Abstract
During oxygenic photosynthesis, photosystem II (PSII) uses light energy for oxidizing water and reducing plastoquinone. It is susceptible to photodamage, and the damaged PSII is repaired through a sophisticated biological process assisted by numerous auxiliary proteins. Here we report the cryogenic electron microscopy structures of four PSII-repair complexes from Chlamydomonas reinhardtii associated with the Thylakoid Enriched Fraction 30 (TEF30, an orthologue of plant MET1) protein—namely, a TEF30–PSII core monomer (TEF30-C), two types of TEF30–PSII core dimers (types I and II, TEF302-C2-I and TEF302-C2-II) and a TEF30-C2S-type PSII–LHCII supercomplex (TEF30-C2S; S, strongly associated light-harvesting complex II trimer). TEF30 mediates the assembly of CP43 with the RC47 module by clamping on the stromal surfaces and prevents the premature association of peripheral antennae with PSII-C. In the transition from TEF302-C2-I to TEF302-C2-II, TEF30-C2S and mature C2S2, one PSII core slides along the dimerization interface against the adjacent one by 22–35 Å, generating a zigzagged surface for accommodating the peripheral antennae. These results suggest that the PSII repair process undergoes multiple TEF30-mediated intermediate states to form intact PSII–LHCII supercomplexes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The cryo-EM maps of the C. reinhardtii TEF30–PSII complexes and the corresponding atomic coordinates have been deposited in the Electron Microscopy Data Bank (EMDB) and the PDB. The accession codes for TEF30-C, TEF302-C2-I, TEF302-C2-II and TEF30-C2S in the EMDB are EMD-60605, EMD-60694, EMD-60748 and EMD-60830, respectively. The accession codes for TEF30-C, TEF302-C2-I, TEF302-C2-II and TEF30-C2S in the PDB are 9IIU, 9IMN, 9IOX and 9IS4, respectively. The previously published PSII cryo-EM map and atomic coordinates used in this study are available under PDB accession code 6KAC and EMDB accession code EMD-9955. Source data are provided with this paper.
References
Barber, J. Photosystem II: an enzyme of global significance. Biochem. Soc. Trans. 34, 619–631 (2006).
Vinyard, D. J., Ananyev, G. M. & Charles Dismukes, G. Photosystem II: the reaction center of oxygenic photosynthesis. Annu. Rev. Biochem. 82, 577–606 (2013).
Barber, J. Photosystem II: the water splitting enzyme of photosynthesis and the origin of oxygen in our atmosphere. Q. Rev. Biophys. 49, e14 (2016).
Umena, Y., Kawakami, K., Shen, J. & Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473, 55–60 (2011).
Caffarri, S., Kouril, R., Kereïche, S., Boekema, E. J. & Croce, R. Functional architecture of higher plant photosystem II supercomplexes. EMBO J. 28, 3052–3063 (2009).
Sheng, X. et al. Structural insight into light harvesting for photosystem II in green algae. Nat. Plants 5, 1320–1330 (2019).
Shen, L. et al. Structure of a C2S2M2N2-type PSII–LHCII supercomplex from the green alga Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 116, 21246–21255 (2019).
Wei, X. et al. Structure of spinach photosystem II–LHCII supercomplex at 3.2 Å resolution. Nature 534, 69–74 (2016).
Su, X. et al. Structure and assembly mechanism of plant C2S2M2-type PSII–LHCII supercomplex. Science 357, 815–820 (2017).
Shen, J. The structure of photosystem II and the mechanism of water oxidation in photosynthesis. Annu. Rev. Plant Biol. 66, 23–48 (2015).
Hu, S., Ding, Y. & Zhu, C. Sensitivity and responses of chloroplasts to heat stress in plants. Front. Plant Sci. 11, 375 (2020).
Landi, M. & Guidi, L. Effects of abiotic stress on photosystem II proteins. Photosynthetica 61, 148–156 (2023).
Murata, N., Allakhverdiev, S. I. & Nishiyama, Y. The mechanism of photoinhibition in vivo: re-evaluation of the roles of catalase, alpha-tocopherol, non-photochemical quenching, and electron transport. Biochim. Biophys. Acta 1817, 1127–1133 (2012).
Erickson, E., Wakao, S. & Niyogi, K. K. Light stress and photoprotection in Chlamydomonas reinhardtii. Plant J. 82, 449–465 (2015).
Murata, N., Takahashi, S., Nishiyama, Y. & Allakhverdiev, S. I. Photoinhibition of photosystem II under environmental stress. Biochim. Biophys. Acta 1767, 414–421 (2007).
Tyystjarvi, E. Photoinhibition of photosystem II and photodamage of the oxygen evolving manganese cluster. Coord. Chem. Rev. 252, 361–376 (2008).
Vass, I. Molecular mechanisms of photodamage in the photosystem II complex. Biochim. Biophys. Acta 1817, 209–217 (2012).
Vavilin, D., Brune, D. C. & Vermaas, W. 15N-labeling to determine chlorophyll synthesis and degradation in Synechocystis sp. PCC 6803 strains lacking one or both photosystems. Biochim. Biophys. Acta 1708, 91–101 (2005).
Yao, D. C. I., Brune, D. C. & Vermaas, W. F. J. Lifetimes of photosystem I and II proteins in the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett. 586, 169–173 (2011).
Nixon, P. J., Michoux, F., Yu, J., Boehm, M. & Komenda, J. Recent advances in understanding the assembly and repair of photosystem II. Ann. Bot. 106, 1–16 (2010).
Nickelsen, J. & Rengstl, B. Photosystem II assembly: from cyanobacteria to plants. Annu. Rev. Plant Biol. 64, 609–635 (2013).
Theis, J. & Schroda, M. Revisiting the photosystem II repair cycle. Plant Signal. Behav. 11, e1218587 (2016).
Nath, K. et al. Towards a critical understanding of the photosystem II repair mechanism and its regulation during stress conditions. FEBS Lett. 587, 3372–3381 (2013).
Bonardi, V. et al. Photosystem II core phosphorylation and photosynthetic acclimation require two different protein kinases. Nature 437, 1179–1182 (2005).
Vainonen, J. P., Hansson, M. & Vener, A. V. STN8 protein kinase in Arabidopsis thaliana is specific in phosphorylation of photosystem II core proteins. J. Biol. Chem. 280, 33679–33686 (2005).
Aro, E. M. et al. Dynamics of photosystem II: a proteomic approach to thylakoid protein complexes. J. Exp. Bot. 56, 347–356 (2005).
Samol, I. et al. Identification of a photosystem II phosphatase involved in light acclimation in Arabidopsis. Plant Cell 24, 2596–2609 (2012).
Lindahl, M. et al. The thylakoid FtsH protease plays a role in the light-induced turnover of the photosystem II D1 protein. Plant Cell 12, 419–431 (2000).
Nixon, P. J., Barker, M., Boehm, M., de Vries, R. & Komenda, J. FtsH-mediated repair of the photosystem II complex in response to light stress. J. Exp. Bot. 56, 357–363 (2005).
Kapri-Pardes, E., Naveh, L. & Adam, Z. The thylakoid lumen protease Deg1 is involved in the repair of photosystem II from photoinhibition in Arabidopsis. Plant Cell 19, 1039–1047 (2007).
Sun, X., Wang, L. & Zhang, L. Involvement of DEG5 and DEG8 proteases in the turnover of the photosystem II reaction center D1 protein under heat stress in Arabidopsis thaliana. Chin. Sci. Bull. 52, 1742–1745 (2007).
Zhang, L., Paakkarinen, V., van Wijk, K. J. & Aro, E. M. Co-translational assembly of the D1 protein into photosystem II. J. Biol. Chem. 274, 16062–16067 (1999).
Nilsson, R., Brunner, J., Hoffman, N. E. & van Wijk, K. J. Interactions of ribosome nascent chain complexes of the chloroplast-encoded D1 thylakoid membrane protein with cpSRP54. EMBO J. 18, 733–742 (1999).
Zhang, L., Paakkarinen, V., Suorsa, M. & Aro, E. M. A SecY homologue is involved in chloroplast-encoded D1 protein biogenesis. J. Biol. Chem. 276, 37809–37814 (2001).
Klostermann, E., Droste Gen Helling, I., Carde, J. P. & Schünemann, D. The thylakoid membrane protein ALB3 associates with the cpSecY-translocase in Arabidopsis thaliana. Biochem. J. 368, 777–781 (2002).
Ossenbühl, F. et al. Efficient assembly of photosystem II in Chlamydomonas reinhardtii requires Alb3.1p, a homolog of Arabidopsis ALBINO3. Plant Cell 16, 1790–1800 (2004).
Diner, B. A., Ries, D. F., Cohen, B. N. & Metz, J. G. COOH-terminal processing of polypeptide D1 of the photosystem II reaction center of Scenedesmus obliquus is necessary for the assembly of the oxygen-evolving complex. J. Biol. Chem. 263, 8972–8980 (1988).
Anbudurai, P. R., Mor, T. S., Ohad, I., Shestakov, S. V. & Pakrasi, H. B. The ctpA gene encodes the C-terminal processing protease for the D1 protein of the photosystem II reaction center complex. Proc. Natl Acad. Sci. USA 91, 8082–8086 (1994).
Mulo, P., Sakurai, I. & Aro, E. M. Strategies for psbA gene expression in cyanobacteria, green algae and higher plants: from transcription to PSII repair. Biochim. Biophys. Acta 1817, 247–257 (2012).
Nowaczyk, M. M. et al. Psb27, a cyanobacterial lipoprotein, is involved in the repair cycle of photosystem II. Plant Cell 18, 3121–3131 (2006).
Liu, H., Huang, R. Y. C., Chen, J., Gross, M. L. & Pakrasi, H. B. Psb27, a transiently associated protein, binds to the chlorophyll binding protein CP43 in photosystem II assembly intermediates. Proc. Natl Acad. Sci. USA 108, 18536–18541 (2011).
Komenda, J. et al. The Psb27 assembly factor binds to the CP43 complex of photosystem II in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 158, 476–486 (2012).
Grasse, N. et al. Role of novel dimeric photosystem II (PSII)–Psb27 protein complex in PSII repair. J. Biol. Chem. 286, 29548–29555 (2011).
Liu, H., Roose, J. L., Cameron, J. C. & Pakrasi, H. B. A genetically tagged Psb27 protein allows purification of two consecutive photosystem II (PSII) assembly intermediates in Synechocystis 6803, a cyanobacterium. J. Biol. Chem. 286, 24865–24871 (2011).
Chen, H. et al. A Psb27 homologue in Arabidopsis thaliana is required for efficient repair of photodamaged photosystem II. Plant Mol. Biol. 61, 567–575 (2006).
Huang, G. et al. Structural insights into a dimeric Psb27–photosystem II complex from a cyanobacterium Thermosynechococcus vulcanus. Proc. Natl Acad. Sci. USA 118, e2018053118 (2021).
Dobáková, M., Sobotka, R., Tichý, M. & Komenda, J. Psb28 protein is involved in the biogenesis of the photosystem II inner antenna CP47 (PsbB) in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 149, 1076–1086 (2009).
Mabbitt, P. D., Wilbanks, S. M. & Eaton-Rye, J. J. Structure and function of the hydrophilic photosystem II assembly proteins: Psb27, Psb28 and Ycf48. Plant Physiol. Biochem. 81, 96–107 (2014).
Jung, K.-H. et al. Identification and functional analysis of light-responsive unique genes and gene family members in rice. PLoS Genet. 4, e1000164 (2008).
Zabret, J. et al. Structural insights into photosystem II assembly. Nat. Plants 7, 524–538 (2021).
Xiao, Y. et al. Structural insights into cyanobacterial photosystem II intermediates associated with Psb28 and Tsl0063. Nat. Plants 7, 1132–1142 (2021).
Fadeeva, M., Klaiman, D., Kandiah, E. & Nelson, N. Structure of native photosystem II assembly intermediate from Chlamydomonas reinhardtii. Front. Plant Sci. 14, 1334608 (2024).
Li, A. et al. Structural basis for an early stage of the photosystem II repair cycle in Chlamydomonas reinhardtii. Nat. Commun. 15, 5211 (2024).
Muranaka, L. S. et al. TEF30 interacts with photosystem II monomers and is involved in the repair of photodamaged photosystem II in Chlamydomonas reinhardtii. Plant Physiol. 170, 821–840 (2016).
Bhuiyan, N. H., Friso, G., Poliakov, A., Ponnala, L. & van Wijk, K. J. MET1 is a thylakoid-associated TPR protein involved in photosystem II supercomplex formation and repair in Arabidopsis. Plant Cell 27, 262–285 (2015).
Garcia-Cerdan, J. G. et al. The PsbW protein stabilizes the supramolecular organization of photosystem II in higher plants. Plant J. 65, 368–381 (2011).
Jamali, K. et al. Automated model building and protein identification in cryo-EM maps. Nature 628, 450–457 (2024).
Krieger-Liszkay, A., Fufezan, C. & Trebst, A. Singlet oxygen production in photosystem II and related protection mechanism. Photosynth. Res. 98, 551–564 (2008).
Sirpio, S. et al. TLP18.3, a novel thylakoid lumen protein regulating photosystem II repair cycle. Biochem. J. 406, 415–425 (2007).
Lima, A. et al. A redox-active FKBP-type immunophilin functions in accumulation of the photosystem II supercomplex in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 103, 12631–12636 (2006).
Fristedt, R. et al. PHOTOSYSTEM II PROTEIN33, a protein conserved in the plastid lineage, is associated with the chloroplast thylakoid membrane and provides stability to photosystem II supercomplexes in Arabidopsis. Plant Physiol. 167, 481–492 (2015).
Shi, L., Lorkovic, Z. J., Oelmuller, R. & Schroder, W. P. The low molecular mass PsbW protein is involved in the stabilization of the dimeric photosystem II complex in Arabidopsis thaliana. J. Biol. Chem. 275, 37945–37950 (2000).
Zheleva, D., Sharma, J., Panico, M., Morris, H. R. & Barber, J. Isolation and characterization of monomeric and dimeric CP47-reaction center Photosystem II complexes. J. Biol. Chem. 273, 16122–16127 (1998).
Müller, B. & Eichacker, L. A. Assembly of the D1 precursor in monomeric photosystem II reaction center precomplexes precedes chlorophyll a-triggered accumulation of reaction center II in barley etioplasts. Plant Cell 11, 2365–2377 (1999).
Wang, L. et al. A chloroplast protein atlas reveals punctate structures and spatial organization of biosynthetic pathways. Cell 186, 3499–3518.e14 (2023).
Cariti, F. et al. Regulation of light harvesting in Chlamydomonas reinhardtii two protein phosphatases are involved in state transitions. Plant Physiol. 183, 1749–1764 (2020).
Kropat, J. et al. A revised mineral nutrient supplement increases biomass and growth rate in Chlamydomonas reinhardtii. Plant J. 66, 770–780 (2011).
Chua, N. H. & Bennoun, P. Thylakoid membrane polypeptides of Chlamydomonas reinhardtii: wild-type and mutant strains deficient in photosystem II reaction center. Proc. Natl Acad. Sci. USA 72, 2175–2179 (1975).
Almagro Armenteros, J. J. et al. Detecting sequence signals in targeting peptides using deep learning. Life Sci. Alliance 2, e201900429 (2019).
Arnon, D. I. Copper enzymes in isolated chloroplasts: polyphenoloxidase in Beta vulgaris. Plant Physiol. 24, 1–15 (1949).
Ladig, R. et al. A high-definition native polyacrylamide gel electrophoresis system for the analysis of membrane complexes. Plant J. 67, 181–194 (2011).
Schägger, H. Tricine–SDS–PAGE. Nat. Protoc. 1, 16–22 (2006).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Mastronarde, D. N. SerialEM: a program for automated tilt series acquisition on Tecnai microscopes using prediction of specimen position. Microsc. Microanal. 9, 1182–1183 (2003).
Wu, C., Huang, X., Cheng, J., Zhu, D. & Zhang, X. High-quality, high-throughput cryo-electron microscopy data collection via beam tilt and astigmatism-free beam-image shift. J. Struct. Biol. 208, 107396 (2019).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Asarnow, D., Palovcak, E. & Cheng, Y. asarnow/pyem: UCSF pyem v0.5. Zenodo https://doi.org/10.5281/zenodo.3576630 (2019).
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Acknowledgements
We thank L.-H. Chen, X.-J. Huang, B.-L. Zhu, B.-X. Huang-Fu, T.-E. Wang, L.-L. Zhang and other staff members at the Center for Biological Imaging, Core Facilities for Protein Science at the Institute of Biophysics, Chinese Academy of Sciences (IBP, CAS), for their support in cryo-EM sample screening and data collection; X. Ding and L.-L. Niu at the CAS Research Platform for Protein Sciences for their support in mass spectrometry data collection and analysis; X.-B. Liang and X.-Y. Liu for technical support on biochemistry, cell culture and sample preparation; Y.-Y. Chen, Z. Yang and B.-X. Zhou (IBP, CAS) for technical support in the BLI assay; D. Duo and Y. Teng (IBP, CAS) for support in the confocal microscopy analysis; H. Liu for discussion on optimizing the conditions for the HDN-PAGE analysis; M. Li and X.-D. Su for their assistance with the oxygen-evolving assay; and Y. Yin from the Plant Science Facility of the Institute of Botany, CAS, for technical support with the flash-induced fluorescence decay and thermoluminescence assays. This work was funded by the National Natural Science Foundation of China (grant no. 31925024 to Z.L.), the Chinese Academy of Sciences Project for Young Scientists in Basic Research (grant no. YSBR-015 to Z.L.) and the Strategic Priority Research Program of CAS (grant no. XDB37020101 to Z.L.).
Author information
Authors and Affiliations
Contributions
Y.W. purified the recombinant TEF30 protein, isolated and obtained the TEF30–PSII complexes from C. reinhardtii, prepared the cryo-EM sample, solved the cryo-EM structures and conducted the biochemical analysis. Y.W., A.L. and Z.L. built and refined the atomic models and analysed the structures. Y.W., A.L. and Z.L. wrote the initial draft of the paper. C.W. constructed the plasmids for expression of recombinant TEF30 proteins (WT and mutants) in E. coli and participated in the initial steps of protein expression and purification, TEF30-C sample preparation, cryo-EM data collection, model building and the BLI assay. All authors were involved in discussing and editing the paper. Z.L. conceived and coordinated the research project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Peter Nixon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of the materials for purification of the TEF30-PSII complexes.
a, The recombinant CrTEF30 protein purified through the cobalt affinity chromatography was analyzed with SDS-PAGE. The expression cassette of recombinant CrTEF30 with 3×Flag and 6×His tags fused at the N-terminal region is shown at bottom. The experiment was repeated independently three times with similar results. b, Images showing the fluorescence of mVenus (colored in green) and chlorophyll (colored in red) from the TEF30-ox cells were recorded with a confocal microscope. The bright field and the merged images are displayed in the bottom row. The experiment was repeated independently three times with similar results. c, Immunodetection of endogenous native TEF30 in the SDG fractions. Thylakoids from the wild type (WT) cells grown under low-light (20 μmol photons m−2 s−1) and high-light (330 μmol photons m−2 s−1) conditions were solubilized with α-DDM and separated through the SDG ultracentrifugation. The SDG fractions were extracted and the endogenous native TEF30 proteins were analyzed using SDS-PAGE and western blot with anti-CrTEF30 antibodies. Note that all TEF30 is from the endogenous source of the WT cells and no exogenous TEF30 was added to the sample. LHCII-M, LHCII monomer; LHCII-T, LHCII trimer.
Extended Data Fig. 2 An overall scheme for the cryo-EM data processing of the TEF30-PSII complexes.
a, The data processing scheme of the TEF30-C complexes from the TEF30-ox cells. The 3D class chosen for further refinement in each step is circled with light pink rounded rectangles. Besides the major class of TEF30-C particles, the sample also contains minor classes of TEF302-C2 and TEF30-C2S complexes, with 2,863 and 2,670 particles respectively, resulting in low-resolution reconstructions. b, The data processing scheme of the TEF302-C2 and TEF30-C2S complexes. The selected 3D classes of PSII-C dimers were circled with blue rounded rectangles and the selected C2S classes were circled with light purple rounded rectangles. c, The gold standard Fourier shell correlation (GSFSC) curves of the cryo-EM maps of the four different TEF30-PSII complexes with a resolution-cutoff threshold at 0.143. The data was exported from CryoSPARC, and the plot was generated using OriginPro. d, Representative orientational distribution plots for the particle sets of the four different TEF30-PSII complexes. e, Estimation of the local resolution in the cryo-EM maps of the TEF30-PSII complexes.
Extended Data Fig. 3 The assembly patterns of the PSII-C dimers and the antenna-binding sites in TEF30 C2-I, TEF302-C2-II, TEF30-C2S and mature PSII-SC.
a, Comparison of the assembly patterns of the PSII-C dimers in TEF302-C2-I, TEF302-C2-II, TEF30-C2S and mature PSII-SC. The models of TEF302-C2-I, TEF302-C2-II, TEF30-C2S and mature dimer (PDB ID: 6KAC) are superposed on the left PSII-C monomer (colored in gray). The peripheral antennae of the TEF30-C2S and mature PSII-SC are omitted for clarity. The distances of the relative shifts of the right PSII-C monomers (colored in gold) along the dimerization interface are labeled nearby the blue arrows which indicate the directions of the translational shifts, and the reference lines were drawn tangent to a loop region (Thr188-Asn189) in the CP43 subunits. The PsbT and PsbM subunits are colored in blue and hot pink respectively. The arrangement patterns of PsbM (pink circles) and PsbT (blue circles) at the interfaces of the dimers are displayed at the bottom. The PsbM and PsbT from the same PSII-C are linked with dashed lines for clarity. b, The hypothetical binding positions of the peripheral antennae in TEF302-C2-I and TEF302-C2-II and the real binding sites of the peripheral antennae in TEF30-C2S and the mature C2S2. In TEF302-C2-I and TEF302-C2-II, the hypothetical positions of CP26, CP29 and S-LHCII trimer are depicted by the dashed lines. CP26 and LHCII trimer are colored in green, and CP29 is colored in purple.
Extended Data Fig. 4 The binding site, domain organization and secondary structures of TEF30.
a, Superposition of the TEF30-C (gray), TEF302-C2-I (sky blue), TEF302-C2-II (pink) and TEF30-C2S (light green) structures on the TEF30-C regions. The other parts of TEF302-C2-I, TEF302-C2-II and TEF30-C2S are omitted for clarity. The cartoon models except that of TEF30 are shown in the transparent mode. b, The cartoon model of TEF30 illustrating its secondary structures. The β-sheets and α-helices are indicated by the arrows and cylinders. c, The TEF30-C structure showing the position of TEF30 in the complex. Left: Stromal view; right: side view. TEF30 is shown as the cartoon and transparent surface models, PSII-C is presented as surface model.
Extended Data Fig. 5 The BLI results on the interactions between the four TEF30 mutants and PSII-C as well as the interactions between the two domains within TEF30.
a, Constructs of TEF30 mutants indicated with cartoon models. Red parts correspond to the deleted regions of the protein. b, The BLI analysis on the kinetics of the interactions between PSII-C and the TEF30 mutants loaded on sensors. The plots were generated using OriginPro. The colored curves are recorded data, and the black curves are the models fitted with the data. The PSII-C concentrations are labeled in the top right corner of the plots. The calculated dissociation constants and R-squared values are labeled below the curves. c, The response signals generated by the interactions of 125 nM PSII-C with the TEF30-immobilized sensors. TEF30-WT–LHCII was included as a negative control, providing the signals from the barely detectable interaction of TEF30-WT with 125 nM LHCII trimer. d, The polar interactions between the H0 helix of PDZ domain and the H3 and H5 helices of TPR domain in TEF30. The amino acid residues involved in the interactions were shown as sticks. The distances (Å) between the two adjacent groups are labeled nearby the dash lines. e, The cryo-EM density and structure model of TEF30 in TEF30-C. The PDZ domain (Pro78-Tyr154) and the TPR domain (Tyr181-Lys320) is colored in lime green and warm pink respectively.
Extended Data Fig. 6 Detailed features of the CP43 module and the D1 DE-loop region in TEF30-PSII complexes.
a, Superposition of the CP43 modules from TEF30-C (light pink) and PSII-SC (PDB ID: 6KAC, green). The local region with differences between the two models in the lumenal Loop E of CP43 is circled by dash lines. A lumenal loop region involved in binding PsbO and PsbQ (Gly313-Gly324) in PSII-SC is colored in yellow, but it is untraceable in TEF30-C. b, The clipped side views showing the D1 DE-loop region of TEF30-C and mature PSII-SC (EMDB ID: 9955, PDB ID: 6KAC) on the left and right sides, respectively. The cryo-EM density and model of TEF30-C within a loop region between the distal traceable residues of the D1 DE-loop (Arg225 and Ala251) are enclosed in a green rectangle and magnified in the top small panel. The cryo-EM density and model of PSII-SC around the intact D1 DE-loop are enclosed in a blue rectangle and magnified in the bottom small panel. Note that the green segment (Glu226-Ala250) of the DE-loop is untraceable in TEF30-C. c, Immunodetection of the D1 degradation products in TEF30-PSII complexes. The TEF30-PSII complexes purified from the high-light adapted TEF30-ox thylakoids with the anti-Flag beads and the PSII-C isolated from the high-light adapted WT thylakoids (as a positive control) were analyzed by SDS-PAGE followed by the western blot using an antibody against a D1 DE-loop fragment (Asn234-Glu242). The experiment was repeated independently three times with similar results.
Extended Data Fig. 7 PsbO and the manganese cluster sites in the TEF30-C2S complex.
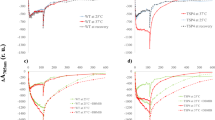
a, Superposition of the TEF30-associated PSII-C (pale turquoise) with the TEF30-free PSII-C (pink) from TEF30-C2S. The residues involved in coordinating the Mn cluster are shown as stick models. The carboxyl-termini of D1 in the two PSII-C monomers are indicated by arrows. b, Superposition of the vacant manganese cluster-binding site in the TEF30-associated PSII-C (pale turquoise) with the corresponding region from the mature C2S2-type PSII-SC (green, PDB ID: 6KAC). The cryo-EM density of TEF30-C2S is shown as transparent blue surface. The residues involved in binding the Mn cluster are shown as stick models. The Mn cluster from the mature PSII-SC is shown as spheres (Mn: purple, O: red, Ca: lime). c, Superposition of the manganese cluster-binding site of the TEF30-free PSII-C in TEF30-C2S (pink) and the one in PSII-SC (green, PDB ID: 6KAC). The cryo-EM density of TEF30-C2S is shown as transparent pink surface. The Mn cluster model from the mature PSII-SC is shown as spheres. d, Superposition of PsbO from TEF30-C2S (white) with the one in PSII-SC (green, PDB ID: 6KAC). The distal residues of the discontinuous loop of PsbO in TEF30-C2S are indicated by the arrows. The unobserved Lys206-Lys238 region in PsbO may be flexible in the absence of PsbP and PsbQ. e, The oxygen-evolving activity assays of SDG1 (TEF30-C), SDG2 (TEF302-C2) and SDG3 (TEF30-C2S). Each curve represents the averaged data from three technical replicates. The PSII-SC sample extracted from the low-light adapted WT cells was included as a positive control. f, The lack of cryo-EM densities in the local region around the Mn cluster-binding sites of TEF30-C, TEF302-C2-I and TEF302-C2-II. The cryo-EM maps are displayed in transparent gray. The Mn cluster models from PSII-SC are placed in the empty sites as a reference.
Extended Data Fig. 8 Identification of PsbW in TEF302-C2-I.
a, The structural model of TEF302-C2-I generated by the ModelAngelo software. The result for the identification of the transmembrane helix at the dimeric boundary in the model is displayed at the bottom. b, The western blots of the SDS-PAGE bands of PsbW and CP47 in the different TEF30-PSII complex samples. The SDG1, SDG2 and SDG3 samples contain TEF30-C, TEF302-C2 and TEF30-C2S, respectively. The PSI-LHCI and PSII-SC samples extracted from the SDG of solubilized WT thylakoids (Extended Data Fig. 1c) were loaded as the negative and positive controls respectively. For each lane, the sample containing 6.8×10−4 μmol target protein complexes was loaded. The experiment was repeated independently four times with similar results. c, The cryo-EM density fitted with the structural model of PsbW in TEF302-C2-I.
Extended Data Fig. 9 The weak densities of Psb27 and TEF14-like proteins in the cryo-EM map of TEF30-C and comparison of TEF30-C with the PSII-C associated with Psb27, Psb28 and Psb34.
a, The unsharpened map of TEF30-C with a low contour level is displayed at the top. The density of TEF30, the mask for Psb27 and the mask for TEF14 are colored in hot pink, blue and light orange, respectively. After the 3D classifications (skipping alignment) with Psb27 or TEF14 masked, the classes of TEF30-Psb27-C and TEF30-PRF1-TEF14-C particles are refined, and the maps are displayed at the bottom. TEF30, Psb27, PRF1 and TEF14 are colored and labeled. b, Superposition of TEF30-C (hot pink) with the PSII-C associated with Psb27, Psb28 and Psb34 (cornflower blue, PDB ID: 7NHP). The two structures are aligned on the D1 subunits. A part of the D1 DE-loop region (Leu223-Glu244) in 7NHP is highlighted in yellow. The overlapping local regions of the binding sites of TEF30 and Psb28 are zoomed in and shown at the bottom.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2 and description of Supplementary Video 1.
Supplementary Video 1 (download MP4 )
A putative model for the TEF30-involved PSII assembly process from the CP43 and RC47 modules to PSII-SC. Structural models of TEF30-C, TEF302-C2-I, TEF302-C2-II, TEF30-C2S and the C2S2-type PSII-SC (PDB ID: 6KAC) were visualized and animated using UCSF ChimeraX.
Source data
Source Data Fig. 1 and Extended Data Figs. 1 and 6 (download PDF )
Unprocessed HDN-PAGE and SDS–PAGE gel images for Fig. 1; unprocessed SDS–PAGE gel images, western blot images and micrographs for Extended Data Fig. 1; and unprocessed western blot images for Extended Data Fig. 6.
Source Data Fig. 3 and Extended Data Figs. 2, 5 and 7 (download XLSX )
Statistical source data.
Rights and permissions
About this article
Cite this article
Wang, Y., Wang, C., Li, A. et al. Roles of multiple TEF30-associated intermediate complexes in the repair and reassembly of photosystem II in Chlamydomonas reinhardtii. Nat. Plants 11, 1455–1469 (2025). https://doi.org/10.1038/s41477-025-02036-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02036-3
This article is cited by
-
A multifunctional LDH nano-platform enhances tobacco (Nicotiana tabacum L.) photosynthesis through phyllospheric spectral conversion and Mg/Mo nutrition
Chemical and Biological Technologies in Agriculture (2025)