Abstract
The human gut microbiota is vital for immune function, metabolism, and resistance to pathogens. Soil-transmitted helminths like Trichuris trichiura can disrupt this microbial community, but the extent and functional significance of these disruptions across diverse regions remain unclear. We investigated the impact of T. trichiura infection on gut microbiota composition and function in three endemic regions—Côte d’Ivoire, Laos, and Tanzania—using standardized, high-resolution metagenomic profiling. Our findings reveal consistent depletion of key short-chain fatty acid (SCFA) producers, including Blautia sp. MSJ 9 and Holdemanella biformis, and enrichment of mucin-degrading genera such as Ruminococcus and Bacteroides. These changes coincided with increased microbial utilization of host-derived carbohydrates and destabilization of microbial networks, notably with the emergence of Segatella copri in infected individuals. Although taxa-level responses varied by region, similar trends in SCFA depletion and mucin degradation were observed across sites, pointing to a potentially shared metabolic response to infection. These alterations suggest compromised gut barrier function and immune modulation, potentially promoting parasite persistence. Our results underscore the potential of microbiome-based strategies, such as targeted probiotics or dietary interventions, to support helminth control by restoring microbial balance and improving host resilience.
Similar content being viewed by others
Introduction
The human gastrointestinal tract harbors a complex and diverse microbial community integral to maintaining health and homeostasis1. These microbes engage in essential processes such as nutrient metabolism, immune modulation, and protection against pathogenic invasion. Disruptions to this delicate ecosystem, known as dysbiosis, have been linked to a wide array of health conditions, including metabolic syndromes, inflammatory diseases, and infections2. Parasitic infections, particularly those caused by soil-transmitted helminths (STHs), are significant drivers of dysbiosis, profoundly altering gut microbial structure and function3,4,5,6. Among these helminths, Trichuris trichiura—commonly referred to as the human whipworm—remains a critical public health concern due to particularly low treatment success compared to other helminth species, especially in low- and middle-income countries7,8. Globally, it is estimated that more than 477 million people are infected with T. trichiura, with the heaviest burden in tropical and subtropical regions where poor sanitation facilitates transmission8. The parasite primarily colonizes the large intestine, where it anchors in the colonic epithelium and mucosa, triggering localized inflammation and immune responses that can alter gut microbiota composition9. Research on T. trichiura infection has highlighted its capacity to influence the microbial community in multiple ways, while the gut microbiota, in turn, plays a role in shaping the parasite’s life cycle and potentially, the treatment outcome10. First, the parasite induces a pronounced Th2-mediated immune response characterized by increased mucus production and altered epithelial barrier function, which can create a niche favorable to certain microbial taxa11. Second, the parasite’s nutrient competition and physical presence in the intestinal lumen may disrupt microbial interactions, contributing to compositional and functional shifts in the microbiota12. Third, T. trichiura infection often leads to broader physiological changes, such as altered bile acid profiles and reduced short-chain fatty acid (SCFA) production, further shaping microbial ecology13. Conversely, the gut microbiota also plays a crucial role in T. trichiura colonization, as microbial communities contribute to egg hatching and larval development, establishing a bidirectional relationship between the parasite and the intestinal microbiome14. This bidirectional interplay underscores the complex relationship between the helminth and the gut microbiota. Previous studies have begun to explore the relationship between T. trichiura infection and gut microbiota, reporting both increased and decreased microbial diversity depending on the population studied and the specific metrics used15. For example, a study using a mouse model of T. muris infection suggests that the presence of the parasite can enrich for specific taxa, including Lactobacillus species16, while another study reports declines in beneficial commensals associated with T. trichiura infection in a small human cohort16. Despite these findings, substantial knowledge gaps remain. There is limited understanding of how T. trichiura affects microbial diversity and function across diverse geographic regions with varying environmental, dietary, and host genetic factors. Functional consequences of these microbiota changes are also underexplored. While a few studies have highlighted alterations in metabolic pathways, such as reduced SCFA production and shifts in carbohydrate and amino acid metabolism, these findings are often based on single populations or limited datasets. Furthermore, the extent to which T. trichiura-induced microbial changes are conserved or vary across regions remains unclear, especially given the significant role that local factors play in shaping the gut microbiota.
We hypothesize that T. trichiura infection leads to systematic but context-dependent microbiome alterations, with regional variation reflecting local dietary and environmental factors. This study aims to address these gaps by systematically characterizing the taxonomic, functional, and ecological impacts of T. trichiura infection on the gut microbiota across multiple countries using a unified technical framework. By focusing on T. trichiura—the only large intestine-colonizing helminth species—this work seeks to provide a comprehensive understanding of the interplay between Trichuris infections and microbial ecology. Our findings are critical for advancing our knowledge of host-microbiota-parasite interactions and may inform the development of targeted therapies to mitigate the impact of helminth infections on gut health.
Results
Sampling results and study characteristics
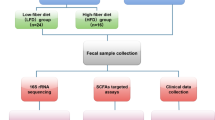
Between September 2018 and December 2019, individuals from the Pujini and Dodo shehias (administrative districts) on Pemba Island, Tanzania; the Dabou and Jacqueville health districts in the Grand Ponts health region, Côte d’Ivoire; and the Nambak district in the Luang Prabang province, Laos, were screened for T. trichiura as described elsewhere17. Of 8,866 screened participants, 160 T. trichiura-negative (60 from Côte d’Ivoire, 50 from Laos, and 50 from Tanzania) and 639 T. trichiura-positive individuals (200, 239, and 200 per site, respectively) were included to represent uninfected and infected groups (Table 1, Fig. 1). 9/799 samples included in this study were dropped due to insufficient sequencing quality. Additional sample metadata is available in Supplementary Data 1. Participants were predominantly young, with mean ages ranging from 14.1 years in Tanzania to 24 years in Laos, and an overall age range of 6–60 years across all regions. Females accounted for roughly half of the infected group, with proportions varying slightly between regions (50.5% in Côte d’Ivoire to 56.6% in Tanzania). T. trichiura infections were mostly of light intensity, with geometric mean eggs per gram (EPG) values ranging from 380 in Laos to 497 in Côte d’Ivoire, while moderate and heavy infections were less common.
Study population screened, number of T. trichiura infected and uninfected participants identified and sequenced samples in each of the three sites.
Community-level taxonomic comparisons
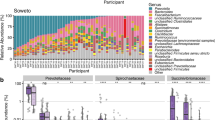
A comparison of alpha diversity metrics between infected and uninfected individuals revealed significant differences across the three countries, as shown in Fig. 2A. Shannon diversity was significantly higher in infected participants from Laos (P < 0.001) but significantly lower in those from Côte d’Ivoire (P < 0.001), with no difference observed in Tanzania (P = 0.11). Analysis of dominance, using the Berger-Parker index, indicated significantly elevated dominance in uninfected individuals in Laos (P < 0.001), suggesting differing community structures between infection states. PERMANOVA analysis (Fig. 2B) revealed that infection status was significantly associated with community composition in all three countries (P < 0.001). PERMDISP analysis revealed significant differences in dispersion around the group centroid in Côte d’Ivoire (P = 0.008) and Laos (P = 0.049), indicating greater heterogeneity in these settings, while no such differences were detected in Tanzania (P = 0.472). Core microbiota comparisons (Fig. 2C) highlighted pronounced differences in microbial composition across countries. Participants in Laos were, on average, older than those from Côte d’Ivoire (P < 0.001) and Tanzania (P < 0.001), whereas age did not differ significantly between participants from Côte d’Ivoire and Tanzania. Therefore, the higher prevalence of Bifidobacterium pseudocatenulatum and B. adolescentis observed in Tanzania is unlikely to be explained by age differences and may instead reflect geographic effects. Clustering of all samples based on species-level abundances (Fig. 2D) revealed distinct country-specific clusters. While infection status did not lead to discernible clustering in Côte d’Ivoire and Tanzania, a significant association was observed in Laos (Chi-square with Yates correction; P < 0.001). Specifically, one sample cluster from Laos contained a disproportionately higher proportion of infected patients (n = 90/92), indicating potential infection-related microbial patterns unique to this setting (Supplementary Fig. 1A).
A Alpha diversity metric stratified by country and infection status. The left panel shows the Shannon diversity index, and the right panel displays the Berger-Parker dominance index. B Non-metric multidimensional scaling (NMDS) plot based on the Bray-Curtis dissimilarity index. Results from community comparisons using PERMANOVA and community dispersion using PERMDISP are shown, with significant results highlighted in bold. C Core microbiome analysis illustrating baseline taxonomic differences across the three study settings. D Heatmap with overall sample clustering based on Robust Aitchison dissimilarity and the Ward method for cladogram construction. The original eggs per gram (EPG) values of stool are displayed at the bottom of each sample.
Species-level taxonomic signal associated with infection status across countries
A prevalence-based comparison using Fisher’s exact test revealed significant differences between infected and uninfected individuals across multiple bacterial taxa (Fig. 3A). A total of 104 species showed variations in prevalence across the three countries studied. Notably, the genus with the highest number of significant differences was Streptococcus, comprising 10 species that were either more or less prevalent depending on infection status. Among these, several species demonstrated consistent differences in prevalence across two or more countries, including S. sanguinis (n = 2), S. parasanguinis (n = 2), S. gallolyticus (n = 2), S. australis (n = 2), and S. gordonii (n = 3). Figure 3B shows species-level differences in relative abundance identified using a stringent multi-method approach combining the Kruskal–Wallis test, LEfSe, and MaAsLin2 (adjusted for country). Species found to be enriched or depleted in this analysis often overlapped with those identified in the prevalence-based model. For example, several taxa from the Actinomyces, Blautia, Clostridium, Roseburia, Ruminococcus, Segatella, and Streptococcus genera showed significant differences in both prevalence and relative abundance models. Species-level taxonomic profiles and differential abundance results are provided in Supplementary Data 2 and 3. Interestingly, many species that were differentially abundant exhibited a country-specific fold-change, which was frequently opposite in direction. Strain-level analysis revealed that the direction of infection-associated abundance changes was closely linked to geographic strain structure. Species exhibiting region-specific strain segregation were significantly more likely to display opposite infection-associated trends across countries (Fisher’s exact P = 0.0044; Supplementary Table 1 and Supplementary Fig. 2), whereas species lacking clear geographic structure tended to show more consistent responses. These findings indicate that strain-level variation partly underlies the observed geographic divergence in microbiome responses to T. trichiura infection. Figure 3C compares the cumulative relative abundance of taxa associated with infection status, distinguishing species enriched or depleted in a single country from those consistently affected in two or more. While species unique to individual countries showed no significant differences in their cumulated abundances, overlapping taxa differed significantly across all three countries (PCI < 0.0001; PLA = 0.0413; PTA = 0.0062), indicating partial cross-regional consistency in microbiome alterations associated with T. trichiura infection.
A Prevalence-based comparison showing phylogenetic relationships of species that were significantly more or less prevalent in at least one country. Statistical significance was determined using Fisher’s exact test with Benjamini-Hochberg false discovery rate (FDR) correction (adjusted P < 0.05). B Comparison of relative abundances between infection status groups, stratified by country. Three statistical tests were applied: Kruskal–Wallis test, LEfSe differential abundance pipeline, and MaAsLin2 (adjusted for the country variable). Only species that were statistically significant across all three tests were retained. The log2 fold-change values, generated using MaAsLin2, are displayed to represent effect sizes. Positive log2 fold-change values indicate higher abundance in uninfected individuals, while negative values indicate higher abundance in infected patients. Taxa highlighted in orange are associated with infection status in two or more settings but show opposite directions of change. Taxa highlighted in green differ in two or more settings and show a concordant direction of change. C Comparison of cumulative relative abundances of species significantly enriched or depleted in one versus two (or more) study sites. Statistical comparisons were performed using the Kruskal-Wallis test, with Bonferroni correction for P-values (adjusted P < 0.05).
Changes in species networks associated with infection status
To assess the impact of T. trichiura infection on microbial community interactions, we performed network analyses with SpiecEasi using taxonomic profiles stratified by infection status and country. Network metrics were calculated from the 200 most abundant species, and visualizations focused on the top 50 species in each group (Fig. 4). Species networks built from infected and uninfected participant samples displayed marked differences in structure across all countries. Key metrics, such as the number of edges (E), network density (De), average path length (PL), diameter (Di), and clustering coefficient (CC), consistently highlighted disrupted microbial interactions associated with infection status. In Côte d’Ivoire, networks from infected participants had an increased number of edges (E = 497 vs. 385), higher density (De = 0.025 vs. 0.019), and greater clustering coefficient (CC = 0.198 vs. 0.147), along with a shorter average path length (PL = 3.5 vs. 4.1). These changes suggest a more fragmented yet locally interconnected microbial network. In Laos, infected participants exhibited a lower number of edges (E = 306 vs. 368), lower density (De = 0.015 vs. 0.018), and shorter average path length (PL = 3.9 vs. 4.6), but an increased clustering coefficient (CC = 0.186 vs. 0.115). This indicates a more modular and less connected network structure. Finally, in Tanzania, the clustering coefficient was higher in infected participants (CC = 0.187 vs. 0.107), while the path length was shorter (PL = 4.5 vs. 5.3) and diameter was reduced (Di = 13 vs. 17). These changes indicate a more compact and tightly clustered microbial network.
SpiecEasi analysis of taxonomic profiles stratified by country and infection status. The visualization was based on the 50 most abundant species in the different groups, while the metrics were calculated using the 200 most abundant species in each group. Network metrics are as follows: E edges, De density, PL average path length, Di diameter, CC clustering coefficient. The numbers in the correlation graphs correspond to the species listed in the table at the bottom of the figure. Inf. infected; Uninf. uninfected.
Networks from infected participants consistently showed reduced inter-species connectivity, with several taxa shifting in their ecological roles. Genera such as Streptococcus, Clostridium, Dorea, and Blautia were central, highly connected nodes in uninfected networks but became peripheral after infection. Conversely, species like Segatella copri emerged as a new hub in infected groups, suggesting increased ecological importance or resilience in the context of infection. Some species—including Faecalibacterium prausnitzii and Eubacterium rectale—became central hubs in infected patients in Côte d’Ivoire and Laos, but not in Tanzania, thus also indicating a widespread, yet locally adapted, ecological response to T. trichiura infection. These changes underscore a reorganization of microbial community dynamics in response to T. trichiura infection. While infection-induced network perturbations were evident across all countries, their magnitude varied, reflecting the influence of local environmental and host factors. Nevertheless, conserved patterns emerged, including reduced connectivity and clustering in infected networks, supporting the hypothesis that T. trichiura infection disrupts microbial community stability.
Functional shifts in microbial communities associated with T. trichiura infection
To investigate functional changes associated with T. trichiura infection, we analyzed abundance profiles based on Enzyme Commission (EC) numbers, which systematically classify enzymes by function, across infection status and countries. A total of 6440 ECs were found, of which 5453 were shared between the three countries, regardless of infection status (Supplementary Fig. 3A). As shown in Supplementary Fig. 3B, functional diversity did not differ significantly between infection groups in any of the three countries. Qualitative differences between infected and uninfected groups were observed in each country, with infected individuals showing 458, 615, and 840 unique ECs in Côte d’Ivoire, Laos, and Tanzania, respectively (Fig. 5A). In contrast, uninfected groups exhibited far fewer unique ECs (122, 38, and 25, respectively), indicating infection-associated diversification of microbial functional potential. NMDS analysis based on Bray–Curtis dissimilarity (Fig. 5B) revealed distinct clustering by infection status and country. PERMANOVA confirmed significant differences between infected and uninfected groups (Côte d’Ivoire: R2 = 0.0076, P = 0.091; Laos: R2 = 0.0217, P = 0.001; Tanzania: R2 = 0.0098, P = 0.014), indicating a stronger infection-related effect on microbial functional potential in Laos. PERMDISP analysis indicated no significant differences in community dispersion in Laos and Tanzania (P > 0.05), whereas Côte d’Ivoire showed marginal significance (F-stat = 3.89, P = 0.035), suggesting greater within-group variability. Volcano plots (Fig. 5C) show ECs significantly enriched or depleted with infection. Overall, the highest number of differentially abundant ECs was found in Laos (n = 1279), followed by Côte d’Ivoire (n = 317) and Tanzania (n = 293). Pathway-level comparisons (Fig. 5D, Supplementary Data 4) revealed infection-associated shifts in core metabolic processes. We identified significant alterations in 67 MetaCyc pathways between Trichuris-infected and uninfected patients using MaAsLin2 while controlling for the “Country” variable (i.e., not stratified by country; threshold: FDR < 0.05). These differences reveal marked country-independent and infection-associated remodeling of microbial glycan metabolism, fermentation capacity, and nitrogen cycling, consistent with epithelial disruption, reduced SCFA availability, and inflammation in the infected gut.
A Venn diagram showing the enzyme commission (EC) numbers found in each study setting, stratified by infection status and showing the overlapping portion. B Non-metric multidimensional scaling (NMDS) plot based on the Bray-Curtis dissimilarity index applied to the EC abundance tables. Results from community comparisons using PERMANOVA and community dispersion using PERMDISP are shown, with significant results highlighted in bold. C Volcano plots based on comparison by infection status of normalized EC abundance tables using MaAsLin2, stratified by country. D Averaged Metacyc superclass abundance comparison by infection status stratified by country. E Fold-change abundance difference of Metacyc pathways between infected and uninfected patients calculated with MaAsLin2, stratified by country. F Fold-change (FC) comparison of Metacyc pathways associated with infection status between countries. To visualize the monotonic trend, we applied LOWESS smoothing (represented by colored triangles with black outlines) using a proportion of 0.6, balancing smoothness with the capture of local variations. IC Côte d’Ivoire, LA Laos, TA Tanzania.
Glycan and mucin-related carbohydrate biosynthesis
Pathways involved in nucleotide-sugar and glycan precursor biosynthesis were among the most depleted in infected individuals. UDP-N-acetyl-D-galactosamine biosynthesis II (log₂FC = −0.493, FDR < 0.001) and UDP-N-acetyl-D-glucosamine biosynthesis II (log₂FC = −0.517, FDR < 0.001), which produce precursors for O-linked glycans and bacterial surface polysaccharides, showed marked reductions, as did dTDP-3-acetamido-α-D-fucose (log₂FC = −0.515, FDR = 0.0047), dTDP-N-acetylviosamine (log₂FC = −0.335, FDR = 0.0182), and dTDP-α-D-ravidosamine biosynthesis (log₂FC = −0.451, FDR = 0.0041). Together, these results indicate broad suppression of bacterial glycan and exopolysaccharide biosynthesis, consistent with impaired mucin metabolism and reduced intestinal barrier support.
Fermentation and SCFA metabolism
Fermentation pathways contributing to SCFA production were consistently reduced in infected individuals. Sucrose degradation II (sucrose synthase; log₂FC = −0.369, FDR < 0.001) supports mixed-acid fermentation yielding acetate, formate, and lactate, whereas TCA cycle VII (acetate producers; log₂FC = −0.259, FDR < 0.012) drives acetate formation from acetyl-CoA under anaerobic conditions. Their concurrent depletion suggests a reduced microbial capacity for carbohydrate fermentation and SCFA generation, which may in turn limit epithelial energy supply and anti-inflammatory signaling.
Nitrogen and amino acid metabolism
Infection was also accompanied by marked alterations in nitrogen cycling. The urea cycle (log₂FC = −0.233, FDR < 0.001) was depleted in infected individuals, suggesting reduced microbial nitrogen recycling and ammonium assimilation. Conversely, several nitrogen- and redox-related pathways were enriched in uninfected individuals, including L-cysteine biosynthesis VI (log₂FC = 0.225, FDR < 0.001), the GABA shunt (log₂FC = 0.244, FDR < 0.016), and the superpathway of allantoin degradation (log₂FC = 0.656, FDR = 0.002). These trends point to enhanced nitrogen turnover and greater metabolic resilience in uninfected microbiomes.
Carbohydrate degradation and general energy metabolism
Pathways involved in carbohydrate breakdown and glycolytic energy production were enriched in uninfected individuals, including starch degradation III (log₂FC = 0.147, FDR = 0.015), the non-oxidative pentose phosphate pathway I (log₂FC = 0.058, FDR = 0.001), and glycolysis IV (log₂FC = 0.098, FDR < 0.001). This enhanced carbohydrate metabolism may increase SCFA precursor availability, thereby supporting epithelial integrity and mucosal immune balance.
Stratified analyses revealed both shared and contrasting microbial responses to T. trichiura infection (Fig. 5E). The superpathway of S-adenosyl-L-methionine biosynthesis, linked to methylation and one-carbon metabolism, was enriched in Côte d’Ivoire (log₂FC = 0.282, FDR = 0.089) but depleted in Laos (log₂FC = −0.689, FDR = 0.001), indicating regional variation in methylation capacity. Energy metabolism showed similar divergence, with TCA cycle V (2-oxoglutarate synthase) depleted in Tanzania (log₂FC = −0.34, FDR = 0.039) but enriched in Laos (log₂FC = 0.247, FDR = 0.071). Carbohydrate utilization also differed: lactose and galactose degradation I was depleted in Tanzania (log₂FC = −0.482, FDR = 0.039) but enriched in Laos (log₂FC = 0.359, FDR = 0.029), while ascorbate degradation II was enriched in Côte d’Ivoire (log₂FC = 0.478, FDR = 0.008) yet strongly depleted in Laos (log₂FC = −1.42, FDR < 0.001). In contrast, several pathways—including purine nucleobase degradation I, glycolysis IV, and the superpathway of N-acetylglucosamine, N-acetylmannosamine, and N-acetylneuraminate degradation—were consistently enriched across countries, reflecting shared mucin- and polyamine-related metabolism.
Across pathways associated with Trichuris infection status in at least one country, the log2FC values were positively correlated between Côte d’Ivoire and Tanzania (Spearman’s ρ = 0.285, P < 0.0001), suggesting a degree of consistency in microbiome functional shifts associated with infection in these two regions (Fig. 5F). However, these values were strongly negatively correlated between Laos and both Côte d’Ivoire (ρ = −0.395, P < 0.0001) and Tanzania (ρ = −0.383, P < 0.0001), indicating opposing patterns of functional pathway enrichment and depletion between the Asian and African cohorts.
Discussion
This study provides a comprehensive cross-regional view of how T. trichiura infection reshapes the human gut microbiome. Across three endemic populations, infection consistently altered microbial metabolism and community structure, combining shared functional disruptions with region-specific adaptations. This study extends previous research on helminth-microbiota interactions15, by demonstrating that T. trichiura induces robust taxonomic and functionally coherent perturbations that transcend geographic boundaries, while still reflecting local environmental influences.
A key observation is the depletion of fermentation and SCFA metabolism, coupled with reprogramming of mucin and glycan processing. Pathways supporting acetate and butyrate production, such as sucrose degradation II and TCA cycle VII (acetate producers), were reduced across regions, alongside losses of SCFA-producing taxa, including Blautia sp. MSJ 9 and Holdemanella biformis. Infection also suppressed bacterial exopolysaccharide and nucleotide-sugar biosynthesis while enriching mucin-degradation pathways. Together, these trends suggest a redirection of microbial metabolism from dietary carbohydrate fermentation toward host-derived substrates, weakening mucosal protection and potentially promoting parasite persistence. The enrichment of mucin-degrading genera such as Ruminococcus and Bacteroides—both linked to mucus thinning and inflammation16,18—supports this interpretation.
Cross-country comparisons revealed that the magnitude and direction of infection-associated changes varied among regions. Functional correlations were positive between Côte d’Ivoire and Tanzania but negative between Laos and both African cohorts, with opposing trends in pathways related to energy metabolism and methylation. Such divergence likely reflects environmental, dietary, and host influences shaping local microbiome responses. Yet, across all settings, infection was characterized by reduced SCFA production and glycan biosynthesis—suggesting a common ecological outcome expressed through regionally adapted community configurations.
Beyond metabolic shifts, T. trichiura infection was associated with marked reorganization of microbial interaction networks. Infected communities showed higher local clustering but lower global connectivity, indicating more modular and less cohesive structures. Central taxa in uninfected networks (Streptococcus, Clostridium, Dorea, Blautia) lost prominence, while Segatella copri emerged as a new hub in infected groups. The gain of centrality by Faecalibacterium prausnitzii and Eubacterium rectale in Côte d’Ivoire and Laos but not in Tanzania highlights both shared and locally adapted ecological responses. These network-level changes suggest reduced cooperation and stability in infected communities, features often linked to lower ecosystem resilience.
The colonic niche of T. trichiura provides a plausible mechanistic context for these findings. Unlike Ascaris lumbricoides and hookworms, which inhabit the small intestine, T. trichiura resides in the cecum and proximal colon—microbial environments rich in fermentation and mucin turnover19,20. Helminth-induced immune modulation is strongest at the site of infection21, meaning that stool-based metagenomics captures ecologically relevant infection signatures22. Collectively, our results suggest T. trichiura as a potent ecological modulator of the large-intestinal microbiota, redirecting microbial energy flow, eroding mucosal protection, and reorganizing community networks in ways that may facilitate its persistence while reducing host resilience.
Nonetheless, this study has several limitations. Its cross-sectional design precludes causal inference between T. trichiura infection and microbiome alterations. Longitudinal studies tracking microbiome dynamics during infection, treatment, and reinfection will be essential to establish temporal relationships and identify mechanisms driving dysbiosis. Although shotgun metagenomics offers broad insight into microbial functional potential, it does not capture transcriptional or metabolic activity; integrating metatranscriptomic and metabolomic approaches would clarify active processes underlying helminth-induced perturbations. The sequencing depth achieved here aligns with standard human gut metagenomics, but deeper coverage could improve detection of rare taxa and low-abundance pathways. Finally, age distribution varied across sites, and age-related microbiome maturation—particularly higher Bifidobacterium abundance in younger participants—may have contributed to some regional differences.
In conclusion, this study provides robust evidence that T. trichiura infection induces both geographically consistent and region-specific disruptions in the gut microbiome, affecting SCFA metabolism, mucin degradation, and microbial network organization. These findings demonstrate that helminth-associated dysbiosis follows a reproducible functional trajectory, modulated by local ecological and dietary factors. A deeper understanding of these host–microbe–parasite interactions will be key to developing microbiome-based strategies to improve gut health and support helminth control in endemic regions.
Methods
Sample collection and microscopy
Stool samples were collected as part of a multi-country randomized controlled trial evaluating the efficacy and safety of albendazole-ivermectin combination therapy against T. trichiura and concomitant helminth infections17. The trial was registered on 17 May 2018 on ClinicalTrials.gov (NCT03527732). Helminth infections were diagnosed by duplicate Kato–Katz thick smears following standard protocols23. For molecular analysis, a small aliquot (< 1 g) of stool was transferred into sterile 2 mL screwcap cryotubes using UV-sterilized plastic spatula and frozen at -20°C. At the end of each trial stage, frozen samples were shipped to Swiss TPH (Allschwil, Switzerland) on dry ice and stored at −80 °C until further analysis.
DNA isolation
DNA was extracted from stool samples using the QIAGEN DNEasy PowerSoil Pro (Qiagen, Hilden, Germany) following the manufacturer’s protocol. DNA concentration was quantified using 2 µL of extracted DNA with the Qubit 4.0 fluorometer and high-sensitivity DNA quantification kits (Thermo Fisher Scientific, Waltham, MA, USA).
Shotgun sequencing
Shallow shotgun sequencing was performed as described previously24, generating 2.8E + 06 to 5.3E + 07 reads per sample. Libraries were prepared using the NEBNext Ultra II FS DNA Library Prep Kit (New England Biolabs, Ipswich, MA, USA) according to the manufacturer’s instructions. Final libraries were quantified and pooled equimolarly before loading onto an Illumina NovaSeq 6000 platform. Sequencing was conducted on S4 flowcell (300 cycles) in paired-end mode (2 × 150 bp). Sample-specific sequencing depth is detailed in Supplementary Data 1.
Taxonomic and functional profiling
Taxonomic profiling of shallow shotgun sequencing data was conducted with MetaPhlAn v4.1.1 (11 Mar 2024)25. The CHOCOPhlAn species-marker database vOct22_CHOCOPhlAnSGB_202403 (October 2022) was used, and mapping was performed with Bowtie2 v2.5.426. The resulting taxonomic relative abundance tables were merged using the built-in “merge_metaphlan_tables.py” and were filtered for low counts (minimum count = 100; prevalence = 20% of samples) and low variance (50% features removed based on interquartile range) using MicrobiomeAnalyst v2.027. Strain-level variation was assessed using StrainPhlAn (MetaPhlAn 4 suite), which reconstructs strain-specific marker gene phylogenies to compare subspecies profiles across infection groups. Strain clusters were inferred from patristic distance matrices retrieved from the trees generated in StrainPhlAn, and species-level geographic structure was quantified as the weighted average of within-cluster country or region (Africa or Asia) homogeneity (H). Species were categorized as country-specific (H_region > 0.8), region-specific (H_region > 0.8 and H_country < 0.8), or non-geographically structured (both H < 0.8), and their association with infection-direction classes was tested using Fisher’s exact test. Functional profiling was performed using HUMAnN4 v4.0.0.alpha.1 with default settings and the UniRef 2019_06 protein database. The unstratified functional abundance tables, including the “reactions” (= abundance tables based on the Enzyme Commission (EC) numbers) tables as well as the “pathabundance” (= abundance tables summarizing the abundance of MetaCyc pathways28 based on EC tables) files, were normalized to copies per million (CPM) using the humann_renorm_table.py script before downstream analyses.
Generation of microbiome-specific metrics
All analyses were conducted in R-studio 2024 using R version 4.4+29. Alpha and beta diversity metrics were calculated to assess microbial diversity within and between samples. Alpha diversity, reflecting the richness and evenness of species within individual samples, was evaluated using the vegan R package v2.6-630. Metrics included the Shannon diversity index for richness and evenness, and the Berger-Parker index to measure community domination. Beta diversity, representing dissimilarity between samples, was assessed using Bray-Curtis dissimilarity, calculated through the “vegdist” function in the same package. Non-metric multidimensional scaling (NMDS) was performed to visualize compositional differences in a reduced-dimensional space, with ordination coordinates calculated using the “metaMDS” function configured with k = 2 and try = 10,000 iterations. To statistically evaluate differences in community-level composition, PERMANOVA (Permutational Multivariate Analysis of Variance) and PERMDISP (Permutation-Based Multivariate Dispersion) analyses were performed using the “adonis2”, “betadisper”, and “permutest” functions. The core microbiome, comprising taxa consistently present across a defined proportion of samples, was identified using the “core” function from the microbiome R package v1.23.131. Taxonomic profiles were visualized with a heatmap generated through an in-house script combining multiple R packages. The ComplexHeatmap package v2.18.032 was used for heatmap generation, with hierarchical clustering performed using the “hclust” function and Ward’s algorithm, and optimized for clarity using the “dendsort” function. The species network correlation analysis was conducted using functions from the SpiecEasi package v1.1.133, the RCy3 package v3.2034, and the igraph package v2.0.335. SpiecEasi (method = “mb”) was used with 20 regularization parameters (nlambda = 20, lambda.min.ratio = 1e − 2). Model selection was performed via the StARS stability selection procedure implemented in Pulsar (rep.num = 100), ensuring that all reported networks and associated metrics reflect stability-adjusted, reproducible structures. Network statistics were calculated using the following functions from the igraph package: “graph.density” for network density, “average.path.length” for the average path length, “diameter” for the network diameter, and “transitivity” for the global clustering coefficient. Centrality measures, including degree, betweenness, closeness, and eigenvector centrality, were calculated using the “degree”, “betweenness”, “closeness”, and “eigen_centrality” functions, respectively.
Statistical analysis
Group comparisons were conducted using Mann–Whitney U tests for two-group comparisons and Kruskal–Wallis tests for comparisons involving three or more groups, and all P values were adjusted using the Bonferroni family-wise correction36. Species prevalence comparisons were performed in R using the “fisher.test” function, and the resulting P-values were adjusted using the Benjamini-Hochberg (BH) procedure37. To compare bacterial species abundances, several statistical methods and tools were employed. Adjusted Kruskal–Wallis tests and the LEfSe package38, available on the MicrobiomeAnalyst platform, were used on country-specific subsets to assess enrichment or depletion of taxa associated with infection status. Additionally, MaAsLin2 v1.18.039 was applied to the complete dataset, controlling for the “Country” variable. All P values resulting from these analyses were adjusted using the BH procedure. To minimize false-positive signals, only species with significant corrected P values across all three statistical methods were considered enriched or depleted. Effect sizes of these differences were visualized using log2 fold-change (log2FC) values generated by MaAsLin2. For comparisons of functional profiles, including ECs and MetaCyc pathway abundances, MaAsLin2 was used to assess statistical differences.
Visualizations
Most visualizations were created and optimized using OriginPro 2024 v10.1.0.178 (OriginLab Corporation, Northampton, MA, USA). Heatmaps were generated directly in R using the ComplexHeatmap package, while Venn diagrams were created using the ggplot2 package v3.5.1.
Ethical approvals
The study was conducted in accordance with the Declaration of Helsinki and the International Council for Harmonization Good Clinical Practice (ICH-GCP) guidelines in effect at the time. Ethical approval was obtained from committees in Côte d’Ivoire (088–18/MSHP/CNESVS-km; ECCI00918), Laos (093/NECHR), Zanzibar, Tanzania (ZAMREC/0003/Feb/2018), and from the institutional research commission of the Swiss TPH and the Ethics Committee of Northwestern and Central Switzerland (EKNZ; O_2023-00066; BASEC Req-2018-00494). Written informed consent for participation and data publication was obtained from adult participants or from parents/guardians of minors, with assent from minors. Participants could withdraw at any time without consequences.
Data availability
Shotgun sequence data supporting the findings of this study have been deposited in the NCBI Short Read Archive with the primary accession code PRJNA1242044.
Code availability
All custom analysis scripts used in this study are publicly available at: https://github.com/STPHxBioinformatics/trichuris_microbiome_patterns.
References
Zmora, N., Suez, J. & Elinav, E. You are what you eat: diet, health and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 16, 35–56 (2019).
Levy, M., Kolodziejczyk, A. A., Thaiss, C. A. & Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 17, 219–232 (2017).
Schneeberger, P. H. et al. Investigations on the interplays between Schistosoma mansoni, praziquantel and the gut microbiome. Parasites vectors 11, 1–12 (2018).
Leung, J. M., Graham, A. L. & Knowles, S. C. Parasite-microbiota interactions with the vertebrate gut: synthesis through an ecological lens. Front. Microbiol. 9, 843 (2018).
Hodžić, A., Dheilly, N. M., Cabezas-Cruz, A. & Berry, D. The helminth holobiont: a multidimensional host–parasite–microbiota interaction. Trends Parasitol. 39, 91–100 (2023).
Loke, P. & Lim, Y. Helminths and the microbiota: parts of the hygiene hypothesis. Parasite Immunol. 37, 314–323 (2015).
Else, K. J. et al. Whipworm and roundworm infections. Nat. Rev. Dis. Prim. 6, 44 (2020).
Jourdan, P. M., Lamberton, P. H., Fenwick, A. & Addiss, D. G. Soil-transmitted helminth infections. Lancet 391, 252–265 (2018).
Lawson, M. A., Roberts, I. S. & Grencis, R. K. The interplay between Trichuris and the microbiota. Parasitology 148, 1806–1813 (2021).
Schneeberger, P. H. H. et al. Different gut microbial communities correlate with efficacy of albendazole-ivermectin against soil-transmitted helminthiases. Nat. Commun. 13, 1063 (2022).
Reynolds, L. A., Finlay, B. B. & Maizels, R. M. Cohabitation in the intestine: interactions among helminth parasites, bacterial microbiota, and host immunity. J. Immunol. 195, 4059–4066 (2015).
Schachter, J. et al. Whipworm infection promotes bacterial invasion, intestinal microbiota imbalance, and cellular immunomodulation. Infect. Immun. 88, e00642-19 (2020).
Chen, H. et al. Dissection of the gut microbiota in mothers and children with chronic Trichuris trichiura infection in Pemba Island, Tanzania. Parasites Vectors 14, 1–13 (2021).
Sargsian S. et al. Clostridia isolated from helminth-colonized humans promote the life cycle of Trichuris species. Cell Rep. 41, 111725 (2022).
Tee, M. Z. et al. Gut microbiome of helminth-infected indigenous Malaysians is context dependent. Microbiome 10, 214 (2022).
Holm, J. B. et al. Chronic Trichuris muris infection decreases diversity of the intestinal microbiota and concomitantly increases the abundance of lactobacilli. PloS One 10, e0125495 (2015).
Hürlimann, E. et al. Efficacy and safety of co-administered ivermectin and albendazole in school-aged children and adults infected with Trichuris trichiura in Côte d’Ivoire, Laos, and Pemba Island, Tanzania: a double-blind, parallel-group, phase 3, randomised controlled trial. Lancet Infect. Dis. 22, 123–135 (2022).
Ramanan, D. et al. Helminth infection promotes colonization resistance via type 2 immunity. Science 352, 608–612 (2016).
Bethony, J. et al. Soil-transmitted helminth infections: ascariasis, trichuriasis, and hookworm. Lancet 367, 1521–1532 (2006).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Bäckhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Maizels, R. M., Smits, H. H. & McSorley, H. J. Modulation of host immunity by helminths: the expanding repertoire of parasite effector molecules. Immunity 49, 801–818 (2018).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Bärenbold, O. et al. Estimating sensitivity of the Kato-Katz technique for the diagnosis of Schistosoma mansoni and hookworm in relation to infection intensity. PLoS Negl. Trop. Dis. 11, e0005953 (2017).
Hillmann B. et al. Evaluating the information content of shallow shotgun metagenomics. mSystems 3, e00069-18 (2018).
Blanco-Míguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. 41, 1633–1644 (2023).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Lu, Y. et al. MicrobiomeAnalyst 2.0: comprehensive statistical, functional and integrative analysis of microbiome data. Nucleic Acids Res. 51, W310–W318 (2023).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes-a 2019 update. Nucleic acids Res. 48, D445–D453 (2020).
R Core Team: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2024).
Oksanen, J. et al. Package ‘vegan’: Community Ecology Package. R package version 2.0-7 2, 1–295 (2013).
Lahti, L., Shetty, S., Blake, T. & Salojarvi, J. Tools for microbiome analysis in R. Version 1, 28 (2017).
Gu, Z. Complex heatmap visualization. Imeta 1, e43 (2022).
Kurtz, Z. D. et al. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput. Biol. 11, e1004226 (2015).
Gustafson, J. A. et al. RCy3: Network biology using Cytoscape from within R. F1000Res. 8, 1773 (2019).
Csárdi, G. et al. igraph: Network Analysis and Visualization in R. R package version 1.4.0 https://doi.org/10.5281/zenodo.7682609 (2023).
Rupert G. Jr. Simultaneous statistical inference. (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, 1–18 (2011).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
World Health Organization. Assessing schistosomiasis and soil-transmitted helminthiases control programmes: monitoring and evaluation framework (World Health Organization, 2024).
Acknowledgements
We are deeply grateful to the dedicated trial team members from the Centre Suisse de Recherches Scientifiques and Université Félix Houphouët-Boigny in Abidjan, Côte d’Ivoire; the Lao Tropical and Public Health Institute in Vientiane, Laos; and the Public Health Laboratory–Ivo de Carneri in Chake Chake, Pemba Island, for their hard work and expertise in both field and laboratory activities. We also extend our heartfelt thanks to the local village and medical authorities in Chake Chake District, Pemba Island; Nambak District, Luang Prabang Province, Laos; and the health districts of Dabou and Jacqueville in Côte d’Ivoire, whose collaboration and support were invaluable in making this work possible. We thank Christian Beisel from the Genomics Facility Basel for their support during the shotgun sequencing experiments. We are grateful for the access and support to/from the sciCORE (https://scicore.unibas.ch) scientific computing center at the University of Basel. We are grateful to the European Research Council (no. 101019223) for financial support.
Author information
Authors and Affiliations
Contributions
P.H.H.S.: study design, research design, project supervision, experimental work, statistical analyses, figure generation, writing of the initial manuscript, manuscript editing; E.H., A.S., S.S., and J.T.C.: study design, conducted field work (sample collection, handling, coordination of treatment, parasitological work, and data curation); J.D. and N.R.: experimental work (DNA isolation), manuscript editing; J.K.: study design, research design, project supervision, writing of the initial manuscript, manuscript editing. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Declaration on the use of AI
In the preparation of this manuscript, AI-based tools (ChatGPT and Copilot) were used to assist in language editing, grammatical correction, and improving the clarity of the text. These tools were employed solely for editorial purposes and did not contribute to the conceptualization, data analysis, or scientific conclusions of this work. All scientific content, interpretations, and conclusions presented in this manuscript are the sole responsibility of the authors, who have thoroughly reviewed and approved the final version of the text.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Schneeberger, P.H.H., Dommann, J., Rahman, N. et al. Profound taxonomic and functional gut microbiota alterations associated with trichuriasis: cross-country and country-specific patterns. npj Biofilms Microbiomes 12, 45 (2026). https://doi.org/10.1038/s41522-026-00911-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-026-00911-1