Abstract
Plants use the plant hormone auxin to incorporate environmental cues into their growth and development to shape the final form. Temperature is an important modulator of all aspects of plant function and growth. In this work, we uncover temperature-regulated accumulation and solubility of members of the AUXIN RESPONSE FACTOR transcription factor family. We determine that ARF7 and ARF19 proteins rapidly hyperaccumulate in response to elevated temperature. Furthermore, we find that diffuse concentrations of ARF protein increase under elevated temperature, consistent with increased solubility. Temperature-driven ARF hyperaccumulation is not fully dependent on the well-established temperature response pathways. We find that natural variation in thermoregulated ARF accumulation is correlated with thermomorphogenesis, suggesting that this is a dial switch in plant temperature response. Regulated ARF thermoaccumulation provides a layer of complexity in shaping and plant growth and form, allowing plants to respond rapidly and persistently to elevated temperatures by modulating levels of nuclear ARF protein accumulation.
Similar content being viewed by others
Introduction
Although plants start their lives with a simple body plan, to adjust their shape to the prevailing conditions, they must incorporate environmental information into developmental and physiological decision-making. A critical cue in this process is environmental temperature, which provides critical information about daily and seasonal rhythms and the risk of stressful conditions. Plants respond to elevated temperature in a suite of growth changes characterized as thermomorphogenesis1. In addition, a myriad of physiological changes enables plants to adjust to changing temperatures2,3. Understanding the mechanisms used by plants to optimize their body shape and physiological responses in response to increased temperature is particularly important as our global temperatures continue to rise.
Elevated temperatures input through multiple pathways, seemingly converging on increased auxin biosynthesis4,5. Indeed, functional auxin receptors TIR1/AFB, which work in conjunction with IAA3, and IAA19 co-receptors, are required for thermomorphogesis6,7,8,9. Auxin-driven association of these Aux/IAA repressors with TIR1/AFB F-box proteins promotes their polyubiquitylation and degradation, relieving ARF proteins from their repression10. Auxin transcriptional output is driven by the AUXIN RESPONSE FACTOR (ARF) transcription factor family, which is repressed by the Aux/IAA proteins in the absence of auxin. Although these auxin-signaling components are necessary for thermomorphogenesis, whether ARFs act simply as interpreters of elevated auxin levels or if they also represent direct targets of temperature cues remains poorly understood.
In addition to Aux/IAA-mediated repression, activity of ARF7 and ARF19, and perhaps additional ARFs, is also attenuated by protein condensation11, which is a rapid and reversible compartmentalization of proteins12. Protein condensation is a concentration-dependent process in which proteins in excess of a saturation concentration are stored in dense bodies, or condensates12. This saturation concentration is modified by environmental conditions, posttranslational modifications, and interaction partners, allowing for rapid and dynamic compartmentalization of cellular components. ARF7 and ARF19 condensation is regulated by unknown developmental regulators11, interaction with MCTP proteins13, and condensate movement14. Further, regulated ARF accumulation is increasingly recognized as a regulator of auxin response15,16,17.
Here, we uncover temperature-regulated changes in ARF protein accumulation, stability and solubility. Further we find that ARF7 and ARF19 temperature-regulated accumulation correlates with thermomorphogenesis in natural Arabidopsis accessions. This ability of ARF proteins to intrinsically respond to rapid changes in temperatures adds a new level of complexity to the auxin transcriptional response and incorporation of temperature cues into plant growth.
Results
Class-A ARF protein accumulation is dynamically regulated by temperature
ARF protein stability is emerging as an important regulator of auxin response15,18. To understand regulators of ARF protein stability, we developed a custom antibody, raised against the ARF7 PB1 domain (Figure S1), which recognizes both ARF7 and ARF19 proteins. Using this antibody, we found that endogenous levels of ARF7 and ARF19 rise with increasing temperatures (Fig. 1a). ARF7 and ARF19 transcripts are unaffected by temperature changes (Supplementary Table 1) and are thus unlikely to be the driver of differential protein accumulation at varying temperatures. Further, when driven behind constitutive promoters, ARF7 and ARF19 protein accumulation varies with temperature (Fig. 1b–f, Figure S2a), suggesting that translation and/or stability of these proteins is temperature-regulated.
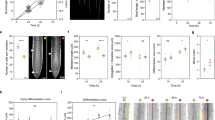
a–c Immunoblot analysis of 4-day-old seedlings after 4-hour treatment with indicated temperature. Each experiment was performed at least three times; additional replicates are in Supplemental Fig. 1b. d Time course fluorescence microscopy images of the hypocotyl of a seedling expressing pUBQ10:YFP-ARF19 after transfer from 22°C to 32 °C. e Quantification of time course fluorescence microscopy images of the hypocotyl of a seedling expressing pUBQ10:YFP-ARF19 after transfer from 22 °C to 32 °C (n = 6). Error bars represent standard deviation of the mean. Arbitrary units are used. P-value was calculated using a t-test of the linear regression with the df = n-2; p = 7.52E-23. f Cladogram of Arabidopsis Class-A ARFs. g Immunoblot analysis of 4-day-old seedlings carrying ARF5:ARF5-GFP, ARF6:ARF6-GFP, or ARF8:ARF8-GFP after 4 h treatment at 32 °C. Each experiment was performed at least three times; additional replicates are in Supplemental Fig. 1b.
In Arabidopsis, there are five Class-A ARFs (Fig. 1f), which are thought to positively drive auxin-responsive gene expression10. ARF7 and 19 are closely-related, with ARF5 as the next most-closely related to ARF7 and 19. ARF6 and 8 are additional Class-A ARFs in a separate clade from ARF7, 19, and 519. We examined these additional Class-A ARFs and found that ARF5, ARF6, and ARF8, similar to ARF7 and ARF19, undergo a dramatic increase in accumulation in response to elevated temperature (Fig. 1g). Thus, all Arabidopsis Class-A ARFs display thermoresponsive protein accumulation. Because our lab has many ARF7- and ARF19-related resources11,14,15,20,21,22, including our newly-developed custom antibody (Figure S1), we proceeded with studies focused primarily on ARF7/19.
ARF protein stability is regulated by temperature
To assess ARF protein stability, we developed a flow cytometry-based ratiometric assay. In this assay in which protoplasts transfected with a ratiometric reporter encoding a P2A self-cleaving peptide (pUBQ10:mNEON-ARF19-P2A-mScarlet) allowed the measurement of two separate fluorescent proteins encoded by a single mRNA transcript (Fig. 2a, b). Using this assay, we found that ARF19 displayed increased stability at 32 °C compared to 22 °C (Fig. 2b).
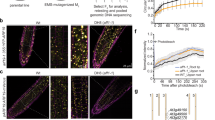
a Graphic depicting the ARF stability ratiometric construct that results in expression of two proteins. 16 hours after protoplast transfection, protoplasts are incubated at 22 °C or 32 °C for 4 h and fluorescence measured. b Mean ( + SE) ratio of mNeon:mScarlet signal after 4-hour treatment (n = 2652 for ARF19 at 22 °C, 1156 for ARF19 at 32 °C, 2840 for empty vector at 22 °C, and 2479 for empty vector at 32 °C), letters indicate a significance of p < 0.001, 2-way ANOVA with Tukey’s HSD test. c Graphic showing cycloheximide (CHX) treatment. d Immunoblot analysis of 5-day-old seedlings treated with the indicated temperature, time, and CHX treatment. e Immunoblot analysis of 4-day-old 35S:ARF7-HA seedlings incubated 4 h at the indicated temperature in the presence or absence of MG132. f Immunoblot analysis of four-day-old wild type (Col-0) and aff1 seedlings treated as indicated. Each immunoblot experiment was performed at least three times; additional replicates are in Supplemental Fig. 2g.
We found that ARF7/19 protein displays a longer half-life at 32 °C compared to 22 °C using a cycloheximide chase experiment (Fig. 2c, d). We then treated seedlings at elevated temperature for four hours prior to a four-hour recovery at 22 °C, with samples taken before and after elevated temperature exposure and after the recovery period (Figure S2b–e). In these assays, ARF7 and ARF19 protein accumulation largely returned to baseline levels after transition back to 22 °C. This incomplete recovery suggests that protein levels do not revert to their original levels at the same rate that they accumulate. Overall, our data show that ARF7 and ARF19 protein accumulation is dynamically regulated as a function of temperature; these effects are driven by altered protein stability.
Next, we examined protein degradation mechanisms that might control temperature-regulated ARF protein accumulation. To date, we know that ARF protein accumulation is regulated by the 26S proteasome18 and by autophagy23. ARF7 levels increased at elevated temperature in both the absence and presence of a proteasome inhibitor (Fig. 2e), suggesting that 26S proteasomal activity is not a major driver of the effects of temperature on ARF stability. Aligned with this result, we found that ARF7/19 accumulation was equally temperature-regulated in wild type and in the aff1 mutant (Figs. 2f, S3b), which is defective in an E3 ubiquitin ligase that targets ARF7 and ARF19 to the proteasome24. Further, temperature effects on ARF7 accumulation were unaffected by the autophagy inhibitor Concanamycin A (Figure S3a), suggesting that this pathway is also unnecessary for the altered ARF accumulation in response to changing temperatures. Finally, to determine if the chaperone protein HSP90 impacted ARF temperature-regulated stability, we assessed ARF7 protein levels in the presence of HSP90 inhibitor Glendamycin A (Figure S3c), finding that ARF7 accumulation remained thermoresponsive in the presence of this inhibitor, suggesting that HSP90 is dispensable for temperature effects on ARF accumulation. Given these results, we postulate that ARF thermostability could be regulated by alternative mechanisms such as interaction with chaperones other than HSP90 or allosteric regulation not captured in the experiments performed.
ARF19 thermoresponsive accumulation is conferred by the DBD and MR regions
We were curious what aspects of ARF7 and ARF19 drove thermoresponsive protein accumulation. To determine which ARF19 regions confer thermoaccumulation properties, we examined ARF19 truncations consisting of the DNA binding Domain (DBD, ARF191–354), the PB1 domain (ARF19958–1086), and the PB1 domain plus the middle region (MR) of the protein (MR + PB1, ARF19355–1086) (Fig. 3a). We found that the ARF19 PB1 domain does not display thermoresponsive accumulation (Fig. 3b). In contrast, both the ARF19 DBD and the ARF19 MR + PB1 displayed thermoresponsive accumulation (Fig. 3b). Because these two proteins share no residues in common, we conclude that more than one region of the protein bolsters protein stability at elevated temperatures.
a Graphic demonstrating examined ARF19 protein truncations. b Immunoblot analysis with anti-YFP antibody of seedlings carrying the indicated full length (FL) or ARF19 truncation incubated for 4 hours at the 22 °C or 37 °C. Two independent transformants shown for each truncation. Asterisk indicates a non-specific band, which displays decreased accumulation at elevated temperature. c Graphic depicting in vivo mutagenesis screen. d Alignment of ARF7 and ARF19 homologs with sites of variation identified. e Model of ARF19 aligned to ARF5 DBD (4LDU51) with ARF19W330L and ARF19R297W variant effects on contact sites. f Mean ( + SE) ratiometric signal of protoplasts expressing ARF19, ARF19W330L, or ARF19R297W ratiometric reporters (n = 5677 for ARF19 at 22 °C, 5904 for ARF19 at 32 °C, 11513 for ARF19W330L at 22 °C, 7373 for ARF19W330L at 32 °C, 7079 for ARF19R297W at 22 °C, 6649 for ARF19R297W at 32 °C, 7877 for empty vector at 22 °C, and 5197 for empty vector at 32 °C,), letters indicate a significance of p < 0.0001, 2-way ANOVA with Tukey’s HSD test. g Immunoblot analysis with anti-GFP antibody of 5-day-old arf7 arf19 seedlings carrying UBQ10:YFP-ARF19, UBQ10:YFP-ARF19W330L, or UBQ10:YFP-ARF19R297W incubated for 4 hours at 22 °C or 32 °C. h Mean (±SE) hypocotyl lengths of 5-day-old seedlings of wildtype (Col-0), arf7 arf19, and arf7 arf19 carrying UBQ10:YFP-ARF19, UBQ10:YFP-ARF19W330L, or UBQ10:YFP-ARF19R297W grown at 22 °C or 32 °C (n = 58 for wild type at 22 °C, 59 for wild type at 32 °C, 107 for arf7 arf19at 22 °C, 109 for arf7 arf19 at 32 °C, 212 for arf7 arf19 carrying UBQ10:YFP-ARF19 at 22 °C, 107 for arf7 arf19 carrying UBQ10:YFP-ARF19 at 32 °C, 134 for arf7 arf19 carrying UBQ10:YFP-ARF19W330L at 22 °C, 212 for arf7 arf19 carrying UBQ10:YFP-ARF19W330L at 32 °C, 118 for arf7 arf19 carrying UBQ10:YFP-ARF19R297W at 22 °C, and 93 for arf7 arf19 carrying UBQ10:YFP-ARF19R297W at 32 °C). Groups not sharing a letter are significantly different from one another (Two-factor ANOVA on the log transformed data; Tukey’s multiple comparison tests, p < 0.05). i Mean (±SE) increase in hypocotyl length of 5-day-old seedlings of wildtype (Col-0), arf7 arf19, and arf7 arf19 carrying UBQ10:YFP-ARF19, UBQ10:YFP-ARF19W330L, or UBQ10:YFP-ARF19R297W grown at 22 °C or 32 °C, calculated from the data in (h). Groups not sharing a letter are significantly different from one another (One-factor ANOVA on the log-transformed data, Tukey’s multiple comparison tests, p < 0.05).
We next sought to identify other residues that could confer stability using an in vivo mutagenesis screen. We randomly mutagenized the pUBQ10:YFP-ARF19 plasmid using an error prone polymerase25 followed by bulk Arabidopsis transformation with these variants (Fig. 3c). We then screened transformants on coverslip-bottom culture plates and a heated stage, identifying ARF19W330L and ARF19R297W variants that failed to display thermoresponsive accumulation (Figure S4). Both mutations are in conserved residues within the ancillary domain of the DBD (Fig. 3d). These mutations are near a region recently reported to regulate ARF stability16. To confirm that these variants impact ARF stability, we assessed variant thermostability using the ratiometric reporter in protoplasts (Fig. 2a), finding that each variant displays attenuated thermoresponsive stability (Fig. 3f).
In the ARF19 protein model, W330 was buried within the ancillary domain of the DBD and had a three-way polar interaction with R331 and E51 (Fig. 3e). ARF19W330L is likely to break the polar interaction with E51. R297 occurs within a beta-sheet of a five-stranded beta-barrel of the ancillary domain and forms a polar contact with E353 at the end of a predicted alpha-helix (Fig. 3e). In ARF19R297W, the polar contacts from the R-group are likely to be ablated. Further, the substitution of a positively charged arginine residue with a bulky aromatic tryptophan could disrupt intramolecular interactions in this region of the protein. Altogether, because thermoresponsive accumulation is displayed by more than one ARF protein region, we speculate that multiple factors contribute to ARF thermostability.
To assess the functional role of ARF19 temperature-responsive accumulation, we created stable rescue lines by transformation arf7 arf19 with UBQ10:YFP-ARF19, UBQ10:YFP-ARF19W330L, or UBQ10:YFP-ARF19R297W. We found that, similar to what we found in our screen (Figure S8), the ARF19W330L and ARF19R297W variants displayed less thermoresponsive protein accumulation than wild type ARF19 (Fig. 3g). However, these stable lines with these variants also displayed decreased accumulation at 22 °C when compared to wild type protein accumulation, suggesting that these variants affect ARF19 protein stability at both ambient and elevated temperature. Expressing UBQ10:YFP-ARF19 in arf7 arf19 mostly rescued its thermomorphogenesis phenotype (Fig. 3h); this partial rescue is likely due to the fact that this was in the double mutant background, and we rescued with only ARF19. Wild type ARF19, ARF19W330L and ARF19R297W each similarly mostly rescued the long hypocotyl developmental phenotype of arf7 arf19 at 22 °C, suggesting that at this temperature, both variants accumulate sufficient protein to compensate for loss of genomic ARF19 (Fig. 3h). In contrast, neither UBQ10:YFP-ARF19W330L nor UBQ10:YFP-ARF19R297W rescued arf7 arf19 thermomorphogenic growth defects (Fig. 3i), suggesting that temperature-regulated ARF19 accumulation contributes to temperature-responsive growth. After finding that ARF protein thermoresponsive stability activity is encoded in both the DBD and disordered middle region, we sought to understand how ARF proteins fit into known temperature pathways, and whether temperature impacted its nucleo-cytoplasmic partitioning.
Temperature-responsive ARF condensation
Given the established relationship between ARF7 and ARF19 function and condensation11, we examined the effects of varying temperatures on ARF7 and 19 condensation and nuclear localization. In phase-separating systems, temperature frequently regulates solubility12 (Fig. 4a). Because diffuse/soluble ARF7 and 19 are localized to the nucleus whereas ARF condensates are localized to the cytoplasm11,26, we examined whether increasing temperature would result in increased soluble/nuclear ARF protein. This question is particularly important because in phase separating systems, increased overall protein concentrations do not necessarily result in increased ‘active’ protein when the protein in the dilute phase is the active form of protein. In these systems, the concentration of dilute phase protein is buffered against changes in overall protein levels. We found that the concentration of nuclear (i.e., diffuse) ARF protein increases with increasing temperatures (Fig. 4b–d). In time course imaging of hypocotyls exposed to elevated temperature, we observed a rapid increase in nuclear signal (8 min post-exposure) prior to the occurrence of cytoplasmic condensates ( ~ 31 min post-exposure), consistent with a model in which elevated temperature conditions allow for increases in soluble/nuclear ARF protein (Fig. 4e, f). Congruent with this result, Borniego et al. (see companion article) demonstrate that ARF7 and ARF19 display increased solubility at elevated temperature when expressed in yeast.
a Graphic of typical temperature-dependent phase diagram for upper critical saturation temperature (UCST) behavior. C1 and C2 correspond to protein critical saturation concentrations. In this example, the saturation concentration of the protein is higher with higher temperature (ie, the protein is more soluble at elevated temperature in UCST systems). b Confocal microscopy images of 4-day-old root tips of seedlings expressing ARF7:ARF7-mVenus after a 4-hour incubation at 22 °C or 37 °C. To the right are closeup views of nuclear signal. c Mean (±SD) fluorescence intensity of root tip nuclei in roots of seedlings pARF7: ARF7-mVenus11 after 4 h incubation at 22 °C and 37 °C (n = 148 nuclei at 22 °C and 143 nuclei 37 °C). ****P = 3.02E-13 using two-tailed unpaired t-test. Arbitrary units are used. d Mean (±SD) fluorescence intensity of root tip nuclei of of 4-day-old seedlings expressing pUBQ10:YFP-ARF19 after 4 h incubation at 22 °C or 28 °C (n = 24 nuclei at 22 °C and 40 nuclei 37 °C). ****P = 8.62E-10 using two-tailed unpaired t-test. Arbitrary units are used. e Time course fluorescence microscopy images of hypocotyl cells of 4-day-old seedlings expressing pUBQ10:YFP-ARF19. Seedlings were mounted on a heated stage at 22 °C that rapidly transitioned to 32 °C. Images were taken at the indicated time points and, in one example cell, the nucleus pointed out with a puce carat and condensates pointed out with orange carats. f Mean (±SD) fluorescence intensity in nuclei at the start of treatment and after 31 min of treatment. (n = 88 nuclei at 0 min and 84 nuclei at 31 min from 5 individual seedlings). ****P = 2.51E-18 using two-tailed unpaired t-test. Arbitrary units are used. g Graphic demonstrating effectors and reporter in the FrankenARF7 system22. h Box plots of median and interquartile range of mScarlet signal found in mNeon positive cells; whiskers represent maximum and minimum values (n = 1340 for the control at 22 °C, 3554 for the control at 32 °C, 2210 for FrARF7 at 22 °C, and 1736 for FrARF7 at 32 °C). P = 1.05E-06 for the FrARF effector and P = 6.37E-10 for the FrARF reporter using two-tailed unpaired t-test. Box plots of median and interquartile range of mNeon signal in transformed protoplasts; whiskers represent maximum and minimum values.
In a phase-separating system12, environmental conditions such as temperature frequently affect the saturation concentration in the system (Fig. 4a). In these systems, the concentration of the dilute phase equals that of the saturation concentration of the system, such that the dilute phase concentration can change with environmental conditions. For the ARF proteins, the dilute ARF protein resides in the nucleus where it is transcriptionally active11,14,26. Thus, the concentration of nuclear ARF protein likely correlates with the solubility of the protein.
To test whether increased ARF solubility and nuclear concentration affected transcriptional output, we directly examined ARF7 transcriptional output using a protoplast-based reporter of transcriptional activity, in which the ARF7 DBD has been replaced with the GAL4 DBD in a FrankenARF protein fused to mVenus22 (Fig. 4g). In the same plasmid, the GAL4 UAS is upstream of an mScarlet reporter to allow for ratiometric analysis of ARF7 transcriptional output. Examination of ARF7 activity in protoplasts at 22 °C and 32 °C reveals increased transcriptional output at this temperature, consistent with elevated ARF7 levels in the nucleus at this temperature (Fig. 4h).
Our data is consistent with the possibility that ARF7 and 19 display upper critical solution temperature (UCST) behavior (Fig. 4a), often displayed by Prion-like domain (PrLD)-containing proteins27,28. In UCST phase separation systems, molecules such as ARF7 and 19 display increased solubility at elevated temperature and thus the dilute concentration of the molecule increases with increasing temperature (Fig. 4a). Enthalpy-driven UCST behavior can be mediated by aromatic sidechain associations. Analysis of the ARF7 and ARF19 intrinsically disordered regions reveals residues compatible with UCST behavior (Figures S5, S6). Indeed, Borniego et al. (accompanying manuscript) observe temperature-regulated ARF7 and ARF19 solubility in root tissues and in yeast. In sum, our data are consistent with a model in which temperature regulates the levels of nuclear ARF7 and ARF19 protein not only by altering protein stability but also by increasing protein solubility.
ARF7 and ARF19 thermoresponsive accumulation is not fully dependent on known thermosensing pathways
PHYTOCHROME INTERACTING FACTOR4 (PIF4) is a central signaling hub of thermomorphogenesis and drives the thermoresponsive expression of auxin biosynthesis genes YUCCA8 (YUC8) and YUC9. ELONGATED HYPOCOTYL 5 (HY5) acts antagonistically to PIF4 by competing for YUC8 binding sites29,30,31,32. The thermosensor EARLY FLOWERING3 (ELF3) also negatively regulates PIF433,34,35,36. The light receptor phyB is a thermosensor that interacts with PIF436,37. Together, these pathways integrate temperature cues to plant physiological adaptations to temperature5. Ultimately, these pathways result in elevated auxin biosynthesis through the YUCCA family (Fig. 5a). To determine whether temperature-regulated auxin biosynthesis was required for our observed ARF7 thermoregulated protein accumulation, we examined ARF7 protein levels in the yucQ and 35S:YUC backgrounds. We found that ARF7 temperature-regulated accumulation was similar to wild type (Figure S7a, b).
a Graphic depicting known temperature response pathways. b Immunoblot analysis of wild type (Col-0) or hy5 seedlings after 4-hour incubation at 22 °C, 32 °C, or 37 °C. c Immunoblot analysis of wild type (Col-0) or phyBpifQ seedlings after 4-hour incubation at 22 °C, 32 °C, or 37 °C. d Immunoblot analysis of wild type (Col-0), elf3 elf4, or seedlings overexpressing ELF3 after 4 hour incubation at 22 °C, 32 °C, or 37 °C. e Immunoblot analysis of wild type (Col-0) or phyB-9 seedlings after 4 h incubation at 22 °C 32 °C. f Immunoblot analysis of wild type (Col-0) or pifq seedlings after 4-hour incubation at 22 °C or 32 °C. Each immunoblot experiment was performed at least three times; additional replicates are in Supplemental Fig. 7b.
To ascertain if ARF thermostability was regulated by PIF4 and HY5, we measured ARF7 protein levels in the hy5 and phyB pifQ mutants, finding that ARF7 thermoresponsive accumulation was not different from wildtype in these backgrounds (Fig. 5b, c). We further assayed ARF7 protein levels in the elf3 elf4 mutant background and in a line overexpressing ELF3 (35S:ELF3). ARF7 thermoresponsive accumulation was unaffected in the elf3 elf4 mutant (Fig. 5d). Conversely, ELF3 overexpression resulted in slightly elevated ARF at all temperatures (Fig. 5d), suggesting a more complicated relationship between ARF7 stability and ELF3, which plays roles not only in temperature sensing36, but also in circadian clock regulation38. To further investigate the effects of PhyB and PIFs on ARF thermoresponsive accumulation, we assessed temperature-regulated ARF7 accumulation in phyB and pifQ, finding that in these mutants, ARF7 thermoresponsive protein accumulation is similar to wild type in these mutant backgrounds (Fig. 5e, f).
Elevated temperature results in elevated auxin levels through the activities of known temperature sensing pathways5.To separate the effects of increased auxin from ARF7/ARF19 activity, we examined temperature-responsive hypocotyl elongation of wild type, the auxin overproducing line 35S:YUC139, and arf7 arf19 with and without the auxin inhibitors L-Kynurenine40 and Yucasin41. Without inhibitors, both auxin overproduction and loss of ARF7 and 19 activity dampened thermomorphogenic response at 28 °C and 32 °C (Figure S8b, c). In 35S:YUC1, auxin levels were likely saturated, limiting further temperature-inducted elongation. Inclusion of auxin biosynthesis inhibitors did not fully block thermomorphogenic hypocotyl elongation in either wild type or in the 35S:YUC1 line (Figure S8b, c). Conversely, inclusion of auxin biosynthesis inhibitors diminished arf7 arf19 hypocotyl thermomorphogenesis (Figure S8b, c). The auxin reporter, DII-Venus confirmed that the auxin inhibitors blocked auxin biosynthesis (Figure S8d, e). The dynamic nature of temperature-regulated Class-A ARF accumulation (Fig. 1), condensation (Fig. 4) and roles in thermomorphogensis (see Borniego et al., companion article) suggests roles for ARF proteins in temperature sensing that are not directly reliant on their roles interpreting auxin levels.
Natural variation in temperature-regulated ARF stability associated with changes in thermomorphogenesis
We were curious if ARF stability was important for thermormorphogenesis in other accessions. Therefore, we leveraged naturally-occurring genetic variation within Arabidopsis thaliana to see if different levels of ARF stability had consequences on hypocotyl thermomorphogenesis. We chose 15 accessions that have evolved in diverse environments (Fig. 6a) and compared hypocotyl elongation at 22 °C and 28 °C. We found a spectrum of hypocotyl elongation in response to the increased temperature (Fig. 6b). We then assayed thermoresponsive ARF7/19 protein accumulation within these accessions using immunoblotting. Some accessions, such as Ler-1, do not display a sharp accumulation in ARF7/19 at elevated temperature whereas other accessions, such as MNF-Che-47, display a robust increase in ARF7/19 accumulation at elevated temperature (Fig. 6c). We then analyzed the relationship between thermomophogenic hypocotyl elongation and thermoresponsive ARF protein accumulation using linear regression and found a correlation between thermoresponsive ARF7/19 protein accumulation and thermormophogenic hypocotyl elongation (Fig. 6d). The strength of this correlation was unexpected, considering the multiple pathways regulating thermomorphogenesis and the broad differences amongst accessions; however, this data suggests that temperature-regulated stability of the ARF7/19 transcription factors, which mediate auxin transcriptional output, contributes to the thermomorphogenic response across Arabidopsis accessions.
a Map of locations where accessions originated. b Representative images of 6-day-old seedlings grown at 22 °C or 28 °C. c Immunoblot analysis of seedlings incubated for four hours at 22 °C or 37 °C. Satellite Data accessed from Wolfram Version 14.3 using GeoListPlot. d Mean (± SEM) increase in hypocotyl lengths of 6-day-old seedlings grown at 28 °C versus 22 °C (n = 16 for Wa-1, 19 for Ha-0, 24 for Qui-0, 21 for Noveg-3, 28 for Ca-0, 24 for MNF-Che-47, 18 for Sq-a, 18 for Ler-1, 23 for Map-35, 19 for Sf-2, 27 for Edi-1, 28 for Xan-1, 27 for Hovdala-2, 23 for Sq-1, 24 for Col-0, and 22 for Ciste-2). Color of bar corresponds to fold change measured in immunoblot analysis. e Linear regression comparing ARF7 protein level increase (X-axis) to increase in hypocotyl elongation (Y-axis) at 28 °C relative to 22 °C. Slope determined to be non-zero with a p-value of 0.0025.
Discussion
Plants rely on the auxin pathway to tailor their physiology to their surroundings. Temperature is a prominent environmental cue plants must register to tune growth, adapt to seasonal rhythms, and to respond to acute stress conditions. Faced with a warming climate, it is paramount to understand ways plant response to elevated and rapidly changing temperatures. Here, we find that the accumulation of Class-A ARFs, which drive many aspects of plant growth and development, is regulated by temperature. Further, although auxin biosynthesis has long been found to be a key component of thermomorphogenesis, our data suggests ARF7 and ARF19 contribute to thermomorphogenesis independent of their role interpreting auxin levels. In particular, thermoresponsive ARF stability and condensation are rapidly altered by changing temperature.
Two well-studied Class-A ARFs include ARF7 and ARF19. These key transcription factors are required for multiple aspects of plant development10. In this work, we determine that ARF7/19 thermoresponsive accumulation is seemingly independent of multiple known temperature response pathways, raising the possibility that either there are yet-to-be discovered termperature sensing pathways or that ARFs directly sense temperature changes. Further, for this increase in ARF protein accumulation to drive a thermal response, this increased ARF must coincide with increased levels of soluble-nuclear protein and transcriptional output. ARF7 and ARF19 undergo phase separation, and ARF cytoplasmic condensates attenuate auxin transcriptional responses11. Our collective data (this work and that of Borniego et al.) point to a model in which ARF7 and ARF19 solubility are temperature-controlled, providing a rapid and reversible system to contributing to the integration of temperature cues into plant biology.
Climate change not only impacts average global temperatures, but also rapid and detrimental shifts in temperature. In these assays, ARF7 and ARF19 dramatically increased at 37 °C, a temperature more representative of acute heat stress in Arabidopsis. Whereas the phenotypes observed in this study focused on seedlings grown at 28 °C, the dramatic protein changes at 37 °C could hint at a possible role for these ARFs in heat stress. Indeed, ARF proteins play a notable role in abiotic stress in plants42. Understanding the behavior of ARF7 and ARF19 protein at elevated and rapidly changing temperatures advances our understanding of how plants respond to elevated temperatures.
Our data and that of Borniego et al. (accompanying manuscript) point to another tunable mechanism contributing to the integration of temperature cues into plant biology. This thermoregulation of ARF stability and condensation occurs rapidly and reversibly, on the time scale of minutes. This rapid shift in ARF7 and ARF19 adds another layer in which plants to respond to temperature shifts and primes cellular competence to respond to temperature-responsive auxin synthesis. In the future, the ARF system could be co-opted to directly and rapidly regulate transcriptional outputs to tune plant growth to varying temperature conditions. How temperature might interface with the recently discovered use of movement-generated force to regulate ARF condensation14 remains to be seen. As our planet warms, understanding how ectotherms integrate cellular temperature into tuned growth output will be critical to predict how these organisms will tolerate novel growth conditions.
Methods
Plant Growth Conditions
Thermomorphogenesis assays: Seedlings were stratified 4 °C overnight and plated on Plant Nutrient Media43 solidified with 0.6% agar and supplemented with 0.5% (w/v) sucrose. To examine the effects of auxin inhibitors on hypocotyl elongation, seedlings were grown for one day under yellow-filtered light prior to grown under dark conditions at 22 °C to encourage hypocotyl elongation. Seedlings were then transferred to media containing a mock treatment (DMSO), or 50 µM Yucasin (gift from Jerry Cohen) and 10 µM L-Kynurenine (Millipore Sigma). Yucasin and L- Kynurenine (L-kyn) are auxin biosynthesis inhibitors that respectively block the enzymatic activities of YUCCA and TAA40,41. Seedlings on mock or auxin inhibitor treatment were incubated at 22 °C, 28 °C, or 32 °C vertically under continuous yellow-filtered light for an additional 3 d prior to imaging and measurement of hypocotyl lengths using FIJI.
For protoplast assays, 7-day-old seedlings were transferred to pots containing Metro-Mix 830 media (Sungro) and placed at 22 °C at 16 h light for an additional 7–10 days. For the growth of T1 agrobacterium transformants, seeds were placed directly in Metro-Mix 830 at 16 h light for 7 days. After 7 days, positive transformants were selected by spraying the seedlings with BASTA. After selection, plants were grown at 16 h light until mature. Transformants were harvested as individuals and their progeny were screened.
Vector Construction
pUBQ10:mNEON-ARF19c-P2A-mScarlet3 was generated using the pUBQ-mNEON-P2A-mScarlet3 backbone. pUBQ10:mNeon-P2A-mScarlet (using 5′-AACGCATGCGGAAGCGGAGC-3′ and 5′-AACACTCCCACTTCCAGAGCC-3′) and ARF19 CDS (using 5′-GGAAGTGGGAGTGTTATGAAAGCTCCATCAAATGG-3′ and 5′-GCTTCCGCATGCGTTTCTGTTGAAAGAAGCTGCAGCAG-3′) were linearized via PCR (Invitrogen, SuperFi II) and the ARF19 CDS was integrated into pUBQ10:mNeon-P2A mScarlet backbone using NEBuilder HiFi DNA Assembly (New England Bioscience). ARF19 variant fragments were generated by twist Bioscience in the pENTR backbone and inserted into the pUBQ10:mNeon-P2A-mScarlet3 backbone using NEBuilder HiFi DNA assembly.
FrARF7 and GAL4 DBD plasmids were generated as previously described22: A gene fragment which encoded the FrARF7 protein, a NOS terminator, and 1000 bp of non-coding DNA including a multiple cloning site was synthesized by Twist in the pENTR Gateway compatible backbone. An additional gene fragment encoding 500 bp of non-coding DNA followed by 5X Gal4 UAS sites, a minimal CaMV 35S promoter, and mScarlett-I fused to Histone 2B was synthesized and put into the pENTR backbone by Twist. The pENTR FrARF7 backbone was linearized using SacI, and the reporter was PCR amplified to include 20 bp overlap with the pENTR FrARF7 multiple cloning site at the 5´ and 3´ ends. The reporter insert was cloned into the linearized FrARF7 vector using NEBuilder HiFi cloning to generate pENTR FrARF7+Reporter clones. The FrARF7+Reporter Entry clone was then cloned into pLCS107, which provided an in-frame mNEON fused to the N-terminus of FrARF7 driven under the UBQ10 promoter and a nos terminator for the mScarlet-I H2B reporter, by gateway cloning. The Gal4 DBD variant was generated by inframe deletions of the ARF7 CDS by PCR linearization and self-assembly with NEBuilder HiFi cloning.
Immunoblot Analysis
Custom polyclonal antibody was generated by injecting rabbits with purified ARF7 PB1 domain21 using a standard 70-day immunization protocol by the company Biomatik.
Fresh tissue from 30 4-day-old seedlings was homogenized using a 2010 Geno/Grinder (SPEX Sample Prep) at 1500 RPM for 30 seconds and then placed at 70°C in 2x NuPage LDS buffer (141 mM Tris, 2% Lithium Dodecyl Sulfide, 0.51 mM EDTA, 10% glycerol, 0.175 mM Phenol Red, 0.22 mM Coomassie Blue) and loaded onto a Bolt 8% Bis-tris protein gel (Thermo Scientific). The gel was run at 80 V for 15 min and then 160 volts for 1 h until proper separation of bands was obtained. The gel was transferred to a nitrocellulose membrane (Amersham Protan 0.45 µm NC) using a wet-transfer system. Loading control was determined with Ponceau solution (0.1% Ponceau w/v, 5% acetic acid) for 2 min, washed in TBS-T solution (200 mM Tris, 1.5 M NaCl, 0.1% Tween-20) until excess Ponceau solution was removed, and imaged for loading control analysis, or with 1:5000 anti-Actin Antibody (Agrisera, AS13 2640). The membrane was blocked in 8% milk (Great Value) in TBS-T buffer for 1 hour. 1:5000 Anti-GFP (Agrisera, AS20 4443) or 1:5000 Anti-ARF7 antibody was added overnight at 4 °C. The membrane was then washed 3 times in TBS-T solution for 5 min before being incubated with anti-rabbit secondary antibody at 4 °C overnight. The signal was detected using a WesternBright ECL HRP substrate kit (Advasta) according to manufacturer’s instructions.
For temperature-dependent assays, 4-day old Arabidopsis tissue was placed in water pre-warmed to the treatment temperature for 4 hours (unless otherwise stated). After 4 hours, seedlings were dabbed dry on a paper towel and flash frozen in liquid nitrogen and used in immunoblot analysis.
To examine the effects of the 26S proteasome on ARF stability, ARF levels in seedlings treated with the proteasomal inhibitor MG132 were examined. MG132 (Sigma-Aldrich) was dissolved in DMSO and supplemented to pre-warmed water to a final concentration of 50 µM. For mock treatments, an equivalent amount of DMSO was supplemented to water. 30 4-day-old seedlings were then added to either treatment and incubated at the described temperature for four hours. After incubation, tissue was dabbed dry on a paper towel and flash frozen in liquid nitrogen for immunoblot analysis.
To examine the effects of the autophagy on ARF stability, ARF levels in seedlings treated with the autophagy inhibitor ConcanamycinA were examined. ConcanamycinA was dissolved in DMSO. ConcanamycinA, Concavlin A, or the equivalent volume of DMSO, was then added to pre-warmed water to a final concentration of 1 µM. Plants were left to incubate at the indicated temperature for 16 h. After incubation, plants were dabbed dry on a paper towel and flash frozen in liquid nitrogen for immunoblot analysis.
To determine the half-lives of ARF proteins at different temperatures, seedlines were pre-treated at their prescribed temperature for one hour in a 6-well plate. Then, cycloheximide, dissolved in water, was added to each well and lightly mixed to ensure even diffusion of the chemical within the well. Plants were then collected from each well at the described time (immediately, 1 h, 2 h, or 4 h later). Tissue was dabbed dry on a paper towel and flash frozen in liquid nitrogen for immunoblot analysis.
Protein Screen
An ARF19 mutagenesis library was created using a protocol adapted from Manel’s 2-plasmid Mutagenesis protocol25. The pUBQ10:YFP-ARF19c binary vector11 was transformed into competent JS200 cells carrying the EP pol I plasmid (Addgene). Overnight cultures were grown statically overnight at 30 °C. The following morning, cultures were shaken for 2 hours at 180 rpm and 2 µl of each culture was used to inoculate 5 ml of 2xYT media with antibiotics. Mutagenesis was stimulated by placing cultures statically at 37 °C for 15 min and then shaken overnight at 37 °C at 180 rpm. Cultures were miniprepped and transformed into JS200 cells and plated at 30 °C in a serial dilution to approximate the number of transformation events present in each culture. All colonies on the plate were pooled together in LB. Cultures were then used to inoculate 5 mL of 2XYT media and a second mutagenesis step was performed. The rate of mutagenesis was determined by transforming plasmids from the error-prone polymerase into NEB5a cells and sequencing. Mutagenized plasmids were then transformed into agrobacterium (GV3101). Agrobacterium colonies were then pooled and used to transform Col-0 plants using the floral dip method44. Positive transformants were selected using BASTA. Transformants were grown as individuals and their progeny were plated on a glass-bottom 24-well plate supplemented with PNS media. After 5 days of growth, seedlings were imaged when their roots grew along the bottom of the glass-bottom plate using a Leica DMI8 Thunder light microscope using an OKO Lab temperature-controlled stage at 32 °C. Seedlings were imaged before and after heat treatment. Mutations in YFP-ARF19 were determined by performing a PCR reaction using primers present only in the binary vector backbone (5′-CAACAATTACCAACAACAAC-3′ and 5′-ACCTAGGATCATCAACC-3′) to prevent amplification of the endogenous ARF19 gene and sequenced.
Protoplast generation and experiments
Arabidopsis mesophyll protoplasts were isolated from 12 to 14-day-old Col-0 leaves using the tape sandwich method45. Approximately 100,000 cells were transformed with 20 ug of plasmid containing pUBQ10:mNEON-ARF19-P2A-mScarlet3, or variants in which ARF19 was replaced by an ARF19 variant and incubated for 16 hours in the dark. Following incubation, protoplasts were either left at ambient temperature or treated at 32°C for four hours. The Beckman Coulter Cytoflex S Flow Cytometer was used to score the transformed protoplasts. Intact protoplasts were identified with a back gating strategy. mNEON levels were collected using the B525 channel (Ex: 488 nm Em: 525 ± 40 nm, 69 gain). mScarlet levels were collected using the Y610 channel (Ex: 561 nm Em: 610 ± 20 nm, 1000 gain). FCS files were generated using CytExpertsoftware and analyzed with FlowKit46, using NumPy47 and Pandas48 packages in Python as previously described. Stability was calculated by dividing the mNEON signal by the mScarlet signal for each cell. Graphs were generated with seaborn49 and Matplotlib50 in Python.
Microscopy
All images were acquired using a Leica DMi8 Fluorescence microscope. Nuclei of seedlings carrying the pARF7:mVenus-ARF7 or pUBQ10:YFP-ARF19 reporter were imaged using a Leica DMi8 Confocal microscope. Nuclei were measured in 4-day-old seedlings in young epidermal cells with or without a four-hour treatment at 37 °C. Nuclei fluorescence intensity was quantified using ImageJ Software.
For time course images depicting pUBQ10:YFP-ARF19 thermal accumulation, 4-day-old seedlings were mounted in water and imaged on a Leica DMI8 Thunder light microscope using an OKO Lab temperature-controlled stage at 32 °C and imaged approximately every 12 min over the course of 4 hours.
Natural Variation Assay
Fifteen Arabidopsis accessions were selected for their geographic diversity. To determine thermomorphogenic hypocotyl elongation, seedlings were grown with continuous illumination under yellow filters for 6 days at either 22 °C or 28 °C and hypocotyls were measured using FIJI. To conduct western blot analysis, 30 seedlings of each genotype were placed in pre-warmed water at 22 °C or 37 °C and incubated for four hours prior to protein extraction. Relative ARF7 protein levels were measured using FIJI by quantifying signal within the 37 °C band and dividing by the 22 °C band.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are either reported in this manuscript or available upon request from the corresponding author. Source data are provided with this paper.
References
Delker, C., Quint, M. & Wigge, P. A. Recent advances in understanding thermomorphogenesis signaling. Curr. Opin. Plant Biol. 68, 102231 (2022).
Ahmad, M., Imtiaz, M., Shoib Nawaz, M., Mubeen, F. & Imran, A. What Did We Learn From Current Progress in Heat Stress Tolerance in Plants? Can Microbes Be a Solution?. Front Plant Sci. 13, 794782 (2022).
Kerbler, S. M. & Wigge, P. A. Temperature Sensing in Plants. Annu Rev. Plant Biol. 74, 341–366 (2023).
Quint, M. et al. Molecular and genetic control of plant thermomorphogenesis. Nat. Plants. 2, 15190 (2016).
Casal, J. J. & Balasubramanian, S. Thermomorphogenesis. Annu Rev. Plant Biol. 70, 321–346 (2019).
Gray, W. M., Östin, Sandberg, A., Romano, G. & Estelle, C. P. M. High temperature promotes auxin-mediated hypocotyl elongation in Arabidopsis. Proc. Natl. Acad. Sci. USA. 95, 7197–7202 (1998).
Wang, R. et al. HSP90 regulates temperature-dependent seedling growth in Arabidopsis by stabilizing the auxin co-receptor F-box protein TIR1. Nat. Commun. 7, 10269 (2016).
Sun, J., Qi, L., Li, Y., Chu, J. & Li, C. PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating Arabidopsis hypocotyl growth. PLoS Genet 8, e1002594 (2012).
PM, M. & S, C. High Temperature Stimulates DWARF4 (DWF4) Expression to Increase Hypocotyl Elongation in Arabidopsis. J. Plant Biol. 54, 425–429 (2011).
Powers, S. K. & Strader, L. C. Regulation of auxin transcriptional responses. Dev. Dyn. 249, 483–495 (2020).
Powers, S. K. et al. Nucleo-cytoplasmic Partitioning of ARF Proteins Controls Auxin Responses in Arabidopsis thaliana. Mol. Cell 76, 177–190.e175 (2019).
Emenecker, R. J., Holehouse, A. S. & Strader, L. C. Biological Phase Separation and Biomolecular Condensates in Plants. Annu Rev. Plant Biol. 72, 17–46 (2021).
Xuan, L. et al. MCTP controls nucleocytoplasmic partitioning of AUXIN RESPONSE FACTORs during lateral root development. Dev. Cell 59, 3229–3244 e3225 (2024).
Chauhan, G. et al. Active transport enables protein condensation in cells. Sci. Adv. 11, e70142 (2025).
Jing, H. et al. Regulation of AUXIN RESPONSE FACTOR condensation and nucleo-cytoplasmic partitioning. Nat. Commun. 13, 4015 (2022).
Prigge, M. J. et al. Comparative mutant analyses reveal a novel mechanism of ARF regulation in land plants. Nat. Plants 11, 821–835 (2025).
de Roij, M., Hernandez Garcia, J., Das, S., Borst, J. W. & Weijers, D. ARF degradation defines a deeply conserved step in auxin response. Nat. Plants 11, 717–724 (2025).
Jing, H. & Strader, L. C. AUXIN RESPONSE FACTOR protein accumulation and function. Bioessays 45, e2300018 (2023).
Mutte, S. K. et al. Origin and evolution of the nuclear auxin response system. Elife. 7, e33399 (2018).
Emenecker, R. J., Holehouse, A. S. & Strader, L. C. Sequence determinants of in cell condensate morphology, dynamics, and oligomerization as measured by number and brightness analysis. Cell Commun. Signal 19, 65 (2021).
Korasick, D. A. et al. Molecular basis for AUXIN RESPONSE FACTOR protein interaction and thesssression. Proc. Natl. Acad. Sci. USA 111, 5427–5432 (2014).
Morffy, N. et al. Plant activation domain identification. Nature 632, 166–173 (2024).
Ebstrup, E. et al. NBR1-mediated selective autophagy of ARF7 modulates root branching. EMBO Rep. https://doi.org/10.1038/s44319-024-00142-5 (2024).
Jing, H. et al. The E3 ubiquitin ligase AFF1 targets ARF19 to the proteasome. Nat. Commun. 13, 4015 (2022).
Camps, M., Naukkarinen, J., Johnson, B. P. & Loeb, L. A. Targeted gene evolution in Escherichia coli using a highly error-prone DNA polymerase I. Proc. Natl. Acad. Sci. USA 100, 9727–9732 (2003).
Kuan, C., Strader, L. C. & Morffy, N. ARF19 Condensation in the Arabidopsis Stomatal Lineage. Micro. Publ. Biol. 2023, https://doi.org/10.17912/micropub.biology.000708 (2023).
Holehouse, A. S., Ginell, G. M., Griffith, D. & Boke, E. Clustering of Aromatic Residues in Prion-like Domains Can Tune the Formation, State, and Organization of Biomolecular Condensates. Biochemistry 60, 3566–3581 (2021).
Pappu, R. V., Cohen, S. R., Dar, F., Farag, M. & Kar, M. Phase Transitions of Associative Biomacromolecules. Chem. Rev. 123, 8945–8987 (2023).
Farag, M. et al. Phase separation of protein mixtures is driven by the interplay of homotypic and heterotypic interactions. Nat. Commun. 14, 5527 (2023).
Delker, C. et al. The DET1-COP1-HY5 pathway constitutes a multipurpose signaling module regulating plant photomorphogenesis and thermomorphogenesis. Cell Rep. 9, 1983–1989 (2014).
Toledo-Ortiz, G. et al. The HY5-PIF regulatory module coordinates light and temperature control of photosynthetic gene transcription. PLoS Genet 10, e1004416 (2014).
Gangappa, S. N. & Kumar, S. V. DET1 and HY5 Control PIF4-Mediated Thermosensory Elongation Growth through Distinct Mechanisms. Cell Rep. 18, 344–351 (2017).
Nieto, C., Lopez-Salmeron, V., Daviere, J. M. & Prat, S. ELF3-PIF4 interaction regulates plant growth independently of the Evening Complex. Curr. Biol. 25, 187–193 (2015).
Box, M. S. et al. ELF3 controls thermoresponsive growth in Arabidopsis. Curr. Biol. 25, 194–199 (2015).
Raschke, A. et al. Natural variants of ELF3 affect thermomorphogenesis by transcriptionally modulating PIF4-dependent auxin response genes. BMC Plant Biol. 15, 197 (2015).
Jung, J. H. et al. A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis. Nature 585, 256–260 (2020).
Legris, M. et al. Phytochrome B integrates light and temperature signals in Arabidopsis. Science 354, 897–900 (2016).
Covington, M. F. et al. ELF3 modulates resetting of the circadian clock in Arabidopsis. Plant Cell 13, 1305–1315 (2001).
Zhao, Y. et al. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291, 306–309 (2001).
He, W. et al. A small-molecule screen identifies L-kynurenine as a competitive inhibitor of TAA1/TAR activity in ethylene-directed auxin biosynthesis and root growth in Arabidopsis. Plant Cell 23, 3944–3960 (2011).
Nishimura, T. et al. Yucasin is a potent inhibitor of YUCCA, a key enzyme in auxin biosynthesis. Plant J. 77, 352–366 (2014).
Jing, H., Wilkinson, E. G., Sageman-Furnas, K. & Strader, L. C. Auxin and abiotic stress responses. J. Exp. Bot. 74, 7000–7014 (2023).
Haughn, G. W. & Somerville, C. Sulfonylurea-resistant mutants of Arabidopsis thaliana. Mol. Gen. Genet. 204, 430–434 (1986).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Wu, F. H. et al. Tape-Arabidopsis Sandwich - a simpler Arabidopsis protoplast isolation method. Plant Methods 5, 16 (2009).
White, S. et al. FlowKit: A Python Toolkit for Integrated Manual and Automated Cytometry Analysis Workflows. Front Immunol. 12, 768541 (2021).
Harris et al. Array programming with NumPy. Nature 585, 357–362 (2020).
pandas-dev/pandas: Pandas (v2.0.2) (Zenodo, 2023).
seaborn: statistical data visualization. (Journal of Open Source Software, 2021).
Matplotlib: A 2D Graphics Environment (Computing in Science & Engineering, 2007).
Boer, D. R. et al. Structural Basis for DNA Binding Specificity by the Auxin-Dependent ARF Transcription Factors. Cell 156, 577–589 (2014).
Acknowledgements
We thank Andrew Willoughby and Nicholas Morffy for critical comments on the manuscript, members of the Casal lab for helpful discussion, Nicholas Morffy with technical assistance in developing the protoplast-based assays, Jerry Cohen for providing the Yucasin, Anna Stepanova for the YUC1-OX seed, Dmitri Nusinow for elf3 elf4 seed, Sheng-Yang He for the 35S: ELF3 seed, and Roosa Laitinen for the natural accession seed stocks. This work was supported by the National Science Foundation (PGRP BIO-2112056 to LCS), the National Institutes of Health (R35 GM136338 to LCS), and Duke Beyond the Horizons funds (to LCS).
Author information
Authors and Affiliations
Contributions
E.G.W., K.S.F., and L.C.S. designed the study. E.G.W. performed microscopy and chemical assays. E.G.W. and K.S.F. performed immunoblot analysis. K.S.F. transfected and analyzed protoplast experiments. E.G.W., K.S.F., M.B.B., M.E.P., and L.C.S. executed seedling phenotypic analysis. E.G.W., K.S.F., and L.C.S. wrote the manuscript. L.C.S. and J.J.C. supervised the project.
Corresponding author
Ethics declarations
Competing interests
L.C.S. is on the scientific advisory board of Prose Foods. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiaofeng Fang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wilkinson, E.G., Sageman-Furnas, K., Pereyra, M.E. et al. AUXIN RESPONSE FACTOR thermostability. Nat Commun 17, 2883 (2026). https://doi.org/10.1038/s41467-026-71012-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71012-y