Abstract
Phenotypic plasticity plays a key role in adaptation to fluctuating environments. However, its evolutionary significance remains debated, with conflicting views on whether it is actively maintained by natural selection or a neutral consequence of molecular constraints. In this study, we examine the evolutionary role of gene expression plasticity in yeast populations exposed to both common, ancestrally experienced stressors (NaCl and heat at 37 °C) and a novel stressor (the antifungal drug clotrimazole). Using mutation accumulation lines to separate the effects of natural selection from genetic drift, we compare gene expression responses (referred to as plastic change) between 22 natural strains, 15 mutation accumulation lines, and their progenitor under non-stressed and stressed conditions. Our results show that, in the antifungal environment, gene expression plasticity is selectively maintained, as indicated by its reduction in magnititude, reversal in directionality, and loss of stress-responsive pathways in mutation accumulation lines. In contrast, plasticity under the common stress conditions appears largely neutral, with random variation across mutation accumulation lines. This study provides direct evidence of the adaptive role of gene expression plasticity in response to novel stressors and offers insights into the molecular mechanisms driving these adaptive responses.
Similar content being viewed by others
Introduction
Phenotypic plasticity, the ability of a genotype to produce different phenotypes in response to environmental changes, is a fundamental feature of biological systems1. While it is widely recognized as an important biological property, its evolutionary origins, maintenance, and role in adaptation remain controversial. A central debate in evolutionary biology concerns whether plasticity is an adaptive mechanism actively maintained by natural selection or a byproduct of molecular constraints and regulatory networks2,3,4,5,6,7. Specifically, the extended evolutionary synthesis challenges the traditional modern synthesis by arguing that plasticity plays a foundational role in adaptation and that its evolutionary importance has been underappreciated8,9,10,11,12. According to the “plasticity-first” hypothesis12, when an environment shifts, the reaction norm of an organism allows it to produce a novel phenotype that may be partially or fully adapted to the new conditions. In other words, plastic phenotypic changes are often necessary for survival in novel environments, providing a crucial buffer that allows populations to persist while genetic adaptation occurs13. For example, plasticity facilitates adaptation by pre-aligning phenotypic traits with their optimal states, thereby serving as a stepping stone for genetic adaptation14. However, an alternative view posits that plastic changes are often reversed rather than reinforced during evolution, suggesting that they may not always be beneficial or necessary for adaptation15.
Gene expression plasticity, a key aspect of phenotypic plasticity, is particularly relevant in microbial populations, where rapid transcriptional responses to environmental stress can directly impact survival and fitness16. By using gene expression levels as focal traits, studying plasticity at the transcriptional level provides an effective and unbiased framework for assessing its evolutionary significance across a group of traits15,17. However, it remains uncertain whether gene expression plasticity is selectively maintained as an adaptive feature or simply represents a transient, nonadaptive response to environmental fluctuations. Moreover, the evolutionary outcomes of plasticity may vary depending on the type of environment encountered, particularly between conditions that have been common throughout an organism’s evolutionary history and those that are novel or rarely experienced. When a stressor has been repeatedly encountered over evolutionary time, such as high salinity (NaCl exposure) or heat stress, organisms are likely to evolve specific adaptive responses to mitigate its effects. Notwithstanding, if such stressors persist over long timescales, plastic responses may eventually become genetically assimilated, resulting in the fixation of adaptive traits and a consequent loss of plasticity5,6,18. In contrast, exposure to novel stressors that were absent from an organism’s evolutionary history poses different challenges. In these cases, organisms may lack specialized adaptive responses but might still cope with stress through mechanisms that overlap with responses to familiar stressors or through broad, general stress–response pathways19.
To address the role of natural selection in the evolution of plasticity under both common and novel stressors, experimental evolution provides a powerful approach by directly assessing the role of selection in shaping gene expression plasticity. Mutation accumulation (MA) experiments, in which populations undergo repeated single-cell bottlenecks to minimize the effects of natural selection, provide a rigorous approach to determine whether plasticity is selectively maintained or subject to genetic drift. If gene expression plasticity is adaptive, selection in natural populations would typically remove mutations that reduce plasticity, preserving an optimal level of gene expression responsiveness. However, in MA lines, where random mutations accumulate primarily due to genetic drift and selection is largely absent, deleterious mutations affecting plasticity would not be purged. As a result, plasticity should systematically decline in MA lines due to the accumulation of mutations that impair its regulatory mechanisms; In contrast, if gene expression plasticity is neutral, meaning it does not provide a selective advantage or disadvantage, its presence or absence in a given strain would be governed by stochastic processes rather than selective pressures. In this case, MA lines would show random deviations from the progenitor in plasticity, with some lines exhibiting higher or lower plasticity compared to the progenitor strain, but without any consistent directional trend. Thus, comparing the magnitude and direction of gene expression plasticity between the progenitor strain and MA lines provides a way to infer whether plasticity is actively maintained by selection.
In this study, we investigate the adaptive significance of gene expression plasticity using yeast MA lines exposed to common and novel stressors. We compare gene expression responses in natural strains, where natural selection remains intact, with MA lines, where selection is largely absent. Our results reveal that gene expression plasticity in the antifungal environment is predominantly adaptive, as indicated by its reduced magnitude in MA lines and the loss of stress-responsive pathways. Conversely, expression plasticity in the common stress conditions (NaCl and heat) appears largely neutral, with no clear pattern of selective maintenance. These findings provide empirical evidence supporting the adaptive role of gene expression plasticity in response to clotrimazole, offering insights into how microbial populations rapidly respond to environmental stressors and how selection shapes these responses over evolutionary timescales.
Results
Study design and transcriptome sequencing of yeast MA lines and natural strains
To investigate the selective pressures acting on expression plasticity, we utilized yeast MA lines generated in a previous study20. These lines were established using BY4741 as the ancestral strain and underwent approximately 1500 generations of MA in the solid nutrient-rich (YPD) medium at 30 °C, which lacked the stressors employed in the current study. To accelerate MA, the mismatch repair gene MSH2 was deleted before MA and reinserted afterward. Consequently, each MA line accumulated an average of ~900 mutations.
To ensure comparability between the common and novel stress conditions, we selected MA lines that were capable of stable growth in both the novel stress and at least one of the common stress environments. Initially, 21 MA lines were chosen. We then conducted whole-genome sequencing to verify the identity of the MA lines. This analysis revealed that 6 lines were contaminated by the progenitor strain, as they carried ≤3 mutations, and were therefore excluded from the study (Supplementary Data 1 and “Methods” for details). As a result, a total of 15 MA lines, their progenitor BY4741, and 22 natural strains21 were collected for analysis (Fig. 1a and Supplementary Data 2). Given the relatively short evolutionary timeframe of the MA experiment, the genetic diversity among the MA lines averaged 1.8 × 10−5, significantly lower than the 3.0 × 10−3 observed in the natural strains (Fig. 1b, c). To test if the plastic response differs between common and novel stressors, transcriptional profiles for each strain were obtained under four conditions: a non-stressful medium (YPD), two common stressors (YPD with 0.5 M NaCl and YPD at 37 °C), and a novel stressor inflicted by a synthetic antifungal drug (YPD with 1 μM clotrimazole (Clo)). NaCl induces osmotic stress22 and distorts intracellular Na+ concentration23, 37 °C induces heat stress, while clotrimazole, an antifungal drug developed in the 1960s24, inhibits ergosterol synthesis in fungal cell membranes25. As expected, growth rates were significantly reduced in stress environments, reaching only 50–70% of those in YPD (Fig. 1d). For each strain, transcriptome sequencing was performed in three biological replicates, yielding an average of 6.9 million reads per sample (Supplementary Data 3 and Supplementary Fig. 1). The high consistency among biological replicates (Pearson’s r = 0.98, P < 10−10; Supplementary Fig. 2) confirmed the robustness of the sequencing data.
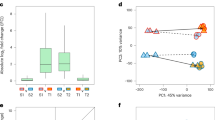
a Schematic representation of the experimental workflow. MA strains, derived from the ancestral strain BY4741 through a laboratory mutation accumulation experiment, and natural strains, collected from natural environments, were cultured in four distinct conditions. Transcriptome were sequenced using a high-throughput platform. The four colors represent four different environments, and the black and gray dots in the scatter plot correspond to natural strains and MA lines. Created in BioRender https://biorender.com/x07m898. b Phylogenetic relationship among the MA lines. c Phylogenetic relationship among the natural strains. Phylogenetic trees were reconstructed using the maximum likelihood method implemented in IQ-TREE (v3.0.1) based on all identified variant sites. The TVM substitution model was applied, as determined by model selection within IQ-TREE. d Growth rate of the ancestor BY4741, 15 MA lines, and 22 natural strains in the four environments, with the ancestral strain BY4741 represented by a black triangle. The center black lines represent the median; the box represents the 25th–75th percentiles; whiskers extend to 1.5× IQR; and points beyond the whiskers indicate outliers. P values were calculated using two-sided t-tests. “***” represents P < 0.001 and “****” represents P < 0.0001. e UMAP visualization of gene expression profiles for the ancestor BY4741, 15 MA lines, and 22 natural strains. Colors represent different environments, and shapes represent distinct strain groups.
Dimensionality reduction analysis of the transcriptome profiles revealed two major transcriptional clusters corresponding to MA and natural strains, with further subgrouping based on environmental conditions (Fig. 1e). These transcriptomic profiles were used to quantify gene expression plasticity by comparing expression levels across YPD and the three stress conditions (Fig. 1a).
Reduced expression plasticity in the antifungal environment after MA
To evaluate whether ancestral expression plasticity was selectively maintained, suppressed, or neutral, we analyzed changes in plasticity across MA lines. If ancestral plasticity was maintained by selection, MA lines would exhibit reduced plasticity due to the gradual erosion of its genetic basis during MA. Conversely, if plasticity was suppressed, MA lines would show increased plasticity, while a neutral model would predict random variation in plasticity levels relative to the progenitor.
Gene expression levels were quantified by normalizing expression levels using log2(FPKM + 1) (Supplementary Data 4 and 5). This transformation is commonly employed to stabilize variance and prevent issues associated with zero counts, ensuring more robust downstream analysis26. Plasticity for each gene was then calculated as the absolute difference in expression levels between stressed and YPD conditions27 (Supplementary Data 6–8). To assess overall plasticity, we summed the absolute plasticity across all genes for each strain (SumAcrossAllGenes(|expressionYPD − expressionstress|). For NaCl/YPD comparisons, 7 of 15 MA lines exhibited lower plasticity compared to the progenitor, a distribution that did not significantly different from randomness (Fig. 2a; two-sided binomial test, proportion = 0.47 (7/15), P = 1.00, 95% confidence interval = 0.21–0.73). A similar pattern was observed for 37/30 °C comparisons (Fig. 2b; two-sided binomial test, proportion = 0.33 (3/9), P = 0.51, 95% confidence interval = 0.07–0.70). However, for Clo/YPD comparisons, all 15 MA lines showed reduced plasticity (Fig. 2c; two-sided binomial test, proportion = 1.00 (15/15), P = 6.1 × 10−5, 95% confidence interval = 0.78–1.00).
a The overall NaCl/YPD plasticity across the MA lines. b The overall 37/30 °C plasticity across the MA lines. c The overall Clo/YPD plasticity across the MA lines. In a–c, overall expression plasticity is defined as SumAcrossAllGenes(|expressionYPD − expressionstress|), and the green dashed line represents the level of plasticity in the progenitor BY4741. d The percentage of genes showing elevated NaCl/YPD plasticity in comparison to the progenitor across all MA lines. e The percentage of genes showing elevated 37/30 °C plasticity in comparison to the progenitor across all MA lines. f The percentage of MA lines with elevated Clo/YPD plasticity compared with the progenitor across all genes. g The percentage of MA lines with elevated NaCl/YPD plasticity compared with the progenitor across all genes. h The percentage of MA lines with elevated 37/30 °C plasticity compared with the progenitor across all genes. i The percentage of MA lines with elevated Clo/YPD plasticity compared with the progenitor across all genes. In d–i, the green dashed line represents 50%. In all panels, the numbers indicate the number of samples or genes on each side of the green line and binomial test was performed between the distribution on the two sides of the green line. Purple bars represent MA lines while gray bars represent natural strains.
To further verify these findings, we conducted an additional analysis that weighted all genes equally. Specifically, for each MA line, we compared the expression plasticity of each gene to that of the progenitor and calculated the percentage of genes exhibiting increased plasticity. For NaCl/YPD, 6 of 15 MA lines had <50% of genes with increased plasticity, while the remaining 9 had >50% of genes with increased plasticity, consistent with neutrality (Fig. 2d; two-sided exact binomial test, proportion = 0.40 (6/15), P = 0.61, 95% confidence interval = 0.16–0.68). Again, the pattern is similar for the plasticity between 37 and 30 °C (Fig. 2e; two-sided binomial test, proportion = 0.56 (5/9), P = 1.00, 95% confidence interval = 0.21–0.86). In contrast, for Clo/YPD, all 15 MA lines had reduced plasticity across the majority of genes, demonstrated a reduced expression plasticity compared with their progenitor (Fig. 2f; two-sided binomial test, proportion = 1.0 (15/15), P = 6.1 × 10−5, 95% confidence interval = 0.78–1.00), suggesting that the overall expression plasticity in the progenitor was selectively maintained.
This pattern held true when analyzing individual gene plasticity across the yeast genome. For each of the 6263 genes, we calculated the proportion of MA lines with increased plasticity compared to the progenitor. No significant bias was detected for NaCl/YPD or 37/30 °C plasticity (Fig. 2g; two-sided binomial test, proportion = 0.50 (3127/6263), P = 0.92, 95% confidence interval = 0.49–0.51; Fig. 2h; two-sided binomial test, proportion = 0.49 (3087/6263), P = 0.27, 95% confidence interval = 0.48–0.51). However, for Clo/YPD, a significant majority of genes in the MA lines showed reduced plasticity, providing strong evidence that plasticity was selectively maintained in the progenitor for the antifungal environment (Fig. 2i; two-sided binomial test, proportion = 0.73 (4572/6263), P = 4.5 × 10−301, 95% confidence interval = 0.72–0.74).
A consistent pattern emerged when comparing MA lines with natural strains, with the largest differences observed in the Clo/YPD plasticity. In all forms of analysis (Fig. 2c, f, i), the magnitude of Clo/YPD plasticity was significantly lower in MA lines compared to natural strains.
Reduced number of significantly plastic genes in the antifungal environment after MA
To confirm that the above findings were not driven by stochastic variation in gene expression, we repeated the analyses using only genes with significant plasticity (SPGs) between YPD and stressed environments (SPGs were identified using DESeq2 with FDR < 0.05 and |log2FoldChange| > 1). In the progenitor, 516 SPGs were identified between NaCl and YPD, with 346–1466 SPGs across MA lines and natural strains (Fig. 3a). Among the 15 MA lines, 4 exhibited fewer SPGs than the progenitor, while 11 showed more, consistent with random expectations (Fig. 3b; two-sided binomial test, proportion = 0.27 (4/15), P = 0.12, 95% confidence interval = 0.08–0.55). Between 37 and 30 °C, 676 SPGs were found in the progenitor, and all the 9 MA lines exhibited more SPGs than the progenitor (Fig. 3c, d; two-sided binomial test, proportion = 0.00 (0/9), P = 0.0039, 95% confidence interval = 0.00–0.34). If anything, it suggest that the plastic expression under heat condition is selectively suppressed in the progenitor.
a Number of SPGs between NaCl and YPD in each strain. b Distribution of number SPGs between NaCl and YPD across the MA lines. c Number of SPGs between 37 and 30 °C in each strain. d Distribution of number SPGs between 37 and 30 °C across the MA lines. e Number of SPGs between clotrimazole and YPD in each strain. f Distribution of number SPGs between clotrimazole and YPD across the MA lines. In a, c, e, orange represents SPGs up-regulated in the stressed environments, while green represents SPGs down-regulated in the stressed environments. In b, d, f, the green dashed line represents the number of SPGs in the progenitor. The numbers indicate the number of MA lines on each side of the green line, and binomial test was performed between the distribution on the two sides of the green line.
Between Clo and YPD conditions, 1876 SPGs were found in the progenitor, while MA lines and natural strains exhibited 234–2237 SPGs (Fig. 3e). Importantly, all 15 MA lines showed reduced numbers of SPGs compared to the progenitor (Fig. 3f; two-sided binomial test, proportion = 1.0 (15/15), P = 6.1 × 10−5, 95% confidence interval = 0.78–1.00), with an average of 605 SPGs in MA lines—less than half of the progenitor’s count. This significant reduction in transcriptional responsiveness highlights the role of selection in maintaining expression plasticity under clotrimazole stress.
Directional reversal of plasticity in the antifungal environment after MA
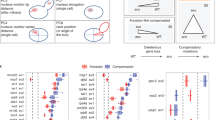
While the above analysis focused on changes in the magnitude of expression plasticity, we further investigated the evolutionary trajectory of plasticity by examining its directionality. To this end, we adapted a modified method from prior studies6,15 to categorize plastic expression changes into two distinct categories: codirectional and reversed. Plastic changes were classified as codirectional if the MA line retained the same directional response as the progenitor, regardless of magnitude (e.g., the expression level of a gene was higher in stressed than non-stressed conditions in both the MA line and the progenitor). Conversely, changes were classified as reversed if the direction of plasticity shifted in the opposite direction relative to the progenitor (e.g., the expression level of a gene was higher in stressed versus non-stressed conditions in the progenitor but lower under the same conditions in the MA line) (Fig. 4a).
a Classification of expression plasticity into codirectional and reversed. On the x-axis, “O” represents the expression level in the original environment and “P” represents the expression level in the new environment. The left subpanel illustrates the concept of ancestral plasticity, and used in the right subpanel as a reference. Orange represents codirectional plasticity, while blue represents reversed plasticity. b The proportions of codirectional and reversed NaCl/YPD plasticity in MA lines and natural strains. c The proportions of codirectional and reversed 37/30 °C plasticity in MA lines and natural strains. d The proportions of codirectional and reversed Clo/YPD plasticity in MA lines and natural strains. e Comparisons of the proportion of codirectional and reversed NaCl/YPD plasticity between 15 MA lines and 22 natural strains. f Comparisons of the proportion of codirectional and reversed 37/30 °C plasticity between 9 MA lines and 22 natural strains. g Comparisons of the proportion of codirectional and reversed Clo/YPD plasticity between 15 MA lines and 22 natural strains. In b–g, purple represents MA lines and yellow represents natural strains. In e–g, t-test is performed between every two bins, with “NS” represents not significant and “***” represents P < 0.001. In e–g, the center line represents the median; the box represents the 25th–75th percentiles; whiskers extend to 1.5× IQR; and points beyond the whiskers indicate outliers. P values were calculated using two-sided t-tests with Benjamini–Hochberg correction.
We hypothesized that if the directionality of ancestral plasticity was selectively maintained, reversed plasticity would occur more frequently than codirectional plasticity. This is because, in the absence of selective constraints, random mutations accumulating in MA lines are more likely to disrupt the regulatory mechanisms that sustain the ancestral plastic response. To test this, we generated a null distribution of genes between the two categories by treating the 22 natural strains as controls and comparing their plastic expression profiles with the progenitor of the MA lines. We classified all genes in the MA lines and natural strains based on plastic expression profiles in the three stressful environments (Fig. 4b–d) and compared the proportions of genes in each category between MA and natural strains.
For NaCl/YPD and 37/30 °C plasticity, no significant differences were found across the two categories between MA and natural strains (Fig. 4e, two sided t-test, P = 0.10; Fig. 4f, two-sided t-test, P = 0.09), consistent with neutral expectation. However, significant differences were observed for Clo/YPD plasticity. The MA lines exhibited significantly fewer genes with codirectional plasticity and significantly more genes with reversed plasticity compared to natural strains (Fig. 4g; two-sided t-test, P = 3.0 × 10−7). The increased levels of reversed plasticity in MA lines further supports the hypothesis that expression plasticity in the progenitor was selectively maintained.
To further dissect these patterns, we refined our classification into six categories, separating ancestral inductions and suppressions under stress into three subtypes each: strengthened, weakened, and reversed (Supplementary Fig. 3a). Strengthened plasticity indicates that the direction of the response is the same in the MA line and progenitor but with a greater magnitude in the MA line, suggesting that ancestral plasticity was selectively suppressed. Conversely, weakened and reversed plasticity imply that ancestral plasticity was selectively maintained. Consistent with our previous findings, the proportion of strengthened plasticity was significantly lower under clotrimazole treatment, whereas the proportions of weakened and reversed plasticity were significantly higher (Supplementary Fig. 3b). Similar trends were observed for ancestral suppressions, although the differences among the three stressful environments were less pronounced (Supplementary Fig. 3c). Collectively, these results indicate that expression plasticity in the clotrimazole environment was selectively maintained.
Shared pathway loss of stress response in the antifungal environment after MA
Our results indicate that gene expression plasticity in the antifungal environment is largely adaptive, suggesting that it provides functional benefits to yeast cells under stress. To identify the specific biological functions associated with this plasticity, we first identified genes with significant plasticity (SPGs) in the progenitor, MA lines, and natural strains. Gene ontology (GO) enrichment analysis was then performed to identify overrepresented biological processes. If a biological process was functionally advantageous in the antifungal environment, it would likely be present in most natural strains but lost in some MA lines. To test this, we compared the presence of progenitor-enriched biological processes between natural strains and MA lines, analyzing up-regulated and down-regulated genes separately. This distinction is important because activators and repressors involved in the same biological process often exhibit opposite patterns of expression change under stress.
For genes up-regulated under clotrimazole environment, we identified 11 biological processes that were significantly enriched in the progenitor and natural strains but largely absent in MA lines (Fig. 5; Fisher’s exact test, P-adjust < 0.05). These processes were frequently detected in natural strains (present in at least 12 of 22 strains) but rarely observed in MA lines (at most in 1 of 15 lines). Notably, processes associated with general stress response, such as response to oxidative stress and response to chemical stimulus, were among the most enriched. Oxidative stress response is commonly induced when yeast cells face diverse environmental stresses19, and azole antifungal drugs, including clotrimazole, are known to cause reactive oxygen species (ROS) accumulation in fungi28,29. Similarly, cellular response to chemical stimulus is frequently activated when cells encounter a wide range of toxic chemicals30,31. These findings suggest that even stressors absent from yeast’s evolutionary history can still be mitigated through general stress–response mechanisms. Other enriched biological processes included pathways involved in carbohydrate metabolism and purine and pyrimidine metabolism. Clotrimazole inhibits glucose metabolism by targeting the key glycolytic enzyme 6-phosphofructo-1-kinase32. This aligns with findings that clotrimazole is more effective against fungal species like C. albicans that rely on glucose rather than alternative carbon sources, such as lactic acid33. Furthermore, elevated ROS levels can lead to DNA damage34, necessitating enhanced nucleotide synthesis for DNA repair.
Heatmap illustrating GO categories enriched in DEGs up-regulated under clotrimazole stress. Each column represents a sample, and each row represents a GO category. Green, purple, and yellow represent the progenitor BY4741, MA lines, and natural strains, respectively. Red highlights GO categories related to the general stress response. Enrichment in progenitor but not in MA lines was assessed using two-sided Fisher’s exact tests, P values were adjusted for multiple comparisons using the Benjamini–Hochberg false discovery rate method.
In contrast, for genes up-regulated under NaCl stress, enriched pathways were mainly related to amino acid metabolism and showed no direct connection to canonical osmotic stress responses, such as the HOG pathway22,35 (Supplementary Fig. 4). No significant enrichment was observed for genes up-regulated under heat stress, nor for genes down-regulated in any of the stressed conditions. In summary, the up-regulated pathways in the progenitor and natural strains under antifungal stress likely conferred functional advantages by activating general stress responses, while their loss in MA lines underscores the selective maintenance of these pathways in natural populations.
Because our transcriptomic analyses revealed that genes associated with the oxidative stress response were among the most frequently lost pathways in MA lines under clotrimazole treatment, we hypothesized that this loss might contribute directly to reduced stress tolerance. To test this, we measured the competitive fitness of 15 MA lines in media containing 1 μM clotrimazole and 0.1 mM hydrogen peroxide (H₂O₂), the latter serving as a well-established inducer of oxidative stress. If the oxidative stress response pathway underlies the adaptive plasticity to clotrimazole, then MA lines impaired in this pathway should show parallel fitness reductions in both environments. Consistent with this prediction, we observed a significant positive correlation between fitness reductions (relative to YPD) across the two conditions (Pearson’s r = 0.62, P = 0.014; Supplementary Fig. 5). These results suggest that the loss of oxidative stress response plasticity in MA lines contributes to decreased fitness under both clotrimazole and oxidative stress, supporting the conclusion that adaptive expression plasticity in the progenitor involves activation of a general oxidative stress response pathway.
Discussion
The results of this study provide compelling evidence that gene expression plasticity is not only a fundamental biological characteristic but also plays an adaptive role in response to environmental stressors, such as antifungal agents. Through a direct comparison between MA lines and natural strains of Saccharomyces cerevisiae, we demonstrated that gene expression plasticity in response to antifungal stress is predominantly adaptive, whereas plasticity in response to osmotic and heat stresses appears neutral, showing no clear evidence of selective maintenance. These findings offer insights into how microbial populations respond to environmental challenges and how natural selection shapes the regulation of gene expression over evolutionary time.
A key finding of this study is the significant reduction in gene expression plasticity observed in the MA lines under antifungal stress. To quantify plasticity, we applied a widely used approach that measures differences in expression levels between stressed and non-stressed conditions27. To ensure the robustness of our conclusions, we also employed a normalized difference metric, where plasticity is calculated as the difference in expression levels between the two conditions divided by the sum of expression levels in both conditions ((|expressionYPD − expressionstress|)/(expressionYPD + expressionstress)). Both approaches yielded consistent results, reinforcing the conclusion that overall expression plasticity was significantly reduced in MA lines in the antifungal environment (Supplementary Fig. 6). Additionally, the ancestral plasticity in the antifungal environment was more frequently reversed (Supplementary Fig. 7). In contrast, under NaCl and heat stresses, no significant evidence of selective maintenance of plasticity was detected (Supplementary Figs. 6 and 7). The transcriptome clustering further supports adaptive plasticity under clotrimazole, as the progenitor clustered with natural strains rather than MA lines (Fig. 1e). This pattern suggests that the progenitor and natural strains share conserved transcriptional responses advantageous for coping with clotrimazole, whereas these responses were largely lost in the MA lines due to relaxed selection.
The loss of plasticity in MA lines under antifungal stress is consistent with the idea that expression plasticity provides a selective advantage by enabling organisms to rapidly adjust their gene expression profiles in response to changing environmental conditions. However, since clotrimazole is a synthetic drug developed in the 1960s and has not been part of the long evolutionary history of budding yeasts, it raises the question of how yeast cells can still maintain adaptive expression plasticity when exposed to clotrimazole. Our analysis of GO categories for genes induced by clotrimazole, which are lost in the MA lines but preserved in the natural strains, identified two key categories: response to oxidative stress and response to chemical stimulus. These categories are integral to the yeast’s general stress response pathway and are often activated under a wide range of stressors19. These findings suggest that, even when exposed to novel stressors not previously encountered by yeast cells, they can still utilize these general stress response pathways to manage such challenges.
Interestingly, our analysis also revealed that plasticity in response to common stressors, such as osmotic stress and heat stress, did not exhibit the same adaptive pattern seen in the antifungal environment. The MA lines showed random deviations in plasticity compared to the progenitors, without any consistent trend toward reduced or enhanced plasticity. This suggests that gene expression plasticity in response to osmotic and heat stress is likely neutral, as there appears to be no clear selective pressure to maintain or alter these plastic responses in the absence of natural selection. While this conclusion might initially seem counterintuitive, it may be explained by recurrent genetic assimilation. Genetic assimilation, first described by Waddington, occurs when environmentally induced phenotypes (i.e., phenotypic plasticity) become fixed in a population over a few generations, eliminating the need for plasticity18. Later, Lande developed a quantitative-genetic model that demonstrated how a transient increase in phenotypic plasticity, followed by genetic assimilation, can drive adaptation to a novel environment5. The strongest experimental evidence for genetic assimilation comes from a recent study on Silene uniflora36, which has repeatedly adapted to zinc-contaminated soils over the last 250 years. By sequencing the transcriptomes of both zinc-sensitive and zinc-tolerant lineages of Silene uniflora in both zinc-containing and zinc-free environments, researchers identified 183 genes whose plastic expression was fixed in the adapted lineages, providing direct evidence for genetic assimilation. In contrast to the recent 250-year exposure to zinc in Silene uniflora, osmotic and heat stressors in yeast have likely occurred intermittently for millions of years during their evolutionary history. Therefore, much of the adaptive ancestral plastic expression to these stressors may have already been fixed through genetic assimilation. This could explain the neutral pattern of plasticity we observed under NaCl and heat stress. The neutral behavior of plasticity in these contexts contrasts with the strongly adaptive response observed under antifungal stress, highlighting how the evolutionary significance of plasticity can vary depending on the environmental context. This finding aligns with a previous study on the expression plasticity of the TDH3 gene in yeast, where evidence of adaptive plasticity was observed in some environments but not others37.
While our study primarily focused on the overall pattern of plastic expression, we acknowledge that certain genes may still exhibit adaptive plasticity under NaCl stress. For example, the core osmoadaptive modules, particularly the HOG pathway, ENA1, and GPD1/GPD2, remain essential and show strong, adaptive induction in response to NaCl stress19,38. However, our data suggest that beyond this essential core, the broader pattern of transcriptome-wide plastic expression changes appears largely random across MA lines, which is consistent with neutral variation in many genes. This observation indicates that while selection continues to preserve the critical plastic responses needed for osmotic balance and ion homeostasis, many peripheral transcriptional responses may evolve under relaxed selection or drift. In this way, adaptive plasticity is maintained where it is functionally necessary, but the entire stress-responsive network is not globally optimized.
In our MA experiment, we used an MSH2 deletion strain to increase the mutation rate, which also alters the mutational spectrum by elevating the frequency of base substitutions and small insertion–deletion events, particularly in repetitive regions. While this spectrum differs somewhat from that of natural yeast populations, the primary purpose of MA experiments is to allow random mutations to accumulate under minimal selection, thereby gradually eroding the genome. In this sense, even though the precise mutation spectrum may differ, it serves the same functional role as naturally occurring mutations in revealing how random genetic changes affect gene expression and plasticity. Because our analyses focus on the selective patterns acting on these phenotypic outcomes rather than the specific mutation mechanisms, the conclusions regarding the selective maintenance or neutrality of expression plasticity should remain valid.
It is worth noting that the neutral pattern of expression plasticity observed under NaCl and heat stress likely reflects the moderate intensity of these treatments, which approximate conditions commonly experienced by yeast in nature. If substantially higher concentrations of NaCl or more extreme temperatures were applied, representing conditions that yeast rarely encounter in their evolutionary history, these stressors might effectively serve as novel environments. Under such circumstances, we might expect to observe a different pattern of selective maintenance, potentially resembling the adaptive response detected under clotrimazole stress. Future studies examining a gradient of stress intensities would therefore be valuable for determining the threshold at which familiar environmental stresses begin to elicit adaptive plastic responses.
Methods
Strains and media
This study utilized the commonly used laboratory yeast strain BY4741, 15 MA lines derived from BY474120, and 22 diverse natural strains21 (Supplementary Data 2). To accumulate mutations more efficiently, the MA lines were generated in a mutator genetic background, with each line accumulating approximately 900 random mutations on average. Following MA, the non-mutator genotype was restored for each MA line20. The natural strains were subjected to sporulation, polyploid selection, and genetic modifications as described by Maclean et al.21. Specifically, the HO and URA3 genes in the natural strains were replaced with antibiotic markers, and all strains were maintained in haploid form.
Yeast strains were cultured in four media: standard YPD, YPD supplemented with 0.5 M NaCl (osmotic stress), YPD at 37 °C (heat stress), and YPD supplemented with 1 µM clotrimazole (antifungal stress). The YPD medium was prepared by dissolving 20 g glucose, 20 g peptone, and 10 g yeast extract in 1 L of distilled deionized water (ddH2O), followed by autoclaving at 121 °C for 30 min. For osmotic stress conditions, 29.25 g of sodium chloride was added to 1 L of YPD medium to achieve a final concentration of 0.5 M NaCl. For antifungal stress conditions, 1.72 g of clotrimazole was dissolved in 50 mL of anhydrous ethanol, filter-sterilized, and added to 1 L of YPD medium to achieve a final concentration of 1 µM.
Sample preparation, transcriptome, and genome sequencing
To prepare samples for transcriptome sequencing, all strains were pre-adapted in the four media (YPD, YPD + NaCl, YPD at 37 °C, and YPD + clotrimazole) for a full growth cycle, from initial inoculation to saturation. A 15 µL aliquot of each culture was inoculated into 985 µL of fresh medium to achieve a final volume of 1 mL. Cultures were incubated at 30 °C, and optical density (OD) was measured every 1 h. Cells were harvested during the logarithmic growth phase (OD 0.4–0.6) by centrifugation at 13,800 × g for 1 min. The supernatant was discarded, and the cell pellets were flash-frozen in liquid nitrogen. Each strain was cultured in triplicate for each medium condition.
RNA extraction, reverse transcription, library preparation, and high-throughput sequencing were performed by Genewiz (https://climsprod.genewiz.com.cn). Sequencing was conducted on the Illumina NovaSeq 6000 platform using a PE150 configuration, generating an average of 6.9 million paired-end reads per sample (Supplementary Data 3).
Genome sequencing was performed on the 15 MA lines initially selected, using the Illumina HiSeq platform with a target output of 1 Gb per sample. Mutation calling was conducted as described previously20. Briefly, sequencing reads were first aligned using BWA39, followed by sorting and duplicate marking with Picard (https://broadinstitute.github.io/picard/). Finally, variants were called using GATK40.
Data analysis
Raw sequencing data were preprocessed using fastp (v0.23.2)41 to remove low-quality reads with default parameters. The Saccharomyces cerevisiae reference genome (R64) and corresponding annotation files were obtained from the Yeast Genome Database (https://www.yeastgenome.org/). Filtered reads were aligned to the reference genome using HISAT2 (v2.2.1)42 and sorted using SAMtools (v1.20)43. Gene-level read counts were quantified using featureCounts44 implemented in the subread package (v2.0.1). Normalized expression levels, expressed as fragments per kilobase per million (FPKM), and significant plastic genes (SPGs) were defined as genes with expression level differ significantly between two environments and were identified using the DESeq2 package (v1.42.1)45 in R. GO enrichment analysis was performed using clusterProfiler (v4.10.1)46.
Expression plasticity was quantified using two methods. First, FPKM values for each gene were normalized using log2(FPKM + 1) to represent gene expression levels (Supplementary Data 4 and 5). Expression plasticity was calculated as the difference between expression levels under stress and non-stress conditions (Supplementary Data 6–8). To ensure robustness, a second method was employed, where expression plasticity was calculated as the normalized difference, which is the difference between stress and non-stress expression levels divided by the sum of the two expression levels.
Growth rate measurement and competition assay
The maximum growth rates of yeast strains in the three media were measured as described previously47. Briefly, strains were pre-adapted in the respective medium for a full growth cycle. Saturated cultures were diluted 10-fold, and 3 µL of the diluted culture was added to 197 µL of fresh medium in a 96-well plate (final volume: 200 µL per well). Plates were incubated at 30 °C in a microplate reader, and OD at 660 nm was recorded every 20 min for 48–72 h (Supplementary Figs. 8 and 9). OD measurements were converted to cell densities using a pre-determined OD-cell density calibration curve. Slopes were calculated for every four measurements spaced 60 min apart along the growth curve by Δln(cell density)/60. To avoid potential artifacts, the two highest slopes were discarded, and the mean of the third to eighth highest slopes was used as the growth rate per minute38,47.
Competition assay was performed following a previous study48. Specifically, YPD and clotrimazole-containing YPD media were prepared as described above. Hydrogen peroxide (H2O2) solution was prepared by adding 0.4 µL of 30% H2O2 to 40 mL of YPD, yielding a final concentration of 0.003‰, which is approximately 0.1 mM. The ADE2 gene was knocked out in BY4741 to generate the Ade2-marked strain, which form pink colonies on low-adenine media. SC medium was used to visualize the pink colonies of the progenitor strain. Competitions were performed in liquid media between each MA strain and the Ade2-marked progenitor, with wildtype BY4741 versus Ade2-marked BY4741 competitions included as controls. Prior to competition, both progenitor and MA strains were pre-cultured in the corresponding environment until reaching an OD600 of approximately 0.3. For each competition, 10 µL of MA culture and 5 µL of Ade2 progenitor culture were inoculated into 1 mL of the corresponding medium. This initial time point was designated as t0. Cultures were incubated at 30 °C with shaking at 250 rpm for 12 h, corresponding to t1. At t1, cells were diluted and plated on SC medium. Plates were incubated at 30 °C for > 2 days and then stored at room temperature for an additional day to allow clear visualization of colonies.
The relative fitness of each MA strain was calculated with respect to the Ade2 progenitor. The relative growth rate, s, was calculated using the formula:
where T1 represents the ratio of MA (white colonies) to Ade2 (pink colonies) at t1, and \(t\) is the duration of the competition experiment, here 12 h. The relative fitness, f, was subsequently calculated as:
Generation times for each environment were determined by monitoring cell number changes of BY4741 over the 12-h period. The estimation of MA strains’ relative fitness finally normalized to the Ade2-marked progenitor versus progenitor across different environmental conditions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided as a Source data file. All sequencing reads generated in this study have been deposited in the China National Genomics Data Center under the accession number CRA033816. Source data are provided with this paper.
References
Sultan, S. E. Phenotypic plasticity as an intrinsic property of organisms. in Phenotypic Plasticity and Evolution (CRC Press, 2021).
Futuyma, D. J. How does phenotypic plasticity fit into evolutionary theory? in Phenotypic Plasticity and Evolution (CRC Press, 2021).
Phenotypic Plasticity and Evolution: Causes, Consequences, Controversies https://doi.org/10.1201/9780429343001 (CRC Press, 2021).
Ghalambor, C. K., McKAY, J. K., Carroll, S. P. & Reznick, D. N. Adaptive versus non-adaptive phenotypic plasticity and the potential for contemporary adaptation in new environments. Funct. Ecol. 21, 394–407 (2007).
Lande, R. Adaptation to an extraordinary environment by evolution of phenotypic plasticity and genetic assimilation. J. Evol. Biol. 22, 1435–1446 (2009).
Chen, P. & Zhang, J. Transcriptomic analysis reveals the rareness of genetic assimilation of gene expression in environmental adaptations. Sci. Adv. 9, eadi3053 (2023).
West-Eberhard, M. J. Developmental Plasticity and Evolution (Oxford University Press, 2003).
Laland, K. N. et al. The extended evolutionary synthesis: its structure, assumptions and predictions. Proc. R. Soc. B Biol. Sci. 282, 20151019 (2015).
Laland, K. et al. Does evolutionary theory need a rethink? Nature 514, 161–164 (2014).
Pfennig, D. W. et al. Phenotypic plasticity’s impacts on diversification and speciation. Trends Ecol. Evol. 25, 459–467 (2010).
Moczek, A. P. et al. The role of developmental plasticity in evolutionary innovation. Proc. R. Soc. B Biol. Sci. 278, 2705–2713 (2011).
Levis, N. A. & Pfennig, D. W. Evaluating ‘plasticity-first’ evolution in nature: key criteria and empirical approaches. Trends Ecol. Evol. 31, 563–574 (2016).
Robinson, B. W. & Dukas, R. The Influence of phenotypic modifications on evolution: the Baldwin effect and modern perspectives. Oikos 85, 582–589 (1999).
Price, T. D., Qvarnström, A. & Irwin, D. E. The role of phenotypic plasticity in driving genetic evolution. Proc. R. Soc. Lond. Ser. B Biol. Sci. 270, 1433–1440 (2003).
Ho, W.-C. & Zhang, J. Evolutionary adaptations to new environments generally reverse plastic phenotypic changes. Nat. Commun. 9, 350 (2018).
López-Maury, L., Marguerat, S. & Bähler, J. Tuning gene expression to changing environments: from rapid responses to evolutionary adaptation. Nat. Rev. Genet. 9, 583–593 (2008).
Ghalambor, C. K. et al. Non-adaptive plasticity potentiates rapid adaptive evolution of gene expression in nature. Nature 525, 372–375 (2015).
Waddington, C. H. Genetic assimilation of an acquired character. Evolution 7, 118–126 (1953).
Gasch, A. P. et al. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 11, 4241–4257 (2000).
Liu, H. & Zhang, J. The rate and molecular spectrum of mutation are selectively maintained in yeast. Nat. Commun. 12, 4044 (2021).
Maclean, C. J. et al. Deciphering the genic basis of yeast fitness variation by simultaneous forward and reverse genetics. Mol. Biol. Evol. 34, 2486–2502 (2017).
Hohmann, S. Osmotic stress signaling and osmoadaptation in yeasts. Microbiol. Mol. Biol. Rev. 66, 300–372 (2002).
Illarionov, A., Lahtvee, P.-J. & Kumar, R. Potassium and sodium salt stress characterization in the yeasts Saccharomyces cerevisiae, Kluyveromyces marxianus, and Rhodotorula toruloides. Appl. Environ. Microbiol. https://doi.org/10.1128/AEM.03100-20 (2021).
Crowley, P. D. & Gallagher, H. C. Clotrimazole as a pharmaceutical: past, present and future. J. Appl. Microbiol. 117, 611–617 (2014).
Haller, I. Mode of action of clotrimazole: implications for therapy. Am. J. Obstet. Gynecol. 152, 939–944 (1985).
Bei, C. et al. Genetically encoded transcriptional plasticity underlies stress adaptation in Mycobacterium tuberculosis. Nat. Commun. 15, 3088 (2024).
Valladares, F., Sanchez-Gomez, D. & Zavala, M. A. Quantitative estimation of phenotypic plasticity: bridging the gap between the evolutionary concept and its ecological applications. J. Ecol. 94, 1103–1116 (2006).
Sellers-Moya, Á, Nuévalos, M., Molina, M. & Martín, H. Clotrimazole-induced oxidative stress triggers novel yeast Pkc1-independent cell wall integrity MAPK pathway circuitry. J. Fungi 7, 647 (2021).
Yardımcı, B. K. & Tarhan, L. Oxidative effects of the human antifungal drug clotrimazole on the eucaryotic model organism Saccharomyces cerevisiae. Arch. Microbiol. 206, 290 (2024).
Chi, W.-C. et al. Autotoxicity mechanism of Oryza sativa: transcriptome response in rice roots exposed to ferulic acid. BMC Genom. 14, 351 (2013).
Li, Y. et al. Using transcriptomics, proteomics and phosphoproteomics as new approach methodology (NAM) to define biological responses for chemical safety assessment. Chemosphere 313, 137359 (2023).
Zancan, P., Rosas, A. O., Marcondes, M. C., Marinho-Carvalho, M. M. & Sola-Penna, M. Clotrimazole inhibits and modulates heterologous association of the key glycolytic enzyme 6-phosphofructo-1-kinase. Biochem. Pharm. 73, 1520–1527 (2007).
Kasper, L. et al. Antifungal activity of clotrimazole against Candida albicans depends on carbon sources, growth phase and morphology. J. Med. Microbiol. 64, 714–723 (2015).
Srinivas, U. S., Tan, B. W. Q., Vellayappan, B. A. & Jeyasekharan, A. D. ROS and the DNA damage response in cancer. Redox Biol. 25, 101084 (2018).
de Nadal, E. & Posas, F. The HOG pathway and the regulation of osmoadaptive responses in yeast. FEMS Yeast Res. 22, foac013 (2022).
Wood, D. P. et al. Genetic assimilation of ancestral plasticity during parallel adaptation to zinc contamination in Silene uniflora. Nat. Ecol. Evol. 7, 414–423 (2023).
Duveau, F., Yuan, D. C., Metzger, B. P. H., Hodgins-Davis, A. & Wittkopp, P. J. Effects of mutation and selection on plasticity of a promoter activity in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 114, E11218–E11227 (2017).
Warringer, J., Ericson, E., Fernandez, L., Nerman, O. & Blomberg, A. High-resolution yeast phenomics resolves different physiological features in the saline response. Proc. Natl. Acad. Sci. USA 100, 15724–15729 (2003).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2, e107 (2023).
Zhang, Y., Park, C., Bennett, C., Thornton, M. & Kim, D. Rapid and accurate alignment of nucleotide conversion sequencing reads with HISAT-3N. Genome Res. 31, 1290–1295 (2021).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Liao, Y., Smyth, G. K. & Shi, W. The subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Xu, H. et al. Synchronization of stochastic expressions drives the clustering of functionally related genes. Sci. Adv. 5, eaax6525 (2019).
Sharon, E. et al. Functional genetic variants revealed by massively parallel precise genome editing. Cell 175, 544–557 (2018).
Acknowledgements
We thank the three anonymous reviewers for their constructive comments. H.L. is supported by the National Key Research and Development Program of China (2024YFA1802500) and the National Natural Science Foundation of China (32470646). P.C. is supported by the National Key Research and Development Program of China (2024YFA1802500) and the National Natural Science Foundation of China (32470650).
Author information
Authors and Affiliations
Contributions
H.L., P.C. conceived the project and designed the experiment. Y.Q., Z.Y., Y.Y., and T.Q. carried out the experiment and analyzed the data. H.L., P.C., and Y.Q. wrote the manuscript. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qian, Y., Yao, Z., Yao, Y. et al. Contrasting selective signatures of gene expression plasticity in response to antifungal clotrimazole and common environmental stresses in yeast. Nat Commun 17, 4467 (2026). https://doi.org/10.1038/s41467-026-71153-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-71153-0