Abstract
Similar to the evolutionary history of all species on Earth, the developmental lineage history of all cells in a multicellular organism is stored in a phylogenetic tree. Mapping the zygote-to-adult developmental cell phylogeny of a complex organism is of tremendous value but technically challenging. We here developed e-SMALT, a powerful cell lineage tracing system integrated with single-cell RNA sequencing, in Arabidopsis thaliana to record the zygote-to-adult developmental lineages of two plant individuals. The system performed efficiently in A. thaliana, with an average of ~50 barcoding mutations accumulated on the 1-kb barcode sequence of each cell in 3-month-old plants. Using the barcoding mutations, we reconstructed the phylogenetic tree for thousands of cells sampled from various shoot branches of each plant, with high statistical confidence and at single-cell resolution. The cell phylogenies show that cells of every shoot branch are derived from exactly three founder cells, each belonging to one of three early-determined lineages. The three-cell pattern holds for primary, secondary and tertiary branches, and even for single flowers/siliques. Incorporating single-cell RNA sequencing data revealed the three founder cells responsible for establishing the three germ layers of each branch/organ, which in turn updates our understanding of plant germ layers to single-cell resolution. We further showed that the three-cell rule reflects an adaptive strategy for an indeterminate plant to manage its stem cell pool, suggesting an analytical framework to unify the distinct strategies between plants and animals in organogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw sequence data of lineage barcode amplicons and single-cell transcriptomics have been deposited in the Genome Sequence Archive in the National Genomics Data Center (https://ngdc.cncb.ac.cn/gsa) with accession number CRA016061.
Code availability
The software sources and core parameters are provided in Methods. The custom data analysis scripts are available via GitHub at https://github.com/XFannv/eSMALT.git.
References
Frumkin, D., Wasserstrom, A., Kaplan, S., Feige, U. & Shapiro, E. Genomic variability within an organism exposes its cell lineage tree. PLoS Comput. Biol. 1, e50 (2005).
Salipante, S. J. & Horwitz, M. S. Phylogenetic fate mapping. Proc. Natl Acad. Sci. USA 103, 5448–5453 (2006).
Behjati, S. et al. Genome sequencing of normal cells reveals developmental lineages and mutational processes. Nature 513, 422–425 (2014).
Park, S. et al. Clonal dynamics in early human embryogenesis inferred from somatic mutation. Nature 597, 393–397 (2021).
Spencer Chapman, M. et al. Lineage tracing of human development through somatic mutations. Nature 595, 85–90 (2021).
Mitchell, E. et al. Clonal dynamics of haematopoiesis across the human lifespan. Nature 606, 343–350 (2022).
Weng, C. et al. Deciphering cell states and genealogies of human haematopoiesis. Nature 627, 389–398 (2024).
Stadler, T., Pybus, O. G. & Stumpf, M. P. H. Phylodynamics for cell biologists. Science 371, eaah6266 (2021).
Domcke, S. & Shendure, J. A reference cell tree will serve science better than a reference cell atlas. Cell 186, 1103–1114 (2023).
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Satina, S., Blakeslee, A. F. & Avery, A. G. Demonstration of the three germ layers in the shoot apex of Datura by means of induced polyploidy in periclinal chimeras. Am. J. Bot. 27, 895–905 (1940).
Burk, L. G., Stewart, R. N. & Dermen, H. Histogenesis and genetics of a plastid-controlled chlorophyll variegation in tobacco. Am. J. Bot. 51, 713–724 (1964).
Stewart, R. N. & Burk, L. G. Independence of tissues derived from apical layers in ontogeny of the tobacco leaf and ovary. Am. J. Bot. 57, 1010–1016 (1970).
Stewart, R. N., Semeniuk, P. & Dermen, H. Competition and accommodation between apical layers and their derivatives in the ontogeny of chimeral shoots of Pelargonium × hortorum. Am. J. Bot. 61, 54–67 (1974).
Stewart, R. N. & Dermen, H. Ontogeny in monocotyledons as revealed by studies of the developmental anatomy of periclinal chloroplast chimeras. Am. J. Bot. 66, 47–58 (1979).
Poethig, R. S. Clonal analysis of cell lineage patterns in plant development. Am. J. Bot. 74, 581–594 (1987).
Furner, I. J., Ainscough, J. F. X., Pumfrey, J. A. & Petty, L. M. Clonal analysis of the late flowering fca mutant of Arabidopsis thaliana: cell fate and cell autonomy. Development 122, 1041–1050 (1996).
Furner, I. J. & Pumfrey, J. E. Cell fate in the shoot apical meristem of Arabidopsis thaliana. Development 115, 755–764 (1992).
Irish, V. F. & Sussex, I. M. A fate map of the Arabidopsis embryonic shoot apical meristem. Development 115, 745–753 (1992).
Stewart, R. N. & Dermen, H. Determination of number and mitotic activity of shoot apical initial cells by analysis of mericlinal chimeras. Am. J. Bot. 57, 816–826 (1970).
Poethig, R. S. & Sussex, I. M. The cellular parameters of leaf development in tobacco: a clonal analysis. Planta 165, 170–184 (1985).
McDaniel, C. N. & Poethig, R. S. Cell-lineage patterns in the shoot apical meristem of the germinating maize embryo. Planta 175, 13–22 (1988).
Bossinger, G. & Smyth, D. R. Initiation patterns of flower and floral organ development in Arabidopsis thaliana. Development 122, 1093–1102 (1996).
Zhai, N. & Xu, L. CRE/LOX-based analysis of cell lineage during root formation and regeneration in Arabidopsis. aBIOTECH 1, 153–156 (2020).
Saulsberry, A., Martin, P. R., O’Brien, T., Sieburth, L. E. & Pickett, F. B. The induced sector Arabidopsis apical embryonic fate map. Development 129, 3403–3410 (2002).
Chan, M. M. et al. Molecular recording of mammalian embryogenesis. Nature 570, 77–82 (2019).
Bowling, S. et al. An engineered CRISPR–Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1410–1422.e27 (2020).
Quinn, J. J. et al. Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts. Science 371, eabc1944 (2021).
Li, L. et al. A mouse model with high clonal barcode diversity for joint lineage, transcriptomic, and epigenomic profiling in single cells. Cell 186, 5183–5199.e22 (2023).
Baron, C. S. & van Oudenaarden, A. Unravelling cellular relationships during development and regeneration using genetic lineage tracing. Nat. Rev. Mol. Cell Biol. 20, 753–765 (2019).
Donà, M. et al. A versatile CRISPR-based system for lineage tracing in living plants. Plant J. 115, 1169–1184 (2023).
McKenna, A. et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016).
Kalhor, R. et al. Developmental barcoding of whole mouse via homing CRISPR. Science 361, eaat9804 (2018).
Salvador-Martínez, I., Grillo, M., Averof, M. & Telford, M. J. Is it possible to reconstruct an accurate cell lineage using CRISPR recorders?. eLife 8, e40292 (2019).
Lu, X. et al. Development of an inducible DNA barcoding system to understand lineage changes in Arabidopsis regeneration. Dev. Cell 60, 305–319.e5 (2025).
Watson, J. M. et al. Germline replications and somatic mutation accumulation are independent of vegetative life span in Arabidopsis. Proc. Natl Acad. Sci. USA 113, 12226–12231 (2016).
Liu, K. et al. Mapping single-cell-resolution cell phylogeny reveals cell population dynamics during organ development. Nat. Methods 18, 1506–1514 (2021).
Deng, S., Gong, H., Zhang, D., Zhang, M. & He, X. A statistical method for quantifying progenitor cells reveals incipient cell fate commitments. Nat. Methods 21, 597–608 (2024).
Lu, Z. et al. Polyclonal-to-monoclonal transition in colorectal precancerous evolution. Nature 636, 233–240 (2024).
Frank, M. H. & Chitwood, D. H. Plant chimeras: the good, the bad, and the ‘Bizzaria’. Dev. Biol. 419, 41–53 (2016).
Grandjean, O. et al. In vivo analysis of cell division, cell growth, and differentiation at the shoot apical meristem in Arabidopsis. Plant Cell 16, 74–87 (2004).
Chandler, J. W., Jacobs, B., Cole, M., Comelli, P. & Werr, W. DORNRÖSCHEN-LIKE expression marks Arabidopsis floral organ founder cells and precedes auxin response maxima. Plant Mol. Biol. 76, 171–185 (2011).
Kwiatkowska, D. Flower primordium formation at the Arabidopsis shoot apex: quantitative analysis of surface geometry and growth. J. Exp. Bot. 57, 571–580 (2006).
Long, J. & Barton, M. K. Initiation of axillary and floral meristems in Arabidopsis. Dev. Biol. 218, 341–353 (2000).
Jenik, P. D. & Irish, V. F. Regulation of cell proliferation patterns by homeotic genes during Arabidopsis floral development. Development 127, 1267–1276 (2000).
Deng, S. & He, X. Tree of life at two levels: from species to cell. Sci. Bull. 68, 2515–2518 (2023).
Xiong, X. et al. Split complementation of base editors to minimize off-target edits. Nat. Plants 9, 1832–1847 (2023).
Fonfara, I., Curth, U., Pingoud, A. & Wende, W. Creating highly specific nucleases by fusion of active restriction endonucleases and catalytically inactive homing endonucleases. Nucleic Acids Res. 40, 847–860 (2012).
Mol, C. D. et al. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: protein mimicry of DNA. Cell 82, 701–708 (1995).
Duan, C. et al. Reduced intrinsic DNA curvature leads to increased mutation rate. Genome Biol. 19, 132 (2018).
Wang, Y. et al. Mesoscale DNA feature in antibody-coding sequence facilitates somatic hypermutation. Cell 186, 2193–2207.e19 (2023).
Niu, Q. et al. Efficient A·T to G·C base conversions in dicots using adenine base editors expressed under the tomato EF1α promoter. Plant Biotechnol. J. 21, 5–7 (2023).
Nguyen, L. T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Spanjaard, B. et al. Simultaneous lineage tracing and cell-type identification using CRISPR–Cas9-induced genetic scars. Nat. Biotechnol. 36, 469–473 (2018).
Liu, Z., Zeng, H., Xiang, H., Deng, S. & He, X. Achieving single-cell-resolution lineage tracing in zebrafish by continuous barcoding mutations during embryogenesis. J. Genet. Genomics 51, 947–956 (2024).
Minh, B. Q., Nguyen, M. A. T. & von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 30, 1188–1195 (2013).
Dolan, L. & Poethig, R. S. The Okra leaf shape mutation in cotton is active in all cell layers of the leaf. Am. J. Bot. 85, 322–327 (1998).
Zeng, M. et al. Stem cell lineage in body layer specialization and vascular patterning of rice root and leaf. Sci. Bull. 61, 847–858 (2016).
Marcotrigiano, M. & Bernatzky, R. Arrangement of cell layers in the shoot apical meristems of periclinal chimeras influences cell fate. Plant J. 7, 193–202 (1995).
Burian, A. Does shoot apical meristem function as the germline in safeguarding against excess of mutations? Front. Plant Sci. 12, 707740 (2021).
Chen, Y., Burian, A. & Johannes, F. Somatic epigenetic drift during shoot branching: a cell lineage-based model. Genetics 227, iyae091 (2024).
Iwasa, Y., Tomimoto, S. & Satake, A. The genetic structure within a single tree is determined by the behavior of the stem cells in the meristem. Genetics 223, iyad020 (2023).
Su, Y. H. et al. Integration of pluripotency pathways regulates stem cell maintenance in the Arabidopsis shoot meristem. Proc. Natl Acad. Sci. USA 117, 22561–22571 (2020).
Cao, X. et al. A self-activation loop maintains meristematic cell fate for branching. Curr. Biol. 30, 1893–1904.e4 (2020).
Shi, B. et al. Two-step regulation of a meristematic cell population acting in shoot branching in Arabidopsis. PLoS Genet. 12, e1006168 (2016).
Landrein, B. et al. Mechanical stress contributes to the expression of the STM homeobox gene in Arabidopsis shoot meristems. eLife 4, e07811 (2015).
Xia, K. et al. The single-cell stereo-seq reveals region-specific cell subtypes and transcriptome profiling in Arabidopsis leaves. Dev. Cell 57, 1299–1310.e4 (2022).
Szymkowiak, E. J. & Irish, E. E. Interactions between jointless and wild-type tomato tissues during development of the pedicel abscission zone and the inflorescence meristem. Plant Cell 11, 159–175 (1999).
Szymkowiak, E. J. & Sussex, I. M. The internal meristem layer (L3) determines floral meristem size and carpel number in tomato periclinal chimeras. Plant Cell 4, 1089–1100 (1992).
Helizon, H. et al. Class I KNOX transcription factors promote differentiation of cambial derivatives into xylem fibers in the Arabidopsis hypocotyl. Development 141, 4311–4319 (2014).
Stanger, B. Z., Tanaka, A. J. & Melton, D. A. Organ size is limited by the number of embryonic progenitor cells in the pancreas but not the liver. Nature 445, 886–891 (2007).
Pineda-Krch, M. & Fagerström, T. On the potential for evolutionary change in meristematic cell lineages through intraorganismal selection. J. Evol. Biol. 12, 681–688 (1999).
Tomimoto, S. & Satake, A. Modelling somatic mutation accumulation and expansion in a long-lived tree with hierarchical modular architecture. J. Theor. Biol. 565, 111465 (2023).
Prusinkiewicz, P., Erasmus, Y., Lane, B., Harder, L. D. & Coen, E. Evolution and development of inflorescence architectures. Science 316, 1452–1456 (2007).
Qin, T. et al. A population of stem cells with strong regenerative potential discovered in deer antlers. Science 379, 840–847 (2023).
Li, J.-F. et al. Comprehensive protein-based artificial microRNA screens for effective gene silencing in plants. Plant Cell 25, 1507–1522 (2013).
Wang, Z. P. et al. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 16, 144 (2015).
Ingham, D. J., Beer, S., Money, S. & Hansen, G. Quantitative real-time PCR assay for determining transgene copy number in transformed plants. Biotechniques 31, 132–140 (2001).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-seq. Genome Biol. 17, 77 (2016).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Zhang, T. Q., Chen, Y. & Wang, J. W. A single-cell analysis of the Arabidopsis vegetative shoot apex. Dev. Cell 56, 1056–1074 (2021).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grant nos 32293191 to X.H., 32530019 to X.H., 32100485 to F.-N.X. and 32200494 to K.L.) and Guangdong Basic and Applied Basic Research Foundation (grant nos 2025A1515010629 to K.L. and 2022A1515010983 to F.-N.X.).
Author information
Authors and Affiliations
Contributions
F.-N.X., K.L., A.L. and Z.L. performed the experiments. F.-N.X., K.L., X.H. and J.W. analysed the data. X.H., J.-F.L. and Z.H. supervised the study. F.-N.X., K.L. and X.H. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Qikun Liu, Ning Zhai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Design of SMALT in Arabidopsis.
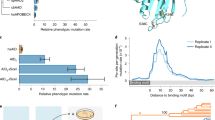
(a) Genotyping of ung msh2 double mutant. (b) Anatomical structure of the shoot apical meristem (SAM) in ung msh2 double mutant. 10 plants were analyzed with one representative image shown. Vibratome sections of the SAM were stained with Calcofluor White to visualize cell walls. Images were acquired using a ZEISS LSM880 confocal microscopy. The boxed region in the left panel is magnified in the middle panel, with cell layers color-coded in the right panel. Bar = 50 um. (c) mRNA expression level of the base editors driven by different promoters. RNA was extracted from young flowers. For each design 3 plants were analyzed. (d) Design of T-DNA containing base editor or lineage barcode. BlpR, phosphinothricin acetyltransferase; HygR, hygromycin B phosphotransferase; LB, left border of T-DNA; RB, right border of T-DNA. (e) Identification of T-DNA insertions of three SMALT lines through Nanopore sequencing. Blue bar indicates T-DNA containing lineage barcode, red bar indicates T-DNA of base editor, grey bar represents genome. Junction reads mapping to insertion sites were displayed below. Arrows indicate primers used for genotyping, with detailed sequence provided in Supplementary Table 8. (f) Sanger sequencing identifying two linked feature sites to decouple Barcode A and Barcode B of Plant-2. (g) An example demonstrating how to decouple barcode sequences of Plant-2 into Barcode A and Barcode B.
Extended Data Fig. 2 Time-course profiling of base editor activity and mutation accumulation.
(a) Expression of the base editor in different developmental stages and organs. The base editor and GFP are linked by 2A peptide and under the driven of SlRPS5a promoter. 12 plants were analyzed with representative images of each organ shown. Images were captured by ZEIZZ LSM880 confocal microscopy. Bar = 100 µm. (b) mRNA expression level of the base editor in different stages and organs. 12 independent SMALT lines were analyzed. House-keeping gene ACTIN serves as an internal control. (c) Time-course mutation analysis. 6 rosette leaves (RL1 to RL6) were collected at the second, forth, and tenth week as indicated. The inflorescence (INF) and siliques (SQ) were also collected as representatives of reproductive organs. White arrowheads indicate collected samples. Bar = 1 cm. 4 individuals (Plant-A, -B, -C, -D) were sampled. For each sample, all sequenced barcodes were counted. Violin plots show mutation number per barcode in each sample. Blue curves indicate LOESS-smoothed fitted values of mutation number in vegetative and reproductive organs, respectively, with shaded regions representing the 99% confidence interval of the fit.
Extended Data Fig. 3 Interpretation of phylogeny under different resolution.
(a) Impact of various mutation rates on reconstructed phylogeny. During the developmental process form a zygote to 64 cells, if mutations accumulated in every cell cycle, a fully-resolved phylogeny can be reconstructed with serial hierarchical bifurcations; if no edits in some cell generations, the reconstructed phylogeny would be partially-resolved with polytomies; variation of mutation accumulation rate per generation affects branch length of the reconstructed phylogeny. (b) Definition and calculation of bifurcation resolving rate (BRR). Upper panel showcases a fully resolved phylogenetic tree with 16 terminal tips and 15 internal nodes. Bottom panel shows a partial resolved tree with 16 terminal tips and 9 internal nodes. The red nodes indicate polytomies caused by lack of resolution. (c) Down sampling of the 1-kb barcode of Plant-1 to reconstruct phylogenies. Down-sampling trees are presented with the full-length tree face-to-face, with the same terminal nodes linked. Cells from each shoot branch were distinguished by color. (d) One-founder and multi-founder models under complete or compromised resolution. Assuming the stem cell pool (only one germ layer is considered here) contains 16 founder cells with known lineage relationships. In the one-founder model (upper-left panel), one cell per germ layer is drawn to initiate a shoot branch. With complete resolution (upper-middle panel), cells of each shoot branch form a single branch-specific clade. If barcoding mutations fail to occur in certain cell generations (red window of the upper-right panel), the phylogeny would comprise polytomies in which branch-specific clades disappear and cells of different shoot branches mix together. In the multi-founder model (bottom-left panel), multiple neighboring founder cells with varying lineage distances initiate a shoot branch (bottom-middle panel). With complete barcoding resolution, multiple branch-specific clades per shoot branch are expected. If barcoding mutations fail to occur in certain cell generations (red window of the bottom-right panel), again, polytomies would be observed and cells of different branches would mix.
Extended Data Fig. 4 Phylogeny analysis of Plant-2.
(a) Schematic drawing showing sample collection of Plant-2. (b) Reconstructed phylogenies of Plant-2. Barcodes from different samples are color-highlighted by shoot branch in separate panels. Barcode A and Barcode B are separately considered. Details of barcoding mutations were presented on the right. The numbers in grey circles show the bootstrap value of each branch-specific clade. (c) Phylogenic topology of the shoot branches in Plant-2. The bootstrap value of each internal node is labeled in grey circles.
Extended Data Fig. 5 The three-cell pattern holds for the secondary and tertiary branches and siliques.
(a) Sample collection and reconstructed phylogeny of the rosette branch 1 (R1) of Plant-1. A total of 888 cell barcodes from the secondary and tertiary branches are combined with the original 1,337 cell barcodes to build a new cell phylogeny. Cell barcodes from R1 and its all secondary and tertiary branches were highlighted in color respectively. Numbers in grey circle show the bootstrap value of each branch-specific clade. (b) Sample collection and reconstructed phylogeny of the cauline branch 2 (C2) of Plant-1. A total of 789 cell barcodes from the secondary and tertiary branches are combined with the original 1,337 cell barcodes to build a new cell phylogeny. Cell barcodes from C2 and its all secondary and tertiary branches were highlighted in color respectively. Numbers in grey circle show the bootstrap value of each branch-specific clade. (c) Cell phylogeny of Plant-1 consisting of inflorescence (INF) barcodes and silique (SQ) barcodes from the indicated six shoot branches. Each panel highlights samples form one shoot branch, with black tips and colored tips indicate INF barcodes and SQ barcodes, respectively. Numbers in grey circle show the bootstrap value of each shoot branch-specific clade.
Extended Data Fig. 6 Single cell library construction and cell atlas.
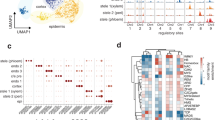
(a) Plant-1 RFP signal examination in planta and protoplasts. (b) Experimental design of four single cell libraries. Each library contains flowers or leaf from four SMALT lines. Each SMALT line contributes two unopen flowers or one rosette leaf to single library. (c) Quality control of four single cell libraries. Number of genes per cell and number of UMIs per cell were displayed in histograms. Shadows indicated the filter windows. (d) Comparison between different data integration methods. Gene-cell matrices of four single cell libraries generated from CellRanger were simply merged into one Seurat object and followed by data normalization and scaling under three different functions. (e) Sankey diagram showing the library/tissue origin of each cell. (f) Global differentiation trajectory based on pseudotime estimation by Monocle 3.
Extended Data Fig. 7 cDNA barcode amplification and recovery.
(a) Schematic diagram demonstrating principle of three amplification strategies. (b) Step-by-step statistics of cDNA barcode recovery. Data were separately counted in TT, TN, or NN amplicons. (c) Number of CCS reads assigned to an original barcode sequence. Each dot represents a UMI-consensus 1-kb barcode sequence. Data were separately counted in TT, TN, or NN amplicons. All quality barcode sequences were counted. (d) Number of UMI-consensus 1-kb barcode sequence assigned to a single cell. Each dot represents a single cell. Data were separately counted in TT, TN, or NN amplicons. All quality cells were counted. (e) Number of UMI-consensus 1-kb barcode sequence recovered per cell. All quality cells were counted. UMI generated from TT, TN, or NN amplicons were integrated by cell. Box plots in (c,d,e) show the median (centre line), the 25th and 75th percentiles (box bounds), and whiskers extending to the most extreme values within 1.5× the interquartile range. Notches indicate the approximate 95% confidence interval of the median. Outliers are not shown. (f) Correlation between barcode expression level and barcode recovery rate. Barcode expression is counted by the 1-kb barcode sequence detected in scRNA-seq. Barcode recovery rate indicates the number of full-length cDNA barcode recovered within a cell. (g) UMAP showing the number and percentage of cells with quality lineage barcode recovered in each library. (h) Number of cells with quality lineage barcode recovered in each plant and library. (i) Mutation spectrum of cDNA barcode. (j) Base editing window of cDNA barcode.
Extended Data Fig. 8 Cell type annotation of three lineages in Plant-2 phylogeny.
Left panel shows the reconstructed phylogenies consisting of both cDNA and DNA barcodes of Plant-2. Black tips indicate cDNA barcodes from single cell libraries. Grey tips represent DNA barcodes from 7 tissue samples in Extended Data Fig. 4. Cell cluster annotation of cDNA barcodes is displayed on the right panel.
Extended Data Fig. 9 Extensive lineage analysis of embryonic and post-embryonic tissues.
(a) A larger phylogeny of Plant-1. 1000 barcode reads were sampled from each shoot branch and subject to phylogeny reconstruction. For samples with less than 1000 barcodes, all available cell barcodes were included. Each panel highlights cells from a single shoot branch. (b and c) Phylogeny analysis of embryonic tissues of Plant-1 (B) and Plant-3 (C). Barcode reads were sampled from shoot branches (post-embryonic tissues), cotyledon, primary root, and lateral root, respectively, for phylogeny reconstruction. The Ln-I, Ln-II, and Ln-III of post-embryonic tissues are highlighted based on mutation profiles. (b) In Plant-1, barcodes from cotyledons and roots are broadly distributed across the entire phylogeny, whereas barcodes from post-embryonic tissues form three major clusters, corresponding to Ln-I, Ln-II, and Ln-III, respectively. Polytomy near the root of the phylogeny indicates insufficient barcoding efficiency during the very early embryonic lineages, preventing interpretation of the relationships among cotyledons, roots, and post-embryonic tissues. (c) In Plant-3, SMALT system was active only at early developmental stages. The reconstructed phylogeny forms three major clades (embryonic clones) with large polytomies, enabling a coarse assignment of embryonic lineage relationships. All barcodes from post-embryonic tissues fall exclusively into Clade-1. Cotyledon barcodes occur both within Clade-1—mixed with post-embryonic barcodes—and in a distinct cluster in Clade-3. Root barcodes occupy the remainder of Clade-3 as well as the entirety of Clade-2. Within Clade-1, the three post-embryonic lineages (Ln-I, Ln-II, Ln-III) can already be distinguished based on mutation signatures learned from single-cell and germline data, suggesting that the three lineages/germ layers of the shoot apical meristem should be established very early during embryogenesis.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–3 and Tables 1–8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xia, FN., Liu, K., Wang, J. et al. Mapping the zygote-to-adult developmental cell phylogeny in Arabidopsis thaliana reveals a three-cell rule of branching. Nat. Plants (2026). https://doi.org/10.1038/s41477-026-02264-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41477-026-02264-1