Abstract
Small interfering RNAs (siRNAs) play a crucial role in plant reproduction, yet their mobility and function remain incompletely understood. We report that a large proportion of siRNAs found in pollen of Capsella rubella relies on mobile siRNAs from maternal sporophytic tissues, highlighting the importance of non-cell-autonomous siRNAs in male gametophyte development. Unlike tapetal siRNAs, which guide DNA methylation and require CLASSY3 and DNA-dependent RNA polymerase IV (Pol IV) activity in the tapetum, we found that Pol IV-dependent mobile siRNAs (PMsiRNAs) mainly function post-transcriptionally and do not guide DNA methylation. Nevertheless, PMsiRNAs share key features with tapetal siRNAs, including Pol IV dependency, clustering and a size range of 21–24 nucleotides. Using a grafting approach, we show that sporophytic Pol IV-dependent siRNAs act as non-cell-autonomous mobile signals that trigger PMsiRNA formation through post-transcriptional gene silencing. This process parallels reproductive phased siRNA biogenesis, which is widespread across angiosperms but has been considered absent in Brassicaceae. Loss of PMsiRNAs causes pollen arrest, underscoring their essential role. Together, these findings highlight siRNAs as long-distance communication signals from maternal sporophytic tissues to the male gametophyte with critical functions in developmental regulation.
Similar content being viewed by others
Main
In plants, the systemic movement of small RNAs (sRNAs), including siRNAs and microRNAs (miRNAs), represents a mechanism for intercellular communication and regulation1. These molecules can traverse long distances and influence gene expression and development in distal tissues2,3. Despite extensive evidence for this systemic transport, the biological impacts of endogenous mobile sRNAs remain poorly characterized, with functional roles attributed to only a limited subset of these molecules.
Transgene-mediated RNA silencing studies have provided important insights into the systemic nature of sRNA mobility. Transgene-induced post-transcriptional gene silencing (PTGS) induced at one site in the plant can systemically move through vascular tissues to silence homologous sequences in distant cells4,5. The results of grafting studies strongly implicate siRNAs as mobile signals of transgene-triggered gene silencing2,6. Systemic silencing of transgenes was shown to involve two RNA silencing pathways. In the PTGS pathway, the primary siRNA derived from a specific transcript triggers the production of secondary siRNAs resembling phased secondary siRNAs (phasiRNAs). This pathway produces 21- and 22-nucleotide siRNAs and involves ARGONAUTE 1 (AGO1), SUPPRESSOR OF GENE SILENCING 3 (SGS3), RNA-DEPENDENT RNA POLYMERASE 6 (RDR6), DICER-LIKE 2 (DCL2) and DICER-LIKE 4 (DCL4)7. The second pathway involves the nuclear components Pol IV and RNA-DEPENDENT RNA POLYMERASE 2 (RDR2), DICER-LIKE 3 (DCL3) and ARGONAUTE 4 (AGO4)8,9,10. This pathway generates 23- and 24-nucleotide siRNAs. The latter directs RNA-dependent DNA methylation (RdDM) in the transcriptional gene silencing pathway11.
Grafting experiments have shown that 24-nucleotide siRNAs can move bidirectionally and induce silencing in recipient tissues2,4,8,12. Silencing involving DNA methylation was predominantly observed for siRNAs moving from shoots to roots2,12. By contrast, systemic silencing by siRNAs travelling from roots to shoots does not consistently involve DNA methylation8,13, suggesting that these mobile siRNAs can mediate gene silencing through PTGS mechanisms.
Mobile siRNAs moving from shoots to roots were shown to constitute a substantial proportion of sRNAs in recipient tissues2,3,12, suggesting that long-distance transport of sRNAs serves to integrate environmental and developmental cues throughout the plant. Conversely, the magnitude and impact of root-to-shoot sRNA transport are less well-defined. Recent data suggest that root-derived siRNAs can induce heritable changes of DNA methylation in the progeny14, implicating an extensive bidirectional transport of sRNAs with possibly heritable consequences.
Reproductive phasiRNAs of 21 and 24 nucleotides accumulate abundantly during male reproduction in many angiosperms15,16,17,18,19. In grasses such as maize and rice, they occur in two principal waves: premeiotic 21-nucleotide phasiRNAs initiated by miR2118 and processed by DCL4, which can direct cis cleavage of intergenic, long non-coding genomic PHAS precursors and trans cleavage of protein-coding transcripts15,16,18; and meiotic 24-nucleotide phasiRNAs triggered by miR2275 and generated by DCL5, which are required to maintain CHH DNA methylation in cis at their source loci17. A third set of premeiotic 24-nucleotide phasiRNAs has also been described that shares genomic origins with meiotic 24-nucleotide phasiRNAs but lacks miRNA triggers and does not mediate PHAS precursor cleavage19. These reproductive phasiRNAs are thought to be produced in the tapetum and to move to meiocytes, where they regulate meiotic progression16,17,18,20. In Arabidopsis, although canonical reproductive phasiRNAs are largely absent, a pattern considered typical of Brassicaceae19, 24-nucleotide siRNAs produced by DCL3 and RDR2 in the tapetum induce DNA methylation in meiocytes, underscoring a conserved role for mobile siRNAs in male germline development21.
Here we report the systemic movement of endogenous siRNAs from roots to shoots and their critical roles in pollen development. We used a Pol IV-deficient mutant in Capsella rubella, carrying a mutation in the largest subunit of Pol IV, NRPD122. In Capsella, loss of NRPD1 causes pollen arrest at the microspore stage22, revealing an essential role of Pol IV-dependent siRNAs in pollen development. Here we demonstrate that mobile siRNAs originating in wild-type roots move to the nrpd1 shoot and restore pollen viability. Those siRNAs function post-transcriptionally and induce the production of a large proportion of all pollen siRNAs. This process parallels reproductive phasiRNA biogenesis, with Pol IV-derived siRNAs acting as a non-cell-autonomous trigger. Our study emphasizes the importance of siRNAs as agents of non-cell-autonomous communication and developmental regulation.
Results
Grafting-mediated rescue of nrpd1 pollen defects
Using CRISPR–Cas9, we previously generated a knockout mutant in Capsella rubella NRPD1, in which the induced deletion caused a frameshift and resulted in a truncated protein without the catalytic site22. Loss of Capsella NRPD1 causes pollen arrest at the microspore stage, connected with depletion of 21- to 24-nucleotide siRNAs in microspores22. To test whether the arrest of microspore development in Capsella nrpd1 mutants could be suppressed by non-cell-autonomous siRNAs, we made hypocotyl grafts of Capsella nrpd1 scions to wild-type (wt) rootstocks (referred to as nrpd1s/wtr) and analysed the effect on pollen development and siRNA formation (Fig. 1a).
a, Scheme of the experimental design for hypocotyl grafting. b, Percentage of mature pollen dissected from anthers at stage 12–13 from wts/wtr, wts/nrpd1r, nrpd1s/nrpd1r and nrpd1s/wtr. Three biological replicates derived from independent plants were analysed. Technical replicates in each biological replicate are denoted by triangles, squares and circles. Statistical significance was assessed using a beta-binomial generalized linear model, followed by Tukey-adjusted pairwise comparisons, two-sided. The asterisks mark statistically significant differences (***P < 0.001). The exact P values are provided in the source data. c,d,f,g,i,j,l,m, Alexander staining testing the viability of mature pollen after grafting wts/wtr (c,d), wts/nrpd1r (f,g), nrpd1s/nrpd1r (i,j) and nrpd1s/wtr (l,m). Scale bars, 100 µm. Arrows indicate aborted pollen. The experiment was repeated twice. e,h,k,n, Seed number per silique in wts/wtr (e), wts/nrpd1r (h), nrpd1s/nrpd1r (k) and nrpd1s/wtr (n). Scale bars, 1 mm. o, Seed number per silique of indicated genotypes. Numbers are based on four biological replicates from individual plants per genotype. Nine or ten siliques were analysed for each plant. Siliques from each replicate are denoted by triangles, squares, circles and crosses. The asterisks mark statistically significant differences (***P < 0.05), Kruskal–Wallis test followed by pairwise Mann–Whitney U-tests with Holm–Bonferroni correction for multiple comparisons. The exact P value is provided in the source data. p, Seed number per silique derived from crossing wt plants with pollen of grafted plants of the indicated genotypes. Seed numbers based on two biological replicates from individual plants per genotype (***P < 0.05), Kruskal–Wallis test followed by pairwise Mann–Whitney U-tests with Holm–Bonferroni correction for multiple comparisons. The exact P value is provided in the source data. In b, o and p, centre lines show the medians; box limits indicate the 25th and 75th percentiles; and whiskers extend 1.5× the interquartile range from the 25th and 75th percentiles. ♀, female; ♂, male; NS, not significant.
As expected, nrpd1s/nrpd1r grafts exhibited a reduced amount of mature pollen compared with wts/wtr and wts/nrpd1r grafts, consistent with the previously reported pollen defect in the Capsella nrpd1 mutant (Fig. 1b–d,f,g,i,j)22. Strikingly, we observed a significant increase in mature pollen in nrpd1s/wtr compared with control nrpd1s/nrpd1r grafts (Fig. 1b,i,j,l,m), which was reflected by a significantly higher seed set in nrpd1s/wtr grafts (Fig. 1e,h,k,n,o). The seed set was similarly increased when pollinating wt plants with pollen from nrpd1s/wtr grafts (Fig. 1p), revealing that the nrpd1 pollen was viable after grafting. Nevertheless, although seed set was significantly increased upon grafting, it was not completely restored, suggesting that viable nrpd1 pollen is less efficient in fertilization. To test whether reduced fertilization efficiency is a consequence of pollen germination or pollen tube elongation defects, we analysed pollen tube growth of pollen derived from different graft combinations using Aniline blue staining. This analysis did not reveal a major germination or elongation defect of nrpd1 mutant pollen (Supplementary Fig. 1a). However, we found a significantly reduced number of ovules being targeted by pollen tubes when pollinated with nrpd1s/nrpd1r pollen (Supplementary Fig. 1a,b), suggesting that loss of NRPD1 impairs pollen tube guidance. The number of targeted ovules increased when pollinated with nrpd1s/wtr-derived pollen, indicating that grafting can restore functionality of nrpd1 pollen.
Our previous work also revealed a maternal nrpd1 effect on seed number22. To test whether this maternal defect could be suppressed by grafting, we pollinated grafted maternal plants with wt pollen. However, the maternal nrpd1 seed defect was not suppressed by grafting; we observed similar frequencies of abnormal seeds when pollinating nrpd1s/wtr plants with wt pollen (Supplementary Fig. 2). Together, these data indicate that a long-distance mobile signal from the wt rootstock to the nrpd1 scion restores pollen viability and contributes to fertilization efficiency.
Grafting restores siRNAs in pollen, but not in the endosperm
To test whether NRPD1-dependent siRNAs can move from roots to shoots and thereby suppress the nrpd1 pollen defect after grafting, we isolated sRNAs of mature pollen from wts/wtr, nrpd1s/nrpd1r and nrpd1s/wtr grafted genotypes. The sRNA profiles were highly correlated among biological replicates of the same genotype, but were clearly distinct between genotypes (Extended Data Fig. 1a,b). Consistent with previous findings22,23, nrpd1s/nrpd1r pollen was depleted for siRNAs in the 21- to 24-nucleotide size range (Fig. 2a–c and Extended Data Fig. 1c). NRPD1-dependent siRNAs accumulated over both transposable elements (TEs) and genes (Fig. 2b,c). Notably, in line with suppression of the nrpd1 pollen defect after grafting, there was a substantial restoration of global siRNA levels in pollen of nrpd1s/wtr grafted plants, which was particularly prominent over genes and TEs (Fig. 2a–c and Extended Data Fig. 1c).
a, An overview of the alignment of the sRNA reads from 21- to 24-nucleotide-long mapping to chromosome 1. Green, wts/wtr; black, nrpd1s/nrpd1r; orange, nrpd1s/wtr. b,c, Abundance of TE-derived (b) and gene-derived (c) sRNAs in a size range of 21 to 24 nucleotides. d, Venn diagram depicting the overlap of PMsiRNA clusters with NRPD1-dependent siRNA clusters. e, Pie chart depicting the proportion of siRNAs generated from the 169 PMsiRNA clusters relative to the total siRNA abundance from all NRPD1-dependent clusters. f, Pie chart displaying the length distribution of siRNAs generated from the 169 PMsiRNA clusters. g, Percentage of PMsiRNA clusters mapping to defined genomic features: gene body, 2 kb upstream of genes, intergenic region and TEs. h, PMsiRNA accumulation (RPM, reads per million) at specific genomic features. Asterisks indicate statistically significant enrichment calculated by a two-sided Fisher’s exact test (***P < 0.001, *P < 0.05). UTR, untranscribed region.
By comparing siRNAs in pollen of nrpd1s/nrpd1r and nrpd1s/wtr grafts, we identified 178 NRPD1-dependent siRNA clusters that were significantly restored after grafting (DESeq2, P < 0.05, log2(fold change) < −1). Nearly all of these (169) overlapped with the 3,100 NRPD1-dependent siRNA clusters identified by comparing nrpd1s/nrpd1r with wts/wtr pollen (Fig. 2d and Supplementary Table 1). These 169 clusters were defined as PMsiRNAs after grafting in pollen. The nine clusters that were not identified as NRPD1-dependent siRNAs in wts/wtr were below the significance threshold, but were expressed in wt pollen (Extended Data Fig. 1d).
Strikingly, the 169 PMsiRNA clusters accounted for approximately half (52%) of all siRNA reads present in the 3,100 NRPD1-dependent siRNA clusters (Fig. 2e). This finding reveals that in pollen, nearly half of the NRPD1-dependent siRNAs are produced from a few specific loci and are either non-cell autonomous or triggered by non-cell-autonomous signals.
The 169 PMsiRNA clusters contained ~19% of 21-nucleotide siRNAs, ~30% of 22-nucleotide siRNAs, ~30% of 23-nucleotide siRNAs and ~21% of 24-nucleotide siRNAs (Fig. 2f), which was similar to the size distribution present in the 3,100 NRPD1-dependent siRNA clusters (Extended Data Fig. 1e) and consistent with the previously reported dependence of 24-nucleotide and 21- or 22-nucleotide siRNAs on NRPD1 in pollen22,23. The 23-nucleotide siRNAs function as passenger strands during 24-nucleotide siRNA loading into AGO4 and are efficiently sliced by AGO424,25. Their accumulation suggests that they are not loaded into AGO4. Nevertheless, because the function of 23-nucleotide siRNAs is coupled to that of 24-nucleotide siRNAs, 23-nucleotide siRNAs were not considered further in our analyses. The majority of PMsiRNA loci accumulated siRNAs on both sense and antisense strands (Extended Data Fig. 1f,g), suggesting that they were derived from Dicer-mediated cleavage of double-stranded RNAs. PMsiRNA clusters mapped to TEs, intergenic regions, as well as gene body and promoter regions (Fig. 2g,h), suggesting that PMsiRNAs play a regulatory role not only on TEs, but also on genes. More than half of genic loci accumulating PMsiRNAs were predominantly expressed in reproductive tissues, whereas only eight TEs had detectable expression in reproductive tissues (Supplementary Fig. 3).
SiRNAs exert their functions upon binding to Argonaute (AGO) proteins, each characterized by distinct 5′ nucleotide preferences26. The 24-nuceotide PMsiRNAs exhibited a 5′ adenine (A) bias, suggesting they are sorted into AGO4 or closely related AGO proteins (Fig. 3a), whereas 21- and 22-nucleotide PMsiRNAs exhibited enrichment of both 5′ uridine (U) and A, suggesting preferential sorting into AGO1 and AGO2 (Fig. 3a).
a, Percentage of 21- to 24-nucleotide PMsiRNAs starting with specified 5′ nucleotides. b, sRNA abundance of AGO1-bound sRNAs in wt and nrpd1 buds. The RPM value is shown. c,d, Percentage of AGO1-bound sRNAs with specified 5′ nucleotides from 21- to 24-nucleotide PMsiRNAs in wt (c) and nrpd1 (d). e, In total, 127 siRNA clusters overlap between 169 PMsiRNA clusters and 2,674 AGO1-bound Pol IV-dependent clusters. f. Examples of loci accumulating PMsiRNAs that are bound by AGO1. The RPM value is shown. RIP, RNA immunoprecipitation.
To directly test whether PMsiRNAs load into AGO1, we performed AGO1-RNA immunoprecipitation from wt and nrpd1 flower bud tissues to profile AGO1-bound siRNAs. Consistent with AGO1’s preference for 21-nucleotide siRNAs27, we found 21-nucleotide siRNAs to be enriched in the AGO1-RNA immunoprecipitation samples of both genotypes (Fig. 3b). Also consistent with the preference of AGO1 for sRNA with a 5′ terminal uridine (U)28, AGO1-bound PMsiRNAs were enriched for U (Fig. 3c,d). We identified clusters of AGO1-bound siRNAs in wt and nrpd1 and tested which of these clusters were significantly enriched in wt tissues (DESeq2, P < 0.05, log2(fold change) < −1). Based on this analysis, we identified 2,674 NRPD1-dependent AGO1-bound siRNA clusters, of which 127 were PMsiRNA clusters (Fig. 3e,f). Based on these data, we conclude that PMsiRNAs can be loaded into AGO1, suggesting that they participate in gene silencing through a PTGS mechanism.
Among those genes overlapping with PMsiRNA clusters were many associated with endosperm development, including AGAMOUS-LIKE genes AGL28, AGL35, AGL62 and paternally expressed imprinted genes, including YUC10, as well as genes involved in pectin metabolism (Extended Data Figs. 2 and 3a and Supplementary Table 2)29,30,31. Several of those genes were previously found to generate siRNAs in the endosperm (sirenRNAs)31, suggesting that PMsiRNAs and sirenRNAs are generated from a similar set of loci. We tested this hypothesis by overlapping PMsiRNA loci with our previously identified siren loci31 in the endosperm and found that nearly all PMsiRNA clusters correspond to siren loci (Extended Data Fig. 3b). Given the strong overlap of PMsiRNAs and sirenRNAs, we tested whether we could restore sirenRNA formation in the endosperm by grafting. We sequenced sRNAs from manually dissected endosperm at 6–7 days after pollination from seeds of grafted genotypes wts/wtr, nrpd1s/nrpd1r and nrpd1s/wtr. We identified 1,031 NRPD1-dependent siRNA clusters in the endosperm by comparing wts/wtr with nrpd1s/nrpd1r (Extended Data Fig. 3c,d). Similar to previously published data31, sirenRNA clusters were enriched for 24-nucleotide siRNAs (Extended Data Fig. 3e), differing from the size distribution of PMsiRNAs that accumulated predominantly 21- and 22-nucleotide siRNAs (~50%) and only ~21% of 24-nucleotide siRNAs (Fig. 2f). The vast majority of PMsiRNA clusters (164 of 169) were also detected in wts/wtr endosperm (Extended Data Fig. 3d). In the endosperm, those clusters accumulated substantially higher levels of siRNAs compared with pollen (Extended Data Fig. 3f–h). However, unlike in pollen, siRNAs were not restored in the endosperm upon grafting (Extended Data Fig. 3f–h). These data reveal that PMsiRNAs are present in both pollen and endosperm, but the mobile signal after grafting is only transmitted from roots to male reproductive cells.
PMsiRNAs do not restore DNA methylation after grafting
Because NRPD1-dependent siRNAs have a functional role in guiding DNA methylation through the RdDM pathway32, we addressed the question of whether PMsiRNAs induced DNA methylation in nrpd1s/wtr and thereby contribute to restoring pollen viability. We conducted bisulfite sequencing of scion leaves, microspores and mature pollen grains (MPG) from three grafted genotypes: wts/wtr, nrpd1s/nrpd1r and nrpd1s/wtr. We observed a global depletion of CHG and CHH methylation on TEs and genes in nrpd1s/nrpd1r and nrpd1s/wtr microspores, pollen and scion leaves, revealing that grafting did not restore global DNA methylation (Fig. 4a,b and Extended Data Figs. 4a,b and 5a,b). Of 169 PMsiRNA loci, only 25 were methylated in the CHG and CHH context in wts/wtr leaves, microspores and pollen; however, their methylation status remained depleted in nrpd1s/wtr (Fig. 4c,d, Extended Data Figs. 4c and 5c and Supplementary Fig. 4). These data suggest that PMsiRNAs restore pollen viability after grafting by functioning through a pathway distinct from RdDM. It furthermore suggests that the restoration of DNA methylation by graft-transmissible siRNAs requires a functional Pol IV, consistent with previous data8. We also tested whether DNA methylation would be restored in the progeny derived after self-fertilization of grafted nrpd1s/wtr by bisulfite sequencing of leaves from 10-day-old seedlings. However, the DNA methylation profile of the graft-derived progeny was depleted for CHG and CHH methylation, similar to leaves from nrpd1s/nrpd1r (Extended Data Fig. 6 and Supplementary Fig. 4), revealing that PMsiRNAs had no transgenerational effects on DNA methylation in the absence of a functional Pol IV.
a,b, Metagene plots showing global DNA methylation levels of genes (a) and TEs (b) at CG, CHG and CHH positions. c, Examples of CHH methylation in microspores at four PMsiRNA loci of indicated genotypes. d, Boxplots showing methylation levels of all PMsiRNA loci (n = 169) of the indicated genotypes. Means of two biological replicates were computed. Centre lines show the medians; box limits indicate the 25th and 75th percentiles; and whiskers extend 1.5× the interquartile range from the 25th and 75th percentiles. The asterisks mark statistically significant differences (***P < 0.001). Statistical significance of differences was determined using the pairwise two-sided Mann–Whitney U-test; exact P values are provided in the source data. kb, kilobase.
Pol IV-dependent siRNAs from roots trigger amplification of PMsiRNAs
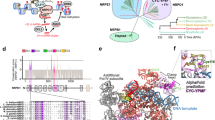
To test whether PMsiRNAs are indeed generated in roots, we isolated and sequenced sRNAs from wt and nrpd1 roots, identifying 9,085 NRPD1-dependent siRNA loci in wt roots, with most of them generating siRNAs of 21 or 22 nucleotides and 24 nucleotides (Extended Data Fig. 7a–c). Only 25 of these loci perfectly overlapped with PMsiRNA loci (Fig. 5a and Extended Data Fig. 7d). However, when allowing for up to three mismatches to their predicted target mRNAs, we identified 707 root loci generating Pol IV-dependent 21- and 22-nucleotide siRNAs that aligned to 126 of 169 PMsiRNA loci (Fig. 5b and Supplementary Tables 3 and 4). In contrast to PMsiRNAs that accumulated over longer genomic regions, root-derived siRNAs were preferentially derived from TEs and accumulated as highly abundant siRNAs over very narrow regions (Fig. 5c,d and Extended Data Fig. 7e,f). The vast majority (84%) were generated from both strands, ruling out that these loci are new miRNAs. Root loci significantly overlapped with helitrons and Harbinger DNA elements (Extended Data Fig. 7g) and were enriched for specific sequence motifs (Extended Data Fig. 7h), suggesting specific binding of transcription factors to those motifs.
a, Examples of CHH methylation and siRNA abundance in roots and microspores. The first two loci are examples of methylated PMsiRNA loci that accumulate siRNAs in roots. The third and fourth loci are unmethylated PMsiRNA loci without siRNA accumulation in roots. b, An example showing siRNAs in roots targeting pollen PMsiRNA loci. The red arrow indicates siRNAs from the original root locus that can be mapped to the PMsiRNA locus with up to three mismatches. c, Example of PMsiRNAs in pollen mapping specifically to exons of a coding gene. d, Examples of root siRNAs accumulating in narrow regions of TEs. e, Circos figure showing an example of root siRNAs from multiple root loci targeting a single PMsiRNA locus in trans. Root siRNAs from 55 loci can trans target a single PMsiRNA cluster indicated by a blue circle (Cluster_17934: Chr. 4, 11082108–11082728). Scaffold indicates chromosome. f, The abundance of 21- and 22-nucleotide and 24-nucleotide PMsiRNA over exons and introns of coding genes. Asterisks indicate statistically significant enrichment calculated by a two-sided Fisher’s exact test (***P < 0.001). RPKM, reads per kilobase of transcript per million mapped reads.
Interestingly, we found that siRNAs from many distinct root loci could target one specific PMsiRNA cluster (Fig. 5e and Supplementary Table 4). Thus, 35 of the 126 PMsiRNA loci were targeted by siRNAs from at least five different root loci. For example, PMsiRNA cluster_2988 (Chr. 1: 12299314–12300170) is trans targeted by siRNAs from 121 root loci, cluster_17934 (Chr. 2: 11082108–11082728) by siRNAs from 55 root loci, cluster_4816 by siRNAs from 27 root loci, and cluster_33527 by siRNAs from 24 root loci. Interestingly, many of the PMsiRNA loci that are targeted by multiple root loci overlap with helitrons: PMsiRNA cluster_2988 contains three helitrons, cluster_17934 overlaps with one helitron, cluster_4816 intersects with gene Carubv10011462m.g, encoding a pectin methylesterase inhibitor that contains a helitron in the intron region, and cluster_33527 intersects with gene Carubv10026508m.g, encoding a member of the ARF GAP domain family and also contains a helitron in the intron region. These data align with the enrichment of root siRNA loci for helitrons (Extended Data Fig. 7g), indicating that helitron-derived root siRNAs target conserved regions in trans.
We speculated that NRPD1-dependent siRNAs produced in roots may target PMsiRNA loci in trans and induce secondary siRNA amplification, as previously proposed33. Consistent with this idea, we found that there was a preference for PMsiRNAs to be generated from exonic regions of genes (Fig. 5f), with several (21) PMsiRNA loci exclusively accumulating over exons (Fig. 5c). We quantified the accumulation of PMsiRNAs over exons and introns and found a comparable exonic bias for 24-nucleotide and 21- or 22-nucleotide siRNAs in pollen of the nrpd1s/wtr grafts (Fig. 5f). Target cleavage by miRNA-loaded AGO1 can trigger the biogenesis of secondary siRNAs that have a distinctive phased configuration34. However, we did not detect elevated phasing scores at PMsiRNA-generating clusters, suggesting that the trigger for secondary siRNA formation is not one distinct cleavage event, but possibly multiple cleavage events causing out-of-phase secondary siRNA formation.
We addressed the question of whether PMsiRNA formation is indeed specifically triggered from root-derived siRNAs or whether the trigger is also present in other sporophytic organs. Indeed, the majority of siRNAs of the 707 root clusters were also present in non-grafted wt leaves (Extended Data Fig. 7i), suggesting that the trigger siRNAs for PMsiRNA formation are not root specific but are generated in different sporophytic organs. Consistently, pollen of wts/nrpd1r grafts developed normally and accumulated PMsiRNAs (Fig. 1f–h and Supplementary Fig. 5). We thus conclude that the trigger for PMsiRNA formation can also be generated in other sporophytic tissues than roots.
Secondary siRNA formation requires the activity of RDR6, which processes cleaved messenger RNA substrates into double-stranded RNA molecules that serve as substrates for DCL2 and DCL4 to produce 22-nucleotide and 21-nucleotide siRNAs, respectively7. To further investigate whether the amplification of PMsiRNAs relies on a conventional PTGS mechanism, we used CRISPR–Cas9 to generate two loss-of-function mutant alleles of the single RDR6 orthologue in Capsella (Supplementary Fig. 6a,b). These alleles featured deletions after amino acids 432 and 434, resulting in truncated proteins that lacked the conserved catalytic Asp867 residue (Supplementary Fig. 6b)35. Both mutant alleles exhibited severely reduced pollen numbers, attributable to impaired anther locule formation and disrupted pollen gametogenesis (Extended Data Fig. 8). Frequently, only one or two of the four anther locules developed, and pollen in these locules was partially inviable (Extended Data Fig. 8m–x). Inviable pollen arrested at various developmental stages, with a predominant arrest at the microspore stage (Extended Data Fig. 8m–r), mirroring the defects observed in Capsella nrpd1 mutants (Extended Data Fig. 8g–j)22. However, unlike nrpd1, which does not have notable tapetal defects (Extended Data Fig. 8g–j)22, rdr6 mutants exhibited a significant delay in tapetal degeneration (Extended Data Fig. 8m–r). In rdr6 mutants, we frequently observed anther locules filled with tapetum-like structures (Extended Data Fig. 8s–x) or undegraded tapetum with no or few pollen grains inside (Extended Data Fig. 8s–x).
Because of the severe defects in pollen formation in rdr6, we focused on isolating and sequencing sRNAs from mature anthers of wt and rdr6 plants rather than from pollen (Extended Data Fig. 9a–c). In wt anthers, siRNA accumulation was detected at 154 of 169 PMsiRNA loci. Surprisingly, PMsiRNA accumulation was also observed in rdr6 anthers of both mutant alleles, with the level of accumulation being substantially higher in rdr6 compared with wt anthers (Extended Data Fig. 9d,e). The general trend was similar between both alleles, with some variation due to variable numbers of developed locules. The increased accumulation of PMsiRNAs in rdr6 anthers is possibly a consequence of the impaired tapetal degradation in rdr6 (Extended Data Fig. 8m–r), suggesting that PMsiRNA generation occurs in the tapetum. By contrast, the formation of RDR6-dependent TAS3 siRNAs1 was strongly impaired in rdr6 mutants (Extended Data Fig. 9f), confirming that Capsella RDR6 functions as a canonical orthologue of RDR6. We thus speculate that other RDRs are involved in this process and possibly act redundantly with RDR6 in PMsiRNA formation32.
Grafting partially restores gene expression in microspores
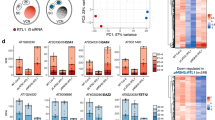
To dissect the functional role of PMsiRNAs in regulating pollen development, we conducted RNA sequencing of microspores of wts/wtr, nrpd1s/nrpd1r and nrpd1s/wtr genotypes. Transcriptome analysis of microspores identified 751 differentially expressed genes in nrpd1s/nrpd1r compared with wts/wtr; of these, 629 genes were downregulated and 122 genes upregulated (Fig. 6a and Supplementary Table 5). The expression of the majority of deregulated genes (454 downregulated genes and 53 upregulated genes) was partially restored in nrpd1s/wtr (Fig. 6b,c and Supplementary Table 5). Transcripts for NRPD1 were not among the restored genes, ruling out that long-distance movement of NRPD1 mRNAs is responsible for the normalization of pollen development. Restored genes had functional roles related to pollen development and pollen tube growth (Fig. 6d) and 131 overlapped with predicted targets of 21- or 22-nucleotide PMsiRNAs (Fig. 6e and Supplementary Table 6). The majority of PMsiRNAs were directed against genes that were downregulated in nrpd1, although 13 targeted upregulated genes (Fig. 6f). The restored expression of those 13 targets was not connected to changes in DNA methylation (Fig. 6g) or to the accumulation of siRNAs, suggesting that PMsiRNAs directly or indirectly contribute to male gametophyte development by mechanisms that are probably independent of RdDM and the slicing of mRNA targets.
a, Volcano plot showing differentially expressed genes in nrpd1/nrpd1 compared with wt/wt. Blue, downregulated genes; red, upregulated genes. b, Heatmap showing genes with recovered expression after grafting in nrpd1s/wtr microspores. c, Graph showing the relative change of upregulated (n = 53 ) and downregulated (n = 454) genes in comparison with wt after grafting. Means of two biological replicates were computed. Centre lines show the medians; box limits indicate the 25th and 75th percentiles; and whiskers extend 1.5× the interquartile range from the 25th and 75th percentiles. d, Enriched Gene Ontology terms of biological processes of normalized genes after grafting in nrpd1s/wtr microspores. The top ten Gene Ontology terms are shown. e, Overlap of normalized genes in nrpd1s/wtr microspores and predicted targets of PMsiRNAs. f, Heatmap showing expression of potentially PMsiRNA targeted genes that are deregulated in nrpd1 and normalized after grafting. The log(transcripts per million (TPM)) value is shown. g, Methylation levels of normalized PMsiRNA target genes (n = 13) shown in f. Means of two biological replicates were computed. Centre lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5× the interquartile range from the 25th and 75th percentiles. P > 0.05. Statistical significance of differences was determined using the pairwise two-sided Mann–Whitney U-test; exact P values are provided in the source data. h, Heatmap showing grafting normalized expression of genes that accumulate PMsiRNAs at the gene body or 2 kb promoter regions. FDR, false discovery rate.
We also considered that the regulation of transcript abundance from PMsiRNA-generating loci themselves might be critical for pollen viability. To investigate this, we examined transcript accumulation at PMsiRNA loci across wts/wtr, nrpd1s/nrpd1r and nrpd1s/wtr genotypes. Our analysis revealed that most loci exhibited transcript hyperaccumulation in nrpd1s/nrpd1r microspores, with approximately half showing reduced transcript levels following grafting (Fig. 6h and Extended Data Fig. 10). We speculate that the ectopic accumulation of transcripts from PMsiRNA loci in nrpd1 pollen may underlie the observed pollen arrest phenotype. We propose that a key function of sporophytically derived siRNAs is to suppress RNA accumulation from PMsiRNA loci, thereby supporting proper pollen development.
Discussion
In this study we report the following new discoveries: (1) Pol IV-dependent siRNAs generated in sporophytic tissues are mobile and act in trans to induce the formation of secondary siRNAs that accumulate in pollen (referred to as Pol IV-dependent mobile siRNAs, PMsiRNAs); (2) the main function of PMsiRNAs is not to induce DNA methylation, but to act post-transcriptionally, by either inducing PTGS in cis and trans and/or to regulate translation; and (3) PMsiRNAs resemble reproductive phasiRNAs (Fig. 7). Reproductive phasiRNAs are broadly present in angiosperms and play a crucial role in male fertility17,18,36. Interestingly, despite their broad presence in angiosperms, they are considered absent in certain eudicot lineages, notably in the Brassicaceae family17,18. Capsella PMsiRNAs identified in this study share several features with reproductive phasiRNAs: both are secondary siRNAs generated from Pol II transcripts, originate from sporophytic tissues, are 21–24 nucleotides in length, and are implicated in male fertility. However, in contrast to reproductive phasiRNAs that frequently require miRNAs for their amplification15,16,17,18, PMsiRNAs are triggered by Pol IV-dependent siRNAs originating in sporophytic tissues. The TE origin of many miRNA loci37 suggests that Pol IV-dependent triggers for PMsiRNAs may represent an ancestral mechanism of phasiRNA formation, preceding the emergence of specialized miRNAs that mediate phasiRNA production in other eudicots. There are three major types of reproductive siRNAs: premeiotic 21-nucleotide and 24-nucleotide phasiRNAs and meiotic 24-nucleotide phasiRNAs. When exactly PMsiRNAs are generated remains to be demonstrated; however, the accumulation PMsiRNAs in the size range of 21–24 nucleotides suggests that their biogenesis may occur during meiosis when nuclear compartmentalization breaks down and DCL enzymes gain concurrent access to target RNAs. Interestingly, a recent study reported the formation of premeiotic 24-nucleotide phasiRNAs in durum wheat independently of miRNA-mediated cleavage38. Disruption of these phasiRNAs led to defects at the microspore stage, resembling the phenotype we observe upon loss of PMsiRNAs. Together, these parallels suggest that PMsiRNAs may fulfil a role analogous to reproductive phasiRNAs in other species.
Pol IV-generated siRNAs trigger RdDM in the root, probably involving the canonical machinery of RdDM11. A subpopulation of Pol IV-dependent root siRNAs are mobile and associate with AGO1 (and possibly other AGOs), inducing the formation of double-stranded transcripts via RDRs and the cleavage of transcripts by DCL2 and/or DCL4, leading to the formation of secondary siRNAs (termed Pol IV-dependent mobile siRNAs, PMsiRNAs).
Previous studies have demonstrated the movement of siRNAs from the tapetum to meiocytes (referred to as tapetal siRNAs)20,21, highlighting the large-scale impact of non-cell-autonomous siRNAs in pollen development. However, our data suggest that the non-cell-autonomous PMsiRNAs identified in this study are distinct from tapetal siRNAs. First, tapetal siRNAs are generated specifically in the tapetum because of the specific expression of CLASSY3 in the tapetum, whereas PMsiRNAs do not require Pol IV function in the tapetum. Second, whereas tapetal siRNAs primarily guide DNA methylation, PMsiRNAs do not restore DNA methylation in nrpd1 pollen and are unlikely to have methylation as their main function. Instead, PMsiRNAs appear to act post-transcriptionally, consistent with previous findings that Pol IV-dependent 21- and 22-nucleotide siRNAs regulate gene expression at the post-transcriptional level33.
Despite these differences, PMsiRNAs and tapetal siRNAs share several common characteristics: they are both Pol IV-dependent, non-cell autonomous, organized in clusters and accumulate in the size range of 21–24 nucleotides. These features make them distinct from classical RdDM-associated siRNAs39. This suggests that pollen development relies on two distinct populations of Pol IV-dependent siRNAs with separate functions: tapetal siRNAs act during meiosis and silence TEs in the germline40, whereas PMsiRNAs appear to function post-meiotically, because nrpd1 pollen arrests at the microspore stage.
The preferential accumulation of PMsiRNAs over exons suggests that they are generated by post-transcriptional mechanisms, which, however, do not involve RDR6 function. This corroborates previous reports showing that Pol IV-dependent biogenesis of 21- and 22-nucleotide siRNAs is independent of RDR633. Similarly, in Arabidopsis pollen, epigenetically activated siRNAs—Pol IV-dependent siRNAs of 21–24 nucleotides—are only weakly affected in rdr6 mutants23, suggesting that their biogenesis may involve a similar, RDR6-independent pathway as discovered here for PMsiRNAs.
Our study reveals that loci generating PMsiRNAs overlap with those producing sirenRNAs, which depend on Pol IV activity in maternal tissues and are thought to move to the central cell or endosperm31,41. Recent data suggest that sirenRNAs are amplified in the endosperm, with amplification requiring siRNAs from surrounding maternal tissues31. This strikingly parallels the proposed formation of PMsiRNAs, suggesting convergence of the sirenRNA and PMsiRNA biogenesis pathways in male and female reproductive tissues. The loss of sirenRNAs and PMsiRNAs results in pollen arrest and endosperm failure in Capsella, underscoring the essential roles of reproductive siRNAs.
siRNAs are known to move bidirectionally between the shoot and root. Although there is substantial evidence for the movement and functional roles of siRNAs translocating from the shoot to the root2,6,12. The mechanism and biological significance of natural siRNA movement from root to shoot remain largely unexplored. Previous work revealed graft-transmissible movement of inverted-repeat-induced siRNA signals into flowers, postulating the existence of endogenous long-distance sRNA transport into meiotically active cells42. Similarly, suppression of MUTS HOMOLOG 1 (MSH1) results in graft-transmissible heritable changes of DNA methylation14,43, supporting the existence of root-to-shoot transport mechanisms of siRNAs. Our study demonstrates the movement of naturally present siRNAs from roots to shoots, where they regulate pollen development. These findings reveal the potential of siRNAs as long-distance signals in reproductive development. To what extent this phenomenon occurs in nature and which signals are induced in roots and transferred to shoots remains an exciting field of future research.
In summary, our data reveal that male reproductive development in Capsella relies on non-cell-autonomous PMsiRNAs that resemble reproductive phasiRNAs present in most angiosperms. PMsiRNAs act post-transcriptionally rather than through DNA methylation, pointing to a siRNA-based mechanism that coordinates gametophytic development with surrounding sporophytic tissues. Future studies will focus on elucidating the precise molecular mechanisms by which PMsiRNAs regulate gene expression.
Methods
Plant material, growth conditions and grafting
The Capsella rubella accession Cr1GR1 was used for all experiments. The nrpd1 mutant of Capsella rubella has been described previously22.
Seeds were sterilized with 70% EtOH, rinsed with distilled water, and sown on half-strength Murashige and Skoog medium with 1% sucrose. Seeds were stratified for 2 days in the dark at 4 °C and then transferred to a growth chamber with a long-day photoperiod (16 h light and 8 h dark, 22 °C, 50% humidity) for 2 weeks. Seedlings were then transferred to soil and grown under long-day conditions in the phytotron under 16 h of light at 21 °C and 8 h of darkness at 18 °C with a light intensity of 150 µE and 70% humidity.
Hypocotyl grafting was conducted 3 days after germination and adjusted from the previously described protocol44. Crosses and seed phenotyping were done as previously described45.
Construction of plasmids and transgenic plants
Generation of the CRISPR–Cas9-induced rdr6 mutant followed the same procedure as previously described22. All primers used for cloning and genotyping are listed in Supplementary Table 7.
Tissue embedding and microscopy
Capsella inflorescences were collected and stored in a fixative containing 3:1 ethanol/acetic acid. Anthers were dissected from floral buds under a stereomicroscope and submerged in Alexander’s staining solution for 30 min as previously described46. For pollen counting, anthers were manually dissected with a needle to release pollen and stained with Alexander’s staining solution. The slides were observed using a Keyence digital microscope.
For Aniline blue staining, siliques were collected 15 h after hand pollination and fixed overnight at 4 °C in ethanol/acetic acid (9:1, v/v). Samples were rehydrated through a graded ethanol series (70%, 50%, 30%; 10 min each). The tissue was then incubated in 8 M NaOH at room temperature overnight. Before microscopy, siliques were stained with 0.1% Aniline blue in phosphate buffer. Individual siliques were mounted on slides in a mounting solution containing phosphate buffer and 50% glycerol, and gently opened under a coverslip and analysed by epifluorescence microscopy using UV excitation.
To generate sections, Capsella inflorescences were harvested and fixed in FAA solution (50% v/v ethanol, 5% v/v acetic acid and 4% v/v formaldehyde) and embedded using a Historesin Embedding Kit (Leica, catalogue no. 702,218,500). Two-micrometre sections were prepared using an HM 355 S microtome (Microm) with glass knives. Sections were stained with 0.1% (w/v) toluidine blue for 1 min, washed five times with distilled water, air dried and observed with an Axio Scope (Zeiss).
Pollen and microspore extraction
MPGs and microspores were extracted as previously described47. Briefly, flower buds up to stage 13 were collected into ice-cold 0.1 M D-mannitol. MPGs were released by vigorously shaking the bottle containing the flower buds in D-mannitol solution. Suspensions containing MPGs were first filtered through a 100-µM nylon mesh, followed by a 53-µM mesh, and the flow through was kept for MPG isolation. The flower buds were shaken and filtered two or three times until the suspensions were clear. The collected inflorescences were used for microspore extraction and the filtered suspensions were used for MPG extraction. MPGs were collected by several rounds of centrifugation at 900g at 4 °C for 10 min. The inflorescences were homogenized using an ice-cold pestle and mortar and then filtered through 100-µM and 53-µM nylon meshes. The suspension was added on top of a 75%/65%/20%/10% Percoll A gradient that after centrifugation allowed separation into three fractions: A1, A2 and A3. The A2 subfraction containing microspores and bicellular pollen was collected and further separated by a Percoll B gradient (55%/45%/35%) into B1, B2 and B3 fractions. The B2 fractions were mainly enriched for microspores as previously described22.
AGO1-RNA immunoprecipitation
Unopened flower buds of Capsella rubella wt and nrpd1 were collected, frozen in liquid nitrogen, and ground using a cold pestle and mortar. Three times the volume of protein extraction buffer (EB) (50 mM Tris HCl, pH 7.5, 150 mM NaCl, 10% glycerol, 5 mM MgCl2, 0.1% Igepal, 10 µM MG132, 2 mM dithiothreitol and 1 tablet of protease inhibitor cocktail (Roche)) was added and the tissue–buffer mixture was incubated on a rotator for 30 min at 4 °C. Tissue debris was removed by centrifuging twice at 12,000g for 15 min and the supernatant was collected. Twice-washed PureProtemo Protectin A Magnetic beads (Merck, LSKMAGA02) were suspended in 1 ml of EB. AGO1 antibodies (Agrisera, AS09527) were added to the tube and incubated for 3 – 4 h at 4 °C to couple to the beads. The beads were separated using a magnetic rack and incubated with the supernatant obtained from the tissue–EB mixture for 2 h at 4 °C. Beads were magnetically separated, washed three times with EB and used directly for subsequent protein and RNA extraction.
sRNA library construction and sequence analysis
Total RNA was extracted using the mirVana miRNA Isolation Kit (Thermo Fisher, AM1560) with phenol. Small RNAs were extracted from total RNAs by size selection of 15 – 30-nucleotide sRNAs using a 15% TBE–urea polyacrylamide gel. sRNA libraries were constructed using the NEBNext Multiplex Small RNA Library Prep Set for Illumina (NEB, catalogue no. E7300). Final libraries were purified from a size selection of bands of about 150 bp on a 6% polyacrylamide gel. Biological duplicates were generated for pollen from grafted plants and anther, and biological triplicates were generated for root samples. sRNA libraries were sequenced on an Illumina NovaSeq 6000 machine in 150 bp pair-end mode.
For the sRNA sequence data, adaptors were removed from the 150-bp paired-end sRNA reads in each library. Adaptors were removed from the first read of the 150-bp read pair of each library using cutadapt, and the resulting 18–25-bp reads were selected. Reads belonging to chloroplasts, mitochondria and structural non-coding RNAs (transfer RNAs, small nuclear RNAs, ribosomal RNAs or small nucleolar RNAs) were removed using bowtie (v.1). The remaining reads were mapped to the Capsella genome v.1.0 (Phytozome) and sRNA loci were annotated with ShortStack v.3.8.548,49. Options used for ShortStack were mismatches, 0; mmap, u; mincov, 0.5 rpm; and pad, 75. Replicates were checked for consistency by principal component analysis using the vegan 2.6-4 package in R50. siRNA clusters were considered NRPD1-dependent by performing a differential accumulation analysis with DESeq251. On the basis of log2(fold change) > 1 and Padj (adjusted P value) < 0.05 in comparisons nrpd1s/nrpd1r versus wts/wtr; nrpd1s/nrpd1r versus nrpd1s/wtr, genomic loci losing siRNAs in nrpd1s/nrpd1r were determined by siRNA clusters intersecting with TEs, genes, and 2 kb upstream of the genes.
RNA library construction and sequence data analysis
The total RNA of microspores was extracted using the Direct-zol RNA Miniprep Kit (Zymo, R2050) and treated using a NEB Poly(A) mRNA Magnetic Isolation Module Kit (NEB, E7490). RNA libraries were prepared from the resulting mRNAs using the NEB Ultra II RNA Library Pre Kit for Illumina (NEB, E7770). Biological duplicates were generated for each genotype. RNA libraries were sequenced on an Illumina NovaSeq 6000 machine in 150-bp pair-end mode.
Adaptor trimming was performed using Trim galore with the following parameters: --three_prime_clip_R1 15 --three_prime_clip_R2 15 --clip_R1 10 --clip_R2 10. Sequencing reads were aligned to the Capsella genome v.1.0 (Phytozome) using HISAT252. Reads were assigned to genes with featureCounts from the Bioconductor Rsubread package53. Differentially regulated genes were detected using DESeq251. Genes were considered as upregulated based on log2(fold change) > 1 and Padj < 0.05 in comparison with the respective control. PCA was performed with the vegan R package, considering normalized gene expression values (TPM)50. Normalization of gene expression shown in Fig. 6c was calculated by using gene expression (TPM value) in wt microspores as standard and comparing the difference between the nrpd1 mutant (\(a=\frac{{nrpd}{1}^{\text{s}}}{{nrpd}{1}^{\text{r}}}-\text{wt}\)) with that of microspores from grafted plants \(b=\frac{{nrpd}{1}^{\text{s}}}{{\text{wt}}^{\text{r}}-\text{wt}}\) to calculate the relative change to \(\text{wt}=(1-(\frac{b}{a}))\times 100{\rm{ \% }}\).
Bisulfite sequencing and data analysis
Genomic DNA from microspores, mature pollen, rosette leaves of 3-month-old grafted plants and 10-day-old seedlings of the progeny of grafted plants were extracted using the DNeasy Plant Mini Kit (Qiagen). Libraries were prepared with the Accel-NGS Methyl-Seq DNA Library Kit (Illumina) and the sequencing was performed on an Illumina NovaSeq 6000 machine in 150-bp paired-end mode. Biological duplicates were generated for each genotype. Libraries were prepared with the Accel-NGS Methyl-Seq DNA Library Kit (Illumina), and the sequencing was performed on an Illumina NovaSeq 6000 machine in 150-bp paired-end mode.
For DNA methylation analysis, the 150-bp paired-end reads were first quality trimmed by removing the first 5 bases from the 5′-end and the last 15 bases from the 3′-end. Reads were mapped to the Cr reference genome in paired-end mode (score_min L, 0–0.6) using Bismark v.0.16.354. The mapped reads were deduplicated, and cytosine methylation values were calculated using the Bismark Methylation Extractor.
Differentially methylated regions in the CG, CHG and CHH contexts were calculated using 50-bp windows across the genome as units. Only hypomethylated regions (wts/wtr > nrpd1s/nrpd1r, nrpd1s/wtr > nrpd1s/nrpd1r) were considered. Windows with differences in fractional methylation below the first decile (Fisher’s exact test P < 0.01) were selected, and these were merged if they occurred within 300 bp.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing data generated in this study are available in the Gene Expression Omnibus in NCBI under the accession numbers GSE291574 (Bisulfite-seq data), GSE291577 (RNA-seq data) and GSE291578 (sRNA-seq data). Source data are provided with this paper.
References
Chen, X. & Rechavi, O. Plant and animal small RNA communications between cells and organisms. Nat. Rev. Mol. Cell Biol. 23, 185–203 (2022).
Molnar, A. et al. Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science 328, 872–875 (2010).
Li, S. et al. Unidirectional movement of small RNAs from shoots to roots in interspecific heterografts. Nat. Plants 7, 50–59 (2021).
Palauqui, J. C., Elmayan, T., Pollien, J. M. & Vaucheret, H. Systemic acquired silencing: transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions. EMBO J. 16, 4738–4745 (1997).
Voinnet, O. & Baulcombe, D. C. Systemic signalling in gene silencing. Nature 389, 553 (1997).
Melnyk, C. W., Molnar, A., Bassett, A. & Baulcombe, D. C. Mobile 24 nt small RNAs direct transcriptional gene silencing in the root meristems of Arabidopsis thaliana. Curr. Biol. 21, 1678–1683 (2011).
Vaucheret, H. & Voinnet, O. The plant siRNA landscape. Plant Cell 36, 246–275 (2024).
Brosnan, C. A. et al. Nuclear gene silencing directs reception of long-distance mRNA silencing in Arabidopsis. Proc. Natl Acad. Sci. USA 104, 14741–14746 (2007).
Dunoyer, P., Himber, C., Ruiz-Ferrer, V., Alioua, A. & Voinnet, O. Intra- and intercellular RNA interference in Arabidopsis thaliana requires components of the microRNA and heterochromatic silencing pathways. Nat. Genet. 39, 848–856 (2007).
Smith, L. M. et al. An SNF2 protein associated with nuclear RNA silencing and the spread of a silencing signal between cells in Arabidopsis. Plant Cell 19, 1507–1521 (2007).
Law, J. A. et al. Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature 498, 385–389 (2013).
Lewsey, M. G. et al. Mobile small RNAs regulate genome-wide DNA methylation. Proc. Natl Acad. Sci. USA 113, E801–E810 (2016).
Mallory, A. C., Mlotshwa, S., Bowman, L. H. & Vance, V. B. The capacity of transgenic tobacco to send a systemic RNA silencing signal depends on the nature of the inducing transgene locus. Plant J. 35, 82–92 (2003).
Kundariya, H. et al. MSH1-induced heritable enhanced growth vigor through grafting is associated with the RdDM pathway in plants. Nat. Commun. 11, 5343 (2020).
Johnson, C. et al. Clusters and superclusters of phased small RNAs in the developing inflorescence of rice. Genome Res. 19, 1429–1440 (2009).
Zhai, J. et al. Spatiotemporally dynamic, cell-type-dependent premeiotic and meiotic phasiRNAs in maize anthers. Proc. Natl Acad. Sci. USA 112, 3146–3151 (2015).
Xia, R. et al. 24-nt reproductive phasiRNAs are broadly present in angiosperms. Nat. Commun. 10, 627 (2019).
Pokhrel, S. et al. Pre-meiotic 21-nucleotide reproductive phasiRNAs emerged in seed plants and diversified in flowering plants. Nat. Commun. 12, 4941 (2021).
Zhan, J. et al. Premeiotic 24-nt phasiRNAs are present in the Zea genus and unique in biogenesis mechanism and molecular function. Proc. Natl Acad. Sci. USA 121, e2402285121 (2024).
Zhou, X. et al. 24-nt phasiRNAs move from tapetal to meiotic cells in maize anthers. New Phytol. 235, 488–501 (2022).
Long, J. et al. Nurse cell-derived small RNAs define paternal epigenetic inheritance in Arabidopsis. Science 373, eabh0556 (2021).
Wang, Z. et al. Polymerase IV plays a crucial role in pollen development in Capsella. Plant Cell 32, 950–966 (2020).
Martinez, G. et al. Paternal easiRNAs regulate parental genome dosage in Arabidopsis. Nat. Genet. 50, 193–198 (2018).
Singh, J., Mishra, V., Wang, F., Huang, H. Y. & Pikaard, C. S. Reaction mechanisms of Pol IV, RDR2, and DCL3 drive RNA channeling in the siRNA-directed DNA methylation pathway. Mol. Cell 75, 576–589 (2019).
Wang, F., Huang, H. Y., Huang, J., Singh, J. & Pikaard, C. S. Enzymatic reactions of AGO4 in RNA-directed DNA methylation: siRNA duplex loading, passenger strand elimination, target RNA slicing, and sliced target retention. Genes Dev. 37, 103–118 (2023).
Mi, S. et al. Sorting of small RNAs into Arabidopsis argonaute complexes is directed by the 5′ terminal nucleotide. Cell 133, 116–127 (2008).
Baumberger, N. & Baulcombe, D. C. Arabidopsis ARGONAUTE1 is an RNA slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl Acad. Sci. USA 102, 11928–11933 (2005).
Takeda, A., Iwasaki, S., Watanabe, T., Utsumi, M. & Watanabe, Y. The mechanism selecting the guide strand from small RNA duplexes is different among argonaute proteins. Plant Cell Physiol. 49, 493–500 (2008).
Wolff, P., Jiang, H., Wang, G., Santos-González, J. & Köhler, C. Paternally expressed imprinted genes establish postzygotic hybridization barriers in Arabidopsis thaliana. eLife 4, e10074 (2015).
Zhang, S. et al. FERTILIZATION-INDEPENDENT SEED-Polycomb Repressive Complex 2 plays a dual role in regulating type I MADS-box genes in early endosperm development. Plant Physiol. 177, 285–299 (2018).
Dziasek, K. et al. Dosage-sensitive maternal siRNAs determine hybridization success in Capsella. Nat. Plants 10, 1969–1983 (2024).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Panda, K., McCue, A. D. & Slotkin, R. K. Arabidopsis RNA Polymerase IV generates 21–22 nucleotide small RNAs that can participate in RNA-directed DNA methylation and may regulate genes. Philos. Trans. R. Soc. Lond. B 375, 20190417 (2020).
Liu, Y., Teng, C., Xia, R. & Meyers, B. C. PhasiRNAs in plants: their biogenesis, genic sources, and roles in stress responses, development, and reproduction. Plant Cell 32, 3059–3080 (2020).
Curaba, J. & Chen, X. Biochemical activities of Arabidopsis RNA-dependent RNA polymerase 6. J. Biol. Chem. 283, 3059–3066 (2008).
Komiya, R. et al. Rice germline-specific Argonaute MEL1 protein binds to phasiRNAs generated from more than 700 lincRNAs. Plant J. 78, 385–397 (2014).
Roberts, J. T., Cardin, S. E. & Borchert, G. M. Burgeoning evidence indicates that microRNAs were initially formed from transposable element sequences. Mob. Genet. Elements 4, e29255 (2014).
Bélanger, S. et al. DICER-LIKE 5 loss causes thermosensitive male sterility in durum wheat and reveals an AU-rich motif guiding 24-nt phasiRNA biogenesis. Proc. Natl Acad. Sci. USA 122, e2504349122 (2025).
Havecker, E. R. et al. The Arabidopsis RNA-directed DNA methylation argonautes functionally diverge based on their expression and interaction with target loci. Plant Cell 22, 321–334 (2012).
Walker, J. et al. Sexual-lineage-specific DNA methylation regulates meiosis in Arabidopsis. Nat. Genet. 50, 130–137 (2017).
Grover, J. W. et al. Abundant expression of maternal siRNAs is a conserved feature of seed development. Proc. Natl Acad. Sci. USA 117, 15305–15315 (2020).
Zhang, W. et al. Graft-transmissible movement of inverted-repeat-induced siRNA signals into flowers. Plant J. 80, 106–121 (2014).
Yang, X. et al. Segregation of an MSH1 RNAi transgene produces heritable non-genetic memory in association with methylome reprogramming. Nat. Commun. 11, 2214 (2020).
Melnyk, C. W. Grafting with Arabidopsis thaliana. Methods Mol. Biol. 1497, 9–18 (2017).
Rebernig, C. A., Lafon-Placette, C., Hatorangan, M. R., Slotte, T. & Köhler, C. Non-reciprocal interspecies hybridization barriers in the Capsella genus are established in the endosperm. PLoS Genet. 11, e1005295 (2015).
Peterson, R., Slovin, J. P. & Chen, C. A simplified method for differential staining of aborted and non-aborted pollen grains. Int. J. Plant Biol. 1, e13 (2010).
Dupl’áková, N., Dobrev, P. I., Renák, D. & Honys, D. Rapid separation of Arabidopsis male gametophyte developmental stages using a Percoll gradient. Nat. Protoc. 11, 1817–1832 (2016).
Axtell, M. J. ShortStack: comprehensive annotation and quantification of small RNA genes. RNA 19, 740–751 (2013).
Johnson, N. R., Yeoh, J. M., Coruh, C. & Axtell, M. J. Improved placement of multi-mapping small RNAs. G3 6, 2103–2111 (2016).
Oksanen, J. et al. Vegan community ecology package v.2.6 (CRAN, 2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Acknowledgements
We thank U. Krause and M. Höhne for supporting cloning and plant handling. We thank A. Froehlich for supporting sectioning. We thank N. Butel for helpful comments on the manuscript. Financial support was received from Knut and Alice Wallenberg Foundation grant no. 2018-0206 (CK), Knut and Alice Wallenberg Foundation grant no. 2019-0062 (CK), Knut and Alice Wallenberg Academy Fellowship 2016-0274 (CWM) and the Max Planck Society, Germany.
Funding
Open access funding provided by Max Planck Society.
Author information
Authors and Affiliations
Contributions
J.Z. and C.K. conceptualized the project, developed the methodology and provided supervision. J.Z., Z.W., A.Z. and T.S. conducted experiments. J.S.-G. and T.H.C.R. performed bioinformatic analyses. B.C.M. and C.W.M. provided intellectual support. C.K. acquired funding and administered the project. J.Z. and C.K. wrote the original paper draft. All authors reviewed and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of sRNA libraries and PMsiRNAs in pollen.
a, Principal component analysis (PCA) of sRNA sequencing samples in pollen. b, Correlogram displays the correlation score matrix across all sequenced sRNA libraries. c, The global level of sRNA abundance from a size range of 21-24-nt. d, The heatmap showing the expression of 9 siRNA clusters present in both wts/wtr and nrpd1s/wtr, which were below the significance threshold in Fig. 2d. e. Percentage of NRPD1-dependent sRNA abundance across different sizes ranging from 21-nt to 24-nt. f, Percentage of PMsiRNA loci accumulating siRNAs at the indicated strand. g. Examples of PMsiRNA accumulation from both sense and anti-sense strands. +, sense strand; -, anti-sense strand.
Extended Data Fig. 2 Expression of PMsiRNAs accumulating in coding genes.
Heatmap shows expression levels of PMsiRNAs in coding genes in pollen. LogRPM values are shown.
Extended Data Fig. 3 Grafting does not restore sirenRNAs in the endosperm.
a, Heatmap shows the accumulation of PMsiRNAs on selected genes associated with endosperm development. b, Venn diagram depicts the overlap of siren loci and PMsiRNAs. c, Abundance of global sRNAs from 21-24-nt in the endosperm. d. Venn diagram shows the overlap of sRNA clusters between wts/wtr endosperm and pollen PMsiRNAs. e, Percentage of endosperm sRNA abundance in sizes ranging from 21-nt to 24-nt. f. Examples of PMsiRNA in pollen and endosperm from grafted plants of the indicated genotypes. RPM value is shown. g, Heatmap displays the accumulation of PMsiRNAs (LogRPM, n = 169) in pollen and endosperm after grafting in wts/wtr, wts/nrpd1r, nrpd1s/nrpd1r and nrpd1s/wtr. h, sRNA abundance in pollen and endosperm after grafting of indicated genotypes. Means of two biological replicates were computed. Center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles. Asterisks indicate statistically significant differences calculated by a Wilcoxon test, two-sided (*** p-value < 0.001, ns, not significant). the exact P values are provided in the Source Data file.
Extended Data Fig. 4 Grafting does not restore DNA methylation at PMsiRNA loci in pollen.
a and b, Metagene plots show pollen global DNA methylation level of genes (a) and TEs (b). c, Boxplots show DNA methylation level of all PMsiRNA loci (n = 169) in pollen. Means of two biological replicates were computed. Center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles. The asterisks mark statistically significant differences (***, P < 0.001, ns, not significant). Statistical significance of differences was determined using the pairwise Mann–Whitney test, two-sided; the exact P values are provided in the Source Data file.
Extended Data Fig. 5 Grafting does not restore DNA methylation at PMsiRNA loci in scion leaves.
a and b, Metagene plots show global DNA methylation levels of genes (a) and TEs (b) in scion leaves. c, Boxplots show DNA methylation ratios of all PMsiRNA loci (n = 169) in scion leaves of grafted plants. Means of two biological replicates were computed. Center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles. The asterisks mark statistically significant differences (***, P < 0.001, ns, not significant). Statistical significance of differences was determined using the pairwise Mann–Whitney test, two-sided; the exact P values are provided in the Source Data file.
Extended Data Fig. 6 DNA methylation is not restored in the progeny of the grafted plants.
a and b, Metagene plots show global DNA methylation levels of genes (a) and TEs (b) in the progeny of grafted plants. Leaves of 10-day-old seedlings were analyzed. c, Boxplots show DNA methylation levels of all PMsiRNA loci (n = 169) in the leaves of the grafted progeny. Means of two biological replicates were computed. Center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles. The asterisks mark statistically significant differences (***, P < 0.001, ns, not significant). Statistical significance of differences was determined using the pairwise Mann–Whitney test, two-sided; the exact P values are provided in the Source Data file.
Extended Data Fig. 7 sRNA profiling in root tissues.
a, Principal component analysis (PCA) of sRNA sequencing samples in roots. b, Abundance of global sRNAs from 21-24-nt in roots. c, Percentage of NRPD1-dependent sRNA abundance across different sizes ranging from 20-24-nt. d, Venn diagram displaying the overlap of root siRNA clusters (9085 clusters) and pollen PMsiRNAs (169 clusters). e, Number of root siRNA clusters mapping to TEs (n = ) and genes (n = ). f, Distribution of siRNA clusters based on length. Significance was tested using a two-sample proportion test: ***, P < 0.001; ** P < 0.01; ns, not significant. g, Percentage of root siRNA clusters mapping to different TE families. Significance was tested using Fisher’s test by comparing 707 root siRNA clusters that target PMsiRNAs in trans vs. random 707 root siRNA clusters, two-sided ***P-value < 0.001. not assigned: repeats not assigned to any TE family; no TEs: clusters not mapped to TEs. h, Motifs identified from 707 root siRNA clusters. E-value < 0.05: significant. i, Abundance (RPM value) of root siRNAs detected in leaves. Means of two biological replicates were computed. siRNAs were mapped to genes (left, n = 72), 2 kb promoters of genes (middle, n = 89), and TEs (right, n = 436).Center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles. The asterisks mark statistically significant differences (***, P < 0.001). Statistical significance of differences was determined using the pairwise Mann–Whitney U test, two-sided; the exact P values are provided in the Source Data file.
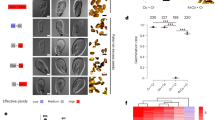
Extended Data Fig. 8 Cr rdr6 shows pollen and anther defects.
a-r, Cross-sections stained with toluidine blue at anther stages 7 to stage 12 of wild type (a-f) nrpd1 (g-i) and rdr6-1 (m-r). Scale bar: 50 μm. Zoom-ins of regions marked with a white frame are shown below each image. Scale bar: 10 μm. s-u, Cross-sections stained with toluidine blue at anther stage 12 of wild type (s), rdr6-1 (t) and rdr6-2 (u). Zoom-ins of regions marked with a white frame are shown below each image. Scale bar: 10 μm. v-x, Alexander staining of aborted and non-aborted pollen grains. (v) wild type (wt), (w) rdr6-1, (x) rdr6-2. Aborted and non-aborted pollen stain in dark blue and magenta, respectively. Scale bar: 100μm. Zoom-ins of regions marked with a white frame are shown below each image. Scale bar: 50 μm. y, Percentage of mature pollen (MP) in wt, rdr6-1 and rdr6-2. Two biological replicates derived from independent plants were analyzed. Statistical significance was assessed using a beta-binomial generalized linear model, followed by Tukey-adjusted pairwise comparisons, two-sided. The asterisks mark statistically significant differences (***, P < 0.001; ns, not significant). Center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles. The exact P values are provided in the Source Data file.
Extended Data Fig. 9 sRNA profiling from anthers of rdr6 mutants.
a, Principal component analysis (PCA) of sRNA sequencing samples of anthers. b and c, Length distribution of sRNA abundance from 21- to 24-nt over genes (b) and TEs (c). sRNA abundance was converted to RPM. d, Example of siRNA accumulation in anthers and pollen of the indicated genotypes. e, Heatmap of siRNA expression on PMsiRNA loci in wt and rdr6 anthers. LogRPM value is shown, f. Abundance of TAS3 siRNAs in wt and rdr6.
Extended Data Fig. 10 Expression of genes accumulating PMsiRNAs.
a and b, Heatmap shows RNA-seq expression levels of genes accumulating PMsiRNAs on the coding region of genes (a) and on the 2 kb promoter region of genes (b), logTPM value is shown.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6 and list of supplementary tables.
Supplementary Table 1 (download XLSX )
List of all siRNA clusters identified in this study.
Supplementary Table 2 (download XLSX )
Genes overlapping with clusters accumulating PMsiRNAs after grafting.
Supplementary Table 3 (download XLSX )
Root siRNAs and their corresponding PMsiRNA target loci.
Supplementary Table 4 (download XLSX )
Number of root siRNAs can target to single PMsiRNA in trans.
Supplementary Table 5 (download XLSX )
Genes with restored expression in microspores of grafted plants.
Supplementary Table 6 (download XLSX )
Predicted targets of PMsiRNAs.
Supplementary Table 7 (download XLSX )
Primers used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, J., Santos-González, J., Wang, Z. et al. Long-distance transport of siRNAs with functional roles in pollen development. Nat. Plants 12, 386–399 (2026). https://doi.org/10.1038/s41477-026-02219-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-026-02219-6