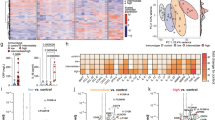

![Fig. 2: TOP TB assay results in respiratory specimens sourced at Boston Medical Center for the anonymized study (May/2013 to September/2013; TOP-positive = 18 [12.3%]) and the longitudinal clinical study (February/2014 to June/2014; TOP-positive = 16 [15.8%]).](http://media.springernature.com/lw685/springer-static/image/art%3A10.1038%2Fs41467-026-70890-6/MediaObjects/41467_2026_70890_Fig2_HTML.png)

