Abstract
Mitochondria are inherited maternally in the vast majority of eukaryotes. Occasional transmission of paternal mitochondria (paternal leakage) can lead to heterochondriomy and recombination between maternal and paternal mitochondrial genomes. Despite its potential physiological and evolutionary consequences, the extent of paternal leakage and the cellular processes governing mitochondrial inheritance remain largely unknown. Here we have established a robust genetic screen to detect paternal mitochondrial inheritance in tobacco (Nicotiana tabacum). Our data reveal an unexpectedly high paternal transmission frequency of 0.18%, which increased markedly to 7.34% when the organellar exonuclease DPD1 was disrupted and pollen development occurred at low temperature. Notably, paternally transmitted mitochondria restored growth, development and male fertility in progeny that inherited dysfunctional mitochondria from the maternal parent. Together, our findings uncover molecular mechanisms underlying maternal mitochondrial inheritance, and highlight the potential of biparental transmission to rescue mitochondrial function and generate novel mitochondrial genotypes through recombination.
Similar content being viewed by others
Main
In photosynthetic eukaryotes, genetic material is stored in three compartments: the nucleus, the plastids and the mitochondria1,2,3. Although plastid and mitochondrial genomes encode only a limited number of proteins, they are essential for key cellular processes including photosynthesis and respiration4,5,6. Consequently, mutations in cytoplasmic genomes can have profound effects on plant growth and development. A striking example is cytoplasmic male sterility (CMS), which arises from mutations in the mitochondrial genome that lead to the failure to produce functional pollen7. As cytoplasmic genomes are maternally inherited in most angiosperms, traits associated with cytoplasmic genomes are typically passed down across generations exclusively through the maternal lineage8,9.
The uniparental inheritance of cytoplasmic genomes stands in stark contrast to the biparental transmission of nuclear DNA10. Although numerous cytological studies support the predominance of maternal inheritance, accumulating evidence suggests that this maternal bias is not absolute. Rare instances of paternal plastid transmission have been reported in species traditionally considered to follow strict maternal inheritance, including Arabidopsis thaliana11, Brassica napus12 and Nicotiana tabacum (tobacco)13. These findings indicate that exceptional cases of paternal leakage can be identified when (1) highly sensitive methods for paternal plastids detection are employed and (2) sufficiently large sample sizes are analysed. Leveraging such sensitive, large-scale screening strategies, we have recently identified genetic and environmental factors that govern plastid inheritance in tobacco14.
In contrast to the progress made with uncovering the mechanisms underlying plastid inheritance, the cellular processes governing mitochondrial inheritance remain largely elusive. A prevailing hypothesis posits that the abundance of mitochondrial DNA in male gametes plays a crucial role in shaping mitochondrial inheritance15. In Arabidopsis, mitochondria are regularly present in sperm cells and can be transmitted to the egg cell upon fertilization16. However, during pollen maturation, mitochondrial DNA is actively degraded by nucleases, thus preventing its paternal transmission17,18,19,20. Although some nucleases involved in mitochondrial DNA degradation have been identified21,22, no instances of paternal mitochondrial transmission could be detected when the corresponding mutants were used as pollen donors. Consequently, the genes that regulate mitochondrial inheritance remain unknown and our understanding of paternal mitochondrial leakage is limited.
This knowledge gap is largely due to the lack of sensitive assays for the detection of paternal transmission events. In particular, the inability to express selectable marker genes from the mitochondrial genome has hindered the development of robust experimental systems for the study of mitochondrial inheritance. Although molecular techniques such as restriction fragment length polymorphism (RFLP) and derived cleaved amplified polymorphic sequence (dCAPS) have been used to specifically detect paternal mitochondria22,23,24, these approaches are labour intensive and lack the throughput required for large-scale screening.
Recent advances in mitochondrial genome engineering have opened up new possibilities to address this major bottleneck25,26,27,28. Using TALEN-mediated targeted mutagenesis, we have generated tobacco mitochondrial mutants that display distinct developmental phenotypes29. By leveraging these phenotypic differences between mutant and wild-type plants, we set out to establish an experimental strategy to detect paternal mitochondrial transmission events. Leveraging this strategy, we report here the identification of factors that control the maternal inheritance of mitochondria. We demonstrate that manipulation of these factors changes the mode of mitochondrial inheritance, thus modulating phenotype and physiology of the progeny. Importantly, our findings show that paternal mitochondria, once transmitted, can enter the germline and become stably inherited across generations, suggesting an impact on the evolution of mitochondrial genomes and speciation processes in plants.
Results
An experimental system to detect paternal mitochondrial transmission
The robust detection of paternal leakage in mitochondrial inheritance requires a high-throughput screening strategy. To identify a screenable phenotypic marker, we evaluated mitochondrial mutants that could be used as maternal parent in crosses with a paternal parent that harbours wild-type mitochondria. Any paternally transmitted mitochondria would then restore a wild-type-like phenotype in the progeny, which ideally, would be distinguishable from the mutant phenotype early upon seed germination. The mitochondrial Δnad9 mutant met these criteria. In this mutant, the nad9 gene has been deleted from the mitochondrial genome (Extended Data Fig. 1), leading to disrupted complex I biogenesis and a range of macroscopic phenotypes, including impaired growth and development, and male sterility29. These Δnad9-associated defects are stably passed on to the progeny through the maternal lineage. Most importantly, the Δnad9 progeny exhibits strongly delayed seed germination and seedling growth. While wild-type seedlings typically produce true leaves after 14 days, Δnad9 seeds germinate at a slower rate, and at day 14, only the cotyledons have emerged (Extended Data Fig. 2). This clear phenotypic difference provides a robust system for distinguishing the two mitochondrial genotypes (wild-type versus Δnad9). We reasoned that after crossing Δnad9 mutants with paternal plants carrying wild-type mitochondria, progeny harbouring paternal (wild-type) mitochondria would germinate and grow faster than the progeny that harbours only maternal (Δnad9) mitochondria. Such a phenotypic restoration would provide an easily screenable readout of paternal mitochondrial leakage events (Fig. 1a).
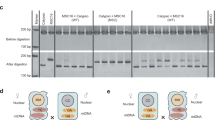
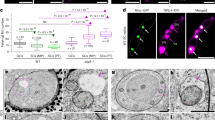
a, Experimental design for the detection of paternal mitochondrial transmission events. The deletion of nad9 from the mitochondrial genome impairs mitochondrial function, leading to delayed seed germination. Upon maternal inheritance, the progeny of Δnad9 mother plants inherit only mutated mitochondrial genomes, thus exhibiting the delayed germination phenotype. This phenotype provides a robust assay for progeny that has inherited functional paternal mitochondria, which restore full respiratory activity and thus wild-type-like germination (encircled seedling). b, Identification of paternal mitochondrial leakage events by germination assays. Pictures were captured on day 10 after sowing. Top: wild-type seeds. Bottom: seeds from crosses using a Δnad9 plant as the maternal parent and a Δdpd1 plant grown at 10 °C (Δdpd1_10 °C) as the pollen donor (Experiment 1). Open circle: a seedling displaying exceptionally fast germination, potentially carrying paternal mitochondria (PMt plantlet). Scale bars, 1 cm. c, Detection of nad9 transcripts by RT–PCR. RNA was extracted from leaf tissues of wild-type (WT), Δnad9 and 5 PMt plants exhibiting exceptionally fast germination. atp9 and nad9 transcripts were co-amplified using gene-specific primers. atp9 served as an internal reference and was detected in all samples. nad9 transcripts were absent from the Δnad9 mutant, but detected in PMt-1, -2, -4 and -5 plants. RT–PCR followed by gel electrophoresis were performed once per sample. d,e, Verification of PMt plants by nuclear and mitochondrial genotyping. Genotypes were analysed by PCR using primers specific to the dpd1 (d) and nad9 (e) deletions. Both deletions were detected in all PMt samples, confirming that the identified PMt plants represent progeny derived from the Δnad9 × Δdpd1_10 °C cross. Simultaneous amplification of the mitochondrial genomic locus encoding the 18S rRNA (rrn18) served as an internal control. PCRs followed by gel electrophoresis were performed once per sample.
To test this strategy, we fertilized emasculated Δnad9 mutant flowers with pollen from chilling-stressed Δdpd1 (defective in pollen organelle DNA degradation 1) mutants carrying wild-type mitochondria. We used the chilling-stressed Δdpd1 mutant as pollen donor, because we recently showed that the combination of low temperature and the Δdpd1 allele significantly promotes paternal plastid transmission14. The DPD1 gene encodes an organellar exonuclease, whose knockout exhibits defective organellar DNA degradation in pollen14,22. We hypothesized that similar conditions would also promote paternal mitochondrial transmission and maximize the likelihood of detecting paternal leakage events. In an initial experiment (Experiment 1, see Methods), a total of 1,000 seeds from the Δnad9 × Δdpd1_10 °C cross were sown, and their germination was monitored by daily inspection. We identified 5 F1 seedlings that germinated at a speed comparable to the wild type (Fig. 1b). When these 5 seedlings had already developed their cotyledons, the rest of the progeny only showed radicle emergence (Fig. 1b). We considered these 5 seedlings as potentially derived from paternal mitochondrial leakage events, and they will hereafter be referred to as PMt plants. To further assess their growth phenotypes, we transferred the PMt seedlings to Magenta boxes. While the PMt-3 plant showed retarded growth and abnormal leaf morphology, closely resembling the characteristic Δnad9 phenotype29, the other four PMt plants displayed growth and development similar to wild-type plants (Extended Data Fig. 3).
To test whether the phenotypic restoration in PMt plants was due to the transmission of paternal mitochondria that provide nad9 expression, RNA was extracted from their leaf and root tissues, followed by polymerase chain reactions with reverse transcription (RT–PCR) with nad9-specific primers to assess nad9 expression. As expected, nad9 transcripts were completely absent from the Δnad9 mutant, but could be readily detected in both leaf and root tissues of PMt-1, -2, -4 and -5 plants. Notably, for PMt-3, nad9 transcripts were detected only in roots (Fig. 1c and Extended Data Fig. 4). The absence of nad9 transcripts from leaf tissues probably explains the leaf developmental defects observed in this line. However, presence of wild-type mitochondria in the roots of PMt-3 suggests that nad9 expression in the root is sufficient to rescue the germination phenotype of the nad9 mutant. Taken together, the presence of nad9 transcripts in all PMt plants provides strong evidence for paternal mitochondrial transmission being the cause of the observed restoration of wild-type-like seed germination.
To exclude the possibility that the PMt plants originated from contamination by wild-type seeds, genotyping experiments were conducted to verify both the nuclear and mitochondrial genotypes of the paternal leakage lines. Since the paternal parent was homozygous for the Δdpd1 allele in the nuclear genome, genuine PMt plants are expected to be heterozygous for Δdpd1. In addition, despite paternal leakage, PMt plants should also inherit Δnad9 mitochondria from the maternal plants. Genotyping experiments confirmed the presence of the nuclear Δdpd1 and the mitochondrial Δnad9 loci in all PMt plants (Fig. 1d,e). Together, these results ultimately confirm that the PMt plants represent bona fide cases of paternal transmission and demonstrate that our experimental strategy is suitable to detect paternal mitochondrial leakage events.
Vegetative segregation of parental mitochondrial genomes
The detection of the Δnad9 locus in the PMt plants (that had been phenotypically rescued by wild-type mitochondria) indicates the coexistence of maternal and paternal mitochondrial genotypes in the tissues examined, a state known as heterochondriomy. To investigate whether plant growth and development is influenced by the relative abundances of the two parental mitochondrial genomes, we transferred PMt plants to soil and cultivated them under standard greenhouse conditions.
We observed three distinct phenotypes in the PMt plants. The first type, exemplified by the PMt-3 plant, exhibited the Δnad9 phenotype characterized by rolled-up leaf margins and curled flower corolla (Fig. 2a). These morphological abnormalities were expected, given the lack of detectable nad9 transcripts in the shoot tissues (Fig. 1c). The second type included PMt-1, -2 and -4, all of which showed wild-type-like growth rates, leaf shape and flower morphology (Fig. 2a), suggesting that nad9 expression originating from paternally inherited mitochondria provided sufficient mitochondrial activity to sustain normal growth and development. The third category, represented by PMt-5, exhibited a chimaeric phenotype. Although nad9 transcripts were detected in its leaf tissues, the PMt-5 plant produced a mix of wild-type-like and Δnad9-like leaves. Notably, the Δnad9 phenotypes became less pronounced as the plant developed, and all emerging flowers resembled those of the wild type (Fig. 2a). This finding suggests a shift in mitochondrial genome ratios over time, with paternally derived nad9 expression eventually reaching levels that restored mitochondrial function to an extent that allowed normal development of reproductive organs.
a, Growth and developmental phenotypes of WT and PMt plants cultivated in the greenhouse. Images were captured at 55 days (top) and 75 days (middle) after sowing. Bottom: close-up views of floral morphology in WT, PMt-3, -4 and -5 plants. PMt-3 displayed a Δnad9-like mutant phenotype, including rolled-up leaf margins, impaired flower development with curly corolla margins and vestigial anthers lacking pollen. By contrast, PMt-4 resembled the WT plant in leaf and floral morphology, and exhibited WT-like pollen production. PMt-5 exhibited signs of heterochondriomy, indicated by a mixture of Δnad9-like (arrowheads) and WT-like leaves (arrows) at early stages (top). However, as the plant continued to grow, later-emerging leaves no longer exhibited the Δnad9 phenotype (middle), and all flowers appeared WT-like with normal pollen production (bottom). Scale bars, 2 cm. b, Confocal microscopy images of pollen from WT, PMt-3 and -4 plants. Pollen viability was assessed using FDA and PI staining. Both WT and PMt-4 plants produce viable pollen, indicated by FDA-derived green fluorescence observed in pollen grains. By contrast, PMt-3 pollen was non-viable, exhibiting only PI fluorescence due to the absence of cytoplasmic esterase activity required for FDA–fluorescein conversion and increased permeability to PI. Scale bars, 50 μm. The pollen viability assay was performed twice per plant and representative images are shown. c, Representative seed capsules that originated from self-pollinated WT and PMt-4 flowers. Except for PMt-3, all PMt plants produced WT-like seed capsules and exhibited normal seed yields. Scale bars, 1 cm.
Paternal mitochondrial transmission reverses cytoplasmic male sterility
Deletion of the nad9 gene from the mitochondrial genome causes CMS in tobacco29. In line with this known phenotypic consequence, we observed a severe reduction in pollen production in the PMt-3 plant that lacked nad9 expression in its shoot tissues. Strikingly, all other PMt plants showed abundant pollen deposition in the anthers (Fig. 2a), suggesting restoration of male fertility. To assess pollen viability, we performed fluorescein diacetate (FDA) and propidium iodide (PI) staining of pollen collected from wild-type and PMt plants. The vast majority of PMt-3 pollen grains were non-viable, as indicated by PI fluorescence (Fig. 2b). By contrast, pollen from wild-type and PMt-4 plants showed predominant green fluorescence from FDA staining, indicating viability (Fig. 2b). We further assessed pollen fertility through pollination assays. Pollen from PMt-1, -2, -4 and -5 plants successfully fertilized flowers upon self- or cross-pollination, resulting in the formation of seed capsules that were similar in size to those of the wild type (Fig. 2c). These findings confirmed the restored male fertility in all PMt plants except PMt-3.
The restoration of male fertility suggests that the paternally derived nad9 gene is present and expressed in reproductive tissues. Hence, it can be expected that the nad9-containing genomes readily enter the female germline and are transmitted to the progeny via maternal inheritance. To investigate whether the paternally derived nad9 genes are inherited to the subsequent F2 generation, we collected individual seed capsules from self-pollinated PMt-4 plants and harvested their seeds separately. Germination assays were performed with seeds from three capsules. While seeds from capsules 1 and 2 showed wild-type-like germination and seedling growth, those of capsule 3 exhibited mixed phenotypes: while most seedlings germinated normally, a subset showed delayed germination and resembled the Δnad9 mutant (Extended Data Fig. 5a). We reasoned that the germ cells that had given rise to capsule 3 contained a mixture of wild-type and Δnad9 mitochondrial genomes. Genotyping experiments revealed that the Δnad9 genome was undetectable in seedlings from capsules 1 and 2, but was present in some seedlings from capsule 3 (Extended Data Fig. 5b). These results indicate that, in PMt plants, the mitochondrial genomes received from both parents continue to segregate during development, and either a single genotype or both genotypes can be incorporated into the germline and transmitted to the next generation.
Unbiased detection of paternal mitochondrial transmission
Our data described above indicated that the relative abundances of parental mitochondrial genomes and their segregation in different cell lineages (somatic versus germline cells) and tissues (root versus shoot) determine the phenotypic outcome of biparental mitochondrial inheritance. We, therefore, reasoned that our phenotype-based screening system, which relies on the detection of restored seed germination time, is inherently influenced by the segregation and proliferation of transmitted paternal mitochondria during embryogenesis. If one assumes that phenotypic restoration requires the presence of a substantial number of paternal mitochondria in the embryo, the sensitivity of our screen may be limited in that only those paternal leakage events are identified that involve a substantial level of paternal mitochondrial transmission. Consequently, the phenotypic screen may underestimate the true frequency of paternal leakage. To address this potential limitation, we set out to develop a more sensitive approach that allows detection of paternal mitochondria also in progeny that do not exhibit phenotypic restoration.
The nuclear genome of tobacco harbours abundant mitochondrial DNA sequences (NUMTs) that originate from intracellular DNA transfer. Some of these NUMTs are very similar in sequence to the mitochondrial nad9 gene (which comes from paternally inherited mitochondria in our screens)29. For this reason, paternal leakage cannot be detected unambiguously by direct amplification of genomic DNA. We, therefore, used nad9 expression as an indicator of paternal mitochondrial leakage. nad9 expression can be very sensitively detected by RT–PCR assays (Fig. 1c), and nad9 transcripts are exclusively derived from mitochondria.
To validate the relationship between nad9 expression levels and paternal leakage, we measured the number of wild-type (paternal) nad9 gene copies and nad9 transcript abundance in additional PMt seedlings (Extended Data Fig. 6a) by quantitative real-time PCR (qPCR) and quantitative PCR with reverse transcription (RT–qPCR), respectively. After quantification and exclusion of nad9-containing NUMTs, we found a strong positive correlation between the number of mitochondrial nad9 gene copies and transcript levels in PMt plants (R2 = 0.9449; Extended Data Fig. 6b). Notably, PMt seedlings with intermediate phenotypes (that is, with some rolled-up leaf margins, resembling the Δnad9 mutant) showed substantial levels of nad9 transcripts (26–55% of the wild-type level), suggesting that seedlings derived from paternal leakage are indistinguishable in morphology from Δnad9 seedlings at low nad9 copy numbers (and transcript levels).
We implemented an RT–PCR-based screening approach, in which we randomly sampled progeny, extracted RNA from whole seedlings and tested for the presence of nad9 transcripts. To increase the number of seedlings that can be assayed while maintaining sensitivity of nad9 transcript detection, we pooled five seedlings per sample for RNA extraction (Fig. 3a). RT–PCR on pooled RNA samples leads to the detection of paternally transmitted nad9 transcripts (Fig. 3b, a representative RT–PCR result showing the detection of nad9 transcripts in 4 out of 19 pools derived from the Δnad9 × Δdpd1_10 °C cross; Experiment 2). In Experiment 1, we detected nad9 transcripts in 8 out of 20 pools derived from the Δnad9 × Δdpd1_10 °C cross, whereas no nad9 transcripts were detected in 20 pools of the control cross (Δnad9 × WT_25 °C). In mitochondria of tobacco, the nad9 mRNA is known to undergo post-transcriptional processing by C-to-U RNA editing at multiple sites28. Sequencing of the amplified RT–PCR products confirmed the presence of the edited nucleotides at the expected sites30, thus unambiguously demonstrating that nad9 was expressed and its mRNA correctly edited inside mitochondria (Fig. 3c). Notably, none of these nad9-positive RNA pools contained seedlings showing phenotypic restoration. These findings demonstrate that the PCR-based screening approach, coupled with validation by complementary DNA (cDNA) sequencing, can reliably and highly sensitively identify paternal leakage events, including events that would go undetected by phenotype-based screening.
a, Experimental design of the PCR-based screening approach for the detection of paternal mitochondrial transmission. Five progeny seedlings were randomly selected and allocated to a single pool for RNA extraction. RT–PCR was performed to assess nad9 expression. Detected nad9 cDNA was subsequently recovered and sequenced to validate transcript identity. b, Example of an RT–PCR analysis performed on RNA pools from progeny derived from Δnad9 × Δdpd1_10 °C crosses (Experiment 2). nad9 transcripts were detected in pools 4, 5, 17 and 18. PCR followed by gel electrophoresis was performed once per sample. c, Sequencing data from a representative nad9-positive sample. Five previously published RNA editing sites (nucleotide positions 298, 328, 368, 398 and 439)29,30 were analysed to assess the RNA editing status of the recovered nad9 cDNA. All five sites were fully edited (arrowheads). d, Quantification of the impact of chilling stress (10 °C), the dpd1 mutation and their combination on paternal mitochondrial inheritance (Experiment 2). Four sets of crosses were analysed, with all 4 using Δnad9 as the maternal parent and the following pollen donors (nuclear background_growth temperature): (1) WT_25 °C (n = 1,105 seedlings); (2) WT_10 °C (n = 386); (3) Δdpd1_25 °C (n = 375); and (4) Δdpd1_10 °C (n = 368). Seedlings displaying WT-like germination were classified as exceptional and were individually processed (that is, without pooling) for nad9 expression analysis. The remaining progenies were pooled before RNA extraction (five seedlings per pool). Positive nad9 amplification signals from pools were considered to derive from a single nad9-positive seedling. The frequency of paternal mitochondrial transmission is the number of nad9-positive seedlings divided by the total number of seedlings analysed. e, Statistical analysis of paternal mitochondrial transmission in Experiment 2. Circles represent the frequency of paternal mitochondrial transmission per group as single data points (no replicates within the experiment). Differences in paternal transmission between each group and the control condition (WT_25 °C) were analysed through a model based on binomial distribution. Individual seedlings are defined as the units of study for the analysis, based on explicit modelling assumptions (see Methods, ‘Modelling and statistics’). Differences were tested simultaneously using two-sided z-tests and corrected for multiple comparisons through the Holm method. Coloured vertical lines: 95% confidence intervals (CI95) of the measured transmission frequency. Effects are expressed as log2 fold change (log2FC). The cross using Δdpd1 plants exposed to 10 °C as pollen donors shows a significant effect compared with the control (log2FC = 5.34; CI95: 2.86–7.82; z = 5.069; P = 1.2 × 10−6), while no significant effects were found for WT_10 °C (log2FC = 2.10; CI95: −0.99 to 5.19; z = 1.600; P = 0.219) and Δdpd1_25 °C (log2FC = 0.55; CI95: −3.59 to 4.71; z = 0.317; P = 0.751). α = 0.05; not significant (NS), P > 0.05; ***P < 0.001.
The exonuclease DPD1 and the temperature during pollen development control mitochondrial inheritance
Leveraging our sensitive PCR-based approach, we next wanted to quantify the effects of low temperature and the Δdpd1 allele on mitochondrial inheritance. To this end, we performed sets of crosses with WT and Δdpd1 plants grown at either 10 °C or 25 °C as pollen donors (Experiment 2; Fig. 3d). The F1 progenies were then screened for paternal leakage using nad9 transcript detection by RT–PCR assays. Contrary to the widely held assumption that mitochondria are strictly maternally inherited, we observed a surprisingly high paternal mitochondrial transmission frequency of 0.18% under standard conditions (that is, in the Δnad9 × WT_25 °C cross). The highest frequency of paternal mitochondrial inheritance, 7.34%, was observed in the Δnad9 × Δdpd1_10 °C cross. By contrast, crosses involving only low temperature (that is, Δnad9 × WT_10 °C) or only the dpd1 deletion allele (Δnad9 × Δdpd1_25 °C) did not show statistically significant differences in paternal transmission frequencies compared with the Δnad9 × WT_25 °C cross (Fig. 3d,e). Together, these results indicate that the combination of low temperature during pollen maturation and Δdpd1 mutation is required to trigger substantial paternal mitochondrial transmission.
Cytological basis of paternal mitochondrial transmission
Given the high paternal transmission frequency observed in the Δnad9 × Δdpd1_10 °C cross, we next sought to identify the cellular processes that are affected in chilling-stressed Δdpd1 pollen. Chilling stress is known to increase plastid abundance in generative cells (GCs), thereby promoting paternal chloroplast transmission14. We therefore speculated that low temperature may likewise enhance mitochondrial abundance in GCs. In addition, we wanted to investigate the potential effects of temperature on mitochondrial integrity by analysing organelle morphology at the ultrastructural level. To this end, we examined mitochondrial distribution and morphology in pollen developed at 10 °C or 25 °C by electron tomography (ET). Three-dimensional GC reconstruction by ET allowed quantitative assessment of the mitochondrial abundance in GCs (Fig. 4a, Extended Data Fig. 7, and Supplementary Videos 1 and 2). Higher numbers of mitochondria were observed in the GCs of pollen grains that had developed at 10 °C versus those at 25 °C (Fig. 4b), suggesting that organelle entry into the GC contributes to the elevated paternal transmission frequency observed in the Δnad9 × Δdpd1_10 °C cross. We did not observe any notable differences in mitochondrial morphology in pollen developed at 10 °C versus those at 25 °C (Fig. 4a and Extended Data Fig. 7), indicating that low temperature does not appreciably affect mitochondrial integrity.
a, Identification of mitochondria in GCs (residing within the cytosol of the vegetative cell) of Δdpd1 pollen developed at 25 °C or 10 °C by electron tomography. Mitochondria were identified on the basis of their characteristic morphology. Serial sections were collected for tomographic analysis. The reconstructed tomograms were used to generate 3D models of the GC with enclosed mitochondria (right). Coloured arrowheads indicate individual mitochondria corresponding to those identified in the respective section layers. Scale bars, 1 μm. For a schematic depiction of a pollen grain with vegetative and generative cell, see Extended Data Fig. 8. b, Number of mitochondria observed in the GC of pollen developed at 25 °C and 10 °C (n = 10 electron tomograms per group). Data points from individual electron tomograms are represented as circles. The difference in means between groups was tested using a t-test for unpaired samples (two-tailed and assuming unequal variances). The effect of low temperature on the abundance of GC mitochondria is significant (difference in means = 3.5; CI95: 1.94–5.06; t = 4.7678; d.f. = 15.272; P = 2.374 × 10−4). α = 0.05; ***P < 0.001. Horizontal lines, median. c, Confocal microscopy images of wild-type (top) and Δdpd1 (bottom) pollen tubes. In vitro-growing pollen tubes were stained with DAPI to visualize nuclear and cytoplasmic DNA. Z-stack images were captured and overlaid to provide an overview of the cytoplasmic DNA abundance in pollen tubes. DAPI signals from the generative cell nucleus were observed in both WT and Δdpd1 pollen tubes (arrowheads), while cytoplasmic DNA was observed exclusively in the Δdpd1 mutant (arrows). Scale bars, 10 μm. Pollen staining followed by microscopy was performed once per genotype and a representative image is shown. d, Confocal microscopy images of a Δdpd1 pollen tube stained with DAPI and MitoTracker. The co-localization of DAPI and MitoTracker signals (arrowheads) suggests retention of mitochondrial DNA in the Δdpd1 mutant. Pollen staining followed by microscopy was performed once and a representative image is shown. Scale bar, 5 μm.
While the presence of mitochondria in GC may enable paternal leakage, the abundance of mitochondrial DNA (mtDNA) in pollen has been proposed as a critical factor governing mitochondrial inheritance15,31. In Arabidopsis, the dually targeted organellar exonuclease DPD1 plays a key role in degrading mtDNA during pollen maturation22, thus potentially restricting paternal transmission. To examine the effect of the dpd1 loss-of-function allele on mtDNA abundance in tobacco pollen, we performed DAPI and MitoTracker staining experiments using in vitro-germinating Δdpd1 pollen tubes. Discrete punctate DAPI fluorescence signals, indicative of cytoplasmic DNA, were observed in Δdpd1 pollen tubes but not in wild-type pollen tubes (Fig. 4c). Importantly, we observed co-localization of the DAPI and MitoTracker fluorescence signals (Fig. 4d), indicating that mtDNA is retained in Δdpd1 pollen.
Together, these findings demonstrate that the combined effects of chilling stress and Δdpd1 mutation lead to an increased abundance of DNA-containing mitochondria in male gametes, thereby promoting paternal mitochondrial transmission.
Discussion
In this study, we report the occurrence of biparental mitochondrial inheritance in tobacco, a species previously believed to inherit its mitochondria strictly maternally. We established phenotypic and RT–PCR-based screens to detect paternal mitochondrial transmission events at high sensitivity. While conventional methods for assaying mitochondrial inheritance have exclusively relied on sequence polymorphisms32,33,34,35,36, our phenotype-based screen identifies paternal leakage through phenotypic restoration, followed by validation through expression analysis and genotyping (Fig. 1). To further improve sensitivity, we developed a PCR-based screening method capable of identifying paternal leakage events even in progeny that do not show phenotypic restoration (Fig. 3). This approach revealed a paternal mitochondrial transmission frequency as high as 7.34% for chilling-stressed dpd1 pollen, which is substantially higher than the 0.5% (5 PMt plants in 1,000 progenies analysed) detected through phenotype-based screening of the same treatment group. A small limitation of this approach lies in the pooling step, where a positive PCR signal from a pooled sample is attributed to a single leakage event, leading to an underestimation of the actual paternal mitochondrial transmission frequency. Correction for this effect (Methods) results in an estimated transmission frequency of 8.7%.
Applying our screening approaches, we showed that the use of chilling-stressed Δdpd1 plants as pollen donors significantly enhances paternal mitochondrial transmission. This finding suggests that the inheritance of both mitochondria and plastids is influenced by a common set of genetic and environmental factors14. However, our findings also reveal differences in the mechanisms controlling the inheritance of the two DNA-containing organelles. For example, neither low temperature nor Δdpd1 mutation alone were sufficient to markedly elevate the paternal transmission frequency of mitochondria (Fig. 3e). This observation stands in stark contrast to what was found previously for plastids, where low temperature and Δdpd1 mutation, on their own, caused a 150-fold and 120-fold increase in paternal transmission, respectively14. The differences between plastid and mitochondrial inheritance observed here are consistent with previous work showing that the inheritance of the two DNA-containing organelles involves distinct mechanisms. For example, in the seed plant cucumber, the mitochondrial genome is inherited paternally, while the chloroplast genome is inherited maternally32.
Chilling stress has been shown to impair the organelle exclusion mechanism in pollen mitosis I—a key barrier that prevents paternal plastid inheritance14. Interference with this mechanism leads to increased inclusion of plastids in the GC14. Our electron tomographic analyses also revealed a higher abundance of mitochondria in the GCs of chilling-stressed pollen than in those developing at 25 °C (Fig. 4b). However, it is worth noting that the exclusion mechanism appears to be inherently less effective for mitochondria than for plastids, as mitochondria are regularly observed in GCs and sperm cells of tobacco37,38. In Arabidopsis, paternal mitochondria have been seen to enter the egg cell upon fertilization16. Interestingly, despite their regular presence in gametes, paternal mitochondrial transmission is considered to be very rare in these species22,29. The prevailing view is that the mere presence of mitochondria in male gametes is insufficient to confer paternal transmission of the mitochondrial genome. Instead, mitochondrial inheritance is primarily governed by mechanisms that regulate mtDNA abundance in male gametes15,39,40,41. Notably, a recent study identified an endonuclease, MTI1, which is expressed in the fertilized egg and selectively degrades maternal mtDNA, thereby causing predominantly paternal mitochondrial inheritance in cucumber42. These findings highlight the critical role of mtDNA abundance in determining inheritance patterns and may explain why an increase in mitochondrial abundance in the GC alone is insufficient to substantially promote paternal transmission (Fig. 3d,e).
In Arabidopsis, DPD1 has been shown to mediate the degradation of both plastid and mitochondrial DNA in maturing pollen22. In line with this, we observed the persistent presence of mtDNA in mature pollen of Δdpd1 tobacco plants (Fig. 4d). Intriguingly, other nucleases have also been implicated in mtDNA degradation during male gametogenesis21. It therefore seems plausible that multiple nucleases act redundantly to mediate mtDNA degradation, and this redundancy may limit the impact of Δdpd1 mutation on mitochondrial inheritance (Fig. 3d,e). By contrast, in chilling-stressed Δdpd1 pollen, the enzymatic activity of all mtDNA-degrading nucleases may be reduced, thus enhancing the mtDNA retention phenotype associated with Δdpd1 mutation. Together with the chilling-induced increase in mitochondrial abundance, the synergistic effect of low temperature and the Δdpd1 allele probably leads to a higher abundance of DNA-containing mitochondria in the GC, accounting for the markedly elevated paternal transmission frequency (Fig. 3d,e).
Unexpectedly, we observed a relatively high paternal mitochondrial transmission frequency of 0.18% under standard conditions, when using greenhouse-grown wild-type plants as pollen donors (Fig. 3d). This frequency is substantially higher than the reported paternal plastid leakage frequency of 0.0016% (ref. 14). Moreover, the combined effect of low temperature and the Δdpd1 mutation causes an even higher paternal leakage for mitochondria (7.34%) than for plastids (2–3%)14 under the same conditions. This could be explained by two factors. First, as discussed above, the organelle exclusion mechanism that effectively prevents paternal plastid transmission appears to be less efficient for mitochondria. Second, we screened for transmission of the paternal nad9 gene, which complements the nad9 defect in the maternal genome. If the paternally transmitted nad9 confers a selective advantage already in the zygote, the paternal mitochondria could preferentially proliferate during embryogenesis. Given that, in the zygote, the paternal organelles are largely outnumbered by the maternal ones15,43, a selective advantage of the transmitted paternal mitochondria could reduce the probability of their loss due to random segregation and sorting out. Previous attempts to detect paternal mitochondrial transmission in Arabidopsis and tobacco have been mostly unsuccessful22,24,44,45, possibly due to the inability to specifically select for paternal mitochondria. Investigating the inheritance of neutral mitochondrial genome variants would offer valuable insight into the possible role of selection in the early maintenance of paternally transmitted mitochondria. Regardless, our findings challenge the widely held belief that paternal leakage is rare and suggest that paternal mitochondrial transmission occurs more frequently than previously assumed.
The occurrence of paternal mitochondrial leakage has profound implications for both plant physiology and evolutionary biology. Consistent with previously reported transgenerational effects of mitochondrial leakage46, our findings show that the inherited paternal mitochondria affect progeny growth and development across generations (Extended Data Fig. 5). Importantly, by promoting paternal mitochondrial transmission, we successfully restored male fertility, offering a novel strategy to manipulate CMS, a trait of great relevance in crop breeding47. CMS typically results from mitochondrial mutations, and the associated male-sterile phenotype cannot be reversed through crosses with wild-type plants due to maternal inheritance48. Paternal mitochondrial transmission offers a novel possibility to restore fertility, which does not even require previous identification of the causal CMS loci or the corresponding nuclear fertility-restorer genes.
From an evolutionary perspective, an intriguing consequence of paternal leakage is the potential recombination between parental mitochondrial genomes upon mitochondrial fusion49,50,51. Genome recombination can introduce genetic diversity and help counteract Muller’s ratchet52,53 especially in taxa harbouring mitochondrial genomes with exceptionally high mutation rates (for example, Silene54 and Plantago55). Interestingly, paternal mitochondrial leakage occurs in Silene vulgaris at rates between 0.2% and 10% (ref. 56). Paternal leakage also has important implications for phylogenetic studies in that it can obscure lineage relationships and thus lead to misinterpretations in phylogenetic analyses57,58.
In summary, we have shown that mitochondrial inheritance is shaped by both environmental and genetic factors. The targeted interference with key cellular processes during male gametogenesis can lead to the breakdown of maternal inheritance, thus opening up new possibilities to employ biparental inheritance as a powerful tool in crop breeding.
Methods
Plant material and growth conditions
Tobacco plants (Nicotiana tabacum cv. Petit Havana) were used for all experiments. The mitochondrial mutant Δnad9 (Δnad9-29 line29) was used as the maternal parent in all crossing experiments. The Δnad9 mutant is homochondriomic, that is, lacks any residual copies of the wild-type mitochondrial genome29. Transplastomic plants (harbouring gfp and aadA expression cassettes in their plastid genome14) with either a dpd1 knockout (Δdpd1) or WT nuclear background were used as paternal parents. The DPD1 exonuclease is encoded by the DPD1S and DPD1T loci in the tobacco nuclear genome. The Δdpd1 mutant used in this study is a doubly homozygous derivative of the previously generated mutant14, and is homozygous for the Δ1/+1 allele in DPD1S and homozygous for a 1.8 kb deletion in DPD1T. In the context of genotyping, detection of the Δdpd1 deletion always refers to the 1.8 kb deletion allele in DPD1T.
Seeds were germinated on agar-solidified Murashige and Skoog (MS) medium59 (0.68% w/v) supplemented with 3% (w/v) sucrose, and the pH adjusted to 5.8. To monitor early plant development, seedlings were transferred to Magenta boxes and cultivated under a 16 h light (25 °C, 50 µmol photons m−2 s−1) and 8 h dark (20 °C) cycle. For continued vegetative and reproductive growth, plants were transferred to soil and grown in a greenhouse under standard conditions (~300 µmol photons m−2 s−1 light intensity) with a 16 h light/8 h dark photoperiod (day/night temperatures: ~25 °C/~20 °C).
Chilling stress experiments were performed in controlled environment chambers maintained at 10 °C (~150 µmol photons m−2 s−1 light intensity, with a 16 h light/8 h dark photoperiod). Paternal plants were first cultivated under standard greenhouse conditions and then transferred to the 10 °C chamber at the onset of flowering. To generate chilling-stressed pollen, the plants were kept at 10 °C throughout the flowering period.
Crosses and screens for paternal mitochondrial transmission
Pollen from Δdpd1 or wild-type plants grown in standard greenhouse conditions or at 10 °C was collected and used for manual pollination. Greenhouse-grown Δnad9 plants were used as the maternal parent. Parental plants were raised and subsequently crossed in two independent experiments: Experiment 1, a small-scale exploratory experiment that included two sets of crosses (Δnad9 × WT_25 °C and Δnad9 × Δdpd1_10 °C), and Experiment 2, a large-scale experiment designed to (1) replicate the findings of Experiment 1 for the two crosses performed and (2) explore the effects of individual factors by including two additional sets of crosses (Δnad9 × WT_10 °C and Δnad9 × Δdpd1_25 °C). Seeds resulting from these four sets of crosses were harvested and screened for paternal mitochondrial inheritance. To prevent self-pollination, all flowers used for crosses were emasculated, including the flowers of the Δnad9 mutant (to make sure that, even if there was low-level leakiness of the CMS trait, there is no risk of self-pollination).
To assess paternal mitochondrial transmission, the progenies were germinated on MS medium solidified with 0.68% (w/v) agar and supplemented with 3% (w/v) sucrose (pH 5.8), at a density of 100 seedlings per 120 mm square plate (10 seeds per row, 10 rows per plate). Plates were incubated in controlled environment chambers under a 16 h light (25 °C, 50 µmol photons m−2 s−1) and 8 h dark (20 °C) cycle. Germination and early seedling development were monitored daily by visual inspection. Paternal leakage was detected using two approaches: phenotype-based screening of the seedlings (by evaluating germination and growth rates) and PCR-based molecular screening (by randomly selecting seedlings for RNA extraction followed by RT–PCR analysis).
RNA extraction and RT–PCR for paternal mitochondrial transmission screens
Leaf and root tissues were harvested from WT, Δnad9 and PMt plants for RNA extraction using the NucleoSpin RNA Plant kit (Macherey-Nagel), following manufacturer protocol except that the DNase digestion step was deliberately omitted. Reverse transcription was carried out using SuperScript III reverse transcriptase (Invitrogen) with oligonucleotides oJF1113 and oJF1144 as primers (Extended Data Table 1), which bind to the 3′ UTRs of nad9 and atp9, respectively. The synthesized cDNA was purified with the NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel) and used as template for PCR with primers P1 (binding to the nad9 CDS), oJF1145 (binding to the atp9 5′ UTR) and P2 (an adapter-specific primer binding to the 5′ terminal sequence of both oligonucleotides oJF1113 and oJF1144, thus excluding amplification of residual DNA present in the RNA preparations; Extended Data Table 1). The expected PCR product sizes were 465 bp for nad9 and 568 bp for atp9. PCR products were resolved and visualized via electrophoresis in 2% agarose gels. Recovered PCR products were analysed by Sanger sequencing to determine the RNA editing pattern of the nad9 cDNA. PCR reactions were performed using GoTaq DNA polymerase (Promega) according to manufacturer instructions. All oligonucleotides used in this work were generated by chemical synthesis and acquired from a commercial provider (Eurofins).
Measurements of nad9 gene copy numbers and relative transcript abundances by qPCR and RT–qPCR
Wild-type, Δnad9 and PMt samples derived from flash-frozen whole seedlings were divided for extraction (in parallel) of genomic DNA and RNA. Total genomic DNA was isolated following a cetyltrimethylammonium bromide (CTAB)-based protocol60. Total RNA was isolated using the NucleoSpin RNA Plant kit (Macherey-Nagel) following manufacturer instructions, including the recommended DNase treatment step. Synthesis of cDNA was carried out with the SuperScript III Reverse Transcriptase kit (Invitrogen) following manufacturer instructions, using 250 ng total RNA and 2.5 μM random hexamer primer in a reaction volume of 40 μl. Negative control reactions (from which the reverse transcriptase was omitted) were done in parallel and included in downstream measurements. All measurements were carried out in technical triplicates using the LightCycler 480 Real-Time PCR System and LightCycler SYBR Green reaction mixtures (Roche) in reaction volumes of 5 μl. Purified genomic DNA (5 ng) or 1/200 of the cDNA reaction was used as template. All oligonucleotide primers target regions within single exons so that amplicons derived from genomic DNA or cDNA templates are of identical size. Transcript levels and relative gene copy numbers of nad9 were calculated relative to those of the nuclear reference genes L25 (ref. 61) or TUBB2, using an efficiency-adjusted ΔCq method62.
Genotyping experiments
DNA was isolated from leaf tissues of WT, Δnad9 and PMt plants, and used as template for PCR reactions. Primer pairs (1) P3–P4, (2) oJF883–oJF884 and (3) oEG327–oEG332 (Extended Data Table 1) were employed to detect Δnad9 mitochondrial genomes, the mitochondrial 18S rRNA gene rrn18 (as an internal control) and the Δdpd1 allele in the nuclear genome, respectively. PCR reactions were performed using DreamTaq DNA polymerase (Thermo Fisher) according to manufacturer instructions.
Pollen viability assay
To assess pollen viability, anthers were detached from WT and PMt flowers at anthesis, and collected in a 5 ml Eppendorf tube. Pollen grains were collected by vortexing with pollen harvesting buffer (100 mM Na3PO4 (pH 7.0) and 1 mM Na2EDTA (pH 8.0)), followed by centrifugation at 1,000 × g for 5 min. The pollen pellet was then resuspended and stained in pollen viability solution (100 mM Na3PO4 (pH 7.0), 1 mM Na2EDTA (pH 8.0), 10 μM PI (Abcam) and 20 μg ml−1 FDA (Sigma)) at room temperature for 30 min. The stained samples were imaged by confocal laser-scanning microscopy (TCS SP8, Leica). FDA was excited with a 488 nm argon laser and the emission signal was detected at 500–550 nm. PI was excited with a 561 nm argon laser and its emission signal was detected at 600–650 nm.
Electron tomography
Pollen samples for electron microscopic analysis were processed through high-pressure freezing, freeze substitution, resin embedding and ultramicrotomy as previously described63. Pollen was collected from Δdpd1 plants grown under standard (22–25 °C) and low-temperature (10 °C) conditions. The pollen samples were placed in 3 mm planchettes for high-pressure freezing, followed by freeze substitution at −80 °C for 24 h, with gradual warming to room temperature over 36 h. After rinsing three times with acetone to remove any excess osmium tetroxide, pollen samples were subsequently embedded in Embed-812 resin (Electron Microscopy Sciences, 14120). Thin sections (90 nm thick) were prepared from sample blocks of each stage and examined with a Hitachi 7400 TEM operated at 80 kV. Serial thick sections (300 nm thick) were collected for tomography. Tilt series were collected from 60° to −60° (1.5° intervals) with a 200 kV Tecnai F20 intermediate voltage electron microscope (FEI). For dual-axis tomography analysis, tilt series around two orthogonal axes were acquired from each section using the microscope’s tomography programme. Tomograms were reconstructed as described64. Tomograms were collected using TOMOGRAPHY v.4.0.3.4538 and reconstructed using Etomo v.5.0.0 10. Tomogram segmentation was performed using IMOD v.5.0.2. For three-dimensional (3D) modelling of generative cells and their enclosed mitochondria, contours were manually traced using the ‘3dmod’ software package. The generative cell membrane, the nucleus and the mitochondria were outlined in every 8th slice, with manual corrections applied as needed. Final 3D models were generated using the ‘imodmesh’ command to mesh the corrected contours.
DAPI and MitoTracker staining and confocal laser-scanning microscopy
To visualize mitochondrial DNA, DAPI and MitoTracker staining was performed with in vitro-germinating pollen tubes. Mature pollen grains were collected from either WT or Δdpd1 plants and suspended in pollen germination buffer (1.6 mM boric acid, 3.0 mM calcium nitrate, 1.0 mM potassium nitrate, 0.8 mM magnesium sulfate, 10 % sucrose, pH 7.4), followed by an incubation at room temperature for 2–3 h to allow pollen tube germination. Pollen tubes were stained with 4,6-diamidino-2-phenylindole (DAPI) solution (100 mM Na₃PO₄ (pH 7.0), 0.1% (v/v) Triton X-100, 1 mM Na2EDTA (pH 8.0) and 1 μg ml−1 DAPI) and 100 nM MitoTracker Red CMXRos (Invitrogen, M7512) for 30 min at room temperature. DAPI and MitoTracker fluorescence signals were imaged by confocal laser-scanning microscopy (TCS SP8, Leica), using (1) a 405 nm laser diode for excitation and a 430–495 nm filter for detection of DAPI fluorescence emission, and (2) a 561 nm laser for excitation and 585–615 nm for detection of MitoTracker fluorescence emission.
Modelling and statistics
All statistical tests and assembly of statistical models were done using R v.4.3.3 (https://www.R-project.org/). The detection of the nad9 transcript in pools and individual samples by RT–PCR was used as a proxy for the transmission of paternal mitochondria. Modelling for analysis of Experiment 2 was performed by considering individual seedlings as the unit of study based on the following assumptions: (1) paternal transmission of mitochondria is the result of a binomially distributed process (that is, a seedling being nad9-positive is one out of two possible outcomes and occurs at a set probability that defines the binomial process), (2) transmission events observed in individual seedlings are independent of each other and (3) for all samples derived from pooled seedlings, the detected nad9 transcripts originated from a single seedling in the pool. Paternal mitochondrial transmission was then calculated as the percentage of nad9-positive seedlings.
The variation in paternal mitochondrial transmission in response to different combinations of nuclear genotype (Δdpd1 or wild-type) and temperature during pollen development (25/20 °C or 10 °C) was analysed through a binomial model constructed with the maximum likelihood method in R. The model was assembled using the glm() function, using ‘log’ as the link function. Parameters in the model represent the difference between a given group (a combination of genotype and temperature) and the control condition (wild type at 25 °C). Hypothesis testing of the differences between these groups was performed using the multcomp v.1.4-25 package. Null hypotheses were stated using the glht() function and tested simultaneously with z-tests using the cftest() function. To account for multiple comparisons, P values were corrected through the Holm method. Simultaneous 95% confidence intervals were obtained using the confint() function. The following packages were used to provide adequate code functionality: ggplot2 v.3.5.1, ggsignif v.0.6.4, rstudioapi v.0.16.0, MASS v.7.3-60.0.1 and dplyr v.1.1.4.
A corrected estimate of the paternal transmission frequency of mitochondria in Experiment 2 (that accounts for multiple transmission events in a single pool) was obtained for the Δdpd1 10 °C group using Monte Carlo simulations in R. For simplicity and only for this calculation, we considered all nad9-positive samples obtained experimentally for this group (7 from individual seedlings and 20 from pools) as pool derived and assumed that the frequency of paternal mitochondrial inheritance (that is, the number of individual nad9-positive seedlings among the total) follows a binomial distribution. After stipulating a set of possible transmission frequencies, a fixed pool size (5 seedlings) and a fixed number of samples (74 pools), we simulated the random allocation of nad9-positive seedlings to pools (10,000 simulations). For each given transmission frequency, we calculated the average of nad9-positive pools (samples) obtained across all simulations. The corrected transmission frequency estimate (8.7%) is the value that produced an average of nad9-positive samples equal to the experimental value (27).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this work are available within the paper, its extended data figures and tables, and Supplementary Videos. NCBI entry NC_006581.1 was used as the reference mitochondrial genome for tobacco. Source data are provided with this paper.
Code availability
The R code used for statistical tests and the model is available via Zenodo at https://doi.org/10.5281/zenodo.18684523 (ref. 65).
References
Camus, M. F., Alexander-Lawrie, B., Sharbrough, J. & Hurst, G. D. Inheritance through the cytoplasm. Heredity 129, 31–43 (2022).
Maliga, P. Engineering the plastid and mitochondrial genomes of flowering plants. Nat. Plants 8, 996–1006 (2022).
Bock, R. Witnessing genome evolution: experimental reconstruction of endosymbiotic and horizontal gene transfer. Annu. Rev. Genet. 51, 1–22 (2017).
Gualberto, J. M. & Newton, K. J. Plant mitochondrial genomes: dynamics and mechanisms of mutation. Annu. Rev. Plant Biol. 68, 225–252 (2017).
Green, B. R. Chloroplast genomes of photosynthetic eukaryotes. Plant J. 66, 34–44 (2011).
Greiner, S. & Bock, R. Tuning a ménage à trois: co-evolution and co-adaptation of nuclear and organellar genomes in plants. BioEssays 35, 354–365 (2013).
Hanson, M. R. Plant mitochondrial mutations and male sterility. Annu. Rev. Genet. 25, 461–486 (1991).
Hagemann, R. The foundation of extranuclear inheritance: plastid and mitochondrial genetics. Mol. Genet. Genomics 283, 199–209 (2010).
Perlman, S. J., Hodson, C. N., Hamilton, P. T., Opit, G. P. & Gowen, B. E. Maternal transmission, sex ratio distortion, and mitochondria. Proc. Natl Acad. Sci. USA 112, 10162–10168 (2015).
Birky, C. W. Jr. The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Annu. Rev. Genet. 35, 125–148 (2001).
Azhagiri, A. K. & Maliga, P. Exceptional paternal inheritance of plastids in Arabidopsis suggests that low-frequency leakage of plastids via pollen may be universal in plants. Plant J. 52, 817–823 (2007).
Schneider, A. et al. Low frequency paternal transmission of plastid genes in Brassicaceae. Transgenic Res. 24, 267–277 (2015).
Ruf, S., Karcher, D. & Bock, R. Determining the transgene containment level provided by chloroplast transformation. Proc. Natl Acad. Sci. USA 104, 6998–7002 (2007).
Chung, K. P., Gonzalez-Duran, E., Ruf, S., Endries, P. & Bock, R. Control of plastid inheritance by environmental and genetic factors. Nat. Plants 9, 68–80 (2023).
Wang, D.-Y. et al. The levels of male gametic mitochondrial DNA are highly regulated in angiosperms with regard to mitochondrial inheritance. Plant Cell 22, 2402–2416 (2010).
Matsushima, R. et al. Mitochondrial dynamics in plant male gametophyte visualized by fluorescent live imaging. Plant Cell Physiol. 49, 1074–1083 (2008).
Chung, K. P. Cytoplasmic inheritance: the transmission of plastid and mitochondrial genomes across cells and generations. Plant Physiol. 198, kiaf168 (2025).
Birky, C. W. Jr. Uniparental inheritance of mitochondrial and chloroplast genes: mechanisms and evolution. Proc. Natl Acad. Sci. USA 92, 11331–11338 (1995).
Nagata, N., Saito, C., Sakai, A., Kuroiwa, H. & Kuroiwa, T. The selective increase or decrease of organellar DNA in generative cells just after pollen mitosis one controls cytoplasmic inheritance. Planta 209, 53–65 (1999).
Kuroiwa, T. Review of cytological studies on cellular and molecular mechanisms of uniparental (maternal or paternal) inheritance of plastid and mitochondrial genomes induced by active digestion of organelle nuclei (nucleoids). J. Plant Res. 123, 207–230 (2010).
Ma, F. et al. The mitochondrial endonuclease M20 participates in the down-regulation of mitochondrial DNA in pollen cells. Plant Physiol. 178, 1537–1550 (2018).
Matsushima, R. et al. A conserved, Mg2+-dependent exonuclease degrades organelle DNA during Arabidopsis pollen development. Plant Cell 23, 1608–1624 (2011).
Medgyesy, P., Páy, A. & Márton, L. Transmission of paternal chloroplasts in Nicotiana. Mol. Gen. Genet. 204, 195–198 (1986).
Thyssen, G., Svab, Z. & Maliga, P. Exceptional inheritance of plastids via pollen in Nicotiana sylvestris with no detectable paternal mitochondrial DNA in the progeny. Plant J. 72, 84–88 (2012).
Arimura, S.-i. & Nakazato, I. Genome editing of plant mitochondrial and chloroplast genomes. Plant Cell Physiol. 65, 477–483 (2024).
Kang, B.-C. et al. Chloroplast and mitochondrial DNA editing in plants. Nat. Plants 7, 899–905 (2021).
Nakazato, I. et al. Targeted base editing in the mitochondrial genome of Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 119, e2121177119 (2022).
Forner, J. et al. Targeted introduction of heritable point mutations into the plant mitochondrial genome. Nat. Plants 8, 245–256 (2022).
Forner, J. et al. Targeted knockout of a conserved plant mitochondrial gene by genome editing. Nat. Plants 9, 1818–1831 (2023).
Fang, J. et al. Dynamic landscape of mitochondrial cytidine-to-uridine RNA editing in tobacco (Nicotiana tabacum) shows its tissue specificity. Plant Cell Tiss. Organ Cult. 148, 363–376 (2022).
Zhang, Q., Liu, Y. & Sodmergen Examination of the cytoplasmic DNA in male reproductive cells to determine the potential for cytoplasmic inheritance in 295 angiosperm species. Plant Cell Physiol. 44, 941–951 (2003).
Park, H.-S. et al. Inheritance of chloroplast and mitochondrial genomes in cucumber revealed by four reciprocal F1 hybrid combinations. Sci. Rep. 11, 2506 (2021).
Pearl, S. A., Welch, M. E. & McCauley, D. E. Mitochondrial heteroplasmy and paternal leakage in natural populations of Silene vulgaris, a gynodioecious plant. Mol. Biol. Evol. 26, 537–545 (2009).
Wagner, D., Dong, J., Carlson, M. & Yanchuk, A. Paternal leakage of mitochondrial DNA in Pinus. Theor. Appl. Genet. 82, 510–514 (1991).
Bulgakova, V., Shatskaya, N., Kosterin, O. & Vasiliev, G. Cases of paternal inheritance and recombination of mitochondrial DNA in peas (Pisum L.). Euphytica 219, 74 (2023).
Reboud, X. & Zeyl, C. Organelle inheritance in plants. Heredity 72, 132–140 (1994).
Yu, H.-S. & Russell, S. D. Occurrence of mitochondria in the nuclei of tobacco sperm cells. Plant Cell 6, 1477–1484 (1994).
Yu, H.-S. & Russell, S. D. Populations of plastids and mitochondria during male reproductive cell maturation in Nicotiana tabacum L.: a cytological basis for occasional biparental inheritance. Planta 193, 115–122 (1994).
Birky, C. W. Uniparental inheritance of organelle genes. Curr. Biol. 18, R692–R695 (2008).
Sodmergen, Z. Q., Zhang, Y., Sakamoto, W. & Kuroiwa, T. Degradation of mitochondrial genome during sperm cell development in Hordeum vulgare: a mechanism for maternal inheritance of mitochondria in angiosperms. Planta 216, 235–244 (2002).
Nagata, N., Saitoa, C., Sakaia, A., Kuroiwa, H. & Kuroiwa, T. Decrease in mitochondrial DNA and concurrent increase in plastid DNA in generative cells of Pharbitis nil during pollen development. Eur. J. Cell Biol. 78, 241–248 (1999).
Shen, J. et al. A nuclear-encoded endonuclease governs the paternal transmission of mitochondria in Cucumis plants. Nat. Commun. 16, 4266 (2025).
Wolff, J. & Gemmell, N. J. Lost in the zygote: the dilution of paternal mtDNA upon fertilization. Heredity 101, 429–434 (2008).
Avni, A. & Edelman, M. Direct selection for paternal inheritance of chloroplasts in sexual progeny of Nicotiana. Mol. Gen. Genet. 225, 273–277 (1991).
Horlow, C., Goujaud, J., Lépingle, A., Missonier, C. & Bourgin, J.-P. Transmission of paternal chloroplasts in tobacco (Nicotiana tabacum). Plant Cell Rep. 9, 249–252 (1990).
Shen, J. et al. Rare maternal and biparental transmission of the cucumber mitochondrial DNA reveals sorting of polymorphisms among progenies. Theor. Appl. Genet. 132, 1223–1233 (2019).
Kim, Y.-J. & Zhang, D. Molecular control of male fertility for crop hybrid breeding. Trends Plant Sci. 23, 53–65 (2018).
Chen, Z. et al. Plant mitochondrial genome evolution and cytoplasmic male sterility. Crit. Rev. Plant Sci. 36, 55–69 (2017).
McCauley, D. E. Paternal leakage, heteroplasmy, and the evolution of plant mitochondrial genomes. New Phytol. 200, 966–977 (2013).
Munasinghe, M. & Ågren, J. A. When and why are mitochondria paternally inherited? Curr. Opin. Genet. Dev. 80, 102053 (2023).
Gurdon, C., Svab, Z., Feng, Y., Kumar, D. & Maliga, P. Cell-to-cell movement of mitochondria in plants. Proc. Natl Acad. Sci. USA 113, 3395–3400 (2016).
Greiner, S., Sobanski, J. & Bock, R. Why are most organelle genomes transmitted maternally? BioEssays 37, 80–94 (2015).
Radzvilavicius, A. & Johnston, I. G. Paternal leakage of organelles can improve adaptation to changing environments. Preprint at bioRxiv https://doi.org/10.1101/2020.12.18.423500 (2020).
Sloan, D. B. et al. Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol. 10, e1001241 (2012).
Cho, Y., Mower, J. P., Qiu, Y. L. & Palmer, J. D. Mitochondrial substitution rates are extraordinarily elevated and variable in a genus of flowering plants. Proc. Natl Acad. Sci. USA 101, 17741–17746 (2004).
Bentley, K. E., Mandel, J. R. & McCauley, D. E. Paternal leakage and heteroplasmy of mitochondrial genomes in Silene vulgaris: evidence from experimental crosses. Genetics 195, 961–968 (2010).
White, D. J., Wolff, J. N., Pierson, M. & Gemmell, N. J. Revealing the hidden complexities of mtDNA inheritance. Mol. Ecol. 17, 4925–4942 (2008).
Wolff, J. N., Gandre, S., Kalinin, A. & Gemmell, N. J. Delimiting the frequency of paternal leakage of mitochondrial DNA in chinook salmon. Genetics 179, 1029–1032 (2008).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Doyle, J. J. & Doyle, J. L. Isolation of plant DNA from fresh tissue. Focus 12, 13–15 (1990).
Schmidt, G. W. & Delaney, S. K. Stable internal reference genes for normalization of real-time RT-PCR in tobacco (Nicotiana tabacum) during development and abiotic stress. Mol. Genet. Genomics 283, 233–241 (2010).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Liang, Z. et al. Electron tomography of prolamellar bodies and their transformation into grana thylakoids in cryofixed Arabidopsis cotyledons. Plant Cell 34, 3830–3843 (2022).
Toyooka, K. & Kang, B.-H. Reconstructing plant cells in 3D by serial section electron tomography. Methods Mol. Biol. 1080, 159–170 (2014).
Gonzalez-Duran, E. R code for statistics and modelling of mitochondrial transmission. Zenodo https://doi.org/10.5281/zenodo.18684523 (2026).
Acknowledgements
We thank A. Nagarkar for help with crosses and germination assays. This research was funded by the Max Planck Society, and grants from the European Research Council (ERC-STG-2024 grant agreement no. 101165903 to K.P.C., and ERC-ADG-2023 grant agreement no. 101140768 to R.B.).
Funding
Open access funding provided by Max Planck Society.
Author information
Authors and Affiliations
Contributions
K.P.C. and R.B. designed the research. E.G.-D., Z.L., W.W., D.K. and K.P.C. performed the experiments. E.G.-D., Z.L., J.F., D.K., W.W., L.J., K.P.C. and R.B. contributed to data analysis and interpretation. K.P.C. wrote the paper draft, R.B. revised it, and E.G.-D., J.F. and Z.L. provided additional input.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Pal Maliga and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic representation of the wild-type and the Δnad9 mitochondrial genomes.
The region from nucleotide position 331,000 to 338,000 (based on the Nicotiana tabacum reference genome; NC_006581) is shown. a, Map of the nad9 region in the wild-type mitochondrial genome. The nad9 coding sequence is located between nucleotide positions 336,111 and 336,683. The nad9-specific primers P1 and P2 were used to detect nad9 transcripts. b, Map of the corresponding genomic region in the mitochondrial genome of the Δnad9 mutant. A 3,809-nucleotide segment of the mitochondrial genome has been deleted (indicated by red arrowheads), leading to the complete loss of the nad9 gene. To detect the deletion, primers P3 and P4 were used that result in amplification of a short 325 bp fragment.
Extended Data Fig. 2 The Δnad9 mutant exhibits delayed seed germination and retarded seedling growth.
Surface-sterilized wild-type and Δnad9 seeds were sown on MS medium supplemented with 3% sucrose, stratified for two days, and then transferred to a growth chamber. Each plate contains 100 seeds. Pictures of wild-type and Δnad9 seedlings were taken on day 14 after sowing. Scale bars, 1 cm.
Extended Data Fig. 3 Vegetative growth phenotypes of PMt plants.
a-g, Seedlings were transferred from germination plates to Magenta boxes. Pictures were captured on day 40 after sowing. Note the stunted growth exhibited by the Δnad9 mutant (b). While PMt-1, PMt-2, PMt-4, and PMt-5 (c,d,f,g) display a growth phenotype similar to wild-type plants, the PMt-3 plant (e) shows a Δnad9-like phenotype, consistent with the absence of nad9 transcripts from its leaf tissues (see Fig. 1c). Scale bars, 1 cm.
Extended Data Fig. 4 Presence of nad9 transcripts in root tissues of PMt plants.
RNA was extracted from the root tissues of wild-type, Δnad9, and PMt plants. atp9 and nad9 transcripts were co-amplified using gene-specific primers (see Methods), with atp9 serving as an internal control. nad9 transcripts are detected in all samples except for the Δnad9 mutant. RT-PCR followed by gel electrophoresis were performed once per sample.
Extended Data Fig. 5 Transmission of paternal mitochondria into the next generation.
a, F2 seedlings derived from individual capsules of the self-pollinated PMt-4 plant growing in MS medium. Pictures were taken on day 14 after sowing. While the majority of seedlings exhibit a WT-like germination and growth rate, some seedlings of capsule 3 show delayed germination and growth (indicated by white circles), suggesting the persistence of Δnad9 mitochondrial genomes in the progeny of this capsule. Scale bars, 1 cm. b, PCR amplification of Δnad9 mitochondrial genomes in the F2 generation. Five seedlings were collected at random and grouped into pools, with three pools (1 – 3) generated per seed capsule. DNA was extracted from each pool, and PCR was performed using primers designed to amplify the nad9 deletion locus (see Extended Data Fig. 1B). The nad9 deletion locus was detected in Pool 3 of Capsule 3, indicating that both paternal and maternal mitochondria are retained in the germ cells of the PMt-4 plant and can be transmitted to the next generation. Amplification of the mitochondrial genomic locus encoding the 18S rRNA (rrn18) served as an internal control.
Extended Data Fig. 6 Correlation between phenotypic restoration, nad9 copy number and transcript abundance in PMt plants.
a, Phenotypes of PMt seedlings (cyan arrows) 28 days after sowing. Plants with partial phenotypic restoration display features reminiscent of the Δnad9 mutant, such as the typical rolled-up leaf margins (white arrows). Scale bars, 1 cm. b, Relationship between relative nad9 transcript abundance and the number of wild-type nad9 gene copies (of paternal origin in PMt plants), as quantified by RT-qPCR and qPCR, respectively. Each circle corresponds to measurements from a whole seedling (biological replicate), and represents mean values from three independent quantification experiments, each using a different combination of nad9 amplicon and reference gene: (i) nad9 amplicon A / L25, (ii) nad9 amplicon B / L25, and (iii) nad9 amplicon A / TUBB2 (Supplementary Table 1). Within one quantification experiment, all measurements were done in triplicate in the same 384-well PCR plate. nad9 transcript abundance was determined relative to the transcript abundance of a nuclear reference gene, and in parallel, preliminary nad9 copy numbers (that is, including nad9-containing NUMTs) were calculated in relation to gene copies of the same reference gene. NUMTs were quantified in the Δnad9 mutants [14.5 ± 0.4; mean ± standard error of the mean (SEM) across experiments] and subtracted from the preliminary values to yield the wild-type mitochondrial nad9 copy numbers per haploid nuclear genome. Error bars: ±SEM of relative transcript abundance (vertical) or gene copy number (horizontal) across experiments.
Extended Data Fig. 7 Electron tomography of generative cells of Δdpd1 pollen developed at 10 °C or 25 °C.
Sections of pollen samples were collected for tomographic analysis. Tomograms were reconstructed from tilt series covering a significant volume of the GCs to quantify mitochondrial abundance. Pollen harvest, followed by sample preparation and electron microscopy, was performed once. Scale bars, 1 μm.
Extended Data Fig. 8 Schematic depiction of a pollen grain showing the generative cell (GC) embedded within the vegetative cell (VC).
The nuclei (Nu) of the two cells are indicated. Plastids (Pt) and mitochondria (Mt) are largely excluded from the GC, but two mitochondria are shown here within the GC to illustrate the paternal leakage reported in this work (cf. Figure 4a).
Supplementary information
Supplementary Video 1 (download MOV )
Serial tomographic sections of Δdpd1 pollen developed at 25 °C. Mature pollen was collected from Δdpd1 plants grown at 25 °C and prepared for electron tomography analysis. Serial thick sections (300 nm) were obtained for tomographicreconstruction. A representative video displaying the serial sections from one tomogram is shown.
Supplementary Video 2 (download MOV )
Serial tomographic sections of Δdpd1 pollen developed at 10 °C. Mature pollen was collected from Δdpd1 plants grown at 10 °C and prepared for electron tomography analysis. Serial thick sections (300 nm) were obtained for tomographicreconstruction. A representative video displaying the serial sections from one tomogram is shown.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed gels for Fig. 1c–e.
Source Data Fig. 3 (download PDF )
Unprocessed gels for Fig. 3b.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed gels.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed gels for Extended Data Fig. 5b.
Source Data Extended Data Fig. 6 (download XLSX )
qPCR data and calculations for Extended Data Fig. 6b.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gonzalez-Duran, E., Liang, Z., Forner, J. et al. High-frequency biparental inheritance of plant mitochondria upon chilling stress and loss of a genome-degrading nuclease. Nat. Plants 12, 571–582 (2026). https://doi.org/10.1038/s41477-026-02242-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-026-02242-7