Abstract
Drought stress affects plant growth and agricultural production, especially in the context of global climate change. Post-drought rehydration is crucial for plant recovery and sustained growth, yet the mechanisms underlying this process remain poorly understood. Nitrogen fertilizer plays a role in optimizing plant growth and enhancing stress resistance, but its role in post-drought recovery has not been fully elucidated. Here we demonstrate that nitrogen enhances post-drought recovery in wheat by modulating TaSnRK2.10-mediated regulation of TaNLP7. Transcriptomic analysis revealed that nitrogen supplementation increased the positive effects of rewatering on gene expression. Nitrogen inhibits the activity of TaSnRK2.10, a kinase involved in abscisic acid signalling. TaSnRK2.10 interacts with and phosphorylates TaNLP7-3A, a master regulator of the nitrate signalling pathway, reducing its nuclear localization and stability. This phosphorylation event suppresses genes involved in nitrate response, inhibiting nitrate-induced growth. Analysis of the nitrogen response levels in a wheat natural population revealed that transcriptional levels of the two haplotypes of TaSnRK2.10-4A respond differently to abscisic acid and nitrate, providing insights into the selection of wheat varieties that may be better suited for different environmental conditions to optimize yield.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analysed during this study are available in both the main text and the Supplementary Information. All wheat genes involved in this study are available in the Ensembl plants database (http://plants.ensembl.org/index.html) and WheatOmics 1.0 (http://202.194.139.32), including TaSnRK2.10-4A (TraesCS4A02G235600), TaSnRK2.10-4B (TraesCS4B02G079300), TaSnRK2.10-4D (TraesCS4D02G078100), TaSnRK2.8-5B (TraesCS5B02G406400), TaSnRK2.9-5D (TraesCS5D02G081700), TaNLP7-3A (TraesCS3A02G159600), TaNRT1.1b (TraesCS1A02G211000, TraesCS1B02G225000, TraesCS1D02G214300), TaNRT2.1 (TraesCS6B02G044300 and TraesCS6A02G031000), TaNR1 (TraesCS6D02G020700), TaGS2 (TraesCS2D02G500600) and TaADPRF (TraesCS3D02G330500). The Arabidopsis genes involved in this study can be found on TAIR (www.arabidopsis.org) under the following accession numbers: AtOST1 (AT4G33950), AtNLP7 (AT4G24020), AtNIA1 (AT1G77760) and AtNRT2.1 (AT1G08090). RNA-seq data that support the findings of this study have been deposited at the National Center for Biotechnology Information Gene Expression Omnibus under the following accession numbers: GSE302351 and GSE302352 for ABA and nitrogen co-regulation of gene expression; GSE302233 for nitrogen-mediated regulation of gene expression during post-drought rehydration. The raw sequencing data are available in the National Center for Biotechnology Information Sequence Read Archive under accession numbers PRJNA1182195 (ref. 47), PRJNA1286945 and PRJNA1286949. Source data are provided with this paper.
References
Bailey-Serres, J., Parker, J. E., Ainsworth, E. A., Oldroyd, G. E. D. & Schroeder, J. I. Genetic strategies for improving crop yields. Nature 575, 109–118 (2019).
Fredes, I., Moreno, S., Díaz, F. P. & Gutiérrez, R. A. Nitrate signaling and the control of Arabidopsis growth and development. Curr. Opin. Plant Biol. 47, 112–118 (2019).
Araus, V., Swift, J., Alvarez, J. M., Henry, A. & Coruzzi, G. M. A balancing act: how plants integrate nitrogen and water signals. J. Exp. Bot. 71, 4442–4451 (2020).
Koua, A. P. et al. Genome‐wide dissection and haplotype analysis identified candidate loci for nitrogen use efficiency under drought conditions in winter wheat. Plant Genome 17, e20394 (2024).
Vadez, V. et al. Crop traits and production under drought. Nat. Rev. Earth Environ. 5, 211–225 (2024).
Sato, H., Mizoi, J., Shinozaki, K. & Yamaguchi-Shinozaki, K. Complex plant responses to drought and heat stress under climate change. Plant J. Cell Mol. Biol. 117, 1873–1892 (2024).
Yang, Z. et al. Genetic and molecular exploration of maize environmental stress resilience: toward sustainable agriculture. Mol. Plant 16, 1496–1517 (2023).
Gessler, A., Bottero, A., Marshall, J. & Arend, M. The way back: recovery of trees from drought and its implication for acclimation. New Phytol. 228, 1704–1709 (2020).
Wang, X. et al. Improving resilience to high temperature in drought: water replenishment enhances sucrose and amino acid metabolisms in maize grain. Plant J. Cell Mol. Biol. 119, 658–675 (2024).
Siopongco, J. D. L. C., Yamauchi, A., Salekdeh, H., Bennett, J. & Wade, L. J. Growth and water use response of doubled-haploid rice linesto drought and rewatering during the vegetative stage. Plant Prod. Sci. 9, 141–151 (2006).
Miyashita, K., Tanakamaru, S., Maitani, T. & Kimura, K. Recovery responses of photosynthesis, transpiration, and stomatal conductance in kidney bean following drought stress. Environ. Exp. Bot. 53, 205–214 (2005).
Song, Y. et al. Nitrogen increases drought tolerance in maize seedlings. Funct. Plant Biol. 46, 350–359 (2019).
Soualiou, S., Duan, F., Li, X. & Zhou, W. Nitrogen supply alleviates cold stress by increasing photosynthesis and nitrogen assimilation in maize seedlings. J. Exp. Bot. 74, 3142–3162 (2023).
Abid, M. et al. Nitrogen nutrition improves the potential of wheat (Triticum aestivum L.) to alleviate the effects of drought stress during vegetative growth periods. Front. Plant Sci. 7, 981 (2016).
Saneoka, H., Moghaieb, R. E. A., Premachandra, G. S. & Fujita, K. Nitrogen nutrition and water stress effects on cell membrane stability and leaf water relations in Agrostis palustris Huds. Environ. Exp. Bot. 52, 131–138 (2004).
Shabbir, R. N. et al. Supplemental exogenous NPK application alters biochemical processes to improve yield and drought tolerance in wheat (Triticum aestivum L.). Environ. Sci. Pollut. Res. 23, 2651–2662 (2016).
Liu, K. et al. Discovery of nitrate–CPK–NLP signalling in central nutrient–growth networks. Nature 545, 311–316 (2017).
Konishi, M. & Yanagisawa, S. Arabidopsis NIN-like transcription factors have a central role in nitrate signalling. Nat. Commun. 4, 1617 (2013).
Castaings, L. et al. The nodule inception‐like protein 7 modulates nitrate sensing and metabolism in Arabidopsis. Plant J. 57, 426–435 (2009).
Liu, K.-H. et al. NIN-like protein 7 transcription factor is a plant nitrate sensor. Science 377, 1419–1425 (2022).
Yu, J. et al. Enhanced OsNLP4‐OsNiR cascade confers nitrogen use efficiency by promoting tiller number in rice. Plant Biotechnol. J. 19, 167–176 (2021).
Wu, J. et al. The OsNLP3/4‐OsRFL module regulates nitrogen‐promoted panicle architecture in rice. New Phytol. 240, 2404–2418 (2023).
Hu, B., Wang, W., Chen, J., Liu, Y. & Chu, C. Genetic improvement toward nitrogen-use efficiency in rice: lessons and perspectives. Mol. Plant 16, 64–74 (2023).
Song, Y. et al. Balanced nitrogen–iron sufficiency boosts grain yield and nitrogen use efficiency by promoting tillering. Mol. Plant 16, 1661–1677 (2023).
Han, M.-L. et al. Decreasing nitrogen assimilation under drought stress by suppressing DST-mediated activation of nitrate reductase 1.2 in rice. Mol. Plant 15, 167–178 (2022).
Ault, T. R. On the essentials of drought in a changing climate. Science 368, 256–260 (2020).
Jia, Z. & Von Wirén, N. Signaling pathways underlying nitrogen-dependent changes in root system architecture: from model to crop species. J. Exp. Bot. 71, 4393–4404 (2020).
Tilman, D., Cassman, K. G., Matson, P. A., Naylor, R. & Polasky, S. Agricultural sustainability and intensive production practices. Nature 418, 671–677 (2002).
Zhu, J. K. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Chen, K. et al. Abscisic acid dynamics, signaling, and functions in plants. J. Integr. Plant Biol. 62, 25–54 (2020).
Zhang, Z. et al. Isolation and characterization of the TaSnRK2.10 gene and its association with agronomic traits in wheat (Triticum aestivum L.). PLoS ONE 12, e0174425 (2017).
Zhang, Y. et al. Wheat TaSnRK2.10 phosphorylates TaERD15 and TaENO1 and confers drought tolerance when overexpressed in rice. Plant Physiol. 191, 1344–1364 (2023).
Li, G.-J., Chen, K., Sun, S. & Zhao, Y. Osmotic signaling releases PP2C-mediated inhibition of Arabidopsis SnRK2s via the receptor-like cytoplasmic kinase BIK1. EMBO J. 43, 6076–6103 (2024).
Marchive, C. et al. Nuclear retention of the transcription factor NLP7 orchestrates the early response to nitrate in plants. Nat. Commun. 4, 1713 (2013).
Wang, H. et al. Regulatory functions of cellular energy sensor SnRK1 for nitrate signalling through NLP7 repression. Nat. Plants 8, 1094–1107 (2022).
Belda-Palazón, B. et al. A dual function of SnRK2 kinases in the regulation of SnRK1 and plant growth. Nat. Plants 6, 1345–1353 (2020).
Belda-Palazón, B., Costa, M., Beeckman, T., Rolland, F. & Baena-González, E. ABA represses TOR and root meristem activity through nuclear exit of the SnRK1 kinase. Proc. Natl Acad. Sci. USA 119, e2204862119 (2022).
Wang, W. et al. SnpHub: an easy-to-set-up web server framework for exploring large-scale genomic variation data in the post-genomic era with applications in wheat. Gigascience 9, giaa060 (2020).
Niu, J. et al. Whole-genome sequencing of diverse wheat accessions uncovers genetic changes during modern breeding in China and the United States. Plant Cell 35, 4199–4216 (2023).
Mao, H. et al. Variation in cis-regulation of a NAC transcription factor contributes to drought tolerance in wheat. Mol. Plant 15, 276–292 (2022).
Gessler, A., Schaub, M. & McDowell, N. G. The role of nutrients in drought-induced tree mortality and recovery. New Phytol. 214, 513–520 (2017).
Han, C. et al. Brassinosteroid signaling and molecular crosstalk with nutrients in plants. J. Genet. Genomics 50, 541–553 (2023).
Cerda, A. & Alvarez, J. M. Insights into molecular links and transcription networks integrating drought stress and nitrogen signaling. New Phytol. 241, 560–566 (2024).
Wang, P. et al. Reactive oxygen species: multidimensional regulators of plant adaptation to abiotic stress and development. J. Integr. Plant Biol. 66, 330–367 (2024).
Chu, X. et al. HBI transcription factor-mediated ROS homeostasis regulates nitrate signal transduction. Plant Cell 33, 3004–3021 (2021).
Yang, Y. & Guo, Y. Unraveling salt stress signaling in plants. J. Integr. Plant Biol. 60, 796–804 (2018).
Yang, F. et al. TabHLH489 suppresses nitrate signaling by inhibiting the function of TaNLP7‐3A in wheat. J. Integr. Plant Biol. 67, 1162–1178 (2025).
Lyu, J. et al. The TaSnRK1‐TabHLH489 module integrates brassinosteroid and sugar signalling to regulate the grain length in bread wheat. Plant Biotechnol. J. 22, 1989–2006 (2024).
Ding, Y. et al. OST1 kinase modulates freezing tolerance by enhancing ICE1 stability in Arabidopsis. Dev. Cell 32, 278–289 (2015).
Chu, X. et al. HBI1‐TCP20 interaction positively regulates the CEPs‐mediated systemic nitrate acquisition. J. Integr. Plant Biol. 63, 902–912 (2021).
Acknowledgements
Thanks to H. Yu, X. Zhao, Y. Guo and S. Wang from the Core Facilities for Life and Environmental Sciences at the State Key Laboratory of Microbial Technology, Shandong University, for the assistance provided in laser scanning confocal microscopy imaging. We appreciate Z. Deng (State Key Laboratory for Managing Biotic and Chemical Threats to the Quality and Safety of Agro-products, Institute of Virology and Biotechnology, Zhejiang Academy of Agricultural Sciences) for the mass spectrophotometric analysis. We gratefully acknowledge Y. Zhao (Center for Excellence in Molecular Plant Sciences, Chinese Academy of Sciences) for generously providing the Phospho-SnRK2.6-S175-pAb antibody (ABclonal, AP1481). This work was supported by grants from the National Natural Science Foundation of China (grant numbers U24A20389, 32461160287, 32325006 and 32070210 to M.-Y.B.; grant number 32270351 to C.H. and grant number U22A6009 to J.X.) and the Agricultural Variety Improvement Project of Shandong Province (grant number 2024LZGC007 to M.F. and grant number 2022LZGC001 to M.-Y.B.).
Author information
Authors and Affiliations
Contributions
J.M., H.W., D.W., J.X., M.F. and M.-Y.B. together designed the experiments. J.M. performed phenotypic analyses under drought and rehydration conditions, as well as RNA-seq analysis. J.M. and H.W. performed subcellular location analysis, kinase assay, western blot and quantitative RT–PCR. J.M. and D.W. performed haplotype classification and natural variations analysis of TaSnRK2.10-4A. J.M. and X.Y. performed statistical analysis of plant growth, genotype identification and plasmid construction. J.M. and X.Y. generated OE-TaSnRK2.10-4AHap-I/II, OE-TaNLP7T260AS284A, OE-TaNLP7T260DS284D, OE-TaSnRK2.10/ost1-3, OE-TaNLP7T260AS284A/nlp7-1, OE-TaNLP7T260DS284D/nlp7-1 transgenic plants and Tasnrk2.10 mutants. F.Y. and J.L. generated OE-TaNLP7 and OE-TaNLP7/nlp7-1 transgenic plants. J.M. and F.Y. performed RNA-seq analysis. J.M., N.S., G.Z., R.Z., B.X. and S.X. performed phenotypic analysis of seedling nitrogen response of different wheat varieties. J.M. performed all other experiments. C.H., G.-M.X., G.L., J.X., M.F. and M.-Y.B. provided critical discussion on the work. J.M. and M.-Y.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Jie Zhou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nitrogen promotes the recovery of gene expression after rewatering by suppressing ABA signal transduction.
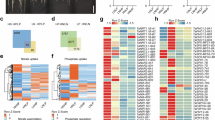
a, Venn diagram showing the overlap between sets of genes regulated by drought, RW5d and RWN5d. b, Hierarchical clustering analysis of the expression profiles of 8,604 genes co-regulated by drought, RW5d, and RWN5d. The color gradient from blue to red represents logarithm (base 2) of the fold change. c, Scatter plot depicting the logarithm (base 2) of the fold change of 8,604 genes co-regulated by drought, RW5d, and RWN5d, comparing expression levels under drought conditions and during RW5d or RWN5d treatments. d, Venn diagram depicting genes regulated by drought within the core ABA signaling pathway. e, Bubble chart illustrating the distribution and number of drought-regulated genes across six expression trends during the rehydration process. f, Hierarchical clustering of ABA signaling and biosynthesis genes exhibiting the ‘Late reduce’ expression pattern. Color gradient (blue to red) represents log2 fold changes in gene expression.
Extended Data Fig. 2 Nitrogen-mediated downregulation of TaSnRK2.10-4A and its homologs facilitates the transcriptional reactivation of PNR genes upon rehydration after drought stress.
a, Heatmap showing the expression profiles of TaSnRK2.8/2.9/2.10 during drought, rehydration, and rehydration with nitrogen supplementation in wheat. b,c, Quantitative RT-PCR analysis of drought effects on TaSnRK2.8/2.9-ABD expression, TaADPRF was used as an internal control. Data are mean ± s.d. (n = 3 biological replicates). Asterisks indicate significant differences (two-tailed, unpaired t-test, *p < 0.05, **p < 0.01). d, Nitrate-mediated suppression of TaSnRK2.8-ABD transcription during rehydration. JW1 wheat seedlings were drought-stressed (5% soil water content) for 1 d, then rewatered with nitrogen-free (RW) or nitrogen-supplemented (RWN) solutions. Samples were collected at indicated time points for quantitative RT-PCR. Data are mean ± s.d. (n = 3 biological replicates). (two-tailed, unpaired t-test, ***p < 0.001, ns, not significant). e,f, Drought-induced expression of TaSnRK2.10-4B / 4D. Error bars represent mean values ± s.d. of three biological replicates. Asterisks indicate statistically significant differences between samples (two-tailed, unpaired t-test, *p < 0.05, **p < 0.01). g,h, Expression dynamics of TaSnRK2.10-4B / 4D during rehydration and nitrogen-supplemented rehydration. Wheat seedlings of the JW1 cultivar were exposed to drought until the soil water content fell below 5%. After one day of drought stress, the seedlings were rewatered with either nitrogen-free (RW) or nitrogen-supplemented (RWN) wheat nutrient solutions for 0 h, 3 h, 1 d, or 5 d. Leaf samples were then collected at each time point for total RNA extraction and quantitative RT-PCR analysis to assess TaSnRK2.10-4B /4D transcript levels. TaADPRF was used as an internal control. The numbers above the column indicate the fold change (RW / RWN). Error bars represent mean values ± s.d. of three biological replicates. Asterisks indicate statistically significant differences between samples (two-tailed, unpaired t-test, *p < 0.05, ns, no significant difference). i, Hierarchical clustering analysis of gene expression profiles for primary nitrate response genes showing the ‘Late reduced’ trend. The color gradient from blue to red indicates the log2 fold change in gene expression. j-l, Quantitative RT-PCR analysis of TaNRT1.1b (j), TaGS2 (k), and TaNR1 (l) expression during drought recovery with nitrogen-free (RW) or nitrogen-supplemented (RWN) solutions. Wheat seedlings of the JW1 cultivar were exposed to drought until the soil water content fell below 5%. After one day of drought stress, the seedlings were rewatered with either nitrogen-free (RW) or nitrogen-supplemented (RWN) wheat nutrient solutions for 0 h, 3 h, 1 d, or 5 d. Leaf samples were then collected at each time point for total RNA extraction. TaADPRF was used as an internal control. The numbers above the column indicate the fold change (RWN / RW). Error bars represent mean values ± s.d. of three biological replicates. Asterisks indicate statistically significant differences between samples (two-tailed, unpaired t-test, *p < 0.05, **p < 0.01, ***p < 0.001, ns, no significant difference). m, Correlation analysis showing that the expression profile of TaSnRK2.10-4A during RW and RWN processes is negatively correlated with nitrogen assimilation related genes. The color gradient from light green to dark blue represents pearson correlation coefficients, and circle size indicates the strength of the correlation.
Extended Data Fig. 3 TaSnRK2.10-4A confers ABA sensitivity to the ost1-3 mutant.
a, immunoblot analysis of TaSnRK2.10-YFP protein levels in different OE-TaSnRK2.10/ost1-3 transgenic lines using an anti-GFP antibody. b, phenotypic assessment of 7-day-old seedlings of Col-0, ost1-3, and OE-TaSnRK2.10/ost1-3 grown on ½ MS medium with or without 0.5 µM ABA. Scale bars, 2 cm. c, TaSnRK2.10-4A partially restores ABA-insensitive seed germination in ost1-3 mutant. Germination rates were measured using a minimum of 50 seeds per replicate. Error bars indicate mean values ± s.d. of three biological replicates. Asterisks indicate statistically significant differences between samples (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, **p < 0.01, ***p < 0.001, ns, no significant difference). d,e, TaSnRK2.10-4A partially restores ABA-insensitive stomatal opening in ost1-3 mutant. Col-0, ost1-3 and OE-TaSnRK2.10/ost1-3 plants were grown on ½ MS medium with or without 10 µM ABA for 7 days, then stomatal aperture was measured following 1 h of light exposure. Scale bars, 20 µm. Error bars indicate mean values ± s.d. (n = 52 independent stomatal). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05).
Extended Data Fig. 4 Nitrate counteracts the dual roles of TaSnRK2.10-4A in growth inhibition and drought tolerance promotion by triggering its protein degradation.
a, The expression levels of TaSnRK2.10-4A in JW1 and various TaSnRK2.10-4A overexpression transgenic wheat lines. TaADPRF was used as an internal control. Error bars indicate mean values ± s.d. of three biological replicates. Statistically significant differences between samples are indicated by different letters above the bars (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). b, CRISPR / Cas9-induced targeted mutagenesis of TaSnRK2.10-4A. Red box indicates sgRNAs designed to target all homologous chromosomes for TaSnRK2.10, with mutation sites highlighted in red. c-g, Plant architecture comparison among of JW1, OE-TaSnRK2.10, and Tasnrk2.10 plants at the heading stage. scale bar, 10 cm. Error bars indicate mean values ± s.d. (n = 12 independent plants for d,f,g; n = 21 (JW1), n = 44 (OE-TaSnRK2.10-6#), n = 37 (OE-TaSnRK2.10-8#), n = 20 (Tasnrk2.10-1#), n = 23 (Tasnrk2.10-20#) independent plants for e). Statistically significant differences between samples are indicated by different letters above the bars (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). h, TaSnRK2.10-4A enhances wheat drought tolerance. Soil water content measurements in JW1, OE-TaSnRK2.10, and Tasnrk2.10 plants at 10, 15, and 17 days post treatment. i,j, ABA enhances TaSnRK2.10-4A phosphorylation. Seven-day-old OE-TaSnRK2.10 seedlings were treated with 50 µM ABA for indicated durations, followed by root protein extraction and immunoblot analysis. Immunoblotting was performed using anti-GFP and Phospho-SnRK2.6-S175-pAb to detect the protein expression and phosphorylation levels of TaSnRK2.10-4A, respectively. The ratio of Phospho-SnRK2.6-S175 to TaSnRK2.10-GFP was quantified using ImageJ software. Error bars represent mean values ± s.d. (n = 3 biological repeats). Different letters above the lines indicate statistically significant differences between samples (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). k,l, Nitrate reduces the protein stability of TaSnRK2.10-4A. Seedlings of OE-TaSnRK2.10 were prepared by removing the endosperm from 3-day-old germinated seeds, followed by 3 days of growth in wheat nutrient culture medium containing 2 mM Ca(NO3)2. After 2 days of nitrogen starvation, seedlings were treated with 2 mM Ca(NO3)2 for varying durations. Western-blot analysis was performed to detect the protein levels and catalytic activity of TaSnRK2.10-4A using anti-GFP and anti-Phospho-Ser175-SnRK2.6 antibodies. The ratio of the immunoprecipitated TaSnRK2.10-GFP to actin was quantified using ImageJ software. Error bars represent mean values ± s.d. (n = 3 biological repeats). Different letters above the bars indicate statistically significant differences between samples (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05).
Extended Data Fig. 5 ABA inhibits nitrate-regulated wheat growth and PNR gene expression.
a-c, Mutation in TaSnRK2.10 reduces the inhibitory effect of ABA on wheat growth. Seedlings derived from 3-day-old germinated seeds with endosperm removed were grown in wheat nutrient culture medium with either 2 mM CaCl2 or 2 mM Ca(NO3)2, under ABA treatment or control conditions for 10 days. Error bars indicate mean values ± s.d. (n = 12 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). d-f, The effects of ABA and nitrate treatment on wheat shoot growth. 3-day-old JW1 seedlings after the endosperm removed were transferred to wheat nutrient culture medium containing 0.05 mM or 5 mM nitrate, and simultaneously treated with ABA at concentrations of 0, 0.5, 1, 3 and 5 µM. After a 10-day cultivation period, phenotypic variations were documented (d), the change index (e) and the shoot fresh weight (f) was quantified and analysis. Scale bars, 5 cm. Error bars indicate mean values ± s.d. (n = 12 independent plants). The numbers above the column in (e) indicate the fold change regulated by nitrate. Error bars in (e) represent s.d. of three biological replicates. Asterisks indicate statistically significant differences between samples (two-tailed, unpaired t-test, *p < 0.05, ***p < 0.001, ****p < 0.0001). The thick dashed lines of violin plots represent median, the thin dashed lines represent first or third quartile. Statistically significant differences between samples in (f) are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). g-i, Quantitative RT-PCR analysis shows the gene expression level of TaNRT2.1 (g), TaNR1 (h), and TaGS2 (i) under nitrate treatment, with or without 5 μM ABA. JW1 seedlings after the endosperm removal from 3-day-old germinated seeds, were grown for 3 days in wheat nutrient culture medium containing 2 mM Ca(NO3)2, subjected to 2 days of nitrogen starvation, and subsequently treated with 2 mM Ca(NO3)2 or 2 mM CaCl2 in the presence or absence of 5 μM ABA for 3 h. TaADPRF served as an internal control. The numbers above the column indicate the fold change regulated by nitrate. Error bars represent mean values ± s.d. of three biological replicates. Asterisks indicate statistically significant differences between samples (unpaired t-test, *p < 0.05, **p < 0.01, ***p < 0.001).
Extended Data Fig. 6 Conserved Interaction and Phosphorylation of TaSnRK2.10 with TaNLP7 in Arabidopsis and Wheat.
a, Yeast-based assay confirms the interaction between OST1 and NLP7. b, Confocal microscopy images from ratiometric bimolecular fluorescence complementation (rBiFC) assays reveal the interaction between OST1 and NLP7 in epidermal cells of tobacco leaves. Scale bar = 20 μm. c, Pull-down assays confirm the direct interaction between OST1 and NLP7 in vitro. Recombinant MBP and MBP-OST1 proteins were incubated with glutathione agarose beads conjugated to GST-NLP7. Immunoblot assay was performed with anti-MBP antibody to detect the proteins bound to GST-NLP7. d, Pull-down assays demonstrate the direct interaction between TaSnRK2.8-5B and TaSnRK2.9-5D with TaNLP7-3A. Recombinant MBP, MBP-TaSnRK2.8-5B and MBP-TaSnRK2.9-5D proteins were incubated with GST-TaNLP7-3A immobilized on glutathione agarose beads, and the interaction was confirmed by immunoblotting with an anti-MBP antibody. e-g, TaSnRK2.8/2.9/2.10 phosphorylates TaNLP7 in vitro. The upper gel image shows ATP-gamma-S labeled proteins, while the following gel image is stained with Coomassie brilliant blue (CBB). Phosphorylation was confirmed by immunoblotting with an anti-TPE antibody. h,i, Mass spectrometry analysis identified OST1-dependent phosphorylation residues within the NLP7 protein. j, The conservation of these OST1-dependent phosphorylation residues across the NLP7 protein family in different plant species is shown in the protein alignment. Protein sequence of NLP7 homologues from monocotyledonous (Triticum aestivum L., TraesCS3A02G159600, TraesCS3B02G190300, and TraesCS3D02G166900, Oryza sativa, Os01t0236700, and BGIOSGA003095, and Zea mays, Zm00001eb002490), and dicotyledonous (Arabidopsis thaliana, At4G24020 and Lotus corniculatus, Lj5g3v1999250) were aligned. Phosphorylation sites mediated by OST1 are indicated with red arrows. Multiple sequence alignment was conducted using DNAMAN software.
Extended Data Fig. 7 Phenotypic analysis of normal growth and nitrogen-related traits in TaNLP7 and its point-mutated materials.
a, Semi-quantitative RT-PCR analysis the transcript levels of TaNLP7 in JW1, OE-TaNLP7, OE-TaNLP7T260AS284A, and OE-TaNLP7T260DS284D plants, with TaADPRF as an internal reference gene. b-d, Comparison of plant architecture at the heading stage among JW1, OE-TaNLP7, OE-TaNLP7T260AS284A and OE-TaNLP7T260DS284D. Scale bar, 5 cm. Error bars indicate mean values ± s.d. (n = 12 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). e,f, Phenotypic images and spike length statistics of wheat during the heading stage for the JW1 variety and transgenic lines OE-TaNLP7, OE-TaNLP7T260AS284A, and OE-TaNLP7T260DS284D plants. The scale bar represents 2 cm. Error bars indicate mean values ± s.d. (n = 12 independent plants). Statistically significant differences among samples are denoted by different letters above the bars (One-way ANOVA and uncorrected Fisher’s LSD multiple comparison test, p < 0.05). g,h, Tiller number of TaNLP7 and TaNLP7T260AS284A overexpression lines grown under low or high nitrogen conditions, indicated by blue arrows. Error bars indicate mean values ± s.d. (n = 8 independent plants). Different letters above the bars indicate statistically significant differences between samples (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparison test, p < 0.05). i-k, Overexpression of TaNLP7T260AS284A mitigates the inhibitory effects of ABA treatment on plant growth. After three days of growth under normal conditions and subsequent endosperm removal, JW1 and different transgenic wheat seedlings were transferred to wheat nutrient culture medium containing 5 mM CaCl2 or Ca(NO3)2, with or without 1 µM ABA, for an additional 10 days. Error bars indicate mean values ± s.d. (n = 12 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05).
Extended Data Fig. 8 TaSnRK2.10-4A-mediated phosphorylation negatively regulates the function of TaNLP7-3A.
a, Semi-quantitative RT-PCR analysis the expression levels of TaNLP7 and mutant version of TaNLP7 in Col-0, nlp7-1, OE-TaNLP7/nlp7-1, OE-TaNLP7T260AS284A/nlp7-1, and OE-TaNLP7T260DS284D/nlp7-1 plants. AtPP2A was used as an internal reference gene. b, Restoration of the nlp7-1 mutant phenotype by TaNLP7 and TaNLP7T260AS284A. Col-0, nlp7-1, OE-TaNLP7/nlp7-1, OE-TaNLP7T260AS284A/nlp7-1, and OE-TaNLP7T260DS284D/nlp7-1 plants were grown under long day conditions (16 h light / 8 h dark) for 3 weeks. Scale bar, 2 cm. c-g, Quantitative analysis of leaf area, primary root length and total lateral root length in Arabidopsis Col-0, nlp7-1 and transgenic plants complemented with different TaNLP7 constructs. Seedlings of Col-0, nlp7-1, OE-TaNLP7/nlp7-1, OE-TaNLP7T260AS284A/nlp7-1, and OE-TaNLP7T260DS284D/nlp7-1 plants were grown in MGRL medium with either 10 mM KCl (Nitrate-free) or 10 mM KNO3 (Nitrate). Leaf area (c,d) was measured with the seeding grown for 7 days, scale bar, 1 cm. Error bars indicate mean values ± s.d. (n = 15 independent plants). Root length (e-g) was measured with the seeding grown for 14 days, scale bar, 2 cm. Error bars indicate mean values ± s.d. (n = 10 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05).
Extended Data Fig. 9 Overexpression of TaNLP7 reduced drought tolerance.
a, Determination of soil moisture content of wheat seedlings (JW1, OE-TaNLP7, OE-TaNLP7T260AS284A, and OE-TaNLP7T260DS284D) under drought stress. Wheat seeds of were germinated and grown in 2 L pots containing wheat nutrient culture medium supplemented with 5 mM nitrate. Soil water content was measured at 10, 15, and 17 days after planting for the following plant materials: JW1, OE-TaNLP7, OE-TaNLP7T260AS284A and OE-TaNLP7T260DS284D. No additional water was provided during the experiment, allowing for natural evaporation. b-d, The Tanlp7 mutants exhibit enhanced drought tolerance. Newly germinated JW1 (wild-type) and Tanlp7 mutant lines were cultivated in 2 L pots containing wheat nutrient culture medium (5 mM nitrate), with no additional water added to allow for natural evaporation. Shoot biomass (c) and soil water content (d) were measured at 15- and 21-days post-planting. Scale bars, 3 cm. Error bars represent mean values ± s.d. (n = 8 independent plants for c; n = 3 biological repeats for d). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). e, TaSnRK2.10-4A interacts with TaSnRK1α1 in yeast. f, Pull-down assays demonstrate the direct interaction between TaSnRK2.10-4A with TaSnRK1α1. Recombinant GST and GST-TaSnRK2.10-4A proteins were incubated with MBP-TaSnRK1α1 immobilized on glutathione agarose beads, and the interaction was confirmed by immunoblotting with an anti-GST antibody. g, Heatmap of TaSnRK1α1-1ABD expression profiles during drought, rehydration, and rehydration with nitrogen supplementation in wheat. Color gradient (blue to red) represents log2 fold changes in gene expression. h,i, Drought stress tolerance of TaSnRK1α1 transgenic plants. Four-day-old wheat seedlings were transplanted into soil (a 1:3 mixture of Pindstrup substrate peat soil and vermiculite) and irrigated with the wheat nutrient solution containing 5 mM nitrate, allowing for natural evapotranspiration. Above-ground biomass was measured at 15- and 21- days post-transplantation. Scale bar, 3 cm. Error bars indicate mean values ± s.d. (n = 9 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05).
Extended Data Fig. 10 Overexpressed TaSnRK2.10-4A from various haplotypes resulted in similar growth inhibition phenotypes.
a, Quantitative RT-PCR analysis of TaSnRK2.10-4A expression levels in JW1 and different TaSnRK2.10-4A overexpression lines. TaADPRF was used as an internal control. Error bars indicate mean values ± s.d. of three biological replicates. Statistically significant differences between samples are indicated by different letters above the bars (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). b-d, Statistical analysis of tiller number (c) and spikelet number (d) of JW1 and transgenic lines overexpressing TaSnRK2.10-4AHap-I/II. Error bars indicate mean values ± s.d. (n = 10 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (One-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). e,f, Overexpressing of two distinct haplotypes of TaSnRK2.10-4A result in similar reductions in nitrogen response in wheat. Error bars represent mean values ± s.d. (n = 16). Different letters above the bars indicate statistically significant differences between samples (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05). g,h, Nitrate exerted distinct effects on the degradation rates of different haplotypes of the TaSnRK2.10 protein. Transgenic lines expressing either p35S:TaSnRK2.10-4AHap-I-YFP or p35S:TaSnRK2.10-4AHap-II-YFP were initially grown on ½ MS medium for 7 days. These lines were then transferred to MGRL medium supplemented with 10 mM nitrate and underwent a time-course treatment at intervals of 0, 6, 12, 24, and 48 h. i-l, Overexpression of TaSnRK2.10-4A from diverse haplotypes both inhibits nitrate-promoted wheat growth. Seedlings derived from 3-day-old germinated seeds, with endosperm removed, were grown in wheat nutrient culture medium containing either 5 mM CaCl2 or 5 mM Ca(NO3)2 for 14 days. Error bars indicate mean values ± s.d. (n = 12 independent plants). Statistically significant differences between samples are indicated by different letters above the bars (Two-way ANOVA followed by uncorrected Fisher’s LSD multiple comparisons test, p < 0.05).
Supplementary information
Supplementary Tables (download XLSX )
Supplementary Tables 1–15.
Supplementary Data 1 (download XLSX )
Statistical analysis of Figs. 1–6 and Extended Data Figs. 1–10.
Source data
Source Data Figs. 1–6 and Extended Data Figs. 1–10 (download XLSX )
Statistical source data and unprocessed western gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mu, J., Wang, H., Wang, D. et al. Nitrogen enhances post-drought recovery in wheat by modulating TaSnRK2.10-mediated regulation of TaNLP7. Nat. Plants 11, 1810–1826 (2025). https://doi.org/10.1038/s41477-025-02083-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02083-w
This article is cited by
-
Nitrate to spark parched plants
Nature Plants (2025)