Abstract
Chorismate is a branch-point metabolite in the biosynthesis of aromatic amino acids, vitamins, antibiotics and various other aromatic products in bacteria, fungi and plants. Although 13 chorismate-utilizing enzymes have been identified in bacteria, only 6 have been described in plants, where an estimated 30% of all photosynthetically fixed carbon passes through chorismate. Here we describe a biosynthetic gene cluster (BGC) consisting of five core genes, including two reductases, two methyltransferases and one glucosyltransferase. Genetic and biochemical evidence shows that these five enzymes collectively give rise to three biosynthetic pathways, each originating from chorismate: two parallel pathways produce a class of non-aromatic, isomeric compounds abundant in the roots of Arabidopsis thaliana, whereas the third pathway produces methylated and glucosylated chorismate derivatives that subsequently react non-enzymatically with glutathione. Genome analysis revealed that variants of this BGC are present in some but not all species in the Brassicaceae family. Taken together, our study uncovered a BGC, containing three chorismate-utilizing enzymes, that controls three distinct post-chorismate pathways in A. thaliana. This work not only advances our understanding of carbon flow in this model plant but also highlights that the biochemical complexity encoded by plant BGCs is greater than previously appreciated.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data are provided with this paper. All other data needed to derive the conclusions in this paper are present in the paper and/or the Supplementary Information.
References
Maeda, H. & Dudareva, N. The shikimate pathway and aromatic amino acid biosynthesis in plants. Annu. Rev. Plant Biol. 63, 73–105 (2012).
Westfall, C. S., Xu, A. & Jez, J. M. Structural evolution of differential amino acid effector regulation in plant chorismate mutases. J. Biol. Chem. 289, 28619–28628 (2014).
Niyogi, K. K. & Fink, G. R. Two anthranilate synthase genes in Arabidopsis: defense-related regulation of the tryptophan pathway. Plant Cell 4, 721–733 (1992).
Lynch, J. H. & Dudareva, N. Aromatic amino acids: a complex network ripe for future exploration. Trends Plant Sci. 25, 670–681 (2020).
Tzin, V. & Galili, G. New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol. Plant 3, 956–972 (2010).
Basset, G. J. C. et al. Foliate synthesis in plants: the p-aminobenzoate branch is initiated by a bifunctional PabA-PabB protein that is targeted to plastids. Proc. Natl Acad. Sci. USA 101, 1496–1501 (2004).
Gross, J. et al. A plant locus essential for phylloquinone (vitamin K1) biosynthesis originated from a fusion of four eubacterial genes. J. Biol. Chem. 281, 17189–17196 (2006).
Verberne, M. C., Sansuk, K., Bol, J. F., Linthorst, H. J. M. & Verpoorte, R. Vitamin K1 accumulation in tobacco plants overexpressing bacterial genes involved in the biosynthesis of salicylic acid. J. Biotechnol. 128, 72–79 (2007).
Garcion, C. et al. Characterization and biological function of the ISOCHORISMATE SYNTHASE2 gene of Arabidopsis. Plant Physiol. 147, 1279–1287 (2008).
Wildermuth, M. C., Dewdney, J., Wu, G. & Ausubel, F. M. Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414, 562–565 (2001).
Rekhter, D. et al. Isochorismate-derived biosynthesis of the plant stress hormone salicylic acid. Science 365, 498–502 (2019).
Torrens-Spence, M. P. et al. PBS3 and EPS1 complete salicylic acid biosynthesis from isochorismate in Arabidopsis. Mol. Plant 12, 1577–1586 (2019).
Scholten, N. et al. In-depth analysis of isochorismate synthase-derived metabolism in plant immunity: identification of meta-substituted benzoates and salicyloyl-malate. J. Biol. Chem. 300, 107667 (2024).
Wu, J. et al. The cytosolic aminotransferase VAS1 coordinates aromatic amino acid biosynthesis and metabolism. Sci. Adv. 10, eadk0738 (2024).
Holland, C. K. et al. Brassicaceae-specific Gretchen Hagen 3 acyl acid amido synthetases conjugate amino acids to chorismate, a precursor of aromatic amino acids and salicylic acid. J. Biol. Chem. 294, 16855–16864 (2019).
Sundin, L. et al. Mutation of the inducible ARABIDOPSIS THALIANA CYTOCHROME P450 REDUCTASE2 alters lignin composition and improves saccharification. Plant Physiol. 166, 1956–1971 (2014).
Van de Wouwer, D. et al. Chemical genetics uncovers novel inhibitors of lignification, including p-iodobenzoic acid targeting CINNAMATE-4-HYDROXYLASE. Plant Physiol. 172, 198–220 (2016).
De Meester, B. et al. Vessel-specific reintroduction of CINNAMOYL-COA REDUCTASE1 (CCR1) in dwarfed ccr1 mutants restores vessel and xylary fiber integrity and increases biomass. Plant Physiol. 176, 611–633 (2018).
Oyarce, P. et al. Introducing curcumin biosynthesis in Arabidopsis enhances lignocellulosic biomass processing. Nat. Plants 5, 225–237 (2019).
Hoengenaert, L. et al. Overexpression of the scopoletin biosynthetic pathway enhances lignocellulosic biomass processing. Sci. Adv. 8, eabo5738 (2022).
Brouckaert, M. et al. QT–GWAS: a novel method for unveiling biosynthetic loci affecting qualitative metabolic traits. Mol. Plant 16, 1212–1227 (2023).
Mano, J. et al. Protection against photooxidative injury of tobacco leaves by 2-alkenal reductase: detoxication of lipid peroxide-derived reactive carbonyls. Plant Physiol. 139, 1773–1783 (2005).
Youn, B. et al. Mechanistic and structural studies of apoform, binary, and ternary complexes of the Arabidopsis alkenal double bond reductase At5g16970. J. Biol. Chem. 281, 40076–40088 (2006).
Zubieta, C. et al. Structural basis for substrate recognition in the salicylic acid carboxyl methyltransferase family. Plant Cell 15, 1704–1716 (2003).
Qu, L., Li, S. & Xing, S. Methylation of phytohormones by the SABATH methyltransferases. Chin. Sci. Bull. 55, 2211–2218 (2010).
Wittstock, U. & Halkier, B. A. Glucosinolate research in the Arabidopsis era. Trends Plant Sci. 7, 263–270 (2002).
Petersen, B. L., Chen, S., Hansen, C. H., Olsen, C. E. & Halkier, B. A. Composition and content of glucosinolates in developing Arabidopsis thaliana. Planta 214, 562–571 (2002).
Noctor, G. et al. Glutathione in plants: an integrated overview. Plant Cell Environ. 35, 454–484 (2012).
Mahanta, N., Fedoseyenko, D., Dairi, T. & Begley, T. P. Menaquinone biosynthesis: formation of aminofutalosine requires a unique radical SAM enzyme. J. Am. Chem. Soc. 135, 15318–15321 (2013).
Mahanta, N. et al. Menaquinone biosynthesis: biochemical and structural studies of chorismate dehydratase. Biochemistry 58, 1837–1840 (2019).
Togninalli, M. et al. AraPheno and the AraGWAS Catalog 2020: a major database update including RNA-Seq and knockout mutation data for Arabidopsis thaliana. Nucleic Acids Res. 48, D1063–D1068 (2020).
Polturak, G. & Osbourn, A. The emerging role of biosynthetic gene clusters in plant defense and plant interactions. PLoS Pathog. 17, e1009698 (2021).
Feng, G. et al. Jasmonate induced alternative splicing responses in Arabidopsis. Plant Direct 4, e00245 (2020).
Le Berre, J.-Y. et al. Transcriptome dynamic of Arabidopsis roots infected with Phytophthora parasitica identifies VQ29, a gene induced during the penetration and involved in the restriction of infection. PLoS ONE 12, e0190341 (2017).
Iven, T. et al. Transcriptional activation and production of tryptophan-derived secondary metabolites in Arabidopsis roots contributes to the defense against the fungal vascular pathogen Verticillium longisporum. Mol. Plant 5, 1389–1402 (2012).
Chen, Y. C. et al. Root defense analysis against Fusarium oxysporum reveals new regulators to confer resistance. Sci. Rep. 4, 5584 (2014).
Lyons, R. et al. Fusarium oxysporum triggers tissue-specific transcriptional reprogramming in Arabidopsis thaliana. PLoS ONE 10, e0121902 (2015).
Thieme, C. J. et al. Endogenous Arabidopsis messenger RNAs transported to distant tissues. Nat. Plants 1, 15025 (2015).
Nishida, S., Kakei, Y., Shimada, Y. & Fujiwara, T. Genome-wide analysis of specific alterations in transcript structure and accumulation caused by nutrient deficiencies in Arabidopsis thaliana. Plant J. 91, 741–753 (2017).
Vanholme, R. et al. COSY catalyses trans-cis isomerization and lactonization in the biosynthesis of coumarins. Nat. Plants 5, 1066–1075 (2019).
Strehmel, N., Böttcher, C., Schmidt, S. & Scheel, D. Profiling of secondary metabolites in root exudates of Arabidopsis thaliana. Phytochemistry 108, 35–46 (2014).
Chai, Y. N. & Schachtman, D. P. Root exudates impact plant performance under abiotic stress. Trends Plant Sci. 27, 80–91 (2022).
Zhan, C. et al. Plant metabolic gene clusters in the multi-omics era. Trends Plant Sci. 27, 981–1001 (2022).
Field, B. & Osbourn, A. E. Metabolic diversification—independent assembly of operon-like gene clusters in different plants. Science 320, 543–547 (2008).
Field, B. et al. Formation of plant metabolic gene clusters within dynamic chromosomal regions. Proc. Natl Acad. Sci. USA 108, 16116–16121 (2011).
Castillo, D. A., Kolesnikova, M. D. & Matsuda, S. P. T. An effective strategy for exploring unknown metabolic pathways by genome mining. J. Am. Chem. Soc. 135, 5885–5894 (2013).
Sohrabi, R. et al. In planta variation of volatile biosynthesis: an alternative biosynthetic route to the formation of the pathogen-induced volatile homoterpene DMNT via triterpene degradation in Arabidopsis roots. Plant Cell 27, 874–890 (2015).
Boutanaev, A. M. et al. Investigation of terpene diversification across multiple sequenced plant genomes. Proc. Natl Acad. Sci. USA 112, E81–E88 (2015).
Nützmann, H.-W., Huang, A. & Osbourn, A. Plant metabolic clusters—from genetics to genomics. N. Phytol. 211, 771–789 (2016).
Liu, H. et al. CRISPR-P 2.0: an improved CRISPR–Cas9 tool for genome editing in plants. Mol. Plant 10, 530–532 (2017).
Payne, R. J. et al. Synthesis and evaluation of 2,5-dihydrochorismate analogues as inhibitors of the chorismate-utilising enzymes. Org. Biomol. Chem. 7, 2421–2429 (2009).
Ife, R. J., Ball, L. F., Lowe, P. & Haslam, E. The shikimate pathway. Part V. Chorismic acid and chorismate mutase. J. Chem. Soc. Perkin Trans. 1, 1776–1783 (1976).
Mitschke, N. et al. Enantiopure 2,9-dideuterodecane—preparation and proof of enantiopurity. Eur. J. Org. Chem. 2021, 3854–3863 (2021).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Hendriks, K. P. et al. Global Brassicaceae phylogeny based on filtering of 1,000-gene dataset. Curr. Biol. 33, 4052–4068 (2023).
Hubrich, F., Müller, M. & Andexer, J. N. Chorismate- and isochorismate converting enzymes: versatile catalysts acting on an important metabolic node. Chem. Commun. 57, 2441–2463 (2021).
Winter, D. et al. An ‘Electronic Fluorescent Pictograph’ browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, e718 (2007).
Haroth, S. et al. The glycosyltransferase UGT76E1 significantly contributes to 12-O-glucopyranosyl-jasmonic acid formation in wounded Arabidopsis thaliana leaves. J. Biol. Chem. 294, 9858–9872 (2019).
Acknowledgements
We thank J. Pollier for the GC–MS analysis, M. Tonelli for managing to obtain far better HSQC spectra of the de5/5a samples than we were able to manage and A. Bleys for critically reading this manuscript. M.P. was supported by the Horizon 2020 Marie Skłodowska-Curie Individual Fellowships (grant no. 897918) and the FWO grant no. G0ACB25N. M.P. and W.B. acknowledge support from the advanced ERC project (POPMET). J.L., X.L. and A.L. were funded by the China Scholarship Council. B.D.M. and M.B. were funded by the Research Foundation – Flanders (FWO project G020618N and FWO personal PhD fellowship grant 1S38920N, respectively). Y.L., V.I.T. and J.R. were funded by the DOE Great Lakes Bioenergy Research Center, US Department of Energy, Office of Science, Biological and Environmental Research Program under Award No. DE-SC0018409. This study used the National Magnetic Resonance Facility at Madison, which is supported by NIH grant no. R24GM141526, and the 600 MHz instrument specifically, NIH S10RR023438. Helium recovery equipment, computers and infrastructure for the data archive were funded by the University of Wisconsin–Madison, NIH grant no. R24GM141526, and National Science Foundation NSF grant no. 1946970 (NSF Mid-Scale Research Infrastructure Big Idea).
Author information
Authors and Affiliations
Contributions
M.P., J.L., R.V. and W.B. designed the project. M.P., J.L., X.L., A.L., B.D.M., G.G., V.I.T. and J.R. performed the experiments. M.P., M.B., K.M., G.G., V.I.T., Y.L., J.R. and R.V. collected and analysed the data. M.P., R.V., J.R. and W.B. wrote the manuscript with the help of all of the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Zhenhua Liu, Jürgen Zeier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chorismate-utilizing enzymes (CUEs) in bacteria and plants.
(a) Thirteen CUEs in bacteria. Two major subtype chorismate mutases (AroQ-CM and AroH-CM) catalyze the same conversion of chorismate to prephenate. AroQ-CM is found in Escherichia coli, whereas AroH-CM is found in Bacillus subtilis. Four subtype chorismatases (CH-I, CH-II, CH-III, and CH-IV) have been characterized. CH-I, exemplified by FkbO in Streptomyces hygroscopicus subsp. ascomyceticus, converts chorismate into 3,4-dihydroxycyclohexa-1,5-diene-1-carboxylic acid (DCDC), whereas CH-II, such as Hyg5 from Streptomyces hygroscopicus subsp. hygroscopicus, catalyzes the conversion of chorismate into 3-hydroxybenzoate (3-HBA). CH-III, such as XanB2 from Xanthomonas campestris pv. campestris, is a bifunctional enzyme producing both 3-HBA and 4-hydroxybenzoate (4-HBA). CH-IV (SmCH-IV in Stenotrophomonas maltophilia) and chorismate pyruvate lyase (CPL, encoded by UbiC in E. coli) also convert chorismate into 4-HBA. Chorismate dehydratase (CDT), encoded by MqnA in Deinococcus radiodurans, converts chorismate into 3-enolpyruvyl-benzoate (3-EPB). For more information on these 13 CUEs, see56. AS, anthranilate synthase; ADCS, 4-amino-4-deoxychorismate (ADC) synthase; ADICS, 4-amino-4-deoxyisochorismate (ADIC) synthase; ICS, isochorismate synthase; SS, salicylate synthase; Trp, tryptophan; Phe, phenylalanine; Tyr, tyrosine. (b) Six CUEs in plants. AS, CM, ADCS and ICS are conserved in plants, whereas GH3.7 and GH3.12 are specific to the Brassicaceae family15. GH3.12 was originally found to be involved in the biosynthesis of salicylic acid by catalyzing the conjugation of isochorismate and glutamate11,12, but later shown to have comparable activities towards chorismate in vitro15. Both GH3.7 and GH3.12 show activity towards chorismate and produce multiple conjugate products in vitro. EPS1, enhanced pseudomonas susceptibility 1, encodes an isochorismoyl-glutamate pyruvoyl-glutamate lyase.
Extended Data Fig. 2 NMR spectra (600 MHz) of de5ae, 5ae, and 6ae (erythro isomers).
(a) Aromatic regions of HSQC NMR spectra (overlaid) for compounds de5ae and 6ae complete with high-resolution projections (for 1D 1H and 13C NMR spectra) if available; the carbon spectra were weak, and that of de5ae is from the low-resolution 2D-projection as it was not possible to obtain the 1D 13C NMR spectrum. (b) Oxygenated-aliphatic regions of NMR spectra (overlaid) for compounds de5ae, 5ae, and 6ae complete with high-resolution projections (for 1D 1H and 13C NMR spectra) if available; the carbon spectrum for 6ae was a DEPT-135 spectrum (CH2s inverted) and is weak, and that of de5ae was from the low-resolution 2D-projection as it was not possible to obtain the 1D 13C NMR spectrum.
Extended Data Fig. 3 NMR spectra (600 MHz) of de5at, 5at, and 6at (threo isomers).
(a) Aromatic regions of NMR spectra (overlaid) for compounds de5at and 6at complete with high-resolution 1D 1H projections; the carbon spectra are low-resolution 2D-projections as it was not possible to obtain the 1D 13C NMR spectra. (b) Aliphatic regions of NMR spectra (overlaid) for compounds de5at, 5at, and 6at complete with high-resolution 1D 1H projections; the carbon spectra are low-resolution 2D-projections as it was not possible to obtain the 1D 13C NMR spectra.
Extended Data Fig. 4 Expression profiles of seven candidate genes in A. thaliana.
(a) The heatmap was generated by using the microarray expression data (log2 transformed) from the eFP browser57. CURE3 expression data were not available. The order of the genes is according to their positions in the genome. (b) Transcript profiling of the candidate genes in four A. thaliana organs via qRT-PCR. Root and shoot organs were collected from 2-week-old seedlings vertically grown in vitro. Rosettes were collected from one-month-old plants grown in soil, and stem materials were harvested from mature plants. CURE3 transcripts were below the detection limit in all four organs. Data are presented as mean abundance ± SD of three biological replicates.
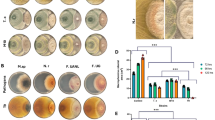
Extended Data Fig. 5 Characterization of knock-out and knock-down mutations in the candidate genes.
(a) Knock-out mutations generated via CRISPR/Cas9 gene editing. The red triangles indicate guide RNAs (gRNA1 and gRNA2), the sequences of which are shown in green in the WT sequence. The PAM site is underlined (NGG in this study). The mutations for each knock-out line are indicated in red and were confirmed by PCR-product sequencing. (b) T-DNA insertion mutant line (dcmt2-2). Primers p1, p2, and LB1 were used for genotyping (1# and 2# indicate two technical repeats, raw image in Supplementary Fig. 33), whereas primers p3 and p4 were used for qRT-PCR. Data are presented as mean ± SD, n = 3.
Extended Data Fig. 6 HSQC NMR spectra (500 MHz) of compounds 4a and 4b.
(a) HSQC spectrum of 4a. Two independent coupling networks, that in the 2,5-dihydrochorismate moiety C and that in the glucoside G are fully annotated. (b) HSQC spectrum of 4b. Two independent coupling networks, that in the 1,2-dihydrochorismate moiety C and that in the glucoside G, are fully annotated.
Extended Data Fig. 7 Characterization of cure1 cure2 cure3 triple mutants and their phenotypes.
(a) cure1 cure2 cure3 triple mutants generated via CRISPR/Cas9 gene editing. One guide RNA (red triangle) was used for each gene, the sequences of which are shown in green in the WT. The underlined bases indicate the PAM site (NGG in this study). The mutations within each CURE gene are indicated in red and were confirmed by PCR-product sequencing. (b) Root tip of cure1 cure2 cure3 triple mutants and WT at 5-day-old seedling stage, stained by propidium iodide and observed by confocal microscopy. One representative of five biological replicates is shown for each line. Scale bar = 20 µm. (c) Twelve-day-old cure1 cure2 cure3 triple mutants seedlings, vertically grown in vitro. Scale bar = 1 cm. (d-e) Primary root length (d) and lateral root density (e) for cure1 cure2 cure3 triple mutants and WT plants shown in (c). Data are shown as mean ± SD. The number of biological replicates is shown in the figure. P values were estimated by two-tailed Student’s t-test. (f) The above-ground part of cure1 cure2 cure3 triple mutants and WT plants grown in soil for one month. One representative of nine biological replicates is shown for each line. Scale bar = 1 cm. (g-h) The root systems of cure1 cure2 cure3 triple mutants and WT plants grown in soil for one month, side view (g) and bottom view (h), respectively. One representative of nine biological replicates is shown for each line (g) and three representatives of nine biological replicates are shown for each line (h). Scale bar = 1 cm.
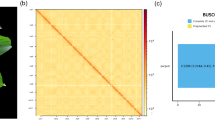
Extended Data Fig. 8 Searching for CURE1, CURE2, DCMT1, UGT76E9 and DCMT2 homologues in 35 plant species.
(a) Summary of homologues for each of the five enzymes obtained by protein BLAST. The homologous proteins and their phylogeny are shown in Supplementary Figs. 26–29. The cut-off identity values, that is, 70%, 60%, 65% and 60% identity for CURE1 and CURE2, DCMT1, UGT76E9, and DCMT2, respectively, are based on the phylogeny of the homologous proteins in A. thaliana (see b-e). Twenty-four other Brassicaceae species include Arabidopsis helleri (Ah), Alyssum linifolium (Alyli), Arabidopsis lyrata (AL), Brassica oleracea capitata (Bol), Brassica rapa (Brara), Boechera stricta (Bostr), Caulanthus amplexicaulis (Caamp), Capsella grandiflora (Cagra), Capsella rubella (Carub), Camelina sativa (Csa), Descurainia sophioides (Desop), Diptychocarpus strictus (Distr), Euclidium salsugineum (Thhalv), Euclidium syriacum (Eusyr), Iberis amara (Ibeam), Isatis tinctorial (Isati), Lunaria annua (Luann), Lepidium sativum (Lesat), Malcolmia maritima (Mamar), Myagrum perfoliatum (Myper), Rorippa islandica (Roisl), Schrenkiella parvula (Sp), Stanleya pinnata (Stapi), and Thlaspi arvense (Thlar). Other abbreviations used in the phylogenetic analysis include Cleome violacea (Clevi), Vitis vinifera (Vitiv), Zea mays (Zm), Oryza sativa (LOC). (b) Maximum likelihood (ML) tree of 11 CURE homologs in A. thaliana. Protein identity shared with CURE1 (left) and CURE2 (right) is shown. P: likely pseudogene. (c) Part of the reported ML tree of SABATH family proteins in A. thaliana25. Protein identity shared with DCMT1 is shown. (d) Part of the reported ML tree of UGT76E family proteins in A. thaliana58. (e) Part of the reported ML tree of SABATH family proteins in A. thaliana25. Protein identity shared with DCMT2 is shown.
Extended Data Fig. 9 Closer view of the clades of CURE1/2-, DCMT1-, UGT76E9- and DCMT2-like proteins.
Detailed view of the orange boxes in Supplementary Figs. 26–29 of the ML trees of CURE1/2 (a), DCMT1(b), UGT76E9 (c) and DCMT2 (d) homologous proteins. For abbreviations of plant species, see legend of Extended Data Fig. 8. The homologous proteins that form a potential BGC in a given plant species are marked by a unique symbol (color and shape). Two potential BGCs are found in A. linifolium (one accession) and are marked by purple stars 1 and 2. Four potential BGCs are found in I. tinctoria (one accession) and are marked by blue diamonds 1-4. * indicates that Isati.4971s0001.1 (located on scaffold 4971) likely forms a BGC with two adjacent members: CURE2-like and UGT76E9-like genes found in the scaffold 8058 (marked by diamond 4). From the current genome data, it is not clear whether scaffolds 4971 and 8058 are adjacent scaffolds.
Extended Data Fig. 10 MS data and a proposed biosynthetic pathway for compound 8.
(a) MS/MS spectrum of the in-source fragment (m/z 151.040) of 8. (b-c) Profiling of 8 in dcmt1 knockout mutants (n = 8 for WT, dcmt1-1 and dcmt1-2) (b) and ugt76e9 knockout mutants (n = 5 for WT, ugt76e9-1 and ugt76e9-2) (c). ND, not detected. Data are shown as mean ± SEM. P values were estimated by two-tailed Student’s t-test. (d) A proposed post-chorismate pathway for the biosynthesis of 8. A bacterial chorismate dehydratase is shown to convert chorismate into 3-enolpyruvyl-benzoate (3-EPB)29, but a plant gene for this reaction has not yet been identified. The dashed line indicates multiple steps, including glycosylation catalyzed by UGT76E9.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–33, Tables 1–9, Methods and additional references.
Supplementary Datasets 1–7 (download XLSX )
Untargeted metabolomics.
Source data
Source Data Figs. 1–3 and Extended Data Figs. 4, 5, 7, 8 and 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Peng, M., Li, J., Liu, X. et al. A biosynthetic gene cluster for three post-chorismate pathways in Arabidopsis. Nat. Plants 12, 205–216 (2026). https://doi.org/10.1038/s41477-025-02185-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02185-5