Abstract
Plant-parasitic nematodes are among the most destructive soil-dwelling pests, posing severe threats to global agriculture. However, the interplay between plant metabolites, rhizosphere microorganisms and their potential role in guiding pathogenic nematodes to their hosts remains poorly understood. Here we explored this gap by investigating the role of benzoxazinoids (BXs), a class of defensive metabolites of maize plants, in influencing the host-seeking behaviour of root-knot nematodes (RKNs). Our findings revealed that, surprisingly, BXs secreted by maize roots, particularly 6-methoxy-benzoxazolin-2-one, not only enhance RKN infection but also serve as powerful attractants. Remarkably, BX effects were observed only in the presence of a soil matrix. Further analysis demonstrated that 6-methoxy-benzoxazolin-2-one modulates the abundance and composition of rhizosphere bacteria, which in turn play a crucial role in RKN attraction and infection. We discovered that rhizosphere bacteria of BX-producing plants emit volatile compounds such as methyl ketones and 2-phenylethanol, which are then used by RKNs to locate host plants. RKNs detect these volatiles through chemosensory genes, including Mi-odr-1, Mi-odr-7 and Mi-gpa-6. Our study provides mechanistic insights into how RKNs use secondary-metabolite-shaped plant–microbe interactions to enhance their host-seeking behaviour and maximize their performance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw bacterial sequencing data (CRA030564, CRA030565, CRA030571, CRA030572 and CRA030576), genome sequences of P. pro and C. fre (GWHGQMN01000000 and GWHGQMO01000000) and gene sequences of FadE and PDh (C_AA120223.1 and C_AA120222.1) have been deposited in the Genome Sequence Archive at the National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, and are publicly accessible at https://ngdc.cncb.ac.cn/gsub. Source data are provided with this paper.
Code availability
The source code for the soil microbiota analysis is available via GitHub at https://github.com/YongxinLiu/EasyAmplicon/releases/tag/v1.12.
References
Coyne, D. L. et al. Plant-parasitic nematodes and food security in sub-Saharan Africa. Annu. Rev. Phytopathol. 56, 381–403 (2018).
Rutter, W. B., Franco, J. & Gleason, C. Rooting out the mechanisms of root-knot nematode–plant interactions. Annu. Rev. Phytopathol. 60, 43–76 (2022).
Siddique, S., Coomer, A., Baum, T. & Williamson, V. M. Recognition and response in plant–nematode interactions. Annu. Rev. Phytopathol. 60, 143–162 (2022).
Carrasco, D., Larsson, M. C. & Anderson, P. Insect host plant selection in complex environments. Curr. Opin. Insect Sci. 8, 1–7 (2015).
Silva, R. & Clarke, A. R. The ‘sequential cues hypothesis’: a conceptual model to explain host location and ranking by polyphagous herbivores. Insect Sci. 27, 1136–1147 (2020).
Johnson, S. N. & Nielsen, U. N. Foraging in the dark—chemically mediated host plant location by belowground insect herbivores. J. Chem. Ecol. 38, 604–614 (2012).
Hu, L. et al. Plant iron acquisition strategy exploited by an insect herbivore. Science 361, 694–697 (2018).
Arce, C. C. M. et al. Plant-associated CO2 mediates long-distance host location and foraging behaviour of a root herbivore. eLife 10, e65575 (2021).
Sikder, M. M. & Vestergard, M. Impacts of root metabolites on soil nematodes. Front. Plant Sci. 10, 1792 (2020).
Oota, M. et al. Identification of naturally occurring polyamines as root-knot nematode attractants. Mol. Plant 13, 658–665 (2020).
Tsai, A. Y. L. et al. Root-knot nematode chemotaxis is positively regulated by l-galactose sidechains of mucilage carbohydrate rhamnogalacturonan-I. Sci. Adv. 7, eabh4182 (2021).
Tsai, A. Y.-L. et al. Regulation of root-knot nematode behavior by seed-coat mucilage-derived attractants. Mol. Plant 12, 99–112 (2019).
Topalovic, O. & Vestergard, M. Can microorganisms assist the survival and parasitism of plant-parasitic nematodes? Trends Parasitol. 37, 947–958 (2021).
La, S. et al. Protective role of native root-associated bacterial consortium against root-knot nematode infection in susceptible plants. Nat. Commun. 15, 6723 (2024).
Zheng, X. et al. Rhizosphere Streptomyces confers dual-mode resistance to root-knot nematodes through nematicidal metabolites and JA-mediated immunity in maize. Soil Ecol. Lett. 7, 250363 (2025).
Sikder, M. M. et al. Genetic disruption of Arabidopsis secondary metabolite synthesis leads to microbiome-mediated modulation of nematode invasion. ISME J. 16, 2230–2241 (2022).
Hou, S. et al. A microbiota–root–shoot circuit favours Arabidopsis growth over defence under suboptimal light. Nat. Plants 7, 1078–1092 (2021).
Korenblum, E. & Aharoni, A. Phytobiome metabolism: beneficial soil microbes steer crop plants’ secondary metabolism. Pest Manage. Sci. 75, 2378–2384 (2019).
Robert, C. A. M. & Mateo, P. The chemical ecology of benzoxazinoids. CHIMIA 76, 928–938 (2022).
Hu, L. et al. Soil chemistry determines whether defensive plant secondary metabolites promote or suppress herbivore growth. Proc. Natl Acad. Sci. USA 118, e2109602118 (2021).
Duan, S. et al. Integrated transcriptome and metabolite profiling highlights the role of benzoxazinoids in wheat resistance against Fusarium crown rot. Crop J. 10, 407–417 (2022).
Hu, L. et al. Root exudate metabolites drive plant–soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat. Commun. 9, 2738 (2018).
Cotton, T. E. A. et al. Metabolic regulation of the maize rhizobiome by benzoxazinoids. ISME J. 13, 1647–1658 (2019).
Kudjordjie, E. N., Sapkota, R., Steffensen, S. K., Fomsgaard, I. S. & Nicolaisen, M. Maize synthesized benzoxazinoids affect the host associated microbiome. Microbiome 7, 59 (2019).
Cadot, S. et al. Specific and conserved patterns of microbiota-structuring by maize benzoxazinoids in the field. Microbiome 9, 103 (2021).
Thoenen, L. et al. Bacterial tolerance to host-exuded specialized metabolites structures the maize root microbiome. Proc. Natl Acad. Sci. USA 120, e2310134120 (2023).
Kong, C. H. et al. Plant neighbor detection and allelochemical response are driven by root-secreted signaling chemicals. Nat. Commun. 9, 3867 (2018).
Gfeller, V. et al. Plant secondary metabolite-dependent plant–soil feedbacks can improve crop yield in the field. eLife 12, e84988 (2023).
Frew, A., Powell, J. R., Glauser, G., Bennett, A. E. & Johnson, S. N. Mycorrhizal fungi enhance nutrient uptake but disarm defences in plant roots, promoting plant-parasitic nematode populations. Soil Biol. Biochem. 126, 123–132 (2018).
Zasada, I. A., Meyer, S. L., Halbrendt, J. M. & Rice, C. Activity of hydroxamic acids from Secale cereale against the plant-parasitic nematodes Meloidogyne incognita and Xiphinema americanum. Phytopathology 95, 1116–1121 (2005).
Sikder, M. M. et al. Benzoxazinoids selectively affect maize root-associated nematode taxa. J. Exp. Bot. 72, 3835–3845 (2021).
Cheng, W. et al. Volatile organic compounds from Paenibacillus polymyxa KM2501-1 control Meloidogyne incognita by multiple strategies. Sci. Rep. 7, 16213 (2017).
Wang, P. et al. Chemotactic responses of the root-knot nematode Meloidogyne incognita to Streptomyces plicatus. FEMS Microbiol. Lett. 366, fnz234 (2019).
Shum, A. & Markovetz, A. Specificity and induction of undecyl acetate esterase from Pseudomonas cepacia grown on 2-tridecanone. J. Bacteriol. 118, 890–897 (1974).
Sakai, M. et al. Production of 2-phenylethanol in roses as the dominant floral scent compound from L-phenylalanine by two key enzymes, a PLP-dependent decarboxylase and a phenylacetaldehyde reductase. Biosci. Biotechnol. Biochem. 71, 2408–2419 (2014).
Vázquez, M. B., Matencio, A., Bianchinotti, M. V., García-Carmona, F. & López-Nicolás, J. M. Enhanced production of 2-phenylethanol by salicylic acid and cyclodextrins in cell suspension cultures of the unexplored filamentous fungus Monochaetinula geoffroeana. J. Sci. Food Agric. 102, 1609–1618 (2021).
Goh, E.-B., Baidoo, E. E. K., Keasling, J. D. & Beller, H. R. Engineering of bacterial methyl ketone synthesis for biofuels. Appl. Environ. Microbiol. 78, 70–80 (2012).
Beltrametti, F. et al. Sequencing and functional analysis of styrene catabolism genes from Pseudomonas fluorescens ST. Appl. Environ. Microbiol. 63, 2232–2239 (1997).
Li, Y., Ren, Q., Bo, T., Mo, M. & Liu, Y. AWA and ASH homologous sensing genes of Meloidogyne incognita contribute to the tomato infection process. Pathogens 11, 1322 (2022).
Shivakumara, T. N. et al. Homologs of Caenorhabditis elegans chemosensory genes have roles in behavior and chemotaxis in the root-knot nematode Meloidogyne incognita. Mol. Plant Microbe Interact. 32, 876–887 (2019).
Meyer, S. L. F., Rice, C. P. & Zasada, I. A. DIBOA: fate in soil and effects on root-knot nematode egg numbers. Soil Biol. Biochem. 41, 1555–1560 (2009).
Liu, X.-L. et al. The molecular basis of host selection in a crucifer-specialized moth. Curr. Biol. 30, 4476–4482. e4475 (2020).
Jiang, Q. et al. Two leucine-rich repeat receptor-like kinases initiate herbivory defense responses in tea plants. Hortic. Res. 12, uhae281 (2025).
Thoenen, L. et al. The lactonase BxdA mediates metabolic specialisation of maize root bacteria to benzoxazinoids. Nat. Commun. 15, 6535 (2024).
Duran, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983. e14 (2018).
Zhang, J. et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat. Biotechnol. 37, 676–684 (2019).
Tytgat, B. et al. Polar lake microbiomes have distinct evolutionary histories. Sci. Adv. 9, eade7130 (2023).
Neu, A. T., Allen, E. E. & Roy, K. Defining and quantifying the core microbiome: challenges and prospects. Proc. Natl Acad. Sci. USA 118, e2104429118 (2021).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Huang, A. C. et al. A specialized metabolic network selectively modulates Arabidopsis root microbiota. Science 364, eaau6389 (2019).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Johnson, J. S. et al. Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 10, 5029 (2019).
Lynch, M. D. J. & Neufeld, J. D. Ecology and exploration of the rare biosphere. Nat. Rev. Microbiol. 13, 217–229 (2015).
Han, G. & Vaishnava, S. Microbial underdogs: exploring the significance of low-abundance commensals in host–microbe interactions. Exp. Mol. Med. 55, 2498–2507 (2023).
Jousset, A. et al. Where less may be more: how the rare biosphere pulls ecosystems strings. ISME J. 11, 853–862 (2017).
Li, B. et al. Chemotaxis of Meloidogyne incognita response to rhizosphere bacteria. Microorganisms 11, 2271 (2023).
Barbosa, P. et al. Nematicidal activity of phytochemicals against the root-lesion nematode Pratylenchus penetrans. Plants 13, 726 (2024).
Zhai, Y. et al. Multiple modes of nematode control by volatiles of Pseudomonas putida 1a00316 from Antarctic soil against Meloidogyne incognita. Front. Microbiol. 9, 253 (2018).
Honda, K., Ômura, H. & Hayashi, N. Identification of floral volatiles from Ligustrum japonicum that stimulate flower-visiting by cabbage butterfly, Pieris rapae. J. Chem. Ecol. 24, 2167–2180 (1998).
Imai, T., Maekawa, M., Tsuchiya, S. & Fujimori, T. Field attraction of Hoplia communis to 2-phenylethanol, a major volatile component from host flowers, Rosa spp. J. Chem. Ecol. 24, 1491–1497 (1998).
Roy, B. A. & Raguso, R. A. Olfactory versus visual cues in a floral mimicry system. Oecologia 109, 414–426 (1997).
Niu, Q. et al. A Trojan horse mechanism of bacterial pathogenesis against nematodes. Proc. Natl Acad. Sci. USA 107, 16631–16636 (2010).
Ali, Q. et al. High killing rate of nematode and promotion of rice growth by synthetic volatiles from Bacillus strains due to enhanced oxidative stress response. Physiol. Plant. 175, e13868 (2023).
Deng, X., Wang, X. & Li, G. Nematicidal effects of volatile organic compounds from microorganisms and plants on plant-parasitic nematodes. Microorganisms 10, 1201 (2022).
Pender, C. L. et al. Perception of a pathogenic signature initiates intergenerational protection. Cell 188, 594–605.e510 (2025).
Schulz-Bohm, K., Martín-Sánchez, L. & Garbeva, P. Microbial volatiles: small molecules with an important role in intra- and inter-kingdom interactions. Front. Microbiol. 8, 2484 (2017).
Hu, L. et al. Herbivory-induced green leaf volatiles increase plant performance through jasmonate-dependent plant–soil feedbacks. Nat. Plants 11, 1001–1017 (2025).
Ruan, W., Zhan, L., Xiao, W. & Chen, S. An improved method for quantification of Heterodera glycines in plant tissues. Nematropica 40, 237–244 (2012).
Zhang, H. et al. Cover crop rotation suppresses root-knot nematode infection by shaping soil microbiota. New Phytol. 245, 363–377 (2025).
Ye, M. et al. A constitutive serine protease inhibitor suppresses herbivore performance in tea (Camellia sinensis). Hortic. Res. 10, uhad178 (2023).
Cesarz, S., Eva Schulz, A., Beugnon, R. & Eisenhauer, N. Testing soil nematode extraction efficiency using different variations of the Baermann-funnel method. Soil Org. 91, 61–72 (2019).
Bargmann, C. I., Hartwieg, E. & Horvitz, H. R. Odorant-selective genes and neurons mediate olfaction in C. elegans. Cell 74, 515–527 (1993).
Zhao, J. et al. A method for estimating nematode body lengths for use in the calculation of biomass via a simplified formula. Soil Biol. Biochem. 134, 36–41 (2019).
Tanwir, F., Dionisio, G., Adhikari, K. B., Fomsgaard, I. S. & Gregersen, P. L. Biosynthesis and chemical transformation of benzoxazinoids in rye during seed germination and the identification of a rye Bx6-like gene. Phytochemistry 140, 95–107 (2017).
Guo, D. et al. Linalool-triggered plant–soil feedback drives defense adaptation in dense maize plantings. Science 389, adv6675 (2025).
Liu, Y. X. et al. A practical guide to amplicon and metagenomic analysis of microbiome data. Protein Cell 12, 315–330 (2021).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahe, F. VSEARCH: a versatile open source tool for metagenomics. Peer J. 4, e2584 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Andrew S. et al. A quality control analysis tool for high throughput sequencing data. GitHub https://github.com/s-andrews/FastQC (2018).
Edgar, R. C. & Flyvbjerg, H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 31, 3476–3482 (2015).
Cole, J. R. et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642 (2014).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. In Proc. of the International AAAI Conference on Web and Social Media Vol. 3, 361–362 (AAAI Press, 2009).
Chen, W. et al. Volatile organic compounds from Bacillus aryabhattai MCCC 1K02966 with multiple modes against Meloidogyne incognita. Molecules 27, 103 (2021).
Jang, J. Y. et al. Biological control of Meloidogyne incognita by Aspergillus niger F22 producing oxalic acid. PLoS ONE 11, e0156230 (2016).
Gfeller, V. et al. Root volatiles in plant–plant interactions I: high root sesquiterpene release is associated with increased germination and growth of plant neighbours. Plant Cell Environ. 42, 1950–1963 (2019).
Ye, M. et al. Indole primes defence signalling and increases herbivore resistance in tea plants. Plant Cell Environ. 44, 1165–1177 (2021).
Li, X. et al. The outer membrane protein OprF and the sigma factor SigX regulate antibiotic production in Pseudomonas fluorescens 2P24. Microbiol. Res. 206, 159–167 (2018).
R v.4.2.0 (R Foundation for Statistical Computing, 2025).
Bates, D., Machler, M., Bolker, B. M. & Walker, S. C. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
car (R Foundation for Statistical Computing, 2012).
emmeans (R Foundation for Statistical Computing, 2021).
multcomp (R Foundation for Statistical Computing, 2016).
RVAideMemoire (R Foundation for Statistical Computing, 2023).
Acknowledgements
We thank J. M. Raaijmakers (Netherlands Institute of Ecology) for his insightful comments and input on the microbial work. This work was supported by the Zhejiang Provincial Natural Science Foundation of China (grant no. LR25D010001), the National Key Research and Development Project of China (grant no. 2021YFD1900200), the National Natural Science Foundation of China (grant no. 42377285), the Science and Technology special fund of Hainan Province (grant no. ZDYF2024XDNY161), the Fundamental Research Funds for the Central Universities (grant no. 226-2025-00049), the 111 Project (grant no. B17039), the China Agriculture Research System (grant no. CARS-01) and China Agriculture Research System of MOF and MARA (grant no. CARS-04).
Author information
Authors and Affiliations
Contributions
L.H. conceived of, designed and supervised the study. Z.W., Z.L., J.L.L.-T., J.X. and L.H. performed the experiments. Z.W., Z.L., W.W., S.Z., L.Z., S.C., X.L., M.E. and L.H. collected and analysed the data. L.H. wrote the initial draft of the paper. Z.W., S.S., A.Y.-L.T., J.L.L.-T. and L.H. revised the paper. All authors discussed the results and approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Klaus Schlaeppi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 BXs promote the growth and infection of RKNs.
a, b, The width (a) and weight (b) of newly hatched J2s from WT and bx1 plants. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks denote significant differences between treatments (two-sided Student’s t-test, **P < 0.01; ***P < 0.001). c, Gall numbers on bx1 plants infected by J2s hatched from WT and bx1 plants. d, Number of J2 hatched from bx1 plants infected by J2s originated from WT and bx1 plants. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, ***P < 0.001). e, Number of newly hatched J2s from WT and bx1 maize roots grown in soils collected from four regions in China, representing Alfisols, Oxisols, Mollisols, and Aridisols. WT plants consistently exhibited significantly more nematode reproduction than bx1 plants across all soil types. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, ***P < 0.001). f, Chemotactic behaviors of J2s choosing between bx1 and WT plants at different time points. Average results are shown in Fig. 1g. Data are presented as mean + s.e.m. There are five biological replicates for each treatment. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, *P < 0.05; ***P < 0.001). g, Gall numbers on W22 and bx1/W22 plants at ten- and fourteen-days post-inoculation. h, Number of hatched J2s from W22 and bx1/W22 plants. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. NS, not significant. Asterisks indicate significant differences between genotypes (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, *P < 0.05; **P < 0.01; ***P < 0.001). i, Chemotactic response of J2 RKNs toward MBOA in unsterilized and sterilized soils. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, ***P < 0.001). j, Shannon, Simpson and Pielou indexes of the bacteria from wild-type (WT) and bx1 rhizospheres. Horizontal lines within boxes represent medians. Tops and bottoms of boxes represent the 75th and 25th percentiles, respectively. The exact number of biological replicates is indicated above the X-axis. Data points represent individual replicates. The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Effects of individual bacteria strains on RKN chemotaxis, mortality and infection.
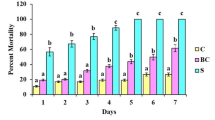
a, Number of J2s migrated toward bacterial cells or respective controls. b, c, Survival rate of J2s treated with different bacterial cells (b) and supernatant (c). Bacterial culture medium or sterile water served as a control. d, Gall numbers on bx1 plants after inoculation with individual bacteria strains. Data are presented as mean ± s.d. The exact number of biological replicates is indicated above the X-axis. Data points represent individual replicates. Asterisks indicate significant differences between the bacteria and control treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, *P < 0.05; **P < 0.01; ***P < 0.001). The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 MBOA promotes RKN attraction across non-BX-producing plant species.
a, Number of the bacteria CFUs migrated toward MBOA (2 μg ml−1). Data are presented as mean ± s.d. There were six biological replicates for each treatment. Data points represent individual replicates. Asterisks indicate significant differences between the bacteria and control treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, *P < 0.05; **P < 0.01; ***P < 0.001). b, c, Chemotactic responses of J2 RKNs toward barley (b) and rice (c) plants with or without MBOA supplementation. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between genotypes (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, ***P < 0.001). d, e, Unconstrained PCoA of rhizosphere bacterial communities in barley (d) or rice (e) following supplementation with synthetic MBOA. There are eight biological replicates for barley and ten biological replicates for rice. Data points represent individual replicates. The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Impacts of synthetic bacterial VOCs on RKN chemotaxis and mortality.
a, Effect of synthetic bacterial VOCs, 1-undecanol, 1-tridecene, 2-undecanone, 2-tridecanone, dimethyl disulfide, 2-phenylethanol, 2-heptanone, and 2-nonanone and 2-decanone, on RKN chemotaxis. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, *P < 0.05; **P < 0.01; ***P < 0.001). b, Survival rate of J2s after treatment with different synthetic bacterial VOCs, including 1-undecanol, 1-tridecene, 2-undecanone, 2-tridecanone, dimethyl disulfide, 2-phenylethanol, 2-heptanone, and 2-nonanone and 2-decanone. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by multiple comparisons of FDR-corrected LSMeans, *P < 0.05; ***P < 0.001). Asterisks indicate overall ANOVA significance (*P < 0.05; **P < 0.01; ***P < 0.001). Exact P values for pairwise comparisons between selected groups are indicated in the figure (two-sided Student’s t-test). NS, not significant. The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Effects of 2-phenylethanol induction on RKN chemotaxis.
a, The released amount of 1-undecanol from P. pro after inhibitor PMSF treatment. b, The released amount of 2-phenylethanol from C. fre after iproniazid treatment. Data are presented as mean ± s.d. The exact number of biological replicates is indicated above the X-axis. Data points represent individual replicates. Asterisks indicate significant differences between the bacteria and control treatments (two-sided Student’s t-test, *P < 0.05; ***P < 0.001). c, d, GC/MS selected ion chromatograms of L-Phe (c) or SA (d)-treated C. fre. e, f, 2-Phenylethanol content released from L-Phe (e) or SA (f)-treated C. fre. Data are presented as mean ± s.d. The exact number of biological replicates is indicated above the X-axis. Data points represent individual replicates. Asterisks indicate significant differences between the bacteria and control treatments (two-sided Student’s t-test, *P < 0.05; **P < 0.01). g, h, J2 RKN chemotactic behavior toward L-Phe (e) or SA (f)-treated C. fre. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, *P < 0.05; **P < 0.01; ***P < 0.001). The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Nucleotide and deduced amino acid sequences as well as predicted domains of FadE gene in Pseudomonas prosekii.
a, Schematic representation of FadE domain composition and organization based on conserved domain analysis. Numbers indicate the positions of each domain. b, Nucleotide and the deduced amino acid sequence of FadE.
Extended Data Fig. 7 P. proΔfadE volatile profiles and RKN attraction.
a, Schematic diagram of the P. pro FadE loci with the deleted regions. Scale bar represents 100 bp. b–d, GC/MS selected ion chromatograms of 1-undecanol (b), 2-undecanone (c), 2-tridecanone (d) released from P. proΔfadE. e–g, Levels of 1-undecanol (e), 2-undecanone (f), 2-tridecanone (g) secreted from P. proΔfadE. Data are presented as mean ± s.d. The exact number of biological replicates is indicated above the X-axis. Data points represent individual replicates. Asterisks indicate significant differences between the bacteria and control treatments (two-sided Student’s t-test, *P < 0.05; ***P < 0.001). h, J2 chemotactic behaviors toward P. proΔfadE. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, **P < 0.01; ***P < 0.001). The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 7.
Extended Data Fig. 8 Nucleotide and deduced amino acid sequences as well as predicted domains of PDh gene in Citrobacter freundii strain2.
a, Schematic representation of PDh domain composition and organization based on conserved domain analysis. Numbers indicate the positions of each domain. b, Nucleotide and the deduced amino acid sequence of PDh.
Extended Data Fig. 9 C. freΔpdh volatile profiles and RKN attraction.
a, Schematic diagram of the C. fre PDh loci with deleted regions. Scale bar represents 100 bp. b, GC/MS selected ion chromatograms of 2-phenylethanol released from C. freΔpdh. c, Levels of 2-phenylethanol secreted from C. freΔpdh. Data are presented as mean ± s.d. The exact number of biological replicates is indicated above the X-axis. Data points represent individual replicates. L.O.D., limit of detection. d, J2 chemotactic behaviors toward C. freΔpdh. Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (ANOVA followed by pairwise comparisons of FDR-corrected LSMeans, ***P < 0.001). The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 9.
Extended Data Fig. 10 Expression of chemosensory genes in RKNs exposed to bacterial or synthetic volatiles.
a, b, Expression levels of twelve chemosensory-associated genes in RKNs exposed to volatiles emitted by P. pro (a) and C. fre (b), as well as their respective mutant strains, P. proΔfadE and C. freΔpdh. c, d, Relative expression levels of twelve chemosensory-associated genes in RKNs following exposure to synthetic 1-undecanol (c) or 2-phenylethanol (d). Data are presented as mean + s.e.m. The exact number of biological replicates is indicated on each bar. Data points represent individual replicates. Asterisks indicate significant differences between treatments (two-sided Student’s t-test, *P < 0.05, **P < 0.01, ***P < 0.001). The raw data and exact P values for all comparisons in this figure are provided in Source Data Extended Data Fig. 10.
Supplementary information
Supplementary Data 1 (download XLSX )
Microbiome statistics. This file contains lists of the taxonomies, sequences and statistical details of the differentially abundant bacterial OTUs in the rhizospheres of WT and bx1 plants.
Supplementary Data 2 (download XLSX )
Sequences of 112 isolated bacteria strains.
Supplementary Data 3 (download XLSX )
Sequences of 28 selected bacteria strains.
Supplementary Data 4 (download XLSX )
The growth of different bacteria strains after MBOA treatment.
Supplementary Data 5 (download XLSX )
Root VOCs of WT and bx1 maize plants.
Supplementary Data 6 (download XLSX )
Peak areas of volatiles released by different bacteria.
Supplementary Data 7 (download XLSX )
Primers used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Z., Liu, Z., Wang, W. et al. Root-knot nematode Meloidogyne incognita uses secondary-metabolite-mediated soil microbiome shifts to locate host plants. Nat. Plants 12, 337–355 (2026). https://doi.org/10.1038/s41477-025-02205-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02205-4