Abstract
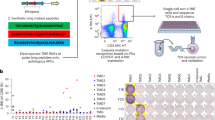
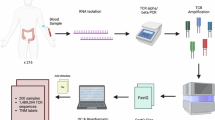
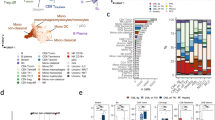
Lynch Syndrome (LS) provides the perfect context to understand DNA mismatch repair deficient carcinogenesis, which is characterized by neoplastic lesions with high rates of shared neoantigens eliciting adaptive immunity through T cell receptor (TCR) recognition. However, the TCR landscape in LS carriers remains unexplored. Here, we perform TCR sequencing of 277 blood samples from LS cancer survivors, previvors, and controls, as well as matching colorectal cancers and pre-cancers. We show that up to 41% of the most expanded TCRβs from colorectal neoplasms are detectable in the blood of LS carriers, while showing minimal expansion in controls. In addition, we develop and validate a classification model that distinguishes LS carriers from controls using circulating TCRβs signatures associated with LS independent of thecancer history and with cancer-free LS previvors. Together, our findings characterize circulating and tissue TCRβs associated with LS, thus representing a step toward identifying blood-based TCR biomarkers for immune surveillance.
Similar content being viewed by others
Data availability
The TCRseq data generated in this study are publicly available in Zenodo (https://zenodo.org/records/13141052) and the scRNA-seq data generated in this study are publicly available in GEO under accession number GSE289646 Source data are provided with this paper.
Code availability
The code used in this study is publicly available at https://github.com/Vilarlab-MDACC/LS_TCR.
References
Abu-Ghazaleh, N., Kaushik, V., Gorelik, A., Jenkins, M. & Macrae, F. Worldwide prevalence of Lynch syndrome in patients with colorectal cancer: Systematic review and meta-analysis. Genet Med 24, 971–985 (2022).
Lynch, H. T., Snyder, C. L., Shaw, T. G., Heinen, C. D. & Hitchins, M. P. Milestones of lynch syndrome: 1895–2015. Nat. Rev. Cancer 15, 181–194 (2015).
Te Paske, I. et al. Noncoding aberrations in mismatch repair genes underlie a substantial part of the missing heritability in lynch syndrome. Gastroenterology 163, 1691–1694 e7 (2022).
Dominguez-Valentin, M. et al. Correction: Cancer risks by gene, age, and gender in 6350 carriers of pathogenic mismatch repair variants: findings from the prospective lynch syndrome database. Genet Med 22, 1569 (2020).
Bolivar, A. M., Duzagac, F., Sinha, K. M. & Vilar, E. Advances in vaccine development for cancer prevention and treatment in Lynch Syndrome. Mol. Asp. Med 93, 101204 (2023).
Ballhausen, A. et al. The shared frameshift mutation landscape of microsatellite-unstable cancers suggests immunoediting during tumor evolution. Nat. Commun. 11, 4740 (2020).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med 372, 2509–2520 (2015).
Roudko, V. et al. Shared Immunogenic poly-epitope frameshift mutations in microsatellite unstable tumors. Cell 183, 1634–1649 e17 (2020).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Alcover, A., Alarcon, B. & Di Bartolo, V. Cell biology of T cell receptor expression and regulation. Annu Rev. Immunol. 36, 103–125 (2018).
Tsunoda, M. et al. Proportional tumor infiltration of T cells via circulation duplicates the T cell receptor repertoire in a bilateral tumor mouse model. Front Immunol. 12, 744381 (2021).
Venturi, V., Price, D. A., Douek, D. C. & Davenport, M. P. The molecular basis for public T-cell responses?. Nat. Rev. Immunol. 8, 231–238 (2008).
Kirsch, I. R. et al. TCR sequencing facilitates diagnosis and identifies mature T cells as the cell of origin in CTCL. Sci. Transl. Med 7, 308ra158 (2015).
Weng, W. K. et al. Minimal residual disease monitoring with high-throughput sequencing of T cell receptors in cutaneous T cell lymphoma. Sci. Transl. Med 5, 214ra171 (2013).
Emerson, R. O. et al. Immunosequencing identifies signatures of cytomegalovirus exposure history and HLA-mediated effects on the T cell repertoire. Nat. Genet 49, 659–665 (2017).
Snyder, T. M. et al. Magnitude and dynamics of the T-cell response to SARS-CoV-2 infection at both individual and population levels. Front. Immunol. 15, 1488860 (2025).
Meier, J. et al. Fractal organization of the human T cell repertoire in health and after stem cell transplantation. Biol. Blood Marrow Transpl. 19, 366–377 (2013).
Oakes, T. et al. Quantitative characterization of the T cell receptor repertoire of naive and memory subsets using an integrated experimental and computational pipeline which is robust, economical, and versatile. Front Immunol. 8, 1267 (2017).
Tickotsky, N., Sagiv, T., Prilusky, J., Shifrut, E. & Friedman, N. McPAS-TCR: a manually curated catalogue of pathology-associated T cell receptor sequences. Bioinformatics 33, 2924–2929 (2017).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Zaslavsky, M. E. et al. Disease diagnostics using machine learning of B cell and T cell receptor sequences. Science 387, eadp2407 2025).
Bolivar, A. M. et al. Genomic Landscape of Lynch Syndrome Colorectal Neoplasia Identifies Shared Mutated Neoantigens for Immunoprevention. Gastroenterology 166, 787–801 e11 (2024).
Wang, X. et al. T cell repertoire in peripheral blood as a potential biomarker for predicting response to concurrent cetuximab and nivolumab in head and neck squamous cell carcinoma. J. Immunother. Cancer 10, e004512 (2022).
Lin, K. R. et al. Circulating CD8(+) T-cell repertoires reveal the biological characteristics of tumors and clinical responses to chemotherapy in breast cancer patients. Cancer Immunol. Immunother. 67, 1743–1752 (2018).
Cui, J. H. et al. TCR repertoire as a novel indicator for immune monitoring and prognosis assessment of patients with cervical cancer. Front Immunol. 9, 2729 (2018).
Liu, Y. Y. et al. Characteristics and prognostic significance of profiling the peripheral blood T-cell receptor repertoire in patients with advanced lung cancer. Int J. Cancer 145, 1423–1431 (2019).
Castro, A. et al. Strength of immune selection in tumors varies with sex and age. Nat. Commun. 11, 4128 (2020).
Sun, X. et al. Longitudinal analysis reveals age-related changes in the T cell receptor repertoire of human T cell subsets. J. Clin. Invest. 132, e158122(2022).
Bai, X. et al. Characteristics of tumor infiltrating lymphocyte and circulating lymphocyte repertoires in pancreatic cancer by the sequencing of T cell receptors. Sci. Rep. 5, 13664 (2015).
Sherwood, A. M. et al. Tumor-infiltrating lymphocytes in colorectal tumors display a diversity of T cell receptor sequences that differ from the T cells in adjacent mucosal tissue. Cancer Immunol. Immunother. 62, 1453–1461 (2013).
Beausang, J. F. et al. T cell receptor sequencing of early-stage breast cancer tumors identifies altered clonal structure of the T cell repertoire. Proc. Natl. Acad. Sci. USA 114, E10409–E10417 (2017).
Cowell, L. G. The diagnostic, prognostic, and therapeutic potential of adaptive immune receptor repertoire profiling in cancer. Cancer Res 80, 643–654 (2020).
Emerson, R. O. et al. High-throughput sequencing of T-cell receptors reveals a homogeneous repertoire of tumour-infiltrating lymphocytes in ovarian cancer. J. Pathol. 231, 433–440 (2013).
Chen, Z. et al. T cell receptor beta-chain repertoire analysis reveals intratumour heterogeneity of tumour-infiltrating lymphocytes in oesophageal squamous cell carcinoma. J. Pathol. 239, 450–458 (2016).
Pavlovic, M. et al. The immuneML ecosystem for machine learning analysis of adaptive immune receptor repertoires. Nat. Mach. Intell. 3, 936–944 (2021).
Rajeh, A. et al. iCAT: diagnostic assessment tool of immunological history using high-throughput T-cell receptor sequencing. F1000Res 10, 65 (2021).
Schwitalle, Y. et al. Immune response against frameshift-induced neopeptides in HNPCC patients and healthy HNPCC mutation carriers. Gastroenterology 134, 988–997 (2008).
von Knebel Doeberitz, M. & Kloor, M. Towards a vaccine to prevent cancer in Lynch syndrome patients. Fam. Cancer 12, 307–312 (2013).
Kloor, M. et al. A frameshift peptide neoantigen-based vaccine for mismatch repair-deficient cancers: a phase I/IIa clinical trial. Clin. Cancer Res 26, 4503–4510 (2020).
Bowen, C. M. et al. Naproxen chemoprevention induces proliferation of cytotoxic lymphocytes in Lynch Syndrome colorectal mucosa. Front Immunol. 14, 1162669 (2023).
ImmunoMind Team. Immunarch: an R package for painless bioinformatics analysis of T-cell and B-cell immune repertoires. Zenodo https://doi.org/10.5281/zenodo.3367200 (2019).
Chiffelle, J. et al. T-cell repertoire analysis and metrics of diversity and clonality. Curr. Opin. Biotechnol. 65, 284–295 (2020).
Acknowledgements
We thank the patients and their families for their participation. We thank the staff of the CPRIT Single Cell Genomics Core at MD Anderson Cancer Center for the assistance with single-cell TCR and RNA sequencing and the staff of the Advanced Technology Genomics Core and Cancer Genomics Laboratory at MD Anderson Cancer Center for their initial assistance in sequencing libraries generated from tissues and blood samples. We would like to acknowledge the support from the University of Kansas Medical Center’s Biospecimen Repository Core Facility staff including Maura Kluthe, Alex Webster, Eric Johnson, and Lauren DiMartino for identifying, collecting, and processing human specimens; the support from Araceli Garcia Gonzalez, Jacklyn Thompson and Pragya Mishra for their assistance in identifying, consenting, collecting and processing human specimens from the MD Anderson cohort. The authors are grateful to Karen Colbert for her editorial assistance in the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
N.D. conducted the bioinformatic pipelines for TCR classifier development. F.D. performed single-cell sequencing and immunological assays. A.M.B. initiated the study, performed bulk TCR sequencing and bioinformatic analyses, and drafted the initial manuscript. N.D., F.D., and K.M.S. contributed to data analysis, manuscript review, editing, and finalization. M.W.T. performed pathological interpretation of tissue section specimens. L.R.U., S.T., L.R., P.M.L., Y.N.Y., and E.V. provided clinical samples and associated clinical information. F.M., M.P., and G.C. critically reviewed the manuscript and provided clinical samples from Spain. A.B. provided clinical samples from Kansas. S.K., P.S., G.A.L., and A.R. provided critical input on study design. E.V. led conceptualization, supervised the study, secured funding, and contributed to data analysis, manuscript review, and editing. E.V. had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Competing interests
Eduardo Vilar (EV) had a consulting or advisory role with Janssen Research and Development, Recursion Pharma, Nouscom, Abbvie, Moderna, Permanence Bio and Parabilis. EV has received research support from Janssen Research and Development and Nouscom. EV has equity in Permanence Bio. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kazuhiko Hashimoto, Santiago Carmona and the other anonymous reviewers for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Deng, N., Duzagac, F., Bolivar, A.M. et al. Genomic analysis of T Cell receptors reveals lynch syndrome specific immune signatures. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71243-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71243-z