Abstract
Hepatocellular carcinoma (HCC) is one of the most prevalent and malignant forms of primary liver cancer, with limited therapeutic options and a poor prognosis. Cellular senescence contributes to the progression of chronic liver disease while creating a microenvironment that supports tumor growth. This study aims to identify dual-purpose therapeutic targets for HCC treatment and cellular senescence intervention, potentially leading to more effective therapeutic strategies. Utilizing the AI-driven target discovery platform PandaOmics, we prioritized 27 high-confidence, and 8 novel HCC targets potentially associated with cellular senescence. Experimental validations confirmed that the knockdown of pre-mRNA-processing factor 19 (PRPF19) or mitogen-activated protein kinase 9 (MAPK9) in HCC cells significantly reduced cell proliferation. Additionally, suppression of PRPF19 or MAPK9 in hepatic stellate cells treated with doxorubicin resulted in a significant decrease in cellular senescence. These findings underscore the pivotal roles of PRPF19 and MAPK9 in both HCC cell proliferation and cellular senescence, suggesting them as promising novel dual-purpose therapeutic targets for HCC treatment and mitigation of senescence-associated pathologies.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is the predominant form of primary liver cancer, accounting for approximately 90% of all liver malignancies, and is associated with a relatively low survival rate1,2. HCC typically develops from a combination of chronic inflammation and genetic alterations, including p53 mutations and dysregulation of the Wnt and β-catenin pathways3,4,5,6. Persistent liver inflammation can lead to liver fibrosis and cirrhosis, creating a microenvironment favorable for tumorigenesis7. In addition, excessive alcohol consumption8, diabetes mellitus9, hyperinsulinemia10, obesity11, and chronic hepatitis infections12, contribute to chronic inflammation and tissue damage, further increasing the risk of developing HCC. Moreover, growing evidence has illustrated the positive correlation between aging and HCC development, particularly in older individuals (aged 65 years and older) 13. Analyses have suggested that aging is an independent risk factor for the progression of HCC, with factors such as liver fibrosis progressing more rapidly in older patients14.
Aging is commonly associated with cellular senescence, a state of permanent cell cycle arrest. Senescence is considered a stress response to the types of cellular injury that accumulate with age15, such as telomere shortening, oxidative stresses, and epigenetic changes16. In the liver, cellular senescence contributes to a decline in liver function, elevated susceptibility to chronic liver diseases, impaired regenerative capacity, and a higher risk of HCC17,18. Oncogenic activation also induces cellular senescence. For instance, co-expression of oncogenic rat sarcoma virus (Ras) and p53 leads to irreversible senescence in fibroblasts19,20. Senescent cells are commonly characterized by mitochondrial dysfunction, resistance to apoptosis, increased lysosomal activity, nuclear enlargement, and the production of senescence-associated secretory phenotype (SASP) associated factors, including pro-inflammatory cytokines, chemokines, and various growth factors21,22. The accumulation of senescent cells can create an SASP-enriched microenvironment, which promotes tumorigenesis and the progression of HCC23,24. In HCC, the immune-mediated clearance of senescent hepatic stellate cells helps resolve fibrotic lesions (pre-cancer stage) and can slow disease progression. At advanced stages, the secretome of senescent hepatocytes may drive carcinogenesis by affecting the proliferation of neighboring parenchymal cells or stem cells25. Furthermore, therapy-induced SASP promotes the dedifferentiation of cancer cells, leading to a stem cell-like gene expression profile that promotes malignancy. For example, doxorubicin-induced senescence in HuH-7 liver cancer cells was shown to increase stemness and tumorigenicity via the Wnt-β-catenin pathway26. This evidence suggests that inhibiting senescence in anti-cancer therapies may provide therapeutic benefits, potentially preventing further tumor progression by mitigating inflammatory responses and treatment-induced senescence in hepatocytes.
HCC frequently results in a poor prognosis due to its complex biology and high degree of tumor heterogeneity, which complicates effective treatment. Although numerous studies have investigated the mechanisms driving HCC progression, only a limited number of drugs have received worldwide approval. The tumor’s heterogeneity, characterized by diverse genetic and molecular profiles within and between patients, contributes to treatment resistance and makes it challenging to develop universally effective therapies. As a result, there is a high demand for novel targets to enable more effective treatment of HCC1. The integration of AI algorithms with experimental, multi-omic, and computational methods has emerged as a promising approach for enhancing the efficiency and effectiveness of therapeutic target identification, leading to the development of more effective treatments for a wide range of diseases27,28,29. PandaOmics–developed by Insilico Medicine–is an AI-driven target discovery platform, prioritizes therapeutic targets for different diseases by leveraging a comprehensive data processing pipeline, disease-specific models, indication prioritization and expansion, validation and evidence-based selection, and integration with robotic platforms and advanced language models to identify and validate potential therapeutic targets and biomarkers30. PandaOmics has been successful in identifying promising targets in aging, cancer, amyotrophic lateral sclerosis and idiopathic pulmonary fibrosis31,32,33,34,35. In this study, we utilized PandaOmics to perform target prioritization in HCC and then intersected the prioritized target genes with the cellular senescence-associated genes. Subsequently, in vitro assays validated that the knockdown of pre-mRNA-processing factor 19 (PRPF19) or mitogen-activated protein kinase 9 (MAPK9) suppressed HCC cell proliferation. Additionally, inhibiting PRPF19 or MAPK9 reduced cellular senescence in hepatic stellate cells. Our findings demonstrate the critical role of PRPF19 and MAPK9 in HCC cell proliferation and cellular senescence, suggesting them as promising dual-purpose therapeutic targets for HCC treatment and mitigation of cellular senescence.
Results
AI-aided prioritization of targets implicated in both HCC and cellular senescence
A total of 11 transcriptomic and methylomic datasets consisting of HCC samples (n = 1047) and adjacent normal samples (n = 563) were collected, including 7 microarrays, 2 RNAseq, and 2 methylation datasets (Table 1). After data processing (see Materials and Methods), each dataset was uploaded to PandaOmics for analysis. After a case-control comparison was performed for each dataset, a meta-analysis including all 11 comparisons was carried out by PandaOmics, leading to the prioritization of the most promising therapeutic targets for HCC. The top 100 novel and 100 high-confidence targets ranked by multiple AI-derived scores were selected after filtering by druggability, safety, and novelty.
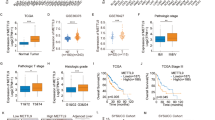
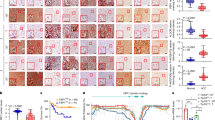
To identify potential dual-purpose HCC and cellular senescence targets, this list of 200 candidates was cross-referenced with genes associated with cellular senescence obtained from multiple databases. This yielded 35 high-confidence and 14 novel targets related to cellular senescence (Fig. 1A). To further narrow down the list to the most promising targets, we assessed the consistency of mRNA expression levels in HCC and control samples across all datasets. Ultimately, 27 high-confidence and 8 novel targets met the defined expression thresholds. High confidence targets (HRAS, CDK1, CDK4, MAPK14, SMARCA4, HDAC1, BRCA1, CDK6, RPS6KA3, PKM, EZH2, CDK2, TERT, TYK2, MAPK3, CHEK1, AURKA, ABL1, RAF1, MAPK9, CHEK2, BRAF, MAPK1, DHX9, SRC, ACLY, ATR) and novel targets (MAPKAPK5, PSMB5, PSMD14, PRPF19, MAPKAPK3, TOP2B, SENP2, DUSP3) were prioritized due to their consistent upregulation in HCC samples across the datasets. Conversely, AKT1, MAP2K1, SIRT1, AR, MAPK8, ATM, HMGB1, MAP2K7, RPS6KA6, DUSP16, NEK6, BLK, CSNK1A1, and KAT6A were excluded from further analysis as they were inconsistently upregulated or downregulated in HCC (Fig. 1B).
A Workflow for AI-aided target prioritization in both HCC and cellular senescence. B Consistency of mRNA expression levels in HCC and controls. Filled colors represent the min-max normalized logFC values in each case-control comparison.
We next conducted a literature review for the shortlisted target genes, examining publications and data on approved drugs, as well as clinical and preclinical studies related to hepatic senescence and HCC. Genes with substantial clinical or preclinical evidence were excluded from our candidate list, as further validation of their therapeutic potential was unnecessary. The remaining candidates were then divided into two batches for in vitro validation of their effects on hepatic cellular senescence and HCC (Table 2). Batch A, consisting of 17 genes with limited evidence supporting their involvement in hepatic senescence, was tested in a hepatic senescence model. Batch B, comprising 6 genes with minimal reported data on both HCC and hepatic senescence, was assessed using both the hepatic cellular senescence model and the HCC cell proliferation assay.
Preliminary validation of genes from Batch A in regulating hepatic stellate cell senescence
To establish a cellular senescence model, we treated hepatic stellate cell-derived LX-2 cells with varying concentrations of doxorubicin (Dox) at 12.5 nM, 25 nM, 50 nM, and 100 nM. The senescence model was confirmed through senescence-associated beta-galactosidase (SA-β-Gal) staining, which revealed a significant increase in both the absolute count and percentage of SA-β-Gal positive cells compared to the DMSO-treated control group (Supp. Figure 1A). While treatment with 12.5 nM Dox did not lead to an increase in SA-β-Gal positive cell count, treatment with 50 and 100 nM Dox dramatically reduced the total cell number and showed significant cellular toxicity. This indicated that 25 nM Dox treatment robustly induced cellular senescence in LX-2 cells (Supp. Figure 1A). To explore potential modulators of cellular senescence, we utilized siRNA to knock down 17 target genes from Batch A including CDK1, CDK6, RAF1, BRAF, DHX9, PRPF19, DUSP3, SENP2, ACLY, BRCA1, MAPK14, TYK2, MAPK1, SRC, ATR, PSMB5, and PSMD14 and investigated their roles in cellular senescence. The efficiency of siRNA knockdown was evaluated (Supp. Figure 1B), and their respective impact on cellular senescence was assessed using SA-β-Gal staining. In the preliminary screening, treatment with siRNA against 7 targets (CDK1, DHX9, DUSP3, ACLY, SRC, PSMB5, and PSMD14) reduced total cell numbers of LX-2 cells, suggesting potential cellular toxicity of target treatment in healthy cells (Supp. Figure 1C). In Dox-induced senescent LX-2 cells, the suppression of CDK1, CDK6, DHX9, ACLY, TYK2, SRC, and PSMD14 resulted in noticeable cellular toxicity, while the knockdown of PRPF19 did not significantly affect the overall cell count compared to the control group (Supp. Figure 1D). In addition, similar to the effects observed with the senolytic agent ABT-263 (also known as navitoclax, a BCL-2 inhibitor)36,37, PRPF19 siRNA treatment led to a marked reduction in both the absolute number and percentage of SA-β-Gal positive cells, indicating that the knockdown of PRPF19 suppressed Dox-induced senescence (Supp. Figure 1E, F). These findings suggest that PRPF19 may serve as a potential cellular senescence target deserving further investigation in the progression of HCC.
PRPF19 emerges as a novel dual-purpose target for HCC and cellular senescence
To further solidify our previous findings, senescent LX-2 cells were transfected with PRPF19 siRNA or non-target siRNA. The knockdown of PRPF19 significantly reduced the number of SA-β-Gal positive cells, demonstrating a comparable effect to the positive control ABT-263 treatment (Fig. 2A). Moreover, PRPF19 knockdown was even more effective in decreasing the percentage of positive SA-β-Gal cells induced by Dox compared to ABT-263 treatment, without impacting the total cell number (Fig. 2B-D). To assess the impact of PRPF19 knockdown on the secretory phenotype of senescent cells, we conducted RNA-Seq analysis (Fig. 2E). The results showed that silencing PRPF19 led to a significant downregulation of SASP-related factors compared to senescent cells transfected with non-target siRNAs. We identified nine SASP factors that were significantly upregulated by DOX and reversed by PRPF19 knockdown (FDR < 0.05) which represent core senescence-associated pathways (Fig. 2F). Notably, inflammatory factors, various cell cycle regulation, and stress response pathways were downregulated, while the G2/M checkpoint pathway was upregulated in Dox-induced senescent cells (Supp. Figure 1H). Furthermore, PRPF19 inhibition upregulated cell cycle regulation pathways (mTORC1 signaling and PI3K-AKT-mTOR), immune response pathways (IL2-STAT5 and IFN signaling responses), cholesterol homeostasis, and stress response pathways (UV response and xenobiotic metabolism) (Supp. Figure 1G).
A–D Evaluation of cell counts, percentage of SA-β-Gal positive cells and total cell number following treatment with non-targeting siRNA, ABT-263, and PRPF19 siRNA. LX-2 cells were exposed to 25 nM doxorubicin (Dox) to induce senescence and further treated with 10 nM non-target siRNA, PRPF19 siRNA, or ABT-263 for quantification purposes. The scale bar is 100 μm. n = 3-9 per group. E, F RNA-seq analysis of senescence-associated secretory phenotype (SASP) related factors in LX-2 cells, senescent LX-2 cells treated with non-target siRNA, and senescent LX-2 cells transfected with siPRPF19. Filled colors from blue to red in E represent relative expression levels (z-score) from low to high. n = 3 per group. Data in F are expressed as mean ± standard deviation (SD). (**p < 0.01, ***p < 0.001, ****p < 0.0001). TPM Transcripts per million.
Previous preclinical studies have shed light on the potential oncogenic roles of PRPF19 in HCC progression38,39. To further support these findings, we investigated the impact of PRPF19 on HCC cell growth using SK-Hep-1 and Huh-1 cell lines. The proliferation indices of these cell lines were tracked over various treatment durations (0, 3, 5, and 7 days) following PRPF-19 siRNA transfection (Supp. Fig. 2A). In SK-HEP-1 and Huh-1 cells, PRPF19 knockdown led to a significant reduction in proliferation compared to the negative control groups (siNC) (Supp. Fig. 2B, C). Additionally, a 10-day colony formation assay post-siRNA transfection confirmed that PRPF19 knockdown effectively inhibited cell colony formation in SK-Hep-1 and Huh-1 cells compared to non-target controls (Supp. Fig. 2D). These results support the notion that PRPF19 inhibition holds promise as a novel dual-purpose therapeutic target in both HCC and cellular senescence warranting further exploration.
Preliminary target validation of genes from Batch B in modulating HCC cell proliferation
To identify potential HCC targets from batch B, comprising HRAS, MAPK3, MAPK9, RPS6KA3, MAPKAPK5, and TOP2B, we conducted a preliminary 7-day CTG assay to evaluate HCC cell viability post-siRNA transfection of the target genes (Supp. Figure 3A). Knockdown of MAPK3, MAPK9, RPS6KA3, TOP2B, and MAPKAPK5 resulted in significant suppression of HCC cell proliferation. However, HRAS knockdown showed relatively limited effects on proliferation inhibition (Supp. Figure 3B). Notably, the infrequency of oncogenic mutations in HRAS in HCC populations40 suggests that HRAS may not be an optimal therapeutic target for HCC. Additionally, concerns regarding the expression of MAPKAPK5 and TOP2B in normal tissues41,42 prompted us to exclude these genes from further analyses. To further validate the anti-tumor effects of the prioritized candidates from batch B (MAPK3, MAPK9, and RPS6KA3), we utilized colony formation assays and CTG assays. The colony formation assays conducted 10 days post-siRNA transfection revealed that knockdown of MAPK3, MAPK9, and RPS6KA3 significantly reduced the number of cell colonies compared to cells transfected with non-targeting siRNAs (Fig. 3A-C). Moreover, the 7-day CTG assay, which monitored cell proliferation at various time points (days 0, 3, 5, and 7), demonstrated substantial growth inhibition in cells transfected with target siRNAs compared to control groups (Fig. 3D-F). To elucidate the mechanisms underlying growth inhibition by MAPK3, MAPK9, and RPS6KA3 knockdowns, we employed Ki67 immunofluorescence staining to validate the effects on cell cycle progression induced by siRNA knockout of target genes. The depletion of MAPK3, MAPK9, and RPS6KA3 led to a significant decrease in Ki67 intensity in HCC cell lines, indicating a considerable inhibition of proliferation (Fig. 3G). These findings support MAPK3, MAPK9, and RPS6KA3 as promising therapeutic targets for combating HCC.
A–C Colony formation assay of HCC cells transfected with non-target or MAPK3, MAPK9, and PRS6KA3 on the 10th day. n = 3 per group. D–F Assessment of the proliferation index of HCC cells transfected with non-target or MAPK3, MAPK9, and PRS6KA3 siRNA on the 0th, 3rd, 5th, and 7th days. The proliferation index was calculated as the fold change in cell viability compared to day 0. n = 3 per group. G Using ki67 staining to evaluate the impact of suppressing MAPK3, MAPK9, and PRS6KA3 in HCC cells on cell proliferation. The scale bar is 200 μm. HCC cells were transfected with 10 nM si-MAPK, si-MAPK9, and si-PRS6KA3. n = 3 per group. Data are expressed as mean ± standard deviation (SD). (*p < 0.05, **p < 0.01, ***p < 0.001).
Dual-purpose target MAPK9 exhibits senomorphic effects
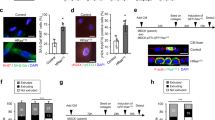
To identify promising dual-purpose therapeutic targets, MAPK3, MAPK9, and RPS6KA3 were assessed for their knockdown efficiency and anti-senescence effects on Dox-induced LX-2 senescent cells (Supp. Figure 3A). Results illustrated that MAPK9 knockdown led to a significant reduction in the absolute cell count of SA-\(\beta\)-Gal positive cells and the percentage of SA-\(\beta\)-Gal positive cells without significantly decreasing the total cell count compared to the negative control (DMSO) group, indicating the absence of significant cytotoxic effects (Fig. 4A-D). Conversely, the knockdown of MAPK3 and RPS6KA3 either had reverse effects or minimal impact on the senescent cells (Fig. 4B-C). These findings suggest that MAPK9 holds promise as a repurposable target for addressing HCC within the realm of cellular senescence.
A Cell count and B Percentage of SA-β-Gal positive cells. C Total cell number. LX-2 cells were stimulated with 10 nM non-targeting control, MAPK3, MAPK9, PRS6KA3 siRNA, and ABT-263. D Images of LX-2 cells after various treatments for SA-β-Gal staining. The scale bar is 200 μm. n = 3–6 per group. Data are expressed as mean ± standard deviation (SD). (*p < 0.05, **p < 0.01, ****p < 0.0001).
Discussion
This study identified and validated dual-purpose therapeutic targets implicated in both HCC and cellular senescence through an AI-driven meta-analysis of transcriptomic and methylomic datasets. By leveraging AI-powered algorithms, the process of target and drug discovery is significantly accelerated, enabling more comprehensive target selection by integrating multi-dimensional data, which yields promising candidates with broader therapeutic potential. In vitro screening using a DOX-induced senescence model in hepatic stellate cells validated the senomorphic effect of the HCC target PRPF19, resulting in a significant decrease in senescent cells. Additionally, MAPK9, along with MAPK3 and RPS6KA3, was identified as an effective inhibitor of HCC proliferation, with MAPK9 alone exhibiting anti-senescent effects. These findings highlight PRPF19 and MAPK9 as promising dual-purpose therapeutic targets for treating HCC and mitigating cellular senescence.
Cellular senescence drives the activation of senescent cancer-associated fibroblasts (CAFs) and endothelial cells (ECs), which release SASP factors that promote immunosuppression, vascular leakage, inflammation, and tumor growth43. Therapy-induced senescence can also lead to the accumulation of senescent tumor cells with drug resistance and heightened invasive and metastatic potential, further accelerating cancer progression44,45,46. This underscores the therapeutic potential of combining senescence inhibition with targeted HCC therapies. In this context, metformin–a drug initially developed to treat type II diabetes–has been shown to inhibit cellular senescence47. Recent studies suggest it may also play a beneficial role in the treatment of HCC, as its use has been associated with reduced cancer metastasis and improved patient survival48,49. Similarly, Navitoclax (ABT263), used as a positive control in this study and initially developed for lymphocytic leukemia, has demonstrated senolytic activity36,37. Navitoclax has also shown promise in treating HCC and other solid tumors50,51,52, further supporting the potential benefits of senescence inhibition in HCC treatment.
Pre-mRNA-processing factor 19 (PRPF19) is an E3 ligase that plays a crucial role in various cellular pathways, including RNA splicing, DNA damage repair, the ubiquitin proteasome pathway, and cell cycle checkpoints53,54,55. Previous research has demonstrated a significant correlation between PRPF19 expression and tumor stage, vascular invasion, and patient prognosis39. Additionally, PRPF19 knockout has been shown to induce cell cycle arrest in HCC cells through the downregulation of CDC5L38. Our findings support the PRPF19’s role in modulating HCC cell proliferation, aligning with previous studies. Furthermore, PRPF19 knockdown reduced both the number and percentage of hepatic senescent cells, as well as the expression of pro-inflammatory SASP factors such as CCL2, CXCL1, IL6, IL1β, and CXCL8. These results suggest a potential senomorphic effect of PRPF19, which may help remodel the chronic inflammatory microenvironment in HCC. We observed a mixed SASP response, where some factors were upregulated while others were downregulated after PRPF19 knockdown. This reflects context-dependent SASP regulation that has been reported by previous studies56,57, and is likely influenced by PRPF19’s roles in splicing and DNA repair. Collectively, our findings highlight PRPF19 as a promising dual-purpose therapeutic for addressing both HCC and cellular senescence, although additional in vivo studies are necessary to further validate the efficacy of inhibiting this target in HCC.
MAPK9 is a serine/threonine protein kinase that regulates cell proliferation, differentiation, migration, transformation, and apoptosis58,59,60,61,62,63. Elevated expression of MAPK9 has been observed in non-small-cell lung carcinoma (NSCLC) tissue, where it promotes NSCLC cell proliferation64. Similarly, MAPK9 enhances the proliferation and migration of glioma cells, with its expression showing a correlation with tumor stage and patient survival65. While the role of MAPK9 in HCC and cellular senescence has not yet been extensively studied, the ERK/MAPK pathway is closely linked to cell proliferation, angiogenesis, and epithelial mesenchymal transition (EMT), all of which contribute to HCC progression66. This suggests that targeting the downstream MAPK9 could offer potential therapeutic benefits for HCC. Moreover, MAPKs such as ERK1/2 and p38 MAPKs are known to regulate senescence-associated gene transcription and mTOR activation67. Our results show that the knockdown of MAPK9 inhibits HCC cell proliferation and reduces the number of hepatic senescent cells. This indicates that MAPK9 could be a promising target for addressing both HCC and cellular senescence. MAPK9 inhibitors like Encorafenib, which are already approved for metastatic melanoma, metastatic non-small cell lung cancer, and metastatic colorectal cancer with BRAF mutations68,69,70, may offer potential for drug repurposing to target HCC and mitigate the pro-tumor inflammatory microenvironment caused by senescent cell accumulation.
Targeting both cellular senescence and HCC holds promise for enhancing the efficacy of cancer therapies. Senescent cells play a role in chronic inflammation and immune suppression by releasing factors associated with SASP71,72, while eliminating these cells has been shown to reduce pro-tumor inflammatory signals and improve immune responses against tumors73. Furthermore, cellular senescence is closely linked to aging. Removing senescent cells from the liver may enhance metabolic and detoxification functions, supporting overall tissue health74,75. Overall, a dual-purpose intervention targeting age-related diseases offers a more comprehensive treatment strategy, addressing root causes, slowing disease progression, and alleviating age-related declines.
Methods
Data selection
Datasets for HCC were screened from publicly available databases including the GeneExpression Omnibus (GEO), ArrayExpress, and The Cancer Genome Atlas (TCGA). Datasets were excluded if i) they consisted fewer than two samples of either cases or controls; ii) the samples were not obtained from the tumors or adjacent sites (for example, serum or plasma); and iii) the samples were obtained from non-human organisms. The raw data for the HCC meta-analysis of this study were then collected with a total of 11 datasets of RNAseq, Microarray, and Methylation data of liver cancer samples compared with adjacent normal samples (GSE36376, GDC-TCGA-LIHC, GSE136319, GSE45267, GSE60502, E-MTAB-5905, GSE6222, GSE107170, GSE102079, GSE89852, GSE54238). A total of 1,047 case samples and 563 control samples were used in the meta-analysis (Table 1).
PandaOmics target prioritization
PandaOmics target identification platform combines a variety of criteria to prioritize the most plausible potential targets for a specific disease, including Omics scores, text-based scores, and KOL scores. Omics scores, which reflect the target-disease association on a molecular level, were calculated from a synthesized approach. Graph-based techniques such as Heterogeneous Graph Walks and Interactome Community scores utilize protein-protein interaction networks and meta-path-guided random walks to rank genes. Matrix Factorization scores employ collaborative filtering to decompose gene-disease associations into latent factors. Expression-based scores normalize omics data to assess differential gene expression. Causal inference and overexpression/knockout scores account for regulatory networks and perturbation-induced transcriptomic data. Mutated submodules and disease submodule scores consider genetic evidence and physical interactions with disease-implicated genes. Mutation burden, pathway analysis, and network neighbors scores were calculated by examining genetic data, pathway activation patterns, and network connectivity. Relevance score integrates clinical trial data to prioritize genes with known drug targets. All scores were normalized to a 0-1 number, indicating the strength of each evidence, with 0 being no evidence and 1 being the highest degree of evidence. PandaOmics also incorporates text-based, financial, and key opinion leader (KOL) scores, which are derived from a comprehensive analysis of scientific publications, grants, patents, and clinical trials. Utilizing advanced natural language processing (NLP) and artificial intelligence (AI) technologies, PandaOmics monitors and analyzes biomedical text data to contextualize mentions of genes, diseases, drugs, and KOLs across a wide array of data sources. For each gene, scores are calculated in a disease-agnostic manner, reflecting the total number of mentions, as well as within specific disease contexts. Attention spikes are calculated exclusively for gene-disease associations, omitting disease-agnostic evaluations. More details can be found in the “PandaOmics User Manual” at https://insilico.com/pandaomics/help.
Manual target selection
Additional filters were manually applied to maximize the therapeutic value of our candidates. Primary filters consisted of druggable protein classes only (enzymes, epigenetic enzymes, ion channels, kinases, nuclear receptors, receptors, secretory proteins, and transporter proteins); a cutoff was set for tissue specificity at n ≥ 0.1 (where 0 indicates that the gene is expressed equally in all tissues and 1 indicates that the gene is exclusively expressed in the selected tissue). In addition, accounting for safety aspects, we excluded genes that have no known small molecule inhibitors or antibodies associated with them. Subsequently, to select novel and high-confidence druggable targets, we applied two distinct filters. For high-confidence targets, we filtered out genes with fewer than 168 related publications and activated key opinion leader (KOL) index scores, financial scores, and text-based scores. For novel targets, we aimed to propose targets based solely on OMICS data calculation. Hence, we excluded genes with more than 168 publications and deactivated KOL, financial, and text-based scores.
Overlap identification for cellular senescence-related HCC targets
PandaOmics prioritized genes that intersected with the cellular senescence-associated gene list. All genes (n = 553) related to the cellular senescence pathway were downloaded from CellAge, GeneOntology, and Reactome (Supp. Table S1). There were 35 high-confidence targets and 14 novel targets that were both prioritized by PandaOmics and associated with cellular senescence.
Expression analysis
The expression performance of each of the overlapped genes was analyzed. We looked for differentially expressed genes (normalized logFC>0.1, combined P value < 0.05) in HCC when compared to normal or healthy samples by linear regression model. We then filtered genes with consistent upregulation in the majority of the datasets for HCC (upregulated in more than 80% of datasets). Min-max normalization was used by PandaOmics to calculate the normalized logFC value.
Cell lines
The normal human hepatic stellate cell line LX-2 was obtained from the American Type Culture Collection (ATCC) and cultured in DMEM medium (Thermo, cat# C11995500BT) supplemented with 10% fetal bovine serum (Gibco, cat# A5669701) and 1% penicillin-streptomycin (P/S, Gibco, cat# 15140122). The cell line was routinely tested for mycoplasma contamination (Lonza, cat# LT07-710) and authenticated with short tandem repeat (STR) assays. Multiple human HCC cell lines were used in this study, including SK-HEP-1, Huh-1, Huh-7, MHCC97H, and Hep-G2 (Table 3). MEM medium was purchased from Procell (cat# PM150411). DMEM medium was purchased from Thermo (cat# C11995500BT). Fetal bovine serum (FBS, cat# A5669701), 1% Non-Essential Amino Acids (cat# 11140050), and 1 mM Sodium Pyruvate (cat# 11360070) were purchased from Gibco.
Reference compounds
Reference compound ABT-263 (cat# HY-10087) was purchased from MedChemExpress (MCE). The compound was dissolved in dimethyl sulfoxide (DMSO, Solarbio, cat# D8371) at a final working concentration of 1 μM.
Doxorubicin-induced senescence model
LX-2 cells were grown in T75 cell culture flasks at a cell confluency of less than 10% and cell viability greater than 90% for senescence model establishment. Cells were seeded into 96-well plates at a volume of 100 μL per well and incubated at 37°C overnight. Doxorubicin (DOX, MCE, cat# HY-15142A) was diluted to 25 nM using a complete cell medium with Dulbecco’s phosphate-buffered saline (DPBS, ThermoFisher, cat# 14190144) as the control. Cell cultures were then treated with Doxorubicin alone, reference compounds or respective siRNAs, and treatment continued for 72 hours. After 72 hours, the SA-β-gal staining was performed.
Senescence-associated beta-galactosidase staining
Senescence-associated beta-galactosidase (SA-β-gal) staining was performed using the SA-β-gal staining kit according to the manufacturer’s protocol (Beyotime, cat# C0602). The cells were fixed with 100 μL fixation buffer and incubated for 15 minutes at room temperature (RT), followed by two washes with 100 μL of PBS for 3 minutes each at room temperature (RT). Subsequently, 100 μL of staining solution was added to each well, and the plates were incubated overnight in a CO2-free incubator at 37°C. The next day, the staining solution was aspirated, and the cells were washed with 100 μL PBS. Then, 100 μL of 4 μg/mL Hoechst solution was added to each well, incubated for 15 minutes at RT, and washed twice with PBS. The plates were then sealed and scanned using the ArrayScan High-Content Screening System (Thermo Fisher Scientific, CX7 LZR) with 25 fields per well and 10x magnification (binning 2×2, 1104×1104) in 5 channels (DAPI: Ex405/Em446/37; SA-β-gal: 590(Amber)-Brightfield, 617(Red)-Brightfield, 447(Blue)-Brightfield, 530(Green)-Brightfield). Each field had a size of 885.54 ×885.54 μm.
siRNA knockdown
Cell suspensions were seeded in 96-well plates at a final cell number of 8000 per well for 3-day treatment or 4000 per well for 7-day treatment. The cells were transfected with siRNAs targeting specific genes and non-targeting control (siNC) using the RNAi duplex-Lipofectamine™ RNAiMAX Kit (ThermoFisher, cat 13778150) for the indicated duration at 37°C and 5% CO2. Three replicates per cell line were performed. All siRNAs were purchased from GENScript and used at a final concentration of 10 nM (Table 4).
Quantitative real-time polymerase chain reaction
Total RNA extraction was performed using the FastPure Cell/Tissue Total RNA Isolation Kit (Vazyme, cat# RC112-01), and complementary DNA (cDNA) was synthesized after reverse transcription of RNA using the HiScript III 1st Strand cDNA Synthesis Kit (Vazyme, cat# R312-01) following the manufacturer’s protocol. Quantitative RT-PCRs were performed in triplicates using the ChamQ SYBR qPCR Master Mix (Vazyme, cat# Q321-03). The primer sequences were listed in Table 5. Samples were run on the LightCycler® 480 real-time PCR Instrument (Roche) with the following program: pre-denaturation (30 s, 95°C), PCR amplification (40 cycles of denaturing at 95°C for 10 s and annealing/extension at 60°C for 30 s). The LightCycler® 480 software was used to compute cycle threshold (CT) values. All values were normalized to those of β-actin.
Colony formation assay
The effect of the knock-down of target genes on the colony formation ability of HCC cells was evaluated by counting the number of colonies. HCC cells were seeded into 24-well plates at a density of 800 cells per well for 24 hours. Subsequently, the medium was replaced with fresh medium containing siNC and siRNAs for each gene. The cells were cultured for 10 days, with the culture media being refreshed every third day. Treated cells were washed with PBS, fixed with 4% paraformaldehyde for 10 minutes, and then stained with 0.05% crystal violet for 30 minutes. Cell images were captured using the Biosystem Azure system. When evaluating functions of different genes, different cell models were applied according to the biological relevance, technical robustness, and phenotypic response in pilot experiments.
Cell viability assay (CTG assay)
For the in vitro cell viability assay, cells were seeded into 96-well plates and transfected with siRNA at indicated time points. After culturing, cell viability was determined using the CellTiter-Glo™ Luminescent Cell Viability Assay Kit (Promega, cat#7573) following the manufacturer’s instructions. The luminescent signal was measured using an Envision multimode microplate reader (PerkinElmer). Cell viability measurements were taken on days 0, 3, 5, and 7, with the values normalized to 1 on the day of seeding. The proliferation index was calculated as the fold change in cell viability compared to day 0.
Cell proliferation assay (Ki67 immunofluorescence staining)
Cells were seeded in 96-well plates and treated with either siRNA or scramble siRNA for specific durations. After treatment, the cells were fixed in 4% paraformaldehyde and incubated at room temperature with an immunofluorescence-blocking buffer (Cell Signaling Technology, cat# 12411S) for one hour. The cell pellets were resuspended in 100 μL PBS (ThermoFisher, cat# 14190144) containing the specified amount of Ki-67 antibody (Abcam, cat# ab16667) and incubated overnight at 4°C in the dark. The next day, cells were washed with PBS and incubated with an Alexa Fluor® 555 secondary antibody (Abcam, cat# ab150078). After an additional PBS wash, the cells were stained with Hoechst 33342 at room temperature for 15 minutes. All samples were then analyzed using an ArrayScan High-Content Screening System (Thermo Fisher Scientific, CX7 LZR) in two channels (Hoechst 33342: Ex346/Em460; Ki67: Ex555/Em565). The analysis was carried out with 20 fields per well at 10× magnification (binning 2×2, 1104×1104 resolution), with each field measuring 885.54 × 885.54 μm.
RNA sequencing sample preparation
For RNA sequencing, LX-2 cells were treated with doxorubicin together with siNC or siPRPF19 for 72 hours. After treatment, RNA was isolated using the Magnetic Tissue/Cell/Blood Total RNA Kit (TIANGEN, cat# DP761) following the manufacturer’s instructions. Total RNA samples were evaluated for integrity using an Agilent 5400 Bioanalyzer. RNA-seq libraries were constructed using the KAPA mRNA HyperPrep Kit (Roche, cat# KR1352) and sequenced on the Illumina NovaSeq NovaSeq6000 platform in the 150 nt, paired-end configuration.
RNA-seq data processing and analysis
Raw RNA-seq count data was normalized by upper quantile normalization. Genes were removed if there were missing values in more than a quarter of the samples. Missing values were imputed using the sample min. Ensembl gene IDs were mapped to HGNC symbols. Normalized count values of genes with the same symbols were combined by geometric mean. Differential gene expression analysis was performed by linear regression integrated in the R package Limma76. Hallmark geneset pathways were obtained from the MSigDB database using the R package msigdbr ( https://igordot.github.io/msigdbr/). Gene Set Enrichment Analysis (GSEA) was performed using the R package fgsea77. The false discovery rate (FDR) was calculated to correct for multiple comparisons. Genes and pathways with FDR < 0.05 were considered significantly differentially expressed and enriched, respectively. SASP factors were selected for analyses according to previous studies78,79,80. The 9 SASP factors were then selected for further analysis based on their robust clinical relevance, critical roles in HCC progression81,82,83,84,85,86,87,88,89, and statistical significance in (1) upregulation in DOX-induced senescence (FDR < 0.05) and (2) downregulation after PRPF19 knockdown (FDR < 0.05).
Statistical analysis
All in vitro experiments were performed in triplicates. All results are expressed as mean ± standard deviation (SD). GraphPad Prism 10.1.0 was used to collect and analyze data. The statistics between two groups were analyzed by non-paired student t-test and for three groups or more, one-way ANOVA tests were performed with Dunnett’s multiple comparison testing applied to adjust the P values. Linear regression was used to calculate the effect size (log2 fold change) of each gene by case-control comparison, followed by calculation of FDR to adjust for multiple comparisons. All statistical findings with adjusted P < 0.05 were considered statistically significant. (*Adj. P < 0.05; **Adj. P < 0.01; ***Adj. P < 0.001; ****Adj. P < 0.0001)
Data availability
The data presented in this article and supplementary material are available from the corresponding authors upon reasonable request.
References
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Prim. 7, 6 (2021).
Greten, T. F. et al. Survival rate in patients with hepatocellular carcinoma: a retrospective analysis of 389 patients. Br. J. Cancer 92, 1862–1868 (2005).
Cooks, T. et al. Mutant p53 prolongs NF-κB activation and promotes chronic inflammation and inflammation-associated colorectal cancer. Cancer Cell 23, 634–646 (2013).
Yan, H. X. et al. DNA damage-induced sustained p53 activation contributes to inflammation-associated hepatocarcinogenesis in rats. Oncogene 32, 4565–4571 (2013).
Lee, J. M. et al. β-Catenin signaling in hepatocellular cancer: Implications in inflammation, fibrosis, and proliferation. Cancer Lett. 343, 90–97 (2014).
Yu, L. X., Ling, Y. & Wang, H. Y. Role of nonresolving inflammation in hepatocellular carcinoma development and progression. NPJ Precis Oncol. 2, 6 (2018).
Refolo, M. G., Messa, C., Guerra, V., Carr, B. I. & D’Alessandro, R. Inflammatory Mechanisms of HCC Development. Cancers (Basel) 12, https://doi.org/10.3390/cancers12030641 (2020).
Huang, D. Q., Mathurin, P., Cortez-Pinto, H. & Loomba, R. Global epidemiology of alcohol-associated cirrhosis and HCC: trends, projections and risk factors. Nat. Rev. Gastroenterol. Hepatol. 20, 37–49 (2023).
Li, X., Wang, X. & Gao, P. Diabetes Mellitus and Risk of Hepatocellular Carcinoma. Biomed. Res Int 2017, 5202684 (2017).
Donadon, V., Balbi, M. & Zanette, G. Hyperinsulinemia and risk for hepatocellular carcinoma in patients with chronic liver diseases and Type 2 diabetes mellitus. Expert Rev. Gastroenterol. Hepatol. 3, 465–467 (2009).
Kew, M. C. Obesity as a cause of hepatocellular carcinoma. Ann. Hepatol. 14, 299–303 (2015).
Bréchot, C., Gozuacik, D., Murakami, Y. & Paterlini-Bréchot, P. Molecular bases for the development of hepatitis B virus (HBV)-related hepatocellular carcinoma (HCC). Semin Cancer Biol. 10, 211–231 (2000).
Asahina, Y. et al. Effect of aging on risk for hepatocellular carcinoma in chronic hepatitis C virus infection. Hepatology 52, 518–527 (2010).
Macias, R. I. R. et al. Impact of aging on primary liver cancer: epidemiology, pathogenesis and therapeutics. Aging (Albany NY) 13, 23416–23434 (2021).
McHugh, D. & Gil, J. Senescence and aging: Causes, consequences, and therapeutic avenues. J. Cell Biol. 217, 65–77 (2017).
Muñoz-Espín, D. & Serrano, M. Cellular senescence: from physiology to pathology. Nat. Rev. Mol. Cell Biol. 15, 482–496 (2014).
Allaire, M. & Gilgenkrantz, H. The aged liver: Beyond cellular senescence. Clin. Res Hepatol. Gastroenterol. 44, 6–11 (2020).
Ogrodnik, M. et al. Cellular senescence drives age-dependent hepatic steatosis. Nat. Commun. 8, 15691 (2017).
Ferbeyre, G. et al. Oncogenic ras and p53 cooperate to induce cellular senescence. Mol. Cell Biol. 22, 3497–3508 (2002).
Adams, P. D. Healing and hurting: molecular mechanisms, functions, and pathologies of cellular senescence. Mol. Cell 36, 2–14 (2009).
Kumari, R. & Jat, P. Mechanisms of Cellular Senescence: Cell Cycle Arrest and Senescence Associated Secretory Phenotype. Front Cell Dev. Biol. 9, 645593 (2021).
Young, A. R. & Narita, M. SASP reflects senescence. EMBO Rep. 10, 228–230 (2009).
Aravinthan, A. D. & Alexander, G. J. M. Senescence in chronic liver disease: Is the future in aging?. J. Hepatol. 65, 825–834 (2016).
Ramakrishna, G. et al. From cirrhosis to hepatocellular carcinoma: new molecular insights on inflammation and cellular senescence. Liver Cancer 2, 367–383 (2013).
Coppé, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu Rev. Pathol. 5, 99–118 (2010).
Karabicici, M., Alptekin, S., Fırtına Karagonlar, Z. & Erdal, E. Doxorubicin-induced senescence promotes stemness and tumorigenicity in EpCAM-/CD133- nonstem cell population in hepatocellular carcinoma cell line, HuH-7. Mol. Oncol. 15, 2185–2202 (2021).
Pun, F. W., Ozerov, I. V. & Zhavoronkov, A. AI-powered therapeutic target discovery. Trends Pharm. Sci. 44, 561–572 (2023).
Pun, F. W. et al. A comprehensive AI-driven analysis of large-scale omic datasets reveals novel dual-purpose targets for the treatment of cancer and aging. Aging Cell 22, e14017 (2023).
Urban, A. et al. Precious1GPT: multimodal transformer-based transfer learning for aging clock development and feature importance analysis for aging and age-related disease target discovery. Aging (Albany NY) 15, 4649–4666 (2023).
Kamya, P. et al. PandaOmics: An AI-Driven Platform for Therapeutic Target and Biomarker Discovery. J. Chem. Inf. Model 64, 3961–3969 (2024).
Pun, F. W. et al. Identification of Therapeutic Targets for Amyotrophic Lateral Sclerosis Using PandaOmics - An AI-Enabled Biological Target Discovery Platform. Front Aging Neurosci. 14, 914017 (2022).
Ren, F. et al. A small-molecule TNIK inhibitor targets fibrosis in preclinical and clinical models. Nat Biotechnol, https://doi.org/10.1038/s41587-024-02143-0 (2024).
Pun, F. W. et al. Hallmarks of aging-based dual-purpose disease and age-associated targets predicted using PandaOmics AI-powered discovery engine. Aging (Albany NY) 14, 2475–2506 (2022).
Olsen, A. et al. Identification of dual-purpose therapeutic targets implicated in aging and glioblastoma multiforme using PandaOmics - an AI-enabled biological target discovery platform. Aging (Albany NY) 15, 2863–2876 (2023).
Mkrtchyan, G. V. et al. High-confidence cancer patient stratification through multiomics investigation of DNA repair disorders. Cell Death Dis. 13, 999 (2022).
Kirkland, J. L. & Tchkonia, T. Senolytic drugs: from discovery to translation. J. Intern Med 288, 518–536 (2020).
Shebl, B. et al. Targeting BCL-XL in fibrolamellar hepatocellular carcinoma. JCI Insight 7, https://doi.org/10.1172/jci.insight.161820 (2022).
Huang, R., Xue, R., Qu, D., Yin, J. & Shen, X. Z. Prp19 Arrests Cell Cycle via Cdc5L in Hepatocellular Carcinoma Cells. Int J Mol Sci 18, https://doi.org/10.3390/ijms18040778 (2017).
Yang, M., Qiu, Y., Yang, Y. & Wang, W. An Integrated Analysis of the Identified PRPF19 as an Onco-immunological Biomarker Encompassing the Tumor Microenvironment, Disease Progression, and Prognoses in Hepatocellular Carcinoma. Front Cell Dev. Biol. 10, 840010 (2022).
Taketomi, A. et al. A rare point mutation in the Ras oncogene in hepatocellular carcinoma. Surg. Today 43, 289–292 (2013).
Ni, H., Wang, X. S., Diener, K. & Yao, Z. MAPKAPK5, a novel mitogen-activated protein kinase (MAPK)-activated protein kinase, is a substrate of the extracellular-regulated kinase (ERK) and p38 kinase. Biochem Biophys. Res Commun. 243, 492–496 (1998).
Austin, C. A. et al. TOP2B: The First Thirty Years. Int J Mol Sci 19, https://doi.org/10.3390/ijms19092765 (2018).
Ye, J. et al. Senescent CAFs Mediate Immunosuppression and Drive Breast Cancer Progression. Cancer Discov. 14, 1302–1323 (2024).
Wang, D. et al. Sunitinib facilitates metastatic breast cancer spreading by inducing endothelial cell senescence. Breast Cancer Res. 22, 103 (2020).
Yasuda, T. et al. Inflammation-driven senescence-associated secretory phenotype in cancer-associated fibroblasts enhances peritoneal dissemination. Cell Rep. 34, 108779 (2021).
Yasuda, T., Baba, H. & Ishimoto, T. Cellular senescence in the tumor microenvironment and context-specific cancer treatment strategies. Febs j. 290, 1290–1302 (2023).
Moiseeva, O. et al. Metformin inhibits the senescence-associated secretory phenotype by interfering with IKK/NF-κB activation. Aging Cell 12, 489–498 (2013).
Vacante, F. et al. Metformin Counteracts HCC Progression and Metastasis Enhancing KLF6/p21 Expression and Downregulating the IGF Axis. Int J. Endocrinol. 2019, 7570146 (2019).
Schulte, L. et al. Treatment with metformin is associated with a prolonged survival in patients with hepatocellular carcinoma. Liver Int 39, 714–726 (2019).
Matsunaga, H. et al. A novel therapeutic combination sequentially targeting aurora B and Bcl-xL in hepatocellular carcinoma. Ann. Surg. Oncol. 22, 3079–3086 (2015).
Gandhi, L. et al. Phase I study of Navitoclax (ABT-263), a novel Bcl-2 family inhibitor, in patients with small-cell lung cancer and other solid tumors. J. Clin. Oncol. 29, 909–916 (2011).
Rudin, C. M. et al. Phase II study of single-agent navitoclax (ABT-263) and biomarker correlates in patients with relapsed small cell lung cancer. Clin. Cancer Res. 18, 3163–3169 (2012).
Hofmann, J. C. et al. The Prp19 complex directly functions in mitotic spindle assembly. PLoS One 8, e74851 (2013).
Yu, J. & Ge, S. PRPF19 functions in DNA damage repair and gemcitabine sensitivity via regulating DDB1 in bladder cancer cells. Cytotechnology 76, 85–96 (2024).
Hildebrandt, A. et al. Interaction profiling of RNA-binding ubiquitin ligases reveals a link between posttranscriptional regulation and the ubiquitin system. Sci. Rep. 7, 16582 (2017).
Acosta, J. C., Banito, A., Wuestefeld, T., Georgilis, A., Zhang, P., Errez, L. & Gil, J. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15(8), 978–990 (2013).
Soto-Gamez, A., Walaszczyk, A. & Demaria, M. Chromatin basis of the senescence-associated secretory phenotype. Trends Cell Biol. 32, 513–526 http://pmc.ncbi.nlm.nih.gov/articles/PMC9106822/ (2022).
Hu, D., Bi, X., Fang, W., Han, A. & Yang, W. GSK3beta is involved in JNK2-mediated beta-catenin inhibition. PLoS One 4, e6640 (2009).
Samak, G., Suzuki, T., Bhargava, A. & Rao, R. K. c-Jun NH2-terminal kinase-2 mediates osmotic stress-induced tight junction disruption in the intestinal epithelium. Am. J. Physiol. Gastrointest. Liver Physiol. 299, G572–584 (2010).
Yu, F. et al. Post-translational modification of RNA m6A demethylase ALKBH5 regulates ROS-induced DNA damage response. Nucleic Acids Res 49, 5779–5797 (2021).
Yoshitane, H. et al. JNK regulates the photic response of the mammalian circadian clock. EMBO Rep. 13, 455–461 (2012).
Mayer, C., Bierhoff, H. & Grummt, I. The nucleolus as a stress sensor: JNK2 inactivates the transcription factor TIF-IA and down-regulates rRNA synthesis. Genes Dev. 19, 933–941 (2005).
De Graeve, F. et al. Role of the ATFa/JNK2 complex in Jun activation. Oncogene 18, 3491–3500 (1999).
Wang, Z. X. et al. CircRNF20 aggravates the progression of non-small-cell lung carcinoma by activating MAPK9. Eur. Rev. Med Pharm. Sci. 24, 9981–9989 (2020).
Yang, X. et al. MAPK9 is Correlated with a Poor Prognosis and Tumor Progression in Glioma. Front Biosci. (Landmark Ed) 28, 63 (2023).
Moon, H. & Ro, S. W. MAPK/ERK Signaling Pathway in Hepatocellular Carcinoma. Cancers (Basel) 13, https://doi.org/10.3390/cancers13123026 (2021).
Sun, Y. et al. Signaling pathway of MAPK/ERK in cell proliferation, differentiation, migration, senescence and apoptosis. J. Receptors Signal Transduct. 35, 600–604 (2015).
Shirley, M. Encorafenib and Binimetinib: First Global Approvals. Drugs 78, 1277–1284 (2018).
Al-Salama, Z. T. Encorafenib: A Review in Metastatic Colorectal Cancer with a BRAF V600E Mutation. Drugs 81, 849–856 (2021).
Fareed, A. et al. Advances in BRAF-targeted therapies for non-small cell lung cancer: the promise of encorafenib and binimetinib. International Journal of Surgery 110 (2024).
Burton, D. G. A. & Stolzing, A. Cellular senescence: Immunosurveillance and future immunotherapy. Ageing Res Rev. 43, 17–25 (2018).
Prieto, L. I. & Baker, D. J. Cellular Senescence and the Immune System in Cancer. Gerontology 65, 505–512 (2019).
Chibaya, L., Snyder, J. & Ruscetti, M. Senescence and the tumor-immune landscape: Implications for cancer immunotherapy. Semin Cancer Biol. 86, 827–845 (2022).
Ogrodnik, M., Salmonowicz, H. & Gladyshev, V. N. Integrating cellular senescence with the concept of damage accumulation in aging: Relevance for clearance of senescent cells. Aging Cell 18, e12841 (2019).
Song, P., An, J. & Zou, M. H. Immune Clearance of Senescent Cells to Combat Ageing and Chronic Diseases. Cells 9, https://doi.org/10.3390/cells9030671 (2020).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Sergushichev, A., Korotkevich, G., Sukhov, V. & Artyomov, M. N. Fast gene set enrichment analysis. BioRxiv, (2019).
Coppé, J.-P., Patil, C. K., Rodier, F., Sun, Y., Muñoz, D. P. & Goldstein, J., et al. Senescence-Associated Secretory Phenotypes Reveal Cell-Nonautonomous Functions of Oncogenic RAS and the p53 Tumor Suppressor. PLoS Biol. 6, e301 (2008).
Krizhanovsky, V., Yon, M., Dickins, R. A., Hearn, S., Simon, J., Miething, C., Yee, H., Zender, L. & Lowe, S. W. Senescence of Activated Stellate Cells Limits Liver Fibrosis. Cell 134, 657–667 (2008).
Okamura, K., Sato, M., Suzuki, T. & Nohara, K. Inorganic arsenic exposure-induced premature senescence and senescence-associated secretory phenotype (SASP) in human hepatic stellate cells. Toxicol. Appl. Pharmacol. 454, 116231 (2022).
Yao, W., Ba, Q., Li, X., Li, H., Zhang, S., Yuan, Y. & Wang, Y. Inhibition of the CCL2 receptor, CCR2, enhances tumor response to immune checkpoint therapy. Commun. Biol. 3, 720 http://www.nature.com/articles/s42003-020-01441-y (2020).
Wang, Y., Zhang, X., Yang, L., Xu, H., Zhang, J., Wang, Q. & Zhang, T. Overexpression of chemokine (C-X-C) ligand 1 (CXCL1) associated with tumor progression and poor prognosis in hepatocellular carcinoma. Med. Oncol. 31, 260 http://pubmed.ncbi.nlm.nih.gov/25298747/ (2014).
Li, H., Wu, K., Tao, K., Chen, L., Zheng, Q., Lu, X. & Wang, G. Co-expression of CXCL8 and HIF-1α is associated with metastasis and poor prognosis in hepatocellular carcinoma. Oncotarget 7, 15738–15749 http://www.oncotarget.com/article/4412/text/ (2016).
Wei, R., Zhu, Y., Zhang, Y., Wang, Z., Wang, J., Zhu, T. & Zhu, W. Comprehensive bioinformatic analysis of MMP1 in hepatocellular carcinoma and establishment of relevant prognostic model. Sci. Rep. 12, 13639 http://www.nature.com/articles/s41598-022-17954-x (2022).
Divella, R., Gadaleta, C. D., Mazzocca, A., Tufaro, A., Paradiso, A., Gadaleta-Caldarola, G. & Sabba, C. Influence of plasminogen activator inhibitor-1 (SERPINE1) 4G/5G polymorphism on circulating SERPINE-1 antigen expression in HCC associated with viral infection. Cancer Genomics Proteom. 9, 193–198 http://pubmed.ncbi.nlm.nih.gov/22798504/ (2012).
Sukowati, C. H. C., El-Khobar, K. E., Ie, S. I., Anfuso, B., Muljono, D. H. & Tiribelli, C. Prognostic value of baseline interleukin 6 levels in liver decompensation and survival in HCC patients undergoing radioembolization. EJNMMI Res. 11, 51 http://pubmed.ncbi.nlm.nih.gov/34080071/ (2021).
Zhuang, Z., Ju, H. Q., Chen, M., Zhang, Z. Y., Wang, L., Sun, X. & Xu, R. H. The novel interplay between CD44 standard isoform and the caspase-1/IL1B pathway to induce hepatocellular carcinoma progression. Cell Death Dis. 11, 999 http://www.nature.com/articles/s41419-020-03158-6 (2020).
Li, Y., Liu, D., Li, F., Li, C., Song, D., Zhang, M. & Wang, C. Loss of tumor suppressor IGFBP4 drives epigenetic reprogramming in hepatic carcinogenesis. Nucleic Acids Res. 46, 8832–8847 http://academic.oup.com/nar/article/46/17/8832/5051112 (2018).
Li, Z., Zhang, Y., Liu, Y., Zhu, J., Xu, Q., Chen, L. & Yu, C. Epigenetic silencing of CDKN1A and CDKN2B by SNHG1 promotes the cell cycle, migration and epithelial-mesenchymal transition progression of HCC. Cell Death Dis. 11, 823 http://www.nature.com/articles/s41419-020-03031-6 (2020).
Acknowledgements
We thank Ms. Elizaveta Ekimova for her technical assistance with figure design. This study received no funding.
Author information
Authors and Affiliations
Contributions
C.R. and F.W.P. designed and administered the project. C.R. and G.H.D.L. performed data collection, curation, and analyses. C.R., P.R.Z., Y.A.M. and Q.Q.T. conducted the experiment, collected, and analyzed the data. C.R. wrote the original draft. G.H.D.L., S.W.Y.L., Y.X., F.W.P. and A.Z. reviewed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.H.D.L., Q.Q.T., P.R.Z., Y.A.M., Y.X., S.W.Y.K., F.W.P. and A.Z. are affiliated with Insilico Medicine, a commercial company that develops and utilizes AI platforms for therapeutic target discovery, drug repurposing, and indication expansion. C.R. declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ren, C., Leung, G.H.D., Tang, Q. et al. AI-aided identification of dual-purpose therapeutic targets PRPF19 and MAPK9 in hepatocellular carcinoma and cellular senescence. npj Aging 11, 101 (2025). https://doi.org/10.1038/s41514-025-00294-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41514-025-00294-1