Abstract
Fruit ripening is a tightly regulated developmental process, in which nuclear gene transcription represents a crucial component of the mechanisms1. Chloroplast-associated protein degradation, a recently discovered pathway for chloroplast protein degradation, has also been reported to control fruit ripening2. Here we report a negative regulator of tomato ripening, termed SlSAD8, which disturbs both nuclear gene transcription and chloroplast-associated protein degradation. As an atypical stearoyl-ACP desaturase (SAD) protein exhibiting dual localization in plastids and the nucleus, SlSAD8 negatively regulates ripening initiation and chloroplast-to-chromoplast transition during fruit ripening. In the nucleus, SlSAD8 interacts with ripening-initiation-associated transcription factor SlNAM1, thereby disturbing the transcriptional activation of ethylene biosynthesis genes. Additionally, SlSAD8 interacts with plastid-transition-associated E3 ligase SlSP1 in the plastid, disturbing the chloroplast-associated protein degradation pathway to elevate chloroplast protein levels. Our findings uncover an unusual ripening regulator that targets distinct subcellular compartments to manipulate gene expression, providing insights into the intricate regulatory networks of fruit ripening.
This is a preview of subscription content, access via your institution
Access options




Similar content being viewed by others
Data availability
All data analysed during this study are included in this Letter and its Supplementary Information. Tomato gene sequences were collected from the Sol Genomics Network (https://solgenomics.net). Additional data related to this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Brumos, J. Gene regulation in climacteric fruit ripening. Curr. Opin. Plant Biol. 63, 102042 (2021).
Ling, Q. et al. The chloroplast-associated protein degradation pathway controls chromoplast development and fruit ripening in tomato. Nat. Plants 7, 655–666 (2021).
Wang, W., Wang, Y., Chen, T., Qin, G. & Tian, S. Current insights into posttranscriptional regulation of fleshy fruit ripening. Plant Physiol. 192, 1785–1798 (2023).
Zhu, G. et al. Rewiring of the fruit metabolome in tomato breeding. Cell 172, 249–261 (2018).
Zhou, L. et al. Redox modification of m6A demethylase SlALKBH2 in tomato regulates fruit ripening. Nat. Plants 11, 218–233 (2025).
Li, X., Wang, X., Zhang, Y., Zhang, A. & You, C.-X. Regulation of fleshy fruit ripening: from transcription factors to epigenetic modifications. Hortic. Res. 9, uhac013 (2022).
Liu, Y. et al. The molecular regulation of ethylene in fruit ripening. Small Methods 4, 1900485 (2020).
Egea, I. et al. Chloroplast to chromoplast transition in tomato fruit: spectral confocal microscopy analyses of carotenoids and chlorophylls in isolated plastids and time-lapse recording on intact live tissue. Ann. Bot. 108, 291–297 (2011).
Ling, Q. et al. Ubiquitin-dependent chloroplast-associated protein degradation in plants. Science 363, eaav4467 (2019).
Haralampidis, K. et al. Temporal and transient expression of stearoyl-ACP carrier protein desaturase gene during olive fruit development. J. Exp. Bot. 49, 1661–1669 (1998).
Taha, R. S., Ismail, I., Zainal, Z. & Abdullah, S. N. A. The stearoyl-acyl-carrier-protein desaturase promoter (Des) from oil palm confers fruit-specific GUS expression in transgenic tomato. J. Plant Physiol. 169, 1290–1300 (2012).
Sedeek, K. E. et al. Amino acid change in an orchid desaturase enables mimicry of the pollinator’s sex pheromone. Curr. Biol. 26, 1505–1511 (2016).
Kachroo, P., Shanklin, J., Shah, J., Whittle, E. J. & Klessig, D. F. A fatty acid desaturase modulates the activation of defense signaling pathways in plants. Proc. Natl Acad. Sci. USA 98, 9448–9453 (2001).
Lu, J. et al. Types of nuclear localization signals and mechanisms of protein import into the nucleus. Cell Commun. Signal. 19, 60 (2021).
Kazaz, S. et al. Differential activation of partially redundant Δ9 stearoyl-ACP desaturase genes is critical for omega-9 monounsaturated fatty acid biosynthesis during seed development in Arabidopsis. Plant Cell 32, 3613–3637 (2020).
Li, Y. et al. Arabidopsis EXECUTER1 interacts with WRKY transcription factors to mediate plastid-to-nucleus singlet oxygen signaling. Plant Cell 35, 827–851 (2023).
Qi, P. et al. A Ralstonia solanacearum effector targets TGA transcription factors to subvert salicylic acid signaling. Plant Cell 34, 1666–1683 (2022).
Deng, H. et al. SlERF.F12 modulates the transition to ripening in tomato fruit by recruiting the co-repressor TOPLESS and histone deacetylases to repress key ripening genes. Plant Cell 34, 1250–1272 (2022).
Cao, H. et al. Tomato transcriptional repressor MYB70 directly regulates ethylene-dependent fruit ripening. Plant J. 104, 1568–1581 (2020).
Gao, Y. et al. A tomato NAC transcription factor, SlNAM1, positively regulates ethylene biosynthesis and the onset of tomato fruit ripening. Plant J. 108, 1317–1331 (2021).
Gao, Y. et al. A NAC transcription factor, NOR-like1, is a new positive regulator of tomato fruit ripening. Hortic. Res. 5, 75 (2018).
Jiang, G. et al. WUSCHEL-related homeobox transcription factor SlWOX13 regulates tomato fruit ripening. Plant Physiol. 194, 2322–2337 (2024).
Liu, Y., Shi, Y., Su, D., Lu, W. & Li, Z. SlGRAS4 accelerates fruit ripening by regulating ethylene biosynthesis genes and SlMADS1 in tomato. Hortic. Res. 8, 3 (2021).
Liu, M., Pirrello, J., Chervin, C., Roustan, J.-P. & Bouzayen, M. Ethylene control of fruit ripening: revisiting the complex network of transcriptional regulation. Plant Physiol. 169, 2380–2390 (2015).
D’Andrea, L. et al. Interference with Clp protease impairs carotenoid accumulation during tomato fruit ripening. J. Exp. Bot. 69, 1557–1568 (2018).
Li, F. et al. A tomato HD-zip I transcription factor, VAHOX1, acts as a negative regulator of fruit ripening. Hortic. Res. 10, uhac236 (2023).
Tang, R. et al. The FvABF3–FvALKBH10B–FvSEP3 cascade regulates fruit ripening in strawberry. Nat. Commun. 15, 10912 (2024).
Li, N. & Jarvis, R. P. Recruitment of Cdc48 to chloroplasts by a UBX-domain protein in chloroplast-associated protein degradation. Nat. Plants 10, 1400–1417 (2024).
Bell, C. C. et al. Comparative cofactor screens show the influence of transactivation domains and core promoters on the mechanisms of transcription. Nat. Genet. 56, 1181–1192 (2024).
Matarasso, N., Schuster, S. & Avni, A. A novel plant cysteine protease has a dual function as a regulator of 1-aminocyclopropane-1-carboxylic acid synthase gene expression. Plant Cell 17, 1205–1216 (2005).
Hu, N. et al. Rapid and user-friendly open-source CRISPR/Cas9 system for single-or multi-site editing of tomato genome. Hortic. Res. 6, 7 (2019).
Tamura, K. et al. MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood evolutionary distance and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Li, Y. et al. MicroTom metabolic network: rewiring tomato metabolic regulatory network throughout the growth cycle. Mol. Plant 13, 1203–1218 (2020).
Huang, K.-L. et al. Fatty acid export protein BnFAX6 functions in lipid synthesis and axillary bud growth in Brassica napus. Plant Physiol. 186, 2064–2077 (2021).
Wu, M. et al. SlMYB72 regulates the metabolism of chlorophylls, carotenoids, and flavonoids in tomato fruit. Plant Physiol. 183, 854–868 (2020).
Xu, X. et al. Auxin and abscisic acid antagonistically regulate ascorbic acid production via the SlMAPK8–SlARF4–SlMYB11 module in tomato. Plant Cell 34, 4409–4427 (2022).
Lin, Y. et al. A conserved fungal effector disturbs Ca2+ sensing and ROS homeostasis to induce plant cell death. Nat. Commun. 16, 3523 (2025).
Gao, G. et al. E3 ligase SlCOP1-1 stabilizes transcription factor SlOpaque2 and enhances fruit resistance to Botrytis cinerea in tomato. Plant Physiol. 196, 1196–1213 (2024).
Acknowledgements
We thank Y. Zhang from the Analytical and Testing Center of Chongqing University for technological guidance. This study was supported by the National Key Research and Development Program of China (grant no. 2022YFD2100100), the National Natural Science Foundation of China (grant nos 32272381 and 32230092), Project of Chongqing Science and Technology Commission (grant no. CSTB2022NSCQ-MSX0959) and Fundamental Research Funds for the Central Universities (grant nos 2024IAIS-ZX005 and 2024CDJXY016).
Author information
Authors and Affiliations
Contributions
Y.C. designed the study. C.X., R.L., X. Chen, Z.F., X. Cui and J.Y. conducted the experiments and analysed the data. Y.C., Z.L., W.D., Y.S. and J.Y. provided project supervision and resources. Y.C. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Hua Zhang and Da-Qi Fu for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
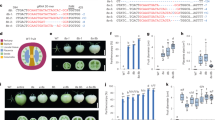
Extended Data Fig. 1 Expression profiling and phylogenetic analysis of tomato SAD genes.
a, The expression profiles of SAD genes in different tomato tissues according to the MMN (Micro-Tom Metabolic Network) database. The expression profiles of SlSAD3 and SlSAD6 were not identified in the MMN database. MG, mature green; Br, breaker; Br + 3-15, 3-15 days after the Br stage. b, Phylogenetic analysis of plant SAD proteins. Phylogenetic tree was generated by the MEGA5 software using the neighbor-joining method.
Extended Data Fig. 2 Arginine (R) and lysine (K) residues in the NLS are essential for SlSAD8’s nuclear localization.
a, Schematic representation of different SlSAD8 mutants in the NLS. Red residues indicate point-mutation sites. b, Nuclear localization proportion of SlSAD8 and its mutants. Data are shown as mean ± s.d. of three biological replicates, and differences were assessed using two-tailed t-test (***, P < 0.001). c, Representative subcellular localization images of SlSAD8 and its mutants. Co-expression of the nuclear marker AtHY5-mCherry was used as a reference for nuclear localization. Bars, 20 μm.
Extended Data Fig. 3 The content of major fatty acids in WT, SlSAD8-KO, and SlSAD8-OE fruits (a) and seeds (b).
Major fatty acids in tomato including palmitic acid (C16:0), stearic acid (C18:0), oleic acid (C18:1), linoleic acid (C18:2), and α-linolenic acid (C18:3). Data are shown as mean ± s.d. of three biological replicates, and differences were assessed using two-tailed t-test (**, P < 0.01; *, P < 0.05; NS, no significant).
Extended Data Fig. 4 LCI assays showed that SlSAD8 did not interact with seven other ripening-associated transcription factors.
a–g, LCI assays were performed in N. benthamiana leaves. All experiments were repeated at three independent times with similar results.
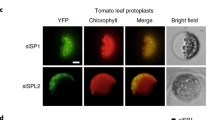
Extended Data Fig. 5 SAD8 did not interact with three other plastid proteins and was not ubiquitinated by SlSP1.
a-c, LCI assays showed that SAD8 did not interact with ClpR1 (a), SlSPL2 (b), and SlCDC48 (c). d, In vitro ubiquitination assay indicated that SlSP1 did not ubiquitinate SlSAD8. In the scenario where all ubiquitination reaction components were fully supplied (rightmost column), the smear bands indicative of SlSAD8 ubiquitination in anti-His Western blotting remained undetected. All experiments were repeated at three independent times with similar results.
Extended Data Fig. 6 The contribution of NLS and cTP to SlSAD8-mediated fruit ripening.
a, Immunoblot analysis was performed to check the subcellular localization of SlSAD8, SlSAD8ΔcTP, and SlSAD8ΔNLS when expressed in SlSAD8-KO (#5) fruits. Toc75 and histone H3 proteins were used as plastid and nuclear markers, respectively. Tot., Total; P., plastids; N., nuclei. Experiments were repeated at three independent times with similar results. b-f, Compared with transient expression of GFP, transient expression of SlSAD8, but not SlSAD8ΔNLS, resulted in a delayed ripening initiation and impaired ethylene biosynthesis. b, Representative ripening phenotypes. Bars, 1 cm. c. Days from anthesis to breaker stage. d, Ethylene production at the Br stage. e, Relative transcript levels of SlACS2. f, Relative transcript levels of SlACS4. g-i, Transient expression of SlSAD8, but not SlSAD8ΔcTP, resulted in delayed color change (g), elevated chlorophyll accumulation (h), and reduced carotenoid levels (i) at the Br + 3 stage. Bars in g, 1 cm. In c, data are shown in a violin plot (n = 50), and differences were assessed using two-tailed t-test (***, P < 0.001; NS, no significant). In d-f, h, and i, data are shown as mean ± s.d. of three biological replicates, and differences were assessed using two-tailed t-test (***, P < 0.001; **, P < 0.01; *, P < 0.05; NS, no significant).
Extended Data Fig. 7 Homologues of SlSAD8 in three other fruit crops exhibited nuclear localization.
a, Sequence alignment of SlSAD8 and its homologues in three other fruit crops, including Actinidia eriantha, Musa acuminata, and Fragaria vesca. NLS, nuclear localization signal. Protein IDs and sequences were obtained from the NCBI database. b, Subcellular localization analysis of SlSAD8 homologues in three other fruit crops. AtHY5-mCherry was used as a reference for nuclear localization. Bars, 20 μm. Experiments were repeated at three independent times with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2.
Source data
Source Data Figs. 1, 3 and 4 and Extended Data Figs. 5 and 6 (download PDF )
Unprocessed western blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, C., Li, R., Chen, X. et al. Tomato ripening regulator SlSAD8 disturbs nuclear gene transcription and chloroplast-associated protein degradation. Nat. Plants 11, 2230–2239 (2025). https://doi.org/10.1038/s41477-025-02134-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02134-2