Abstract
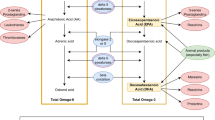
Genome-wide association studies (GWAS) have successfully identified genetic loci associated with major depressive disorder (MDD), yet the complex gene networks underpinning this polygenic risk remain largely uncharacterised. Here, we elucidate the neurobiological mechanisms of MDD by analyzing co-expression networks of 94 risk genes in the human prefrontal cortex. By linking these networks to individual symptoms, we identify the FADS1 (fatty acid desaturase 1) network as a central integrator across symptom clusters. We find that the FADS1 network functions primarily in astrocytes to regulate fatty acid metabolism and influence oligodendrocyte-related cell states. Furthermore, we identify FGF2 as a synaptic effector of this pathway and highlight PPARα (peroxisome proliferator-activated receptor alpha) as a putative therapeutic target. These results establish astrocyte fatty acid metabolism as a critical mechanistic contributor to MDD and a promising avenue for treatment.
Similar content being viewed by others
Data availability
Co-expression networks are available from https://zenodo.org/records/10181579. GWAS summary statistics are available from the PGC consortium website: https://pgc.unc.edu/for-researchers/download-results/. GTEx data are available through the online portal or through dbGAP for protected datasets. See https://www.gtexportal.org/home/datasets for more information. The MDD summary statistics were downloaded from iPSYCH as described in the original publication15 (https://ipsych.dk/en/research/downloads/). Hi-C data from the DLPFC16 were obtained from https://zenodo.org/record/6382668#.Y9PSG3bMJPY. Single nucleus RNA seq data from Nagy et al. and Maitra et al. (accession numbers GSE144136 and GSE213982 respectively). GnomAD data were downloaded from https://gnomad.broadinstitute.org/downloads.
Code availability
Code used for co-expression was adapted from Kim et al.9. All code used in this study is available at https://zenodo.org/records/10181579.
References
Ferrari, A. Global, regional, and national burden of 12 mental disorders in 204 countries and territories, 1990–2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet Psychiatry 9, 137–150 (2022).
Abbafati, C. et al. Global burden of 369 diseases and injuries in 204 countries and territories, 1990–2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet 396, 1204–1222 (2020).
Rush, A. J. et al. Acute and longer-term outcomes in depressed outpatients requiring one or several treatment steps: a STAR*D report. Am. J. Psychiatry 163, 1905–1917 (2006).
Levey, D. F. et al. Bi-ancestral depression GWAS in the Million Veteran Program and meta-analysis in >1.2 million individuals highlight new therapeutic directions. Nat. Neurosci. 24, 954–963 (2021).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: from polygenic to omnigenic. Cell 169, 1177 (2017).
Sinnott-Armstrong, N., Naqvi, S., Rivas, M. & Pritchard, J. K. GWAS of three molecular traits highlights core genes and pathways alongside a highly polygenic background. Elife 10, 1–35 (2021).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 1–13 (2008).
Willsey, A. J. et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 155, 997 (2013).
Kim, M. et al. Brain gene co-expression networks link complement signaling with convergent synaptic pathology in schizophrenia. Nat. Neurosci. 24, 799–809 (2021).
Liao, C. et al. Convergent coexpression of autism-associated genes suggests some novel risk genes may not be detectable in large-scale genetic studies. Cell Genom. 3, 100277 (2023).
Cole, E. J. et al. Stanford neuromodulation therapy (SNT): a double-blind randomized controlled. Trial 179, 132–141 (2021).
Otte, C. et al. Major depressive disorder. Nat. Rev. Dis. Prim. 2, 1–20 (2016).
Nagy, C. et al. Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat. Neurosci. 23, 771–781 (2020).
Maitra, M. et al. Cell type specific transcriptomic differences in depression show similar patterns between males and females but implicate distinct cell types and genes. Nat. Commun. 14, 1–18 (2023).
Als, T. D. et al. Depression pathophysiology, risk prediction of recurrence and comorbid psychiatric disorders using genome-wide analyses. Nat. Med. 29, 1832–1844 (2023).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, 2018 (1979).
Sey, N. Y. A. et al. A computational tool (H-MAGMA) for improved prediction of brain-disorder risk genes by incorporating brain chromatin interaction profiles. Nat. Neurosci. 23, 583–593 (2020).
Pergola, G. et al. Prefrontal coexpression of schizophrenia risk genes is associated with treatment response in patients. Biol. Psychiatry 86, 45–55 (2019).
Pergola, G. et al. Consensus molecular environment of schizophrenia risk genes in coexpression networks shifting across age and brain regions. Sci. Adv. 9, eade2812 (2023).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Radulescu, E. et al. Identification and prioritization of gene sets associated with schizophrenia risk by co-expression network analysis in human brain. Mol. Psychiatry 25, 791–804 (2020).
Walker, R. L. et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell 179, 750–771.e22 (2019).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Punzi, G. et al. Genetics and brain transcriptomics of completed suicide. Am. J. Psychiatry. 179, 226–241 (2022).
Hartl, C. L. et al. Coexpression network architecture reveals the brain-wide and multiregional basis of disease susceptibility. Nat. Neurosci. 24, 1313–1323 (2021).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Singh, T. et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature 604, 509–516 (2022).
Scarpa, J. R. et al. Shared transcriptional signatures in major depressive disorder and mouse chronic stress models. Biol. Psychiatry 88, 159–168 (2020).
Mullins, N. et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat. Genet. 53, 817–829 (2021).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Nievergelt, C. M. et al. International meta-analysis of PTSD genome-wide association studies identifies sex- and ancestry-specific genetic risk loci. Nat. Commun. 10, 1–16 (2019).
Cross-Disorder Group of the Psychiatric Genomics Consortium et al. Genomic relationships, novel loci, and pleiotropic mechanisms across eight psychiatric disorders. Cell 179, 1469–1482.e11 (2019).
Watson, H. J. et al. Genome-wide association study identifies eight risk loci and implicates metabo-psychiatric origins for anorexia nervosa. Nat. Genet. 51, 1207–1214 (2019).
Otowa, T. et al. Meta-analysis of genome-wide association studies of anxiety disorders. Mol. Psychiatry 21, 1391–1399 (2016).
Labonté, B. et al. Sex-specific transcriptional signatures in human depression. Nat. Med. 23, 1102 (2017).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49, D605–D612 (2021).
Akil, H. et al. Treatment resistant depression: a multi-scale, systems biology approach. Neurosci. Biobehav. Rev. 84, 272–288 (2018).
Bagot, R. C. C. et al. Circuit-wide transcriptional profiling reveals brain region-specific gene networks regulating depression susceptibility. Neuron 90, 969–983 (2016).
Blaveri, E. et al. Expression profiling of a genetic animal model of depression reveals novel molecular pathways underlying depressive-like behaviours. PLoS ONE 5, e12596 (2010).
Chang, L. C. et al. A Cconserved BDNF, Glutamate- and GABA-enriched gene module related to human depression identified by coexpression meta-analysis and DNA variant genome-wide association studies. PLoS ONE 9, e90980 (2014).
Camargo, N. et al. Oligodendroglial myelination requires astrocyte-derived lipids. PLoS Biol. 15, e1002605 (2017).
Sacchet, M. D. & Gotlib, I. H. Myelination of the brain in major depressive disorder: an in vivo quantitative magnetic resonance imaging study. Sci. Rep. 7, 1–14 (2017).
Edinoff, A. N. et al. Brexanolone, a GABAA modulator, in the treatment of postpartum depression in adults: a comprehensive review. Front. Psychiatry 12, 1331 (2021).
Dion-Albert, L. et al. Vascular and blood-brain barrier-related changes underlie stress responses and resilience in female mice and depression in human tissue. Nat. Commun. 13, 1–18 (2022).
Menard, C. et al. Social stress induces neurovascular pathology promoting depression. Nat. Neurosci. 20, 1752 (2017).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2021).
Soto, J. S. et al. Astrocyte–neuron subproteomes and obsessive–compulsive disorder mechanisms. Nature 616, 764–773 (2023).
Koopmans, F. et al. SynGO: an evidence-based, expert-curated knowledge base for the synapse. Neuron 103, 217–234.e4 (2019).
Wang, L. et al. A cross-species proteomic map reveals neoteny of human synapse development. Nature 622, 112–119 (2023).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2019).
Choi, G. E. et al. Prenatal glucocorticoid exposure selectively impairs neuroligin 1-dependent neurogenesis by suppressing astrocytic FGF2–neuronal FGFR1 axis. Cell. Mol. Life Sci. 79, 1–23 (2022).
Kajitani, N. et al. Antidepressant acts on astrocytes leading to an increase in the expression of neurotrophic/growth factors: differential regulation of FGF-2 by Noradrenaline. PLoS ONE 7, e51197 (2012).
Kirby, E. D. et al. Acute stress enhances adult rat hippocampal neurogenesis and activation of newborn neurons via secreted astrocytic FGF2. Elife 2, e00362 (2013).
Turner, C. A., Watson, S. J. & Akil, H. The fibroblast growth factor family: neuromodulation of affective behavior. Neuron 76, 160 (2012).
Evans, S. J. et al. Dysregulation of the fibroblast growth factor system in major depression. Proc. Natl. Acad. Sci. USA 101, 15506–15511 (2004).
Deng, Z., Deng, S., Zhang, M. R. & Tang, M. M. Fibroblast growth factors in depression. Front. Pharm. 10, 60 (2019).
Svoboda, D. L., Saddler, T. & Auerbach, S. S. An overview of national toxicology program’s toxicogenomic applications: DrugMatrix and ToxFX. Chall Adv. Comput. Chem. Phys. 30, 141–157 (2019).
Pilarczyk, M. et al. Connecting omics signatures and revealing biological mechanisms with iLINCS. Nat. Commun. 13, 1–13 (2022).
Tyagi, S., Gupta, P., Saini, A., Kaushal, C. & Sharma, S. The peroxisome proliferator-activated receptor: a family of nuclear receptors role in various diseases. J. Adv. Pharm. Technol. Res. 2, 236 (2011).
Yao, X. et al. Integrative analysis of genome-wide association studies identifies novel loci associated with neuropsychiatric disorders. Transl. Psychiatry 11, 69 (2021).
Flint, J. The genetic basis of major depressive disorder. Mol. Psychiatry 28. https://doi.org/10.1038/s41380-023-01957-9. (2023).
Wainberg, M. et al. Opportunities and challenges for transcriptome-wide association studies. Nat. Genet. 51, 592–599 (2019).
Ernst, C. et al. Alternative Splicing, methylation state, and expression profile of tropomyosin-related kinase B in the frontal cortex of suicide completers. Arch. Gen. Psychiatry 66, 22–32 (2009).
Nagy, C. et al. Astrocytic abnormalities and global DNA methylation patterns in depression and suicide. Mol. Psychiatry 20, 320–328 (2014).
O’Leary, L. A. & Mechawar, N. Implication of cerebral astrocytes in major depression: a review of fine neuroanatomical evidence in humans. Glia 69, 2077–2099 (2021).
Codeluppi, S. A. et al. Chronic stress alters astrocyte morphology in mouse prefrontal cortex. Int. J. Neuropsychopharmacol. 24, 842–853 (2021).
Rajkowska, G., Hughes, J., Stockmeier, C. A., Javier Miguel-Hidalgo, J. & Maciag, D. Coverage of blood vessels by astrocytic endfeet is reduced in major depressive disorder. Biol. Psychiatry 73, 613–621 (2013).
Ernst, C. et al. Dysfunction of astrocyte connexins 30 and 43 in dorsal lateral prefrontal cortex of suicide completers. Biol. Psychiatry 70, 312–319 (2011).
Hamilton, P. J. et al. Chronic stress and antidepressant treatment alter purine metabolism and beta oxidation within mouse brain and serum. Sci. Rep. 10, 1–14 (2020).
Bazinet, R. P. & Layé, S. Polyunsaturated fatty acids and their metabolites in brain function and disease. Nat. Rev. Neurosci. 15, 771–785 (2014).
Salvati, S. et al. Eicosapentaenoic acid stimulates the expression of myelin proteins in rat brain. J. Neurosci. Res. 86, 776–784 (2008).
Sublette, M. E., Ellis, S. P., Geant, A. L. & Mann, J. J. Meta-analysis of the effects of eicosapentaenoic acid (EPA) in clinical trials in depression. J. Clin. Psychiatry 72, 1577–1584 (2011).
Yu, H. L. et al. N-palmitoylethanolamide, an endocannabinoid, exhibits antidepressant effects in the forced swim test and the tail suspension test in mice. Pharmacol. Rep. 63, 834–839 (2011).
Locci, A. & Pinna, G. Stimulation of peroxisome proliferator-activated receptor-α by N-palmitoylethanolamine engages allopregnanolone biosynthesis to modulate emotional behavior. Biol. Psychiatry 85, 1036–1045 (2019).
Umathe, S. N., Manna, S. S. S. & Jain, N. S. Involvement of endocannabinoids in antidepressant and anti-compulsive effect of fluoxetine in mice. Behav. Brain Res. 223, 125–134 (2011).
Ghazizadeh-Hashemi, M. et al. Palmitoylethanolamide as adjunctive therapy in major depressive disorder: a double-blind, randomized and placebo-controlled trial. J. Affect Disord. 232, 127–133 (2018).
Jiang, B. et al. Antidepressant-like effects of fenofibrate in mice via the hippocampal brain-derived neurotrophic factor signalling pathway. Br. J. Pharm. 174, 177–194 (2017).
Jiang, B., Huang, C., Zhu, Q., Tong, L. J. & Zhang, W. WY14643 produces anti-depressant-like effects in mice via the BDNF signaling pathway. Psychopharmacology 232, 1629–1642 (2015).
Luo, R. et al. Activation of PPARA-mediated autophagy reduces Alzheimer disease-like pathology and cognitive decline in a murine model. Autophagy 16, 52–69 (2020).
Patel, D. et al. Aspirin binds to PPARα to stimulate hippocampal plasticity and protect memory. Proc. Natl. Acad. Sci. USA 115, E7408–E7417 (2018).
Roy, A. et al. Regulation of Cyclic AMP response element binding and hippocampal plasticity-related genes by peroxisome proliferator-activated receptor α. Cell Rep. 4, 724–737 (2013).
Esmaeili, M. A. et al. Preferential PPAR-α activation reduces neuroinflammation, and blocks neurodegeneration in vivo. Hum. Mol. Genet. 25, 317–327 (2016).
Wada, Y. et al. Peroxisome proliferator-activated receptor α as a novel therapeutic target for schizophrenia. EBioMedicine 62, 103130 (2020).
D’Agostino, G. et al. Peroxisome proliferator-activated receptor alpha plays a crucial role in behavioral repetition and cognitive flexibility in mice. Mol. Metab. 4, 528–536 (2015).
Roy, A. et al. Identification and characterization of PPARα ligands in the hippocampus. Nat. Chem. Biol. 12, 1075–1083 (2016).
Staels, B. et al. Mechanism of action of fibrates on lipid and lipoprotein metabolism. Circulation 98, 2088–2093 (1998).
Miao, G. et al. Plasma lipidomic profile of depressive symptoms: a longitudinal study in a large sample of community-dwelling American Indians in the strong heart study. Mol. Psychiatry 28 https://doi.org/10.1038/s41380-023-01948-w. (2023).
Van Reedt Dortland, A. K. B. et al. Associations between serum lipids and major depressive disorder: results from the netherlands study of depression and anxiety (NESDA). J. Clin. Psychiatry 71, 1549 (2009).
Tkachev, A. et al. Lipid alteration signature in the blood plasma of individuals with schizophrenia, depression, and bipolar disorder. JAMA Psychiatry https://doi.org/10.1001/JAMAPSYCHIATRY.2022.4350 (2023).
Chen, Y. et al. Genomic atlas of the plasma metabolome prioritizes metabolites implicated in human diseases. Nat. Genet. 55, 44–53 (2023).
Davyson, E. et al. Metabolomic investigation of major depressive disorder identifies a potentially causal association with polyunsaturated fatty acids. Biol. Psychiatry 94, 630–639 (2023).
Lalovic, A., Klempan, T., Sequeira, A., Luheshi, G. & Turecki, G. Altered expression of lipid metabolism and immune response genes in the frontal cortex of suicide completers. J. Affect Disord. 120, 24–31 (2010).
Lanekoff, I., Sharma, V. V. & Marques, C. Single-cell metabolomics: where are we and where are we going? Curr. Opin. Biotechnol. 75, 102693 (2022).
Borcuk, C. et al. Network-wide risk convergence in gene co-expression identifies reproducible genetic hubs of schizophrenia risk. Neuron 112, 3551–3566.e6 (2024).
Pergola, G. et al. DRD2 co-expression network and a related polygenic index predict imaging, behavioral and clinical phenotypes linked to schizophrenia. Transl. Psychiatry 7, e1006–e1006 (2017).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Aguet, F. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882 (2012).
Shen, L. GeneOverlap: R package for testing and visualizing gene list overlaps. https://github.com/shenlab-sinai/geneoverlap.
Korotkevich, G., Sukhov, V. & Sergushichev, A. Fast gene set enrichment analysis. Preprint at https://doi.org/10.1101/060012. (2019).
Durinck, S. et al. BioMart and bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
pheatmap: Pretty Heatmaps version 1.0.12 from CRAN. https://rdrr.io/cran/pheatmap/.
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498 (2003).
igraph for R: R interface of the igraph library for graph theory and network analysis. https://doi.org/10.5281/ZENODO.8240644.
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, e29 (2021).
Squair, J. W. et al. Confronting false discoveries in single-cell differential expression. Nat. Commun. 12, 1–15 (2021).
Mountjoy, E. et al. An open approach to systematically prioritize causal variants and genes at all published human GWAS trait-associated loci. Nat. Genet. 53, 1527–1533 (2021).
R Core Team. R: The R Project for Statistical Computing. Preprint at (2021).
RStudio Team. RStudio: Integrated Development for R. Preprint at (2020).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis. (Springer-Verlag New York, 2016).
Acknowledgements
This work was funded through a Hope for Depression Research Foundation grant to MJM. The Genotype-Tissue Expression (GTEx) Project was supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. The data used for the analyses described in this manuscript were obtained from: the GTEx Portal or dbGaP accession number phs000424.v9.p2. Schematics were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
E.F. conceived the study, with input from M.J.M., E.F., N.O.T., and I.P. analysed the data. E.F. interpreted the data and wrote the manuscript with input from E.J.N, G.T., C.N., and M.J.M. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fitzgerald, E., O’Toole, N., Pokhvisneva, I. et al. Astrocyte fatty acid metabolism as a driver of risk for major depressive disorder. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71542-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71542-5